Nucleic Acids Research
|
|
|
- Lucas Armstrong
- 6 years ago
- Views:
Transcription
1 Volume 16 Number Nucleic Acids Research Variable rrna gene copies in extreme halobacteria J.L.Sanz, I.Marin, L.Ramirez, J.P.Abad1, C.L.Smith2 and R.Amils* Centro de Biologfa Molecular, UAM, Cantoblanco, Madrid, Spain, 'Department of Genetics and Development and 2Microbiology and Psychiatry, College of Physicians and Surgeons, Columbia University, New York, NY 10032, USA Received June 21, 1988; Revised and Accepted July 20, 1988 ABSTRACT Using PFG electrophoresis techniques, we have examined the organization of rrna gene in halobacterium species. The results show that the organization of rrna genes among closely related halobacteria is quite heterogeneous. This contrasts with the high degree of conservation of rrna sequence (1). The possible mechanism of such rrna gene amplification and its evolutionary implications are discussed. INTRODUCTION A fundamental question regarding archaebacterial phylogeny is how their genome compares within this group of microorganisms and with eubacteria and eukaryotic organisms. The genomic organization of rrna is one of the systems used to address this basic question. It was highly surprising to find that the first archaebacterial species analyzed, H. halobium (2), had only one rrna gene as opposed to the multiple copies found in eubacteria (seven for E. coli) and the high number of copies in the eukaryotic kingdom. Studies of rrna gene organization with sulphur dependent archaebacteria showed that the single-rrna gene structure is more generalized than previously had been thought (3), although this type of organization is not characteristic of all archaebacteria. For instance, the methanogenic branch of archaebacteria showed a more complex structure containing two rrna genes for Mb. thermoautotrophicum (4) and four genes for Mc. vannielli (5). Large DNA technology and PFG electrophoresis have allowed an efficient and accurate means for analyzing the genomic organization of halobacteria. I R L Press Limited, Oxford, England. 7827
2 MATERIAL AND METHODS Haloarcula californiae ATCC 33799, "Halobacterium marismortui" Haloferax gibbonsli ATCC Halococcus morhuae, Halobacterium halobium NCMB 777, Halobacterium salinarium CCM 2148 were used. Details of the DNA preparation, digestion and electrophoresis are as described in references 6., 7. DNA obtained from each preparation was prerun with 100 second pulses in order to remove plasmids and degraded DNA. Macrorestriction fragments were resolved by PFG electrophoresis in the conditions described in the legends of the figures. The PFG electrophoretic gels were blotted and hybridizated (8) with 32p labeled H. mediterranei 16S and 23S rrna. RESULTS AND DISCUSSION Autoradiography of the blotted PFG electrophoresis of the macrorestriction fragments correspondent to different halobacteria revealed hybridization to various fragments (fig 1 and 2). Table 1 shows that the number of fragments detected is constant for each organism and independent of the restriction enzymes (DraI, NotI, SfiI, BamHI) and probe used (16S or 23S rrna). Detection of multiple fragments using rrna probes is not due to partial digestion of the genomic DNA by the restriction enzymes. Close examination of the ethidium bromide photograph of PFG experiments like those shown in Figure 1 and 2 reveal no evidence of partial digestion and the same number of TABLE 1. Summary of PFG experiments using chromosomal DNA from different halobacteria, cut with different restriction enzymes and hybridized with 23S and 16S rrna probes. Microorganism Enzymes used Minimum gene number NotI DraI Sf ii BamHI Haloarcula caiiforniae ATCC * * * 4 Haloferax gibbonsii ATCC * * * * 4 Halobacterium halobium NCMB 777 * * * 3 "Halobacterium marismortui" * * * 3 Halococcus morhuae NCMB 757 * * * * 2 Halobacterium salinarium CCM 2148 * * * *
3 hybridization fragments is obtained with the various restriction enzymes (Table 1). The digestion patterns are reproducible and no further digestion is obtained with addition of fresh enzyme and/or longer incubation times. Detection of multiple fragments using rrna gene probe is not due to the presence of restriction site in 16S or 23S rdna because: i) the same hybridization pattern is obtained with both rrna probes, ii) the highly homologous 16S rrna sequences of three taxonomically distant halobacteria H. halobium H. volcanii and H. morrhuae, lack restriction sites for any of the AH XB 7 a >H >B FIG. 1. PULSED FIELD GEL ELECTROPHORESIS OF HALOBACTERIAL DNA (A) AND SOUTHERN HYBRIDIZATION WITH H. mediterranei 16S rrna (B). Samples were separated in a 55 cm double inhomogeneus PFG apparatus for 72 hours at 500 volts with 5 second pulses between 900 field reorientations. The lanes are: 1 Saccharomyces cerevisiae, 2 and 7 Haloarcula californiae; 3 and 8 "Halobacterium marismortui";4 and 9 Haloferax gibbonsii; 5 and 10 Halococcus morrhuae; 6 and 11 Halobacterium halobium digested with NotI Tlanes 2 to 6) and DraI (lanes 7 to 11). -XH and Xe *are digestions Of \ vir (48.5 Kb) with HindIII and EcoRI (,X) and BstI (Xtl). 7829
4 r_~~~~~~~ -350 kb kb -250 kb kb kh 150 kb 00k b FIG. 2. PULSED FIELD GEL ELECTROPHORESIS OF HALOBACTERIAL DNA and SOUTHERN HYBRIDIZATION WITH H. mediterranei 23S rrna. Samples were run in the same conditions as described in Fig. 1 with the exception of the pulse time that was 25 seconds. The lanes are: 1., Halobacterium salinarium; 2 and 4, H. gibbnsi; 3 and 5 Hc. morrhuae; 6 H. halobium digested with BamIii (lanes 1 to 3) and NotI (lanes 4 to 6). Numbers indicate the sizes of the restriction bands in kilobases obtained using cancatemers of\~ ir (48.5 Kb monomer). enzymes used in this work (1), and iii) synthesized 5S rrna oligonucleotides, which lack restriction sites for any of the enzymes used, produce the same hybridization patterns as those obtained with 16S and 23S rrna probes (data not shown). The gene numbers shown in Table 1 are the minimal values. It is possible that the megabase restriction fragments detected in the experiment shown in Figure 1 and 2 contain more than one copy of rrna genes, which might explain why in the southern hybridization of Figure 1 some bands do not show equivalent density, suggesting the presence of more than one rrna gene in some of the bands. Our results show that there is considerable variation in the number of rrna genes in Halobacteria. The one rrna gene detected for H. salinariunu CCM 2148 in this study is the same as was reported for Hi. halobium strain Rl (2) and H. cutirubrum (9) and with the taxonomic concept that all three are indeed the same species (10). Three rrna genes have been detected for 7830
5 H. halobium NCMB 777, a different species from the H. halobium Rl, which corroborates the taxonomic incongruence of giving the same name (11) to two very different species. There is a reference to the possibility of two rrna genes for H. volcanii (12), which agrees with the gradient of rrna genomic organization reported in this work. Earlier studies showed that the sulfur dependent branch of archaebacteria is uniform in having only one copy of its rrna genes, although some differences in their arrangements (3) were reported. The situation is quite different in the methanogenichalobacterial branch. In contrast to the presence of a single gene in H. halobium Rl and H. cutirubrum, the reported number in the methanogens varies: two for M. thermoautotrophicum and four in Mc. vannielli. It is obvious that the presumed difference in rrna gene organization between halobacteria and methanogens disappears when the diverse numbers of halobacterial rrna gene copies are considered. The characteristic differences between rrna gene organization of of the sulfur dependent and the methanogenic-halobacterial branches of archaebacteria may be of phylogenetic relevance. It has been reported that halobacteria have high numbers of insertion elements (12), these mobile elements might facilitate chromosome rearrangement. Current literature contains many references to the evolutionary importance of the gradient of rrna genes ranging from one in the sulfur dependent archaebacteria to between one and four in the methanogenic-halobacteria, seven in the eubacterial gram negative E. coli, and several hundred in eukaryotic systems (13). The most frequent explanation attributes this variability to the adjustment of the transcription of rrna and the translation of the ribosomal proteins needed for the adequate assembly of the ribosomes. So far it has been difficult to correlate this gene amplification with any known advantage other than relaxation of the type of selective pressure which would exist when only one copy of rrna gene is present. Clearly further studies of halobacterial chromosomal organization are needed to determine the mechanism of rdna amplification, as well as how stable these multiple 7831
