Supplementary Materials for
|
|
|
- Magdalene Higgins
- 5 years ago
- Views:
Transcription
1 Supplementary Materials for Estrogen Alters the Splicing of Type 1 Corticotropin-Releasing Hormone Receptor in Breast Cancer Cells Suchita Lal, Anna Allan, Danijela Markovic, Rosemary Walker, James Macartney, Nick Europe-Finner, Alison Tyson-Capper, Dimitris K. Grammatopoulos* *Corresponding author. d.grammatopoulos@warwick.ac.uk Published 2 July 2013, Sci. Signal. 6, ra53 (2013) DOI: /scisignal The PDF file includes: Fig. S1. CRH-regulated genes involved in estrogen signaling in MCF7 cells and in silico predicted integrated network. Fig. S2. Use of receptor antagonists to identify CRH-R subtypes mediating CRH effects in MCF7 cells Fig. S3. CRH-regulated β-catenin expression in MCF7 cells. Fig. S4. Kinetics of basal and CRH regulated MCF7 cell invasion. Fig. S5. CRH-R1 and GPR30 mrna and protein expression in MCF7 cells. Fig. S6. Effect of knockdown of SRp55 on CRH- and E 2 -induced transcriptional effects in MCF7 cells. Fig. S7. Subcellular localization of SRp55 expression in E 2 -treated cells.
2 SUPPLEMENTARY MATERIALS Figure S1: CRH-regulated genes involved in estrogen signaling in MCF7 cells and in silico predicted integrated network. (A to B) Analysis using the Human Estrogen Receptor Signaling in Breast Cancer RT² Profiler PCR Array and target-specific real-time RT-PCR. (A) Heat map of genes altered 24 hours after treatment with 100nM CRH compared with 1% acetic acid (control vehicle). The maximum expression of any given gene is shown in red; minimum expression is shown in green. Representative results from
3 three independent biological replicates are shown. (B) Volcano plot of gene expression in (A), showing expression in response to CRH versus vehicle. Each circle in the plot represents one gene. Genes which are increased at least 2-fold are in red whereas the genes which are decreased at least 2-fold are in green; genes represented by black circles were not differentially regulated (P<0.05, paired Student s t-test N=3). (C) In silico predicted integrated network of genes with significantly altered expression in response to CRH. : genes identified as increased in ERα(+) breast tumors (55). : genes whose expression is weakly inhibited by E 2 in MCF7 cells (54). Solid lines denote distinct regulatory relationships as indicated, whereas dotted lines indicate predicted interactions. (D to F) Real-time RT- PCR for (D) ESR1 and GPER, (E) ESR2, or (F) CRH-R1 and CRH-R2 in MCF7 cells stimulated with CRH as in (A). Data are mean ± SEM % or fold change relative to control or basal conditions against GAPDH expression from at least three independent biological replicates (*P<0.05, Mann-Whitney U test; N=3).
4 Figure S2: Use of receptor antagonists to identify CRH-R subtypes mediating CRH effects in MCF7 cells. (A and B) Western blots for the phosphorylation of GSK3β at Ser 9, Akt, and p38 in (A) MCF7 cells stimulated with CRH for 5 minutes and pre-treated with 1µM NBI (CRH-R1-specific antagonist), 1µM anti-sauvagine-30 (CRH-R2 antagonist) or vehicle (control) for 2 hours or (B) unstimulated cells. Data are mean ± SEM % change versus controls from at least three independent biological replicates (*P <0.05, Mann-Whitney U test; N=3). (C) Real-time RT-PCR for KRT18 and GNAS expression in MCF7 cells stimulated with 100nM CRH for 24hours and pre-treated with specific CRH-R antagonists as in (A) or vehicle. Data are mean ± SEM fold change relative to basal conditions, normalized to GAPDH, of at least three independent biological replicates (*P <0.05, paired Student s t-test N=3).
5 Figure S3: CRH-regulated β-catenin expression in MCF7 cells. (A C) Western blots with lysates from MCF7 cells treated with (A) 40 nm LiCl, (B) 100nM CRH for up to 120 min, or (C) 300nM sfrp1, 100nM CRH, or combined for 3 hours,. Immunoblots shown are representative of three independent experiments. Bar graphs show means ± SEM of % change relative to basal (untreated) of at least three independent biological replicates. (*P <0.05, Mann-Whitney U test; N=3; + P<0.05, Kruskal-Wallis ANOVA followed by Bonferoni test, N=3).
6 Figure S4: Kinetics of basal and CRH regulated MCF7 cell invasion. Kinetics of invasion was measured with the xcelligence System. Cells were either treated with 1% acetic acid (control; squares) or treated with 100 nm CRH (circles), and invasion was measured over 19 hours at frequent time intervals. Results shown are means ± SEM of the Cell Index curves of at least three independent biological replicates.
7 Figure S5: CRH-R1 and GPR30 mrna and protein expression in MCF7 cells. (A) Conventional RT-PCR for CRH-R1α or CRH-R1( 12) in MCF7 cells. PCR products were subsequently electrophoresed and visualized by ethidium bromide. Photomicrographs shown are representative of three independent experiments. (B) Confocal microscopy images of CRH-R1 in MCF7 cells transfected with or without CRH-R2 sirna prior to stimulation with 100nM E 2 for 24 hours. Photomicrographs shown are representative of three independent experiments. (C) Real time PCR of GPR30 expression from RNA isolated from MCF7 cells stimulated with E 2. Data are mean ± SEM of % change versus control, normalized togapdh mrna, from at least three independent biological replicates (*P<0.05, Mann-Whitney U test, N=3). Representative confocal microscopy images of GPR30 localization in MCF7 cells treated with E 2 (inset, C). Photomicrographs shown are representative of three independent experiments.
8 Figure S6: Effect of knockdown of SRp55 on CRH- and E 2 -induced transcriptional effects in MCF7 cells. (A-B) Real time PCR of (A) KRT18 and GNAS or (B) CRH-R2 expression in MCF7 cells transfected with or without SRSF6 sirna prior to stimulation with either 100nM CRH for 24 hours (A) or 100nM E 2 for 24 hours (B). Data are mean ± SEM fold change relative to control from at least three independent biological replicates, normalized to GAPDH expression. (*P<0.05, Mann-Whitney U test, N=3).
9 Figure S7: Subcellular localization of SRp55 expression in E 2 -treated cells. Representative confocal microscopy images of immunofluorescent-labelled SRp55 localization in MCF7 cells treated with or without 100nM E 2 for 2 hours. Photomicrographs shown are representative of three independent experiments.
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/5/233/ra50/dc1 Supplementary Materials for Epidermal Growth Factor Receptor Is Essential for Toll-Like Receptor 3 Signaling Michifumi Yamashita, Saurabh Chattopadhyay,
www.sciencesignaling.org/cgi/content/full/5/233/ra50/dc1 Supplementary Materials for Epidermal Growth Factor Receptor Is Essential for Toll-Like Receptor 3 Signaling Michifumi Yamashita, Saurabh Chattopadhyay,
T H E J O U R N A L O F C E L L B I O L O G Y
 T H E J O U R N A L O F C E L L B I O L O G Y Supplemental material Rainero et al., http://www.jcb.org/cgi/content/full/jcb.201109112/dc1 Figure S1. The expression of DGK- is reduced upon transfection
T H E J O U R N A L O F C E L L B I O L O G Y Supplemental material Rainero et al., http://www.jcb.org/cgi/content/full/jcb.201109112/dc1 Figure S1. The expression of DGK- is reduced upon transfection
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/4/198/ra74/dc1 Supplementary Materials for Short RNA Duplexes Elicit RIG-I Mediated Apoptosis in a Cell Type and Length-Dependent Manner Osamu Ishibashi, Md. Moksed
www.sciencesignaling.org/cgi/content/full/4/198/ra74/dc1 Supplementary Materials for Short RNA Duplexes Elicit RIG-I Mediated Apoptosis in a Cell Type and Length-Dependent Manner Osamu Ishibashi, Md. Moksed
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/8/362/ra12/dc1 Supplementary Materials for FOXP1 potentiates Wnt/β-catenin signaling in diffuse large B cell lymphoma Matthew P. Walker, Charles M. Stopford, Maria
www.sciencesignaling.org/cgi/content/full/8/362/ra12/dc1 Supplementary Materials for FOXP1 potentiates Wnt/β-catenin signaling in diffuse large B cell lymphoma Matthew P. Walker, Charles M. Stopford, Maria
Supplementary Figure 1 PARP1 is involved in regulating the stability of mrnas from pro-inflammatory cytokine/chemokine mediators.
