Paper DP04 Statistical Analysis of Gene Expression Micro Arrays John Schwarz, University of Louisville, Louisville, KY
|
|
|
- Milo Shields
- 6 years ago
- Views:
Transcription
1 Paper DP04 Statistical Analysis of Gene Expression Micro Arrays John Schwarz, University of Louisville, Louisville, KY ABSTRACT Advancements in genetic research have led to increased amounts of data often without efficient analysis techniques. One such area of genetics that has developed a great deal in the past several years has been micro arrays. One area of micro array experimentation is that of gene expression. Gene expression uses arrays of thousands of genes with one to two targeted strands of DNA that are fluorescently tagged and used to identify which genes are expressed. This experiment can identify which conditions cause certain genes to be activated in different cells. It can also be used to track certain cellular changes through the expression of genes. The data gathered from the micro array experimentation is contained in a large matrix containing the data results from gene expression analysis on a micro array plate. Some of the standard analysis techniques and programs can be inefficient and inconclusive. The purpose of this analysis is to arrive at conclusive results with efficient methods. The primary statistical software used in the analysis was SAS/Stat system version 8.2. A series of statistical tests was applied to the data set to determine the meaningfulness of the results and the efficiency of the tests. INTRODUCTION Every living organism is made up of a single cell or many groups of cells. The identity of these different cells and the function of these cells are determined by genes. Genes are segments of DNA providing the code for producing proteins. Different organisms contain different numbers of genes. For example, a human being contains 30,000 genes as estimated and the fruit fly contains only about 13,000 genes. The identification of genes depends upon the knowledge of the DNA sequence made up of different alleles (different forms of a gene). The human genome has recently been fully mapped. However, not every gene has been fully expressed in the DNA alleles. Certain organisms have been fully mapped; this refers mainly to smaller organisms with a fewer number of different genes. The advantages of having the knowledge of the full sequenced genes will be discussed later with micro arrays. Understanding genes is important; unfortunately, genes can often be hidden within compacted DNA strands and are not easily identifiable. Genes also interact in different ways; some have similar properties and responsibilities while other genes may be totally different. Genes that are different, however, may be involved in the same reactions inside a cell and, equivalently, genes that have similar properties may not be involved in the same reactions. Micro arrays offer an efficient method of comparing multiple genes quickly and easily. Micro arrays, however, require several analyses upon completion of an experiment. The analysis of micro arrays offers substantial evidence of genes that may or may not be related in a cell. BACKGROUND Gene expression is important in cellular identification and gene function. With new technologies and research, gene expression and identification have become an ever growing area in biotechnologies with the opportunity for new, more efficient analyses available. The field of cellular genetics has shown that changing ph and temperature causes certain genes to be expressed and not expressed. It is possible to alter these settings in a lab and the expressed genes can be identified. Most genes are known by the proteins they produce and the function of these proteins. It is possible to analyze large groups of proteins as well as genes. This process will be discussed later. Analyzing different genes expressed can determine when certain reactions take place in the body, or determine what processes some of the genes are responsible for. Understanding which genes are expressed is important to different gene interactions and cellular identities. According to Campbell, In all organisms, the expression of specific genesis are most commonly regulated at the level of transcription by DNA-binding proteins that also interact with other proteins and often with external signals. For that reason, the term gene expression is often equated with gene activity- that is, transcription- for both prokaryotes (cells lacking membrane enclosed nucleus and membrane enclosed organelles) and eukaryotes (cell with membrane enclosed nucleus and membrane enclosed organelles). However, the greater complexity of eukaryotic cell structure and function provides opportunities for controlling gene expression at additional stages (Campbell 2002, p. 362).
2 Gene expression can be broken down into several different stages and identified at any time during one of the specific stages. These range from gene to functional protein activity. The first stage in expression is the unpacking of chromatin (complex of DNA and proteins that makes up a chromosome) DNA (Figure 1). In chromatin form, DNA is closely packed and regulatory proteins used in the transcription process often are unable to access certain portions of the DNA, and consequently certain genes are not expressed. The differences in chromatin packing vary in each different cell type and thus, some cells inhibit or allow the expression of certain genes. The chromatin packing is known as regulatory function for gene expression. Figure 1- Cellular Structure (Brazma 2003, p. 1) Even though entire DNA strands exist in chromatin form at times, they can be unpacked to allow access to certain strands for protein production and replication. Methylation occurs, meaning the process when the DNA is unpacked and methyl groups (CH 3) groups are placed on the ends of the DNA strand for the gene being used, and identifies the start and finish of the gene to replication compounds used for protein construction. Transcription occurs and the DNA is copied into RNA to change certain alleles and is then copied to mrna for protein production outside the nucleolus. The mrna is moved to the cytoplasm where protein translation is accomplished. Polypeptide (polymer chain in which amino acids are linked together with the peptide bonds) groups are created from the mrna code and, after cleaving and modification, are known as proteins. From this stage the proteins are sent to certain areas of the cell for purposes identified by the type of protein. The mrna strand breaks down after replicating. Usually several proteins in the cytoplasm and the allele groups are absorbed and recycled for later protein production. This whole process is known as the gene expression, and again any stage of this process can be used to identify the gene expression (Campbell 2002, p. 364). Cellular identification is determined by the genes expressed and the proteins produced. Different cells have different roles in the body and in turn produce a number of different proteins. By learning which conditions activate certain genes and deactivate other genes, more can be understood about certain cellular identities. Furthermore, much can be understood about cellular life and death and whether death occurs as a consequence of time or as a malfunction (Campbell 2002). Cells generally have the ability to regenerate and reproduce themselves. These processes are marked with the expression of certain genes. The understanding of when and how these processes take place, and more importantly, why some cells do not go through this process can be better understood through gene expression analysis. The main objective in micro array analysis is to identify gene expression and the comparison of several genes at once. The process of identifying and comparing the genes expressed in a cell or culture is a complex process resulting in large amounts of data. The process can be simplified into six main steps (See Figure 2): Selecting the cell culture for analysis Identifying the specific DNA gene sequence Radioactively tagging the DNA sequences Hybridization of the array (Hybridization is the process in which the fluorescently tagged cdna is applied to the array) Laser intensity readings from the plate Interpreting the results of the hybridized array Of course, only fully sequenced genes can be used in this experimentation since known DNA sequences are hybridized to an array of thousands of different genes at one time. The idea behind creating this array is to identify genes at certain points in an expression and isolate conditions for the certain genes to be expressed. There are several different types of micro array experimentations. The first type is gene expressions using DNA. This method uses the DNA sequences of specific genes that are applied to an arrayed, cultured plate with the goal of identifying genes in the cells and comparing their interactions. Initially, the mrna is used in this experiment. The
3 mrna, messenger RNA (synthesized DNA), is used in the production of proteins. The mrna is copied directly from DNA in the cell. When the mrna is copied, different gene alleles are used that differ from normal DNA. The mrna sequences are known to be unstable and to deteriorate after a short amount of time, making them useless for hybridization reactions. For this reason cdna (DNA copied from mrna with enzyme reverse transcriptase) is copied from the mrna for its ease of use and compatibility (Fortina 2000). Figure 2- Microarray process (Buhler 2003, p. 1) The cdna, complementary DNA, is copied from the mrna and uses the original alleles as DNA. The cdna is then tagged with fluorescent markers and applied to an array of cellular cultures. Another type of micro array experimentation uses DNA sequences. A DNA sequence, again fluorescently tagged, can be used so that the same sequence can be identified in different cells across an array. This identification can be used to understand evolutionary traits across species. The third process uses proteins. Proteins are tagged fluorescently and applied to a slide consisting of an environment with protein-carrying structures. This method can be used to determine the relationships of many proteins at once and to show how they interact with one another in specific areas of the body. Protein analysis can help to gain an understanding for which proteins have certain relationships or even tracing it back to certain genes that produce the proteins. The process of the creation of an array plate generates thousands of closely contained DNA segments. The DNA segments are placed within a small region on a glass slide and glued to prevent washing away during the gene reaction process (Campbell 2002). The DNA segments are placed in a grid pattern for identification once the hybridization reaction has taken place. The use of certain genes depends upon what is known about the organism. Some organisms have fully sequenced genes so that all the genes can be used in the expression. Organisms that have a great number of genes may require the use of several array plates since not all the genes can fit on a single array. Continuing with the other steps involved in micro array, the analysis will be described using the techniques of gene expression micro arrays. Hybridization is the next step to micro array. In addition, realization of such developments will facilitate early diagnosis and evaluation of treatment strategies. The assay is template-dependent and involves primer extension by a radioactive- or dye-labeled dideoxynucleotide terminator (ddntp) with the tag of the incorporated base revealing the identity of the template complementary nucleotide imme-diately 3 to the primer. Solution-phase extensions followed by electrophoresis on gels as well as microtiter plate-based assays have been described (Fortina 2000, p. 884). Each spot on the array contains more than one sample strand of DNA. The ability for more than one interaction at each spot is possible. Thus the intensity of the fluorescent marking can differ from spot to spot and cannot be used to determine how accurately the strength of the binding is to the sequence. The different sequences are marked with different colored fluorescent tags. Using different colors allows for multiple detection of gene expression (Figure 3). The different colors can also determine similar genes and gene interactions. Figure 3- Microarray plate(leming 2003, p. 1) Following the hybridization process, a washing stage is carried out. The purpose of the washing is to eliminate any fluorescently labeled sequences that did not hybridize to gene spots. That is why it is important to secure the spots before the hybridization process. This security leads to a clearer interpretation of which gene spots hybridize with the applied marked gene sequences.
