Supplemental Figure 1
|
|
|
- Warren Haynes
- 6 years ago
- Views:
Transcription
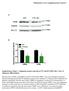
1 Supplemental Figure 1 A B 50 * NeurofilamentM/βactin Relative expression control D neurogenic cond. PPARγ/ Relative expression C PPARγ Hprt OilRedO adipogenic cond. control 125 *
2 Supplemental Figure 2 rat dental epithelium (SF2 cells) phase (SF2) GFP pef6/gfppdgftmmycha selection SF2/GFP cocultured with cells SF2/GFP cells expressing GFPmycHA tag
3 Supplemental Figure 3 A B SF224 ips Clone No. Ambn Rat dental epithelium (SF2 cell) Ambn CK14 Klf4 Sox2 Oct3/4 Nanog
4 Supplemental Figure 4 A B m totalrna r totalrna (ng) (ng) mambn ips MEF tooth germ mambn rambn
5 Supplemental Figure 5 SF224iPS (days) Ambn Epiprofin/Sp6
6 SUPPLEMENTAL INFORMATION Supplemental Figure 1. Adipogenic and neurogenic induction of cells. (A) Adipocytespecific OilRedO staining of,, and cells in regular or adipogenic induction culture (adipogenic cond.) medium. (B) RTPCR and realtime PCR analyses of the adipocyte marker gene PPARγ in,, and cells in adipogenic induction culture medium. (C) Phase images of, and cells cultured in regular or neurogenic induction culture (neurogenic cond.) medium. (D) Western blotting of,, and cells cultured in regular () or neurogenic induction culture () medium was performed using the antineurofilamentm antibody. The relative intensities of bands obtained with western blots were scanned and quantified using an LAS UV4000 system (FujifilmGE Healthcare, Tokyo, Japan). Supplemental Figure 2. Schematic diagram of establishment of SF2 cells expressing PDGFtmGFPmycHA. Ratderived dental epithelial SF2 cells were transfected with a GFPPDGFtmmycHA expression vector and selected with antibiotics. GFP in the SF2 cells was visualized using fluorescence microscopy. SF2/GFP cells were cocultured with cells (bottom) and separated with the antiha antibody. Supplemental Figure 3. Gene expression of subcloned SF2 cell lines and ips cells. (A) Ambn expression in dental epithelial cells subcloned from SF2 cells. Clone SF224 expressed Ambn at the highest level and was used for coculturing with ips cells. (B) SF224 cells expressed the ameloblast marker genes CK14 and Ambn, but not stem cell markers. In contrast, stem cell markers were observed in ips cells, but not ameloblast markers. Supplemental Figure 4. Design of mousespecific Ambn primer. Mouse and rat Ambn mrna showed a high degree of homology (94.2%), thus regular Ambn primers for RTPCR are able to precisely distinguish between them. To resolve this problem, we employed a mouse Ambn LNA primer. The accuracy of the mousespecific Ambn LNA primer was confirmed by PCR with gradient cdna mixtures containing serial dilution ratios of mouse and/or rat cdnas. (A) The expected size of the amplicon using the mouse LNA primer was observed only with mouse cdna, while the intensity of the band increased in a cdna dosedependent manner. (B) Undifferentiated ips cells and MEF feeder cells did not express Ambn mrna. A mouse tooth germ sample served as a positive control for the mouse LNA primer. Supplemental Figure 5. Expression of epiprofin/sp6 transcription factor in ips cells cocultured with SF224 cells. Time course analyses of Ambn and epiprofin/sp6 gene expressions in ips cells cocultured with SF224 cells for 1 (1D), 7 (7D) and 10 days (10D) were analyzed by RTPCR.
7 Supplemental Table1 Gene name forward primer reverse primer Size bp (mouse/rat) mouse/rat Ambn 5 GCGTTTCCAAGAGCCCTGATAAC3 5 AAGAAGCAGTGTCACATTTCCTGG3 366/368 mouse/rat CK14 5 GAGAAGAACCGCAAGGATGC3 5 AGGTTATTCTCCAGGGATGCTTTC3 191 mouse/rat 5 GGAGCGAGACCCCACTAACATC3 5 CTCGTGGTTCACACCCATCAC3 181 mouse Ambn 5 GCAGGTGGCACCATCCGALA3 5 AAGAAGCAGTGTCACATTTCCTGG rat Ambn 5 GCTACCCTTCCACAGGGAAGGTLA3 5 AAGAAGCGGTGTCACATTTCCTGG3 267 mouse Enam 5 GGGGACCACCAACAGCGTTTGGACGGCCA3 5 CCCGGAGGTAGGAGAGGGAGGTTTACC3 614 rat Enam 5 CAGGACCACCAACAGGGTTCGGACGA3 5 TCTTGAGGTAGGAGGGGAAGGTTTGT3 626 mouse CK14 5 GGCTGCCGATGACTTCCGGAC3 5 CAGCAGTATCTGCGTCCACGCA3 909 rat CK14 5 AGACTACAGCCCCTACTTCAAGACC3 5 AGGTTATTCTCCAGGGATGCTTTC3 597 mouse p63 5 GGCATTTCAGCACTATTTAGGTGG3 5 CACTGGTGTGAGGAGACAAACTGTG3 110 mouse Epiprofin 5 TCAGCCTGCTTTGGGAGGATAC3 5 ATGCTTCTTCTTGCCCCCATCG3 313 Klf4 5 CACCATGGACCCGGGCGTGGCTGCCAGAAA3 5 TTAGGCTGTTCTTTTCCGGGGCCACGA3 739 Sox2 5 GGTTACCTCTTCCTCCCACTCCAG3 5 TCACATGTGCGACAGGGGCAG3 193 Oct3/4 5 TCTTTCCACCAGGCCCCCGGCTC3 5 TGCGGGCGGACATGGGGAGATCC3 224 Nanog 5 CAGGTGTTTGAGGGTAGCTC3 5 CGGTTCATCATGGTACAGTC3 223 Fgf4 5 CGTGGTGAGCATCTTCGGAGTGG3 5 CCTTCTTGGTCCGCCCGTTCTTA3 197 Gdf3 5 GTTCCAACCTGTGCCTCGCGTCTT3 5 AGCGAGGCATGGAGAGAGCGGAGCAG3 570 Cdx2 5 GGCGAAACCTGTGCGAGTGGATGCGGAA3 5 GATTGCTGTGCCGCCGCCGCTTCAGACC3 493/490 Gata6 5 ACCTTATGGCGTAGAAATGCTGAGGGTG3 5 CTGAATACTTGAGGTCACTGTTCTCGGG3 334/328 Brachyury 5 ATGCCAAAGAAAGAAACGAC3 5 AGAGGCTGTAGAACATGATT3 835 B2 5 GGGACCCGCTGTCTTCTAGT3 5 TCAACTCAAATTCGCTGAGGAC3 154 B4 5 ACTGCCGCAGCTTCTCTGAG3 5 TTCTCCAGATGTTCTTCGTG3 486 Osteocalcin 5 CCTCTTGAAAGAGTGGGCTG3 5 CCTCGGGAGACAAACAACAT3 268 Osteonectin 5 GTCTCACTGGCTGTGTTGGA3 5 AAGACTTGCCATGTGGGTTC3 264 Runx2 5 GACGTTCCCAAGCATTTCAT3 5 ACTCTGGCTTTGGGAAGAG3 181 Dspp 5'GGAACTGCAGCACAGAATGA3' 5'CAGTGTTCCCCTGTTCGTTT3' 199/193 Bcrp1 5' GGTGAATCTCAGAACCATTGGGC3 5'GCTGTTGTCCGTTACATTGAATCC 3' 139 PPARγ 5 CTGATGCACTGCCTATGAGC3 5 CAGACTCGGCACTCAATGGC3 373 Hprt 5 GTTAAGCAGTACAGCCCCAAA3 5 AGGGCATATCCAACAACAAACTT3 131 Gapdh (Realtime) 5 CCATCACCATCTTCCAGGAG3 5 GCATGGACTGTGGTCATGAG3 322
SUPPLEMENTARY INFORMATION
 doi:.38/nature08267 Figure S: Karyotype analysis of ips cell line One hundred individual chromosomal spreads were counted for each of the ips samples. Shown here is an spread at passage 5, predominantly
doi:.38/nature08267 Figure S: Karyotype analysis of ips cell line One hundred individual chromosomal spreads were counted for each of the ips samples. Shown here is an spread at passage 5, predominantly
Supplementary Data. Supplementary Methods Three-step protocol for spontaneous differentiation of mouse induced pluripotent stem (embryonic stem) cells
 Supplementary Data Supplementary Methods Three-step protocol for spontaneous differentiation of mouse induced pluripotent stem (embryonic stem) cells Mouse induced pluripotent stem cells (ipscs) were cultured
Supplementary Data Supplementary Methods Three-step protocol for spontaneous differentiation of mouse induced pluripotent stem (embryonic stem) cells Mouse induced pluripotent stem cells (ipscs) were cultured
gacgacgaggagaccaccgctttg aggcacattgaaggtctcaaacatg
 Supplementary information Supplementary table 1: primers for cloning and sequencing cloning for E- Ras ggg aat tcc ctt gag ctg ctg ggg aat ggc ttt gcc ggt cta gag tat aaa gga agc ttt gaa tcc Tpbp Oct3/4
Supplementary information Supplementary table 1: primers for cloning and sequencing cloning for E- Ras ggg aat tcc ctt gag ctg ctg ggg aat ggc ttt gcc ggt cta gag tat aaa gga agc ttt gaa tcc Tpbp Oct3/4
SUPPLEMENTARY INFORMATION
 SUPPLEMENTRY INFORMTION DOI:.38/ncb Kdmb locus kb Long isoform Short isoform Long isoform Jmj XX PHD F-box LRR Short isoform XX PHD F-box LRR Target Vector 3xFlag Left H Neo Right H TK LoxP LoxP Kdmb Locus
SUPPLEMENTRY INFORMTION DOI:.38/ncb Kdmb locus kb Long isoform Short isoform Long isoform Jmj XX PHD F-box LRR Short isoform XX PHD F-box LRR Target Vector 3xFlag Left H Neo Right H TK LoxP LoxP Kdmb Locus
Cell proliferation was measured with Cell Counting Kit-8 (Dojindo Laboratories, Kumamoto, Japan).
