ChIP-seq analysis. adapted from J. van Helden, M. Defrance, C. Herrmann, D. Puthier, N. Servant
|
|
|
- Horatio Chapman
- 6 years ago
- Views:
Transcription
1 ChIP-seq analysis adapted from J. van Helden, M. Defrance, C. Herrmann, D. Puthier, N. Servant
2 A model of transcriptional regulation
3 Chromatin constraints Each diploid cell contains about 2 meters of DNA High level of compaction required Accessibility required Replication Transcription DNA repair Specific machinery required
4 Chromatin has highly complex structure with several levels of organization Genetics: A Conceptual Approach, 2nd ed.
5 Beads on a string Figure 4: Chromatin fibers purified from chicken erythrocytes. Each nucleosome (~12-15 nm) is well resolved, along with the linker DNA between the nucleosomes. Given the resolution, other components, if present, such as a transcribing RNA polymerase or transcription factor complexes, should be resolvable
6 Histones and nucleosomes Histones Small proteins (11-22 kda) Highly conserved Basic (Arginine et Lysine) N-terminal tails subject to post translational modification Nucleosome Octamers of histone (H2A,H2B,H3,H4) x 2 146bp DNA
7 Nucleosome structure
8 Histone post translational modification Lysine acetylation Lysine methylation Arginine methylation Serine phosphorylation Threonine phosphorylation ADP-ribosylation Ubiquitylation Sumoylation...
9 Some alternative modifications
10 The Brno nomenclature The nomenclature set out here was devised following the first meeting of the Epigenome Network of Excellence (NoE), at the Mendel Abbey in Brno, Czech Republic. For this reason, it can be referred to as the Brno nomenclature.
11 Epigenetic Epigenetics involves genetic control by factors other than an individual's DNA sequence Histone modifications DNA methylation Epigenetic modifications may be inherited mitotically or meiotically
12 Chromatine immuno-precipitation (ChIP) Used for: TF localization Histone modifications
13 ChIP-Seq: technical considerations Quality of antibodies: one of the most important factors ('ChIP grade') High sensitivity High specificity The specificity of an antibody can be directly addressed by immunoblot analysis (knockdown by RNA-mediated interference or genetic knockout) Polyclonal antibodies may be prefered Fivefold enrichment by ChIP-PCR at several positive-control regions Offer the flexibility of the recognition of multiple epitopes Cell Number Typically (e.g, RNA polymerase II/histone modifications) (less-abundant proteins)
14 ChIP-Seq: technical considerations Open chromatin regions are easier to shear Higher background signals Two solutions Isotype control antibodies Immunoprecipitate much less DNA than specific antibodies Overamplification of particular genomic regions during the library construction step (PCR) Input Non-ChIP genomic DNA Better control
15 ChIP-seq signal for transcription factors ChIP seq on DNA binding TF read densities on +/- strand We expect to see a typical strand asymmetry in read densities ChIP peak recognition pattern
16 ChIP-seq signal for transcription factors treatment read density (=WIG) input read density (=WIG) peak (=BED) aligned reads + strand (=BAM) aligned reads - strand (=BAM) (this is the data you are going to manipulate...)
17 ChIP-seq signal for transcription factors ChIP seq on DNA binding TF read densities on +/- strand Binding of several TF as complexes tend to blur this asymmetry
18 ChIP-seq signal for histone marks ChIP seq on histone modifications read densities on +/- strand The strand asymmetry is completely lost when considering ChIP datasets for diffuse histone modifications
19 Real example of ChIP-seq signal ESR1 input H3K4me1 ESR1 reads H3K4me1 reads
20 Keys aspects of peak finding Treating the reads Modelling noise levels Scaling datasets Detecting enriched/peak regions Dealing with replicates
21 From aligned reads to binding sites Tag shifting vs. extension positive/negative strand read peaks do not represent the true location of the binding site reads can be shifted by d/2 where d is the band size (MACS) increased resolution reads can be elongated to a size of d (FindPeaks, PeakSeq,...) d can be estimate from the data (MACS) or given as input parameter example of MACS model building using top enriched regions
22 From aligned reads to binding sites d/2 Tag shifting shifted position initial position read densities on +/- strand Each tag is shifted by d/2 (i.e. towards the middle of the IP fragment) where d represent the fragment length
23 From aligned reads to binding sites Tag elongation read densities on +/- strand Each tag is computationaly extended in 3' to a total length of d
24 Modelling noise levels ChIP-seq dataset (=treatment) = signal + background noise How do we estimate the noise?
25 Modelling noise levels noise is not uniform (chromatin conformation, local biases, mappability) input dataset is mandatory for reliable local estimation! (although some algorithms do not require it :-( ) chr1:114,720, ,746, kb treatment input?
26 Modelling noise levels random distribution of reads in a window of size w modelled using a theoretical distribution Poisson distribution 1 parameter : λ = expected number of reads in window k P( X =k )=e λk k!
27 Scaling unequal datasets treatment (=signal + noise) and input (=noise) datasets generally do not have the same sequencing depth need for normalization input dataset should model the noise level in the treatment dataset naïve approach : upscale/downscale the smaller/larger dataset Input : N reads ChIP-seq dataset M > N reads scale by library size : M M' = N Problem : signal influences scaling factor More signal (but equal noise) artificial noise over-estimation
28 Scaling unequal datasets by library size input 1 area ~ number of reads = treatment 1 area ~ number of reads = = 18 Scaling by library size : upscale input by 18/10 = treatment 10 estimated noise level 1 Noise level is over-estimated! 10
29 Scaling unequal datasets by library size input 1 area ~ number of reads = treatment 1 area ~ number of reads = = 23 Scaling by library size : upscale input by 23/10 = treatment 10 estimated noise level 1 10
30 Scaling unequal datasets by library size input 1 area ~ number of reads = treatment 1 area ~ number of reads = = 23 Scaling by library size : upscale input by 23/10 = treatment 10 estimated noise level 1 10
31 Scaling unequal datasets more advanced : linear regression by exclusing peak regions (PeakSeq) read counts in 1Mb regions in input and treatment all regions excluding enriched (=signal) regions
32 Defining peaks Determining enriched regions sliding window across the genome At each location, evaluate the enrichment of the Signal vs background based on Poisson distribution retain regions with P-values below threshold evaluate FDR Pval < 1e-20 Pval ~ 0.6
33 MACS [Zhang et al. Genome Biol. 2008] Step 1 : estimating fragment length d slide a window of size BANDWIDTH retain top regions with MFOLD enrichment of treatment vs. input plot average +/- strand read densities estimate d enrichment > MFOLD treatment control
34 MACS [Zhang et al. Genome Biol. 2008] Step 2 : identification of local noise parameter slide a window of size 2*d across treatment and input estimate parameter λlocal of Poisson distribution 1 kb 5 kb 10 kb full genome estimate λ over diff. range take the max
35 MACS [Zhang et al. Genome Biol. 2008] Step 3 : identification of enriched/peak regions determine regions with P-values < PVALUE determine summit position inside enriched regions as max density P-val = 1e-30
36 MACS [Zhang et al. Genome Biol. 2008] Step 4 : estimating FDR positive peaks (P-values) swap treatment and input; call negative peaks (P-value) FDR(p) = # negative peaks with Pval < p # positive peaks with Pval < p increasing P-value FDR = 2/25=0.08
37 Peak-Calling: WTD Window Tag Density (SPP package) pd pd= positive downstream pu= positive upstream pu nd = negative downstream nd nu nu = negative upstream
38 Peak-Calling: MTC Mirror Tag Correlation (SPP package) Strand cross-correlation profile
39
40 Histone modification profiles
41 DNase-Seq
42 Nucleosome positioning The consensus distribution of nucleosomes (grey ovals) around all yeast genes is shown, aligned by the beginning and end of every gene. The resulting two plots were fused in the genic region. The peaks and valleys represent similar positioning relative to the transcription start site (TSS). The arrow under the green circle near the 5' nucleosome-free region (NFR) represents the TSS. The green -blue shading in the plot represents the transitions observed in nucleosome composition and phasing (green represents high H2A.Z levels, acetylation, H3K4 methylation and phasing, whereas blue represents low levels of these modifications). The red circle indicates transcriptional termination within the 3' NFR. Figure is reproduced, with permission, from ref. 20 (2008) Cold Spring Harbor Laboratory Press.