4 The fluorescent tags can be identified by computer analysis. Since hybridization can occur on only a single sequence of DNA in each spot, visual detection cannot be accurate. Furthermore, the fluorescent tags used in the hybridization process cannot be visually identified unaided. The tags used require certain frequencies provided by laser treatment after hybridization and washing has occurred. Consequently, there are thousands and thousands of data results from each micro array hybridization experiment. Simple visual analysis in most cases is time consuming and inefficient. Analysis by computer is the most efficient and accurate option. For this reason, different statistical methods must be employed to understand the relation between different genes. Several different programs exist currently designed to analyze the large sets of data produced by the micro array process. Some of these include the simple frequency counts from each of the different experiments and genes. Other methods include simple statistical tests to understand small portions of the large array. METHODS The first step in the analysis process was to examine the properties of each gene that reacted with the micro array plate. The method used for modeling the data was kernel density estimation. Two-dimensional graphical representations of kernel density estimators resemble a smoothed histogram. The kernel density estimator approximates a probability density function allowing for specific values to be accentuated. The kernel density estimator uses all the values in the data. This differs from the histogram in that data are separated into certain sections and allow for the maximum value to be emphasized, not just the section in which this maximum occurs. When dealing with such large data sets for visual analysis, certain portions of the data are shortened by bound adjustments. This allows for the most prominent portions of the data to be observed clearly. The equation used for kernel density estimation is as follows: The value n is the sample size for the dataset, a n is a constant based on the sample size. Inside the summation, k is a known density function, and x is any value within the domain of the density function f. The kernel density can be analyzed visually to allow for preliminary relationships to be determined. Since this process is visual, optimal graphical representations must be achieved. Optimal graphical representation is done through bandwidth adjustment. Several different methods for bandwidth adjustment are available through SAS, and each method produces different results. For determining the best bandwidth adjustment, the different methods are tested individually at different percentage levels. Using SAS and kernel density estimation, the KDE procedure returns output in table form. The table lists 401 rows of the density function, data position, and count. From the table form, the kernel density model can be plotted and then visually examined. The specified SAS code is as follows: Proc kde data=_proj_.john gridl=0 gridu=954 method=srot bwm=1.00 out=outkde1; Var CJ_BaP1; Run; The first section of the code designates procedure. Next, the lower and upper bounds are declared for the data. Then the method of bandwidth adjustments is specified. The second line begins with the percentage for the bandwidth adjustment and finishes with the location of the output file where the analysis will be stored. The third line declares the variable that is analyzed. For this data set, the method of bandwidth was Silvermans Rule of Thumb Method. This method is modeled by the following equation: (SAS Institute 2003, p. 1) In this equation, n is the sample size and σ is the sample standard deviation. The Q values represent the first and third quartiles. This method proved visually optimal for the data set at 100%. The 100% bandwidth gives a smooth representation of the data which with multiple graphs allows for a simple distinct comparison. The lower bound was set at zero and the upper bound was determined based on the analysis of each individual variable. Each upper bound had the same density endpoint value.
5 RESULTS The resulting kernel density analysis produced a series of tables with three columns and four hundred rows plus one. For graphical analysis, the first two columns, value and density, were used to plot solutions. The graphs from each different variable could then be combined and simple visual interpretations could be made. Each single variable plot was similar in pattern but maximum regions often differed from variable to variable. Figure 4 is an example of a plotted graph using the variable of CJ_BaP1. Figure 4-Gene CJ_BaP1 Kernel Density Plot There were several different methods involving the comparison of the separate graphs. The first was to plot all of the variables on one graph. The resulting plot is illustrated in figure 5. The y axis represents the probability density and the x axis defines the frequency of the light recorded. The boxes indicate the different concentrations of genes. Figure 5-Comparison of all genes using kernel density. Upon visual analysis, it can be determined that there are two major concentrations of variables that are the different genes run through the experiment, and several outliers. The upper concentration is made up of 26 gene variables. The lower concentration is made up of 14 variables and the remaining 2 variables look to be outliers. From this point, several different comparisons were made. The concentrations represent possible groups of related genes. First each concentration was examined more thoroughly by removing the other variables from the plot. This was repeated several times, and smaller concentrations throughout the group could be examined individually. An example of a smaller concentration is illustrated in figure 6 and the clusters within are identified by the boxes on the
6 plot. Figure 6-Upper concentration of genes analyzed. Similarly the analysis of the concentration was repeated for the lower concentration and again smaller clusters could be identified and interpreted. An example from the lower concentration is illustrated here figure 7 and the clusters identified by the boxes.. Figure 7-Lower concentration of genes analyzed. Next, an examination of the difference in the concentrations and outliers was undertaken. Several variables were selected from each cluster nearly at random, and plotted to observed differences in position and behavior. Plotting variables against one another helped to distinguish outliers and cluster groups. Figure 8 is an example of this analysis.
7 Figure 8- Example of genes that differ in expression. The difference in concentrations was the initial step to understand gene relationships and which genes could possibly be associated with one another. Using this information, the individual clusters could help identify more exactly which genes had a higher probability of being related to one another. This is demonstrated in figure 9. Figure 9-Example of genes that are similar in expression. CONCLUSION The visual comparison has allowed for a greater understanding of the relations between the genes. As discussed before in the results section, there were two major concentrations of the gene variables when analyzed by SAS kernel density estimator (Fig 2). The larger concentration contained 26 different genes and when examined again was shown to contain 2 smaller clusters within the concentration (Fig. 3). The larger of the clusters contained 14 separate gene variables and the smaller cluster contained 12 different gene variables. These results suggest that the groups of genes contained in each cluster have relations within the cell and may have similar expression properties. The smaller concentration that contained 14 from the original gene variables also contained two smaller clusters (Fig 4). Each of these smaller clusters contained 7 gene variables each. Again, these results suggest that the groups of genes contained in each cluster have a chance of being related and share similar expressions. The two gene variables that appear to be outliers were not clustered together; thus they appear to have no relation with any other gene run in the experiment. The kernel density estimation allowed for easier visual interpretation of the larger data set. By using this method, comparing multiple variables at once becomes much easier and possible. This also has allowed for easier presentation of the results. The bandwidth gave the ability to have a clear presentation and easier interpretation. Further analysis of the data using SAS could determine exact relationships and even confidence intervals within the different clusters and groups of the data. More advanced techniques such as mixed models could also be explored
8 as options to analyze the data set and identify relations between different genes and their expression within the cell. REFERENCES Brazma, Alvis; Parkinson, Helen; Schlitt, Thomas; Shojatalab, Mohammadreza. Quick Introduction to Elements of Biology (December 2003). Buhler, Jeremy. Anatomy of a Comparative Gene Expression Study (December 2003). Campbell, Neil A.; Reece, Jane B. Biology. Pearson Education, San Francisco, Ca Fortina, Paolo; Delgrosso, Kathleen; Sakazume, Taku; Santacroce, Rosa; Moutereau, Stephane; Su, Hung-Ju; Graves, David; McKenzie, Steven; Surrey, Saul. Simple two-color array-based approach for mutation detection European Journal of Human Genetics; Nov2000, Vol. 8 Issue 11, p884, 11p. Leming Shi, PHD. DNA Microarray. (March 2004). SAS Institute Inc SAS Online Doc, Version 8.2, SAS Institute Inc., Cary, North Carolina, USA ACKNOWLEDGMENTS I acknowledge the help from my advisor Dr. Patricia Cerrito. CONTACT INFORMATION Your comments and questions are valued and encouraged. Contact the author at: John Schwarz University of Louisville 1416 Sylvan Way Louisville, KY Work Phone: jcschw02@louisville.edu SAS and all other SAS Institute Inc. product or service names are registered trademarks or trademarks of SAS Institute Inc. in the USA and other countries. fi indicates USA registration. Other brand and product names are trademarks of their respective companies.
