SUPPLEMENTARY INFORMATION
|
|
|
- Jean Bradford
- 6 years ago
- Views:
Transcription
1 DOI: /ncb2239 Hepatocytes (Endoderm) Foreskin fibroblasts (Mesoderm) Melanocytes (Ectoderm) +Dox rtta TetO CMV O,S,M,K & N H1 hes H7 hes H9 hes H1 hes-derived EBs H7 hes-derived EBs H9 hes-derived EBs Affymetrix Human Gene ST 1.0 arrays Pluripotency validation for the Hepatocyte-iPS cells derived and used for the microarray analysis (Fig 1) Comparison of transcriptional profiles between hescs, hipscs derived from different cell types & differentiated cells (Fig 2) Analysis of transcriptional somatic cell memory in each type of hipsc (Fig 3) Correlation between differential gene expression and DNA methylation/bisulfite sequencing analysis (Fig 4) Meta-analysis of DNA methylation-associated transcriptional memory in independent datasets (Fig 5) Functional analysis of the somatic cell memory gene C9orf64 during reprogramming (Fig 6) Proximity in the genome affects efficiency of gene silencing in ips cells (Fig 7) Model for the role of DNA methylation in differences between human ips cells and ES cells (Fig 8) Figure S1 Flowchart of analysis and data reported in the manuscript. Human somatic cells representative of the 3 embryonic germ layers were reprogrammed to pluripotency using the dox-inducible lentiviral transgene system for overexpression of the reprogramming factors OCT4 (O), SOX2 (S), cmyc (M), KLF4 (K) and NANOG (N). Hepatocyte-derived ips cells, newborn foreskin fibroblast-derived ips cells and melanocyte-derived ips cells represented the endodermal, mesodermal and ectodermal lineages, respectively. All 3 types of ips cells, their parental somatic cell counterparts, 3 lines of human ES cells (H1, H7, H9) and Embryoid Bodies derived from these ES cells were hybridized to Affymetrix Human Gene ST 1.0 arrays and transcriptionally profiled in parallel. Triplicates of independent clones were used for all cell types except for the somatic cells, since each somatic cell represents a single clone. For all 3 types of parental somatic cells, technical triplicates were used for the analysis. Shown in the box beneath the flowchart is a brief description of each of the figures presented in the main text of the manuscript. 1
2 Figure S2 Pluripotency validation for the Fibroblast-iPS cells derived and used for the microarray studies. (a) All 3 Fib-iPS clones used in this analysis showed strong, positive staining for the human ES cell specific-markers SSEA-3, SSEA-4, Tra1-60 and Tra1-81, comparable to that observed in control H9 ES cells. Scale bar represents 300 μm. (b) qrt-pcr was used to confirm both high expression levels of endogenous pluripotency genes in all 3 Fib-iPS cell clones, as well as negligible levels of transgene expression. Values were standardized to GAPDH and Ubb, then normalized to H9 ES cells (endogenous) or 5-factor infected BJ fibroblasts + dox for 4 days (viral). (c) All clones of Fib-iPS cells formed embryoid bodies in vitro when grown under non-attachment conditions. Shown here are d8 EBs alongside with control ES cell-derived EBs. Scale bar represents 200 μm. (d) The pluripotent nature of all ips cells used in our analysis was confirmed by their ability to form EBs comprised of tissues derivative of the 3 germ layers in vitro (also see Fig. 1c and Supplementary Figure 2c). qrt-pcr analysis on d8 EBs derived from all types of ips cells and ES cell controls confirmed the presence of the 3 embryonic germ layers. Values were standardized to GAPDH and Ubb. Expression fold changes shown in the graph are relative to H9 ES cells on d0. (e) Hematoxylin and eosin stain of teratomas generated from fibroblastderived ips cells injected subcutaneously into immunocompromised SCID mice. Structures derivative of all three germ layers could be identified. (i) Neural tissue (ectoderm), (ii) Striated muscle and mesenchyme (mesoderm), (iii) gut-like epithelium (endoderm). In b and d, data are from triplicate reactions and error bars represent standard deviations. 2
3 a b Figure S3 ips cells retain a transcriptional memory of the original somatic cell. (a) Similar to Fig. 3a, these figures show LOESS curves fitted to the scatter plots of t-test log p-values for fibroblast and melanocyte: -log(p) and log(p) are plotted for fold changes greater than 1 and less than 1, respectively. The black line is a curve fitted to our data, and other curves are fitted to the 1000 bootstrap simulation datasets obtained by assuming identically distributed ips and ES cell expression levels. The black line shows clear deviation from the null hypothesis ips=es and thus reflects the trend that the transcriptional memory of the originating cell type is retained in ips cells: genes that were higher (or lower) in the somatic cell than in ES cells tend to be significantly repressed (or induced) during reprogramming, but nevertheless remain higher (or lower) in ips cells than in ES cells. (b) The partition of differentially expressed genes between ips cells and ES cells according to their expression status in somatic cells. It is seen that ~50% of the genes were already differentially expressed between the corresponding somatic cell type and ES cells, while ~10% were differentially expressed only in the corresponding somatic cell type and not other cell types relative to ES cells. Cell type-specific somatic expression can thus explain ~10% of the observed incomplete reprogramming. 3
4 b c Figure S4 Targeted differentiation of ips and ES cells towards an endodermal fate. (a) A targeted differentiation approach based on a previously published protocol by Kroon et al., 2008 was used to differentiate 2 clones each of Fib-iPS and Hep-iPS cells and 2 lines of ES cells (H1 and H9) to the definitive endoderm stage (d3 of the protocol) and to the primitive gut tube stage (d6 of the protocol) using the growth conditions shown. (b) On d3 of the differentiation assay, qrt-pcr was used to analyze the expression levels of 3 definitive endoderm markers (SOX17, FOXA2, CXCR4) in each of the cell types analyzed. SOX7 (extraembryonic endoderm-specific marker) expression was used to monitor the formation of definitive endoderm since SOX17, FOXA2 and CXCR4 can be found in all endodermal tissues. Low SOX7 levels indicated that the tissue generated was definitive endoderm and low OCT4 expression levels indicated the loss of pluripotency in ips and ES cells during differentiation. (c) On d6, the expression level of 2 primitive gut tube markers, HNF1B and HNF4A, was analyzed by qrt-pcr. The same controls as on d3 were used in this analysis. In both graphs, values were standardized to Ubb and TBP, and the expression fold changes for all genes in each cell type are referenced to a corresponding d0 sample. In b and c, data are from triplicate reactions and error bars represent standard deviations. 4
5 a D3 - Definitive endoderm stage: b D6 - Primitive gut tube stage: 58.4% 76.0% 92.3% 86.0% 72.7% 87.7% 72.1% 76.3% 83.9% 84.2% 76.4% 93.1% 77.2% 76.7% 51.8% 69.9% 34.8% 44.9% 43.6% 51.9% 66.2% 52.8% 57.8% 56.8% Figure S5 Immunohistochemical analysis of ips and ES cells differentiated towards endoderm. (a) On d3 of the targeted differentiation protocol, the same 2 clones of Fib-iPS (BJ 1 and 2) and Hep-iPS (Hep 1 and 2) cells, as well as 2 human ES cell lines (H1 and H9), analyzed by qrt-pcr in Supplementary Figure 4b were stained for the definitive endoderm markers FOXA2 and SOX17. The number of FOXA2- and SOX17-positive cells relative to the total number of DAPI-positive nuclei were quantified for each cell type analyzed. Values in green, percentage of FOXA2-positive cells; values in red, percentage of SOX17-postiive cells. (b) On d6 of the targeted differentiation protocol, the same cell lines were stained for FOXA2 and the primitive gut marker, HNF1b. The number of FOXA2- and HNF1b-positive cells relative to the total number of DAPI-positive nuclei were quantified for each cell type analyzed. Values in green, percentage of FOXA2-positive cells; values in red, percentage of HNF1b-postiive cells. Scale bars represent 200 μm. 5