43 Data processing & file formats
44 Fastq file format Header Sequence + (optional header) Quality (default HWUSI EAS1691:3:1:17036:13000#0/1 PF=0 length=36 GGGGGTCATCATCATTTGATCTGGGAAAGGCTACTG + HWUSI EAS1691:3:1:17257:12994#0/1 PF=1 length=36 TGTACAACAACAACCTGAATGGCATACTGGTTGCTG + DDDD<BDBDB??BB*DD:D#################
45 Solid output Read sequence in color (csfasta) >1831_573_1004_F3 T >1831_573_1567_F3 T Quality scores (qual) >1831_573_1004_F >1831_573_1567_F
46 Solid output in fastq T _573_1004 T _573_1004
47 Illumina sequence identifiers Sequences from the Illumina software use a systematic HWI EAS434:4:1:1:1701 length=36 NAATCGGAAATTTTATTTGTTCAGTACACCAAATAG +SRR sra.2 HWI EAS434:4:1:1:1701 length=36!0<<;:::<<<<<<<<<<<<<<;;;<<<<<<<<;76 HWI EAS Unique instrument name Flowcell lane Tile number within the flow cell 'x'-coordinate of the cluster within the tile 'y'-coordinate #0 Index number for a multiplexed sample (opt.) /1 /1 or /2 for paired-end and maite-pair sequencing (opt.)
48 Sanger quality score Sanger quality score (Phred quality score): Measure the quality of each base call Based on p, the probality of error (the probability that the corresponding base call is incorrect) Qsanger= -10*log10(p) p = 0.01 <=> Qsanger 20 Quality score are in ASCII 33 Note that SRA has adopted Sanger quality score although original fastq files may use different quality score (see:
49 ASCII 33 Storing PHRED scores as single characters gave a simple and space efficient encoding: Character! means a quality of 0 Range 0-40
50 Quality control for high throughput sequence data FastQC GUI / command line ShortRead Bioconductor package
51 Trimming Essential step (at least when using bowtie) Almost mandatory when using tophat FASTX-Toolkit Sickle ShortRead Window-based trimming (unpublished) Bioconductor package csfasta_quality_filter.pl SOLiD Mean quality Continuous run of bad colors at the end of the read
52 Quality control with FastQC Quality Position in read
53 Quality control with FastQC Position in read
54 Quality control with FastQC Nb Reads Mean Phred Score
55 Mapping reads to genome: general softwares a Work well for Sanger and 454 reads, allowing gaps and clipping. b Paired end mapping. c Make use of base quality in alignment.dbwa trims the primer base and the first color for a color read. e Long-read alignment implemented in the BWA-SW module. fmaq only does gapped alignment for Illumina paired-end reads. g Free executable for non-profit projects only.
56 Bowtie principle Use highly efficient compressing and mapping algorithms based on Burrows Wheeler Transform (BWT) The Burrows-Wheeler Transform of a text T, BWT(T), can be constructed as follows. The character $ is appended to T, where $ is a character not in T that is lexicographically less than all characters in T. The Burrows-Wheeler Matrix of T, BWM(T), is obtained by computing the matrix whose rows comprise all cyclic rotations of T sorted lexicographically. T acaacg$ acaacg$ caacg$a aacg$ac acg$aca cg$acaa g$acaac $acaacg $acaacg aacg$ac acaacg$ acg$aca caacg$a cg$acaa g$acaac BWT (T) gc$aaac
57 Bowtie principle Burrows-Wheeler Matrices have a property called the Last First (LF) Mapping. The ith occurrence of character c in the last column corresponds to the same text character as the ith occurrence of c in the first column. Example: searching AAC in ACAACG
58 Storing alignment: SAM Format Store information related to alignement Read ID CIGAR String Bitwise FLAG read paired read mapped in proper pair read unmapped,... Alignment position Mapping quality...
59 The extended CIGAR string Exemple flags: M alignment match (can be a sequence match or mismatch) I insertion to the reference D deletion from the reference ATTCAGATGCAGTA ATTCA--TGCAGTA 5M2D7M
60 Mapping reads Main Issues: Number of multi hits PCR duplicates Issue with short reads mappability Warning with ChIP-Seq (library complexity) Number of allowed mismatches Depend on sequence size (sometimes heterogeneous length) Depend of the aligner
61 Mappability Sequence uniqueness of the reference These tracks display the level of sequence uniqueness of the reference NCBI36/hg18 genome assembly. They were generated using different window sizes, and high signal will be found in areas where the sequence is unique.
62 Compressing and indexing files Needed before visualization in Genome Browser samtools view output.bam # output SAM format [u@m] samtools sort output.bam output.sorted [u@m] samtools index output.sorted.bam Or use Galaxy or IGVtools
63 Sequence read Archive (SRA) The SRA archives high-throughput sequencing data that are associated with: RNA-Seq, ChIP-Seq, and epigenomic data that are submitted to GEO
64 SRA growth
65 SRA Concepts Data submitted to SRA is organized using a metadata model consisting of six objects: Study A set of experiments with an overall goal and literature references. Experiment An experiment is a consistent set of laboratory operations on input material with an expected result. Sample An experiment targets one or more samples. Results are expressed in terms of individual samples or bundles of samples as defined by the experiment. Run Results are called runs. Runs comprise the data gathered for a sample or sample bundle and refer to a defining experiment.
66 Getting fastq files using SRA toolkit *.sra to fastq conversion Fastq-dump fastq dump A SRRxxxx.sra Note: use split-files argument for paired-end library
67 Merci
Introduction to transcriptome analysis using High Throughput Sequencing technologies. D. Puthier 2012
 Introduction to transcriptome analysis using High Throughput Sequencing technologies D. Puthier 2012 A typical RNA-Seq experiment Library construction Protocol variations Fragmentation methods RNA: nebulization,
Introduction to transcriptome analysis using High Throughput Sequencing technologies D. Puthier 2012 A typical RNA-Seq experiment Library construction Protocol variations Fragmentation methods RNA: nebulization,
Introduction to transcriptome analysis using High Throughput Sequencing technologies. D. Puthier 2012
 Introduction to transcriptome analysis using High Throughput Sequencing technologies D. Puthier 2012 Transcriptome: the old school Cyanine 5 (Cy5) Cy-3: - Excitation 550nm - Emission 570nm Cy-5: - Excitation
Introduction to transcriptome analysis using High Throughput Sequencing technologies D. Puthier 2012 Transcriptome: the old school Cyanine 5 (Cy5) Cy-3: - Excitation 550nm - Emission 570nm Cy-5: - Excitation
ChIP-seq data analysis with Chipster. Eija Korpelainen CSC IT Center for Science, Finland
 ChIP-seq data analysis with Chipster Eija Korpelainen CSC IT Center for Science, Finland chipster@csc.fi What will I learn? Short introduction to ChIP-seq Analyzing ChIP-seq data Central concepts Analysis
ChIP-seq data analysis with Chipster Eija Korpelainen CSC IT Center for Science, Finland chipster@csc.fi What will I learn? Short introduction to ChIP-seq Analyzing ChIP-seq data Central concepts Analysis
ChIP-seq analysis 2/28/2018
 ChIP-seq analysis 2/28/2018 Acknowledgements Much of the content of this lecture is from: Furey (2012) ChIP-seq and beyond Park (2009) ChIP-seq advantages + challenges Landt et al. (2012) ChIP-seq guidelines
ChIP-seq analysis 2/28/2018 Acknowledgements Much of the content of this lecture is from: Furey (2012) ChIP-seq and beyond Park (2009) ChIP-seq advantages + challenges Landt et al. (2012) ChIP-seq guidelines
Genomic DNA ASSEMBLY BY REMAPPING. Course overview
 ASSEMBLY BY REMAPPING Laurent Falquet, The Bioinformatics Unravelling Group, UNIFR & SIB MA/MER @ UniFr Group Leader @ SIB Course overview Genomic DNA PacBio Illumina methylation de novo remapping Annotation
ASSEMBLY BY REMAPPING Laurent Falquet, The Bioinformatics Unravelling Group, UNIFR & SIB MA/MER @ UniFr Group Leader @ SIB Course overview Genomic DNA PacBio Illumina methylation de novo remapping Annotation
Charles Girardot, Furlong Lab. MACS, CisGenome, SISSRs and other peak calling algorithms: differences and practical use