3.1.4 DNA Microarray Technology
 3.1.4 DNA Microarray Technology Scientists have discovered that one of the differences between healthy and cancer is which genes are turned on in each. Scientists can compare the gene expression patterns
3.1.4 DNA Microarray Technology Scientists have discovered that one of the differences between healthy and cancer is which genes are turned on in each. Scientists can compare the gene expression patterns
Molecular Cell Biology - Problem Drill 11: Recombinant DNA
 Molecular Cell Biology - Problem Drill 11: Recombinant DNA Question No. 1 of 10 1. Which of the following statements about the sources of DNA used for molecular cloning is correct? Question #1 (A) cdna
Molecular Cell Biology - Problem Drill 11: Recombinant DNA Question No. 1 of 10 1. Which of the following statements about the sources of DNA used for molecular cloning is correct? Question #1 (A) cdna
Bio 101 Sample questions: Chapter 10
 Bio 101 Sample questions: Chapter 10 1. Which of the following is NOT needed for DNA replication? A. nucleotides B. ribosomes C. Enzymes (like polymerases) D. DNA E. all of the above are needed 2 The information
Bio 101 Sample questions: Chapter 10 1. Which of the following is NOT needed for DNA replication? A. nucleotides B. ribosomes C. Enzymes (like polymerases) D. DNA E. all of the above are needed 2 The information
The Polymerase Chain Reaction. Chapter 6: Background
 The Polymerase Chain Reaction Chapter 6: Background Invention of PCR Kary Mullis Mile marker 46.58 in April of 1983 Pulled off the road and outlined a way to conduct DNA replication in a tube Worked for
The Polymerase Chain Reaction Chapter 6: Background Invention of PCR Kary Mullis Mile marker 46.58 in April of 1983 Pulled off the road and outlined a way to conduct DNA replication in a tube Worked for
Make the protein through the genetic dogma process.
 Make the protein through the genetic dogma process. Coding Strand 5 AGCAATCATGGATTGGGTACATTTGTAACTGT 3 Template Strand mrna Protein Complete the table. DNA strand DNA s strand G mrna A C U G T A T Amino
Make the protein through the genetic dogma process. Coding Strand 5 AGCAATCATGGATTGGGTACATTTGTAACTGT 3 Template Strand mrna Protein Complete the table. DNA strand DNA s strand G mrna A C U G T A T Amino
Feature Selection of Gene Expression Data for Cancer Classification: A Review
 Available online at www.sciencedirect.com ScienceDirect Procedia Computer Science 50 (2015 ) 52 57 2nd International Symposium on Big Data and Cloud Computing (ISBCC 15) Feature Selection of Gene Expression
Available online at www.sciencedirect.com ScienceDirect Procedia Computer Science 50 (2015 ) 52 57 2nd International Symposium on Big Data and Cloud Computing (ISBCC 15) Feature Selection of Gene Expression
Year III Pharm.D Dr. V. Chitra
 Year III Pharm.D Dr. V. Chitra 1 Genome entire genetic material of an individual Transcriptome set of transcribed sequences Proteome set of proteins encoded by the genome 2 Only one strand of DNA serves
Year III Pharm.D Dr. V. Chitra 1 Genome entire genetic material of an individual Transcriptome set of transcribed sequences Proteome set of proteins encoded by the genome 2 Only one strand of DNA serves
Gene Expression Transcription/Translation Protein Synthesis
 Gene Expression Transcription/Translation Protein Synthesis 1. Describe how genetic information is transcribed into sequences of bases in RNA molecules and is finally translated into sequences of amino
Gene Expression Transcription/Translation Protein Synthesis 1. Describe how genetic information is transcribed into sequences of bases in RNA molecules and is finally translated into sequences of amino
Recent technology allow production of microarrays composed of 70-mers (essentially a hybrid of the two techniques)
 Microarrays and Transcript Profiling Gene expression patterns are traditionally studied using Northern blots (DNA-RNA hybridization assays). This approach involves separation of total or polya + RNA on
Microarrays and Transcript Profiling Gene expression patterns are traditionally studied using Northern blots (DNA-RNA hybridization assays). This approach involves separation of total or polya + RNA on
Methods of Biomaterials Testing Lesson 3-5. Biochemical Methods - Molecular Biology -
 Methods of Biomaterials Testing Lesson 3-5 Biochemical Methods - Molecular Biology - Chromosomes in the Cell Nucleus DNA in the Chromosome Deoxyribonucleic Acid (DNA) DNA has double-helix structure The
Methods of Biomaterials Testing Lesson 3-5 Biochemical Methods - Molecular Biology - Chromosomes in the Cell Nucleus DNA in the Chromosome Deoxyribonucleic Acid (DNA) DNA has double-helix structure The
Controlling DNA. Ethical guidelines for the use of DNA technology. Module Type: Discussion, literature review, and debate
 Ethical guidelines for the use of DNA technology Author: Tara Cornelisse, Ph.D. Candidate, Environmental Studies, University of California Santa Cruz. Field-tested with: 11 th -12 th grade students in
Ethical guidelines for the use of DNA technology Author: Tara Cornelisse, Ph.D. Candidate, Environmental Studies, University of California Santa Cruz. Field-tested with: 11 th -12 th grade students in
Non-Organic-Based Isolation of Mammalian microrna using Norgen s microrna Purification Kit
 Application Note 13 RNA Sample Preparation Non-Organic-Based Isolation of Mammalian microrna using Norgen s microrna Purification Kit B. Lam, PhD 1, P. Roberts, MSc 1 Y. Haj-Ahmad, M.Sc., Ph.D 1,2 1 Norgen
Application Note 13 RNA Sample Preparation Non-Organic-Based Isolation of Mammalian microrna using Norgen s microrna Purification Kit B. Lam, PhD 1, P. Roberts, MSc 1 Y. Haj-Ahmad, M.Sc., Ph.D 1,2 1 Norgen
Multiple choice questions (numbers in brackets indicate the number of correct answers)
 1 Multiple choice questions (numbers in brackets indicate the number of correct answers) February 1, 2013 1. Ribose is found in Nucleic acids Proteins Lipids RNA DNA (2) 2. Most RNA in cells is transfer
1 Multiple choice questions (numbers in brackets indicate the number of correct answers) February 1, 2013 1. Ribose is found in Nucleic acids Proteins Lipids RNA DNA (2) 2. Most RNA in cells is transfer
AP Biology Gene Expression/Biotechnology REVIEW
 AP Biology Gene Expression/Biotechnology REVIEW Multiple Choice Identify the choice that best completes the statement or answers the question. 1. Gene expression can be a. regulated before transcription.
AP Biology Gene Expression/Biotechnology REVIEW Multiple Choice Identify the choice that best completes the statement or answers the question. 1. Gene expression can be a. regulated before transcription.