6 Figure S6 Genome browser tracks of the 4 most incompletely silenced genes showing differential methylation between H1 ES cells and IMR90 cells. (a) Upper panel, a putative gene C9orf64 on chromosome 9, showing browser tracks for CpG islands, and DNA methylation levels for H1 ES and IMR90 cells assayed by MethylC-Seq (Lister et al. 2009). In the H1 ES and IMR90 cells methylation tracks, the Y-axis displays methylation scores of individual sites (CG, CHG and CHH). Methylation score is defined by the number of Cs and Ts at that position from MethylC-seq reads using the following formula: score = number of Cs / (C+T) * Data from both strands are combined. Scores range between -500 (unmethylated) and 500 (methylated), and the zero line is equivalent to 50% methylated. Negative scores are displayed as green bars and positive scores are displayed as orange bars. Lower panel, a close-up of the promoter near the region (black rectangle) that was assessed for methylation by bisulfite sequencing. (b-d) 3 additional genes which exhibit differential methylation between ES cells and IMR90 cells, and which lack epigenetic reprogramming in ips cells (Supplementary Data 2) (b) CSRP1, (c) TRIM4, (d) COMT. 6
7 Supplementary Figure 7 a b c Figure S7 DNA methylation and incompletely reprogrammed genes. (a) The high expression level of C9orf64 and TRIM4 in somatic cells and their ips cell counterparts, relative to ES cells (in this case, H9 ES cells), was confirmed by qrt-pcr. Values were standardized to GAPDH and Ubb. Data are from triplicate reactions. Error bars represent standard deviations. (b) The genes which maintain higher expression levels in Hep-iPS cells and Mel-iPS cells compared to ES cells tend to be also methylated at higher levels in H1 ES cells compared to the fibroblast cell line IMR90. The Pearson correlation coefficient between the log expression fold change and single-nucleotide resolution differences in CpG island methylation was 0.37 and 0.74 for Hep-iPS cells and Mel-iPS cells, respectively. (c) We performed additional bisulfite sequence analysis in 4 human ips cells (Hep-iPS 6, Adult Fib-iPS 1, BJ-RiPS 1.2 and BJ-RiPS 1.3), that were derived from different donor somatic cells and by reprogramming strategies different from those that were originally transcriptionally profiled by microarrays. Four additional ES cell lines (HSF6, ESI03, HSF12 and HSF13) were also analyzed, in parallel, as controls. The results are essentially the same as depicted in Figure 4c, while there is some variability in C9orf64 in ES cells, indicating that our findings are not clone- or methodology-dependent. Shown in the graphs is the % methylation observed at the promoters of C9orf64, TRIM4, CSRP1 and COMT in all the original cells that were transcriptionally-profiled by microarray, combined with the results from the newly analyzed ips and ES cell lines. 7
8 Figure S8 C9orf64 overexpression rescues the deficiency in ips colony number that results from C9orf64 inhibition. The number of ips colonies was counted on d21 after infection of BJ foreskin fibroblasts with 4f alone (control), 4f+non-targeting shrna (pgipz-nt) and 4f+C9orf64 shrna (pgipz-c9orf64 i1/2/3). For each condition, C9orf64 was also overexpressed by plove-c9orf64 lentivirus infection, and plove-gfp was used as a negative control. Infections were performed in triplicate, and error bars represent standard deviations. C9orf64 overexpression resulted in a significant rescue of the reduction in number of ips colonies by C9orf64 shrna2 and shrna3, which target the UTR region of the C9orf64 mrna. No rescue was observed for C9orf64 shrna1, which targets the ORF region of the C9orf64 mrna. 8
9 Supplementary Table 1 Gene RefSeq Log 2 (Fib/ES) expression change CpG ES>IMR90 CpG IMR90>ES C9orf64 NM_ TRIM4 NM_ DNAJA4 NM_ IQCA1 NM_ COMT NM_ CES1 NM_ Supplementary Table 1. 6 genes differentially expressed between ESC and all 3 ipsc. DEDS 5% FDR cutoff was used to determine differential expression. 9
10 Supplementary Table 2 Gene name Coordinates(hg18 version) Bisulfite primer names Sequence C9orf64 chr9:85,761,380-85,761,819 Forward TGTAGTTAAGGTAAAGGTTTTTTTT Reverse ACTCAATCCTCAACACCCAAATCTAC CSRP1 chr1:199,742, ,742,959 Forward GTGTTTAGGAAGTTTAGGAAGGTT Reverse CAATATACAAAACCCACTAATTAAC TRIM4 chr7:99,354,907-99,355,386 Forward ATAGTTTAGGTAGATGGGGTAGGTTAATTT Reverse CCTAAACCCTCAAACTTAAAAAAAA COMT chr22:18,309,072-18,309,357 Forward TTTGAGTAAGATTAGATTAAGAGGT Reverse ACAACCCTAACTACCCCAAAAACCC Supplementary Table 2. Sequences for bisulfite primers used for methylation analysis. 10
11 Supplementary Table 3 Gene name Forward primer sequence Reverse primer sequence GAPDH CAATGACCCCTTCATTGACC GACAAGCTTCCCGTTCTCAG Ubb TTGTTGGGTGAGCTTGTTTG GTCTTGCCGGTAAGGGTTTT TBP TGTGCACAGGAGCCAAGAGT ATTTTCTTGCTGCCAGTCTGG C9orf64 AGTGGGTACTGGTCCCTGTG GTCGCGTAGTACGAGGCACT Endogenous OCT4 TGTACTCCTCGGTCCCTTTC TCCAGGTTTTCTTTCCCTAGC Endogenous SOX2 GCTAGTCTCCAAGCGACGAA GCAAGAAGCCTCTCCTTGAA Endogenous cmyc CGGAACTCTTGTGCGTAAGG CTCAGCCAAGGTTGTGAGGT Endogenous KLF4 TATGACCCACACTGCCAGAA TGGGAACTTGACCATGATTG Endogenous NANOG CAGTCTGGACACTGGCTGAA CTCGCTGATTAGGCTCCAAC Lentiviral OCT4 CCCCTGTCTCTGTCACCACT CCACATAGCGTAAAAGGAGCA Lentiviral SOX2 ACACTGCCCCTCTCACACAT CATAGCGTAAAAGGAGCAACA Lentiviral cmyc AAGAGGACTTGTTGCGGAAA TTGTAATCCAGAGGTTGATTATCG Lentiviral KLF4 GACCACCTCGCCTTACACAT CATAGCGTAAAAGGAGCAACA Lentiviral NANOG ACATGCAACCTGAAGACGTG CACATAGCGTAAAAGGAGCAA TRIM4 GAAGTGAAGAACGCCACACA TCAACCAGGAAGTTGTGCAG SOX17 GGCGCAGCAGAATCCAGA CCACGACTTGCCCAGCAT FOXA2 GGGAGCGGTGAAGATGGA TCATGTTGCTCACGGAGGAGTA MSX1 CGAGAGGACCCCGTGGATGCAGAG GGCGGCCATCTTCAGCTTCTCCAG BRACHYURY TGCTTCCCTGAGACCCAGTT GATCACTTCTTTCCTTTGCATCAAG NCAM AGGAGACAGAAACGAAGCCA GGTGTTGGAAATGCTCTGGT SOX1 ATGCACCGCTACGACATGG CTCATGTAGCCCTGCGAGTTG OCT4 T GCATAGTCGCTGCTTGATCG TGGGCTCGAGAACCATGTG CXCR4 CACCGCATCTGGAGAACCA GCCCATTTCCTCGGTGTAGTT SOX7 ACGCCGAGCTCAGCAAGAT TCCACGTACGGCCTCTTCTG HNF1B TCACAGATACCAGCAGCATCAGT GGGCATCACCAGGCTTGTA HNF4A CATGGCCAAGATTGACAACCT TTCCCATATGTTCCTGCATCAG Supplementary Table 3. Sequences for qrt-pcr primers used in our study. *OCT4 T primer set was used only for the targeted differentiation analysis. 11
Xeno-Free Systems for hesc & hipsc. Facilitating the shift from Stem Cell Research to Clinical Applications
 Xeno-Free Systems for hesc & hipsc Facilitating the shift from Stem Cell Research to Clinical Applications NutriStem Defined, xeno-free (XF), serum-free media (SFM) specially formulated for growth and
Xeno-Free Systems for hesc & hipsc Facilitating the shift from Stem Cell Research to Clinical Applications NutriStem Defined, xeno-free (XF), serum-free media (SFM) specially formulated for growth and
Replacement of Oct4 by Tet1 during ipsc Induction Reveals an Important Role of DNA Methylation and Hydroxymethylation in Reprogramming
 Hydroxymethylation in Reprogramming, (2013), http://dx.doi.org/10.1016/j.stem.2013.02.005 Article Replacement of Oct4 by Tet1 during ipsc Induction Reveals an Important Role of DNA Methylation and Hydroxymethylation
Hydroxymethylation in Reprogramming, (2013), http://dx.doi.org/10.1016/j.stem.2013.02.005 Article Replacement of Oct4 by Tet1 during ipsc Induction Reveals an Important Role of DNA Methylation and Hydroxymethylation
Chemically defined conditions for human ipsc derivation and culture
 Nature Methods Chemically defined conditions for human ipsc derivation and culture Guokai Chen, Daniel R Gulbranson, Zhonggang Hou, Jennifer M Bolin, Victor Ruotti, Mitchell D Probasco, Kimberly Smuga-Otto,
Nature Methods Chemically defined conditions for human ipsc derivation and culture Guokai Chen, Daniel R Gulbranson, Zhonggang Hou, Jennifer M Bolin, Victor Ruotti, Mitchell D Probasco, Kimberly Smuga-Otto,
Fundamental properties of Stem Cells
 Stem cells Learning Goals: Define what a stem cell is and describe its general properties, using hematopoietic stem cells as an example. Describe to a non-scientist the current progress of human stem cell
Stem cells Learning Goals: Define what a stem cell is and describe its general properties, using hematopoietic stem cells as an example. Describe to a non-scientist the current progress of human stem cell
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature11435 Supplementary Figures Figure S1. The scheme of androgenetic haploid embryonic stem (ahes) cells derivation. WWW.NATURE.COM/NATURE 1 RESEARCH SUPPLEMENTARY INFORMATION Figure S2.