 Charles Girardot, Furlong Lab MACS, CisGenome, SISSRs and other peak calling algorithms: differences and practical use ChIP-Seq signal properties Only 5 ends of ChIPed fragments are sequenced Shifted read
Charles Girardot, Furlong Lab MACS, CisGenome, SISSRs and other peak calling algorithms: differences and practical use ChIP-Seq signal properties Only 5 ends of ChIPed fragments are sequenced Shifted read
ChIP. November 21, 2017
 ChIP November 21, 2017 functional signals: is DNA enough? what is the smallest number of letters used by a written language? DNA is only one part of the functional genome DNA is heavily bound by proteins,
ChIP November 21, 2017 functional signals: is DNA enough? what is the smallest number of letters used by a written language? DNA is only one part of the functional genome DNA is heavily bound by proteins,
Applications of short-read
 Applications of short-read sequencing: RNA-Seq and ChIP-Seq BaRC Hot Topics March 2013 George Bell, Ph.D. http://jura.wi.mit.edu/bio/education/hot_topics/ Sequencing applications RNA-Seq includes experiments
Applications of short-read sequencing: RNA-Seq and ChIP-Seq BaRC Hot Topics March 2013 George Bell, Ph.D. http://jura.wi.mit.edu/bio/education/hot_topics/ Sequencing applications RNA-Seq includes experiments
RNAseq Applications in Genome Studies. Alexander Kanapin, PhD Wellcome Trust Centre for Human Genetics, University of Oxford
 RNAseq Applications in Genome Studies Alexander Kanapin, PhD Wellcome Trust Centre for Human Genetics, University of Oxford RNAseq Protocols Next generation sequencing protocol cdna, not RNA sequencing
RNAseq Applications in Genome Studies Alexander Kanapin, PhD Wellcome Trust Centre for Human Genetics, University of Oxford RNAseq Protocols Next generation sequencing protocol cdna, not RNA sequencing
NEXT GENERATION SEQUENCING. Farhat Habib
 NEXT GENERATION SEQUENCING HISTORY HISTORY Sanger Dominant for last ~30 years 1000bp longest read Based on primers so not good for repetitive or SNPs sites HISTORY Sanger Dominant for last ~30 years 1000bp
NEXT GENERATION SEQUENCING HISTORY HISTORY Sanger Dominant for last ~30 years 1000bp longest read Based on primers so not good for repetitive or SNPs sites HISTORY Sanger Dominant for last ~30 years 1000bp
Sequencing applications. Today's outline. Hands-on exercises. Applications of short-read sequencing: RNA-Seq and ChIP-Seq
 Sequencing applications Applications of short-read sequencing: RNA-Seq and ChIP-Seq BaRC Hot Topics March 2013 George Bell, Ph.D. http://jura.wi.mit.edu/bio/education/hot_topics/ RNA-Seq includes experiments
Sequencing applications Applications of short-read sequencing: RNA-Seq and ChIP-Seq BaRC Hot Topics March 2013 George Bell, Ph.D. http://jura.wi.mit.edu/bio/education/hot_topics/ RNA-Seq includes experiments
Mapping Next Generation Sequence Reads. Bingbing Yuan Dec. 2, 2010
 Mapping Next Generation Sequence Reads Bingbing Yuan Dec. 2, 2010 1 What happen if reads are not mapped properly? Some data won t be used, thus fewer reads would be aligned. Reads are mapped to the wrong
Mapping Next Generation Sequence Reads Bingbing Yuan Dec. 2, 2010 1 What happen if reads are not mapped properly? Some data won t be used, thus fewer reads would be aligned. Reads are mapped to the wrong
NGS Approaches to Epigenomics
 I519 Introduction to Bioinformatics, 2013 NGS Approaches to Epigenomics Yuzhen Ye (yye@indiana.edu) School of Informatics & Computing, IUB Contents Background: chromatin structure & DNA methylation Epigenomic
I519 Introduction to Bioinformatics, 2013 NGS Approaches to Epigenomics Yuzhen Ye (yye@indiana.edu) School of Informatics & Computing, IUB Contents Background: chromatin structure & DNA methylation Epigenomic
DATA FORMATS AND QUALITY CONTROL
 HTS Summer School 12-16th September 2016 DATA FORMATS AND QUALITY CONTROL Romina Petersen, University of Cambridge (rp520@medschl.cam.ac.uk) Luigi Grassi, University of Cambridge (lg490@medschl.cam.ac.uk)
HTS Summer School 12-16th September 2016 DATA FORMATS AND QUALITY CONTROL Romina Petersen, University of Cambridge (rp520@medschl.cam.ac.uk) Luigi Grassi, University of Cambridge (lg490@medschl.cam.ac.uk)
Introduction to Next Generation Sequencing
 The Sequencing Revolution Introduction to Next Generation Sequencing Dena Leshkowitz,WIS 1 st BIOmics Workshop High throughput Short Read Sequencing Technologies Highly parallel reactions (millions to
The Sequencing Revolution Introduction to Next Generation Sequencing Dena Leshkowitz,WIS 1 st BIOmics Workshop High throughput Short Read Sequencing Technologies Highly parallel reactions (millions to
Introduction to ChIP Seq data analyses. Acknowledgement: slides taken from Dr. H
 Introduction to ChIP Seq data analyses Acknowledgement: slides taken from Dr. H Wu @Emory ChIP seq: Chromatin ImmunoPrecipitation it ti + sequencing Same biological motivation as ChIP chip: measure specific
Introduction to ChIP Seq data analyses Acknowledgement: slides taken from Dr. H Wu @Emory ChIP seq: Chromatin ImmunoPrecipitation it ti + sequencing Same biological motivation as ChIP chip: measure specific
ChIP-seq and RNA-seq. Farhat Habib
 ChIP-seq and RNA-seq Farhat Habib fhabib@iiserpune.ac.in Biological Goals Learn how genomes encode the diverse patterns of gene expression that define each cell type and state. Protein-DNA interactions
ChIP-seq and RNA-seq Farhat Habib fhabib@iiserpune.ac.in Biological Goals Learn how genomes encode the diverse patterns of gene expression that define each cell type and state. Protein-DNA interactions
Introduction to NGS analyses
 Introduction to NGS analyses Giorgio L Papadopoulos Institute of Molecular Biology and Biotechnology Bioinformatics Support Group 04/12/2015 Papadopoulos GL (IMBB, FORTH) IMBB NGS Seminar 04/12/2015 1
Introduction to NGS analyses Giorgio L Papadopoulos Institute of Molecular Biology and Biotechnology Bioinformatics Support Group 04/12/2015 Papadopoulos GL (IMBB, FORTH) IMBB NGS Seminar 04/12/2015 1
Galaxy Platform For NGS Data Analyses
 Galaxy Platform For NGS Data Analyses Weihong Yan wyan@chem.ucla.edu Collaboratory Web Site http://qcb.ucla.edu/collaboratory http://collaboratory.lifesci.ucla.edu Workshop Outline ü Day 1 UCLA galaxy
Galaxy Platform For NGS Data Analyses Weihong Yan wyan@chem.ucla.edu Collaboratory Web Site http://qcb.ucla.edu/collaboratory http://collaboratory.lifesci.ucla.edu Workshop Outline ü Day 1 UCLA galaxy
The ChIP-Seq project. Giovanna Ambrosini, Philipp Bucher. April 19, 2010 Lausanne. EPFL-SV Bucher Group
 The ChIP-Seq project Giovanna Ambrosini, Philipp Bucher EPFL-SV Bucher Group April 19, 2010 Lausanne Overview Focus on technical aspects Description of applications (C programs) Where to find binaries,
The ChIP-Seq project Giovanna Ambrosini, Philipp Bucher EPFL-SV Bucher Group April 19, 2010 Lausanne Overview Focus on technical aspects Description of applications (C programs) Where to find binaries,
C3BI. VARIANTS CALLING November Pierre Lechat Stéphane Descorps-Declère
 C3BI VARIANTS CALLING November 2016 Pierre Lechat Stéphane Descorps-Declère General Workflow (GATK) software websites software bwa picard samtools GATK IGV tablet vcftools website http://bio-bwa.sourceforge.net/
C3BI VARIANTS CALLING November 2016 Pierre Lechat Stéphane Descorps-Declère General Workflow (GATK) software websites software bwa picard samtools GATK IGV tablet vcftools website http://bio-bwa.sourceforge.net/
ChIP-seq and RNA-seq
 ChIP-seq and RNA-seq Biological Goals Learn how genomes encode the diverse patterns of gene expression that define each cell type and state. Protein-DNA interactions (ChIPchromatin immunoprecipitation)
ChIP-seq and RNA-seq Biological Goals Learn how genomes encode the diverse patterns of gene expression that define each cell type and state. Protein-DNA interactions (ChIPchromatin immunoprecipitation)
Introduction to genome biology