Pre-Lab: Molecular Biology
 Pre-Lab: Molecular Biology Name 1. What are the three chemical parts of a nucleotide. Draw a simple sketch to show how the three parts are arranged. 2. What are the rules of base pairing? 3. In double
Pre-Lab: Molecular Biology Name 1. What are the three chemical parts of a nucleotide. Draw a simple sketch to show how the three parts are arranged. 2. What are the rules of base pairing? 3. In double
Lecture Four. Molecular Approaches I: Nucleic Acids
 Lecture Four. Molecular Approaches I: Nucleic Acids I. Recombinant DNA and Gene Cloning Recombinant DNA is DNA that has been created artificially. DNA from two or more sources is incorporated into a single
Lecture Four. Molecular Approaches I: Nucleic Acids I. Recombinant DNA and Gene Cloning Recombinant DNA is DNA that has been created artificially. DNA from two or more sources is incorporated into a single
CHAPTER 20 DNA TECHNOLOGY AND GENOMICS. Section A: DNA Cloning
 Section A: DNA Cloning 1. DNA technology makes it possible to clone genes for basic research and commercial applications: an overview 2. Restriction enzymes are used to make recombinant DNA 3. Genes can
Section A: DNA Cloning 1. DNA technology makes it possible to clone genes for basic research and commercial applications: an overview 2. Restriction enzymes are used to make recombinant DNA 3. Genes can
Fig Ch 17: From Gene to Protein
 Fig. 17-1 Ch 17: From Gene to Protein Basic Principles of Transcription and Translation RNA is the intermediate between genes and the proteins for which they code Transcription is the synthesis of RNA
Fig. 17-1 Ch 17: From Gene to Protein Basic Principles of Transcription and Translation RNA is the intermediate between genes and the proteins for which they code Transcription is the synthesis of RNA
Chapter 13. From DNA to Protein
 Chapter 13 From DNA to Protein Proteins All proteins consist of polypeptide chains A linear sequence of amino acids Each chain corresponds to the nucleotide base sequenceof a gene The Path From Genes to
Chapter 13 From DNA to Protein Proteins All proteins consist of polypeptide chains A linear sequence of amino acids Each chain corresponds to the nucleotide base sequenceof a gene The Path From Genes to
CHAPTER 17 FROM GENE TO PROTEIN. Section C: The Synthesis of Protein
 CHAPTER 17 FROM GENE TO PROTEIN Section C: The Synthesis of Protein 1. Translation is the RNA-directed synthesis of a polypeptide: a closer look 2. Signal peptides target some eukaryotic polypeptides to
CHAPTER 17 FROM GENE TO PROTEIN Section C: The Synthesis of Protein 1. Translation is the RNA-directed synthesis of a polypeptide: a closer look 2. Signal peptides target some eukaryotic polypeptides to
Transcription. Unit: DNA. Central Dogma. 2. Transcription converts DNA into RNA. What is a gene? What is transcription? 1/7/2016
 Warm Up Questions 1. Where is DNA located? 2. Name the 3 parts of a nucleotide. 3. Enzymes can catalyze many different reactions (T or F) 4. How many variables should you have in an experiment? 5. A red
Warm Up Questions 1. Where is DNA located? 2. Name the 3 parts of a nucleotide. 3. Enzymes can catalyze many different reactions (T or F) 4. How many variables should you have in an experiment? 5. A red
Gene Expression Technology
 Gene Expression Technology Bing Zhang Department of Biomedical Informatics Vanderbilt University bing.zhang@vanderbilt.edu Gene expression Gene expression is the process by which information from a gene
Gene Expression Technology Bing Zhang Department of Biomedical Informatics Vanderbilt University bing.zhang@vanderbilt.edu Gene expression Gene expression is the process by which information from a gene
Gene Expression and Heritable Phenotype. CBS520 Eric Nabity
 Gene Expression and Heritable Phenotype CBS520 Eric Nabity DNA is Just the Beginning DNA was determined to be the genetic material, and the structure was identified as a (double stranded) double helix.
Gene Expression and Heritable Phenotype CBS520 Eric Nabity DNA is Just the Beginning DNA was determined to be the genetic material, and the structure was identified as a (double stranded) double helix.
Bio11 Announcements. Ch 21: DNA Biology and Technology. DNA Functions. DNA and RNA Structure. How do DNA and RNA differ? What are genes?
 Bio11 Announcements TODAY Genetics (review) and quiz (CP #4) Structure and function of DNA Extra credit due today Next week in lab: Case study presentations Following week: Lab Quiz 2 Ch 21: DNA Biology
Bio11 Announcements TODAY Genetics (review) and quiz (CP #4) Structure and function of DNA Extra credit due today Next week in lab: Case study presentations Following week: Lab Quiz 2 Ch 21: DNA Biology
DNA and RNA. Chapter 12
 DNA and RNA Chapter 12 History of DNA Late 1800 s scientists discovered that DNA is in the nucleus of the cell 1902 Walter Sutton proposed that hereditary material resided in the chromosomes in the nucleus
DNA and RNA Chapter 12 History of DNA Late 1800 s scientists discovered that DNA is in the nucleus of the cell 1902 Walter Sutton proposed that hereditary material resided in the chromosomes in the nucleus
Division Ave. High School AP Biology
 Control of Eukaryotic Genes 2007-2008 The BIG Questions n How are genes turned on & off in eukaryotes? n How do cells with the same genes differentiate to perform completely different, specialized functions?
Control of Eukaryotic Genes 2007-2008 The BIG Questions n How are genes turned on & off in eukaryotes? n How do cells with the same genes differentiate to perform completely different, specialized functions?
TRANSCRIPTION AND PROCESSING OF RNA
 TRANSCRIPTION AND PROCESSING OF RNA 1. The steps of gene expression. 2. General characterization of transcription: steps, components of transcription apparatus. 3. Transcription of eukaryotic structural
TRANSCRIPTION AND PROCESSING OF RNA 1. The steps of gene expression. 2. General characterization of transcription: steps, components of transcription apparatus. 3. Transcription of eukaryotic structural
Review of Protein (one or more polypeptide) A polypeptide is a long chain of..
 Gene expression Review of Protein (one or more polypeptide) A polypeptide is a long chain of.. In a protein, the sequence of amino acid determines its which determines the protein s A protein with an enzymatic
Gene expression Review of Protein (one or more polypeptide) A polypeptide is a long chain of.. In a protein, the sequence of amino acid determines its which determines the protein s A protein with an enzymatic
DNA replication: Enzymes link the aligned nucleotides by phosphodiester bonds to form a continuous strand.
 DNA replication: Copying genetic information for transmission to the next generation Occurs in S phase of cell cycle Process of DNA duplicating itself Begins with the unwinding of the double helix to expose
DNA replication: Copying genetic information for transmission to the next generation Occurs in S phase of cell cycle Process of DNA duplicating itself Begins with the unwinding of the double helix to expose
Recombinant DNA Technology. The Role of Recombinant DNA Technology in Biotechnology. yeast. Biotechnology. Recombinant DNA technology.
 PowerPoint Lecture Presentations prepared by Mindy Miller-Kittrell, North Carolina State University C H A P T E R 8 Recombinant DNA Technology The Role of Recombinant DNA Technology in Biotechnology Biotechnology?
PowerPoint Lecture Presentations prepared by Mindy Miller-Kittrell, North Carolina State University C H A P T E R 8 Recombinant DNA Technology The Role of Recombinant DNA Technology in Biotechnology Biotechnology?