 1 2 3 4 5 6 7 8 Supplemental Materials and Methods Cell proliferation assay Cell proliferation was measured with Cell Counting Kit-8 (Dojindo Laboratories, Kumamoto, Japan). GCs were plated at 96-well
1 2 3 4 5 6 7 8 Supplemental Materials and Methods Cell proliferation assay Cell proliferation was measured with Cell Counting Kit-8 (Dojindo Laboratories, Kumamoto, Japan). GCs were plated at 96-well
SUPPLEMENTARY INFORMATION
 Supplementary Figure S1 Colony-forming efficiencies using transposition of inducible mouse factors. (a) Morphologically distinct growth foci appeared from rtta-mefs 6-8 days following PB-TET-mFx cocktail
Supplementary Figure S1 Colony-forming efficiencies using transposition of inducible mouse factors. (a) Morphologically distinct growth foci appeared from rtta-mefs 6-8 days following PB-TET-mFx cocktail
Nature Biotechnology: doi: /nbt.4166
 Supplementary Figure 1 Validation of correct targeting at targeted locus. (a) by immunofluorescence staining of 2C-HR-CRISPR microinjected embryos cultured to the blastocyst stage. Embryos were stained
Supplementary Figure 1 Validation of correct targeting at targeted locus. (a) by immunofluorescence staining of 2C-HR-CRISPR microinjected embryos cultured to the blastocyst stage. Embryos were stained
Figure S2. Response of mouse ES cells to GSK3 inhibition. Mentioned in discussion
 Stem Cell Reports, Volume 1 Supplemental Information Robust Self-Renewal of Rat Embryonic Stem Cells Requires Fine-Tuning of Glycogen Synthase Kinase-3 Inhibition Yaoyao Chen, Kathryn Blair, and Austin
Stem Cell Reports, Volume 1 Supplemental Information Robust Self-Renewal of Rat Embryonic Stem Cells Requires Fine-Tuning of Glycogen Synthase Kinase-3 Inhibition Yaoyao Chen, Kathryn Blair, and Austin
Supplementary data. sienigma. F-Enigma F-EnigmaSM. a-p53
 Supplementary data Supplemental Figure 1 A sienigma #2 sienigma sicontrol a-enigma - + ++ - - - - - - + ++ - - - - - - ++ B sienigma F-Enigma F-EnigmaSM a-flag HLK3 cells - - - + ++ + ++ - + - + + - -
Supplementary data Supplemental Figure 1 A sienigma #2 sienigma sicontrol a-enigma - + ++ - - - - - - + ++ - - - - - - ++ B sienigma F-Enigma F-EnigmaSM a-flag HLK3 cells - - - + ++ + ++ - + - + + - -
Schematic representation of the endogenous PALB2 locus and gene-disruption constructs
 Supplementary Figures Supplementary Figure 1. Generation of PALB2 -/- and BRCA2 -/- /PALB2 -/- DT40 cells. (A) Schematic representation of the endogenous PALB2 locus and gene-disruption constructs carrying
Supplementary Figures Supplementary Figure 1. Generation of PALB2 -/- and BRCA2 -/- /PALB2 -/- DT40 cells. (A) Schematic representation of the endogenous PALB2 locus and gene-disruption constructs carrying
Nanog-Luc. R-Luc. 40 HA-Klf Relative protein level of Klf USP21. Vector USP2 USP21 *** *** *** ***
 :Flag Relative protein level of Relative protein level of : Flag Vec 21LV 21SV 2 : HA HA- HA- HA- HA-Klf4 Relative mrna expression Relative protein level of Relative protein level of Relative protein level
:Flag Relative protein level of Relative protein level of : Flag Vec 21LV 21SV 2 : HA HA- HA- HA- HA-Klf4 Relative mrna expression Relative protein level of Relative protein level of Relative protein level
HUMAN EMBRYONIC STEM CELL (hesc) ASSESSMENT BY REAL-TIME POLYMERASE CHAIN REACTION (PCR)
 HUMAN EMBRYONIC STEM CELL (hesc) ASSESSMENT BY REAL-TIME POLYMERASE CHAIN REACTION (PCR) OBJECTIVE: is performed to analyze gene expression of human Embryonic Stem Cells (hescs) in culture. Real-Time PCR
HUMAN EMBRYONIC STEM CELL (hesc) ASSESSMENT BY REAL-TIME POLYMERASE CHAIN REACTION (PCR) OBJECTIVE: is performed to analyze gene expression of human Embryonic Stem Cells (hescs) in culture. Real-Time PCR
Supplementary Figure 1. Soft fibrin gels promote growth and organized mesodermal differentiation. Representative images of single OGTR1 ESCs cultured
 Supplementary Figure 1. Soft fibrin gels promote growth and organized mesodermal differentiation. Representative images of single OGTR1 ESCs cultured in 90-Pa 3D fibrin gels for 5 days in the presence
Supplementary Figure 1. Soft fibrin gels promote growth and organized mesodermal differentiation. Representative images of single OGTR1 ESCs cultured in 90-Pa 3D fibrin gels for 5 days in the presence
SUPPLEMENTARY INFORMATION
 doi: 10.1038/nature06403 Supplementary Information: Nanog safeguards pluripotency and mediates germline development Supplementary Tables SUPPLEMENTARY INFORMATION mrna Forward primer Reverse primer Annealing
doi: 10.1038/nature06403 Supplementary Information: Nanog safeguards pluripotency and mediates germline development Supplementary Tables SUPPLEMENTARY INFORMATION mrna Forward primer Reverse primer Annealing
Human induced Pluripotent Stem Cell (ipsc) Line: GM23225*B
 Certificate of Analysis NIGMS Human tic Cell Repository Human induced Pluripotent Stem Cell (ipsc) Line: GM23225*B Diagnosis Mutation Reprogramming method Publication(s) describing ipsc line establishment
Certificate of Analysis NIGMS Human tic Cell Repository Human induced Pluripotent Stem Cell (ipsc) Line: GM23225*B Diagnosis Mutation Reprogramming method Publication(s) describing ipsc line establishment
NIA Aging Cell Repository
 Certificate of Analysis NIA Aging Cell Repository Human induced Pluripotent Stem Cell (ipsc) Line: AG25370*B Diagnosis Alzheimer Disease, Familial, Type 4 Mutation Reprogramming method Publication(s) describing
Certificate of Analysis NIA Aging Cell Repository Human induced Pluripotent Stem Cell (ipsc) Line: AG25370*B Diagnosis Alzheimer Disease, Familial, Type 4 Mutation Reprogramming method Publication(s) describing
Chemically defined conditions for human ipsc derivation and culture
 Nature Methods Chemically defined conditions for human ipsc derivation and culture Guokai Chen, Daniel R Gulbranson, Zhonggang Hou, Jennifer M Bolin, Victor Ruotti, Mitchell D Probasco, Kimberly Smuga-Otto,
Nature Methods Chemically defined conditions for human ipsc derivation and culture Guokai Chen, Daniel R Gulbranson, Zhonggang Hou, Jennifer M Bolin, Victor Ruotti, Mitchell D Probasco, Kimberly Smuga-Otto,
Disease Clinical features Genotype
 Disease Clinical features Genotype Alpha 1 Antitrypsin deficiency (A1ATD) Patient 1 65 year old caucasian male, liver transplant recipient, explant biopsy confirmed A1ATD induced liver cirrhosis Patient
Disease Clinical features Genotype Alpha 1 Antitrypsin deficiency (A1ATD) Patient 1 65 year old caucasian male, liver transplant recipient, explant biopsy confirmed A1ATD induced liver cirrhosis Patient
Nature Biotechnology: doi: /nbt Supplementary Figure 1
 Supplementary Figure 1 Schematic and results of screening the combinatorial antibody library for Sox2 replacement activity. A single batch of MEFs were plated and transduced with doxycycline inducible
Supplementary Figure 1 Schematic and results of screening the combinatorial antibody library for Sox2 replacement activity. A single batch of MEFs were plated and transduced with doxycycline inducible
Supplementary Table 1. Primers used to construct full-length or various truncated mutants of ISG12b2.
