Nature Immunology: doi: /ni Supplementary Figure 1. Zranb1 gene targeting.
|
|
|
- Julius McBride
- 6 years ago
- Views:
Transcription
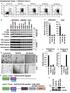
1 Supplementary Figure 1 Zranb1 gene targeting. (a) Schematic picture of Zranb1 gene targeting using an FRT-LoxP vector, showing the first 6 exons of Zranb1 gene (exons 7-9 are not shown). Targeted mice were crossed with FRT deleter (Rosa26-FLPe) mice to generate Zranb1-floxed mice, which were further crossed with different Cre mice to generate germline KO (CMV-Cre), T-cKO (Cd4-Cre), or DC-cKO (Cd11c-Cre) mice. (b) Genotyping PCR analysis of germline Zranb1 wild-type (WT), KO (KO), and heterozygous (Het) mice using P1/P2 primer pair for wild-type allele and P1/P4 primer pair for KO allele. (c,d) Genotyping PCR of T cell-conditional Zranb1 wild-type (WT, Zranb1+/+Cd4-Cre), T-cKO (Zranb1f/fCd4-Cre), and heterozygous (T-cKO Het, Zranb1+/fCd4-Cre) mice (c) or DC-conditional Zranb1 wild-type (WT, Zranb1+/+Cd11c-Cre), DC-cKO (Zranb1f/fCd11c-Cre), and heterozygous (DC-cKO Het, Zranb1+/fCd11c-Cre) mice (d) using P3/P4 primer pair to amplify WT and floxed alleles and Cre-specific primers for the Cd4-Cre and Cd11c-Cre DNA. (e) RT-PCR was performed to measure the Zranb1 mrna level in the indicated cell types.
2 Supplementary Figure 2 Development of cells of the immune system in mice with germline deletion of Zranb1. (a) Flow cytometry analysis of the percentage (numbers in quadrants) of CD4 + CD8 + double-positive (DP), CD4 + and CD8 + single-positive (SP) thymocytes in the thymus (Thy) and B220 + B cells (B) and CD4 + and CD8 + T cells in the spleen (Spl) of wild-type (WT) and KO mice. (b) Flow cytometry analysis of naïve (N, CD62 hi CD44 lo ) and memory (M, CD62 lo CD44 hii ) CD4 + T cells and naïve (N), central memory (CM, CD62L hi CD44 hi ), and effector memory (EM, CD62 lo CD44 hi ) CD8 + T cells in the spleen of wild-type and KO mice. (c) Flow cytometry analysis of the Foxp3 + Treg cells in the spleen and thymus of wild-type (WT) or KO mice. (d) Flow cytometry analysis of conventional DC (cdc, CD11c + CD11b hi B220 ) and plasmacytoid DC (pdc, CD11c + CD11b B220 + ) percentage in total CD11c-positive cells of Bone marrow (BM) and Spleen (Spl). Data in all panels are presented as a representative FACS plots (left) and mean ± SEM values based on multiple samples (right). Similar results were obtained in three independent experiments. *P<0.05.
3 Supplementary Figure 3 Trabid is dispensable in T cells for T cell activation and EAE induction. Age- and sex-matched wild-type (WT) and T-cKO mice were subjected to MOG induced EAE. (a) Flow cytometry analysis of the CD45 + immune cells (CD4 + and CD8 + T cells and CD11b + monocytes) infiltrated into the CNS (brain and spinal cord) (n = 3, day 14 post-immunization), presented as a representative plot (left) and summary graph (right). (b) Flow cytometry analysis of the IFN- producing T H 1 and IL-17A-producing T H 17 cells in the CNS (percent of CD4 + CD45 + cells) and draining lymph nodes (percent of CD4 + T cells) (n = 3, day 14 post-immunization). Data are presented as a representative plot (left, CNS only) and summary graph of absolute cell numbers (right). (c) ELISA analysis of the indicated cytokines in the supernatants of cultures of splenocytes isolated from WT and T-cKO EAE mice (day 14 after immunization) and restimulated in vitro with MOG peptide (20 g/ml for 48 h). (d) Flow cytometry analysis of the frequency of T H 1, T H 17, and T REG cells in wild-type and T-cKO naive CD4 + T cells, stimulated for 4 days with platebound anti-cd3 and anti-cd28 antibodies under T H 0, T H 1, T H 17, or T REG conditions. (e) Flow cytometry analysis of cell proliferation, based on CFSE dilution, of naïve T cells stimulated with plate-bound anti-cd3 (5 g/ml) and anti-cd28 (1 g/ml) for 48 h. (f) ELISA of IFN- and IL-2 using supernatants of the cell cultures described in e. Data are representative of three independent experiments. Error bars are mean ± SEM values.
4 Supplementary Figure 4 Effect of Trabid deficiency on the function of DCs in regulating T cell differentiation. (a,b) Flow cytometry analysis of Foxp3 + T REG cells (a) or IFN- + T H 1 and IL-17 + T H 17 cells (b) in wild-type naïve CD4 + T cells stimulated for 4 days with plate-bound anti-cd3 and anti-cd28 plus supernatant of LPS-activated wild-type (WT) or DC-cKO BMDCs (LPS-DC) in the absence ( ) or presence (+) of the indicated cytokines.
5 Supplementary Figure 5 Trabid regulates LPS-stimulated c-rel recruitment and histone modifications at the promoters of Il12a and Il12b. (a) ChIP assays of c-rel recruitment to the promoter of the indicated genes using wild-type (WT) and DC-cKO BMDCs that were either unstimulated (0 h) or stimulated for 6 h with LPS. Data are presented as percentage of the total input DNA, as determined by QPCR assays. (b) Luciferase assays using HEK293 cells transfected with an Il12b-luciferase reporter plasmid in the presence (+) or absence ( ) of an empty vector (Vector) or expression vectors encoding c-rel, Trabid, or a catalytically inactive Trabid mutant, C443 (Mut Trabid). Luciferase activity is presented as fold based on empty vector. (c) ChIP analyses of H3K9 dimethylation (H3K9me2) and trimethylation (H3K9me3) in the Il12a promoter using wild-type or DC-cKO BMDCs that were untreated (NT) or stimulated with LPS for 6 h. The Y axis is percentage (%) of total H3. Data are presented as mean ± SEM values and representative of at least three independent experiments. Statistical analyses represent variations in technical replicates *P < 0.05; **P < 0.01.
6 Supplementary Figure 6 Trabid deficiency suppresses the recruitment of Pol II complex components to the Il12b promoter. ChIP analysis of the recruitment of Pol II, Pol II ps5, TFIIB, and TFIID to the indicated promoters in wild-type (WT) or DC-cKO BMDCs. The Y axis is percentage (%) of the DNA bound to the specific factors based on total input DNA. Data are presented as mean ± SEM values and representative of at least three independent experiments. Statistical analyses represent variations in technical replicates. *P < 0.05; **P < 0.01.
7 Supplementary Figure 7 Jmjd2d is regulated by Trabid and mediates induction of Il12 genes. (a,b) ChIP analysis of the binding of indicated Jmjd2 family members to different regions of Il12b locus in wild-type (WT) and DC-cKO BMDCs (a) or wild-type and Rel-KO BMDCs (b) that were either not treated (NT) or stimulated with LPS for 6 h. (c) Immunoblotting analysis of Jmjd2d in wild-type BMDCs transduced with pgipz lentiviral vectors encoding a non-silencing control shrna (C) or two different Kdm4d-specific shrnas. The cells were stimulated with LPS for 6 h to induce the level of Jmjd2d. (d) qrt-pcr analysis of the indicated genes in LPS-stimulated BMDCs described in c. Data are presented as fold relative to the Actb mrna level. (e) Immunoblotting analysis of HA-tagged Trabid (or Trabid C443A) and FLAG-tagged Jmjd2d in anti-flag (Jmjd2d) immunoprecipitates (top two panels) or lysates (lower panels) of HEK293 cells transfected with the indicated plasmids. (f,g) Immunoblotting analysis of Jmjd2d (f) and qrt-pcr analysis of the indicated genes in DC-cKO BMDCs reconstituted with pclxsn(gfp) (Vec) or pclxsn(gfp)- HA-Jmjd2d (Jmjd2d). Data are presented as mean ± SEM values and representative of at least three independent experiments. *P < 0.05; **P<0.01.
8 Supplementary Figure 8 A model of Trabid function in TLR-mediated induction of Il12b. LPS induces the expression of the Jmjd2d gene (Kdm4d) via an unknown signaling pathway. Jmjd2d mediates removal of H3K9me2 and H3K9me3 from the Il12b promoter, thereby promoting c-rel binding to the B element and induction of Il12b expression. The fate of Jmjd2d in TLR-stimulated DCs is tightly controlled by ubiquitination. An unknown E3 ubiquitin ligase mediates Jmjd2d ubiquitination and degradation, whereas Trabid deubiquitinates Jmjd2d to prevent its degradation. Trabid deficiency causes elevated ubiquitination and degradation of Jmjd2d and, thus, reduces the level of Il12b induction. Similar mechanisms may apply to the induction of Il12a and Il23a genes.