 Supplementary Figure 1 PARP1 is involved in regulating the stability of mrnas from pro-inflammatory cytokine/chemokine mediators. (a) A graphic depiction of the approach to determining the stability of
Supplementary Figure 1 PARP1 is involved in regulating the stability of mrnas from pro-inflammatory cytokine/chemokine mediators. (a) A graphic depiction of the approach to determining the stability of
Supplementary Figure 1. Confirmation of sirna in PC3 and H1299 cells PC3 (a) and H1299 (b) cells were transfected with sirna oligonucleotides
 Supplementary Figure 1. Confirmation of sirna in PC3 and H1299 cells PC3 (a) and H1299 (b) cells were transfected with sirna oligonucleotides targeting RCP (SMARTPool (RCP) or two individual oligos (RCP#1
Supplementary Figure 1. Confirmation of sirna in PC3 and H1299 cells PC3 (a) and H1299 (b) cells were transfected with sirna oligonucleotides targeting RCP (SMARTPool (RCP) or two individual oligos (RCP#1
Supplementary Information. A novel human endogenous retroviral protein inhibits cell-cell fusion. Supplementary Figures:
 Supplementary Information A novel human endogenous retroviral protein inhibits cell-cell fusion Jun Sugimoto, Makiko Sugimoto, Helene Bernstein, Yoshihiro Jinno and Danny J. Schust Supplementary Figures:
Supplementary Information A novel human endogenous retroviral protein inhibits cell-cell fusion Jun Sugimoto, Makiko Sugimoto, Helene Bernstein, Yoshihiro Jinno and Danny J. Schust Supplementary Figures:
Supplementary Figure 1. Characterization of EVs (a) Phase-contrast electron microscopy was used to visualize resuspended EV pellets.
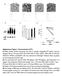 Supplementary Figure 1. Characterization of EVs (a) Phase-contrast electron microscopy was used to visualize resuspended EV pellets. Scale bar represent 100 nm. The sizes of EVs from MDA-MB-231-D3H1 (D3H1),
Supplementary Figure 1. Characterization of EVs (a) Phase-contrast electron microscopy was used to visualize resuspended EV pellets. Scale bar represent 100 nm. The sizes of EVs from MDA-MB-231-D3H1 (D3H1),
Heparan Sulphate in Breast Cancer Low, Y.L.A 1 and Yip, C.W.G 2
 Heparan Sulphate in Breast Cancer Low, Y.L.A 1 and Yip, C.W.G 2 Department of Anatomy, Yong Loo Lin School of Medicine, National University of Singapore MD10, 4 Medical Drive, Singapore 117597 ABSTRACT
Heparan Sulphate in Breast Cancer Low, Y.L.A 1 and Yip, C.W.G 2 Department of Anatomy, Yong Loo Lin School of Medicine, National University of Singapore MD10, 4 Medical Drive, Singapore 117597 ABSTRACT
Stargazin regulates AMPA receptor trafficking through adaptor protein. complexes during long term depression
 Supplementary Information Stargazin regulates AMPA receptor trafficking through adaptor protein complexes during long term depression Shinji Matsuda, Wataru Kakegawa, Timotheus Budisantoso, Toshihiro Nomura,
Supplementary Information Stargazin regulates AMPA receptor trafficking through adaptor protein complexes during long term depression Shinji Matsuda, Wataru Kakegawa, Timotheus Budisantoso, Toshihiro Nomura,
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb3363 Supplementary Figure 1 Several WNTs bind to the extracellular domains of PKD1. (a) HEK293T cells were co-transfected with indicated plasmids. Flag-tagged proteins were immunoprecipiated
DOI: 10.1038/ncb3363 Supplementary Figure 1 Several WNTs bind to the extracellular domains of PKD1. (a) HEK293T cells were co-transfected with indicated plasmids. Flag-tagged proteins were immunoprecipiated
Dual PI3K/ERK inhibition induces necroptotic cell death of Hodgkin Lymphoma cells through IER3 downregulation
 Dual PI3K/ERK inhibition induces necroptotic cell death of Hodgkin Lymphoma cells through IER3 downregulation *Silvia Laura Locatelli, 1 Giuseppa Careddu, 1 Giuliano Giuseppe Stirparo, 1 Luca Castagna,
Dual PI3K/ERK inhibition induces necroptotic cell death of Hodgkin Lymphoma cells through IER3 downregulation *Silvia Laura Locatelli, 1 Giuseppa Careddu, 1 Giuliano Giuseppe Stirparo, 1 Luca Castagna,
Revision Checklist for Science Signaling Research Manuscripts: Data Requirements and Style Guidelines
 Revision Checklist for Science Signaling Research Manuscripts: Data Requirements and Style Guidelines Further information can be found at: http://stke.sciencemag.org/sites/default/files/researcharticlerevmsinstructions_0.pdf.
Revision Checklist for Science Signaling Research Manuscripts: Data Requirements and Style Guidelines Further information can be found at: http://stke.sciencemag.org/sites/default/files/researcharticlerevmsinstructions_0.pdf.
Supplementary Figure 1. Co-localization of GLUT1 and DNAL4 in BeWo cells cultured
 Supplementary Figure 1. Co-localization of GLUT1 and DNAL4 in BeWo cells cultured under static conditions. Cells were seeded in the chamber area of the device and cultured overnight without medium perfusion.
Supplementary Figure 1. Co-localization of GLUT1 and DNAL4 in BeWo cells cultured under static conditions. Cells were seeded in the chamber area of the device and cultured overnight without medium perfusion.