 Supplementary Table 1. Primers used to construct full-length or various truncated mutants of ISG12b2. Construct name ISG12b2 (No tag) HA-ISG12b2 (N-HA) ISG12b2-HA (C-HA; FL-HA) 94-283-HA (FL-GFP) 93-GFP
Supplementary Table 1. Primers used to construct full-length or various truncated mutants of ISG12b2. Construct name ISG12b2 (No tag) HA-ISG12b2 (N-HA) ISG12b2-HA (C-HA; FL-HA) 94-283-HA (FL-GFP) 93-GFP
Molecular Cell Biology - Problem Drill 11: Recombinant DNA
 Molecular Cell Biology - Problem Drill 11: Recombinant DNA Question No. 1 of 10 1. Which of the following statements about the sources of DNA used for molecular cloning is correct? Question #1 (A) cdna
Molecular Cell Biology - Problem Drill 11: Recombinant DNA Question No. 1 of 10 1. Which of the following statements about the sources of DNA used for molecular cloning is correct? Question #1 (A) cdna
Rapid differentiation of human pluripotent stem cells into functional motor neurons by
 Supplementary Information Rapid differentiation of human pluripotent stem cells into functional motor neurons by mrnas encoding transcription factors Sravan Kumar Goparaju, Kazuhisa Kohda, Keiji Ibata,
Supplementary Information Rapid differentiation of human pluripotent stem cells into functional motor neurons by mrnas encoding transcription factors Sravan Kumar Goparaju, Kazuhisa Kohda, Keiji Ibata,
Online Supplementary Information
 Online Supplementary Information NLRP4 negatively regulates type I interferon signaling by targeting TBK1 for degradation via E3 ubiquitin ligase DTX4 Jun Cui 1,4,6,7, Yinyin Li 1,5,6,7, Liang Zhu 1, Dan
Online Supplementary Information NLRP4 negatively regulates type I interferon signaling by targeting TBK1 for degradation via E3 ubiquitin ligase DTX4 Jun Cui 1,4,6,7, Yinyin Li 1,5,6,7, Liang Zhu 1, Dan
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature11070 Supplementary Figure 1 Purification of FLAG-tagged proteins. a, Purification of FLAG-RNF12 by FLAG-affinity from nuclear extracts of wild-type (WT) and two FLAG- RNF12 transgenic
doi:10.1038/nature11070 Supplementary Figure 1 Purification of FLAG-tagged proteins. a, Purification of FLAG-RNF12 by FLAG-affinity from nuclear extracts of wild-type (WT) and two FLAG- RNF12 transgenic
TITLE: Induced Pluripotent Stem Cells as Potential Therapeutic Agents in NF1
 AD Award Number: W81XWH-10-1-0181 TITLE: Induced Pluripotent Stem Cells as Potential Therapeutic Agents in NF1 PRINCIPAL INVESTIGATOR: Jonathan Chernoff, M.D., Ph.D. CONTRACTING ORGANIZATION: Institute
AD Award Number: W81XWH-10-1-0181 TITLE: Induced Pluripotent Stem Cells as Potential Therapeutic Agents in NF1 PRINCIPAL INVESTIGATOR: Jonathan Chernoff, M.D., Ph.D. CONTRACTING ORGANIZATION: Institute
Supplementary information for: Ten-Eleven Translocation-2 (Tet2) Is Involved in Myogenic Differentiation of Skeletal Myoblast Cells in
 Supplementary information for: Ten-Eleven Translocation-2 (Tet2) Is Involved in Myogenic Differentiation of Skeletal Myoblast Cells in Vitro Xia Zhong*, Qian-Qian Wang*, Jian-Wei Li, Yu-Mei Zhang, Xiao-Rong
Supplementary information for: Ten-Eleven Translocation-2 (Tet2) Is Involved in Myogenic Differentiation of Skeletal Myoblast Cells in Vitro Xia Zhong*, Qian-Qian Wang*, Jian-Wei Li, Yu-Mei Zhang, Xiao-Rong
Generation of ips-derived model cells for analyses of hair shaft differentiation
 Supplementary Material Generation of ips-derived model cells for analyses of hair shaft differentiation Takumi Kido, Tomoatsu Horigome, Minori Uda, Naoki Adachi, Yohei Hirai Department of Biomedical Chemistry,
Supplementary Material Generation of ips-derived model cells for analyses of hair shaft differentiation Takumi Kido, Tomoatsu Horigome, Minori Uda, Naoki Adachi, Yohei Hirai Department of Biomedical Chemistry,
Authors: Takuro Horii, Masamichi Yamamoto, Sumiyo Morita, Mika Kimura, Yasumitsu Nagao, Izuho Hatada *
 SUPPLEMENTARY INFORMATION Title: p53 Suppresses Tetraploid Development in Mice Authors: Takuro Horii, Masamichi Yamamoto, Sumiyo Morita, Mika Kimura, Yasumitsu Nagao, Izuho Hatada * Supplementary Methods
SUPPLEMENTARY INFORMATION Title: p53 Suppresses Tetraploid Development in Mice Authors: Takuro Horii, Masamichi Yamamoto, Sumiyo Morita, Mika Kimura, Yasumitsu Nagao, Izuho Hatada * Supplementary Methods
Supplementary Data. Supplementary Table S1.