9 Supplementary Table 1 List of antibodies used in biochemical assays. Antibody Product number Vender RelB C- 19 Santa Cruz Biotechnology Lamin B C- 20 Santa Cruz Biotechnology TRAF2 C- 20 Santa Cruz Biotechnology TRAF3 H- 122 Santa Cruz Biotechnology ERK K- 23 Santa Cruz Biotechnology JNK1 C- 17 Santa Cruz Biotechnology p38 H- 147 Santa Cruz Biotechnology IKKα H744 Santa Cruz Biotechnology Ubiquitin P4D1 Santa Cruz Biotechnology p105/p50 C- 19 Santa Cruz Biotechnology cebp- β C- 19 Santa Cruz Biotechnology STAT3 C- 20 Santa Cruz Biotechnology c- Rel C Santa Cruz Biotechnology TFIIB (SI- 1)X Santa Cruz Biotechnology TFIID (SI- 1)X Santa Cruz Biotechnology Pol II (N- 20)X Santa Cruz Biotechnology HSP60 H- 1 Santa Cruz Biotechnology Rabbit IgG sc Santa Cruz Biotechnology Phospho- ERK (Y204) E- 4 Santa Cruz Biotechnology Phospho- IκBα (S32) 14D4 Cell Signaling Technology Phospho- JNK (T180/T185) 81E11 Cell Signaling Technology Phospho- p38 (T180/T182) 9211 Cell Signaling Technology Phospho- p105 (S933) 18E6 Cell Signaling Technology Phospho- IKKα/β (S176/S180) 16A6 Cell Signaling Technology Actin C- 4 Sigma- Aldrich HRP- anti- HA HA- 7 Sigma- Aldrich HRP- anti- FLAG M2 Sigma- Aldrich Ubiquityl- H2B Millipore H3K4me Millipore H3K27me Millipore H2B ab1790 Abcam H3K9me2 ab1220 Abcam H3K9me3 ab8898 Abcam JMJD2A ab Abcam JMJD2B ab27531 Abcam JMJD2C ab85454 Abcam JMJD2D ab93694 Abcam Pol II Phospho- Ser 5 ab5131 Abcam p100/p52 TB4 NCI Preclinical Repository
10 Supplementary Table 2 Gene-specific primers used in qrt-pcr experiments Gene Forward primer Reverse primer Il1b AAGCCTCGTGCTGTCGGACC TGAGGCCCAAGGCCACAGGT Il6 CACAGAGGATACCACTCCCAACA TCCACGATTTCCCAGAGAACA Tnf CATCTTCTCAAAATTCGAGTGACAA CCAGCTGCTCCTCCACTTG Il12a ACTAGAGAGACTTCTTCCACAACAAGAG GCACAGGGTCATCATCAAAGAC Il12b GGAGACACCAGCAAAACGAT TCCAGATTCAGACTCCAGGG Il23a GCCAAGAAGAC CATTCCCGA TCAGTGCTACAATCTTCTTCAGAGGACA Il10 CCAGAGCCACATGCTCCTAGA GGTCCTTTGTTTGAAAGAAAGTCTTC Nos2 GTGGTGACAAGCACATTTGG AAGGCCAAACACAGCATACC Kdm4a GGAGCCTTGCTGAGCATCAC TCACTGAAGGAGCCGTCGTC Kdm4b GGCTTTAACTGCGCTGAGTC GTGTGGTCCAGCACTGTGAG Kdm4c CACGGAGGACATGGATCTCT CGAAGGGAATGCCATACTTC Kdm4d Zranb1 GTCTTGGTCGTCGTCCTTGT GGCTATTCCTCAATGCCTGT AATCCCCCTTCAGAAGCTGT TGTCTCCTCCCGATGACTT
11 Supplementary Table 3 QPCR primers used in ChIP assays. Gene Forward primer Reverse primer Il1b pro (κb site) GTTCCGCACATCCTGACTTA CAATTGTGCAGATGGTGTCAA Il6 pro (κb site) TTTCCAATCAGCCCCACC CAGAATGAGCTACAGACATCCC Il12a pro (κb site) ACGCACTTGTCCTTGAGATG CTGACCTTGGGAGACACATTT Il12b pro (κb site) CATTTCCTCTTAACCTGGGATTTC CTGCTCCTGGTGCTTATATACT Il12b pro AGGCCAGGGACAGGAAT AAGAGGTGTGCAATCCTGTAAG Il12b pro TGCACACCTCTTTGCAATTT GAGGGCGTCTCTTATGCTTAT Il12b pro CATAAGAGACGCCCTCAAA GTTACAGGCCCAAAGAATAAAG Il12b pro GGGCCTGTAACACCTACTTATTT ACGTCGAAATCCCAGGTTAAG Il12b pro CATTTCCTCTTAACCTGGGATTTC CTGCTCCTGGTGCTTATATACT Il1b TATA GAATTCCCATCAAGCTTCTCC AGACAGAGAGAGAGAGAGACTTAC Il6 TATA CCCACCCTCCAACAAAGATT ACTCCTCTCTCACAGTCTCAATA Il12a TATA CTGGCGTCTACACTGCTG TTTGGAAATGGGAATCAACACAG Il12b TATA CTGCTCCTGGTGCTTATATACT CTGCTCCTGGTGCTTATATACT Il23a TATA AGGCACTAGGAAAGAGGATCTA GTTCATACCTGGAGGAGTTGG
Supplementary Materials for
 advances.sciencemag.org/cgi/content/full/4/9/eaat5401/dc1 Supplementary Materials for GLK-IKKβ signaling induces dimerization and translocation of the AhR-RORγt complex in IL-17A induction and autoimmune
advances.sciencemag.org/cgi/content/full/4/9/eaat5401/dc1 Supplementary Materials for GLK-IKKβ signaling induces dimerization and translocation of the AhR-RORγt complex in IL-17A induction and autoimmune
TRIM31 is recruited to mitochondria after infection with SeV.
 Supplementary Figure 1 TRIM31 is recruited to mitochondria after infection with SeV. (a) Confocal microscopy of TRIM31-GFP transfected into HEK293T cells for 24 h followed with SeV infection for 6 h. MitoTracker
Supplementary Figure 1 TRIM31 is recruited to mitochondria after infection with SeV. (a) Confocal microscopy of TRIM31-GFP transfected into HEK293T cells for 24 h followed with SeV infection for 6 h. MitoTracker
SUPPLEMENTARY INFORMATION
 doi: 1.138/nature875 a promoter firefly luciferase CNS b Supplementary Figure 1. Dual luciferase assays on enhancer activity of CNS1, 2, and 3. a. promoter sequence was inserted upstream of firefly luciferase
doi: 1.138/nature875 a promoter firefly luciferase CNS b Supplementary Figure 1. Dual luciferase assays on enhancer activity of CNS1, 2, and 3. a. promoter sequence was inserted upstream of firefly luciferase
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb3240 Supplementary Figure 1 GBM cell lines display similar levels of p100 to p52 processing but respond differentially to TWEAK-induced TERT expression according to TERT promoter mutation
DOI: 10.1038/ncb3240 Supplementary Figure 1 GBM cell lines display similar levels of p100 to p52 processing but respond differentially to TWEAK-induced TERT expression according to TERT promoter mutation
Supplementary Figure 1. Characterization of the POP2 transcriptional and post-transcriptional regulatory elements. (A) POP2 nucleotide sequence
 1 5 6 7 8 9 10 11 1 1 1 Supplementary Figure 1. Characterization of the POP transcriptional and post-transcriptional regulatory elements. (A) POP nucleotide sequence depicting the consensus sequence for
1 5 6 7 8 9 10 11 1 1 1 Supplementary Figure 1. Characterization of the POP transcriptional and post-transcriptional regulatory elements. (A) POP nucleotide sequence depicting the consensus sequence for
Revised: RG-RV2 by Fukuhara et al.