Introduction to BioMEMS & Medical Microdevices DNA Microarrays and Lab-on-a-Chip Methods
 Introduction to BioMEMS & Medical Microdevices DNA Microarrays and Lab-on-a-Chip Methods Companion lecture to the textbook: Fundamentals of BioMEMS and Medical Microdevices, by Prof., http://saliterman.umn.edu/
Introduction to BioMEMS & Medical Microdevices DNA Microarrays and Lab-on-a-Chip Methods Companion lecture to the textbook: Fundamentals of BioMEMS and Medical Microdevices, by Prof., http://saliterman.umn.edu/
DNA is the genetic material. DNA structure. Chapter 7: DNA Replication, Transcription & Translation; Mutations & Ames test
 DNA is the genetic material Chapter 7: DNA Replication, Transcription & Translation; Mutations & Ames test Dr. Amy Rogers Bio 139 General Microbiology Hereditary information is carried by DNA Griffith/Avery
DNA is the genetic material Chapter 7: DNA Replication, Transcription & Translation; Mutations & Ames test Dr. Amy Rogers Bio 139 General Microbiology Hereditary information is carried by DNA Griffith/Avery
RNA & PROTEIN SYNTHESIS
 RNA & PROTEIN SYNTHESIS DNA & RNA Genes are coded DNA instructions that control the production of proteins within the cell. The first step in decoding these genetic messages is to copy part of the nucleotide
RNA & PROTEIN SYNTHESIS DNA & RNA Genes are coded DNA instructions that control the production of proteins within the cell. The first step in decoding these genetic messages is to copy part of the nucleotide
Rapid Learning Center Presents. Teach Yourself AP Biology in 24 Hours
 Rapid Learning Center Chemistry :: Biology :: Physics :: Math Rapid Learning Center Presents Teach Yourself AP Biology in 24 Hours 1/35 *AP is a registered trademark of the College Board, which does not
Rapid Learning Center Chemistry :: Biology :: Physics :: Math Rapid Learning Center Presents Teach Yourself AP Biology in 24 Hours 1/35 *AP is a registered trademark of the College Board, which does not
Protein Synthesis & Gene Expression
 DNA provides the instructions for how to build proteins Each gene dictates how to build a single protein in prokaryotes The sequence of nucleotides (AGCT) in DNA dictates the order of amino acids that
DNA provides the instructions for how to build proteins Each gene dictates how to build a single protein in prokaryotes The sequence of nucleotides (AGCT) in DNA dictates the order of amino acids that
What is DNA??? DNA = Deoxyribonucleic acid IT is a molecule that contains the code for an organism s growth and function
 Review DNA and RNA 1) DNA and RNA are important organic compounds found in cells, called nucleic acids 2) Both DNA and RNA molecules contain the following chemical elements: carbon, hydrogen, oxygen, nitrogen
Review DNA and RNA 1) DNA and RNA are important organic compounds found in cells, called nucleic acids 2) Both DNA and RNA molecules contain the following chemical elements: carbon, hydrogen, oxygen, nitrogen
CHAPTER 21 LECTURE SLIDES
 CHAPTER 21 LECTURE SLIDES Prepared by Brenda Leady University of Toledo To run the animations you must be in Slideshow View. Use the buttons on the animation to play, pause, and turn audio/text on or off.
CHAPTER 21 LECTURE SLIDES Prepared by Brenda Leady University of Toledo To run the animations you must be in Slideshow View. Use the buttons on the animation to play, pause, and turn audio/text on or off.
DNA RNA PROTEIN SYNTHESIS -NOTES-
 DNA RNA PROTEIN SYNTHESIS -NOTES- THE COMPONENTS AND STRUCTURE OF DNA DNA is made up of units called nucleotides. Nucleotides are made up of three basic components:, called deoxyribose in DNA In DNA, there
DNA RNA PROTEIN SYNTHESIS -NOTES- THE COMPONENTS AND STRUCTURE OF DNA DNA is made up of units called nucleotides. Nucleotides are made up of three basic components:, called deoxyribose in DNA In DNA, there
DNA. translation. base pairing rules for DNA Replication. thymine. cytosine. amino acids. The building blocks of proteins are?
 2 strands, has the 5-carbon sugar deoxyribose, and has the nitrogen base Thymine. The actual process of assembling the proteins on the ribosome is called? DNA translation Adenine pairs with Thymine, Thymine
2 strands, has the 5-carbon sugar deoxyribose, and has the nitrogen base Thymine. The actual process of assembling the proteins on the ribosome is called? DNA translation Adenine pairs with Thymine, Thymine
Protein Synthesis
 HEBISD Student Expectations: Identify that RNA Is a nucleic acid with a single strand of nucleotides Contains the 5-carbon sugar ribose Contains the nitrogen bases A, G, C and U instead of T. The U is
HEBISD Student Expectations: Identify that RNA Is a nucleic acid with a single strand of nucleotides Contains the 5-carbon sugar ribose Contains the nitrogen bases A, G, C and U instead of T. The U is
3 Designing Primers for Site-Directed Mutagenesis
 3 Designing Primers for Site-Directed Mutagenesis 3.1 Learning Objectives During the next two labs you will learn the basics of site-directed mutagenesis: you will design primers for the mutants you designed
3 Designing Primers for Site-Directed Mutagenesis 3.1 Learning Objectives During the next two labs you will learn the basics of site-directed mutagenesis: you will design primers for the mutants you designed
DNA Transcription. Visualizing Transcription. The Transcription Process
 DNA Transcription By: Suzanne Clancy, Ph.D. 2008 Nature Education Citation: Clancy, S. (2008) DNA transcription. Nature Education 1(1) If DNA is a book, then how is it read? Learn more about the DNA transcription
DNA Transcription By: Suzanne Clancy, Ph.D. 2008 Nature Education Citation: Clancy, S. (2008) DNA transcription. Nature Education 1(1) If DNA is a book, then how is it read? Learn more about the DNA transcription
Introduction to Microarray Data Analysis and Gene Networks. Alvis Brazma European Bioinformatics Institute
 Introduction to Microarray Data Analysis and Gene Networks Alvis Brazma European Bioinformatics Institute A brief outline of this course What is gene expression, why it s important Microarrays and how
Introduction to Microarray Data Analysis and Gene Networks Alvis Brazma European Bioinformatics Institute A brief outline of this course What is gene expression, why it s important Microarrays and how
DNA Arrays Affymetrix GeneChip System
 DNA Arrays Affymetrix GeneChip System chip scanner Affymetrix Inc. hybridization Affymetrix Inc. data analysis Affymetrix Inc. mrna 5' 3' TGTGATGGTGGGAATTGGGTCAGAAGGACTGTGGGCGCTGCC... GGAATTGGGTCAGAAGGACTGTGGC
DNA Arrays Affymetrix GeneChip System chip scanner Affymetrix Inc. hybridization Affymetrix Inc. data analysis Affymetrix Inc. mrna 5' 3' TGTGATGGTGGGAATTGGGTCAGAAGGACTGTGGGCGCTGCC... GGAATTGGGTCAGAAGGACTGTGGC
Chromosomes. Chromosomes. Genes. Strands of DNA that contain all of the genes an organism needs to survive and reproduce
 Chromosomes Chromosomes Strands of DNA that contain all of the genes an organism needs to survive and reproduce Genes Segments of DNA that specify how to build a protein genes may specify more than one
Chromosomes Chromosomes Strands of DNA that contain all of the genes an organism needs to survive and reproduce Genes Segments of DNA that specify how to build a protein genes may specify more than one
From Gene to Protein. Chapter 17. Biology Eighth Edition Neil Campbell and Jane Reece. PowerPoint Lecture Presentations for
 Chapter 17 From Gene to Protein PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley with contributions from Joan Sharp
Chapter 17 From Gene to Protein PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley with contributions from Joan Sharp
BIOLOGY LTF DIAGNOSTIC TEST DNA to PROTEIN & BIOTECHNOLOGY
 Biology Multiple Choice 016074 BIOLOGY LTF DIAGNOSTIC TEST DNA to PROTEIN & BIOTECHNOLOGY Test Code: 016074 Directions: Each of the questions or incomplete statements below is followed by five suggested
Biology Multiple Choice 016074 BIOLOGY LTF DIAGNOSTIC TEST DNA to PROTEIN & BIOTECHNOLOGY Test Code: 016074 Directions: Each of the questions or incomplete statements below is followed by five suggested
Genetics Lecture 21 Recombinant DNA
 Genetics Lecture 21 Recombinant DNA Recombinant DNA In 1971, a paper published by Kathleen Danna and Daniel Nathans marked the beginning of the recombinant DNA era. The paper described the isolation of
Genetics Lecture 21 Recombinant DNA Recombinant DNA In 1971, a paper published by Kathleen Danna and Daniel Nathans marked the beginning of the recombinant DNA era. The paper described the isolation of
2054, Chap. 14, page 1
 2054, Chap. 14, page 1 I. Recombinant DNA technology (Chapter 14) A. recombinant DNA technology = collection of methods used to perform genetic engineering 1. genetic engineering = deliberate modification
2054, Chap. 14, page 1 I. Recombinant DNA technology (Chapter 14) A. recombinant DNA technology = collection of methods used to perform genetic engineering 1. genetic engineering = deliberate modification
Chapter 20: Biotechnology