 Supplementary Data Supplementary Table S1. Primer Sequences for Real-Time Reverse Transcription-Polymerase Chain Reaction Gene Forward 5?3 Reverse 3?5 Housekeeping gene 36B4 TCCAGGCTTTGGGCATCA CTTTATCAGCTGCACATCACTCAGA
Supplementary Data Supplementary Table S1. Primer Sequences for Real-Time Reverse Transcription-Polymerase Chain Reaction Gene Forward 5?3 Reverse 3?5 Housekeeping gene 36B4 TCCAGGCTTTGGGCATCA CTTTATCAGCTGCACATCACTCAGA
Electromagnetic Fields Mediate Efficient Cell Reprogramming Into a Pluripotent State
 Electromagnetic Fields Mediate Efficient Cell Reprogramming Into a Pluripotent State Soonbong Baek, 1 Xiaoyuan Quan, 1 Soochan Kim, 2 Christopher Lengner, 3 Jung- Keug Park, 4 and Jongpil Kim 1* 1 Lab
Electromagnetic Fields Mediate Efficient Cell Reprogramming Into a Pluripotent State Soonbong Baek, 1 Xiaoyuan Quan, 1 Soochan Kim, 2 Christopher Lengner, 3 Jung- Keug Park, 4 and Jongpil Kim 1* 1 Lab
Table S1. Primer sequences
 Table S1. Primer sequences Primers for quantitative PCR Tgf 1 Forward Tgf 1 Reverse Tgf 2Forward Tgf 2Reverse Tgf 3 Forward Tgf 3 Reverse Tgf r1 Forward Tgf r1 Reverse Tgf r2 Forward Tgf r2 Reverse Thbs1
Table S1. Primer sequences Primers for quantitative PCR Tgf 1 Forward Tgf 1 Reverse Tgf 2Forward Tgf 2Reverse Tgf 3 Forward Tgf 3 Reverse Tgf r1 Forward Tgf r1 Reverse Tgf r2 Forward Tgf r2 Reverse Thbs1
Supplemental material
 Supplemental material THE JOURNAL OF CELL BIOLOGY Taylor et al., http://www.jcb.org/cgi/content/full/jcb.201403021/dc1 Figure S1. Representative images of Cav 1a -YFP mutants with and without LMB treatment.
Supplemental material THE JOURNAL OF CELL BIOLOGY Taylor et al., http://www.jcb.org/cgi/content/full/jcb.201403021/dc1 Figure S1. Representative images of Cav 1a -YFP mutants with and without LMB treatment.
Supplementary Figure 1
 Supplementary Figure A Name Location Start position End position Oct4P Chr 3 29433673 29435 Annotation Mus musculus predicted gene 572 (Gm572), noncoding RNA RefSeq accession number NR_33594. Oct4P2 Chr
Supplementary Figure A Name Location Start position End position Oct4P Chr 3 29433673 29435 Annotation Mus musculus predicted gene 572 (Gm572), noncoding RNA RefSeq accession number NR_33594. Oct4P2 Chr
Supplementary Figure 1, related to Figure 1. GAS5 is highly expressed in the cytoplasm of hescs, and positively correlates with pluripotency.
 Supplementary Figure 1, related to Figure 1. GAS5 is highly expressed in the cytoplasm of hescs, and positively correlates with pluripotency. (a) Transfection of different concentration of GAS5-overexpressing
Supplementary Figure 1, related to Figure 1. GAS5 is highly expressed in the cytoplasm of hescs, and positively correlates with pluripotency. (a) Transfection of different concentration of GAS5-overexpressing
Nature Biotechnology: doi: /nbt Supplementary Figure 1. Generation of NSCs from hpscs in SDC medium.
 Supplementary Figure 1 Generation of NSCs from hpscs in SDC medium. (a) Q-PCR of pluripotent markers (OCT4, NANOG), neural markers (SOX1, PAX6, N-Cadherin), markers for the other germ layers (T, EOMES,
Supplementary Figure 1 Generation of NSCs from hpscs in SDC medium. (a) Q-PCR of pluripotent markers (OCT4, NANOG), neural markers (SOX1, PAX6, N-Cadherin), markers for the other germ layers (T, EOMES,
SUPPLEMENTARY INFORMATION
 Supplementary Figure a 3 factors 4 factors Nanog-GFP phase b c.5 Endogenous Oct3/4 2 Total Oct3/4.5 Relative expression (ES = ).5.5 6 3 Endogenous Sox2 Nanog Relative expression (ES = ) 5 25 2 Total Sox2
Supplementary Figure a 3 factors 4 factors Nanog-GFP phase b c.5 Endogenous Oct3/4 2 Total Oct3/4.5 Relative expression (ES = ).5.5 6 3 Endogenous Sox2 Nanog Relative expression (ES = ) 5 25 2 Total Sox2
Supplementary Figure 1
 Supplementary Figure 1 Supplementary Figure 1. Functional assays of hepatocyte-like cells induced from H1 hescs (a) P450 and AAT staining, PAS staining, LDL uptake assay, and ICG uptake and release assay
Supplementary Figure 1 Supplementary Figure 1. Functional assays of hepatocyte-like cells induced from H1 hescs (a) P450 and AAT staining, PAS staining, LDL uptake assay, and ICG uptake and release assay
Gene Forward Primer Reverse Primer GAPDH ATCATCCCTGCCTCTACTGG GTCAGGTCCACCACTGACAC SSB1 AACTTCAGTGAGCCAAACCC GTTCTCAGAGGCTGGAGAGG
 Supplemental Data EXPERIMENTAL PROCEDURES Plasmids and Antibodies- Full length cdna of INT11 or INT12 were cloned into ps- Flag-SBP vector respectively. Anti-RNA pol II (RPB1) was purchased from Santa
Supplemental Data EXPERIMENTAL PROCEDURES Plasmids and Antibodies- Full length cdna of INT11 or INT12 were cloned into ps- Flag-SBP vector respectively. Anti-RNA pol II (RPB1) was purchased from Santa
Fig. S1. Effect of p120-catenin overexpression on the interaction of SCUBE2 with E-cadherin. The expression plasmid encoding FLAG.
 Fig. S1. Effect of p120-catenin overexpression on the interaction of SCUBE2 with E-cadherin. The expression plasmid encoding FLAG.SCUBE2, E-cadherin.Myc, or HA.p120-catenin was transfected in a combination
Fig. S1. Effect of p120-catenin overexpression on the interaction of SCUBE2 with E-cadherin. The expression plasmid encoding FLAG.SCUBE2, E-cadherin.Myc, or HA.p120-catenin was transfected in a combination
Supplementary Figure 1.