6 copies are, and the phenotypic difference, if any, that multiple rrna genes confer on the various species. Currently we are constructing chromosomal physical maps of these organisms in order to investigate these important questions. Acknowledgments This work was supported by grants from DOE, DE-FG02-87ER-gD852, the MacArthur Foundation, Pharmacia-LKB; CSIC and a institutional grant from the Fondo de Investigaciones Sanitarias. J.L. Sanz and I. Marnn are fellows at the CSIC, and L. Ramirez is a fellow at the Ministerio de Universidades e Investigacion. *To whom correspondence should be addressed REFERENCE 1. Huysmans,E. and De Wachter,R. (1986) Nucleic Acids Res. 14 (Supplement) r73-r Hofman,J.D., Lau,R.H. and Doolite,W.F. (1979). Nucleic Acids Res. 7, Bock,A., Hummel,H., Jarsch,M. and Wich,G. (1986) Biology of Anaerobic Bacteria. Ed: H.C. Dubourguier et al. Elsevier Science Publ., Amsterdam Lechner,K., Wich,G. and Bock,A. (1985) Syst. Appl. Microbiol Neumann,H., Gierl,A., Tu,J., Leibrock,J., Staiger,G. and Zillig,W. (1983) Molec. Gen. Genet. 192, Smith,C.L. and Cantor,C.R. (1987) (R. Wu, ed) (Academic Press, New York) pp (1987). 7. Smith,C.L., Lawrance,S.K., Gillespie,G.A., Cantor,C.R. Weessman,S.M. and Collins,F.S. (1987) Methods in Enzymology 151, , M. Gottesman, ed. Academic Press, New York. 8. Southern, E.J. (1975), Mol. Biol. 98, Hui,I. and Dennis,P.P. (1985) J. Biol. Chem. 260, Larsen.H. (1984) Bergey's Manual of Systematic Bacteriology gth Edit (Vol. 1) N. R. Krieg & J. G. Holt Edit. 11. Ross,H.N. & Grant,W.D. (1985) J. Gen. Microbiol. 131J DoolittleyW.F. (1985) In "The Bacteria" Archaebacteria. Vol VIII, , C.R. Woese and R.S. Wolfe Eds. Academic Press. Inc., New York. 13. Dyson,P. and Sherratt,D. (1985) In The Evolution of Genome Size, , Ed: T. Cavalier-Smith. Wiley-Interscience Publication. 7832
Molecular Cell Biology - Problem Drill 11: Recombinant DNA
 Molecular Cell Biology - Problem Drill 11: Recombinant DNA Question No. 1 of 10 1. Which of the following statements about the sources of DNA used for molecular cloning is correct? Question #1 (A) cdna
Molecular Cell Biology - Problem Drill 11: Recombinant DNA Question No. 1 of 10 1. Which of the following statements about the sources of DNA used for molecular cloning is correct? Question #1 (A) cdna
BS 50 Genetics and Genomics Week of Nov 29
 BS 50 Genetics and Genomics Week of Nov 29 Additional Practice Problems for Section Problem 1. A linear piece of DNA is digested with restriction enzymes EcoRI and HinDIII, and the products are separated
BS 50 Genetics and Genomics Week of Nov 29 Additional Practice Problems for Section Problem 1. A linear piece of DNA is digested with restriction enzymes EcoRI and HinDIII, and the products are separated
Production and Characterization of Protease from an Extremely Halophilic Halobacterium sp. PB407
 Kasetsart J. (Nat. Sci.) 38 : 15-20 (2004) Production and Characterization of Protease from an Extremely Halophilic Halobacterium sp. PB407 Werasit Kanlayakrit 1, Preeyanuch Bovornreungroj 1, Takuji Oka
Kasetsart J. (Nat. Sci.) 38 : 15-20 (2004) Production and Characterization of Protease from an Extremely Halophilic Halobacterium sp. PB407 Werasit Kanlayakrit 1, Preeyanuch Bovornreungroj 1, Takuji Oka
Computational Biology I LSM5191
 Computational Biology I LSM5191 Lecture 5 Notes: Genetic manipulation & Molecular Biology techniques Broad Overview of: Enzymatic tools in Molecular Biology Gel electrophoresis Restriction mapping DNA
Computational Biology I LSM5191 Lecture 5 Notes: Genetic manipulation & Molecular Biology techniques Broad Overview of: Enzymatic tools in Molecular Biology Gel electrophoresis Restriction mapping DNA
Microbial Diversity and Assessment (III) Spring, 2007 Guangyi Wang, Ph.D. POST103B
 Microbial Diversity and Assessment (III) Spring, 2007 Guangyi Wang, Ph.D. POST103B guangyi@hawaii.edu http://www.soest.hawaii.edu/marinefungi/ocn403webpage.htm Overview of Last Lecture Taxonomy (three
Microbial Diversity and Assessment (III) Spring, 2007 Guangyi Wang, Ph.D. POST103B guangyi@hawaii.edu http://www.soest.hawaii.edu/marinefungi/ocn403webpage.htm Overview of Last Lecture Taxonomy (three
Providing clear solutions to microbiological challenges TM. cgmp/iso CLIA. Polyphasic Microbial Identification & DNA Fingerprinting
 Providing clear solutions to microbiological challenges TM Cert. No. 2254.01 Polyphasic Microbial Identification & DNA Fingerprinting Microbial Contamination Tracking & Trending cgmp/iso-17025-2005 CLIA
Providing clear solutions to microbiological challenges TM Cert. No. 2254.01 Polyphasic Microbial Identification & DNA Fingerprinting Microbial Contamination Tracking & Trending cgmp/iso-17025-2005 CLIA
2054, Chap. 14, page 1
 2054, Chap. 14, page 1 I. Recombinant DNA technology (Chapter 14) A. recombinant DNA technology = collection of methods used to perform genetic engineering 1. genetic engineering = deliberate modification
2054, Chap. 14, page 1 I. Recombinant DNA technology (Chapter 14) A. recombinant DNA technology = collection of methods used to perform genetic engineering 1. genetic engineering = deliberate modification
Lecture Four. Molecular Approaches I: Nucleic Acids
 Lecture Four. Molecular Approaches I: Nucleic Acids I. Recombinant DNA and Gene Cloning Recombinant DNA is DNA that has been created artificially. DNA from two or more sources is incorporated into a single
Lecture Four. Molecular Approaches I: Nucleic Acids I. Recombinant DNA and Gene Cloning Recombinant DNA is DNA that has been created artificially. DNA from two or more sources is incorporated into a single
Genetic Engineering & Recombinant DNA
 Genetic Engineering & Recombinant DNA Chapter 10 Copyright The McGraw-Hill Companies, Inc) Permission required for reproduction or display. Applications of Genetic Engineering Basic science vs. Applied
Genetic Engineering & Recombinant DNA Chapter 10 Copyright The McGraw-Hill Companies, Inc) Permission required for reproduction or display. Applications of Genetic Engineering Basic science vs. Applied
Nucleic Acids Research
 Volume 16 Number 24 1988 Nucleic Acids Research Rapid physical mapping of the Mycoplasma mobile genome by two-dimensional field inversion gel electrophoresis techniques Wilfried Bautsch Zentrum Biochemie,
Volume 16 Number 24 1988 Nucleic Acids Research Rapid physical mapping of the Mycoplasma mobile genome by two-dimensional field inversion gel electrophoresis techniques Wilfried Bautsch Zentrum Biochemie,
Multiple choice questions (numbers in brackets indicate the number of correct answers)
 1 Multiple choice questions (numbers in brackets indicate the number of correct answers) February 1, 2013 1. Ribose is found in Nucleic acids Proteins Lipids RNA DNA (2) 2. Most RNA in cells is transfer
1 Multiple choice questions (numbers in brackets indicate the number of correct answers) February 1, 2013 1. Ribose is found in Nucleic acids Proteins Lipids RNA DNA (2) 2. Most RNA in cells is transfer