 Name Period The AP Biology exam has reached into this chapter for essay questions on a regular basis over the past 15 years. Student responses show that biotechnology is a difficult topic. This chapter
Name Period The AP Biology exam has reached into this chapter for essay questions on a regular basis over the past 15 years. Student responses show that biotechnology is a difficult topic. This chapter
Chapter 13 - Concept Mapping
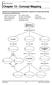 Chapter 13 - Concept Mapping Using the terms and phrases provided below, complete the concept map showing the discovery of DNA structure. amount of base pairs five-carbon sugar purine DNA polymerases Franklin
Chapter 13 - Concept Mapping Using the terms and phrases provided below, complete the concept map showing the discovery of DNA structure. amount of base pairs five-carbon sugar purine DNA polymerases Franklin
Chapter 3. DNA Replication & The Cell Cycle
 Chapter 3 DNA Replication & The Cell Cycle DNA Replication and the Cell Cycle Before cells divide, they must duplicate their DNA // the genetic material DNA is organized into strands called chromosomes
Chapter 3 DNA Replication & The Cell Cycle DNA Replication and the Cell Cycle Before cells divide, they must duplicate their DNA // the genetic material DNA is organized into strands called chromosomes
Transcription in Eukaryotes
 Transcription in Eukaryotes Biology I Hayder A Giha Transcription Transcription is a DNA-directed synthesis of RNA, which is the first step in gene expression. Gene expression, is transformation of the
Transcription in Eukaryotes Biology I Hayder A Giha Transcription Transcription is a DNA-directed synthesis of RNA, which is the first step in gene expression. Gene expression, is transformation of the
DNA Replication and Protein Synthesis
 DNA Replication and Protein Synthesis DNA is Deoxyribonucleic Acid. It holds all of our genetic information which is passed down through sexual reproduction DNA has three main functions: 1. DNA Controls
DNA Replication and Protein Synthesis DNA is Deoxyribonucleic Acid. It holds all of our genetic information which is passed down through sexual reproduction DNA has three main functions: 1. DNA Controls
Nucleic acids deoxyribonucleic acid (DNA) ribonucleic acid (RNA) nucleotide
 Nucleic Acids Nucleic acids are molecules that store information for cellular growth and reproduction There are two types of nucleic acids: - deoxyribonucleic acid (DNA) and ribonucleic acid (RNA) These
Nucleic Acids Nucleic acids are molecules that store information for cellular growth and reproduction There are two types of nucleic acids: - deoxyribonucleic acid (DNA) and ribonucleic acid (RNA) These
DNA Structure and Analysis. Chapter 4: Background
 DNA Structure and Analysis Chapter 4: Background Molecular Biology Three main disciplines of biotechnology Biochemistry Genetics Molecular Biology # Biotechnology: A Laboratory Skills Course explorer.bio-rad.com
DNA Structure and Analysis Chapter 4: Background Molecular Biology Three main disciplines of biotechnology Biochemistry Genetics Molecular Biology # Biotechnology: A Laboratory Skills Course explorer.bio-rad.com
Answers to Module 1. An obligate aerobe is an organism that has an absolute requirement of oxygen for growth.
 Answers to Module 1 Short Answers 1) What is an obligate aerobe? An obligate aerobe is an organism that has an absolute requirement of oxygen for growth. What about facultative anaerobe? 2) Distinguish
Answers to Module 1 Short Answers 1) What is an obligate aerobe? An obligate aerobe is an organism that has an absolute requirement of oxygen for growth. What about facultative anaerobe? 2) Distinguish
BIOLOGY. Chapter 16 GenesExpression
 BIOLOGY Chapter 16 GenesExpression CAMPBELL BIOLOGY TENTH EDITION Reece Urry Cain Wasserman Minorsky Jackson 18 Gene Expression 2014 Pearson Education, Inc. Figure 16.1 Differential Gene Expression results
BIOLOGY Chapter 16 GenesExpression CAMPBELL BIOLOGY TENTH EDITION Reece Urry Cain Wasserman Minorsky Jackson 18 Gene Expression 2014 Pearson Education, Inc. Figure 16.1 Differential Gene Expression results
DNA DNA Profiling 18. Discuss the stages involved in DNA profiling 19. Define the process of DNA profiling 20. Give two uses of DNA profiling
 Name: 2.5 Genetics Objectives At the end of this sub section students should be able to: 2.5.1 Heredity and Variation 1. Discuss the diversity of organisms 2. Define the term species 3. Distinguish between
Name: 2.5 Genetics Objectives At the end of this sub section students should be able to: 2.5.1 Heredity and Variation 1. Discuss the diversity of organisms 2. Define the term species 3. Distinguish between
Replication Review. 1. What is DNA Replication? 2. Where does DNA Replication take place in eukaryotic cells?
 Replication Review 1. What is DNA Replication? 2. Where does DNA Replication take place in eukaryotic cells? 3. Where does DNA Replication take place in the cell cycle? 4. 4. What guides DNA Replication?
Replication Review 1. What is DNA Replication? 2. Where does DNA Replication take place in eukaryotic cells? 3. Where does DNA Replication take place in the cell cycle? 4. 4. What guides DNA Replication?
Name Class Date. Practice Test
 Name Class Date 12 DNA Practice Test Multiple Choice Write the letter that best answers the question or completes the statement on the line provided. 1. What do bacteriophages infect? a. mice. c. viruses.
Name Class Date 12 DNA Practice Test Multiple Choice Write the letter that best answers the question or completes the statement on the line provided. 1. What do bacteriophages infect? a. mice. c. viruses.
Prokaryotic Transcription
 Prokaryotic Transcription Transcription Basics DNA is the genetic material Nucleic acid Capable of self-replication and synthesis of RNA RNA is the middle man Nucleic acid Structure and base sequence are
Prokaryotic Transcription Transcription Basics DNA is the genetic material Nucleic acid Capable of self-replication and synthesis of RNA RNA is the middle man Nucleic acid Structure and base sequence are
8/21/2014. From Gene to Protein
 From Gene to Protein Chapter 17 Objectives Describe the contributions made by Garrod, Beadle, and Tatum to our understanding of the relationship between genes and enzymes Briefly explain how information
From Gene to Protein Chapter 17 Objectives Describe the contributions made by Garrod, Beadle, and Tatum to our understanding of the relationship between genes and enzymes Briefly explain how information
M I C R O B I O L O G Y WITH DISEASES BY TAXONOMY, THIRD EDITION
 M I C R O B I O L O G Y WITH DISEASES BY TAXONOMY, THIRD EDITION Chapter 7 Microbial Genetics Lecture prepared by Mindy Miller-Kittrell, University of Tennessee, Knoxville The Structure and Replication
M I C R O B I O L O G Y WITH DISEASES BY TAXONOMY, THIRD EDITION Chapter 7 Microbial Genetics Lecture prepared by Mindy Miller-Kittrell, University of Tennessee, Knoxville The Structure and Replication
Gene Expression Transcription
 Why? ene Expression Transcription How is mrn synthesized and what message does it carry? DN is often referred to as a genetic blueprint. In the same way that blueprints contain the instructions for construction
Why? ene Expression Transcription How is mrn synthesized and what message does it carry? DN is often referred to as a genetic blueprint. In the same way that blueprints contain the instructions for construction
NON MENDELIAN GENETICS. DNA, PROTEIN SYNTHESIS, MUTATIONS DUE DECEMBER 8TH
 NON MENDELIAN GENETICS. DNA, PROTEIN SYNTHESIS, MUTATIONS DUE DECEMBER 8TH MONDAY TUESDAY WEDNESDAY THURSDAY FRIDAY 11/14 11/15 11/16 11/17 11/18 Non-Mendelian Genetics DNA Structure and Replication 11/28
NON MENDELIAN GENETICS. DNA, PROTEIN SYNTHESIS, MUTATIONS DUE DECEMBER 8TH MONDAY TUESDAY WEDNESDAY THURSDAY FRIDAY 11/14 11/15 11/16 11/17 11/18 Non-Mendelian Genetics DNA Structure and Replication 11/28
GENE REGULATION slide shows by Kim Foglia modified Slides with blue edges are Kim s
 GENE REGULATION slide shows by Kim Foglia modified Slides with blue edges are Kim s 2007-2008 Bacterial metabolism Bacteria need to respond quickly to changes in their environment STOP GO if they have
GENE REGULATION slide shows by Kim Foglia modified Slides with blue edges are Kim s 2007-2008 Bacterial metabolism Bacteria need to respond quickly to changes in their environment STOP GO if they have
Chapter 15 Gene Technologies and Human Applications
 Chapter Outline Chapter 15 Gene Technologies and Human Applications Section 1: The Human Genome KEY IDEAS > Why is the Human Genome Project so important? > How do genomics and gene technologies affect
Chapter Outline Chapter 15 Gene Technologies and Human Applications Section 1: The Human Genome KEY IDEAS > Why is the Human Genome Project so important? > How do genomics and gene technologies affect
What we ll do today. Types of stem cells. Do engineered ips and ES cells have. What genes are special in stem cells?