Supplemental Table 1 Gene Symbol FDR corrected p-value PLOD1 CSRP2 PFKP ADFP ADM C10orf10 GPI LOX PLEKHA2 WIPF1
 Supplemental Table 1 Gene Symbol FDR corrected p-value PLOD1 4.52E-18 PDK1 6.77E-18 CSRP2 4.42E-17 PFKP 1.23E-14 MSH2 3.79E-13 NARF_A 5.56E-13 ADFP 5.56E-13 FAM13A1 1.56E-12 FAM29A_A 1.22E-11 CA9 1.54E-11
Supplemental Table 1 Gene Symbol FDR corrected p-value PLOD1 4.52E-18 PDK1 6.77E-18 CSRP2 4.42E-17 PFKP 1.23E-14 MSH2 3.79E-13 NARF_A 5.56E-13 ADFP 5.56E-13 FAM13A1 1.56E-12 FAM29A_A 1.22E-11 CA9 1.54E-11
Supplementary Information
 Supplementary Information Supplementary Figures Supplementary Figure 1. MLK1-4 phosphorylate MEK in the presence of RAF inhibitors. (a) H157 cells were transiently transfected with Flag- or HA-tagged MLK1-4
Supplementary Information Supplementary Figures Supplementary Figure 1. MLK1-4 phosphorylate MEK in the presence of RAF inhibitors. (a) H157 cells were transiently transfected with Flag- or HA-tagged MLK1-4
Supplemental Fig. 1: PEA-15 knockdown efficiency assessed by immunohistochemistry and qpcr
 Supplemental figure legends Supplemental Fig. 1: PEA-15 knockdown efficiency assessed by immunohistochemistry and qpcr A, LβT2 cells were transfected with either scrambled or PEA-15 sirna. Cells were then
Supplemental figure legends Supplemental Fig. 1: PEA-15 knockdown efficiency assessed by immunohistochemistry and qpcr A, LβT2 cells were transfected with either scrambled or PEA-15 sirna. Cells were then
JCB. Supplemental material THE JOURNAL OF CELL BIOLOGY. Kimura et al.,
 Supplemental material JCB Kimura et al., http://www.jcb.org/cgi/content/full/jcb.201503023/dc1 THE JOURNAL OF CELL BIOLOGY Figure S1. TRIMs regulate IFN-γ induced autophagy. (A and B) HC image analysis
Supplemental material JCB Kimura et al., http://www.jcb.org/cgi/content/full/jcb.201503023/dc1 THE JOURNAL OF CELL BIOLOGY Figure S1. TRIMs regulate IFN-γ induced autophagy. (A and B) HC image analysis
supplementary information
 DOI: 1.138/ncb1839 a b Control 1 2 3 Control 1 2 3 Fbw7 Smad3 1 2 3 4 1 2 3 4 c d IGF-1 IGF-1Rβ IGF-1Rβ-P Control / 1 2 3 4 Real-time RT-PCR Relative quantity (IGF-1/ mrna) 2 1 IGF-1 1 2 3 4 Control /
DOI: 1.138/ncb1839 a b Control 1 2 3 Control 1 2 3 Fbw7 Smad3 1 2 3 4 1 2 3 4 c d IGF-1 IGF-1Rβ IGF-1Rβ-P Control / 1 2 3 4 Real-time RT-PCR Relative quantity (IGF-1/ mrna) 2 1 IGF-1 1 2 3 4 Control /
Supplementary Information
 Supplementary Information MLL histone methylases regulate expression of HDLR- in presence of estrogen and control plasma cholesterol in vivo Khairul I. Ansari 1, Sahba Kasiri 1, Imran Hussain 1, Samara
Supplementary Information MLL histone methylases regulate expression of HDLR- in presence of estrogen and control plasma cholesterol in vivo Khairul I. Ansari 1, Sahba Kasiri 1, Imran Hussain 1, Samara
Supplementary Figure Legend
 Supplementary Figure Legend Supplementary Figure S1. Effects of MMP-1 silencing on HEp3-hi/diss cell proliferation in 2D and 3D culture conditions. (A) Downregulation of MMP-1 expression in HEp3-hi/diss
Supplementary Figure Legend Supplementary Figure S1. Effects of MMP-1 silencing on HEp3-hi/diss cell proliferation in 2D and 3D culture conditions. (A) Downregulation of MMP-1 expression in HEp3-hi/diss
Supplementary Figure1: ClustalW comparison between Tll, Dsf and NR2E1.
 P-Box Dsf -----------------MG-TAG--DRLLD-IPCKVCGDRSSGKHYGIYSCDGCSGFFKR 39 NR2E1 -----------------MSKPAGSTSRILD-IPCKVCGDRSSGKHYGVYACDGCSGFFKR 42 Tll MQSSEGSPDMMDQKYNSVRLSPAASSRILYHVPCKVCRDHSSGKHYGIYACDGCAGFFKR
P-Box Dsf -----------------MG-TAG--DRLLD-IPCKVCGDRSSGKHYGIYSCDGCSGFFKR 39 NR2E1 -----------------MSKPAGSTSRILD-IPCKVCGDRSSGKHYGVYACDGCSGFFKR 42 Tll MQSSEGSPDMMDQKYNSVRLSPAASSRILYHVPCKVCRDHSSGKHYGIYACDGCAGFFKR
TE5 KYSE510 TE7 KYSE70 KYSE140
 TE5 KYSE5 TT KYSE7 KYSE4 Supplementary Figure. Hockey stick plots showing input normalized, rank ordered H3K7ac signals for the candidate SE-associated lncrnas in this study. Rpm Rpm Rpm Chip-seq H3K7ac
TE5 KYSE5 TT KYSE7 KYSE4 Supplementary Figure. Hockey stick plots showing input normalized, rank ordered H3K7ac signals for the candidate SE-associated lncrnas in this study. Rpm Rpm Rpm Chip-seq H3K7ac
Supplementary Table 1. Sequences for BTG2 and BRCA1 sirnas.
 Supplementary Table 1. Sequences for BTG2 and BRCA1 sirnas. Target Gene Non-target / Control BTG2 BRCA1 NFE2L2 Target Sequence ON-TARGET plus Non-targeting sirna # 1 (Cat# D-001810-01-05) sirna1: GAACCGACAUGCUCCCGGA
Supplementary Table 1. Sequences for BTG2 and BRCA1 sirnas. Target Gene Non-target / Control BTG2 BRCA1 NFE2L2 Target Sequence ON-TARGET plus Non-targeting sirna # 1 (Cat# D-001810-01-05) sirna1: GAACCGACAUGCUCCCGGA
Emanuela Tumini, Sonia Barroso, Carmen Pérez Calero and Andrés Aguilera
 SUPPLEMENTARY INFORMATION Roles of human POLD1 and POLD3 in genome stability Emanuela Tumini, Sonia Barroso, Carmen Pérez Calero and Andrés Aguilera SUPPLEMENTARY METHODS Cell proliferation After sirna
SUPPLEMENTARY INFORMATION Roles of human POLD1 and POLD3 in genome stability Emanuela Tumini, Sonia Barroso, Carmen Pérez Calero and Andrés Aguilera SUPPLEMENTARY METHODS Cell proliferation After sirna
monoclonal antibody. (a) The specificity of the anti-rhbdd1 monoclonal antibody was examined in
 Supplementary information Supplementary figures Supplementary Figure 1 Determination of the s pecificity of in-house anti-rhbdd1 mouse monoclonal antibody. (a) The specificity of the anti-rhbdd1 monoclonal
Supplementary information Supplementary figures Supplementary Figure 1 Determination of the s pecificity of in-house anti-rhbdd1 mouse monoclonal antibody. (a) The specificity of the anti-rhbdd1 monoclonal
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb3164 Supplementary Figure 1 Validation of effective Gnas deletion and epithelial thickness. a, Representative genotyping in mice treated or not with tamoxifen to show Gnas deletion. To
DOI: 10.1038/ncb3164 Supplementary Figure 1 Validation of effective Gnas deletion and epithelial thickness. a, Representative genotyping in mice treated or not with tamoxifen to show Gnas deletion. To
Supplementary Materials for
 advances.sciencemag.org/cgi/content/full/4/9/eaat5401/dc1 Supplementary Materials for GLK-IKKβ signaling induces dimerization and translocation of the AhR-RORγt complex in IL-17A induction and autoimmune
advances.sciencemag.org/cgi/content/full/4/9/eaat5401/dc1 Supplementary Materials for GLK-IKKβ signaling induces dimerization and translocation of the AhR-RORγt complex in IL-17A induction and autoimmune
Supplementary Fig. 1. (A) Working model. The pluripotency transcription factor OCT4
 SUPPLEMENTARY FIGURE LEGENDS Supplementary Fig. 1. (A) Working model. The pluripotency transcription factor OCT4 directly up-regulates the expression of NIPP1 and CCNF that together inhibit protein phosphatase
SUPPLEMENTARY FIGURE LEGENDS Supplementary Fig. 1. (A) Working model. The pluripotency transcription factor OCT4 directly up-regulates the expression of NIPP1 and CCNF that together inhibit protein phosphatase
T H E J O U R N A L O F C E L L B I O L O G Y
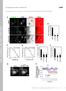 T H E J O U R N A L O F C E L L B I O L O G Y Supplemental material Yamaguchi et al., http://www.jcb.org/cgi/content/full/jcb.201009126/dc1 S1 Figure S2. The expression levels of GFP-Akt1-PH and localization
T H E J O U R N A L O F C E L L B I O L O G Y Supplemental material Yamaguchi et al., http://www.jcb.org/cgi/content/full/jcb.201009126/dc1 S1 Figure S2. The expression levels of GFP-Akt1-PH and localization
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/9/429/ra54/dc1 Supplementary Materials for Dephosphorylation of the adaptor LAT and phospholipase C by SHP-1 inhibits natural killer cell cytotoxicity Omri Matalon,
www.sciencesignaling.org/cgi/content/full/9/429/ra54/dc1 Supplementary Materials for Dephosphorylation of the adaptor LAT and phospholipase C by SHP-1 inhibits natural killer cell cytotoxicity Omri Matalon,
Supplementary Figure 1, related to Figure 1. GAS5 is highly expressed in the cytoplasm of hescs, and positively correlates with pluripotency.