 Supplemental Figure 1 The generation of Spns2 conditional knockout mice. (A) Schematic representation of the wild type Spns2 locus (Spns2 + ), the targeted allele, the floxed allele (Spns2 f ) and the
Supplemental Figure 1 The generation of Spns2 conditional knockout mice. (A) Schematic representation of the wild type Spns2 locus (Spns2 + ), the targeted allele, the floxed allele (Spns2 f ) and the
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature10772 Supplementary Figures: Supplementary Figure 1. Location of CNS1 within of the Foxp3 locus highlighting CNS1 and indicating transcription factor binding motifs downstream of TCR,
doi:10.1038/nature10772 Supplementary Figures: Supplementary Figure 1. Location of CNS1 within of the Foxp3 locus highlighting CNS1 and indicating transcription factor binding motifs downstream of TCR,
Supplementary Figure 1
 Supplementary Figure 1 Virus infection induces RNF128 expression. (a,b) RT-PCR analysis of Rnf128 (RNF128) mrna expression in mouse peritoneal macrophages (a) and THP-1 cells (b) upon stimulation with
Supplementary Figure 1 Virus infection induces RNF128 expression. (a,b) RT-PCR analysis of Rnf128 (RNF128) mrna expression in mouse peritoneal macrophages (a) and THP-1 cells (b) upon stimulation with
isolated from ctr and pictreated mice. Activation of effector CD4 +
 Supplementary Figure 1 Bystander inflammation conditioned T reg cells have normal functional suppressive activity and ex vivo phenotype. WT Balb/c mice were treated with polyi:c (pic) or PBS (ctr) via
Supplementary Figure 1 Bystander inflammation conditioned T reg cells have normal functional suppressive activity and ex vivo phenotype. WT Balb/c mice were treated with polyi:c (pic) or PBS (ctr) via
Supplemental Figure 1
 Supplemental Figure 1 A IL-12p7 (pg/ml) 7 6 4 3 2 1 Medium then TLR ligands MDP then TLR ligands Medium then TLR ligands + MDP MDP then TLR ligands + MDP B IL-12p4 (ng/ml) 1.2 1..8.6.4.2. Medium MDP Medium
Supplemental Figure 1 A IL-12p7 (pg/ml) 7 6 4 3 2 1 Medium then TLR ligands MDP then TLR ligands Medium then TLR ligands + MDP MDP then TLR ligands + MDP B IL-12p4 (ng/ml) 1.2 1..8.6.4.2. Medium MDP Medium
Nature Immunology: doi: /ni Supplementary Figure 1
 Supplementary Figure 1 EPRS expression in multiple cell lines upon viral infection. (a,b) EPRS is slightly induced upon viral induction. qpcr of Eprs mrna (a) and immunoblot analysis of corresponding endogenous
Supplementary Figure 1 EPRS expression in multiple cell lines upon viral infection. (a,b) EPRS is slightly induced upon viral induction. qpcr of Eprs mrna (a) and immunoblot analysis of corresponding endogenous
Table S1. Oligonucleotide primer sequences
 Table S1. Oligonucleotide primer sequences Primer Name DNA Sequence (5 -> 3 ) Description CD19c CD19d Cre7 Sfpi1lox1 Sfpi1lox2 Sfpi1lox3 5 -AACCAGTCAACACCCTTCC-3 5 -CCAGACTAGATACAGACCAG-3 5 -TCAGCTACACCAGAGACGG-3
Table S1. Oligonucleotide primer sequences Primer Name DNA Sequence (5 -> 3 ) Description CD19c CD19d Cre7 Sfpi1lox1 Sfpi1lox2 Sfpi1lox3 5 -AACCAGTCAACACCCTTCC-3 5 -CCAGACTAGATACAGACCAG-3 5 -TCAGCTACACCAGAGACGG-3
Supplementary Figure 1. TRIM9 does not affect AP-1, NF-AT or ISRE activity. (a,b) At 24h post-transfection with TRIM9 or vector and indicated
 Supplementary Figure 1. TRIM9 does not affect AP-1, NF-AT or ISRE activity. (a,b) At 24h post-transfection with TRIM9 or vector and indicated reporter luciferase constructs, HEK293T cells were stimulated
Supplementary Figure 1. TRIM9 does not affect AP-1, NF-AT or ISRE activity. (a,b) At 24h post-transfection with TRIM9 or vector and indicated reporter luciferase constructs, HEK293T cells were stimulated
Supplementary Figure 1
 Supplementary Figure 1 Ex2 promotor region Cre IRES cherry pa Ex4 Ex5 Ex1 untranslated Ex3 Ex5 untranslated EYFP pa Rosa26 STOP loxp loxp Cre recombinase EYFP pa Rosa26 loxp 1 kb Interleukin-9 fate reporter
Supplementary Figure 1 Ex2 promotor region Cre IRES cherry pa Ex4 Ex5 Ex1 untranslated Ex3 Ex5 untranslated EYFP pa Rosa26 STOP loxp loxp Cre recombinase EYFP pa Rosa26 loxp 1 kb Interleukin-9 fate reporter
USP19 modulates autophagy and antiviral immune responses by. deubiquitinating Beclin-1
 USP19 modulates autophagy and antiviral immune responses by deubiquitinating Beclin-1 Shouheng Jin 1,2,, Shuo Tian 1,, Yamei Chen 1,, Chuanxia Zhang 1,2, Weihong Xie, 1 Xiaojun Xia 3,4, Jun Cui 1,3* &
USP19 modulates autophagy and antiviral immune responses by deubiquitinating Beclin-1 Shouheng Jin 1,2,, Shuo Tian 1,, Yamei Chen 1,, Chuanxia Zhang 1,2, Weihong Xie, 1 Xiaojun Xia 3,4, Jun Cui 1,3* &
a Lamtor1 (gene) b Lamtor1 (mrna) c WT Lamtor1 Lamtor1 flox Lamtor2 A.U. p = LysM-Cre Lamtor3 Lamtor4 Lamtor5 BMDMs: Φ WT Φ KO β-actin WT BMDMs
 a Lamtor (gene) b Lamtor (mrna) c BMDMs: Φ WT Φ KO Lamtor flox 8 bp LysM-Cre 93 bp..5 p =.4 WT BMDMs: Φ WT Φ KO Lamtor (protein) BMDMs: Φ WT Φ KO Lamtor 8 kda Lamtor 4 kda Lamtor3 4 kda Lamtor4 kda Lamtor5.5
a Lamtor (gene) b Lamtor (mrna) c BMDMs: Φ WT Φ KO Lamtor flox 8 bp LysM-Cre 93 bp..5 p =.4 WT BMDMs: Φ WT Φ KO Lamtor (protein) BMDMs: Φ WT Φ KO Lamtor 8 kda Lamtor 4 kda Lamtor3 4 kda Lamtor4 kda Lamtor5.5
Supplementary Information
 Supplementary Information Supplementary Figure 1: Identification of new regulators of MuSC by a proteome-based shrna screen. (a) FACS plots of GFP + and GFP - cells from Pax7 ICN -Z/EG (upper panel) and
Supplementary Information Supplementary Figure 1: Identification of new regulators of MuSC by a proteome-based shrna screen. (a) FACS plots of GFP + and GFP - cells from Pax7 ICN -Z/EG (upper panel) and
Supplementary Methods Plasmid constructs
 Supplementary Methods Plasmid constructs. Mouse cdna encoding SHP-1, amplified from mrna of RAW264.7 macrophages with primer 5'cgtgcctgcccagacaaactgt3' and 5'cggaattcagacgaatgcccagatcacttcc3', was cloned
Supplementary Methods Plasmid constructs. Mouse cdna encoding SHP-1, amplified from mrna of RAW264.7 macrophages with primer 5'cgtgcctgcccagacaaactgt3' and 5'cggaattcagacgaatgcccagatcacttcc3', was cloned
Supplementary Figure S1
 Supplementary Figure S1 Supplementary Figure S1. CD11b expression on different B cell subsets in anti-snrnp Ig Tg mice. (a) Highly purified follicular (FO) and marginal zone (MZ) B cells were sorted from
Supplementary Figure S1 Supplementary Figure S1. CD11b expression on different B cell subsets in anti-snrnp Ig Tg mice. (a) Highly purified follicular (FO) and marginal zone (MZ) B cells were sorted from
Supplementary Figure 1. GST pull-down analysis of the interaction of GST-cIAP1 (A, B), GSTcIAP1
 Legends Supplementary Figure 1. GST pull-down analysis of the interaction of GST- (A, B), GST mutants (B) or GST- (C) with indicated proteins. A, B, Cell lysate from untransfected HeLa cells were loaded
Legends Supplementary Figure 1. GST pull-down analysis of the interaction of GST- (A, B), GST mutants (B) or GST- (C) with indicated proteins. A, B, Cell lysate from untransfected HeLa cells were loaded
SUPPLEMENTARY INFORMATION
 doi:.8/nature85 Supplementary Methods Plasmid construction Murine RIG-I, HMGB and Rab5 cdnas were obtained by polymerase chain reaction with reverse transcription (RT-PCR) on total RNA from, and then cloned
doi:.8/nature85 Supplementary Methods Plasmid construction Murine RIG-I, HMGB and Rab5 cdnas were obtained by polymerase chain reaction with reverse transcription (RT-PCR) on total RNA from, and then cloned
Supplementary Figures and Figure legends
 Supplementary Figures and Figure legends 3 Supplementary Figure 1. Conditional targeting construct for the murine Satb1 locus with a modified FLEX switch. Schematic of the wild type Satb1 locus; the conditional
Supplementary Figures and Figure legends 3 Supplementary Figure 1. Conditional targeting construct for the murine Satb1 locus with a modified FLEX switch. Schematic of the wild type Satb1 locus; the conditional
Real-time PCR. Total RNA was isolated from purified splenic or LP macrophages using
 Supplementary Methods Real-time PCR. Total RNA was isolated from purified splenic or LP macrophages using the Qiagen RNeasy Mini Kit, according to the manufacturer s protocol with on-column DNase digestion
Supplementary Methods Real-time PCR. Total RNA was isolated from purified splenic or LP macrophages using the Qiagen RNeasy Mini Kit, according to the manufacturer s protocol with on-column DNase digestion
Nature Biotechnology: doi: /nbt.4086
 Ag (-) anti-cd3 p815 p815-hcd2 Ag (-) anti-cd3 p815 p815-hcd2 Ag (-) anti-cd3 p815 p815-hcd2 Ag (-) anti-cd3 p815 p815-hcd2 Ag (-) anti-cd3 p815 p815-hcd2 Ag (-) anti-cd3 p815 p815-hcd2 Ag (-) anti-cd3
Ag (-) anti-cd3 p815 p815-hcd2 Ag (-) anti-cd3 p815 p815-hcd2 Ag (-) anti-cd3 p815 p815-hcd2 Ag (-) anti-cd3 p815 p815-hcd2 Ag (-) anti-cd3 p815 p815-hcd2 Ag (-) anti-cd3 p815 p815-hcd2 Ag (-) anti-cd3
Supplementary Methods
 Supplementary Methods Microarray Data Analysis Gene expression data were obtained by hybridising a total of 24 samples from 6 experimental groups (n=4 per group) to Illumina HumanHT-12 Expression BeadChips.