 Do engineered ips and ES cells have similar molecular signatures? What we ll do today Research questions in stem cell biology Comparing expression and epigenetics in stem cells asuring gene expression
Do engineered ips and ES cells have similar molecular signatures? What we ll do today Research questions in stem cell biology Comparing expression and epigenetics in stem cells asuring gene expression
Chapter 8: DNA and RNA
 Chapter 8: DNA and RNA Lecture Outline Enger, E. D., Ross, F. C., & Bailey, D. B. (2012). Concepts in biology (14th ed.). New York: McGraw- Hill. 1 8-1 DNA and the Importance of Proteins Proteins play
Chapter 8: DNA and RNA Lecture Outline Enger, E. D., Ross, F. C., & Bailey, D. B. (2012). Concepts in biology (14th ed.). New York: McGraw- Hill. 1 8-1 DNA and the Importance of Proteins Proteins play
Chapter 18: Regulation of Gene Expression. 1. Gene Regulation in Bacteria 2. Gene Regulation in Eukaryotes 3. Gene Regulation & Cancer
 Chapter 18: Regulation of Gene Expression 1. Gene Regulation in Bacteria 2. Gene Regulation in Eukaryotes 3. Gene Regulation & Cancer Gene Regulation Gene regulation refers to all aspects of controlling
Chapter 18: Regulation of Gene Expression 1. Gene Regulation in Bacteria 2. Gene Regulation in Eukaryotes 3. Gene Regulation & Cancer Gene Regulation Gene regulation refers to all aspects of controlling
Do engineered ips and ES cells have similar molecular signatures?
 Do engineered ips and ES cells have similar molecular signatures? Comparing expression and epigenetics in stem cells George Bell, Ph.D. Bioinformatics and Research Computing 2012 Spring Lecture Series
Do engineered ips and ES cells have similar molecular signatures? Comparing expression and epigenetics in stem cells George Bell, Ph.D. Bioinformatics and Research Computing 2012 Spring Lecture Series
DNA/RNA STUDY GUIDE. Match the following scientists with their accomplishments in discovering DNA using the statement in the box below.
 Name: Period: Date: DNA/RNA STUDY GUIDE Part A: DNA History Match the following scientists with their accomplishments in discovering DNA using the statement in the box below. Used a technique called x-ray
Name: Period: Date: DNA/RNA STUDY GUIDE Part A: DNA History Match the following scientists with their accomplishments in discovering DNA using the statement in the box below. Used a technique called x-ray
Total Test Questions: 66 Levels: Grades Units of Credit: 1.0 STANDARD 2. Demonstrate appropriate use of personal protective devices.
 DESCRIPTION Biotechnology is designed to create an awareness of career possibilities in the field of biotechnology. Students are introduced to diagnostic and therapeutic laboratory procedures that support
DESCRIPTION Biotechnology is designed to create an awareness of career possibilities in the field of biotechnology. Students are introduced to diagnostic and therapeutic laboratory procedures that support
Themes: RNA and RNA Processing. Messenger RNA (mrna) What is a gene? RNA is very versatile! RNA-RNA interactions are very important!
 Themes: RNA is very versatile! RNA and RNA Processing Chapter 14 RNA-RNA interactions are very important! Prokaryotes and Eukaryotes have many important differences. Messenger RNA (mrna) Carries genetic
Themes: RNA is very versatile! RNA and RNA Processing Chapter 14 RNA-RNA interactions are very important! Prokaryotes and Eukaryotes have many important differences. Messenger RNA (mrna) Carries genetic
DNA makes RNA makes Proteins. The Central Dogma
 DNA makes RNA makes Proteins The Central Dogma TRANSCRIPTION DNA RNA transcript RNA polymerase RNA PROCESSING Exon RNA transcript (pre-mrna) Intron Aminoacyl-tRNA synthetase NUCLEUS CYTOPLASM FORMATION
DNA makes RNA makes Proteins The Central Dogma TRANSCRIPTION DNA RNA transcript RNA polymerase RNA PROCESSING Exon RNA transcript (pre-mrna) Intron Aminoacyl-tRNA synthetase NUCLEUS CYTOPLASM FORMATION
Do you remember. What is a gene? What is RNA? How does it differ from DNA? What is protein?
 Lesson 1 - RNA Do you remember What is a gene? What is RNA? How does it differ from DNA? What is protein? Gene Segment of DNA that codes for building a protein DNA code is copied into RNA form, and RNA
Lesson 1 - RNA Do you remember What is a gene? What is RNA? How does it differ from DNA? What is protein? Gene Segment of DNA that codes for building a protein DNA code is copied into RNA form, and RNA
Chapter 8 Lecture Outline. Transcription, Translation, and Bioinformatics
 Chapter 8 Lecture Outline Transcription, Translation, and Bioinformatics Replication, Transcription, Translation n Repetitive processes Build polymers of nucleotides or amino acids n All have 3 major steps
Chapter 8 Lecture Outline Transcription, Translation, and Bioinformatics Replication, Transcription, Translation n Repetitive processes Build polymers of nucleotides or amino acids n All have 3 major steps
DNA and RNA. Chapter 12
 DNA and RNA Chapter 12 Warm Up Exercise Test Corrections Make sure to indicate your new answer and provide an explanation for why this is the correct answer. Do this with a red pen in the margins of your
DNA and RNA Chapter 12 Warm Up Exercise Test Corrections Make sure to indicate your new answer and provide an explanation for why this is the correct answer. Do this with a red pen in the margins of your
Chapter 10 - Molecular Biology of the Gene
 Bio 100 - Molecular Genetics 1 A. Bacterial Transformation Chapter 10 - Molecular Biology of the Gene Researchers found that they could transfer an inherited characteristic (e.g. the ability to cause pneumonia),
Bio 100 - Molecular Genetics 1 A. Bacterial Transformation Chapter 10 - Molecular Biology of the Gene Researchers found that they could transfer an inherited characteristic (e.g. the ability to cause pneumonia),
Chapter 5 DNA and Chromosomes
 Chapter 5 DNA and Chromosomes DNA as the genetic material Heat-killed bacteria can transform living cells S Smooth R Rough Fred Griffith, 1920 DNA is the genetic material Oswald Avery Colin MacLeod Maclyn
Chapter 5 DNA and Chromosomes DNA as the genetic material Heat-killed bacteria can transform living cells S Smooth R Rough Fred Griffith, 1920 DNA is the genetic material Oswald Avery Colin MacLeod Maclyn
PROTEIN SYNTHESIS. copyright cmassengale
 PROTEIN SYNTHESIS 1 DNA and Genes 2 Roles of RNA and DNA DNA is the MASTER PLAN RNA is the BLUEPRINT of the Master Plan 3 RNA Differs from DNA RNA has a sugar ribose DNA has a sugar deoxyribose 4 Other
PROTEIN SYNTHESIS 1 DNA and Genes 2 Roles of RNA and DNA DNA is the MASTER PLAN RNA is the BLUEPRINT of the Master Plan 3 RNA Differs from DNA RNA has a sugar ribose DNA has a sugar deoxyribose 4 Other
PROTEIN SYNTHESIS Flow of Genetic Information The flow of genetic information can be symbolized as: DNA RNA Protein
 PROTEIN SYNTHESIS Flow of Genetic Information The flow of genetic information can be symbolized as: DNA RNA Protein This is also known as: The central dogma of molecular biology Protein Proteins are made
PROTEIN SYNTHESIS Flow of Genetic Information The flow of genetic information can be symbolized as: DNA RNA Protein This is also known as: The central dogma of molecular biology Protein Proteins are made
Protein Synthesis. OpenStax College