 Supplementary Figure 1. (A) UCSC Genome Browser view of region immediately downstream of the NEUROG3 start codon. All candidate grna target sites which meet the G(N 19 )NGG constraint are aligned to illustrate
Supplementary Figure 1. (A) UCSC Genome Browser view of region immediately downstream of the NEUROG3 start codon. All candidate grna target sites which meet the G(N 19 )NGG constraint are aligned to illustrate
Isolation of human ips cells using EOS lentiviral vectors to select for pluripotency
 nature methods Isolation of human ips cells using EOS lentiviral vectors to select for pluripotency Akitsu Hotta, Aaron Y L Cheung, Natalie Farra, Kausalia Vijayaragavan, Cheryle A Séguin, Jonathan S Draper,
nature methods Isolation of human ips cells using EOS lentiviral vectors to select for pluripotency Akitsu Hotta, Aaron Y L Cheung, Natalie Farra, Kausalia Vijayaragavan, Cheryle A Séguin, Jonathan S Draper,
all samples of a band with a molecular weight close to that expected for the endogenous!-
 SUPPLEMENTAL FIGURE LEGENDS Supplemental Figure 1 : Specificity of anti-!-arrestin Abs and levels of!-arrestin in WHIM wt leukocytes. (A and B) HEK 293T cells were transiently transfected using the reagent
SUPPLEMENTAL FIGURE LEGENDS Supplemental Figure 1 : Specificity of anti-!-arrestin Abs and levels of!-arrestin in WHIM wt leukocytes. (A and B) HEK 293T cells were transiently transfected using the reagent
Cat# Product Name amounts h NANOG (RFP-Bsd) inducible particles
 Premade Lentiviral Particles for ips Stem Factors For generating induced pluripotent stem (ips) cells or other applications. LVP003 LVP004 LVP005 LVP006 LVP007 LVP008 RESEARCH USE ONLY, not intend for
Premade Lentiviral Particles for ips Stem Factors For generating induced pluripotent stem (ips) cells or other applications. LVP003 LVP004 LVP005 LVP006 LVP007 LVP008 RESEARCH USE ONLY, not intend for
Supporting Online Material for
 www.sciencemag.org/cgi/content/full/1154040/dc1 Supporting Online Material for Selective Blockade of MicroRNA Processing by Lin-28 Srinivas R. Viswanathan, George Q. Daley,* Richard I. Gregory* *To whom
www.sciencemag.org/cgi/content/full/1154040/dc1 Supporting Online Material for Selective Blockade of MicroRNA Processing by Lin-28 Srinivas R. Viswanathan, George Q. Daley,* Richard I. Gregory* *To whom
Protocol Using the Reprogramming Ecotropic Retrovirus Set: Mouse OSKM to Reprogram MEFs into ips Cells
 STEMGENT Page 1 OVERVIEW The following protocol describes the reprogramming of one well of mouse embryonic fibroblast (MEF) cells into induced pluripotent stem (ips) cells in a 6-well format. Transduction
STEMGENT Page 1 OVERVIEW The following protocol describes the reprogramming of one well of mouse embryonic fibroblast (MEF) cells into induced pluripotent stem (ips) cells in a 6-well format. Transduction
Regulation of transcription by the MLL2 complex and MLL complex-associated AKAP95
 Supplementary Information Regulation of transcription by the complex and MLL complex-associated Hao Jiang, Xiangdong Lu, Miho Shimada, Yali Dou, Zhanyun Tang, and Robert G. Roeder Input HeLa NE IP lot:
Supplementary Information Regulation of transcription by the complex and MLL complex-associated Hao Jiang, Xiangdong Lu, Miho Shimada, Yali Dou, Zhanyun Tang, and Robert G. Roeder Input HeLa NE IP lot:
Supplemental Materials and Methods
 Supplemental Materials and Methods In situ hybridization In situ hybridization analysis of HFE2 and genin mrna in rat liver tissues was performed as previously described (1). Briefly, the digoxigenin-labeled
Supplemental Materials and Methods In situ hybridization In situ hybridization analysis of HFE2 and genin mrna in rat liver tissues was performed as previously described (1). Briefly, the digoxigenin-labeled
A cost-effective system for differentiation of intestinal epithelium from human induced pluripotent stem cells
 A cost-effective system for differentiation of intestinal epithelium from human induced pluripotent stem cells Soichiro Ogaki 1, 2, 3, 4, 5, *, ayu orooka 1,*, Kaito Otera 1, 1, 3 and Shoen Kume Supplementary
A cost-effective system for differentiation of intestinal epithelium from human induced pluripotent stem cells Soichiro Ogaki 1, 2, 3, 4, 5, *, ayu orooka 1,*, Kaito Otera 1, 1, 3 and Shoen Kume Supplementary
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb3363 Supplementary Figure 1 Several WNTs bind to the extracellular domains of PKD1. (a) HEK293T cells were co-transfected with indicated plasmids. Flag-tagged proteins were immunoprecipiated
DOI: 10.1038/ncb3363 Supplementary Figure 1 Several WNTs bind to the extracellular domains of PKD1. (a) HEK293T cells were co-transfected with indicated plasmids. Flag-tagged proteins were immunoprecipiated
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb3164 Supplementary Figure 1 Validation of effective Gnas deletion and epithelial thickness. a, Representative genotyping in mice treated or not with tamoxifen to show Gnas deletion. To
DOI: 10.1038/ncb3164 Supplementary Figure 1 Validation of effective Gnas deletion and epithelial thickness. a, Representative genotyping in mice treated or not with tamoxifen to show Gnas deletion. To
Fatchiyah
 Fatchiyah Email: fatchiya@yahoo.co.id RNAs: mrna trna rrna RNAi DNAs: Protein: genome DNA cdna mikro-makro mono-poly single-multi Analysis: Identification human and animal disease Finger printing Sexing
Fatchiyah Email: fatchiya@yahoo.co.id RNAs: mrna trna rrna RNAi DNAs: Protein: genome DNA cdna mikro-makro mono-poly single-multi Analysis: Identification human and animal disease Finger printing Sexing
Supplementary Figure 1. Generation of B2M -/- ESCs. Nature Biotechnology: doi: /nbt.3860
 Supplementary Figure 1 Generation of B2M -/- ESCs. (a) Maps of the B2M alleles in cells with the indicated B2M genotypes. Probes and restriction enzymes used in Southern blots are indicated (H, Hind III;
Supplementary Figure 1 Generation of B2M -/- ESCs. (a) Maps of the B2M alleles in cells with the indicated B2M genotypes. Probes and restriction enzymes used in Southern blots are indicated (H, Hind III;
5.4 PERIODONTAL LIGAMENT STEM CELL (PDLSC) ISOLATION. Having validated that the isolated PDL stromal cells possessed the major phenotypic
 5.4 PERIODONTAL LIGAMENT STEM CELL (PDLSC) ISOLATION Having validated that the isolated PDL stromal cells possessed the major phenotypic characteristics of their tissue of origin, we next focused on identifying
5.4 PERIODONTAL LIGAMENT STEM CELL (PDLSC) ISOLATION Having validated that the isolated PDL stromal cells possessed the major phenotypic characteristics of their tissue of origin, we next focused on identifying
Supplemental Figure 1 Human REEP family of proteins can be divided into two distinct subfamilies. Residues (single letter amino acid code) identical
 Supplemental Figure Human REEP family of proteins can be divided into two distinct subfamilies. Residues (single letter amino acid code) identical in all six REEPs are highlighted in green. Additional
Supplemental Figure Human REEP family of proteins can be divided into two distinct subfamilies. Residues (single letter amino acid code) identical in all six REEPs are highlighted in green. Additional
PrimePCR Assay Validation Report
 Gene Information Gene Name keratin 3 Gene Symbol Organism Gene Summary Gene Aliases RefSeq Accession No. UniGene ID Ensembl Gene ID KRT3 Human The protein encoded by this gene is a member of the keratin
Gene Information Gene Name keratin 3 Gene Symbol Organism Gene Summary Gene Aliases RefSeq Accession No. UniGene ID Ensembl Gene ID KRT3 Human The protein encoded by this gene is a member of the keratin
PrimePCR Assay Validation Report
 Gene Information Gene Name Gene Symbol Organism Gene Summary Gene Aliases RefSeq Accession No. UniGene ID Ensembl Gene ID Glyceraldehyde-3-phosphate dehydrogenase Gapdh Rat This gene encodes a member of