doi:10.1038/nature11435 Supplementary Figures Figure S1. The scheme of androgenetic haploid embryonic stem (ahes) cells derivation. WWW.NATURE.COM/NATURE 1 RESEARCH SUPPLEMENTARY INFORMATION Figure S2.
SUPPLEMENTAL METHODS Reverse transcriptase PCR (RT-PCR) and quantitative real-time PCR (Q-RT-PCR) RNA was isolated with TRIZOL (Invitrogen), and
 SUPPLEMENTAL METHODS Reverse transcriptase PCR (RT-PCR) and quantitative real-time PCR (Q-RT-PCR) RNA was isolated with TRIZOL (Invitrogen), and first strand cdna was synthesized using 1 μg of RNA and
SUPPLEMENTAL METHODS Reverse transcriptase PCR (RT-PCR) and quantitative real-time PCR (Q-RT-PCR) RNA was isolated with TRIZOL (Invitrogen), and first strand cdna was synthesized using 1 μg of RNA and
John Gurdon was testing the hypothesis of genomic equivalence or that when cells divide they retain a full genomic compliment.
 1. (15 pts) John Gurdon won the 2012 Nobel Prize in Physiology or Medicine for work he did in the 1960 s. What was the major developmental hypothesis he set out to test? What techniques did he development
1. (15 pts) John Gurdon won the 2012 Nobel Prize in Physiology or Medicine for work he did in the 1960 s. What was the major developmental hypothesis he set out to test? What techniques did he development
SUPPLEMENTARY INFORMATION
 a before amputation regeneration regenerated limb DERMIS SKELETON MUSCLE SCHWANN CELLS EPIDERMIS DERMIS SKELETON MUSCLE SCHWANN CELLS EPIDERMIS developmental origin: lateral plate mesoderm presomitic mesoderm
a before amputation regeneration regenerated limb DERMIS SKELETON MUSCLE SCHWANN CELLS EPIDERMIS DERMIS SKELETON MUSCLE SCHWANN CELLS EPIDERMIS developmental origin: lateral plate mesoderm presomitic mesoderm
Directed Differentiation of Human Induced Pluripotent Stem Cells. into Fallopian Tube Epithelium
 Directed Differentiation of Human Induced Pluripotent Stem Cells into Fallopian Tube Epithelium Nur Yucer 1,2, Marie Holzapfel 2, Tilley Jenkins Vogel 2, Lindsay Lenaeus 1, Loren Ornelas 1, Anna Laury
Directed Differentiation of Human Induced Pluripotent Stem Cells into Fallopian Tube Epithelium Nur Yucer 1,2, Marie Holzapfel 2, Tilley Jenkins Vogel 2, Lindsay Lenaeus 1, Loren Ornelas 1, Anna Laury
A15871 A15872 A15876 A15870
 TaqMan hpsc Scorecard Kits TaqMan hpsc Scorecard Panels A15871 A15872 A15876 A15870 Product Qty Cat. No. TaqMan hpsc Scorecard Kit 2 x 96w FAST 1 kit A15871 TaqMan hpsc Scorecard Kit 384w 1 kit A15872
TaqMan hpsc Scorecard Kits TaqMan hpsc Scorecard Panels A15871 A15872 A15876 A15870 Product Qty Cat. No. TaqMan hpsc Scorecard Kit 2 x 96w FAST 1 kit A15871 TaqMan hpsc Scorecard Kit 384w 1 kit A15872
hesc Otx2 B-T Sox17 Actin M r (K)
 DOI:.38/ncb357 In the format provided by the authors and unedited. a Nanog OCT b hesc hipsc hipsc SSEA TRA -6 c hesc hipsc Nanog OCT Otx hesc SSEA TRA -6 B-T d f Sox7 M r (K) M r (K) Actin Actin e OCR
DOI:.38/ncb357 In the format provided by the authors and unedited. a Nanog OCT b hesc hipsc hipsc SSEA TRA -6 c hesc hipsc Nanog OCT Otx hesc SSEA TRA -6 B-T d f Sox7 M r (K) M r (K) Actin Actin e OCR
Supplementary Fig. 1 related to Fig. 1 Clinical relevance of lncrna candidate
 Supplementary Figure Legends Supplementary Fig. 1 related to Fig. 1 Clinical relevance of lncrna candidate BC041951 in gastric cancer. (A) The flow chart for selected candidate lncrnas in 660 up-regulated
Supplementary Figure Legends Supplementary Fig. 1 related to Fig. 1 Clinical relevance of lncrna candidate BC041951 in gastric cancer. (A) The flow chart for selected candidate lncrnas in 660 up-regulated
What we ll do today. Types of stem cells. Do engineered ips and ES cells have. What genes are special in stem cells?
 Do engineered ips and ES cells have similar molecular signatures? What we ll do today Research questions in stem cell biology Comparing expression and epigenetics in stem cells asuring gene expression
Do engineered ips and ES cells have similar molecular signatures? What we ll do today Research questions in stem cell biology Comparing expression and epigenetics in stem cells asuring gene expression
Do engineered ips and ES cells have similar molecular signatures?