 OpenStax-CNX module: m46032 1 Protein Synthesis OpenStax College This work is produced by OpenStax-CNX and licensed under the Creative Commons Attribution License 3.0 By the end of this section, you will
OpenStax-CNX module: m46032 1 Protein Synthesis OpenStax College This work is produced by OpenStax-CNX and licensed under the Creative Commons Attribution License 3.0 By the end of this section, you will
Introduction to Bioinformatics
 Introduction to Bioinformatics Contents Cell biology Organisms and cells Building blocks of cells How genes encode proteins? Bioinformatics What is bioinformatics? Practical applications Tools and databases
Introduction to Bioinformatics Contents Cell biology Organisms and cells Building blocks of cells How genes encode proteins? Bioinformatics What is bioinformatics? Practical applications Tools and databases
CHAPTERS , 17: Eukaryotic Genetics
 CHAPTERS 14.1 14.6, 17: Eukaryotic Genetics 1. Review the levels of DNA packing within the eukaryote nucleus. Label each level. (A similar diagram is on pg 188 of your textbook.) 2. How do the coding regions
CHAPTERS 14.1 14.6, 17: Eukaryotic Genetics 1. Review the levels of DNA packing within the eukaryote nucleus. Label each level. (A similar diagram is on pg 188 of your textbook.) 2. How do the coding regions
M Keramatipour 2. M Keramatipour 1. M Keramatipour 4. M Keramatipour 3. M Keramatipour 5. M Keramatipour
 Molecular Cloning Methods Mohammad Keramatipour MD, PhD keramatipour@tums.ac.ir Outline DNA recombinant technology DNA cloning co Cell based PCR PCR-based Some application of DNA cloning Genomic libraries
Molecular Cloning Methods Mohammad Keramatipour MD, PhD keramatipour@tums.ac.ir Outline DNA recombinant technology DNA cloning co Cell based PCR PCR-based Some application of DNA cloning Genomic libraries
Gene Expression: Transcription
 Gene Expression: Transcription The majority of genes are expressed as the proteins they encode. The process occurs in two steps: Transcription = DNA RNA Translation = RNA protein Taken together, they make
Gene Expression: Transcription The majority of genes are expressed as the proteins they encode. The process occurs in two steps: Transcription = DNA RNA Translation = RNA protein Taken together, they make
HiPer RT-PCR Teaching Kit
 HiPer RT-PCR Teaching Kit Product Code: HTBM024 Number of experiments that can be performed: 5 Duration of Experiment: Protocol: 4 hours Agarose Gel Electrophoresis: 45 minutes Storage Instructions: The
HiPer RT-PCR Teaching Kit Product Code: HTBM024 Number of experiments that can be performed: 5 Duration of Experiment: Protocol: 4 hours Agarose Gel Electrophoresis: 45 minutes Storage Instructions: The
Nucleic acids and protein synthesis
 THE FUNCTIONS OF DNA Nucleic acids and protein synthesis The full name of DNA is deoxyribonucleic acid. Every nucleotide has the same sugar molecule and phosphate group, but each nucleotide contains one
THE FUNCTIONS OF DNA Nucleic acids and protein synthesis The full name of DNA is deoxyribonucleic acid. Every nucleotide has the same sugar molecule and phosphate group, but each nucleotide contains one
If Dna Has The Instructions For Building Proteins Why Is Mrna Needed
 If Dna Has The Instructions For Building Proteins Why Is Mrna Needed if a strand of DNA has the sequence CGGTATATC, then the complementary each strand of DNA contains the info needed to produce the complementary
If Dna Has The Instructions For Building Proteins Why Is Mrna Needed if a strand of DNA has the sequence CGGTATATC, then the complementary each strand of DNA contains the info needed to produce the complementary
2. From the first paragraph in this section, find three ways in which RNA differs from DNA.
 Name Chapter 17: From Gene to Protein Begin reading at page 328 Basic Principles of Transcription and Translation. Work on this chapter a single concept at a time, and expect to spend at least 6 hours
Name Chapter 17: From Gene to Protein Begin reading at page 328 Basic Principles of Transcription and Translation. Work on this chapter a single concept at a time, and expect to spend at least 6 hours
From Gene to Protein Transcription and Translation
 Name: Hour: From Gene to Protein Transcription and Translation Introduction: In this activity you will learn how the genes in our DNA influence our characteristics. For example, how can a gene cause albinism
Name: Hour: From Gene to Protein Transcription and Translation Introduction: In this activity you will learn how the genes in our DNA influence our characteristics. For example, how can a gene cause albinism
TRANSCRIPTION AND TRANSLATION
 TRANSCRIPTION AND TRANSLATION Bell Ringer (5 MINUTES) 1. Have your homework (any missing work) out on your desk and ready to turn in 2. Draw and label a nucleotide. 3. Summarize the steps of DNA replication.
TRANSCRIPTION AND TRANSLATION Bell Ringer (5 MINUTES) 1. Have your homework (any missing work) out on your desk and ready to turn in 2. Draw and label a nucleotide. 3. Summarize the steps of DNA replication.
DNA Structure and Properties Basic Properties Predicting Melting Temperature. Dinesh Yadav
 DNA Structure and Properties Basic Properties Predicting Melting Temperature Dinesh Yadav Nucleic Acid Structure Question: Is this RNA or DNA? Molecules of Life, pp. 15 2 Nucleic Acid Bases Molecules of
DNA Structure and Properties Basic Properties Predicting Melting Temperature Dinesh Yadav Nucleic Acid Structure Question: Is this RNA or DNA? Molecules of Life, pp. 15 2 Nucleic Acid Bases Molecules of
RNA and Protein Synthesis
 RNA and Protein Synthesis CTE: Agriculture and Natural Resources: C5.3 Understand various cell actions, such as osmosis and cell division. C5.4 Compare and contrast plant and animal cells, bacteria, and
RNA and Protein Synthesis CTE: Agriculture and Natural Resources: C5.3 Understand various cell actions, such as osmosis and cell division. C5.4 Compare and contrast plant and animal cells, bacteria, and
Self-test Quiz for Chapter 12 (From DNA to Protein: Genotype to Phenotype)
 Self-test Quiz for Chapter 12 (From DNA to Protein: Genotype to Phenotype) Question#1: One-Gene, One-Polypeptide The figure below shows the results of feeding trials with one auxotroph strain of Neurospora
Self-test Quiz for Chapter 12 (From DNA to Protein: Genotype to Phenotype) Question#1: One-Gene, One-Polypeptide The figure below shows the results of feeding trials with one auxotroph strain of Neurospora
American Society of Cytopathology Core Curriculum in Molecular Biology
 American Society of Cytopathology Core Curriculum in Molecular Biology American Society of Cytopathology Core Curriculum in Molecular Biology Chapter 3 Molecular Techniques Separation and Detection, Part
American Society of Cytopathology Core Curriculum in Molecular Biology American Society of Cytopathology Core Curriculum in Molecular Biology Chapter 3 Molecular Techniques Separation and Detection, Part
The common structure of a DNA nucleotide. Hewitt
 GENETICS Unless otherwise noted* the artwork and photographs in this slide show are original and by Burt Carter. Permission is granted to use them for non-commercial, non-profit educational purposes provided
GENETICS Unless otherwise noted* the artwork and photographs in this slide show are original and by Burt Carter. Permission is granted to use them for non-commercial, non-profit educational purposes provided