 Supplementary Figure 1, related to Figure 1. GAS5 is highly expressed in the cytoplasm of hescs, and positively correlates with pluripotency. (a) Transfection of different concentration of GAS5-overexpressing
Supplementary Figure 1, related to Figure 1. GAS5 is highly expressed in the cytoplasm of hescs, and positively correlates with pluripotency. (a) Transfection of different concentration of GAS5-overexpressing
Supplementary Figure 1. NORAD expression in mouse (A) and dog (B). The black boxes indicate the position of the regions alignable to the 12 repeat
 Supplementary Figure 1. NORAD expression in mouse (A) and dog (B). The black boxes indicate the position of the regions alignable to the 12 repeat units in the human genome. Annotated transposable elements
Supplementary Figure 1. NORAD expression in mouse (A) and dog (B). The black boxes indicate the position of the regions alignable to the 12 repeat units in the human genome. Annotated transposable elements
Cell proliferation was measured with Cell Counting Kit-8 (Dojindo Laboratories, Kumamoto, Japan).
 1 2 3 4 5 6 7 8 Supplemental Materials and Methods Cell proliferation assay Cell proliferation was measured with Cell Counting Kit-8 (Dojindo Laboratories, Kumamoto, Japan). GCs were plated at 96-well
1 2 3 4 5 6 7 8 Supplemental Materials and Methods Cell proliferation assay Cell proliferation was measured with Cell Counting Kit-8 (Dojindo Laboratories, Kumamoto, Japan). GCs were plated at 96-well
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb2743 Figure S1 stabilizes cellular protein level, post-transcriptionally. (a, b) and DDR1 were RNAi-depleted from HEK.293.-CBG cells. Western blots with indicated antibodies (a). RT-PCRs
DOI: 10.1038/ncb2743 Figure S1 stabilizes cellular protein level, post-transcriptionally. (a, b) and DDR1 were RNAi-depleted from HEK.293.-CBG cells. Western blots with indicated antibodies (a). RT-PCRs
Supplemental Table 1 Primers used in study. Human. Mouse
 Supplemental Table 1 Primers used in study Human Forward primer region(5-3 ) Reverse primer region(5-3 ) RT-PCR GAPDH gagtcaacggatttggtcgt ttgattttggagggatctcg Raftlin atgggttgcggattgaacaagttaga ctgaggtataacaccaacgaatttcaggc
Supplemental Table 1 Primers used in study Human Forward primer region(5-3 ) Reverse primer region(5-3 ) RT-PCR GAPDH gagtcaacggatttggtcgt ttgattttggagggatctcg Raftlin atgggttgcggattgaacaagttaga ctgaggtataacaccaacgaatttcaggc
Supplementary Figure 1. (A) Cell proliferative ability and (B) the invasiveness of
 LEGEND FOR SUPPLEMENTARY FIGURES Supplementary Figure 1. (A) Cell proliferative ability and (B) the invasiveness of LLC-1, LLC-3, and LLC-5 cell line series were quantified by BrdU assay after 72 h of
LEGEND FOR SUPPLEMENTARY FIGURES Supplementary Figure 1. (A) Cell proliferative ability and (B) the invasiveness of LLC-1, LLC-3, and LLC-5 cell line series were quantified by BrdU assay after 72 h of
Journal of Cell Science Supplementary Material
 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 SUPPLEMENTARY FIGURE LEGENDS Figure S1: Eps8 is localized at focal adhesions and binds directly to FAK (A) Focal
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 SUPPLEMENTARY FIGURE LEGENDS Figure S1: Eps8 is localized at focal adhesions and binds directly to FAK (A) Focal
Inhibition of Twist1 mediated invasion by Chk2 promotes premature senescence in p53 defective cancer cells
 Inhibition of Twist1 mediated invasion by Chk2 promotes premature senescence in p53 defective cancer cells Debasis Nayak, a,1 Anmol Kumar, b Souneek Chakraborty, a,1 Reyaz ur Rasool, a,1 Hina Amin, a Archana
Inhibition of Twist1 mediated invasion by Chk2 promotes premature senescence in p53 defective cancer cells Debasis Nayak, a,1 Anmol Kumar, b Souneek Chakraborty, a,1 Reyaz ur Rasool, a,1 Hina Amin, a Archana
Supplementary Figure 1.
 Supplementary Figure 1. Quantification of western blot analysis of fibroblasts (related to Figure 1) (A-F) Quantification of western blot analysis for control and IR-Mut fibroblasts. Data are expressed
Supplementary Figure 1. Quantification of western blot analysis of fibroblasts (related to Figure 1) (A-F) Quantification of western blot analysis for control and IR-Mut fibroblasts. Data are expressed
Supplementary figures and tables Table of content
 Supplementary figures: Supplementary figures and tables Table of content Figure S1: The minimal i model written with BIOCHAM reaction rules. Figure S2: Ordinary differential equation (ODE) set of the model.
Supplementary figures: Supplementary figures and tables Table of content Figure S1: The minimal i model written with BIOCHAM reaction rules. Figure S2: Ordinary differential equation (ODE) set of the model.