2. Outline the levels of DNA packing in the eukaryotic nucleus below next to the diagram provided.
 AP Biology Reading Packet 6- Molecular Genetics Part 2 Name Chapter 19: Eukaryotic Genomes 1. Define the following terms: a. Euchromatin b. Heterochromatin c. Nucleosome 2. Outline the levels of DNA packing
AP Biology Reading Packet 6- Molecular Genetics Part 2 Name Chapter 19: Eukaryotic Genomes 1. Define the following terms: a. Euchromatin b. Heterochromatin c. Nucleosome 2. Outline the levels of DNA packing
Biotechnology and Genomics in Public Health. Sharon S. Krag, PhD Johns Hopkins University
 This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike License. Your use of this material constitutes acceptance of that license and the conditions of use of materials on this
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike License. Your use of this material constitutes acceptance of that license and the conditions of use of materials on this
Genetics Lecture 21 Recombinant DNA
 Genetics Lecture 21 Recombinant DNA Recombinant DNA In 1971, a paper published by Kathleen Danna and Daniel Nathans marked the beginning of the recombinant DNA era. The paper described the isolation of
Genetics Lecture 21 Recombinant DNA Recombinant DNA In 1971, a paper published by Kathleen Danna and Daniel Nathans marked the beginning of the recombinant DNA era. The paper described the isolation of
Manipulation of Purified DNA
 Manipulation of Purified DNA To produce the recombinant DNA molecule, the vector, as well as the DNA to be cloned, must be cut at specific points and then joined together in a controlled manner by DNA
Manipulation of Purified DNA To produce the recombinant DNA molecule, the vector, as well as the DNA to be cloned, must be cut at specific points and then joined together in a controlled manner by DNA
Cloning and Sequencing of the Gene Encoding Curvaticin FS47, an Anti- Listerial Bacteriocin Produced by Lactobacillus curvatus FS47
 Cloning and Sequencing of the Gene Encoding Curvaticin FS47, an Anti- Listerial Bacteriocin Produced by Lactobacillus curvatus FS47 S. Macwana, L. Ma, M.A. Cousin, and P.M. Muriana Story in Brief Curvaticin
Cloning and Sequencing of the Gene Encoding Curvaticin FS47, an Anti- Listerial Bacteriocin Produced by Lactobacillus curvatus FS47 S. Macwana, L. Ma, M.A. Cousin, and P.M. Muriana Story in Brief Curvaticin
2. In Figure 10-4, why is edna made only from mrna and not also from trnas and ribosomal RNAs?
 2. In Figure 10-4, why is edna made only from mrna and not also from trnas and ribosomal RNAs? Answer: edna is made from mrna and not from trnas or rrnas because polyt primers are used to prime the first
2. In Figure 10-4, why is edna made only from mrna and not also from trnas and ribosomal RNAs? Answer: edna is made from mrna and not from trnas or rrnas because polyt primers are used to prime the first
Contents... vii. List of Figures... xii. List of Tables... xiv. Abbreviatons... xv. Summary... xvii. 1. Introduction In vitro evolution...
 vii Contents Contents... vii List of Figures... xii List of Tables... xiv Abbreviatons... xv Summary... xvii 1. Introduction...1 1.1 In vitro evolution... 1 1.2 Phage Display Technology... 3 1.3 Cell surface
vii Contents Contents... vii List of Figures... xii List of Tables... xiv Abbreviatons... xv Summary... xvii 1. Introduction...1 1.1 In vitro evolution... 1 1.2 Phage Display Technology... 3 1.3 Cell surface
Manipulating DNA. Nucleic acids are chemically different from other macromolecules such as proteins and carbohydrates.
 Lesson Overview 14.3 Studying the Human Genome Nucleic acids are chemically different from other macromolecules such as proteins and carbohydrates. Nucleic acids are chemically different from other macromolecules
Lesson Overview 14.3 Studying the Human Genome Nucleic acids are chemically different from other macromolecules such as proteins and carbohydrates. Nucleic acids are chemically different from other macromolecules
Synthetic Biology for
 Synthetic Biology for Plasmids and DNA Digestion Plasmids Plasmids are small DNA molecules that are separate from chromosomal DNA They are most commonly found as double stranded, circular DNA Typical plasmids
Synthetic Biology for Plasmids and DNA Digestion Plasmids Plasmids are small DNA molecules that are separate from chromosomal DNA They are most commonly found as double stranded, circular DNA Typical plasmids
Recombinant DNA Technology. The Role of Recombinant DNA Technology in Biotechnology. yeast. Biotechnology. Recombinant DNA technology.
 PowerPoint Lecture Presentations prepared by Mindy Miller-Kittrell, North Carolina State University C H A P T E R 8 Recombinant DNA Technology The Role of Recombinant DNA Technology in Biotechnology Biotechnology?
PowerPoint Lecture Presentations prepared by Mindy Miller-Kittrell, North Carolina State University C H A P T E R 8 Recombinant DNA Technology The Role of Recombinant DNA Technology in Biotechnology Biotechnology?
Chromosome Map of the Thermophilic Archaebacterium Thermococcus celer
 JOURNAL OF BACTERIOLOGY, Dec. 1989, p. 6720-6725 0021-9193/89/126720-06$02.00/0 Copyright 1989, American Society for Microbiology Vol. 171, No. 12 Chromosome Map of the Thermophilic Archaebacterium Thermococcus
JOURNAL OF BACTERIOLOGY, Dec. 1989, p. 6720-6725 0021-9193/89/126720-06$02.00/0 Copyright 1989, American Society for Microbiology Vol. 171, No. 12 Chromosome Map of the Thermophilic Archaebacterium Thermococcus
Contents. 1 Basic Molecular Microbiology of Bacteria... 1 Exp. 1.1 Isolation of Genomic DNA Introduction Principle...
 Contents 1 Basic Molecular Microbiology of Bacteria... 1 Exp. 1.1 Isolation of Genomic DNA... 1 Introduction... 1 Principle... 1 Reagents Required and Their Role... 2 Procedure... 3 Observation... 4 Result
Contents 1 Basic Molecular Microbiology of Bacteria... 1 Exp. 1.1 Isolation of Genomic DNA... 1 Introduction... 1 Principle... 1 Reagents Required and Their Role... 2 Procedure... 3 Observation... 4 Result
Microbiology 微生物学 Spring-Summer
 Microbiology 微生物学 2017 Spring-Summer Relevant Information and Resources Course slides can be found at http://mypage.zju.edu.cn/haichun 教学工作 Course-related questions will be answered through emails. Textbook:
Microbiology 微生物学 2017 Spring-Summer Relevant Information and Resources Course slides can be found at http://mypage.zju.edu.cn/haichun 教学工作 Course-related questions will be answered through emails. Textbook:
pamp, pkan, or pblu?