 Introduction to genome biology Lisa Stubbs Deep transcritpomes for traditional model species from ENCODE (and modencode) Deep RNA-seq and chromatin analysis on 147 human cell types, as well as tissues,
Introduction to genome biology Lisa Stubbs Deep transcritpomes for traditional model species from ENCODE (and modencode) Deep RNA-seq and chromatin analysis on 147 human cell types, as well as tissues,
Introduction to genome biology
 Introduction to genome biology Lisa Stubbs We ve found most genes; but what about the rest of the genome? Genome size* 12 Mb 95 Mb 170 Mb 1500 Mb 2700 Mb 3200 Mb #coding genes ~7000 ~20000 ~14000 ~26000
Introduction to genome biology Lisa Stubbs We ve found most genes; but what about the rest of the genome? Genome size* 12 Mb 95 Mb 170 Mb 1500 Mb 2700 Mb 3200 Mb #coding genes ~7000 ~20000 ~14000 ~26000
Francisco García Quality Control for NGS Raw Data
 Contents Data formats Sequence capture Fasta and fastq formats Sequence quality encoding Quality Control Evaluation of sequence quality Quality control tools Identification of artifacts & filtering Practical
Contents Data formats Sequence capture Fasta and fastq formats Sequence quality encoding Quality Control Evaluation of sequence quality Quality control tools Identification of artifacts & filtering Practical
ChIP-Seq Data Analysis. J Fass UCD Genome Center Bioinformatics Core Wednesday December 17, 2014
 ChIP-Seq Data Analysis J Fass UCD Genome Center Bioinformatics Core Wednesday December 17, 2014 What s the Question? Where do Transcription Factors (TFs) bind genomic DNA 1? (Where do other things bind
ChIP-Seq Data Analysis J Fass UCD Genome Center Bioinformatics Core Wednesday December 17, 2014 What s the Question? Where do Transcription Factors (TFs) bind genomic DNA 1? (Where do other things bind
TECH NOTE Ligation-Free ChIP-Seq Library Preparation
 TECH NOTE Ligation-Free ChIP-Seq Library Preparation The DNA SMART ChIP-Seq Kit Ligation-free template switching technology: Minimize sample handling in a single-tube workflow >> Simplified protocol with
TECH NOTE Ligation-Free ChIP-Seq Library Preparation The DNA SMART ChIP-Seq Kit Ligation-free template switching technology: Minimize sample handling in a single-tube workflow >> Simplified protocol with
ChIP-Seq Data Analysis. J Fass UCD Genome Center Bioinformatics Core Wednesday 15 June 2015
 ChIP-Seq Data Analysis J Fass UCD Genome Center Bioinformatics Core Wednesday 15 June 2015 What s the Question? Where do Transcription Factors (TFs) bind genomic DNA 1? (Where do other things bind DNA
ChIP-Seq Data Analysis J Fass UCD Genome Center Bioinformatics Core Wednesday 15 June 2015 What s the Question? Where do Transcription Factors (TFs) bind genomic DNA 1? (Where do other things bind DNA
ChIP-Seq Tools. J Fass UCD Genome Center Bioinformatics Core Wednesday September 16, 2015
 ChIP-Seq Tools J Fass UCD Genome Center Bioinformatics Core Wednesday September 16, 2015 What s the Question? Where do Transcription Factors (TFs) bind genomic DNA 1? (Where do other things bind DNA or
ChIP-Seq Tools J Fass UCD Genome Center Bioinformatics Core Wednesday September 16, 2015 What s the Question? Where do Transcription Factors (TFs) bind genomic DNA 1? (Where do other things bind DNA or
Caroline Townsend December 2012 Biochem 218 A critical review of ChIP-seq enrichment analysis tools
 Caroline Townsend December 2012 Biochem 218 A critical review of ChIP-seq enrichment analysis tools Introduction Transcriptional regulation, chromatin states, and genome stability pathways are largely
Caroline Townsend December 2012 Biochem 218 A critical review of ChIP-seq enrichment analysis tools Introduction Transcriptional regulation, chromatin states, and genome stability pathways are largely
Parts of a standard FastQC report
 FastQC FastQC, written by Simon Andrews of Babraham Bioinformatics, is a very popular tool used to provide an overview of basic quality control metrics for raw next generation sequencing data. There are
FastQC FastQC, written by Simon Andrews of Babraham Bioinformatics, is a very popular tool used to provide an overview of basic quality control metrics for raw next generation sequencing data. There are
Bioinformatics in next generation sequencing projects
 Bioinformatics in next generation sequencing projects Rickard Sandberg Assistant Professor Department of Cell and Molecular Biology Karolinska Institutet May 2013 Standard sequence library generation Illumina
Bioinformatics in next generation sequencing projects Rickard Sandberg Assistant Professor Department of Cell and Molecular Biology Karolinska Institutet May 2013 Standard sequence library generation Illumina
Next Generation Sequencing: An Overview
 Next Generation Sequencing: An Overview Cavan Reilly November 13, 2017 Table of contents Next generation sequencing NGS and microarrays Study design Quality assessment Burrows Wheeler transform Next generation
Next Generation Sequencing: An Overview Cavan Reilly November 13, 2017 Table of contents Next generation sequencing NGS and microarrays Study design Quality assessment Burrows Wheeler transform Next generation
H3K36me3 polyclonal antibody
 H3K36me3 polyclonal antibody Cat. No. C15410192 Type: Polyclonal ChIP-grade/ChIP-seq grade Source: Rabbit Lot #: A1845P Size: 50 µg/32 µl Concentration: 1.6 μg/μl Specificity: Human, mouse, Arabidopsis,
H3K36me3 polyclonal antibody Cat. No. C15410192 Type: Polyclonal ChIP-grade/ChIP-seq grade Source: Rabbit Lot #: A1845P Size: 50 µg/32 µl Concentration: 1.6 μg/μl Specificity: Human, mouse, Arabidopsis,
Experimental Design. Sequencing. Data Quality Control. Read mapping. Differential Expression analysis
 -Seq Analysis Quality Control checks Reproducibility Reliability -seq vs Microarray Higher sensitivity and dynamic range Lower technical variation Available for all species Novel transcript identification
-Seq Analysis Quality Control checks Reproducibility Reliability -seq vs Microarray Higher sensitivity and dynamic range Lower technical variation Available for all species Novel transcript identification
Lecture 7. Next-generation sequencing technologies
 Lecture 7 Next-generation sequencing technologies Next-generation sequencing technologies General principles of short-read NGS Construct a library of fragments Generate clonal template populations Massively
Lecture 7 Next-generation sequencing technologies Next-generation sequencing technologies General principles of short-read NGS Construct a library of fragments Generate clonal template populations Massively
CS273B: Deep learning for Genomics and Biomedicine
 CS273B: Deep learning for Genomics and Biomedicine Lecture 2: Convolutional neural networks and applications to functional genomics 09/28/2016 Anshul Kundaje, James Zou, Serafim Batzoglou Outline Anatomy
CS273B: Deep learning for Genomics and Biomedicine Lecture 2: Convolutional neural networks and applications to functional genomics 09/28/2016 Anshul Kundaje, James Zou, Serafim Batzoglou Outline Anatomy
Computational Analysis of Ultra-high-throughput sequencing data: ChIP-Seq
 Computational Analysis of Ultra-high-throughput sequencing data: ChIP-Seq Philipp Bucher Wednesday January 21, 2009 SIB graduate school course EPFL, Lausanne Data flow in ChIP-Seq data analysis Level 1:
Computational Analysis of Ultra-high-throughput sequencing data: ChIP-Seq Philipp Bucher Wednesday January 21, 2009 SIB graduate school course EPFL, Lausanne Data flow in ChIP-Seq data analysis Level 1:
Data and Metadata Models Recommendations Version 1.2 Developed by the IHEC Metadata Standards Workgroup
 Data and Metadata Models Recommendations Version 1.2 Developed by the IHEC Metadata Standards Workgroup 1. Introduction The data produced by IHEC is illustrated in Figure 1. Figure 1. The space of epigenomic
Data and Metadata Models Recommendations Version 1.2 Developed by the IHEC Metadata Standards Workgroup 1. Introduction The data produced by IHEC is illustrated in Figure 1. Figure 1. The space of epigenomic
Supplemental Figure 1.