Nature Neuroscience: doi: /nn Supplementary Figure 1
 Supplementary Figure 1 PCR-genotyping of the three mouse models used in this study and controls for behavioral experiments after semi-chronic Pten inhibition. a-c. DNA from App/Psen1 (a), Pten tg (b) and
Supplementary Figure 1 PCR-genotyping of the three mouse models used in this study and controls for behavioral experiments after semi-chronic Pten inhibition. a-c. DNA from App/Psen1 (a), Pten tg (b) and
Gene Sequence Fragment size ΔEGFR F 5' GGGCTCTGGAGGAAAAGAAAG GT 3' 116 bp R 5' CTTCTTACACTTGCGGACGC 3'
 Supplementary Table 1: Real-time PCR primer sequences for ΔEGFR, wtegfr, IL-6, LIF and GAPDH. Gene Sequence Fragment size ΔEGFR F 5' GGGCTCTGGAGGAAAAGAAAG GT 3' 116 bp R 5' CTTCTTACACTTGCGGACGC 3' wtegfr
Supplementary Table 1: Real-time PCR primer sequences for ΔEGFR, wtegfr, IL-6, LIF and GAPDH. Gene Sequence Fragment size ΔEGFR F 5' GGGCTCTGGAGGAAAAGAAAG GT 3' 116 bp R 5' CTTCTTACACTTGCGGACGC 3' wtegfr
Supplementary Figure 1. (a) The qrt-pcr for lnc-2, lnc-6 and lnc-7 RNA level in DU145, 22Rv1, wild type HCT116 and HCT116 Dicer ex5 cells transfected
 Supplementary Figure 1. (a) The qrt-pcr for lnc-2, lnc-6 and lnc-7 RNA level in DU145, 22Rv1, wild type HCT116 and HCT116 Dicer ex5 cells transfected with the sirna against lnc-2, lnc-6, lnc-7, and the
Supplementary Figure 1. (a) The qrt-pcr for lnc-2, lnc-6 and lnc-7 RNA level in DU145, 22Rv1, wild type HCT116 and HCT116 Dicer ex5 cells transfected with the sirna against lnc-2, lnc-6, lnc-7, and the
Supporting Information
 Supporting Information Chakrabarty et al. 10.1073/pnas.1018001108 SI Materials and Methods Cell Lines. All cell lines were purchased from the American Type Culture Collection. Media and FBS were purchased
Supporting Information Chakrabarty et al. 10.1073/pnas.1018001108 SI Materials and Methods Cell Lines. All cell lines were purchased from the American Type Culture Collection. Media and FBS were purchased
SureSilencing sirna Array Technology Overview
 SureSilencing sirna Array Technology Overview Pathway-Focused sirna-based RNA Interference Topics to be Covered Who is SuperArray? Brief Introduction to RNA Interference Challenges Facing RNA Interference
SureSilencing sirna Array Technology Overview Pathway-Focused sirna-based RNA Interference Topics to be Covered Who is SuperArray? Brief Introduction to RNA Interference Challenges Facing RNA Interference
Supplementary Fig. 1. Multiple five micron sections of liver tissues of rats treated
 Supplementary Figure Legends Supplementary Fig. 1. Multiple five micron sections of liver tissues of rats treated with either vehicle (left; n=3) or CCl 4 (right; n=3) were co-immunostained for NRP-1 (green)
Supplementary Figure Legends Supplementary Fig. 1. Multiple five micron sections of liver tissues of rats treated with either vehicle (left; n=3) or CCl 4 (right; n=3) were co-immunostained for NRP-1 (green)
Supplementary Figure 1. Related to Figure 1. Characterization of centrosome fragmentation in mitotically delayed RPE1 cells. (a) Cells transiently
 Supplementary Figure 1. Related to Figure 1. Characterization of centrosome fragmentation in mitotically delayed RPE1 cells. (a) Cells transiently transfected with EGFP centrin-2 (Green), synchronized
Supplementary Figure 1. Related to Figure 1. Characterization of centrosome fragmentation in mitotically delayed RPE1 cells. (a) Cells transiently transfected with EGFP centrin-2 (Green), synchronized
Supplementary Figure 1. Adipogenic protein expression in WT and KO MEFs after 7 days of adipogenic differentiation.
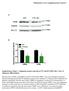 Merkestein et al. Supplementary Figure 1 A PLIN WT FTO KO 72 kda FABP4 17 kda HSC70 72 kda B Supplementary Figure 1. Adipogenic protein expression in WT and KO MEFs after 7 days of adipogenic differentiation.
Merkestein et al. Supplementary Figure 1 A PLIN WT FTO KO 72 kda FABP4 17 kda HSC70 72 kda B Supplementary Figure 1. Adipogenic protein expression in WT and KO MEFs after 7 days of adipogenic differentiation.
T H E J O U R N A L O F C E L L B I O L O G Y
 T H E J O U R N A L O F C E L L B I O L O G Y Supplemental material Han et al., http://www.jcb.org/cgi/content/full/jcb.201311007/dc1 Figure S1. SIVA1 interacts with PCNA. (A) HEK293T cells were transiently
T H E J O U R N A L O F C E L L B I O L O G Y Supplemental material Han et al., http://www.jcb.org/cgi/content/full/jcb.201311007/dc1 Figure S1. SIVA1 interacts with PCNA. (A) HEK293T cells were transiently
Supplementary Fig. 1 related to Fig. 1 Clinical relevance of lncrna candidate
 Supplementary Figure Legends Supplementary Fig. 1 related to Fig. 1 Clinical relevance of lncrna candidate BC041951 in gastric cancer. (A) The flow chart for selected candidate lncrnas in 660 up-regulated
Supplementary Figure Legends Supplementary Fig. 1 related to Fig. 1 Clinical relevance of lncrna candidate BC041951 in gastric cancer. (A) The flow chart for selected candidate lncrnas in 660 up-regulated
Supplementary Figure S1 Evi/Wls and Wnt3 are overexpressed in epithelial cells in colon cancers. (a) Validation of the specificity of the Wnt3
 Supplementary Figure S1 Evi/Wls and Wnt3 are overexpressed in epithelial cells in colon cancers. (a) Validation of the specificity of the Wnt3 antibody. HCT116 cells were reverse transfected with sicontrol
Supplementary Figure S1 Evi/Wls and Wnt3 are overexpressed in epithelial cells in colon cancers. (a) Validation of the specificity of the Wnt3 antibody. HCT116 cells were reverse transfected with sicontrol
(A-B) P2ry14 expression was assessed by (A) genotyping (upper arrow: WT; lower
 Supplementary Figures S1. (A-B) P2ry14 expression was assessed by (A) genotyping (upper arrow: ; lower arrow: KO) and (B) q-pcr analysis with Lin- cells, The white vertical line in panel A indicates that
Supplementary Figures S1. (A-B) P2ry14 expression was assessed by (A) genotyping (upper arrow: ; lower arrow: KO) and (B) q-pcr analysis with Lin- cells, The white vertical line in panel A indicates that
Supplementary Information
 Supplementary Information promotes cancer cell invasion and proliferation by receptor-mediated endocytosis-dependent and -independent mechanisms, respectively Kensaku Shojima, Akira Sato, Hideaki Hanaki,
Supplementary Information promotes cancer cell invasion and proliferation by receptor-mediated endocytosis-dependent and -independent mechanisms, respectively Kensaku Shojima, Akira Sato, Hideaki Hanaki,
Supplementary Figure 1. ips_3y+mir-302b. ips_3y+mir-372. ips_3y+mir-302b + mir-372. ips_4y. ips_4y+mir-302b. ips_4y + mir-372
 Supplementary Figure 1 ips_3y+mir-32b ips_3y+ ips_3y+mir-32b + ips_4y ips_4y+mir-32b ips_4y + Nature Biotechnology: doi:1.138/nb.t.1862 TRA-1-6 DAPI Oct3/4 Supplementary Figure 1: ips cells derived from
Supplementary Figure 1 ips_3y+mir-32b ips_3y+ ips_3y+mir-32b + ips_4y ips_4y+mir-32b ips_4y + Nature Biotechnology: doi:1.138/nb.t.1862 TRA-1-6 DAPI Oct3/4 Supplementary Figure 1: ips cells derived from
SUPPLEMENTAL FIGURE LEGENDS
 SUPPLEMENTAL FIGURE LEGENDS Suppl. Fig. 1. Notch and myostatin expression is unaffected in the absence of p65 during postnatal development. A & B. Myostatin and Notch-1 expression levels were determined
SUPPLEMENTAL FIGURE LEGENDS Suppl. Fig. 1. Notch and myostatin expression is unaffected in the absence of p65 during postnatal development. A & B. Myostatin and Notch-1 expression levels were determined
0.9 5 H M L E R -C tr l in T w is t1 C M
 a. b. c. d. e. f. g. h. 2.5 C elltiter-g lo A ssay 1.1 5 M T S a s s a y Lum inescence (A.U.) 2.0 1.5 1.0 0.5 n s H M L E R -C tr l in C tr l C M H M L E R -C tr l in S n a il1 C M A bsorbance (@ 490nm