 pamp, pkan, or pblu? Introduction In my final biotechnology lab project, I was given an unidentified plasmid, in which I was required to perform restriction digests using restriction enzymes. A plasmid,
pamp, pkan, or pblu? Introduction In my final biotechnology lab project, I was given an unidentified plasmid, in which I was required to perform restriction digests using restriction enzymes. A plasmid,
HiPer RT-PCR Teaching Kit
 HiPer RT-PCR Teaching Kit Product Code: HTBM024 Number of experiments that can be performed: 5 Duration of Experiment: Protocol: 4 hours Agarose Gel Electrophoresis: 45 minutes Storage Instructions: The
HiPer RT-PCR Teaching Kit Product Code: HTBM024 Number of experiments that can be performed: 5 Duration of Experiment: Protocol: 4 hours Agarose Gel Electrophoresis: 45 minutes Storage Instructions: The
7 Gene Isolation and Analysis of Multiple
 Genetic Techniques for Biological Research Corinne A. Michels Copyright q 2002 John Wiley & Sons, Ltd ISBNs: 0-471-89921-6 (Hardback); 0-470-84662-3 (Electronic) 7 Gene Isolation and Analysis of Multiple
Genetic Techniques for Biological Research Corinne A. Michels Copyright q 2002 John Wiley & Sons, Ltd ISBNs: 0-471-89921-6 (Hardback); 0-470-84662-3 (Electronic) 7 Gene Isolation and Analysis of Multiple
cgmp/iso CLIA Experience Unsurpassed Quality
 Cert. No. 2254.01 cgmp/iso-17025-2005 CLIA Experience Unsurpassed Quality Polyphasic Microbial Identification & DNA Fingerprinting Microbial Contamination Tracking & Trending Microbial Identification
Cert. No. 2254.01 cgmp/iso-17025-2005 CLIA Experience Unsurpassed Quality Polyphasic Microbial Identification & DNA Fingerprinting Microbial Contamination Tracking & Trending Microbial Identification
Rapid Learning Center Presents. Teach Yourself AP Biology in 24 Hours
 Rapid Learning Center Chemistry :: Biology :: Physics :: Math Rapid Learning Center Presents Teach Yourself AP Biology in 24 Hours 1/35 *AP is a registered trademark of the College Board, which does not
Rapid Learning Center Chemistry :: Biology :: Physics :: Math Rapid Learning Center Presents Teach Yourself AP Biology in 24 Hours 1/35 *AP is a registered trademark of the College Board, which does not
Name_BS50 Exam 3 Key (Fall 2005) Page 2 of 5
 Name_BS50 Exam 3 Key (Fall 2005) Page 2 of 5 Question 1. (14 points) Several Hfr strains derived from the same F + strain were crossed separately to an F - strain, giving the results indicated in the table
Name_BS50 Exam 3 Key (Fall 2005) Page 2 of 5 Question 1. (14 points) Several Hfr strains derived from the same F + strain were crossed separately to an F - strain, giving the results indicated in the table
Year III Pharm.D Dr. V. Chitra
 Year III Pharm.D Dr. V. Chitra 1 Genome entire genetic material of an individual Transcriptome set of transcribed sequences Proteome set of proteins encoded by the genome 2 Only one strand of DNA serves
Year III Pharm.D Dr. V. Chitra 1 Genome entire genetic material of an individual Transcriptome set of transcribed sequences Proteome set of proteins encoded by the genome 2 Only one strand of DNA serves
AP Biology. Chapter 20. Biotechnology: DNA Technology & Genomics. Biotechnology. The BIG Questions. Evolution & breeding of food plants
 What do you notice about these phrases? radar racecar Madam I m Adam Able was I ere I saw Elba a man, a plan, a canal, Panama Was it a bar or a bat I saw? Chapter 20. Biotechnology: DNA Technology & enomics
What do you notice about these phrases? radar racecar Madam I m Adam Able was I ere I saw Elba a man, a plan, a canal, Panama Was it a bar or a bat I saw? Chapter 20. Biotechnology: DNA Technology & enomics
DNA Structure and Analysis. Chapter 4: Background
 DNA Structure and Analysis Chapter 4: Background Molecular Biology Three main disciplines of biotechnology Biochemistry Genetics Molecular Biology # Biotechnology: A Laboratory Skills Course explorer.bio-rad.com
DNA Structure and Analysis Chapter 4: Background Molecular Biology Three main disciplines of biotechnology Biochemistry Genetics Molecular Biology # Biotechnology: A Laboratory Skills Course explorer.bio-rad.com
BIOLOGY 163 LABORATORY. RESTRICTION MAPPING OF PLASMID DNA (Revised Fall 2017)
 BIOLOGY 163 LABORATORY RESTRICTION MAPPING OF PLASMID DNA (Revised Fall 2017) Physical mapping of genomes is an important part of modern molecular genetics. As it's name implies, physical mapping seeks
BIOLOGY 163 LABORATORY RESTRICTION MAPPING OF PLASMID DNA (Revised Fall 2017) Physical mapping of genomes is an important part of modern molecular genetics. As it's name implies, physical mapping seeks
HiPer Random Amplification of Polymorphic DNA (RAPD) Teaching Kit
 HiPer Random Amplification of Polymorphic DNA (RAPD) Teaching Kit Product Code: HTBM031 Number of experiments that can be performed: 5 Duration of Experiment: Protocol: 3.5 hours Agarose Gel Electrophoresis:
HiPer Random Amplification of Polymorphic DNA (RAPD) Teaching Kit Product Code: HTBM031 Number of experiments that can be performed: 5 Duration of Experiment: Protocol: 3.5 hours Agarose Gel Electrophoresis:
Chapter 11. Restriction mapping. Objectives
 Restriction mapping Restriction endonucleases (REs) are part of bacterial defense systems. REs recognize and cleave specific sites in DNA molecules. REs are an indispensable tool in molecular biology for
Restriction mapping Restriction endonucleases (REs) are part of bacterial defense systems. REs recognize and cleave specific sites in DNA molecules. REs are an indispensable tool in molecular biology for
R1 12 kb R1 4 kb R1. R1 10 kb R1 2 kb R1 4 kb R1
 Bcor101 Sample questions Midterm 3 1. The maps of the sites for restriction enzyme EcoR1 (R1) in the wild type and mutated cystic fibrosis genes are shown below: Wild Type R1 12 kb R1 4 kb R1 _ _ CF probe
Bcor101 Sample questions Midterm 3 1. The maps of the sites for restriction enzyme EcoR1 (R1) in the wild type and mutated cystic fibrosis genes are shown below: Wild Type R1 12 kb R1 4 kb R1 _ _ CF probe
Sequence Analysis Lab Protocol
 Sequence Analysis Lab Protocol You will need this handout of instructions The sequence of your plasmid from the ABI The Accession number for Lambda DNA J02459 The Accession number for puc 18 is L09136
Sequence Analysis Lab Protocol You will need this handout of instructions The sequence of your plasmid from the ABI The Accession number for Lambda DNA J02459 The Accession number for puc 18 is L09136
Section A: Prokaryotes Types and Structure 1. What is microbiology?
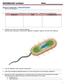 Section A: Prokaryotes Types and Structure 1. What is microbiology? 2. Compare and contrast characteristics of each bacterial type: Eubacteria and Archaebacteria. Eubacteria Both Archaebacteria 3. Label
Section A: Prokaryotes Types and Structure 1. What is microbiology? 2. Compare and contrast characteristics of each bacterial type: Eubacteria and Archaebacteria. Eubacteria Both Archaebacteria 3. Label
10 Restriction Analysis of Genomic DNA
 10 Restriction Analysis of Genomic DNA Objectives: A) To determine the rough location of restriction sites of an unknown restriction enzyme and B) to use this information to determine the identity of this
10 Restriction Analysis of Genomic DNA Objectives: A) To determine the rough location of restriction sites of an unknown restriction enzyme and B) to use this information to determine the identity of this
Supplemental Materials. DNA preparation. Dehalogenimonas lykanthroporepellens strain BL-DC-9 T (=ATCC
 Supplemental Materials DNA preparation. Dehalogenimonas lykanthroporepellens strain BL-DC-9 T (=ATCC BAA-1523 = JCM 15061) was grown in defined basal medium amended with 0.5 mm 1,1,2- trichloroethane (1,1,2-TCA)
Supplemental Materials DNA preparation. Dehalogenimonas lykanthroporepellens strain BL-DC-9 T (=ATCC BAA-1523 = JCM 15061) was grown in defined basal medium amended with 0.5 mm 1,1,2- trichloroethane (1,1,2-TCA)
Genomes summary. Bacterial genome sizes
 Genomes summary 1. >930 bacterial genomes sequenced. 2. Circular. Genes densely packed. 3. 2-10 Mbases, 470-7,000 genes 4. Genomes of >200 eukaryotes (45 higher ) sequenced. 5. Linear chromosomes 6. On
Genomes summary 1. >930 bacterial genomes sequenced. 2. Circular. Genes densely packed. 3. 2-10 Mbases, 470-7,000 genes 4. Genomes of >200 eukaryotes (45 higher ) sequenced. 5. Linear chromosomes 6. On
Supplementary Information
 Supplementary Information Deletion of the B-B and C-C regions of inverted terminal repeats reduces raav productivity but increases transgene expression Qingzhang Zhou 1, Wenhong Tian 2, Chunguo Liu 3,
Supplementary Information Deletion of the B-B and C-C regions of inverted terminal repeats reduces raav productivity but increases transgene expression Qingzhang Zhou 1, Wenhong Tian 2, Chunguo Liu 3,
This article reprinted from: Dooley, M. M Restriction endonuclease digestion of a plasmid.