 Supplemental Data. Charron et al. Dynamic landscapes of four histone modifications during de-etiolation in Arabidopsis. Plant Cell (2009). 10.1105/tpc.109.066845 Supplemental Figure 1. Immunodetection
Supplemental Data. Charron et al. Dynamic landscapes of four histone modifications during de-etiolation in Arabidopsis. Plant Cell (2009). 10.1105/tpc.109.066845 Supplemental Figure 1. Immunodetection
Green Center Computational Core ChIP- Seq Pipeline, Just a Click Away
 Green Center Computational Core ChIP- Seq Pipeline, Just a Click Away Venkat Malladi Computational Biologist Computational Core Cecil H. and Ida Green Center for Reproductive Biology Science Introduc
Green Center Computational Core ChIP- Seq Pipeline, Just a Click Away Venkat Malladi Computational Biologist Computational Core Cecil H. and Ida Green Center for Reproductive Biology Science Introduc
Bioinformatics of Transcriptional Regulation
 Bioinformatics of Transcriptional Regulation Carl Herrmann IPMB & DKFZ c.herrmann@dkfz.de Wechselwirkung von Maßnahmen und Auswirkungen Einflussmöglichkeiten in einem Dialog From genes to active compounds
Bioinformatics of Transcriptional Regulation Carl Herrmann IPMB & DKFZ c.herrmann@dkfz.de Wechselwirkung von Maßnahmen und Auswirkungen Einflussmöglichkeiten in einem Dialog From genes to active compounds
Gene Expression analysis with RNA-Seq data
 Gene Expression analysis with RNA-Seq data C3BI Hands-on NGS course November 24th 2016 Frédéric Lemoine Plan 1. 2. Quality Control 3. Read Mapping 4. Gene Expression Analysis 5. Splicing/Transcript Analysis
Gene Expression analysis with RNA-Seq data C3BI Hands-on NGS course November 24th 2016 Frédéric Lemoine Plan 1. 2. Quality Control 3. Read Mapping 4. Gene Expression Analysis 5. Splicing/Transcript Analysis
Ecole de Bioinforma(que AVIESAN Roscoff 2014 GALAXY INITIATION. A. Lermine U900 Ins(tut Curie, INSERM, Mines ParisTech
 GALAXY INITIATION A. Lermine U900 Ins(tut Curie, INSERM, Mines ParisTech How does Next- Gen sequencing work? DNA fragmentation Size selection and clonal amplification Massive parallel sequencing ACCGTTTGCCG
GALAXY INITIATION A. Lermine U900 Ins(tut Curie, INSERM, Mines ParisTech How does Next- Gen sequencing work? DNA fragmentation Size selection and clonal amplification Massive parallel sequencing ACCGTTTGCCG
APPLICATION NOTE. Abstract. Introduction
 From minuscule amounts to magnificent results: reliable ChIP-seq data from 1, cells with the True MicroChIP and the MicroPlex Library Preparation kits Abstract Diagenode has developed groundbreaking solutions
From minuscule amounts to magnificent results: reliable ChIP-seq data from 1, cells with the True MicroChIP and the MicroPlex Library Preparation kits Abstract Diagenode has developed groundbreaking solutions
Why QC? Next-Generation Sequencing: Quality Control. Illumina data format. Fastq format:
 Why QC? Next-Generation Sequencing: Quality Control BaRC Hot Topics January 2017 Bioinformatics and Research Computing Whitehead Institute Do you want to include the reads with low quality base calls?
Why QC? Next-Generation Sequencing: Quality Control BaRC Hot Topics January 2017 Bioinformatics and Research Computing Whitehead Institute Do you want to include the reads with low quality base calls?
2/10/17. Contents. Applications of HMMs in Epigenomics
 2/10/17 I529: Machine Learning in Bioinformatics (Spring 2017) Contents Applications of HMMs in Epigenomics Yuzhen Ye School of Informatics and Computing Indiana University, Bloomington Spring 2017 Background:
2/10/17 I529: Machine Learning in Bioinformatics (Spring 2017) Contents Applications of HMMs in Epigenomics Yuzhen Ye School of Informatics and Computing Indiana University, Bloomington Spring 2017 Background:
Next-Generation Sequencing: Quality Control
 Next-Generation Sequencing: Quality Control Bingbing Yuan BaRC Hot Topics January 2017 Bioinformatics and Research Computing Whitehead Institute http://barc.wi.mit.edu/hot_topics/ Why QC? Do you want to
Next-Generation Sequencing: Quality Control Bingbing Yuan BaRC Hot Topics January 2017 Bioinformatics and Research Computing Whitehead Institute http://barc.wi.mit.edu/hot_topics/ Why QC? Do you want to
Next- genera*on Sequencing. Lecture 13
 Next- genera*on Sequencing Lecture 13 ChIP- seq Applica*ons iden%fy sequence varia%ons DNA- seq Iden%fy Pathogens RNA- seq Kahvejian et al, 2008 Protein-DNA interaction DNA is the informa*on carrier of
Next- genera*on Sequencing Lecture 13 ChIP- seq Applica*ons iden%fy sequence varia%ons DNA- seq Iden%fy Pathogens RNA- seq Kahvejian et al, 2008 Protein-DNA interaction DNA is the informa*on carrier of
Sanger vs Next-Gen Sequencing
 Tools and Algorithms in Bioinformatics GCBA815/MCGB815/BMI815, Fall 2017 Week-8: Next-Gen Sequencing RNA-seq Data Analysis Babu Guda, Ph.D. Professor, Genetics, Cell Biology & Anatomy Director, Bioinformatics
Tools and Algorithms in Bioinformatics GCBA815/MCGB815/BMI815, Fall 2017 Week-8: Next-Gen Sequencing RNA-seq Data Analysis Babu Guda, Ph.D. Professor, Genetics, Cell Biology & Anatomy Director, Bioinformatics
Incorporating Molecular ID Technology. Accel-NGS 2S MID Indexing Kits
 Incorporating Molecular ID Technology Accel-NGS 2S MID Indexing Kits Molecular Identifiers (MIDs) MIDs are indices used to label unique library molecules MIDs can assess duplicate molecules in sequencing
Incorporating Molecular ID Technology Accel-NGS 2S MID Indexing Kits Molecular Identifiers (MIDs) MIDs are indices used to label unique library molecules MIDs can assess duplicate molecules in sequencing
Short Read Alignment to a Reference Genome
 Short Read Alignment to a Reference Genome Shamith Samarajiwa CRUK Summer School in Bioinformatics Cambridge, September 2018 Aligning to a reference genome BWA Bowtie2 STAR GEM Pseudo Aligners for RNA-seq
Short Read Alignment to a Reference Genome Shamith Samarajiwa CRUK Summer School in Bioinformatics Cambridge, September 2018 Aligning to a reference genome BWA Bowtie2 STAR GEM Pseudo Aligners for RNA-seq
BST 226 Statistical Methods for Bioinformatics David M. Rocke. March 10, 2014 BST 226 Statistical Methods for Bioinformatics 1
 BST 226 Statistical Methods for Bioinformatics David M. Rocke March 10, 2014 BST 226 Statistical Methods for Bioinformatics 1 NGS Technologies Illumina Sequencing HiSeq 2500 & MiSeq PacBio Sequencing PacBio
BST 226 Statistical Methods for Bioinformatics David M. Rocke March 10, 2014 BST 226 Statistical Methods for Bioinformatics 1 NGS Technologies Illumina Sequencing HiSeq 2500 & MiSeq PacBio Sequencing PacBio
Applications of HMMs in Epigenomics
 I529: Machine Learning in Bioinformatics (Spring 2013) Applications of HMMs in Epigenomics Yuzhen Ye School of Informatics and Computing Indiana University, Bloomington Spring 2013 Contents Background:
I529: Machine Learning in Bioinformatics (Spring 2013) Applications of HMMs in Epigenomics Yuzhen Ye School of Informatics and Computing Indiana University, Bloomington Spring 2013 Contents Background:
Nature Genetics: doi: /ng Supplementary Figure 1. H3K27ac HiChIP enriches enhancer promoter-associated chromatin contacts.