 Do engineered ips and ES cells have similar molecular signatures? Comparing expression and epigenetics in stem cells George Bell, Ph.D. Bioinformatics and Research Computing 2012 Spring Lecture Series
Do engineered ips and ES cells have similar molecular signatures? Comparing expression and epigenetics in stem cells George Bell, Ph.D. Bioinformatics and Research Computing 2012 Spring Lecture Series
Supplementary Figure 1: Derivation and characterization of RN ips cell lines. (a) RN ips cells maintain expression of pluripotency markers OCT4 and
 Supplementary Figure 1: Derivation and characterization of RN ips cell lines. (a) RN ips cells maintain expression of pluripotency markers OCT4 and SSEA4 after 10 passages in mtesr 1 medium. (b) Schematic
Supplementary Figure 1: Derivation and characterization of RN ips cell lines. (a) RN ips cells maintain expression of pluripotency markers OCT4 and SSEA4 after 10 passages in mtesr 1 medium. (b) Schematic
Supplemental Information
 Cell Stem Cell, Volume 15 Supplemental Information Generation of Naive Induced Pluripotent Stem Cells from Rhesus Monkey Fibroblasts Riguo Fang, Kang Liu, Yang Zhao, Haibo Li, Dicong Zhu, Yuanyuan Du,
Cell Stem Cell, Volume 15 Supplemental Information Generation of Naive Induced Pluripotent Stem Cells from Rhesus Monkey Fibroblasts Riguo Fang, Kang Liu, Yang Zhao, Haibo Li, Dicong Zhu, Yuanyuan Du,
Supporting Information
 Supporting Information Ho et al. 1.173/pnas.81288816 SI Methods Sequences of shrna hairpins: Brg shrna #1: ccggcggctcaagaaggaagttgaactcgagttcaacttccttcttgacgnttttg (TRCN71383; Open Biosystems). Brg shrna
Supporting Information Ho et al. 1.173/pnas.81288816 SI Methods Sequences of shrna hairpins: Brg shrna #1: ccggcggctcaagaaggaagttgaactcgagttcaacttccttcttgacgnttttg (TRCN71383; Open Biosystems). Brg shrna
HUMAN EMBRYONIC STEM CELL (hesc) ASSESSMENT BY REAL-TIME POLYMERASE CHAIN REACTION (PCR)
 HUMAN EMBRYONIC STEM CELL (hesc) ASSESSMENT BY REAL-TIME POLYMERASE CHAIN REACTION (PCR) OBJECTIVE: is performed to analyze gene expression of human Embryonic Stem Cells (hescs) in culture. Real-Time PCR
HUMAN EMBRYONIC STEM CELL (hesc) ASSESSMENT BY REAL-TIME POLYMERASE CHAIN REACTION (PCR) OBJECTIVE: is performed to analyze gene expression of human Embryonic Stem Cells (hescs) in culture. Real-Time PCR
A Murine ESC-like State Facilitates Transgenesis and Homologous Recombination in Human Pluripotent Stem Cells
 Article A Murine ESC-like State Facilitates Transgenesis and Homologous Recombination in Human Pluripotent Stem Cells Christa Buecker, 1,4 Hsu-Hsin Chen, 1,4 Jose Maria Polo, 1,2,4 Laurence Daheron, 1,4
Article A Murine ESC-like State Facilitates Transgenesis and Homologous Recombination in Human Pluripotent Stem Cells Christa Buecker, 1,4 Hsu-Hsin Chen, 1,4 Jose Maria Polo, 1,2,4 Laurence Daheron, 1,4
Single cell transcriptional profiling reveals heterogeneity of human induced pluripotent stem cells
 Brief report Single cell transcriptional profiling reveals heterogeneity of human induced pluripotent stem cells Kazim H. Narsinh, 1,2,3 Ning Sun, 1,2 Veronica Sanchez-Freire, 1,2 Andrew S. Lee, 1,2 Patricia
Brief report Single cell transcriptional profiling reveals heterogeneity of human induced pluripotent stem cells Kazim H. Narsinh, 1,2,3 Ning Sun, 1,2 Veronica Sanchez-Freire, 1,2 Andrew S. Lee, 1,2 Patricia
Guided differentiation of ES cell into mesenchymal stem cell
 Guided differentiation of ES cell into mesenchymal stem cell Takumi Era Division of Molecular Neurobiology Institute of Molecular Embryology and Genetics, Kumamoto University Differentiation pathways of
Guided differentiation of ES cell into mesenchymal stem cell Takumi Era Division of Molecular Neurobiology Institute of Molecular Embryology and Genetics, Kumamoto University Differentiation pathways of
Microarrays and Stem Cells
 Microarray Background Information Microarrays and Stem Cells Stem cells are the building blocks that allow the body to produce new cells and repair tissues. Scientists are actively investigating the potential
Microarray Background Information Microarrays and Stem Cells Stem cells are the building blocks that allow the body to produce new cells and repair tissues. Scientists are actively investigating the potential
DB3230 Midterm 1 11/15/2013 Name:
 1. (15 pts) Nuclear cloning by John Gurdon was rarely successful in producing fertile adults. Why not? Explain why serial transplantation improves the success rate. What else could you do to improve the
1. (15 pts) Nuclear cloning by John Gurdon was rarely successful in producing fertile adults. Why not? Explain why serial transplantation improves the success rate. What else could you do to improve the
camp and EPAC Signaling Functionally Replace OCT4 During Induced Pluripotent Stem Cell Reprogramming
 original article camp and EPAC Signaling Functionally Replace OCT4 During Induced Pluripotent Stem Cell Reprogramming Ashley L Fritz 1, Maroof M Adil 1, Sunnie R Mao 1 and David V Schaffer 1 3 1 Department
original article camp and EPAC Signaling Functionally Replace OCT4 During Induced Pluripotent Stem Cell Reprogramming Ashley L Fritz 1, Maroof M Adil 1, Sunnie R Mao 1 and David V Schaffer 1 3 1 Department
The Effects of Serum Starvation on Cell Cycle Synchronization
 OSR Journal of Student Research Volume 1 Article 4 2013 The Effects of Serum Starvation on Cell Cycle Synchronization Negin Baghdadchi California State University, San Bernardino Follow this and additional
OSR Journal of Student Research Volume 1 Article 4 2013 The Effects of Serum Starvation on Cell Cycle Synchronization Negin Baghdadchi California State University, San Bernardino Follow this and additional
ADVANCED MEDIA TECHNOLOGY
 ADVANCED MEDIA TECHNOLOGY For Human ES and ips Cell Research The right medium makes all the difference in ensuring successful ES and ips cell culture. Stemgent offers high-quality, chemically-defined,
ADVANCED MEDIA TECHNOLOGY For Human ES and ips Cell Research The right medium makes all the difference in ensuring successful ES and ips cell culture. Stemgent offers high-quality, chemically-defined,
Supplementary Figure 1
 Supplementary Figure 1 Supplementary Figure 1: Vector maps of TRMPV and TRMPVIR variants. Many derivatives of TRMPV have been generated and tested. Unless otherwise noted, experiments in this paper use
Supplementary Figure 1 Supplementary Figure 1: Vector maps of TRMPV and TRMPVIR variants. Many derivatives of TRMPV have been generated and tested. Unless otherwise noted, experiments in this paper use
Heterokaryon-Based Reprogramming of Human B Lymphocytes for Pluripotency Requires Oct4 but Not Sox2
 Heterokaryon-Based Reprogramming of Human B Lymphocytes for Pluripotency Requires Oct4 but Not Sox2 Carlos F. Pereira 1,Rémi Terranova 1, Natalie K. Ryan 1, Joana Santos 1, Kelly J. Morris 1, Wei Cui 2,
Heterokaryon-Based Reprogramming of Human B Lymphocytes for Pluripotency Requires Oct4 but Not Sox2 Carlos F. Pereira 1,Rémi Terranova 1, Natalie K. Ryan 1, Joana Santos 1, Kelly J. Morris 1, Wei Cui 2,
Stem Cell Services. Driving Innovation for Stem Cell Researchers
 Driving Innovation for Stem Cell Researchers Stem Cell Services Partner with us and have access to the most advanced and comprehensive stem cell services available today. 675 W. Kendall St. Cambridge,
Driving Innovation for Stem Cell Researchers Stem Cell Services Partner with us and have access to the most advanced and comprehensive stem cell services available today. 675 W. Kendall St. Cambridge,
Effects of Multiple Probesets in Affymetrix GeneChips on Identifying Differentially Expressed Genes in ips Cells
 The Fourth International Conference on Computational Systems Biology (ISB2010) Suzhou, China, September 9 11, 2010 Copyright 2010 ORSC & APORC, pp. 187 195 Effects of Multiple Probesets in Affymetrix GeneChips
The Fourth International Conference on Computational Systems Biology (ISB2010) Suzhou, China, September 9 11, 2010 Copyright 2010 ORSC & APORC, pp. 187 195 Effects of Multiple Probesets in Affymetrix GeneChips
Chapter 20: Biotechnology
 Name Period The AP Biology exam has reached into this chapter for essay questions on a regular basis over the past 15 years. Student responses show that biotechnology is a difficult topic. This chapter
Name Period The AP Biology exam has reached into this chapter for essay questions on a regular basis over the past 15 years. Student responses show that biotechnology is a difficult topic. This chapter
qbiomarker PCR Array Handbook
 March 2011 qbiomarker PCR Array Handbook qbiomarker Screening PCR Array qbiomarker Validation PCR Array For induced pluripotent stem cell screening, validation, and differentiation Sample & Assay Technologies