Supplementary Methods Microarray Data Analysis Gene expression data were obtained by hybridising a total of 24 samples from 6 experimental groups (n=4 per group) to Illumina HumanHT-12 Expression BeadChips.
Online Supplementary Information
 Online Supplementary Information NLRP4 negatively regulates type I interferon signaling by targeting TBK1 for degradation via E3 ubiquitin ligase DTX4 Jun Cui 1,4,6,7, Yinyin Li 1,5,6,7, Liang Zhu 1, Dan
Online Supplementary Information NLRP4 negatively regulates type I interferon signaling by targeting TBK1 for degradation via E3 ubiquitin ligase DTX4 Jun Cui 1,4,6,7, Yinyin Li 1,5,6,7, Liang Zhu 1, Dan
To determine the effect of N1IC in the susceptibility of T cells to the tolerogenic effect of tumorassociated
 Supplementary Methods Tolerogenic effect of MDSC To determine the effect of N1IC in the susceptibility of T cells to the tolerogenic effect of tumorassociated MDSC in vivo, we used a model described previously
Supplementary Methods Tolerogenic effect of MDSC To determine the effect of N1IC in the susceptibility of T cells to the tolerogenic effect of tumorassociated MDSC in vivo, we used a model described previously
SI Appendix. Tumor-specific CD8 + Tc9 cells are superior effector than Tc1 cells for. adoptive immunotherapy of cancers
 SI Appendix Tumor-specific CD8 + Tc9 cells are superior effector than Tc1 cells for adoptive immunotherapy of cancers Yong Lu, Bangxing Hong, Haiyan Li, Yuhuan Zheng, Mingjun Zhang, Siqing Wang, Jianfei
SI Appendix Tumor-specific CD8 + Tc9 cells are superior effector than Tc1 cells for adoptive immunotherapy of cancers Yong Lu, Bangxing Hong, Haiyan Li, Yuhuan Zheng, Mingjun Zhang, Siqing Wang, Jianfei
Supplementary Figure 1 Validate the expression of mir-302b after bacterial infection by northern
 Supplementary Figure 1 Validate the expression of mir-302b after bacterial infection by northern blot. Northern blot analysis of mir-302b expression following infection with PAO1, PAK and Kp in (A) lung
Supplementary Figure 1 Validate the expression of mir-302b after bacterial infection by northern blot. Northern blot analysis of mir-302b expression following infection with PAO1, PAK and Kp in (A) lung
Flowcytometry-based purity analysis of peritoneal macrophage culture.
 Liao et al., KLF4 regulates macrophage polarization Revision of Manuscript 45444-RG- Supplementary Figure Legends Figure S Flowcytometry-based purity analysis of peritoneal macrophage culture. Thioglycollate
Liao et al., KLF4 regulates macrophage polarization Revision of Manuscript 45444-RG- Supplementary Figure Legends Figure S Flowcytometry-based purity analysis of peritoneal macrophage culture. Thioglycollate
Supplementary Material & Methods
 Supplementary Material & Methods Affymetrix micro-array Total RNA was extracted using Trizol reagent (Invitrogen, Carlsbad, USA). RNA concentration and quality was determined using the ND-1000 spectrophotometer
Supplementary Material & Methods Affymetrix micro-array Total RNA was extracted using Trizol reagent (Invitrogen, Carlsbad, USA). RNA concentration and quality was determined using the ND-1000 spectrophotometer
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature09732 Supplementary Figure 1: Depletion of Fbw7 results in elevated Mcl-1 abundance. a, Total thymocytes from 8-wk-old Lck-Cre/Fbw7 +/fl (Control) or Lck-Cre/Fbw7 fl/fl (Fbw7 KO) mice
doi:10.1038/nature09732 Supplementary Figure 1: Depletion of Fbw7 results in elevated Mcl-1 abundance. a, Total thymocytes from 8-wk-old Lck-Cre/Fbw7 +/fl (Control) or Lck-Cre/Fbw7 fl/fl (Fbw7 KO) mice
Supplementary Figure 1
 Supplementary Figure 1 PTEN promotes virus-induced expression of IFNB1 and its downstream genes. (a) Quantitative RT-PCR analysis of IFNB1 mrna (left) and ELISA of IFN-β (right) in HEK 293 cells (2 10
Supplementary Figure 1 PTEN promotes virus-induced expression of IFNB1 and its downstream genes. (a) Quantitative RT-PCR analysis of IFNB1 mrna (left) and ELISA of IFN-β (right) in HEK 293 cells (2 10
Nature Medicine doi: /nm.3554
 SUPPLEMENTARY FIGURES LEGENDS Supplementary Figure 1: Generation, purification and characterization of recombinant mouse IL-35 (ril-35). High-Five insect cells expressing high levels of the bicistronic
SUPPLEMENTARY FIGURES LEGENDS Supplementary Figure 1: Generation, purification and characterization of recombinant mouse IL-35 (ril-35). High-Five insect cells expressing high levels of the bicistronic
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb3562 In the format provided by the authors and unedited. Supplementary Figure 1 Glucose deficiency induced FH-ATF2 interaction. In b-m, immunoblotting or immunoprecipitation analyses were
DOI: 10.1038/ncb3562 In the format provided by the authors and unedited. Supplementary Figure 1 Glucose deficiency induced FH-ATF2 interaction. In b-m, immunoblotting or immunoprecipitation analyses were
Supplementary Figure 1 Generation of migg1-yf mice. (A) Targeting strategy. Upper panel: schematic organization of the murine ɣ1 immunoglobulin
 Supplementary Figure 1 Generation of migg1-yf mice. (A) Targeting strategy. Upper panel: schematic organization of the murine ɣ1 immunoglobulin locus. The EcoRI restriction site between exons M1 and M2
Supplementary Figure 1 Generation of migg1-yf mice. (A) Targeting strategy. Upper panel: schematic organization of the murine ɣ1 immunoglobulin locus. The EcoRI restriction site between exons M1 and M2
Supplementary Fig. 1 Generation and genotyping of ThPOK / mice
 Supplementary Fig. 1 Generation and genotyping of ThPOK / mice (a). The strategy used to target the ThPOK locus is schematically depicted. A targeting construct was generated by inserting a loxp site (filled
Supplementary Fig. 1 Generation and genotyping of ThPOK / mice (a). The strategy used to target the ThPOK locus is schematically depicted. A targeting construct was generated by inserting a loxp site (filled
OX40 MARKET LANDSCAPE
 OX40 Agonist OX40 MARKET LANDSCAPE OX40 (a.k.a. CD134) is a costimulatory molecule belonging to the TNF receptor family expressed primarily on activated effector T (T eff) cells and naive regulatory T
OX40 Agonist OX40 MARKET LANDSCAPE OX40 (a.k.a. CD134) is a costimulatory molecule belonging to the TNF receptor family expressed primarily on activated effector T (T eff) cells and naive regulatory T
Sarker et al. Supplementary Material. Subcellular Fractionation
 Supplementary Material Subcellular Fractionation Transfected 293T cells were harvested with phosphate buffered saline (PBS) and centrifuged at 2000 rpm (500g) for 3 min. The pellet was washed, re-centrifuged
Supplementary Material Subcellular Fractionation Transfected 293T cells were harvested with phosphate buffered saline (PBS) and centrifuged at 2000 rpm (500g) for 3 min. The pellet was washed, re-centrifuged
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb3209 Supplementary Figure 1 IR induces the association of FH with chromatin. a, U2OS cells synchronized by thymidine double block (2 mm) underwent no release (G1 phase) or release for 2