a. b. c. d. e. f. g. h. 2.5 C elltiter-g lo A ssay 1.1 5 M T S a s s a y Lum inescence (A.U.) 2.0 1.5 1.0 0.5 n s H M L E R -C tr l in C tr l C M H M L E R -C tr l in S n a il1 C M A bsorbance (@ 490nm
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/6/259/ra5/dc1 Supplementary Materials for Systems Biology Approach Identifies the Kinase Csnk1a1 as a Regulator of the DNA Damage Response in Embryonic Stem Cells
www.sciencesignaling.org/cgi/content/full/6/259/ra5/dc1 Supplementary Materials for Systems Biology Approach Identifies the Kinase Csnk1a1 as a Regulator of the DNA Damage Response in Embryonic Stem Cells
Supplementary Data. For generation of stable cell lines, U87MG cells posttransfection
 Supplementary Data Supplementary Materials and Methods Chromatin immunoprecipitation In this study, chromatin immunoprecipitation (ChIP) experiments were performed with the SimpleChIP Enzymatic Chromatin
Supplementary Data Supplementary Materials and Methods Chromatin immunoprecipitation In this study, chromatin immunoprecipitation (ChIP) experiments were performed with the SimpleChIP Enzymatic Chromatin
Supplementary Figure 1. Reconstitution of human-acquired lymphoid system in
 Supplementary Figure 1. Reconstitution of human-acquired lymphoid system in mouse NOD/SCID/Jak3 null mice were transplanted with human CD34 + hematopoietic stem cells. (Top) Four weeks after the transplantation
Supplementary Figure 1. Reconstitution of human-acquired lymphoid system in mouse NOD/SCID/Jak3 null mice were transplanted with human CD34 + hematopoietic stem cells. (Top) Four weeks after the transplantation
Supplementary Figure 1. APP cleavage assay. HEK293 cells were transfected with various
 Supplementary Figure 1. APP cleavage assay. HEK293 cells were transfected with various GST-tagged N-terminal truncated APP fragments including GST-APP full-length (FL), APP (123-695), APP (189-695), or
Supplementary Figure 1. APP cleavage assay. HEK293 cells were transfected with various GST-tagged N-terminal truncated APP fragments including GST-APP full-length (FL), APP (123-695), APP (189-695), or
Supplementary Figure 1. Soft fibrin gels promote growth and organized mesodermal differentiation. Representative images of single OGTR1 ESCs cultured
 Supplementary Figure 1. Soft fibrin gels promote growth and organized mesodermal differentiation. Representative images of single OGTR1 ESCs cultured in 90-Pa 3D fibrin gels for 5 days in the presence
Supplementary Figure 1. Soft fibrin gels promote growth and organized mesodermal differentiation. Representative images of single OGTR1 ESCs cultured in 90-Pa 3D fibrin gels for 5 days in the presence
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb3240 Supplementary Figure 1 GBM cell lines display similar levels of p100 to p52 processing but respond differentially to TWEAK-induced TERT expression according to TERT promoter mutation
DOI: 10.1038/ncb3240 Supplementary Figure 1 GBM cell lines display similar levels of p100 to p52 processing but respond differentially to TWEAK-induced TERT expression according to TERT promoter mutation
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/3/146/ra80/dc1 Supplementary Materials for DNMT1 Stability Is Regulated by Proteins Coordinating Deubiquitination and Acetylation-Driven Ubiquitination Zhanwen
www.sciencesignaling.org/cgi/content/full/3/146/ra80/dc1 Supplementary Materials for DNMT1 Stability Is Regulated by Proteins Coordinating Deubiquitination and Acetylation-Driven Ubiquitination Zhanwen
Nature Immunology: doi: /ni.3015
 Supplementary Figure 1 Role of RIP1-RIP3 and PGAM5 in RNA virus induced inflammasome activation. (a) LDH release from LPS-primed BMDMs from wild-type mice (WT), Rip3 -/- or Nlrp3 -/- mice infected with
Supplementary Figure 1 Role of RIP1-RIP3 and PGAM5 in RNA virus induced inflammasome activation. (a) LDH release from LPS-primed BMDMs from wild-type mice (WT), Rip3 -/- or Nlrp3 -/- mice infected with
Supplementary Figure 1. Expressions of stem cell markers decreased in TRCs on 2D plastic. TRCs were cultured on plastic for 1, 3, 5, or 7 days,
 Supplementary Figure 1. Expressions of stem cell markers decreased in TRCs on 2D plastic. TRCs were cultured on plastic for 1, 3, 5, or 7 days, respectively, and their mrnas were quantified by real time
Supplementary Figure 1. Expressions of stem cell markers decreased in TRCs on 2D plastic. TRCs were cultured on plastic for 1, 3, 5, or 7 days, respectively, and their mrnas were quantified by real time
supplementary information
 Figure S1 ZEB1 full length mrna. (a) Analysis of the ZEB1 mrna using the UCSC genome browser (http://genome.ucsc.edu) revealed truncation of the annotated Refseq sequence (NM_030751). The probable terminus
Figure S1 ZEB1 full length mrna. (a) Analysis of the ZEB1 mrna using the UCSC genome browser (http://genome.ucsc.edu) revealed truncation of the annotated Refseq sequence (NM_030751). The probable terminus
Hypoxia-induced carbonic anhydrase IX facilitates lactate flux in human breast cancer cells by non-catalytic function
 Hypoxia-induced carbonic anhydrase IX facilitates lactate flux in human breast cancer cells by non-catalytic function Somayeh Jamali 1, Michael Klier 1,2, Samantha Ames 1,2, L. Felipe Barros 3, Robert
Hypoxia-induced carbonic anhydrase IX facilitates lactate flux in human breast cancer cells by non-catalytic function Somayeh Jamali 1, Michael Klier 1,2, Samantha Ames 1,2, L. Felipe Barros 3, Robert
(a) Immunoblotting to show the migration position of Flag-tagged MAVS
 Supplementary Figure 1 Characterization of six MAVS isoforms. (a) Immunoblotting to show the migration position of Flag-tagged MAVS isoforms. HEK293T Mavs -/- cells were transfected with constructs expressing
Supplementary Figure 1 Characterization of six MAVS isoforms. (a) Immunoblotting to show the migration position of Flag-tagged MAVS isoforms. HEK293T Mavs -/- cells were transfected with constructs expressing
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1. Analyses of ECTRs by C-circle and T-circle assays.
 Supplementary Figure 1 Analyses of ECTRs by C-circle and T-circle assays. (a) C-circle and (b) T-circle amplification reactions using genomic DNA from different cell lines in the presence (+) or absence
Supplementary Figure 1 Analyses of ECTRs by C-circle and T-circle assays. (a) C-circle and (b) T-circle amplification reactions using genomic DNA from different cell lines in the presence (+) or absence
Supplementary information
 Supplementary information Supplementary Figure 1 (a) EM image of the pyramidal layer of wt mice. CA3 pyramidal neurons were selected according to their typical alignment, size and shape for subsequent
Supplementary information Supplementary Figure 1 (a) EM image of the pyramidal layer of wt mice. CA3 pyramidal neurons were selected according to their typical alignment, size and shape for subsequent
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION Supplementary Figure 1. Detecting ESR1-CCDC170 and P2RY6-ARHGEF17 chimerical transcripts in breast cancer cell lines by RT-PCR. (a) RT-PCR results using a forward primer (AACAGGCTCGAAAGGTCCATGC)
SUPPLEMENTARY INFORMATION Supplementary Figure 1. Detecting ESR1-CCDC170 and P2RY6-ARHGEF17 chimerical transcripts in breast cancer cell lines by RT-PCR. (a) RT-PCR results using a forward primer (AACAGGCTCGAAAGGTCCATGC)
Primers used for PCR of conductin, SGK1 and GAPDH have been described in (Dehner et al,
 Supplementary METHODS Flow Cytometry (FACS) For FACS analysis, trypsinized cells were fixed in ethanol, rehydrated in PBS and treated with 40μg/ml propidium iodide and 10μ/ml RNase for 30 min at room temperature.