March 2011 qbiomarker PCR Array Handbook qbiomarker Screening PCR Array qbiomarker Validation PCR Array For induced pluripotent stem cell screening, validation, and differentiation Sample & Assay Technologies
Supplementary Information
 Supplementary Figure 1. Inverse correlation between MeDIP-seq and MRE-seq signals: (a) sperm, (b) 2.5 hpf, (c) 3.5 hpf, (d) 4.5 hpf, (e) 6 hpf and (f) 24 hpf embryos. a Sperm b 2.5 hpf c 3.5 hpf d 4.5
Supplementary Figure 1. Inverse correlation between MeDIP-seq and MRE-seq signals: (a) sperm, (b) 2.5 hpf, (c) 3.5 hpf, (d) 4.5 hpf, (e) 6 hpf and (f) 24 hpf embryos. a Sperm b 2.5 hpf c 3.5 hpf d 4.5
A simple choice. Essential 8 Media
 A simple choice Essential 8 Media Your stem cells thrive with the essentials Gibco Essential 8 Medium is a feeder-free, xeno-free medium originally developed in the laboratory of stem cell research pioneer
A simple choice Essential 8 Media Your stem cells thrive with the essentials Gibco Essential 8 Medium is a feeder-free, xeno-free medium originally developed in the laboratory of stem cell research pioneer
Cloning. 1. What is cloning: Natural and artificial 2. Cloning of what? 3. Embryonic development of multi-cellular organisms:
 Cloning 1. What is cloning: Natural and artificial 2. Cloning of what? 3. Embryonic development of multi-cellular organisms: cell division, morphogenesis, differentiation 2. Plant cloning 3. Animal cloning
Cloning 1. What is cloning: Natural and artificial 2. Cloning of what? 3. Embryonic development of multi-cellular organisms: cell division, morphogenesis, differentiation 2. Plant cloning 3. Animal cloning
ANAT 2341 Embryology Lecture 18 Stem Cells
 ANAT 2341 Embryology Lecture 18 Stem Cells 29 September 2010 Dr Antonio Lee Neuromuscular & Regenera
ANAT 2341 Embryology Lecture 18 Stem Cells 29 September 2010 Dr Antonio Lee Neuromuscular & Regenera
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb2774 Figure S1 TRF2 dosage modulates the tumorigenicity of mouse and human tumor cells. (a) Left: immunoblotting with antibodies directed against the Myc tag of the transduced TRF2 forms
DOI: 10.1038/ncb2774 Figure S1 TRF2 dosage modulates the tumorigenicity of mouse and human tumor cells. (a) Left: immunoblotting with antibodies directed against the Myc tag of the transduced TRF2 forms
Agilent Seahorse XF Live-Cell Metabolism Solutions for STEM CELL RESEARCH
 Agilent Seahorse XF Live-Cell Metabolism Solutions for STEM CELL RESEARCH MONITOR CELL FATE TRANSITIONS Improve Differentiation and Reprogramming Outcomes Adult Fibroblast Cells Reprogramming (ips Cells)
Agilent Seahorse XF Live-Cell Metabolism Solutions for STEM CELL RESEARCH MONITOR CELL FATE TRANSITIONS Improve Differentiation and Reprogramming Outcomes Adult Fibroblast Cells Reprogramming (ips Cells)
A cost-effective system for differentiation of intestinal epithelium from human induced pluripotent stem cells
 A cost-effective system for differentiation of intestinal epithelium from human induced pluripotent stem cells Soichiro Ogaki 1, 2, 3, 4, 5, *, ayu orooka 1,*, Kaito Otera 1, 1, 3 and Shoen Kume Supplementary
A cost-effective system for differentiation of intestinal epithelium from human induced pluripotent stem cells Soichiro Ogaki 1, 2, 3, 4, 5, *, ayu orooka 1,*, Kaito Otera 1, 1, 3 and Shoen Kume Supplementary
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1.
 Supplementary Figure 1. Characterization and high expression of Lnc-β-Catm in liver CSCs. (a) Heatmap of differently expressed lncrnas in Liver CSCs (CD13 + CD133 + ) and non-cscs (CD13 - CD133 - ) according
Supplementary Figure 1. Characterization and high expression of Lnc-β-Catm in liver CSCs. (a) Heatmap of differently expressed lncrnas in Liver CSCs (CD13 + CD133 + ) and non-cscs (CD13 - CD133 - ) according
Introducing new DNA into the genome requires cloning the donor sequence, delivery of the cloned DNA into the cell, and integration into the genome.
 Key Terms Chapter 32: Genetic Engineering Cloning describes propagation of a DNA sequence by incorporating it into a hybrid construct that can be replicated in a host cell. A cloning vector is a plasmid
Key Terms Chapter 32: Genetic Engineering Cloning describes propagation of a DNA sequence by incorporating it into a hybrid construct that can be replicated in a host cell. A cloning vector is a plasmid
Nature Protocols: doi: /nprot Supplementary Figure 1. Efficient genome editing in human pluripotent stem cells.
 Supplementary Figure 1 Efficient genome editing in human pluripotent stem cells. Our genome editing protocol was highly efficient in an additional pluripotent human cell line (CHB10) and at several different
Supplementary Figure 1 Efficient genome editing in human pluripotent stem cells. Our genome editing protocol was highly efficient in an additional pluripotent human cell line (CHB10) and at several different
Gene Expression Technology
 Gene Expression Technology Bing Zhang Department of Biomedical Informatics Vanderbilt University bing.zhang@vanderbilt.edu Gene expression Gene expression is the process by which information from a gene
Gene Expression Technology Bing Zhang Department of Biomedical Informatics Vanderbilt University bing.zhang@vanderbilt.edu Gene expression Gene expression is the process by which information from a gene
Single-cell RNA-seq reveals novel regulators of human embryonic stem cell differentiation to definitive endoderm
 Chu et al. Genome Biology (2016) 17:173 DOI 10.1186/s13059-016-1033-x RESEARCH Open Access Single-cell RNA-seq reveals novel regulators of human embryonic stem cell differentiation to definitive endoderm
Chu et al. Genome Biology (2016) 17:173 DOI 10.1186/s13059-016-1033-x RESEARCH Open Access Single-cell RNA-seq reveals novel regulators of human embryonic stem cell differentiation to definitive endoderm
Concepts and Methods in Developmental Biology
 Biology 4361 Developmental Biology Concepts and Methods in Developmental Biology June 16, 2009 Conceptual and Methodological Tools Concepts Genomic equivalence Differential gene expression Differentiation/de-differentiation
Biology 4361 Developmental Biology Concepts and Methods in Developmental Biology June 16, 2009 Conceptual and Methodological Tools Concepts Genomic equivalence Differential gene expression Differentiation/de-differentiation
SPERMATOGENESIS FROM STEM CELLS
 SPERMATOGENESIS FROM STEM CELLS Anna Veiga Centre de Medicina Regenerativa de Barcelona Hospital Universitari Quiron Dexeus. Barcelona Nothing to disclose Disclosure Hochedlinger, Development 2009 (adapted
SPERMATOGENESIS FROM STEM CELLS Anna Veiga Centre de Medicina Regenerativa de Barcelona Hospital Universitari Quiron Dexeus. Barcelona Nothing to disclose Disclosure Hochedlinger, Development 2009 (adapted
NPTEL Biotechnology Tissue Engineering. Stem cells
 Stem cells S. Swaminathan Director Centre for Nanotechnology & Advanced Biomaterials School of Chemical & Biotechnology SASTRA University Thanjavur 613 401 Tamil Nadu Joint Initiative of IITs and IISc
Stem cells S. Swaminathan Director Centre for Nanotechnology & Advanced Biomaterials School of Chemical & Biotechnology SASTRA University Thanjavur 613 401 Tamil Nadu Joint Initiative of IITs and IISc
Genetic Basis of Development & Biotechnologies
 Genetic Basis of Development & Biotechnologies 1. Steps of embryonic development: cell division, morphogenesis, differentiation Totipotency and pluripotency 2. Plant cloning 3. Animal cloning Reproductive
Genetic Basis of Development & Biotechnologies 1. Steps of embryonic development: cell division, morphogenesis, differentiation Totipotency and pluripotency 2. Plant cloning 3. Animal cloning Reproductive
Best Practices Maintaining Pluripotency. Anne R. Greenlee, PhD
 Best Practices Maintaining Pluripotency Anne R. Greenlee, PhD HESI-Workshop Cary, NC 27 Feb 2007 OHSU La Grande Campus Research Directions Past Fertility Risk Factor Study Pesticide effects on embryo development
Best Practices Maintaining Pluripotency Anne R. Greenlee, PhD HESI-Workshop Cary, NC 27 Feb 2007 OHSU La Grande Campus Research Directions Past Fertility Risk Factor Study Pesticide effects on embryo development
Quantitative Real Time PCR USING SYBR GREEN
 Quantitative Real Time PCR USING SYBR GREEN SYBR Green SYBR Green is a cyanine dye that binds to double stranded DNA. When it is bound to D.S. DNA it has a much greater fluorescence than when bound to
Quantitative Real Time PCR USING SYBR GREEN SYBR Green SYBR Green is a cyanine dye that binds to double stranded DNA. When it is bound to D.S. DNA it has a much greater fluorescence than when bound to
Activity 47.1 What common events occur in the early development of animals? 1. What key events occur at each stage of development?