Gene Information Gene Name Gene Symbol Organism Gene Summary Gene Aliases RefSeq Accession No. UniGene ID Ensembl Gene ID Glyceraldehyde-3-phosphate dehydrogenase Gapdh Rat This gene encodes a member of
Supplementary Figure 1. Nature Structural & Molecular Biology: doi: /nsmb.3494
 Supplementary Figure 1 Pol structure-function analysis (a) Inactivating polymerase and helicase mutations do not alter the stability of Pol. Flag epitopes were introduced using CRISPR/Cas9 gene targeting
Supplementary Figure 1 Pol structure-function analysis (a) Inactivating polymerase and helicase mutations do not alter the stability of Pol. Flag epitopes were introduced using CRISPR/Cas9 gene targeting
GFP CCD2 GFP IP:GFP
 D1 D2 1 75 95 148 178 492 GFP CCD1 CCD2 CCD2 GFP D1 D2 GFP D1 D2 Beclin 1 IB:GFP IP:GFP Supplementary Figure 1: Mapping domains required for binding to HEK293T cells are transfected with EGFP-tagged mutant
D1 D2 1 75 95 148 178 492 GFP CCD1 CCD2 CCD2 GFP D1 D2 GFP D1 D2 Beclin 1 IB:GFP IP:GFP Supplementary Figure 1: Mapping domains required for binding to HEK293T cells are transfected with EGFP-tagged mutant
Supplementary Figure 1. Expressions of stem cell markers decreased in TRCs on 2D plastic. TRCs were cultured on plastic for 1, 3, 5, or 7 days,
 Supplementary Figure 1. Expressions of stem cell markers decreased in TRCs on 2D plastic. TRCs were cultured on plastic for 1, 3, 5, or 7 days, respectively, and their mrnas were quantified by real time
Supplementary Figure 1. Expressions of stem cell markers decreased in TRCs on 2D plastic. TRCs were cultured on plastic for 1, 3, 5, or 7 days, respectively, and their mrnas were quantified by real time
FastGene Optima. Products for PCR. proof-reading. ReadyMix. cdna SNP. routine PCR. polymerase. optimized blend complex templates. N gene knock out B O
 Products for PCR www.nippongenetics.eu efficiency polymerase convenience endpoint PCR incl. loading dye direct PCR incl. dntps engineered enzyme archeal type B polymerase high GC content FastGene tissuebest
Products for PCR www.nippongenetics.eu efficiency polymerase convenience endpoint PCR incl. loading dye direct PCR incl. dntps engineered enzyme archeal type B polymerase high GC content FastGene tissuebest
Protocol for Genome Editing via the RNA-guided Cas9 Nuclease in. Zebrafish Embryos 1
 Protocol for Genome Editing via the RNA-guided Cas9 Nuclease in Zebrafish Embryos 1 1. In vitro synthesis of capped Cas9 mrna The full length of humanized Cas9 cdnas with double NLS were cloned into pxt7
Protocol for Genome Editing via the RNA-guided Cas9 Nuclease in Zebrafish Embryos 1 1. In vitro synthesis of capped Cas9 mrna The full length of humanized Cas9 cdnas with double NLS were cloned into pxt7
Supplementary Figure 1 a. d 0.8 CON LPS PAN. 2nd ab nephrin podocin CON LPS PAN. upar. -tubulin. upar. upar / -tubulin CON LPS PAN
 Supplementary Figure 1 a Efferent arteriole Podocytes Afferent arteriole FP Endothelium GBM Glomerular filtration barrier b 188kD HEK + GFP HEK + GFP-Nphs1 Differentiated Podocytes HEK + GFP HEK + GFP-Nphs2
Supplementary Figure 1 a Efferent arteriole Podocytes Afferent arteriole FP Endothelium GBM Glomerular filtration barrier b 188kD HEK + GFP HEK + GFP-Nphs1 Differentiated Podocytes HEK + GFP HEK + GFP-Nphs2
SUPPLEMENTAL MATERIALS SIRTUIN 1 PROMOTES HYPEROXIA-INDUCED LUNG EPITHELIAL DEATH INDEPENDENT OF NRF2 ACTIVATION
 SUPPLEMENTAL MATERIALS SIRTUIN PROMOTES HYPEROXIA-INDUCED LUNG EPITHELIAL DEATH INDEPENDENT OF NRF ACTIVATION Haranatha R. Potteti*, Subbiah Rajasekaran*, Senthilkumar B. Rajamohan*, Chandramohan R. Tamatam,
SUPPLEMENTAL MATERIALS SIRTUIN PROMOTES HYPEROXIA-INDUCED LUNG EPITHELIAL DEATH INDEPENDENT OF NRF ACTIVATION Haranatha R. Potteti*, Subbiah Rajasekaran*, Senthilkumar B. Rajamohan*, Chandramohan R. Tamatam,
8Br-cAMP was purchased from Sigma (St. Louis, MO). Silencer Negative Control sirna #1 and
 1 Supplemental information 2 3 Materials and Methods 4 Reagents and animals 5 8Br-cAMP was purchased from Sigma (St. Louis, MO). Silencer Negative Control sirna #1 and 6 Silencer Select Pre-designed sirna
1 Supplemental information 2 3 Materials and Methods 4 Reagents and animals 5 8Br-cAMP was purchased from Sigma (St. Louis, MO). Silencer Negative Control sirna #1 and 6 Silencer Select Pre-designed sirna
Supplementary Figure 1 Characterization of OKSM-iNSCs
 Supplementary Figure 1 Characterization of OKSM-iNSCs (A) Gene expression levels of Sox1 and Sox2 in the indicated cell types by microarray gene expression analysis. (B) Dendrogram cluster analysis of
Supplementary Figure 1 Characterization of OKSM-iNSCs (A) Gene expression levels of Sox1 and Sox2 in the indicated cell types by microarray gene expression analysis. (B) Dendrogram cluster analysis of
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature12119 SUPPLEMENTARY FIGURES AND LEGENDS pre-let-7a- 1 +14U pre-let-7a- 1 Ddx3x Dhx30 Dis3l2 Elavl1 Ggt5 Hnrnph 2 Osbpl5 Puf60 Rnpc3 Rpl7 Sf3b3 Sf3b4 Tia1 Triobp U2af1 U2af2 1 6 2 4 3
doi:10.1038/nature12119 SUPPLEMENTARY FIGURES AND LEGENDS pre-let-7a- 1 +14U pre-let-7a- 1 Ddx3x Dhx30 Dis3l2 Elavl1 Ggt5 Hnrnph 2 Osbpl5 Puf60 Rnpc3 Rpl7 Sf3b3 Sf3b4 Tia1 Triobp U2af1 U2af2 1 6 2 4 3
Biology Open (2015): doi: /bio : Supplementary information
 Fig. S1. Verification of optimal conditions for the generation of LIF-dependent inscs Representative images (from five repeat experiments) of human primary fibroblasts undergoing reprogramming for 21 days
Fig. S1. Verification of optimal conditions for the generation of LIF-dependent inscs Representative images (from five repeat experiments) of human primary fibroblasts undergoing reprogramming for 21 days
e15 e14 3utr i13 K36me3 / H i4 ** ** ** ** ** tss IIIb K36me2 / H IIIb BCOR (% input)
 a Epithelial cells (2) Mesenchymal cells () Alternatively spliced region i13 e14 e15 / CycA 25 15 1 5 e14 e13-e14 - - K27me3 / H3 b 2.5 c v5 K36me3 / H3 d 3.5 2.8 2.1 1.4.7 e 3.5 2.8 2.1 1.4.7 v5 K9me3
a Epithelial cells (2) Mesenchymal cells () Alternatively spliced region i13 e14 e15 / CycA 25 15 1 5 e14 e13-e14 - - K27me3 / H3 b 2.5 c v5 K36me3 / H3 d 3.5 2.8 2.1 1.4.7 e 3.5 2.8 2.1 1.4.7 v5 K9me3
A Novel Platform to Enable the High-Throughput Derivation and Characterization of Feeder-Free Human ipscs
 Supplementary Information A Novel Platform to Enable the High-Throughput Derivation and Characterization of Feeder-Free Human ipscs Bahram Valamehr, Ramzey Abujarour, Megan Robinson, Thuy Le, David Robbins,
Supplementary Information A Novel Platform to Enable the High-Throughput Derivation and Characterization of Feeder-Free Human ipscs Bahram Valamehr, Ramzey Abujarour, Megan Robinson, Thuy Le, David Robbins,
Table S1. List of DNA constructs and primers, part 1 Construct
 SUPPLEMENTARY TABLES: Table S1. List of DNA constructs and primers, part 1 Construct Comment (Expressed protein) Vector, PCR primers and template or source of Restriction Sites name Resistance construct
SUPPLEMENTARY TABLES: Table S1. List of DNA constructs and primers, part 1 Construct Comment (Expressed protein) Vector, PCR primers and template or source of Restriction Sites name Resistance construct
Gene name forward primer 5->3 reverse primer 5->3 product GCCCATA. Nfatc1 AGGTGCAGCCCAAGTCTCAC GTGGCCATCTGGAGCCTTC CTGT GATGC GCT ACCAA
 Supplementary able 1 is presenting the PCR primer list used. ene name forward primer 5->3 reverse primer 5->3 product size cdna VE-Cadherin AACCCAAAAA CACCCCAAAAC 260bp CCA CCCAA PECAM-1 CACCACAA CCCCCACC
Supplementary able 1 is presenting the PCR primer list used. ene name forward primer 5->3 reverse primer 5->3 product size cdna VE-Cadherin AACCCAAAAA CACCCCAAAAC 260bp CCA CCCAA PECAM-1 CACCACAA CCCCCACC
Supplementary Figure 1. Adipogenic protein expression in WT and KO MEFs after 7 days of adipogenic differentiation.