 This article reprinted from: Dooley, M. M. 2008. Restriction endonuclease digestion of a plasmid. Pages 389-392, in Tested Studies for Laboratory Teaching, Volume 29 (K.L. Clase, Editor). Proceedings of
This article reprinted from: Dooley, M. M. 2008. Restriction endonuclease digestion of a plasmid. Pages 389-392, in Tested Studies for Laboratory Teaching, Volume 29 (K.L. Clase, Editor). Proceedings of
DNA Technology. B. Using Bacteria to Clone Genes: Overview:
 DNA Technology A. Basic Vocabulary: is DNA from 2 different sources that is combined. is the direct manipulation of genes for practical purposes. literally means or in a test tube or flask. is the manipulation
DNA Technology A. Basic Vocabulary: is DNA from 2 different sources that is combined. is the direct manipulation of genes for practical purposes. literally means or in a test tube or flask. is the manipulation
SYLLABUS: 53:154 ENVIRONMENTAL MICROBIOLOGY Assoc. Prof. Timothy E. Mattes Phone: Lectures: Mon/Wed/Fri 12:30 1:20 p.m.
 SYLLABUS: 53:154 ENVIRONMENTAL MICROBIOLOGY Assoc. Prof. Timothy E. Mattes Phone: 335-5065 Lectures: Mon/Wed/Fri 12:30 1:20 p.m. 3026 SC Labs: Wednesdays 3:30 5:30 p.m. 1246 SC Office Hours: Tues/Thurs
SYLLABUS: 53:154 ENVIRONMENTAL MICROBIOLOGY Assoc. Prof. Timothy E. Mattes Phone: 335-5065 Lectures: Mon/Wed/Fri 12:30 1:20 p.m. 3026 SC Labs: Wednesdays 3:30 5:30 p.m. 1246 SC Office Hours: Tues/Thurs
LECTURE TOPICS 3) DNA SEQUENCING, RNA SEQUENCING, DNA SYNTHESIS 5) RECOMBINANT DNA CONSTRUCTION AND GENE CLONING
 Page 1 of 25 Chapter 5 Notes Biochemistry 461 Fall 2010 CHAPTER 5, EXPLORING GENES: LECTURE TOPICS 1) RESTRICTION ENZYMES 2) GEL ELECTROPHORESIS OF DNA 3) DNA SEQUENCING, RNA SEQUENCING, DNA SYNTHESIS
Page 1 of 25 Chapter 5 Notes Biochemistry 461 Fall 2010 CHAPTER 5, EXPLORING GENES: LECTURE TOPICS 1) RESTRICTION ENZYMES 2) GEL ELECTROPHORESIS OF DNA 3) DNA SEQUENCING, RNA SEQUENCING, DNA SYNTHESIS
Supplemental Materials and Methods
 Supplemental Materials and Methods Proteins and reagents Proteins were purified as described previously: RecA, RecQ, and SSB proteins (Harmon and Kowalczykowski 1998); RecF protein (Morimatsu and Kowalczykowski
Supplemental Materials and Methods Proteins and reagents Proteins were purified as described previously: RecA, RecQ, and SSB proteins (Harmon and Kowalczykowski 1998); RecF protein (Morimatsu and Kowalczykowski
NAME TA SEC Problem Set 4 FRIDAY October 15, Answers to this problem set must be inserted into the box outside
 MIT Biology Department 7.012: Introductory Biology - Fall 2004 Instructors: Professor Eric Lander, Professor Robert A. Weinberg, Dr. Claudette Gardel NAME TA SEC 7.012 Problem Set 4 FRIDAY October 15,
MIT Biology Department 7.012: Introductory Biology - Fall 2004 Instructors: Professor Eric Lander, Professor Robert A. Weinberg, Dr. Claudette Gardel NAME TA SEC 7.012 Problem Set 4 FRIDAY October 15,
Chapter 15 Gene Technologies and Human Applications
 Chapter Outline Chapter 15 Gene Technologies and Human Applications Section 1: The Human Genome KEY IDEAS > Why is the Human Genome Project so important? > How do genomics and gene technologies affect
Chapter Outline Chapter 15 Gene Technologies and Human Applications Section 1: The Human Genome KEY IDEAS > Why is the Human Genome Project so important? > How do genomics and gene technologies affect
Bacteria. Bacteria. Chapter 27. Bacteria 7/18/2016
 Chapter 27 Prokaryotes Most numerous organisms on earth Earliest life forms (fossils: 2.5 billion years old) Contain ribosomes Surrounded by protective cell wall containing peptidoglycan (protein-carbohydrate)
Chapter 27 Prokaryotes Most numerous organisms on earth Earliest life forms (fossils: 2.5 billion years old) Contain ribosomes Surrounded by protective cell wall containing peptidoglycan (protein-carbohydrate)
Genomic Structure of Candida stellatoidea: Extra Chromosomes
 INFECTION AND IMMUNITY, Apr. 1990, p. 949-954 0019-9567/90/040949-06$02.00/0 Copyright C 1990, American Society for Microbiology Vol. 58, No. 4 Genomic Structure of Candida stellatoidea: Extra Chromosomes
INFECTION AND IMMUNITY, Apr. 1990, p. 949-954 0019-9567/90/040949-06$02.00/0 Copyright C 1990, American Society for Microbiology Vol. 58, No. 4 Genomic Structure of Candida stellatoidea: Extra Chromosomes
Non-Organic-Based Isolation of Mammalian microrna using Norgen s microrna Purification Kit
 Application Note 13 RNA Sample Preparation Non-Organic-Based Isolation of Mammalian microrna using Norgen s microrna Purification Kit B. Lam, PhD 1, P. Roberts, MSc 1 Y. Haj-Ahmad, M.Sc., Ph.D 1,2 1 Norgen
Application Note 13 RNA Sample Preparation Non-Organic-Based Isolation of Mammalian microrna using Norgen s microrna Purification Kit B. Lam, PhD 1, P. Roberts, MSc 1 Y. Haj-Ahmad, M.Sc., Ph.D 1,2 1 Norgen
Improved method for assembly of linear yeast expression cassettes using NEBuilder HiFi DNA Assembly Master Mix
 DNA CLONING DNA AMPLIFICATION & PCR Improved method for assembly of linear yeast expression cassettes using NEBuilder HiFi DNA Assembly Master Mix EPIGENETICS RNA ANALYSIS LIBRARY PREP FOR NEXT GEN SEQUENCING
DNA CLONING DNA AMPLIFICATION & PCR Improved method for assembly of linear yeast expression cassettes using NEBuilder HiFi DNA Assembly Master Mix EPIGENETICS RNA ANALYSIS LIBRARY PREP FOR NEXT GEN SEQUENCING
Genome Sequence Assembly
 Genome Sequence Assembly Learning Goals: Introduce the field of bioinformatics Familiarize the student with performing sequence alignments Understand the assembly process in genome sequencing Introduction:
Genome Sequence Assembly Learning Goals: Introduce the field of bioinformatics Familiarize the student with performing sequence alignments Understand the assembly process in genome sequencing Introduction:
Escherichia coli plasmids (Si nuclease/gel electrophoresis/electron microscope)
 Proc. Natl. Acad. Sci. USA Vol. 73, No. 7, pp. 2316-2320, July 1976 Biochemistry Isolation of inverted repeat sequences, including ISi, IS2, and IS3, in Escherichia coli plasmids (Si nuclease/gel electrophoresis/electron