 Notes to Instructors Chapter 47 Animal Development What is the focus of this activity? Chapter 21 provided a review of how genes act to control development. Chapter 47 reviews some of the major morphological
Notes to Instructors Chapter 47 Animal Development What is the focus of this activity? Chapter 21 provided a review of how genes act to control development. Chapter 47 reviews some of the major morphological
Naked Mole Rat Induced Pluripotent Stem Cells and Their Contribution
 Stem Cell eports, Volume 9 Supplemental Information Naked Mole at Induced Pluripotent Stem Cells and Their Contribution to Interspecific Chimera Sang-Goo Lee, Aleksei E. Mikhalchenko, Sun Hee Yim, Alexei
Stem Cell eports, Volume 9 Supplemental Information Naked Mole at Induced Pluripotent Stem Cells and Their Contribution to Interspecific Chimera Sang-Goo Lee, Aleksei E. Mikhalchenko, Sun Hee Yim, Alexei
V4: cellular reprogramming
 V4: cellular reprogramming Embryonic development Stem cells Differentiation ips cells Oct4 Paper4 Model of a human cell, Spiegel Online 3D-Model of the human genome, Spiegel Online Some human cells Astrocyte
V4: cellular reprogramming Embryonic development Stem cells Differentiation ips cells Oct4 Paper4 Model of a human cell, Spiegel Online 3D-Model of the human genome, Spiegel Online Some human cells Astrocyte
Human embryonic stem cells for in-vitro developmental toxicity testing
 Human embryonic stem cells for in-vitro developmental toxicity testing HESI workshop on alternatives assays for developmental toxicity Raimund Strehl Cellartis The company Founded in early 2001, University
Human embryonic stem cells for in-vitro developmental toxicity testing HESI workshop on alternatives assays for developmental toxicity Raimund Strehl Cellartis The company Founded in early 2001, University
A molecular roadmap for the emergence of early-embryonic-like cells in culture
 SUPPLEMENTARY INFORMATION Articles https://doi.org/10.1038/s41588-017-0016-5 In the format provided by the authors and unedited. A molecular roadmap for the emergence of early-embryonic-like cells in culture
SUPPLEMENTARY INFORMATION Articles https://doi.org/10.1038/s41588-017-0016-5 In the format provided by the authors and unedited. A molecular roadmap for the emergence of early-embryonic-like cells in culture
Supplementary Figure 1
 number of cells, normalized number of cells, normalized number of cells, normalized Supplementary Figure CD CD53 Cd3e fluorescence intensity fluorescence intensity fluorescence intensity Supplementary
number of cells, normalized number of cells, normalized number of cells, normalized Supplementary Figure CD CD53 Cd3e fluorescence intensity fluorescence intensity fluorescence intensity Supplementary
GENESDEV/2007/ Supplementary Figure 1 Elkabetz et al.,
 GENESDEV/2007/089581 Supplementary Figure 1 Elkabetz et al., GENESDEV/2007/089581 Supplementary Figure 2 Elkabetz et al., GENESDEV/2007/089581 Supplementary Figure 3 Elkabetz et al., GENESDEV/2007/089581
GENESDEV/2007/089581 Supplementary Figure 1 Elkabetz et al., GENESDEV/2007/089581 Supplementary Figure 2 Elkabetz et al., GENESDEV/2007/089581 Supplementary Figure 3 Elkabetz et al., GENESDEV/2007/089581
Supplementary Figures Montero et al._supplementary Figure 1
 Montero et al_suppl. Info 1 Supplementary Figures Montero et al._supplementary Figure 1 Montero et al_suppl. Info 2 Supplementary Figure 1. Transcripts arising from the structurally conserved subtelomeres
Montero et al_suppl. Info 1 Supplementary Figures Montero et al._supplementary Figure 1 Montero et al_suppl. Info 2 Supplementary Figure 1. Transcripts arising from the structurally conserved subtelomeres
Cell culture and drug treatment. Lineage - Sca-1+ CD31+ EPCs were cultured on
 Supplemental Material Detailed Methods Cell culture and drug treatment. Lineage - Sca-1+ CD31+ EPCs were cultured on 5µg/mL human fibronectin coated plates in DMEM supplemented with 10% FBS and penicillin/streptomycin
Supplemental Material Detailed Methods Cell culture and drug treatment. Lineage - Sca-1+ CD31+ EPCs were cultured on 5µg/mL human fibronectin coated plates in DMEM supplemented with 10% FBS and penicillin/streptomycin
Supporting Online Material for
 www.sciencemag.org/cgi/content/full/1137999/dc1 Supporting Online Material for Disrupting the Pairing Between let-7 and Enhances Oncogenic Transformation Christine Mayr, Michael T. Hemann, David P. Bartel*
www.sciencemag.org/cgi/content/full/1137999/dc1 Supporting Online Material for Disrupting the Pairing Between let-7 and Enhances Oncogenic Transformation Christine Mayr, Michael T. Hemann, David P. Bartel*
Efficient Feeder-Free Episomal Reprogramming with Small Molecules
 Efficient Feeder-Free Episomal Reprogramming with Small Molecules Junying Yu*, Kevin Fongching Chau, Maxim A. Vodyanik, Jinlan Jiang, Yong Jiang Advanced Development Programs, Cellular Dynamics International,
Efficient Feeder-Free Episomal Reprogramming with Small Molecules Junying Yu*, Kevin Fongching Chau, Maxim A. Vodyanik, Jinlan Jiang, Yong Jiang Advanced Development Programs, Cellular Dynamics International,
Technical Review. Real time PCR
 Technical Review Real time PCR Normal PCR: Analyze with agarose gel Normal PCR vs Real time PCR Real-time PCR, also known as quantitative PCR (qpcr) or kinetic PCR Key feature: Used to amplify and simultaneously
Technical Review Real time PCR Normal PCR: Analyze with agarose gel Normal PCR vs Real time PCR Real-time PCR, also known as quantitative PCR (qpcr) or kinetic PCR Key feature: Used to amplify and simultaneously
supplementary information
 DOI: 1.138/ncb1839 a b Control 1 2 3 Control 1 2 3 Fbw7 Smad3 1 2 3 4 1 2 3 4 c d IGF-1 IGF-1Rβ IGF-1Rβ-P Control / 1 2 3 4 Real-time RT-PCR Relative quantity (IGF-1/ mrna) 2 1 IGF-1 1 2 3 4 Control /
DOI: 1.138/ncb1839 a b Control 1 2 3 Control 1 2 3 Fbw7 Smad3 1 2 3 4 1 2 3 4 c d IGF-1 IGF-1Rβ IGF-1Rβ-P Control / 1 2 3 4 Real-time RT-PCR Relative quantity (IGF-1/ mrna) 2 1 IGF-1 1 2 3 4 Control /
The many faces of Pluripotency: in vitro adaptations of a continuum of in vivo states
 Morgani et al. BMC Developmental Biology (2017) 17:7 DOI 10.1186/s12861-017-0150-4 REVIEW The many faces of Pluripotency: in vitro adaptations of a continuum of in vivo states Sophie Morgani 1,2, Jennifer
Morgani et al. BMC Developmental Biology (2017) 17:7 DOI 10.1186/s12861-017-0150-4 REVIEW The many faces of Pluripotency: in vitro adaptations of a continuum of in vivo states Sophie Morgani 1,2, Jennifer
Supplemental Table 1 Gene Symbol FDR corrected p-value PLOD1 CSRP2 PFKP ADFP ADM C10orf10 GPI LOX PLEKHA2 WIPF1
 Supplemental Table 1 Gene Symbol FDR corrected p-value PLOD1 4.52E-18 PDK1 6.77E-18 CSRP2 4.42E-17 PFKP 1.23E-14 MSH2 3.79E-13 NARF_A 5.56E-13 ADFP 5.56E-13 FAM13A1 1.56E-12 FAM29A_A 1.22E-11 CA9 1.54E-11