 Supplementary Figure 1 H3K27ac HiChIP enriches enhancer promoter-associated chromatin contacts. (a) Schematic of chromatin contacts captured in H3K27ac HiChIP. (b) Loop call overlap for cohesin HiChIP
Supplementary Figure 1 H3K27ac HiChIP enriches enhancer promoter-associated chromatin contacts. (a) Schematic of chromatin contacts captured in H3K27ac HiChIP. (b) Loop call overlap for cohesin HiChIP
Supplemental Figure 1 A
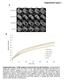 Supplemental Figure A prebleach postbleach 2 min 6 min 3 min mh2a.-gfp mh2a.2-gfp mh2a2-gfp GFP-H2A..9 Relative Intensity.8.7.6.5 mh2a. GFP n=8.4 mh2a.2 GFP n=4.3 mh2a2 GFP n=2.2 GFP H2A n=24. GFP n=7.
Supplemental Figure A prebleach postbleach 2 min 6 min 3 min mh2a.-gfp mh2a.2-gfp mh2a2-gfp GFP-H2A..9 Relative Intensity.8.7.6.5 mh2a. GFP n=8.4 mh2a.2 GFP n=4.3 mh2a2 GFP n=2.2 GFP H2A n=24. GFP n=7.
Next-generation sequencing and quality control: An introduction 2016
 Next-generation sequencing and quality control: An introduction 2016 s.schmeier@massey.ac.nz http://sschmeier.com/bioinf-workshop/ Overview Typical workflow of a genomics experiment Genome versus transcriptome
Next-generation sequencing and quality control: An introduction 2016 s.schmeier@massey.ac.nz http://sschmeier.com/bioinf-workshop/ Overview Typical workflow of a genomics experiment Genome versus transcriptome
Reference genomes and common file formats
 Reference genomes and common file formats Overview Reference genomes and GRC Fasta and FastQ (unaligned sequences) SAM/BAM (aligned sequences) Summarized genomic features BED (genomic intervals) GFF/GTF
Reference genomes and common file formats Overview Reference genomes and GRC Fasta and FastQ (unaligned sequences) SAM/BAM (aligned sequences) Summarized genomic features BED (genomic intervals) GFF/GTF
RNA-Seq de novo assembly training
 RNA-Seq de novo assembly training Training session aims Give you some keys elements to look at during read quality check. Transcriptome assembly is not completely a strait forward process : Multiple strategies
RNA-Seq de novo assembly training Training session aims Give you some keys elements to look at during read quality check. Transcriptome assembly is not completely a strait forward process : Multiple strategies
ChIP-seq/Functional Genomics/Epigenomics. CBSU/3CPG/CVG Next-Gen Sequencing Workshop. Josh Waterfall. March 31, 2010
 ChIP-seq/Functional Genomics/Epigenomics CBSU/3CPG/CVG Next-Gen Sequencing Workshop Josh Waterfall March 31, 2010 Outline Introduction to ChIP-seq Control data sets Peak/enriched region identification
ChIP-seq/Functional Genomics/Epigenomics CBSU/3CPG/CVG Next-Gen Sequencing Workshop Josh Waterfall March 31, 2010 Outline Introduction to ChIP-seq Control data sets Peak/enriched region identification
L3: Short Read Alignment to a Reference Genome
 L3: Short Read Alignment to a Reference Genome Shamith Samarajiwa CRUK Autumn School in Bioinformatics Cambridge, September 2017 Where to get help! http://seqanswers.com http://www.biostars.org http://www.bioconductor.org/help/mailing-list
L3: Short Read Alignment to a Reference Genome Shamith Samarajiwa CRUK Autumn School in Bioinformatics Cambridge, September 2017 Where to get help! http://seqanswers.com http://www.biostars.org http://www.bioconductor.org/help/mailing-list
Data Basics. Josef K Vogt Slides by: Simon Rasmussen Next Generation Sequencing Analysis
 Data Basics Josef K Vogt Slides by: Simon Rasmussen 2017 Generalized NGS analysis Sample prep & Sequencing Data size Main data reductive steps SNPs, genes, regions Application Assembly: Compare Raw Pre-
Data Basics Josef K Vogt Slides by: Simon Rasmussen 2017 Generalized NGS analysis Sample prep & Sequencing Data size Main data reductive steps SNPs, genes, regions Application Assembly: Compare Raw Pre-
Figure 7.1: PWM evolution: The sequence affinity of TFBSs has evolved from single sequences, to PWMs, to larger and larger databases of PWMs.
 Chapter 7 Discussion This thesis presents dry and wet lab techniques to elucidate the involvement of transcription factors (TFs) in the regulation of the cell cycle and myogenesis. However, the techniques
Chapter 7 Discussion This thesis presents dry and wet lab techniques to elucidate the involvement of transcription factors (TFs) in the regulation of the cell cycle and myogenesis. However, the techniques
ChIP-seq guidlines used by ENCODE AND modencode consortia. 2012/11/15 DJEKIDEL Mohamed Nadhir
 ChIP-seq guidlines used by ENCODE AND modencode consortia 2012/11/15 DJEKIDEL Mohamed Nadhir Paper & authors ENCODE and modencode performed: >1000 ChIP-seq experiment. 140 different factors. 100 cell lines
ChIP-seq guidlines used by ENCODE AND modencode consortia 2012/11/15 DJEKIDEL Mohamed Nadhir Paper & authors ENCODE and modencode performed: >1000 ChIP-seq experiment. 140 different factors. 100 cell lines
Next Generation Sequencing Lecture Saarbrücken, 19. March Sequencing Platforms
 Next Generation Sequencing Lecture Saarbrücken, 19. March 2012 Sequencing Platforms Contents Introduction Sequencing Workflow Platforms Roche 454 ABI SOLiD Illumina Genome Anlayzer / HiSeq Problems Quality
Next Generation Sequencing Lecture Saarbrücken, 19. March 2012 Sequencing Platforms Contents Introduction Sequencing Workflow Platforms Roche 454 ABI SOLiD Illumina Genome Anlayzer / HiSeq Problems Quality
2/19/13. Contents. Applications of HMMs in Epigenomics
 2/19/13 I529: Machine Learning in Bioinformatics (Spring 2013) Contents Applications of HMMs in Epigenomics Yuzhen Ye School of Informatics and Computing Indiana University, Bloomington Spring 2013 Background:
2/19/13 I529: Machine Learning in Bioinformatics (Spring 2013) Contents Applications of HMMs in Epigenomics Yuzhen Ye School of Informatics and Computing Indiana University, Bloomington Spring 2013 Background:
DNA:CHROMATIN INTERACTIONS
 DNA:CHROMATIN INTERACTIONS Exploring transcription factor binding and the epigenomic landscape Chris Seward Introductions Cell and Developmental Biology PhD Candidate in Dr. Lisa Stubbs Laboratory Currently
DNA:CHROMATIN INTERACTIONS Exploring transcription factor binding and the epigenomic landscape Chris Seward Introductions Cell and Developmental Biology PhD Candidate in Dr. Lisa Stubbs Laboratory Currently
Genome 373: Mapping Short Sequence Reads II. Doug Fowler
 Genome 373: Mapping Short Sequence Reads II Doug Fowler The final Will be in this room on June 6 th at 8:30a Will be focused on the second half of the course, but will include material from the first half
Genome 373: Mapping Short Sequence Reads II Doug Fowler The final Will be in this room on June 6 th at 8:30a Will be focused on the second half of the course, but will include material from the first half
Introduc)on to Bioinforma)cs of next- genera)on sequencing. Sequence acquisi)on and processing; genome mapping and alignment manipula)on
 Introduc)on to Bioinforma)cs of next- genera)on sequencing Sequence acquisi)on and processing; genome mapping and alignment manipula)on Ruslan Sadreyev Director of Bioinformatics Department of Molecular
Introduc)on to Bioinforma)cs of next- genera)on sequencing Sequence acquisi)on and processing; genome mapping and alignment manipula)on Ruslan Sadreyev Director of Bioinformatics Department of Molecular
The first thing you will see is the opening page. SeqMonk scans your copy and make sure everything is in order, indicated by the green check marks.