Supplementary METHODS Flow Cytometry (FACS) For FACS analysis, trypsinized cells were fixed in ethanol, rehydrated in PBS and treated with 40μg/ml propidium iodide and 10μ/ml RNase for 30 min at room temperature.
Supplementary Figures and Legends.
 Supplementary Figures and Legends. Supplementary Figure 1: Impact of injury on Rb1 and PPARϒ expression. Following ipsilateral axotomy injury, adult DRG expression of Rb1 mrna declined (*p
Supplementary Figures and Legends. Supplementary Figure 1: Impact of injury on Rb1 and PPARϒ expression. Following ipsilateral axotomy injury, adult DRG expression of Rb1 mrna declined (*p
Transcriptional regulation of BRCA1 expression by a metabolic switch: Di, Fernandez, De Siervi, Longo, and Gardner. H3K4Me3
 ChIP H3K4Me3 enrichment.25.2.15.1.5 H3K4Me3 H3K4Me3 ctrl H3K4Me3 + E2 NS + E2 1. kb kb +82 kb Figure S1. Estrogen promotes entry of MCF-7 into the cell cycle but does not significantly change activation-associated
ChIP H3K4Me3 enrichment.25.2.15.1.5 H3K4Me3 H3K4Me3 ctrl H3K4Me3 + E2 NS + E2 1. kb kb +82 kb Figure S1. Estrogen promotes entry of MCF-7 into the cell cycle but does not significantly change activation-associated
TRIM31 is recruited to mitochondria after infection with SeV.
 Supplementary Figure 1 TRIM31 is recruited to mitochondria after infection with SeV. (a) Confocal microscopy of TRIM31-GFP transfected into HEK293T cells for 24 h followed with SeV infection for 6 h. MitoTracker
Supplementary Figure 1 TRIM31 is recruited to mitochondria after infection with SeV. (a) Confocal microscopy of TRIM31-GFP transfected into HEK293T cells for 24 h followed with SeV infection for 6 h. MitoTracker
Nature Structural and Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Distribution of mirnas between lncrna and protein-coding genes. Pie chart showing distribution of human mirna between protein coding and lncrna genes. To the right, lncrna mirna
Supplementary Figure 1 Distribution of mirnas between lncrna and protein-coding genes. Pie chart showing distribution of human mirna between protein coding and lncrna genes. To the right, lncrna mirna
Sarker et al. Supplementary Material. Subcellular Fractionation
 Supplementary Material Subcellular Fractionation Transfected 293T cells were harvested with phosphate buffered saline (PBS) and centrifuged at 2000 rpm (500g) for 3 min. The pellet was washed, re-centrifuged
Supplementary Material Subcellular Fractionation Transfected 293T cells were harvested with phosphate buffered saline (PBS) and centrifuged at 2000 rpm (500g) for 3 min. The pellet was washed, re-centrifuged
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/6/304/ra104/dc1 Supplementary Materials for Lysine Methylation Promotes VEGFR-2 Activation and Angiogenesis Edward J. Hartsough, Rosana D. Meyer, Vipul Chitalia,
www.sciencesignaling.org/cgi/content/full/6/304/ra104/dc1 Supplementary Materials for Lysine Methylation Promotes VEGFR-2 Activation and Angiogenesis Edward J. Hartsough, Rosana D. Meyer, Vipul Chitalia,
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/10/496/eaam6291/dc1 Supplementary Materials for Regulation of autophagy, NF-κB signaling, and cell viability by mir-124 in KRAS mutant mesenchymal-like NSCLC cells
www.sciencesignaling.org/cgi/content/full/10/496/eaam6291/dc1 Supplementary Materials for Regulation of autophagy, NF-κB signaling, and cell viability by mir-124 in KRAS mutant mesenchymal-like NSCLC cells
Intestinal Epithelial Cell-Specific Deletion of PLD2 Alleviates DSS-Induced Colitis by. Regulating Occludin
 Intestinal Epithelial Cell-Specific Deletion of PLD2 Alleviates DSS-Induced Colitis by Regulating Occludin Chaithanya Chelakkot 1,ǂ, Jaewang Ghim 2,3,ǂ, Nirmal Rajasekaran 4, Jong-Sun Choi 5, Jung-Hwan
Intestinal Epithelial Cell-Specific Deletion of PLD2 Alleviates DSS-Induced Colitis by Regulating Occludin Chaithanya Chelakkot 1,ǂ, Jaewang Ghim 2,3,ǂ, Nirmal Rajasekaran 4, Jong-Sun Choi 5, Jung-Hwan
Figure S1. Optimization of reverse-transfection protocols using GAPDH sirnas. MDA- MB-231 cells were seeded in 96-well plates at a density of 5000
 Figure S1. Optimization of reverse-transfection protocols using GAPDH sirnas. MDA- MB-231 cells were seeded in 96-well plates at a density of 5000 cells per well, and reverse transfected with the indicated
Figure S1. Optimization of reverse-transfection protocols using GAPDH sirnas. MDA- MB-231 cells were seeded in 96-well plates at a density of 5000 cells per well, and reverse transfected with the indicated
Supplementary Figure Legends
 Supplementary Figure Legends Figure S1 gene targeting strategy for disruption of chicken gene, related to Figure 1 (f)-(i). (a) The locus and the targeting constructs showing HpaI restriction sites. The
Supplementary Figure Legends Figure S1 gene targeting strategy for disruption of chicken gene, related to Figure 1 (f)-(i). (a) The locus and the targeting constructs showing HpaI restriction sites. The
Supplemental Figure 1: Podocyte specific knockdown of Klf4 in Podocin-Cre Klf4 flox/flox mice was confirmed. (A) Primary glomerular epithelial cells
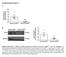 Supplemental Figure 1: Podocyte specific knockdown of Klf4 in Podocin-Cre Klf4 flox/flox mice was confirmed. (A) Primary glomerular epithelial cells were isolated from 10-week old Podocin-Cre Klf4 flox/flox
Supplemental Figure 1: Podocyte specific knockdown of Klf4 in Podocin-Cre Klf4 flox/flox mice was confirmed. (A) Primary glomerular epithelial cells were isolated from 10-week old Podocin-Cre Klf4 flox/flox
14_integrins_EGFR
 α1 Integrin α1 -/- fibroblasts from integrin α1 knockout animals Figure S1. Serum-starved fibroblasts from α -/- 1 and +/+ mice were stimulated with 10% FBS for 30 min. (FBS) or plated on collagen I (CI)
α1 Integrin α1 -/- fibroblasts from integrin α1 knockout animals Figure S1. Serum-starved fibroblasts from α -/- 1 and +/+ mice were stimulated with 10% FBS for 30 min. (FBS) or plated on collagen I (CI)
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/5/244/ra72/dc1 Supplementary Materials for An Interaction Between BZR1 and DELLAs Mediates Direct Signaling Crosstalk Between Brassinosteroids and Gibberellins
www.sciencesignaling.org/cgi/content/full/5/244/ra72/dc1 Supplementary Materials for An Interaction Between BZR1 and DELLAs Mediates Direct Signaling Crosstalk Between Brassinosteroids and Gibberellins
Regulation of axonal and dendritic growth by the extracellular calcium-sensing
 Regulation of axonal and dendritic growth by the extracellular calcium-sensing receptor (CaSR). Thomas N. Vizard, Gerard W. O Keeffe, Humberto Gutierrez, Claudine H. Kos, Daniela Riccardi, Alun M. Davies
Regulation of axonal and dendritic growth by the extracellular calcium-sensing receptor (CaSR). Thomas N. Vizard, Gerard W. O Keeffe, Humberto Gutierrez, Claudine H. Kos, Daniela Riccardi, Alun M. Davies
Supplementary Fig S1 Nutlin-3a treatment does not affect cell cycle progression in the absence
 Supplementary Figure Legends Supplementary Fig S1 Nutlin-3a treatment does not affect cell cycle progression in the absence of p53 or p21. HCT116 cells which were null for either p53 (A) or p21 (B) were
Supplementary Figure Legends Supplementary Fig S1 Nutlin-3a treatment does not affect cell cycle progression in the absence of p53 or p21. HCT116 cells which were null for either p53 (A) or p21 (B) were
Supplemental figures Supplemental Figure 1: Fluorescence recovery for FRAP experiments depicted in Figure 1.