Proc. Natl. Acad. Sci. USA Vol. 73, No. 7, pp. 2316-2320, July 1976 Biochemistry Isolation of inverted repeat sequences, including ISi, IS2, and IS3, in Escherichia coli plasmids (Si nuclease/gel electrophoresis/electron
Positively Charged Membrane
 BIOBOND NYLON MEMBRANES ProductInformation Technical Bulletin No. MB-570 June 1999 Size Quantity Positively Charged Membrane Neutral Membrane 30 cm x 3.5 m 1 roll N4781 N1031 30 cm x 12 m 1 roll N4906
BIOBOND NYLON MEMBRANES ProductInformation Technical Bulletin No. MB-570 June 1999 Size Quantity Positively Charged Membrane Neutral Membrane 30 cm x 3.5 m 1 roll N4781 N1031 30 cm x 12 m 1 roll N4906
Molecular Scissors: Lambda Digest Student Materials
 Molecular Scissors: Lambda Digest Student Materials Introduction 2 Pre-Lab Questions. 5 Lab Protocol 6 Data Collection Worksheet. 9 Post-Lab Questions and Analysis.. 10 Plasmid Maps. 13 Last updated: August
Molecular Scissors: Lambda Digest Student Materials Introduction 2 Pre-Lab Questions. 5 Lab Protocol 6 Data Collection Worksheet. 9 Post-Lab Questions and Analysis.. 10 Plasmid Maps. 13 Last updated: August
CHAPTER 9 DNA Technologies
 CHAPTER 9 DNA Technologies Recombinant DNA Artificially created DNA that combines sequences that do not occur together in the nature Basis of much of the modern molecular biology Molecular cloning of genes
CHAPTER 9 DNA Technologies Recombinant DNA Artificially created DNA that combines sequences that do not occur together in the nature Basis of much of the modern molecular biology Molecular cloning of genes
AP Biology Gene Expression/Biotechnology REVIEW
 AP Biology Gene Expression/Biotechnology REVIEW Multiple Choice Identify the choice that best completes the statement or answers the question. 1. Gene expression can be a. regulated before transcription.
AP Biology Gene Expression/Biotechnology REVIEW Multiple Choice Identify the choice that best completes the statement or answers the question. 1. Gene expression can be a. regulated before transcription.
Isolation of Candidatus Liberibacter Genes by RAPD and New PCR Detection Technique
 Fourteenth IOCV Conference, 2000 Short Communications Isolation of Candidatus Liberibacter Genes by RAPD and New PCR Detection Technique A. Hocquellet, J. M. Bové, and M. Garnier ABSTRACT. The RAPD technique
Fourteenth IOCV Conference, 2000 Short Communications Isolation of Candidatus Liberibacter Genes by RAPD and New PCR Detection Technique A. Hocquellet, J. M. Bové, and M. Garnier ABSTRACT. The RAPD technique
Pulsed Field Gel Electrophoresis Map It Out
 Pulsed Field Gel Electrophoresis Pulsed Field Gel Electrophoresis Map It Out The Direct Route to Your Large Molecule Applications PFGE Is Used in Many Research Areas Pulsed field gel electrophoresis (PFGE)
Pulsed Field Gel Electrophoresis Pulsed Field Gel Electrophoresis Map It Out The Direct Route to Your Large Molecule Applications PFGE Is Used in Many Research Areas Pulsed field gel electrophoresis (PFGE)
Concepts: What are RFLPs and how do they act like genetic marker loci?
 Restriction Fragment Length Polymorphisms (RFLPs) -1 Readings: Griffiths et al: 7th Edition: Ch. 12 pp. 384-386; Ch.13 pp404-407 8th Edition: pp. 364-366 Assigned Problems: 8th Ch. 11: 32, 34, 38-39 7th
Restriction Fragment Length Polymorphisms (RFLPs) -1 Readings: Griffiths et al: 7th Edition: Ch. 12 pp. 384-386; Ch.13 pp404-407 8th Edition: pp. 364-366 Assigned Problems: 8th Ch. 11: 32, 34, 38-39 7th
Functional vs Organismal views of Ecology. One organism: Population genetics Many organisms: Ecology No organisms: Ecosystems
 Functional vs Organismal views of Ecology One organism: Population genetics Many organisms: Ecology No organisms: Ecosystems The trade-off between precision and relevance Another trade-off exists: resolution
Functional vs Organismal views of Ecology One organism: Population genetics Many organisms: Ecology No organisms: Ecosystems The trade-off between precision and relevance Another trade-off exists: resolution
Regents Biology REVIEW 5: GENETICS
 Period Date REVIEW 5: GENETICS 1. Chromosomes: a. Humans have chromosomes, or homologous pairs. Homologous: b. Chromosome pairs carry genes for the same traits. Most organisms have two copies of the gene
Period Date REVIEW 5: GENETICS 1. Chromosomes: a. Humans have chromosomes, or homologous pairs. Homologous: b. Chromosome pairs carry genes for the same traits. Most organisms have two copies of the gene
3. human genomics clone genes associated with genetic disorders. 4. many projects generate ordered clones that cover genome
 Lectures 30 and 31 Genome analysis I. Genome analysis A. two general areas 1. structural 2. functional B. genome projects a status report 1. 1 st sequenced: several viral genomes 2. mitochondria and chloroplasts
Lectures 30 and 31 Genome analysis I. Genome analysis A. two general areas 1. structural 2. functional B. genome projects a status report 1. 1 st sequenced: several viral genomes 2. mitochondria and chloroplasts
Total Test Questions: 66 Levels: Grades Units of Credit: 1.0 STANDARD 2. Demonstrate appropriate use of personal protective devices.
 DESCRIPTION Biotechnology is designed to create an awareness of career possibilities in the field of biotechnology. Students are introduced to diagnostic and therapeutic laboratory procedures that support
DESCRIPTION Biotechnology is designed to create an awareness of career possibilities in the field of biotechnology. Students are introduced to diagnostic and therapeutic laboratory procedures that support
CHAPTER 21 LECTURE SLIDES
 CHAPTER 21 LECTURE SLIDES Prepared by Brenda Leady University of Toledo To run the animations you must be in Slideshow View. Use the buttons on the animation to play, pause, and turn audio/text on or off.
CHAPTER 21 LECTURE SLIDES Prepared by Brenda Leady University of Toledo To run the animations you must be in Slideshow View. Use the buttons on the animation to play, pause, and turn audio/text on or off.