Supplemental Table 1 Gene Symbol FDR corrected p-value PLOD1 4.52E-18 PDK1 6.77E-18 CSRP2 4.42E-17 PFKP 1.23E-14 MSH2 3.79E-13 NARF_A 5.56E-13 ADFP 5.56E-13 FAM13A1 1.56E-12 FAM29A_A 1.22E-11 CA9 1.54E-11
Developmental Biology 3230 Exam 1 (Feb. 6) NAME
 DevelopmentalBiology3230Exam1(Feb.6)NAME 1. (10pts) What is a Fate Map? How would you experimentally acquire the data to draw a Fate Map? Explain what a Fate Map does and does not tell you about the mechanisms
DevelopmentalBiology3230Exam1(Feb.6)NAME 1. (10pts) What is a Fate Map? How would you experimentally acquire the data to draw a Fate Map? Explain what a Fate Map does and does not tell you about the mechanisms
DNA methylation-based body fluid typing
 DNA methylation-based body fluid typing Hwan Young Lee Department of Forensic Medicine Yonsei University College of Medicine, Seoul, Korea Forensic Body Fluid Identification Body fluid identification can
DNA methylation-based body fluid typing Hwan Young Lee Department of Forensic Medicine Yonsei University College of Medicine, Seoul, Korea Forensic Body Fluid Identification Body fluid identification can
Single Cell Analysis Reveals the Stochastic Phase of Reprogramming to Pluripotency Is an Ordered Probabilistic Process
 University of Connecticut DigitalCommons@UConn Open Access Author Fund Awardees' Articles UConn Library 4-17-2014 Single Cell Analysis Reveals the Stochastic Phase of Reprogramming to Pluripotency Is an
University of Connecticut DigitalCommons@UConn Open Access Author Fund Awardees' Articles UConn Library 4-17-2014 Single Cell Analysis Reveals the Stochastic Phase of Reprogramming to Pluripotency Is an
Supplemental Figure 1
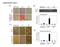 Supplemental Figure 1 A B 50 * 40 30 20 10 0 NeurofilamentM/βactin Relative expression control D neurogenic cond. PPARγ/ Relative expression C PPARγ Hprt OilRedO adipogenic cond. control 125 * 100 75 50
Supplemental Figure 1 A B 50 * 40 30 20 10 0 NeurofilamentM/βactin Relative expression control D neurogenic cond. PPARγ/ Relative expression C PPARγ Hprt OilRedO adipogenic cond. control 125 * 100 75 50
Supplemental Information. Tissue Mechanics Orchestrate Wnt-Dependent. Human Embryonic Stem Cell Differentiation
 Cell Stem Cell, Volume 19 Supplemental Information Tissue Mechanics Orchestrate Wnt-Dependent Human Embryonic Stem Cell Differentiation Laralynne Przybyla, Johnathon N. Lakins, and Valerie M. Weaver Supplemental
Cell Stem Cell, Volume 19 Supplemental Information Tissue Mechanics Orchestrate Wnt-Dependent Human Embryonic Stem Cell Differentiation Laralynne Przybyla, Johnathon N. Lakins, and Valerie M. Weaver Supplemental
Human Fibroblast Reprogramming to Pluripotent Stem Cells Regulated by the mir19a/b-pten Axis
 Human Fibroblast Reprogramming to Pluripotent Stem Cells Regulated by the mir19a/b-pten Axis Xiaoping He 1, Yang Cao 1, Lihua Wang 1, Yingli Han 1, Xiuying Zhong 1, Guixiang Zhou 2, Yongping Cai 3, Huafeng
Human Fibroblast Reprogramming to Pluripotent Stem Cells Regulated by the mir19a/b-pten Axis Xiaoping He 1, Yang Cao 1, Lihua Wang 1, Yingli Han 1, Xiuying Zhong 1, Guixiang Zhou 2, Yongping Cai 3, Huafeng
Figure S2. Response of mouse ES cells to GSK3 inhibition. Mentioned in discussion
 Stem Cell Reports, Volume 1 Supplemental Information Robust Self-Renewal of Rat Embryonic Stem Cells Requires Fine-Tuning of Glycogen Synthase Kinase-3 Inhibition Yaoyao Chen, Kathryn Blair, and Austin
Stem Cell Reports, Volume 1 Supplemental Information Robust Self-Renewal of Rat Embryonic Stem Cells Requires Fine-Tuning of Glycogen Synthase Kinase-3 Inhibition Yaoyao Chen, Kathryn Blair, and Austin
mir-24-mediated down-regulation of H2AX suppresses DNA repair
 Supplemental Online Material mir-24-mediated down-regulation of H2AX suppresses DNA repair in terminally differentiated blood cells Ashish Lal 1,4, Yunfeng Pan 2,4, Francisco Navarro 1,4, Derek M. Dykxhoorn
Supplemental Online Material mir-24-mediated down-regulation of H2AX suppresses DNA repair in terminally differentiated blood cells Ashish Lal 1,4, Yunfeng Pan 2,4, Francisco Navarro 1,4, Derek M. Dykxhoorn
 Constitutive Reprogramming Lentiviral Vectors Expressing Polycistronic Human OSKM or Monocistronic Human Oct4, Sox2, Klf4, Myc, Lin28 or Nanog for Somatic Cell Reprogramming www.vectalys.com/products/
Constitutive Reprogramming Lentiviral Vectors Expressing Polycistronic Human OSKM or Monocistronic Human Oct4, Sox2, Klf4, Myc, Lin28 or Nanog for Somatic Cell Reprogramming www.vectalys.com/products/
Endogenous mirna Sponge lincrna-ror Regulates Oct4, Nanog, and Sox2 in Human Embryonic Stem Cell Self-Renewal
 Stem Cell Self-Renewal, (2013), http://dx.doi.org/10.1016/j.devcel.2013.03.002 Article Endogenous mirna Sponge lincrna-ror Regulates Oct4, Nanog, and Sox2 in Human Embryonic Stem Cell Self-Renewal Yue
Stem Cell Self-Renewal, (2013), http://dx.doi.org/10.1016/j.devcel.2013.03.002 Article Endogenous mirna Sponge lincrna-ror Regulates Oct4, Nanog, and Sox2 in Human Embryonic Stem Cell Self-Renewal Yue
Supplementary Information Design of small molecule-responsive micrornas based on structural requirements for Drosha processing
 Supplementary Information Design of small molecule-responsive micrornas based on structural requirements for Drosha processing Chase L. Beisel, Yvonne Y. Chen, Stephanie J. Culler, Kevin G. Hoff, & Christina
Supplementary Information Design of small molecule-responsive micrornas based on structural requirements for Drosha processing Chase L. Beisel, Yvonne Y. Chen, Stephanie J. Culler, Kevin G. Hoff, & Christina
PluriQ TM Serum Replacement (PluriQ TM SR)
 PluriQ TM Serum Replacement (PluriQ TM SR) (PluriQ SR) is a defined, serum-free supplement formulated as a direct replacement for FBS (fetal bovine serum) used in the growth and maintenance of undifferentiated
PluriQ TM Serum Replacement (PluriQ TM SR) (PluriQ SR) is a defined, serum-free supplement formulated as a direct replacement for FBS (fetal bovine serum) used in the growth and maintenance of undifferentiated
Stem Cells and Cell Culture Developments Within ThermoFisher Scientific
 The world leader in serving science Stem Cells and Cell Culture Developments Within ThermoFisher Scientific Katrina Kidd European Application Scientist Cellomics ThermoFisher Scientific A Typical Life
The world leader in serving science Stem Cells and Cell Culture Developments Within ThermoFisher Scientific Katrina Kidd European Application Scientist Cellomics ThermoFisher Scientific A Typical Life
Human ips cell derived alveolar epithelium repopulates lung extracellular matrix
 Technical advance Human ips cell derived alveolar epithelium repopulates lung extracellular matrix Mahboobe Ghaedi, 1 Elizabeth A. Calle, 1 Julio J. Mendez, 1 Ashley L. Gard, 1 Jenna Balestrini, 1 Adam
Technical advance Human ips cell derived alveolar epithelium repopulates lung extracellular matrix Mahboobe Ghaedi, 1 Elizabeth A. Calle, 1 Julio J. Mendez, 1 Ashley L. Gard, 1 Jenna Balestrini, 1 Adam