 Supplemental figures Supplemental Figure 1: Fluorescence recovery for FRAP experiments depicted in Figure 1. Percent of original fluorescence was plotted as a function of time following photobleaching
Supplemental figures Supplemental Figure 1: Fluorescence recovery for FRAP experiments depicted in Figure 1. Percent of original fluorescence was plotted as a function of time following photobleaching
Figure S1- Targeting strategy for generating SIK1-T182A, SIK2-T175A and SIK3-T163A KI mice.
 Figure S1- Targeting strategy for generating SIK1-T12A, SIK2-T175A and SIK3-T163A mice. Left panel represents a depiction of the genomic loci of each gene, the targeting strategy and the final recombined
Figure S1- Targeting strategy for generating SIK1-T12A, SIK2-T175A and SIK3-T163A mice. Left panel represents a depiction of the genomic loci of each gene, the targeting strategy and the final recombined
Supplementary Figure Legends. Figure-S1 Molecular basis of ASS1 deficiency in myxofibrosarcoma: (A)
 Supplementary Figure Legends Figure-S1 Molecular basis of ASS1 deficiency in myxofibrosarcoma: (A) High-resolution oligonucleotide-based array comparative genomic hybridization shows normal DNA copy number
Supplementary Figure Legends Figure-S1 Molecular basis of ASS1 deficiency in myxofibrosarcoma: (A) High-resolution oligonucleotide-based array comparative genomic hybridization shows normal DNA copy number
Figure 1: TDP-43 is subject to lysine acetylation within the RNA-binding domain a) QBI-293 cells were transfected with TDP-43 in the presence or
 Figure 1: TDP-43 is subject to lysine acetylation within the RNA-binding domain a) QBI-293 cells were transfected with TDP-43 in the presence or absence of the acetyltransferase CBP and acetylated TDP-43
Figure 1: TDP-43 is subject to lysine acetylation within the RNA-binding domain a) QBI-293 cells were transfected with TDP-43 in the presence or absence of the acetyltransferase CBP and acetylated TDP-43
Transient silencing of the CDKN2A gene was achieved using a pool of four sirna duplexes
 Supplementary Methods Small interfering RNA (sirna) knockdown of CDKN2A (p16 INK4a /p14 ARF ) Transient silencing of the CDKN2A gene was achieved using a pool of four sirna duplexes that each targeted
Supplementary Methods Small interfering RNA (sirna) knockdown of CDKN2A (p16 INK4a /p14 ARF ) Transient silencing of the CDKN2A gene was achieved using a pool of four sirna duplexes that each targeted
Supplementary Information
 Supplementary Information Negative regulation of initial steps in skeletal myogenesis by mtor and other kinases Raphael A. Wilson 1*, Jing Liu 1*, Lin Xu 1, James Annis 2, Sara Helmig 1, Gregory Moore
Supplementary Information Negative regulation of initial steps in skeletal myogenesis by mtor and other kinases Raphael A. Wilson 1*, Jing Liu 1*, Lin Xu 1, James Annis 2, Sara Helmig 1, Gregory Moore
Supplementary Figure 1 a. d 0.8 CON LPS PAN. 2nd ab nephrin podocin CON LPS PAN. upar. -tubulin. upar. upar / -tubulin CON LPS PAN
 Supplementary Figure 1 a Efferent arteriole Podocytes Afferent arteriole FP Endothelium GBM Glomerular filtration barrier b 188kD HEK + GFP HEK + GFP-Nphs1 Differentiated Podocytes HEK + GFP HEK + GFP-Nphs2
Supplementary Figure 1 a Efferent arteriole Podocytes Afferent arteriole FP Endothelium GBM Glomerular filtration barrier b 188kD HEK + GFP HEK + GFP-Nphs1 Differentiated Podocytes HEK + GFP HEK + GFP-Nphs2
SUPPLEMENTARY MATERIALS
 Anti-SSTR2 peptide based targeted delivery of potent PLGA encapsulated 3,3 -diindolylmethane nanoparticles through blood brain barrier prevents glioma progression SUPPLEMENTARY MATERIALS Supplementary
Anti-SSTR2 peptide based targeted delivery of potent PLGA encapsulated 3,3 -diindolylmethane nanoparticles through blood brain barrier prevents glioma progression SUPPLEMENTARY MATERIALS Supplementary
Supplementary Data. Supplementary Materials and Methods Quantification of delivered sirnas. Fluorescence microscopy analysis
 Supplementary Data Supplementary Materials and Methods Quantification of delivered sirnas After transfection, cells were washed in three times with phosphate buffered saline and total RNA was extracted
Supplementary Data Supplementary Materials and Methods Quantification of delivered sirnas After transfection, cells were washed in three times with phosphate buffered saline and total RNA was extracted
hnrnp C promotes APP translation by competing with FMRP for APP mrna recruitment to P bodies
 hnrnp C promotes APP translation by competing with for APP mrna recruitment to P bodies Eun Kyung Lee 1, Hyeon Ho Kim 1, Yuki Kuwano 1, Kotb Abdelmohsen 1, Subramanya Srikantan 1, Sarah S. Subaran 2, Marc
hnrnp C promotes APP translation by competing with for APP mrna recruitment to P bodies Eun Kyung Lee 1, Hyeon Ho Kim 1, Yuki Kuwano 1, Kotb Abdelmohsen 1, Subramanya Srikantan 1, Sarah S. Subaran 2, Marc
Li et al., Supplemental Figures
 Li et al., Supplemental Figures Fig. S1. Suppressing TGM2 expression with TGM2 sirnas inhibits migration and invasion in A549-TR cells. A, A549-TR cells transfected with negative control sirna (NC sirna)
Li et al., Supplemental Figures Fig. S1. Suppressing TGM2 expression with TGM2 sirnas inhibits migration and invasion in A549-TR cells. A, A549-TR cells transfected with negative control sirna (NC sirna)
Supplementary Figure 1. Generation of B2M -/- ESCs. Nature Biotechnology: doi: /nbt.3860
 Supplementary Figure 1 Generation of B2M -/- ESCs. (a) Maps of the B2M alleles in cells with the indicated B2M genotypes. Probes and restriction enzymes used in Southern blots are indicated (H, Hind III;
Supplementary Figure 1 Generation of B2M -/- ESCs. (a) Maps of the B2M alleles in cells with the indicated B2M genotypes. Probes and restriction enzymes used in Southern blots are indicated (H, Hind III;