Supplementary Material
 Supplementary Material Repressor CopG prevents access of RN polymerase to promoter and actively dissociates open complexes na M. Hernández-rriaga, Tania S. Rubio-Lepe, Manuel Espinosa and Gloria del Solar
Supplementary Material Repressor CopG prevents access of RN polymerase to promoter and actively dissociates open complexes na M. Hernández-rriaga, Tania S. Rubio-Lepe, Manuel Espinosa and Gloria del Solar
SuperScript IV Reverse Transcriptase as a better alternative to AMV-based enzymes
 WHITE PAPER SuperScript IV Reverse Transcriptase SuperScript IV Reverse Transcriptase as a better alternative to AMV-based enzymes Abstract Reverse transcriptases (RTs) from avian myeloblastosis virus
WHITE PAPER SuperScript IV Reverse Transcriptase SuperScript IV Reverse Transcriptase as a better alternative to AMV-based enzymes Abstract Reverse transcriptases (RTs) from avian myeloblastosis virus
DNA Replication in Prokaryotes and Eukaryotes
 DNA Replication in Prokaryotes and Eukaryotes 1. Overall mechanism 2. Roles of Polymerases & other proteins 3. More mechanism: Initiation and Termination 4. Mitochondrial DNA replication DNA replication
DNA Replication in Prokaryotes and Eukaryotes 1. Overall mechanism 2. Roles of Polymerases & other proteins 3. More mechanism: Initiation and Termination 4. Mitochondrial DNA replication DNA replication
High Pure PCR Template Preparation Kit for preparation of 100 nucleic acid samples Cat. No
 for preparation of 100 nucleic acid samples Cat. No. 1 796 88 Principle Cells are lysed during a short incubation with Proteinase K in the presence of a chaotropic salt (guanidine HCl), which immediately
for preparation of 100 nucleic acid samples Cat. No. 1 796 88 Principle Cells are lysed during a short incubation with Proteinase K in the presence of a chaotropic salt (guanidine HCl), which immediately
Chapter 9 Genetic Engineering
 Chapter 9 Genetic Engineering Biotechnology: use of microbes to make a protein product Recombinant DNA Technology: Insertion or modification of genes to produce desired proteins Genetic engineering: manipulation
Chapter 9 Genetic Engineering Biotechnology: use of microbes to make a protein product Recombinant DNA Technology: Insertion or modification of genes to produce desired proteins Genetic engineering: manipulation
Technician: Dionne Lutz, BS: Biology & MsED Office: Kanbar Center, Room 704, 41 Cooper Sq. (212) (office)
 BIO101: Molecular and Cellular Biology (WITH LABS!) Meeting Mondays, 6-9pm, in room 101 or in Kanbar Center on select dates (see schedule). (3 credits) Instructor: Oliver Medvedik, Ph.D Office: Room 206,
BIO101: Molecular and Cellular Biology (WITH LABS!) Meeting Mondays, 6-9pm, in room 101 or in Kanbar Center on select dates (see schedule). (3 credits) Instructor: Oliver Medvedik, Ph.D Office: Room 206,
The GeneEditor TM in vitro Mutagenesis System: Site- Directed Mutagenesis Using Altered Beta-Lactamase Specificity
 Promega Notes Magazine Number 62, 1997, p. 02 The GeneEditor TM in vitro Mutagenesis System: Site- Directed Mutagenesis Using Altered Beta-Lactamase Specificity By Christine Andrews and Scott Lesley Promega
Promega Notes Magazine Number 62, 1997, p. 02 The GeneEditor TM in vitro Mutagenesis System: Site- Directed Mutagenesis Using Altered Beta-Lactamase Specificity By Christine Andrews and Scott Lesley Promega
Construction of plant complementation vector and generation of transgenic plants
 MATERIAL S AND METHODS Plant materials and growth conditions Arabidopsis ecotype Columbia (Col0) was used for this study. SALK_072009, SALK_076309, and SALK_027645 were obtained from the Arabidopsis Biological
MATERIAL S AND METHODS Plant materials and growth conditions Arabidopsis ecotype Columbia (Col0) was used for this study. SALK_072009, SALK_076309, and SALK_027645 were obtained from the Arabidopsis Biological
Biology 252 Nucleic Acid Methods
 Fall 2015 Biology 252 Nucleic Acid Methods COURSE OUTLINE Prerequisites: One semester of college biology (BIO 101 or BIO 173) and one semester of college English (ENG 111); completion of CHM 111is recommended.
Fall 2015 Biology 252 Nucleic Acid Methods COURSE OUTLINE Prerequisites: One semester of college biology (BIO 101 or BIO 173) and one semester of college English (ENG 111); completion of CHM 111is recommended.
Bio 101 Sample questions: Chapter 10
 Bio 101 Sample questions: Chapter 10 1. Which of the following is NOT needed for DNA replication? A. nucleotides B. ribosomes C. Enzymes (like polymerases) D. DNA E. all of the above are needed 2 The information
Bio 101 Sample questions: Chapter 10 1. Which of the following is NOT needed for DNA replication? A. nucleotides B. ribosomes C. Enzymes (like polymerases) D. DNA E. all of the above are needed 2 The information
Site-specific ribosomal DNA insertion dements in Anopheles gambiae and A.anbiensis: nucleotide sequence of gene-element boundaries
 volume 17 Number 20 1989 Nucleic Acids Research Site-specific ribosomal DNA insertion dements in Anopheles gambiae and A.anbiensis: nucleotide sequence of gene-element boundaries Susan M.Paskewrtz* and
volume 17 Number 20 1989 Nucleic Acids Research Site-specific ribosomal DNA insertion dements in Anopheles gambiae and A.anbiensis: nucleotide sequence of gene-element boundaries Susan M.Paskewrtz* and
Sept 2. Structure and Organization of Genomes. Today: Genetic and Physical Mapping. Sept 9. Forward and Reverse Genetics. Genetic and Physical Mapping
 Sept 2. Structure and Organization of Genomes Today: Genetic and Physical Mapping Assignments: Gibson & Muse, pp.4-10 Brown, pp. 126-160 Olson et al., Science 245: 1434 New homework:due, before class,
Sept 2. Structure and Organization of Genomes Today: Genetic and Physical Mapping Assignments: Gibson & Muse, pp.4-10 Brown, pp. 126-160 Olson et al., Science 245: 1434 New homework:due, before class,
Pulse-Field gel-electrophoretic analysis of the amplification and copy-number stability of an integrational plasmid in Bacillus subtilis
 Appl Microbiol Biotechnol (1996) 46 :55 60 Springer-Verlag 1996 ORIGINAL PAPER C. Va zquez-cruz J. C. Ochoa-Sánchez G. Olmedo-Alvarez Pulse-Field gel-electrophoretic analysis of the amplification and copy-number
Appl Microbiol Biotechnol (1996) 46 :55 60 Springer-Verlag 1996 ORIGINAL PAPER C. Va zquez-cruz J. C. Ochoa-Sánchez G. Olmedo-Alvarez Pulse-Field gel-electrophoretic analysis of the amplification and copy-number
MOLECULAR GENETICS: TRANSFORMATION AND CLONING adapted by Dr. D. L. Vogelien
 Introduction MOLECULAR GENETICS: TRANSFORMATION AND CLONING adapted by Dr. D. L. Vogelien The field of molecular genetics has resulted in a number of practical applications that have been of tremendous
Introduction MOLECULAR GENETICS: TRANSFORMATION AND CLONING adapted by Dr. D. L. Vogelien The field of molecular genetics has resulted in a number of practical applications that have been of tremendous
Comparing the Agilent 2100 Bioanalyzer Performance to Traditional DNA Analysis Techniques
 Comparing the Agilent 2100 Bioanalyzer Performance to Traditional DNA Analysis Techniques Application Note Author Deborah Vitale Agilent Technologies, Inc. Palo Alto, CA, USA Abstract This Application
Comparing the Agilent 2100 Bioanalyzer Performance to Traditional DNA Analysis Techniques Application Note Author Deborah Vitale Agilent Technologies, Inc. Palo Alto, CA, USA Abstract This Application
BA, BSc, and MSc Degree Examinations
 Examination Candidate Number: Desk Number: BA, BSc, and MSc Degree Examinations 2017-8 Department : BIOLOGY Title of Exam: Genetics Time Allowed: 1 hour and 30 minutes Marking Scheme: Total marks available