DOI: 10.1038/ncb3209 Supplementary Figure 1 IR induces the association of FH with chromatin. a, U2OS cells synchronized by thymidine double block (2 mm) underwent no release (G1 phase) or release for 2
Pei et al. Supplementary Figure S1
 Pei et al. Supplementary Figure S1 C H-CUL9: + + + + + Myc-ROC1: - - + + + U2OS/pcDN3 U2OS/H-CUL9 U2OS/ + H-CUL9 IP: -H -myc input -H -myc 1 2 3 4 5 H-CUL9 Myc-ROC1 -H -H -H -H H-CUL9: wt RR myc-roc1:
Pei et al. Supplementary Figure S1 C H-CUL9: + + + + + Myc-ROC1: - - + + + U2OS/pcDN3 U2OS/H-CUL9 U2OS/ + H-CUL9 IP: -H -myc input -H -myc 1 2 3 4 5 H-CUL9 Myc-ROC1 -H -H -H -H H-CUL9: wt RR myc-roc1:
Supplemental Information. Inflammatory Ly6C high Monocytes Protect. against Candidiasis through IL-15-Driven. NK Cell/Neutrophil Activation
 Immunity, Volume 46 Supplemental Information Inflammatory Ly6C high Monocytes Protect against Candidiasis through IL-15-Driven NK Cell/Neutrophil Activation Jorge Domínguez-Andrés, Lidia Feo-Lucas, María
Immunity, Volume 46 Supplemental Information Inflammatory Ly6C high Monocytes Protect against Candidiasis through IL-15-Driven NK Cell/Neutrophil Activation Jorge Domínguez-Andrés, Lidia Feo-Lucas, María
Table S1. Antibodies and recombinant proteins used in this study
 Table S1. Antibodies and recombinant proteins used in this study Labeled Antibody Clone Cat. no. Streptavidin-PerCP BD Biosciences 554064 Biotin anti-mouse CD25 7D4 BD Biosciences 553070 PE anti-mouse
Table S1. Antibodies and recombinant proteins used in this study Labeled Antibody Clone Cat. no. Streptavidin-PerCP BD Biosciences 554064 Biotin anti-mouse CD25 7D4 BD Biosciences 553070 PE anti-mouse
Supplementary Fig. 1 Kinetics of appearence of the faster migrating form of Bcl-10.
 α-cd3 + α-cd28: Time (min): + + + + + + + + + 0 5 15 30 60 120 180 240 300 360 360 n.s. Supplementary Fig. 1 Kinetics of appearence of the faster migrating form of. Immunoblot of lysates from Jurkat cells
α-cd3 + α-cd28: Time (min): + + + + + + + + + 0 5 15 30 60 120 180 240 300 360 360 n.s. Supplementary Fig. 1 Kinetics of appearence of the faster migrating form of. Immunoblot of lysates from Jurkat cells
Nature Immunology: doi: /ni Supplementary Figure 1
 Supplementary Figure 1 Sox2 localizes in neutrophils of mouse spleen. (a) Microscopy analysis of Sox2 and MPO in wild-type mouse spleen. Nucl, nucleus. (b) Immunohistochemistry staining of Sox2 and MPO
Supplementary Figure 1 Sox2 localizes in neutrophils of mouse spleen. (a) Microscopy analysis of Sox2 and MPO in wild-type mouse spleen. Nucl, nucleus. (b) Immunohistochemistry staining of Sox2 and MPO
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:10.1038/nature11809 Supplementary Figure 1. Antibiotic treatment reduces intestinal bacterial load and allows access of non-pathogenic bacteria to the MLN, inducing intestinal
SUPPLEMENTARY INFORMATION doi:10.1038/nature11809 Supplementary Figure 1. Antibiotic treatment reduces intestinal bacterial load and allows access of non-pathogenic bacteria to the MLN, inducing intestinal
Supplementary information.
 Supplementary information. Construction of JMJD3 plasmids and other vectors plasmid, pfksd346, encoding the ORF of human JMJD3 (KI346) was obtained from the Kazusa DN Research Institute, Japan. Sequence
Supplementary information. Construction of JMJD3 plasmids and other vectors plasmid, pfksd346, encoding the ORF of human JMJD3 (KI346) was obtained from the Kazusa DN Research Institute, Japan. Sequence
Supplementary Fig S1: Bone marrow macrophages and liver Kupffer cells increase IL-1 in response to adenosine signaling. (a) Murine bone marrow
 Supplementary Fig S1: Bone marrow macrophages and liver Kupffer cells increase IL-1 in response to adenosine signaling. (a) Murine bone marrow derived macrophages (BMDM) were primed with LPS for 16 hrs,
Supplementary Fig S1: Bone marrow macrophages and liver Kupffer cells increase IL-1 in response to adenosine signaling. (a) Murine bone marrow derived macrophages (BMDM) were primed with LPS for 16 hrs,
Supplementary Information
 Supplementary Information Mutual reinforcement of inflammation and carcinogenesis by the Helicobacter pylori CagA oncoprotein Nobumi Suzuki, Naoko Murata-Kamiya, Kohei Yanagiya, Wataru Suda, Masahira Hattori,
Supplementary Information Mutual reinforcement of inflammation and carcinogenesis by the Helicobacter pylori CagA oncoprotein Nobumi Suzuki, Naoko Murata-Kamiya, Kohei Yanagiya, Wataru Suda, Masahira Hattori,
b alternative classical none
 Supplementary Figure. 1: Related to Figure.1 a d e b alternative classical none NIK P-IkBa Total IkBa Tubulin P52 (Lighter) P52 (Darker) RelB (Lighter) RelB (Darker) HDAC1 Control-Sh RelB-Sh NF-kB2-Sh
Supplementary Figure. 1: Related to Figure.1 a d e b alternative classical none NIK P-IkBa Total IkBa Tubulin P52 (Lighter) P52 (Darker) RelB (Lighter) RelB (Darker) HDAC1 Control-Sh RelB-Sh NF-kB2-Sh
Nature Immunology: doi: /ni Supplementary Figure 1. Construction of lnckdm2b RFP reporter and lnckdm2b-knockout mice.
 Supplementary Figure 1 Construction of lnckdm2b RFP reporter and lnckdm2b-knockout mice. (a) ILC3s (Lin CD45 lo CD90 hi ) were isolated from small intestines, followed by immunofluorescence staining. LncKdm2b
Supplementary Figure 1 Construction of lnckdm2b RFP reporter and lnckdm2b-knockout mice. (a) ILC3s (Lin CD45 lo CD90 hi ) were isolated from small intestines, followed by immunofluorescence staining. LncKdm2b
Supporting Online Material. 3. Analysis of in vivo TLR4-mediated response in TRIF-deficient mice
 Supporting Online Material 1. Materials and Methods 2. Generation of TRIF-deficient mice 3. Analysis of in vivo TLR4-mediated response in TRIF-deficient mice 4. Measurement of LPS-induced IRAK-1 kinase
Supporting Online Material 1. Materials and Methods 2. Generation of TRIF-deficient mice 3. Analysis of in vivo TLR4-mediated response in TRIF-deficient mice 4. Measurement of LPS-induced IRAK-1 kinase
Supplemental Table S1. RT-PCR primers used in this study
 Supplemental Table S1. RT-PCR primers used in this study -----------------------------------------------------------------------------------------------------------------------------------------------
Supplemental Table S1. RT-PCR primers used in this study -----------------------------------------------------------------------------------------------------------------------------------------------
(A-B) P2ry14 expression was assessed by (A) genotyping (upper arrow: WT; lower
 Supplementary Figures S1. (A-B) P2ry14 expression was assessed by (A) genotyping (upper arrow: ; lower arrow: KO) and (B) q-pcr analysis with Lin- cells, The white vertical line in panel A indicates that
Supplementary Figures S1. (A-B) P2ry14 expression was assessed by (A) genotyping (upper arrow: ; lower arrow: KO) and (B) q-pcr analysis with Lin- cells, The white vertical line in panel A indicates that
Anti-ERK1/2, anti-p-erk1/2, anti-p38 anti-p-msk1 (Thr581) and anti-p-msk2
 Supplementary methods Antibodies Anti-ERK1/2, anti-p-erk1/2, anti-p38 anti-p-msk1 (Thr581) and anti-p-msk2 polyclonal were from Cell Signalling and the anti-p-histone H3(Ser1) antibody was from Upstate.