 Merkestein et al. Supplementary Figure 1 A PLIN WT FTO KO 72 kda FABP4 17 kda HSC70 72 kda B Supplementary Figure 1. Adipogenic protein expression in WT and KO MEFs after 7 days of adipogenic differentiation.
Merkestein et al. Supplementary Figure 1 A PLIN WT FTO KO 72 kda FABP4 17 kda HSC70 72 kda B Supplementary Figure 1. Adipogenic protein expression in WT and KO MEFs after 7 days of adipogenic differentiation.
Bootcamp: Molecular Biology Techniques and Interpretation
 Bootcamp: Molecular Biology Techniques and Interpretation Bi8 Winter 2016 Today s outline Detecting and quantifying nucleic acids and proteins: Basic nucleic acid properties Hybridization PCR and Designing
Bootcamp: Molecular Biology Techniques and Interpretation Bi8 Winter 2016 Today s outline Detecting and quantifying nucleic acids and proteins: Basic nucleic acid properties Hybridization PCR and Designing
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION DOI: 1.138/NMAT3777 Biophysical regulation of epigenetic state and cell reprogramming Authors: Timothy L. Downing 1,2, Jennifer Soto 1,2, Constant Morez 2,3,, Timothee Houssin
SUPPLEMENTARY INFORMATION DOI: 1.138/NMAT3777 Biophysical regulation of epigenetic state and cell reprogramming Authors: Timothy L. Downing 1,2, Jennifer Soto 1,2, Constant Morez 2,3,, Timothee Houssin
Figure S1. Phenotypic characterization of AND-1_WASKO cell lines. AND- 1_WASKO_C1.1 (WASKO_C1.1) and AND-1_WASKO_C1.2 (WASKO_C1.
 LEGENDS TO SUPPLEMENTARY FIGURES Figure S1. Phenotypic characterization of AND-1_WASKO cell lines. AND- 1_WASKO_C1.1 (WASKO_C1.1) and AND-1_WASKO_C1.2 (WASKO_C1.2) were stained with the antibodies oct3/4
LEGENDS TO SUPPLEMENTARY FIGURES Figure S1. Phenotypic characterization of AND-1_WASKO cell lines. AND- 1_WASKO_C1.1 (WASKO_C1.1) and AND-1_WASKO_C1.2 (WASKO_C1.2) were stained with the antibodies oct3/4
Supplementary Figure 1. Quantitative RT-PCR experimental validation of CRISPR/Cas9 and sgrnas expression in HEK293A transfected cells.
 Supplementary Figure 1. Quantitative RT-PCR experimental validation of CRISPR/Cas9 and sgrnas expression in HEK293A transfected cells. HEK293A cells were transfected with the indicated combinations of
Supplementary Figure 1. Quantitative RT-PCR experimental validation of CRISPR/Cas9 and sgrnas expression in HEK293A transfected cells. HEK293A cells were transfected with the indicated combinations of
Protocol Reprogramming MEFs using the Dox Inducible Reprogramming Lentivirus Set: Mouse OKSM
 STEMGENT Page 1 OVERVIEW The following protocol describes the reprogramming of one well of mouse embryonic fibroblasts (MEFs) into induced pluripotent stem (ips) cells in a 6-well format. Transduction
STEMGENT Page 1 OVERVIEW The following protocol describes the reprogramming of one well of mouse embryonic fibroblasts (MEFs) into induced pluripotent stem (ips) cells in a 6-well format. Transduction
Critical Factors For Successful Real-Time RT-PCR
 Critical Factors For Successful Real-Time RT-PCR Andreas Missel, PhD Senior Scientist R&D Dept. Modification/Amplification QIAGEN Annealing Specific product High yield High sensitivity Nonspecific product
Critical Factors For Successful Real-Time RT-PCR Andreas Missel, PhD Senior Scientist R&D Dept. Modification/Amplification QIAGEN Annealing Specific product High yield High sensitivity Nonspecific product
MeCP2. MeCP2/α-tubulin. GFP mir1-1 mir132
 Conservation Figure S1. Schematic showing 3 UTR (top; thick black line), mir132 MRE (arrow) and nucleotide sequence conservation (vertical black lines; http://genome.ucsc.edu). a GFP mir1-1 mir132 b GFP
Conservation Figure S1. Schematic showing 3 UTR (top; thick black line), mir132 MRE (arrow) and nucleotide sequence conservation (vertical black lines; http://genome.ucsc.edu). a GFP mir1-1 mir132 b GFP
Fig. S1. eif6 expression in HEK293 transfected with shrna against eif6 or pcmv-eif6 vector.
 Fig. S1. eif6 expression in HEK293 transfected with shrna against eif6 or pcmv-eif6 vector. (a) Western blotting analysis and (b) qpcr analysis of eif6 expression in HEK293 T cells transfected with either
Fig. S1. eif6 expression in HEK293 transfected with shrna against eif6 or pcmv-eif6 vector. (a) Western blotting analysis and (b) qpcr analysis of eif6 expression in HEK293 T cells transfected with either
Table S1. Primers used in the study
 Table S1. Primers used in the study Primer name Application Sequence I1F16 Genotyping GGCAAGTGAGTGAGTGCCTA I1R11 Genotyping CCCACTCGTATTGACGCTCT V19 Genotyping GGGTCTCAAAGTCAGGGTCA D18Mit184-F Genotyping
Table S1. Primers used in the study Primer name Application Sequence I1F16 Genotyping GGCAAGTGAGTGAGTGCCTA I1R11 Genotyping CCCACTCGTATTGACGCTCT V19 Genotyping GGGTCTCAAAGTCAGGGTCA D18Mit184-F Genotyping
Supplemental Table S1. RT-PCR primers used in this study
 Supplemental Table S1. RT-PCR primers used in this study -----------------------------------------------------------------------------------------------------------------------------------------------
Supplemental Table S1. RT-PCR primers used in this study -----------------------------------------------------------------------------------------------------------------------------------------------
Supplementary Figures
 Supplementary Figures Fig. S1. Specificity of perilipin antibody in SAT compared to various organs. (A) Images of SAT at E18.5 stained with perilipin and CD31 antibody. Scale bars: 100 μm. SAT stained
Supplementary Figures Fig. S1. Specificity of perilipin antibody in SAT compared to various organs. (A) Images of SAT at E18.5 stained with perilipin and CD31 antibody. Scale bars: 100 μm. SAT stained
Supplementary Fig.1. Over-expression of RNase L in stable polyclonal cell line
 Supplemental Data mrna Protein A kda 75 40 NEO/vector NEO/RNase L RNASE L β-actin RNASE L β-actin B % of control (neo vector), normalized 240 ** 200 160 120 80 40 0 Neo Neo/RNase L RNase L Protein Supplementary
Supplemental Data mrna Protein A kda 75 40 NEO/vector NEO/RNase L RNASE L β-actin RNASE L β-actin B % of control (neo vector), normalized 240 ** 200 160 120 80 40 0 Neo Neo/RNase L RNase L Protein Supplementary
ExProfile TM Human Osteogenesis Related Gene qpcr Array
 ExProfile TM Human Osteogenesis Related Gene qpcr Array For focused group profiling of human osteogenesis genes expression Cat. No. QG046-A (1 x 96-well plate, Format A) Cat. No. QG046-B (1 x 96-well plate,
ExProfile TM Human Osteogenesis Related Gene qpcr Array For focused group profiling of human osteogenesis genes expression Cat. No. QG046-A (1 x 96-well plate, Format A) Cat. No. QG046-B (1 x 96-well plate,
Genomic DNA fragments identified by ChIP-chip assay for MR [28] corresponding to two
![Genomic DNA fragments identified by ChIP-chip assay for MR [28] corresponding to two Genomic DNA fragments identified by ChIP-chip assay for MR [28] corresponding to two](/thumbs/89/98549522.jpg) Supplemental Materials and Methods Genomic DNA fragments identified by ChIP-chip assay for MR [28] corresponding to two upstream regions of the human Klf9 gene (-5139 to -5771 bp and -3875 to -4211 bp)
Supplemental Materials and Methods Genomic DNA fragments identified by ChIP-chip assay for MR [28] corresponding to two upstream regions of the human Klf9 gene (-5139 to -5771 bp and -3875 to -4211 bp)
IRE1a-XBP1 is a novel branch in the transcriptional regulation of Ucp1 in brown adipocytes.