Lectures 28 and 29 applications of recombinant technology I. Manipulate gene of interest
 Lectures 28 and 29 applications of recombinant technology I. Manipulate gene of interest C A. site-directed mutagenesis A C A T A DNA B. in vitro mutagenesis by PCR T A 1. anneal primer 1 C A 1. fill in
Lectures 28 and 29 applications of recombinant technology I. Manipulate gene of interest C A. site-directed mutagenesis A C A T A DNA B. in vitro mutagenesis by PCR T A 1. anneal primer 1 C A 1. fill in
Scaffolding for human pluripotent stem cell lineage specification
 University of Arkansas, Fayetteville ScholarWorks@UARK Biological and Agricultural Engineering Undergraduate Honors Theses Biological and Agricultural Engineering 5-2013 Scaffolding for human pluripotent
University of Arkansas, Fayetteville ScholarWorks@UARK Biological and Agricultural Engineering Undergraduate Honors Theses Biological and Agricultural Engineering 5-2013 Scaffolding for human pluripotent
Genetics - Problem Drill 19: Dissection of Gene Function: Mutational Analysis of Model Organisms
 Genetics - Problem Drill 19: Dissection of Gene Function: Mutational Analysis of Model Organisms No. 1 of 10 1. The mouse gene knockout is based on. (A) Homologous recombination (B) Site-specific recombination
Genetics - Problem Drill 19: Dissection of Gene Function: Mutational Analysis of Model Organisms No. 1 of 10 1. The mouse gene knockout is based on. (A) Homologous recombination (B) Site-specific recombination
Supplementary Fig. 1. Isolation and in vitro expansion of EpCAM + cholangiocytes. For collagenase perfusion, enzyme solution was injected from the
 Supplementary Fig. 1. Isolation and in vitro expansion of EpCAM + cholangiocytes. For collagenase perfusion, enzyme solution was injected from the portal vein for digesting adult livers, whereas it was
Supplementary Fig. 1. Isolation and in vitro expansion of EpCAM + cholangiocytes. For collagenase perfusion, enzyme solution was injected from the portal vein for digesting adult livers, whereas it was
Theoretical cloning project
 Theoretical cloning project Needed to get credits Make it up yourself, don't copy Possible to do in groups of 2-4 students If you need help or an idea, ask! If you have no idea what to clone, I can give
Theoretical cloning project Needed to get credits Make it up yourself, don't copy Possible to do in groups of 2-4 students If you need help or an idea, ask! If you have no idea what to clone, I can give
Culture and differentiation of embryonic stem cells. Hong-Lin Su Department of Life Sciences, National Chung-Hsing University
 Culture and differentiation of embryonic stem cells Hong-Lin Su Department of Life Sciences, National Chung-Hsing University Topics-ES cell maintenance Establishment Culture condition, including the serum,
Culture and differentiation of embryonic stem cells Hong-Lin Su Department of Life Sciences, National Chung-Hsing University Topics-ES cell maintenance Establishment Culture condition, including the serum,
Stem Cells Express, published online December 18, 2008; doi: /stemcells
 Stem Cells Express, published online December 18, 2008; doi:10.1634/stemcells.2008-1075 STEM CELLS EMBRYONIC STEM CELLS/INDUCED PLURIPOTENT STEM CELLS ips Cell Generation Using a Single Lentiviral Stem
Stem Cells Express, published online December 18, 2008; doi:10.1634/stemcells.2008-1075 STEM CELLS EMBRYONIC STEM CELLS/INDUCED PLURIPOTENT STEM CELLS ips Cell Generation Using a Single Lentiviral Stem
HUMAN INDUCED PLURIPOTENT STEM CELL DERIVED EXTRACELLULAR VESICLES REVERSE HEPATIC STELLATE CELL ACTIVATION
 HUMAN INDUCED PLURIPOTENT STEM CELL DERIVED EXTRACELLULAR VESICLES REVERSE HEPATIC STELLATE CELL ACTIVATION Dr. Davide POVERO LABORATORY of Prof. ARIEL E. FELDSTEIN, M.D. UNIVERSITY OF CALIFORNIA SAN DIEGO
HUMAN INDUCED PLURIPOTENT STEM CELL DERIVED EXTRACELLULAR VESICLES REVERSE HEPATIC STELLATE CELL ACTIVATION Dr. Davide POVERO LABORATORY of Prof. ARIEL E. FELDSTEIN, M.D. UNIVERSITY OF CALIFORNIA SAN DIEGO
Cornell Probability Summer School 2006
 Colon Cancer Cornell Probability Summer School 2006 Simon Tavaré Lecture 5 Stem Cells Potten and Loeffler (1990): A small population of relatively undifferentiated, proliferative cells that maintain their
Colon Cancer Cornell Probability Summer School 2006 Simon Tavaré Lecture 5 Stem Cells Potten and Loeffler (1990): A small population of relatively undifferentiated, proliferative cells that maintain their
An investigation of the role of integrin alpha-6 in human induced pluripotent stem cell development and pluripotency
 An investigation of the role of integrin alpha-6 in human induced pluripotent stem cell development and pluripotency Submitted by Genna Wilber Biology and English To The Honors College Oakland University
An investigation of the role of integrin alpha-6 in human induced pluripotent stem cell development and pluripotency Submitted by Genna Wilber Biology and English To The Honors College Oakland University
Int. J. Mol. Sci. 2016, 17, 1259; doi: /ijms
 S1 of S5 Supplementary Materials: Fibroblast-Derived Extracellular Matrix Induces Chondrogenic Differentiation in Human Adipose-Derived Mesenchymal Stromal/Stem Cells in Vitro Kevin Dzobo, Taegyn Turnley,
S1 of S5 Supplementary Materials: Fibroblast-Derived Extracellular Matrix Induces Chondrogenic Differentiation in Human Adipose-Derived Mesenchymal Stromal/Stem Cells in Vitro Kevin Dzobo, Taegyn Turnley,
Nature Biotechnology: doi: /nbt Supplementary Figure 1
 Supplementary Figure 1 An extended version of Figure 2a, depicting multi-model training and reverse-complement mode To use the GPU s full computational power, we train several independent models in parallel
Supplementary Figure 1 An extended version of Figure 2a, depicting multi-model training and reverse-complement mode To use the GPU s full computational power, we train several independent models in parallel
Gene Regulation Solutions. Microarrays and Next-Generation Sequencing
 Gene Regulation Solutions Microarrays and Next-Generation Sequencing Gene Regulation Solutions The Microarrays Advantage Microarrays Lead the Industry in: Comprehensive Content SurePrint G3 Human Gene
Gene Regulation Solutions Microarrays and Next-Generation Sequencing Gene Regulation Solutions The Microarrays Advantage Microarrays Lead the Industry in: Comprehensive Content SurePrint G3 Human Gene
Induced pluripotent stem cells: current progress and potential for regenerative medicine
 Review Induced pluripotent stem cells: current progress and potential for regenerative medicine Giovanni Amabile 1,2 and Alexander Meissner 1,2 1 Department of Stem Cell and Regenerative Biology, Harvard
Review Induced pluripotent stem cells: current progress and potential for regenerative medicine Giovanni Amabile 1,2 and Alexander Meissner 1,2 1 Department of Stem Cell and Regenerative Biology, Harvard