 Open Seqmonk Launch SeqMonk The first thing you will see is the opening page. SeqMonk scans your copy and make sure everything is in order, indicated by the green check marks. SeqMonk Analysis Page 1 Create
Open Seqmonk Launch SeqMonk The first thing you will see is the opening page. SeqMonk scans your copy and make sure everything is in order, indicated by the green check marks. SeqMonk Analysis Page 1 Create
RNA-seq Data Analysis
 Lecture 3. Clustering; Function/Pathway Enrichment analysis RNA-seq Data Analysis Qi Sun Bioinformatics Facility Biotechnology Resource Center Cornell University Lecture 1. Map RNA-seq read to genome Lecture
Lecture 3. Clustering; Function/Pathway Enrichment analysis RNA-seq Data Analysis Qi Sun Bioinformatics Facility Biotechnology Resource Center Cornell University Lecture 1. Map RNA-seq read to genome Lecture
A more efficient, sensitive and robust method of chromatin immunoprecipitation (ChIP)
 A more efficient, sensitive and robust method of chromatin immunoprecipitation (ChIP) ADVANCEMENTS IN EPIGENETICS Introducing ChIP and Chromatrap Chromatrap is a more efficient, sensitive and robust method
A more efficient, sensitive and robust method of chromatin immunoprecipitation (ChIP) ADVANCEMENTS IN EPIGENETICS Introducing ChIP and Chromatrap Chromatrap is a more efficient, sensitive and robust method
Astrocyte GCRB/BICF Workflow for ChIP-Seq Analysis. Venkat Beibei
 Astrocyte GCRB/BICF Workflow for ChIP-Seq Analysis Venkat Malladi @GCRB Beibei Chen @BICF What%is%ChIP+Seq?% Chromatin immunoprecipitation followed by Sequencing (ChIP-Seq): Identify the binding sites
Astrocyte GCRB/BICF Workflow for ChIP-Seq Analysis Venkat Malladi @GCRB Beibei Chen @BICF What%is%ChIP+Seq?% Chromatin immunoprecipitation followed by Sequencing (ChIP-Seq): Identify the binding sites
Introduction of RNA-Seq Analysis
 Introduction of RNA-Seq Analysis Jiang Li, MS Bioinformatics System Engineer I Center for Quantitative Sciences(CQS) Vanderbilt University September 21, 2012 Goal of this talk 1. Act as a practical resource
Introduction of RNA-Seq Analysis Jiang Li, MS Bioinformatics System Engineer I Center for Quantitative Sciences(CQS) Vanderbilt University September 21, 2012 Goal of this talk 1. Act as a practical resource
Analysis of RNA-seq Data. Feb 8, 2017 Peikai CHEN (PHD)
 Analysis of RNA-seq Data Feb 8, 2017 Peikai CHEN (PHD) Outline What is RNA-seq? What can RNA-seq do? How is RNA-seq measured? How to process RNA-seq data: the basics How to visualize and diagnose your
Analysis of RNA-seq Data Feb 8, 2017 Peikai CHEN (PHD) Outline What is RNA-seq? What can RNA-seq do? How is RNA-seq measured? How to process RNA-seq data: the basics How to visualize and diagnose your
Deep Sequencing technologies
 Deep Sequencing technologies Gabriela Salinas 30 October 2017 Transcriptome and Genome Analysis Laboratory http://www.uni-bc.gwdg.de/index.php?id=709 Microarray and Deep-Sequencing Core Facility University
Deep Sequencing technologies Gabriela Salinas 30 October 2017 Transcriptome and Genome Analysis Laboratory http://www.uni-bc.gwdg.de/index.php?id=709 Microarray and Deep-Sequencing Core Facility University
Genome 373: High- Throughput DNA Sequencing. Doug Fowler
 Genome 373: High- Throughput DNA Sequencing Doug Fowler Tasks give ML unity We learned about three tasks that are commonly encountered in ML Models/Algorithms Give ML Diversity Classification Regression
Genome 373: High- Throughput DNA Sequencing Doug Fowler Tasks give ML unity We learned about three tasks that are commonly encountered in ML Models/Algorithms Give ML Diversity Classification Regression
De Novo Assembly of High-throughput Short Read Sequences
 De Novo Assembly of High-throughput Short Read Sequences Chuming Chen Center for Bioinformatics and Computational Biology (CBCB) University of Delaware NECC Third Skate Genome Annotation Workshop May 23,
De Novo Assembly of High-throughput Short Read Sequences Chuming Chen Center for Bioinformatics and Computational Biology (CBCB) University of Delaware NECC Third Skate Genome Annotation Workshop May 23,
NGS in Pathology Webinar
 NGS in Pathology Webinar NGS Data Analysis March 10 2016 1 Topics for today s presentation 2 Introduction Next Generation Sequencing (NGS) is becoming a common and versatile tool for biological and medical
NGS in Pathology Webinar NGS Data Analysis March 10 2016 1 Topics for today s presentation 2 Introduction Next Generation Sequencing (NGS) is becoming a common and versatile tool for biological and medical
Assay Standards Working Group Nov 2012 Assay Standards Working Group Recommendations, November 2012
 Assay Standards Working Group Recommendations, November 2012 Contents Assay Standards Working Group Recommendations, August 2012... 1 Contents... 1 Introduction... 2 1: Reference Epigenome Criteria...
Assay Standards Working Group Recommendations, November 2012 Contents Assay Standards Working Group Recommendations, August 2012... 1 Contents... 1 Introduction... 2 1: Reference Epigenome Criteria...
Discovering gene regulatory control using ChIP-chip and ChIP-seq. An introduction to gene regulatory control, concepts and methodologies
 Discovering gene regulatory control using ChIP-chip and ChIP-seq An introduction to gene regulatory control, concepts and methodologies Ian Simpson ian.simpson@.ed.ac.uk bit.ly/bio2_2012 The Central Dogma
Discovering gene regulatory control using ChIP-chip and ChIP-seq An introduction to gene regulatory control, concepts and methodologies Ian Simpson ian.simpson@.ed.ac.uk bit.ly/bio2_2012 The Central Dogma
Reads to Discovery. Visualize Annotate Discover. Small DNA-Seq ChIP-Seq Methyl-Seq. MeDIP-Seq. RNA-Seq. RNA-Seq.