 IREaXBP is a novel branch in the transcriptional regulation of Ucp in brown adipocytes. Supplementary Information Rie Asada, Soshi Kanemoto, Koji Matsuhisa, Kenta Hino, Min Cui, Xiang Cui, Masayuki Kaneko,
IREaXBP is a novel branch in the transcriptional regulation of Ucp in brown adipocytes. Supplementary Information Rie Asada, Soshi Kanemoto, Koji Matsuhisa, Kenta Hino, Min Cui, Xiang Cui, Masayuki Kaneko,
Supplemental Fig. 1: PEA-15 knockdown efficiency assessed by immunohistochemistry and qpcr
 Supplemental figure legends Supplemental Fig. 1: PEA-15 knockdown efficiency assessed by immunohistochemistry and qpcr A, LβT2 cells were transfected with either scrambled or PEA-15 sirna. Cells were then
Supplemental figure legends Supplemental Fig. 1: PEA-15 knockdown efficiency assessed by immunohistochemistry and qpcr A, LβT2 cells were transfected with either scrambled or PEA-15 sirna. Cells were then
Table S1. Components for knockout serum replacer medium (KSR medium) (500 ml) Component Supplier Catalogue number
 Table S1. Components for knockout serum replacer medium (KSR medium) (500 ml) Component Supplier Catalogue number Stock Concentration Final concentration Volume used Advanced DMEM/F12 Invitrogen 12634010-78%
Table S1. Components for knockout serum replacer medium (KSR medium) (500 ml) Component Supplier Catalogue number Stock Concentration Final concentration Volume used Advanced DMEM/F12 Invitrogen 12634010-78%
Primers used for PCR of conductin, SGK1 and GAPDH have been described in (Dehner et al,
 Supplementary METHODS Flow Cytometry (FACS) For FACS analysis, trypsinized cells were fixed in ethanol, rehydrated in PBS and treated with 40μg/ml propidium iodide and 10μ/ml RNase for 30 min at room temperature.
Supplementary METHODS Flow Cytometry (FACS) For FACS analysis, trypsinized cells were fixed in ethanol, rehydrated in PBS and treated with 40μg/ml propidium iodide and 10μ/ml RNase for 30 min at room temperature.
monoclonal antibody. (a) The specificity of the anti-rhbdd1 monoclonal antibody was examined in
 Supplementary information Supplementary figures Supplementary Figure 1 Determination of the s pecificity of in-house anti-rhbdd1 mouse monoclonal antibody. (a) The specificity of the anti-rhbdd1 monoclonal
Supplementary information Supplementary figures Supplementary Figure 1 Determination of the s pecificity of in-house anti-rhbdd1 mouse monoclonal antibody. (a) The specificity of the anti-rhbdd1 monoclonal
Applications and Uses. (adapted from Roche RealTime PCR Application Manual)
 What Can You Do With qpcr? Applications and Uses (adapted from Roche RealTime PCR Application Manual) What is qpcr? Real time PCR also known as quantitative PCR (qpcr) measures PCR amplification as it
What Can You Do With qpcr? Applications and Uses (adapted from Roche RealTime PCR Application Manual) What is qpcr? Real time PCR also known as quantitative PCR (qpcr) measures PCR amplification as it
Premade Lentiviral Particles for human ips Stem Factors
 Premade Lentiviral Particles for human ips Stem Factors For generating induced pluripotent stem (ips) cells or other applications RESEARCH USE ONLY, not for diagnostics or therapeutics Cat# Product Name
Premade Lentiviral Particles for human ips Stem Factors For generating induced pluripotent stem (ips) cells or other applications RESEARCH USE ONLY, not for diagnostics or therapeutics Cat# Product Name
SUPPLEMENTARY INFORMATION
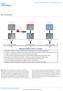 DOI: 10.1038/ncb2239 Hepatocytes (Endoderm) Foreskin fibroblasts (Mesoderm) Melanocytes (Ectoderm) +Dox rtta TetO CMV O,S,M,K & N H1 hes H7 hes H9 hes H1 hes-derived EBs H7 hes-derived EBs H9 hes-derived
DOI: 10.1038/ncb2239 Hepatocytes (Endoderm) Foreskin fibroblasts (Mesoderm) Melanocytes (Ectoderm) +Dox rtta TetO CMV O,S,M,K & N H1 hes H7 hes H9 hes H1 hes-derived EBs H7 hes-derived EBs H9 hes-derived
i-stop codon positions in the mcherry gene
 Supplementary Figure 1 i-stop codon positions in the mcherry gene The grnas (green) that can potentially generate stop codons from Trp (63 th and 98 th aa, upper panel) and Gln (47 th and 114 th aa, bottom
Supplementary Figure 1 i-stop codon positions in the mcherry gene The grnas (green) that can potentially generate stop codons from Trp (63 th and 98 th aa, upper panel) and Gln (47 th and 114 th aa, bottom
A novel tool for monitoring endogenous alpha-synuclein transcription by NanoLuciferase
 A novel tool for monitoring endogenous alpha-synuclein transcription by NanoLuciferase tag insertion at the 3 end using CRISPR-Cas9 genome editing technique Sambuddha Basu 1, 3, Levi Adams 1, 3, Subhrangshu
A novel tool for monitoring endogenous alpha-synuclein transcription by NanoLuciferase tag insertion at the 3 end using CRISPR-Cas9 genome editing technique Sambuddha Basu 1, 3, Levi Adams 1, 3, Subhrangshu
SUPPLEMENTARY INFORMATION
 Secondary mutations as a mechanism of cisplatin resistance in BRCA2-mutated cancers Wataru Sakai, Elizabeth M. Swisher, Beth Y. Karlan, Mukesh K. Agarwal, Jake Higgins, Cynthia Friedman, Emily Villegas,
Secondary mutations as a mechanism of cisplatin resistance in BRCA2-mutated cancers Wataru Sakai, Elizabeth M. Swisher, Beth Y. Karlan, Mukesh K. Agarwal, Jake Higgins, Cynthia Friedman, Emily Villegas,
Supplementary Fig. 1. (A) Working model. The pluripotency transcription factor OCT4
 SUPPLEMENTARY FIGURE LEGENDS Supplementary Fig. 1. (A) Working model. The pluripotency transcription factor OCT4 directly up-regulates the expression of NIPP1 and CCNF that together inhibit protein phosphatase
SUPPLEMENTARY FIGURE LEGENDS Supplementary Fig. 1. (A) Working model. The pluripotency transcription factor OCT4 directly up-regulates the expression of NIPP1 and CCNF that together inhibit protein phosphatase
Supplementary Information
 Supplementary Information Super-resolution imaging of fluorescently labeled, endogenous RNA Polymerase II in living cells with CRISPR/Cas9-mediated gene editing Won-Ki Cho 1, Namrata Jayanth 1, Susan Mullen
Supplementary Information Super-resolution imaging of fluorescently labeled, endogenous RNA Polymerase II in living cells with CRISPR/Cas9-mediated gene editing Won-Ki Cho 1, Namrata Jayanth 1, Susan Mullen
RNA-Guided Gene Activation by CRISPR-Cas9-Based Transcription Factors
 Supplementary Information RNA-Guided Gene Activation by CRISPR-Cas9-Based Transcription Factors Pablo Perez-Pinera 1, Daniel D. Kocak 1, Christopher M. Vockley 2,3, Andrew F. Adler 1, Ami M. Kabadi 1,
Supplementary Information RNA-Guided Gene Activation by CRISPR-Cas9-Based Transcription Factors Pablo Perez-Pinera 1, Daniel D. Kocak 1, Christopher M. Vockley 2,3, Andrew F. Adler 1, Ami M. Kabadi 1,