Supplementary methods Antibodies Anti-ERK1/2, anti-p-erk1/2, anti-p38 anti-p-msk1 (Thr581) and anti-p-msk2 polyclonal were from Cell Signalling and the anti-p-histone H3(Ser1) antibody was from Upstate.
DRG Pituitary Cerebral Cortex
 Liver Spinal cord Pons Atg5 -/- Atg5 +/+ DRG Pituitary Cerebral Cortex WT KO Supplementary Figure S1 Ubiquitin-positive IBs accumulate in Atg5 -/- tissues. Atg5 -/- neonatal tissues were fixed and decalcified.
Liver Spinal cord Pons Atg5 -/- Atg5 +/+ DRG Pituitary Cerebral Cortex WT KO Supplementary Figure S1 Ubiquitin-positive IBs accumulate in Atg5 -/- tissues. Atg5 -/- neonatal tissues were fixed and decalcified.
Supplementary Methods
 Supplementary Methods Mice injections. C57BL/6 female mice 6-10 weeks of age were purchased from the Jackson Laboratory. Soluble rapamycin (Sigma) was diluted in PBS and administered i.p. to mice at 1.5
Supplementary Methods Mice injections. C57BL/6 female mice 6-10 weeks of age were purchased from the Jackson Laboratory. Soluble rapamycin (Sigma) was diluted in PBS and administered i.p. to mice at 1.5
Yalin Emre, Magali Irla, Isabelle Dunand- Sauthier, Romain Ballet, Mehdi Meguenani, Stephane Jemelin, Christian Vesin, Walter Reith and Beat A.
 Supplementary information Thymic epithelial cell expansion through matricellular protein CYR61 boosts progenitor homing and T- cell output Yalin Emre, Magali Irla, Isabelle Dunand- Sauthier, Romain Ballet,
Supplementary information Thymic epithelial cell expansion through matricellular protein CYR61 boosts progenitor homing and T- cell output Yalin Emre, Magali Irla, Isabelle Dunand- Sauthier, Romain Ballet,
Supplementary Information (Ha, et. al) Supplementary Figures Supplementary Fig. S1
 Supplementary Information (Ha, et. al) Supplementary Figures Supplementary Fig. S1 a His-ORMDL3 ~ 17 His-ORMDL3 GST-ORMDL3 - + - + IPTG GST-ORMDL3 ~ b Integrated Density (ORMDL3/ -actin) 0.4 0.3 0.2 0.1
Supplementary Information (Ha, et. al) Supplementary Figures Supplementary Fig. S1 a His-ORMDL3 ~ 17 His-ORMDL3 GST-ORMDL3 - + - + IPTG GST-ORMDL3 ~ b Integrated Density (ORMDL3/ -actin) 0.4 0.3 0.2 0.1
SUPPLEMENTARY INFORMATION
 doi:.38/nature899 Supplementary Figure Suzuki et al. a c p7 -/- / WT ratio (+)/(-) p7 -/- / WT ratio Log X 3. Fold change by treatment ( (+)/(-)) Log X.5 3-3. -. b Fold change by treatment ( (+)/(-)) 8
doi:.38/nature899 Supplementary Figure Suzuki et al. a c p7 -/- / WT ratio (+)/(-) p7 -/- / WT ratio Log X 3. Fold change by treatment ( (+)/(-)) Log X.5 3-3. -. b Fold change by treatment ( (+)/(-)) 8
Supplementary Figure 1. Xbp1 deficiency does not alter hematopoietic cellularity.
 Supplementary Figure 1 Xbp1 deficiency does not alter hematopoietic cellularity. Absolute number of leukocytes obtained from bone marrow (BM, 2 tibias and 2 femurs per mouse) of Xbp1 f/f and Xbp1 Vav1
Supplementary Figure 1 Xbp1 deficiency does not alter hematopoietic cellularity. Absolute number of leukocytes obtained from bone marrow (BM, 2 tibias and 2 femurs per mouse) of Xbp1 f/f and Xbp1 Vav1
Fig. S1. eif6 expression in HEK293 transfected with shrna against eif6 or pcmv-eif6 vector.
 Fig. S1. eif6 expression in HEK293 transfected with shrna against eif6 or pcmv-eif6 vector. (a) Western blotting analysis and (b) qpcr analysis of eif6 expression in HEK293 T cells transfected with either
Fig. S1. eif6 expression in HEK293 transfected with shrna against eif6 or pcmv-eif6 vector. (a) Western blotting analysis and (b) qpcr analysis of eif6 expression in HEK293 T cells transfected with either
Supplementary Table 1. Sequences for BTG2 and BRCA1 sirnas.
 Supplementary Table 1. Sequences for BTG2 and BRCA1 sirnas. Target Gene Non-target / Control BTG2 BRCA1 NFE2L2 Target Sequence ON-TARGET plus Non-targeting sirna # 1 (Cat# D-001810-01-05) sirna1: GAACCGACAUGCUCCCGGA
Supplementary Table 1. Sequences for BTG2 and BRCA1 sirnas. Target Gene Non-target / Control BTG2 BRCA1 NFE2L2 Target Sequence ON-TARGET plus Non-targeting sirna # 1 (Cat# D-001810-01-05) sirna1: GAACCGACAUGCUCCCGGA
HPV E6 oncoprotein targets histone methyltransferases for modulating specific. Chih-Hung Hsu, Kai-Lin Peng, Hua-Ci Jhang, Chia-Hui Lin, Shwu-Yuan Wu,
 1 HPV E oncoprotein targets histone methyltransferases for modulating specific gene transcription 3 5 Chih-Hung Hsu, Kai-Lin Peng, Hua-Ci Jhang, Chia-Hui Lin, Shwu-Yuan Wu, Cheng-Ming Chiang, Sheng-Chung
1 HPV E oncoprotein targets histone methyltransferases for modulating specific gene transcription 3 5 Chih-Hung Hsu, Kai-Lin Peng, Hua-Ci Jhang, Chia-Hui Lin, Shwu-Yuan Wu, Cheng-Ming Chiang, Sheng-Chung
ClustalW algorithm. The alignment statistics are 96.9% for both pairwise identity and percent identical sites.
 Identity Human Mouse Supplemental Figure 1. Amino acid sequence alignment of -MyHC. The alignment was created using the ClustalW algorithm. The alignment statistics are 96.9% for both pairwise identity
Identity Human Mouse Supplemental Figure 1. Amino acid sequence alignment of -MyHC. The alignment was created using the ClustalW algorithm. The alignment statistics are 96.9% for both pairwise identity
Supplementary Figure 1: Analysis of monocyte subsets and lineage relationships. (a) Gating strategy for definition of MDP and cmop populations in BM
 Supplementary Figure 1: Analysis of monocyte subsets and lineage relationships. (a) Gating strategy for definition of MDP and cmop populations in BM of Cx3cr1 GFP/+ mice related to Fig. 1a. MDP was defined
Supplementary Figure 1: Analysis of monocyte subsets and lineage relationships. (a) Gating strategy for definition of MDP and cmop populations in BM of Cx3cr1 GFP/+ mice related to Fig. 1a. MDP was defined
Supplemental figure 1: Phenotype of IMC and MDSC after purification. A. Gating
 Supplemental Figure Legend: Supplemental figure 1: Phenotype of IMC and MDSC after purification. A. Gating strategy for mouse MDSC. CD11b + Ly6C high Ly6G - cells are defined as M-MDSC. CD11b + Ly6C low
Supplemental Figure Legend: Supplemental figure 1: Phenotype of IMC and MDSC after purification. A. Gating strategy for mouse MDSC. CD11b + Ly6C high Ly6G - cells are defined as M-MDSC. CD11b + Ly6C low
used at a final concentration of 5 ng/ml. Rabbit anti-bim and mouse anti-mkp2 antibodies were
 1 Supplemental Methods Reagents and chemicals: TGFβ was a generous gift from Genzyme Inc. (Cambridge, MA) and was used at a final concentration of 5 ng/ml. Rabbit anti-bim and mouse anti-mkp2 antibodies
1 Supplemental Methods Reagents and chemicals: TGFβ was a generous gift from Genzyme Inc. (Cambridge, MA) and was used at a final concentration of 5 ng/ml. Rabbit anti-bim and mouse anti-mkp2 antibodies
Nature Immunology: doi: /ni.3694
 Supplementary Figure 1 Expression of Bhlhe41 and Bhlhe40 in B cell development and mature B cell subsets. (a) Scatter plot showing differential expression of genes between splenic B-1a cells and follicular
Supplementary Figure 1 Expression of Bhlhe41 and Bhlhe40 in B cell development and mature B cell subsets. (a) Scatter plot showing differential expression of genes between splenic B-1a cells and follicular
Stabilization of the Transcription Factor Foxp3 by the Deubiquitinase USP7 Increases Treg-Cell-Suppressive Capacity
 Immunity, Volume 39 Supplemental Information Stabilization of the Transcription Factor Foxp3 by the Deubiquitinase USP7 Increases Treg-Cell-Suppressive Capacity Jorg van Loosdregt, Veerle Fleskens, Juan
Immunity, Volume 39 Supplemental Information Stabilization of the Transcription Factor Foxp3 by the Deubiquitinase USP7 Increases Treg-Cell-Suppressive Capacity Jorg van Loosdregt, Veerle Fleskens, Juan
indicated numbers of pups at day of life (DOL) 10, or embryonic day (ED) B. Male mice of
 SUPPLEMENTRY FIGURE LEGENDS Figure S1. USP44 loss leads to chromosome missegregation.. Genotypes obtained from the indicated numbers of pups at day of life (DOL) 10, or embryonic day (ED) 13.5.. Male mice
SUPPLEMENTRY FIGURE LEGENDS Figure S1. USP44 loss leads to chromosome missegregation.. Genotypes obtained from the indicated numbers of pups at day of life (DOL) 10, or embryonic day (ED) 13.5.. Male mice
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature11326 Supplementary Figure 1: Histone exchange increases over the ORF in a set2 mutant. (a) Gene average analysis. Schematic representation of the bin distribution over the coding and
doi:10.1038/nature11326 Supplementary Figure 1: Histone exchange increases over the ORF in a set2 mutant. (a) Gene average analysis. Schematic representation of the bin distribution over the coding and
Nature Genetics: doi: /ng Supplementary Figure 1. ChIP-seq genome browser views of BRM occupancy at previously identified BRM targets.