 Reads to Discovery RNA-Seq Small DNA-Seq ChIP-Seq Methyl-Seq RNA-Seq MeDIP-Seq www.strand-ngs.com Analyze Visualize Annotate Discover Data Import Alignment Vendor Platforms: Illumina Ion Torrent Roche
Reads to Discovery RNA-Seq Small DNA-Seq ChIP-Seq Methyl-Seq RNA-Seq MeDIP-Seq www.strand-ngs.com Analyze Visualize Annotate Discover Data Import Alignment Vendor Platforms: Illumina Ion Torrent Roche
Gene expression analysis. Biosciences 741: Genomics Fall, 2013 Week 5. Gene expression analysis
 Gene expression analysis Biosciences 741: Genomics Fall, 2013 Week 5 Gene expression analysis From EST clusters to spotted cdna microarrays Long vs. short oligonucleotide microarrays vs. RT-PCR Methods
Gene expression analysis Biosciences 741: Genomics Fall, 2013 Week 5 Gene expression analysis From EST clusters to spotted cdna microarrays Long vs. short oligonucleotide microarrays vs. RT-PCR Methods
Hitting the mark: specificity analysis of histone antibodies
 APPLICATION NOTE Histone modification antibodies Hitting the mark: specificity analysis of histone antibodies Introduction The nucleosome, composed of the histones H2A, H2B, H3, and H4, is the fundamental
APPLICATION NOTE Histone modification antibodies Hitting the mark: specificity analysis of histone antibodies Introduction The nucleosome, composed of the histones H2A, H2B, H3, and H4, is the fundamental
Next-Generation Sequencing. Technologies
 Next-Generation Next-Generation Sequencing Technologies Sequencing Technologies Nicholas E. Navin, Ph.D. MD Anderson Cancer Center Dept. Genetics Dept. Bioinformatics Introduction to Bioinformatics GS011062
Next-Generation Next-Generation Sequencing Technologies Sequencing Technologies Nicholas E. Navin, Ph.D. MD Anderson Cancer Center Dept. Genetics Dept. Bioinformatics Introduction to Bioinformatics GS011062
Atelier Chip-Seq. Stéphanie Le Gras, IGBMC Strasbourg Violaine Saint-André, Institut Curie Paris Morgane Thomas-Chollier, ENS Paris
 Atelier Chip-Seq Stéphanie Le Gras, IGBMC Strasbourg Violaine Saint-André, Institut Curie Paris Morgane Thomas-Chollier, ENS Paris École de bioinformatique AVIESAN-IFB 2017 Get connected to the server
Atelier Chip-Seq Stéphanie Le Gras, IGBMC Strasbourg Violaine Saint-André, Institut Curie Paris Morgane Thomas-Chollier, ENS Paris École de bioinformatique AVIESAN-IFB 2017 Get connected to the server
Y1 Biology 131 Syllabus - Academic Year
 Y1 Biology 131 Syllabus - Academic Year 2016-2017 Monday 28/11/2016 DNA Packaging Week 11 Tuesday 29/11/2016 Regulation of gene expression Wednesday 22/9/2014 Cell cycle Sunday 4/12/2016 Tutorial Monday
Y1 Biology 131 Syllabus - Academic Year 2016-2017 Monday 28/11/2016 DNA Packaging Week 11 Tuesday 29/11/2016 Regulation of gene expression Wednesday 22/9/2014 Cell cycle Sunday 4/12/2016 Tutorial Monday
02 Agenda Item 03 Agenda Item
 01 Agenda Item 02 Agenda Item 03 Agenda Item SOLiD 3 System: Applications Overview April 12th, 2010 Jennifer Stover Field Application Specialist - SOLiD Applications Workflow for SOLiD Application Application
01 Agenda Item 02 Agenda Item 03 Agenda Item SOLiD 3 System: Applications Overview April 12th, 2010 Jennifer Stover Field Application Specialist - SOLiD Applications Workflow for SOLiD Application Application
ChampionChIP Quick, High Throughput Chromatin Immunoprecipitation Assay System
 ChampionChIP Quick, High Throughput Chromatin Immunoprecipitation Assay System Liyan Pang, Ph.D. Application Scientist 1 Topics to be Covered Introduction What is ChIP-qPCR? Challenges Facing Biological
ChampionChIP Quick, High Throughput Chromatin Immunoprecipitation Assay System Liyan Pang, Ph.D. Application Scientist 1 Topics to be Covered Introduction What is ChIP-qPCR? Challenges Facing Biological
Analysis of ChIP-seq data with R / Bioconductor
 Analysis of ChIP-seq data with R / Bioconductor Martin Morgan Bioconductor / Fred Hutchinson Cancer Research Center Seattle, WA, USA 8-10 June 2009 ChIP-seq Chromatin immunopreciptation to enrich sample
Analysis of ChIP-seq data with R / Bioconductor Martin Morgan Bioconductor / Fred Hutchinson Cancer Research Center Seattle, WA, USA 8-10 June 2009 ChIP-seq Chromatin immunopreciptation to enrich sample
Illumina Sequencing Error Profiles and Quality Control
 Illumina Sequencing Error Profiles and Quality Control RNA-seq Workflow Biological samples/library preparation Sequence reads FASTQC Adapter Trimming (Optional) Splice-aware mapping to genome Counting
Illumina Sequencing Error Profiles and Quality Control RNA-seq Workflow Biological samples/library preparation Sequence reads FASTQC Adapter Trimming (Optional) Splice-aware mapping to genome Counting
Chromatin. Structure and modification of chromatin. Chromatin domains
 Chromatin Structure and modification of chromatin Chromatin domains 2 DNA consensus 5 3 3 DNA DNA 4 RNA 5 ss RNA forms secondary structures with ds hairpins ds forms 6 of nucleic acids Form coiling bp/turn
Chromatin Structure and modification of chromatin Chromatin domains 2 DNA consensus 5 3 3 DNA DNA 4 RNA 5 ss RNA forms secondary structures with ds hairpins ds forms 6 of nucleic acids Form coiling bp/turn
Systematic evaluation of the impact of ChIP-seq read designs on genome coverage, peak identification, and allele-specific binding detection
 Zhang et al. BMC Bioinformatics (216) 17:96 DOI 1.1186/s12859-16-957-1 RESEARCH ARTICLE Open Access Systematic evaluation of the impact of ChIP-seq read designs on genome coverage, peak identification,
Zhang et al. BMC Bioinformatics (216) 17:96 DOI 1.1186/s12859-16-957-1 RESEARCH ARTICLE Open Access Systematic evaluation of the impact of ChIP-seq read designs on genome coverage, peak identification,
Whole Transcriptome Analysis of Illumina RNA- Seq Data. Ryan Peters Field Application Specialist
 Whole Transcriptome Analysis of Illumina RNA- Seq Data Ryan Peters Field Application Specialist Partek GS in your NGS Pipeline Your Start-to-Finish Solution for Analysis of Next Generation Sequencing Data
Whole Transcriptome Analysis of Illumina RNA- Seq Data Ryan Peters Field Application Specialist Partek GS in your NGS Pipeline Your Start-to-Finish Solution for Analysis of Next Generation Sequencing Data
Differential gene expression analysis using RNA-seq
 https://abc.med.cornell.edu/ Differential gene expression analysis using RNA-seq Applied Bioinformatics Core, March 2018 Friederike Dündar with Luce Skrabanek & Paul Zumbo Day 1: Introduction into high-throughput
https://abc.med.cornell.edu/ Differential gene expression analysis using RNA-seq Applied Bioinformatics Core, March 2018 Friederike Dündar with Luce Skrabanek & Paul Zumbo Day 1: Introduction into high-throughput
Browser Exercises - I. Alignments and Comparative genomics
 Browser Exercises - I Alignments and Comparative genomics 1. Navigating to the Genome Browser (GBrowse) Note: For this exercise use http://www.tritrypdb.org a. Navigate to the Genome Browser (GBrowse)
Browser Exercises - I Alignments and Comparative genomics 1. Navigating to the Genome Browser (GBrowse) Note: For this exercise use http://www.tritrypdb.org a. Navigate to the Genome Browser (GBrowse)
About Strand NGS. Strand Genomics, Inc All rights reserved.
 About Strand NGS Strand NGS-formerly known as Avadis NGS, is an integrated platform that provides analysis, management and visualization tools for next-generation sequencing data. It supports extensive
About Strand NGS Strand NGS-formerly known as Avadis NGS, is an integrated platform that provides analysis, management and visualization tools for next-generation sequencing data. It supports extensive
Plant Molecular and Cellular Biology Lecture 9: Nuclear Genome Organization: Chromosome Structure, Chromatin, DNA Packaging, Mitosis Gary Peter
 Plant Molecular and Cellular Biology Lecture 9: Nuclear Genome Organization: Chromosome Structure, Chromatin, DNA Packaging, Mitosis Gary Peter 9/16/2008 1 Learning Objectives 1. List and explain how DNA
Plant Molecular and Cellular Biology Lecture 9: Nuclear Genome Organization: Chromosome Structure, Chromatin, DNA Packaging, Mitosis Gary Peter 9/16/2008 1 Learning Objectives 1. List and explain how DNA
NUCLEIC ACIDS. DNA (Deoxyribonucleic Acid) and RNA (Ribonucleic Acid): information storage molecules made up of nucleotides.
 NUCLEIC ACIDS DNA (Deoxyribonucleic Acid) and RNA (Ribonucleic Acid): information storage molecules made up of nucleotides. Base Adenine Guanine Cytosine Uracil Thymine Abbreviation A G C U T DNA RNA 2
NUCLEIC ACIDS DNA (Deoxyribonucleic Acid) and RNA (Ribonucleic Acid): information storage molecules made up of nucleotides. Base Adenine Guanine Cytosine Uracil Thymine Abbreviation A G C U T DNA RNA 2
Integrated NGS Sample Preparation Solutions for Limiting Amounts of RNA and DNA. March 2, Steven R. Kain, Ph.D. ABRF 2013
 Integrated NGS Sample Preparation Solutions for Limiting Amounts of RNA and DNA March 2, 2013 Steven R. Kain, Ph.D. ABRF 2013 NuGEN s Core Technologies Selective Sequence Priming Nucleic Acid Amplification
Integrated NGS Sample Preparation Solutions for Limiting Amounts of RNA and DNA March 2, 2013 Steven R. Kain, Ph.D. ABRF 2013 NuGEN s Core Technologies Selective Sequence Priming Nucleic Acid Amplification