Examination Candidate Number: Desk Number: BA, BSc, and MSc Degree Examinations 2017-8 Department : BIOLOGY Title of Exam: Genetics Time Allowed: 1 hour and 30 minutes Marking Scheme: Total marks available
Lecture 25 (11/15/17)
 Lecture 25 (11/15/17) Reading: Ch9; 328-332 Ch25; 990-995, 1005-1012 Problems: Ch9 (study-guide: applying); 1,2 Ch9 (study-guide: facts); 7,8 Ch25 (text); 1-3,5-7,9,10,13-15 Ch25 (study-guide: applying);
Lecture 25 (11/15/17) Reading: Ch9; 328-332 Ch25; 990-995, 1005-1012 Problems: Ch9 (study-guide: applying); 1,2 Ch9 (study-guide: facts); 7,8 Ch25 (text); 1-3,5-7,9,10,13-15 Ch25 (study-guide: applying);
Gel Electrophoresis: Quantitative length and mass measurements of DNA
 BIO440 Genetics Lab Humboldt State University Gel Electrophoresis: Quantitative length and mass measurements of DNA Electrophoresis, and in particular agarose gel electrophoresis, is an integral analysis
BIO440 Genetics Lab Humboldt State University Gel Electrophoresis: Quantitative length and mass measurements of DNA Electrophoresis, and in particular agarose gel electrophoresis, is an integral analysis
The right answer, every time
 MICROSEQ MICROBIAL IDENTIFICATION SYSTEM The right answer, every time Definitive identification of bacterial and fungal isolates. Accuracy adds confidence. When microbial contamination poses problems,
MICROSEQ MICROBIAL IDENTIFICATION SYSTEM The right answer, every time Definitive identification of bacterial and fungal isolates. Accuracy adds confidence. When microbial contamination poses problems,
Antisense RNA Insert Design for Plasmid Construction to Knockdown Target Gene Expression
 Vol. 1:7-15 Antisense RNA Insert Design for Plasmid Construction to Knockdown Target Gene Expression Ji, Tom, Lu, Aneka, Wu, Kaylee Department of Microbiology and Immunology, University of British Columbia
Vol. 1:7-15 Antisense RNA Insert Design for Plasmid Construction to Knockdown Target Gene Expression Ji, Tom, Lu, Aneka, Wu, Kaylee Department of Microbiology and Immunology, University of British Columbia
240EQ222 - Genetic Engineering
 Coordinating unit: Teaching unit: Academic year: Degree: ECTS credits: 2017 295 - EEBE - Barcelona East School of Engineering 713 - EQ - Department of Chemical Engineering MASTER'S DEGREE IN CHEMICAL ENGINEERING
Coordinating unit: Teaching unit: Academic year: Degree: ECTS credits: 2017 295 - EEBE - Barcelona East School of Engineering 713 - EQ - Department of Chemical Engineering MASTER'S DEGREE IN CHEMICAL ENGINEERING
Enzymatic assembly of DNA molecules up to several hundred kilobases
 nature methods Enzymatic assembly of DNA molecules up to several hundred kilobases Daniel G Gibson, Lei Young, Ray-Yuan Chuang, J Craig Venter, Clyde A Hutchison III & Hamilton O Smith Supplementary figures
nature methods Enzymatic assembly of DNA molecules up to several hundred kilobases Daniel G Gibson, Lei Young, Ray-Yuan Chuang, J Craig Venter, Clyde A Hutchison III & Hamilton O Smith Supplementary figures
DNA replication: Enzymes link the aligned nucleotides by phosphodiester bonds to form a continuous strand.
 DNA replication: Copying genetic information for transmission to the next generation Occurs in S phase of cell cycle Process of DNA duplicating itself Begins with the unwinding of the double helix to expose
DNA replication: Copying genetic information for transmission to the next generation Occurs in S phase of cell cycle Process of DNA duplicating itself Begins with the unwinding of the double helix to expose
PULSE FIELD GEL ELECTROPHORESIS FOR COMPARATIVE STUDIES OF BACTERIA DIVERSITY
 PULSE FIELD GEL ELECTROPHORESIS FOR COMPARATIVE STUDIES OF BACTERIA DIVERSITY ROSABEL BONG Department of Microbiology and Immunology, UBC The technique of pulse-field gel electrophoresis (PFGE) makes use
PULSE FIELD GEL ELECTROPHORESIS FOR COMPARATIVE STUDIES OF BACTERIA DIVERSITY ROSABEL BONG Department of Microbiology and Immunology, UBC The technique of pulse-field gel electrophoresis (PFGE) makes use
RFLP: Restriction Fragment Length Polymorphism
 RFLP: Restriction Fragment Length Polymorphism Various endonucleases: 6 cutters and 4 cutters Enzyme Source Recognition Sequence Cut EcoRI Escherichia coli 5'GAATTC 5'---G/AATTC---3' EcoRII Escherichia
RFLP: Restriction Fragment Length Polymorphism Various endonucleases: 6 cutters and 4 cutters Enzyme Source Recognition Sequence Cut EcoRI Escherichia coli 5'GAATTC 5'---G/AATTC---3' EcoRII Escherichia
1. A brief overview of sequencing biochemistry
 Supplementary reading materials on Genome sequencing (optional) The materials are from Mark Blaxter s lecture notes on Sequencing strategies and Primary Analysis 1. A brief overview of sequencing biochemistry
Supplementary reading materials on Genome sequencing (optional) The materials are from Mark Blaxter s lecture notes on Sequencing strategies and Primary Analysis 1. A brief overview of sequencing biochemistry
Introduction to Bioinformatics and Gene Expression Technologies
 Introduction to Bioinformatics and Gene Expression Technologies Utah State University Fall 2017 Statistical Bioinformatics (Biomedical Big Data) Notes 1 1 Vocabulary Gene: hereditary DNA sequence at a
Introduction to Bioinformatics and Gene Expression Technologies Utah State University Fall 2017 Statistical Bioinformatics (Biomedical Big Data) Notes 1 1 Vocabulary Gene: hereditary DNA sequence at a
Chapter 20: Biotechnology
 Name Period The AP Biology exam has reached into this chapter for essay questions on a regular basis over the past 15 years. Student responses show that biotechnology is a difficult topic. This chapter
Name Period The AP Biology exam has reached into this chapter for essay questions on a regular basis over the past 15 years. Student responses show that biotechnology is a difficult topic. This chapter
Biology 142 Advanced Topics in Genetics and Molecular Biology Course Syllabus Spring 2006
 Biology 142 Advanced Topics in Genetics and Molecular Biology Course Syllabus Spring 2006 Faculty Information: Dr. Nitya Jacob, Office: Room 104, Pierce Hall; Phone: 770-784-8346 Office Hours: T 9:30-10:30
Biology 142 Advanced Topics in Genetics and Molecular Biology Course Syllabus Spring 2006 Faculty Information: Dr. Nitya Jacob, Office: Room 104, Pierce Hall; Phone: 770-784-8346 Office Hours: T 9:30-10:30
J. Eco. Heal. Env. 4, No. 1, 1-5 (2016) 1
 J. Eco. Heal. Env. 4, No. 1, 1-5 (2016) 1 Journal of Ecology of Health & Environment An International Journal http://dx.doi.org/10.18576/jehe/040101 Characterization and Molecular Identification of Unknown
J. Eco. Heal. Env. 4, No. 1, 1-5 (2016) 1 Journal of Ecology of Health & Environment An International Journal http://dx.doi.org/10.18576/jehe/040101 Characterization and Molecular Identification of Unknown
Karyotypes of Saccharomyces sensu lato species
 International Journal of Systematic Bacteriology (1999), 49, 1925 1931 Printed in Great Britain Karyotypes of Saccharomyces sensu lato species Randi Føns Petersen, 1,2 Torsten Nilsson-Tillgren 2 and Jure
International Journal of Systematic Bacteriology (1999), 49, 1925 1931 Printed in Great Britain Karyotypes of Saccharomyces sensu lato species Randi Føns Petersen, 1,2 Torsten Nilsson-Tillgren 2 and Jure
DNA Technology. Asilomar Singer, Zinder, Brenner, Berg
 DNA Technology Asilomar 1973. Singer, Zinder, Brenner, Berg DNA Technology The following are some of the most important molecular methods we will be using in this course. They will be used, among other
DNA Technology Asilomar 1973. Singer, Zinder, Brenner, Berg DNA Technology The following are some of the most important molecular methods we will be using in this course. They will be used, among other