 Supplementary Figure 1 ChIP-seq genome browser views of BRM occupancy at previously identified BRM targets. Gene structures are shown underneath each panel. Supplementary Figure 2 pref6::ref6-gfp complements
Supplementary Figure 1 ChIP-seq genome browser views of BRM occupancy at previously identified BRM targets. Gene structures are shown underneath each panel. Supplementary Figure 2 pref6::ref6-gfp complements
Autocrine Complement Inhibits IL10-Dependent T-Cell Mediated. Antitumor Immunity to Promote Tumor Progression
 Supplemental Information Autocrine Complement Inhibits IL10-Dependent T-Cell Mediated Antitumor Immunity to Promote Tumor Progression Yu Wang, Sheng-Nan Sun, Qing Liu, Yang-Yang Yu, Jian Guo, Kun Wang,
Supplemental Information Autocrine Complement Inhibits IL10-Dependent T-Cell Mediated Antitumor Immunity to Promote Tumor Progression Yu Wang, Sheng-Nan Sun, Qing Liu, Yang-Yang Yu, Jian Guo, Kun Wang,
Bcl-2 family member Bcl-G is not a pro-apoptotic BH3-only protein
 Bcl-2 family member Bcl-G is not a pro-apoptotic BH3-only protein Maybelline Giam 1,2, Toru Okamoto 1,2,3, Justine D. Mintern 1,2,4, Andreas Strasser 1,2 and Philippe Bouillet 1, 2 1 The Walter and Eliza
Bcl-2 family member Bcl-G is not a pro-apoptotic BH3-only protein Maybelline Giam 1,2, Toru Okamoto 1,2,3, Justine D. Mintern 1,2,4, Andreas Strasser 1,2 and Philippe Bouillet 1, 2 1 The Walter and Eliza
Supplementary figures
 Mucida et al. Supplementary material Supplementary figures Supplementary Figure 1. Oral administration of OVA suppresses Th2 differentiation, Germinal Center (GC) formation and immunoglobulin class switching
Mucida et al. Supplementary material Supplementary figures Supplementary Figure 1. Oral administration of OVA suppresses Th2 differentiation, Germinal Center (GC) formation and immunoglobulin class switching
Supplementary Figure 1
 Supplementary Figure 1 Supplementary Fig. 1 shrna mediated knockdown of ZRSR2 in K562 and 293T cells. (a) ZRSR2 transcript levels in stably transduced K562 cells were determined using qrt-pcr. GAPDH was
Supplementary Figure 1 Supplementary Fig. 1 shrna mediated knockdown of ZRSR2 in K562 and 293T cells. (a) ZRSR2 transcript levels in stably transduced K562 cells were determined using qrt-pcr. GAPDH was
Table S1. Primer sequences
 Table S1. Primer sequences Primers for quantitative PCR Tgf 1 Forward Tgf 1 Reverse Tgf 2Forward Tgf 2Reverse Tgf 3 Forward Tgf 3 Reverse Tgf r1 Forward Tgf r1 Reverse Tgf r2 Forward Tgf r2 Reverse Thbs1
Table S1. Primer sequences Primers for quantitative PCR Tgf 1 Forward Tgf 1 Reverse Tgf 2Forward Tgf 2Reverse Tgf 3 Forward Tgf 3 Reverse Tgf r1 Forward Tgf r1 Reverse Tgf r2 Forward Tgf r2 Reverse Thbs1
Cell death analysis using the high content bioimager BD PathwayTM 855 instrument (BD
 Supplemental information Materials and Methods: Cell lines, reagents and antibodies: Wild type (A3) and caspase-8 -/- (I9.2) Jurkat cells were cultured in RPMI 164 medium (Life Technologies) supplemented
Supplemental information Materials and Methods: Cell lines, reagents and antibodies: Wild type (A3) and caspase-8 -/- (I9.2) Jurkat cells were cultured in RPMI 164 medium (Life Technologies) supplemented
Figure S1: GM-CSF upregulates CCL17 expression in in vitro-derived and ex vivo murine macrophage populations
 Figure S1 A B C Figure S1: GM-CSF upregulates CCL17 expression in in vitro-derived and ex vivo murine macrophage populations (A-B) Murine bone cells were cultured in either M-CSF (5,000 U/ml) or GM-CSF
Figure S1 A B C Figure S1: GM-CSF upregulates CCL17 expression in in vitro-derived and ex vivo murine macrophage populations (A-B) Murine bone cells were cultured in either M-CSF (5,000 U/ml) or GM-CSF
Table S1. Primers used in RT-PCR studies (all in 5 to 3 direction)
 Table S1. Primers used in RT-PCR studies (all in 5 to 3 direction) Epo Fw CTGTATCATGGACCACCTCGG Epo Rw TGAAGCACAGAAGCTCTTCGG Jak2 Fw ATCTGACCTTTCCATCTGGGG Jak2 Rw TGGTTGGGTGGATACCAGATC Stat5A Fw TTACTGAAGATCAAGCTGGGG
Table S1. Primers used in RT-PCR studies (all in 5 to 3 direction) Epo Fw CTGTATCATGGACCACCTCGG Epo Rw TGAAGCACAGAAGCTCTTCGG Jak2 Fw ATCTGACCTTTCCATCTGGGG Jak2 Rw TGGTTGGGTGGATACCAGATC Stat5A Fw TTACTGAAGATCAAGCTGGGG
Tumour 2. Tumour 9. Tumour 25 Tumour 26. Tumour 27. t(6;16) doi: /nature06159 SUPPLEMENTARY INFORMATION Supplementary Figure 1
 doi: 1.18/nature6159 SUPPLEMENTARY INFORMATION Supplementary Figure 1 Tumour Tumour 9 t(6;16) Tumour 5 Tumour 6 Tumour 7 www.nature.com/nature 1 doi: 1.18/nature6159 SUPPLEMENTARY INFORMATION Supplementary
doi: 1.18/nature6159 SUPPLEMENTARY INFORMATION Supplementary Figure 1 Tumour Tumour 9 t(6;16) Tumour 5 Tumour 6 Tumour 7 www.nature.com/nature 1 doi: 1.18/nature6159 SUPPLEMENTARY INFORMATION Supplementary
Supplementary Methods
 Supplementary Methods Reverse transcribed Quantitative PCR. Total RNA was isolated from bone marrow derived macrophages using RNeasy Mini Kit (Qiagen), DNase-treated (Promega RQ1), and reverse transcribed
Supplementary Methods Reverse transcribed Quantitative PCR. Total RNA was isolated from bone marrow derived macrophages using RNeasy Mini Kit (Qiagen), DNase-treated (Promega RQ1), and reverse transcribed
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/5/233/ra50/dc1 Supplementary Materials for Epidermal Growth Factor Receptor Is Essential for Toll-Like Receptor 3 Signaling Michifumi Yamashita, Saurabh Chattopadhyay,
www.sciencesignaling.org/cgi/content/full/5/233/ra50/dc1 Supplementary Materials for Epidermal Growth Factor Receptor Is Essential for Toll-Like Receptor 3 Signaling Michifumi Yamashita, Saurabh Chattopadhyay,
ASPP1 Fw GGTTGGGAATCCACGTGTTG ASPP1 Rv GCCATATCTTGGAGCTCTGAGAG
 Supplemental Materials and Methods Plasmids: the following plasmids were used in the supplementary data: pwzl-myc- Lats2 (Aylon et al, 2006), pretrosuper-vector and pretrosuper-shp53 (generous gift of
Supplemental Materials and Methods Plasmids: the following plasmids were used in the supplementary data: pwzl-myc- Lats2 (Aylon et al, 2006), pretrosuper-vector and pretrosuper-shp53 (generous gift of
monoclonal antibody. (a) The specificity of the anti-rhbdd1 monoclonal antibody was examined in
 Supplementary information Supplementary figures Supplementary Figure 1 Determination of the s pecificity of in-house anti-rhbdd1 mouse monoclonal antibody. (a) The specificity of the anti-rhbdd1 monoclonal
Supplementary information Supplementary figures Supplementary Figure 1 Determination of the s pecificity of in-house anti-rhbdd1 mouse monoclonal antibody. (a) The specificity of the anti-rhbdd1 monoclonal
SUPPLEMENTARY INFORMATION
 DOI: 1.138/ncb37 a mrna relative expression 1.3 1..8.5.3. CTR NGPS OCT4 SOX2 KLF4 c-myc b BANF1 mrna levels 1.2 1..8.6.4.2. plko.1 shbanf1 c Tra-1-6+ colonies 3 1 plko.1 shbanf1 d BANF1 locus Plasmid donor
DOI: 1.138/ncb37 a mrna relative expression 1.3 1..8.5.3. CTR NGPS OCT4 SOX2 KLF4 c-myc b BANF1 mrna levels 1.2 1..8.6.4.2. plko.1 shbanf1 c Tra-1-6+ colonies 3 1 plko.1 shbanf1 d BANF1 locus Plasmid donor
Supplementary Figures S1-S5. a b c
 Supplementary Figures S1-S5 a b c Supplementary Figure S1. Generation of IL-17RD-deficient mice. (a) Schematic shows the murine il-17rd gene and the genetrap targeting vector, containing a promoter-less
Supplementary Figures S1-S5 a b c Supplementary Figure S1. Generation of IL-17RD-deficient mice. (a) Schematic shows the murine il-17rd gene and the genetrap targeting vector, containing a promoter-less
Nanog-Luc. R-Luc. 40 HA-Klf Relative protein level of Klf USP21. Vector USP2 USP21 *** *** *** ***
 :Flag Relative protein level of Relative protein level of : Flag Vec 21LV 21SV 2 : HA HA- HA- HA- HA-Klf4 Relative mrna expression Relative protein level of Relative protein level of Relative protein level
:Flag Relative protein level of Relative protein level of : Flag Vec 21LV 21SV 2 : HA HA- HA- HA- HA-Klf4 Relative mrna expression Relative protein level of Relative protein level of Relative protein level
(a) Immunoblotting to show the migration position of Flag-tagged MAVS
 Supplementary Figure 1 Characterization of six MAVS isoforms. (a) Immunoblotting to show the migration position of Flag-tagged MAVS isoforms. HEK293T Mavs -/- cells were transfected with constructs expressing
Supplementary Figure 1 Characterization of six MAVS isoforms. (a) Immunoblotting to show the migration position of Flag-tagged MAVS isoforms. HEK293T Mavs -/- cells were transfected with constructs expressing
Supplementary Information
 Supplementary Information Supplementary Figures Supplementary Figure 1. MLK1-4 phosphorylate MEK in the presence of RAF inhibitors. (a) H157 cells were transiently transfected with Flag- or HA-tagged MLK1-4
Supplementary Information Supplementary Figures Supplementary Figure 1. MLK1-4 phosphorylate MEK in the presence of RAF inhibitors. (a) H157 cells were transiently transfected with Flag- or HA-tagged MLK1-4
Supplemental Information. Myeloid-Derived Suppressor Cells Are. Controlled by Regulatory T Cells via TGF-b. during Murine Colitis
 Cell Reports, Volume 17 Supplemental Information Myeloid-Derived Suppressor Cells Are Controlled by Regulatory T Cells via TGF-b during Murine Colitis Cho-Rong Lee, Yewon Kwak, Taewoo Yang, Jung Hyun Han,
Cell Reports, Volume 17 Supplemental Information Myeloid-Derived Suppressor Cells Are Controlled by Regulatory T Cells via TGF-b during Murine Colitis Cho-Rong Lee, Yewon Kwak, Taewoo Yang, Jung Hyun Han,
Figure 1: TDP-43 is subject to lysine acetylation within the RNA-binding domain a) QBI-293 cells were transfected with TDP-43 in the presence or
 Figure 1: TDP-43 is subject to lysine acetylation within the RNA-binding domain a) QBI-293 cells were transfected with TDP-43 in the presence or absence of the acetyltransferase CBP and acetylated TDP-43
Figure 1: TDP-43 is subject to lysine acetylation within the RNA-binding domain a) QBI-293 cells were transfected with TDP-43 in the presence or absence of the acetyltransferase CBP and acetylated TDP-43
Figure S1- Targeting strategy for generating SIK1-T182A, SIK2-T175A and SIK3-T163A KI mice.
 Figure S1- Targeting strategy for generating SIK1-T12A, SIK2-T175A and SIK3-T163A mice. Left panel represents a depiction of the genomic loci of each gene, the targeting strategy and the final recombined
Figure S1- Targeting strategy for generating SIK1-T12A, SIK2-T175A and SIK3-T163A mice. Left panel represents a depiction of the genomic loci of each gene, the targeting strategy and the final recombined
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb3206 Supplementary Figure 1 Autophagy-related gene expression in murine and human intestinal tumors. (a) Representative LC3 immunostaining in colonic adenoma and adjacent non tumoral tissue
DOI: 10.1038/ncb3206 Supplementary Figure 1 Autophagy-related gene expression in murine and human intestinal tumors. (a) Representative LC3 immunostaining in colonic adenoma and adjacent non tumoral tissue
Supplemental Information. The TRAIL-Induced Cancer Secretome. Promotes a Tumor-Supportive Immune. Microenvironment via CCR2
 Molecular Cell, Volume 65 Supplemental Information The TRAIL-Induced Cancer Secretome Promotes a Tumor-Supportive Immune Microenvironment via CCR2 Torsten Hartwig, Antonella Montinaro, Silvia von Karstedt,
Molecular Cell, Volume 65 Supplemental Information The TRAIL-Induced Cancer Secretome Promotes a Tumor-Supportive Immune Microenvironment via CCR2 Torsten Hartwig, Antonella Montinaro, Silvia von Karstedt,
Supplementary figures
 Relative intensity Relative intensity Relative intensity Supplementary figures a None Caffeine None Caffeine c None Caffeine 6 6 6 ISG 6 6 6 UBE1L 6 6 6 UBCH8 6 6 6 EFP 1 1 DOX (h) 1 1 CPT (h) 1 1 UV (h)
Relative intensity Relative intensity Relative intensity Supplementary figures a None Caffeine None Caffeine c None Caffeine 6 6 6 ISG 6 6 6 UBE1L 6 6 6 UBCH8 6 6 6 EFP 1 1 DOX (h) 1 1 CPT (h) 1 1 UV (h)
supplementary information
 DOI: 1.138/ncb1839 a b Control 1 2 3 Control 1 2 3 Fbw7 Smad3 1 2 3 4 1 2 3 4 c d IGF-1 IGF-1Rβ IGF-1Rβ-P Control / 1 2 3 4 Real-time RT-PCR Relative quantity (IGF-1/ mrna) 2 1 IGF-1 1 2 3 4 Control /
DOI: 1.138/ncb1839 a b Control 1 2 3 Control 1 2 3 Fbw7 Smad3 1 2 3 4 1 2 3 4 c d IGF-1 IGF-1Rβ IGF-1Rβ-P Control / 1 2 3 4 Real-time RT-PCR Relative quantity (IGF-1/ mrna) 2 1 IGF-1 1 2 3 4 Control /
Supplementary Figure Legends
 Supplementary Figure Legends Figure S1 gene targeting strategy for disruption of chicken gene, related to Figure 1 (f)-(i). (a) The locus and the targeting constructs showing HpaI restriction sites. The
Supplementary Figure Legends Figure S1 gene targeting strategy for disruption of chicken gene, related to Figure 1 (f)-(i). (a) The locus and the targeting constructs showing HpaI restriction sites. The
Supplementary Information
 Supplementary Information Supplementary Figure 1: Over-expression of CD300f in NIH3T3 cells enhances their capacity to phagocytize AC. (a) NIH3T3 cells were stably transduced by EV, CD300f WT or CD300f
Supplementary Information Supplementary Figure 1: Over-expression of CD300f in NIH3T3 cells enhances their capacity to phagocytize AC. (a) NIH3T3 cells were stably transduced by EV, CD300f WT or CD300f
