SUPPLEMENTARY INFORMATION
|
|
|
- Leslie Piers Neal
- 6 years ago
- Views:
Transcription
1 A diphtheria toxin resistance marker for in vitro and in vivo selection of stably transduced human cells Gabriele Picco, Consalvo Petti, Livio Trusolino, Andrea Bertotti and Enzo Medico SUPPLEMENTARY INFORMATION
2
3
4 CRC cell lines (n=151) CRC PDX (n=515) DPH2, mrna expression (log2) TCGA_Colon (n=365) TCGA_H&N (n=498) TCGA_Glioblastoma (n=166) TCGA_Pancreas (n=179) HBEGF, mrna expression (log2) Picco et al., Supplementary Figure 1. HBEGF and DPH2 are well expressed in CRC cell lines, PDXs and tumors, and in other tumor types. Dot plots displaying for each sample, on the x-axis expression of HBEGF, and on the y-axis expression of DPH2. Each dot plot is annotated for the dataset from which expression levels have been obtained.
5 a HBEGF expression ME:MALME3M LC:NCIH460 CNS:SNB19 CO:HCT116 LC:NCI-H23 ME:SKMEL2 OV:OVCAR4 OV:OVCAR8 CNS:SF268 CNS:SF539 HBEGF Z-score b Diphtheria toxin (ng/ml) Picco et al., Supplementary Figure 2. HBEGF expression does not affect sensitivity to DT in human cancer cell lines. (a) Histogram representing HBEGF mrna expression (Z-score) in the NCI60 panel of human cancer cell lines (data from the cbbioportal: Red bars point out the ten cell lines from different tissues selected for DT treatment. (b) Crystal violet staining of selected cell lines, grown for one week in the presence of variable DT concentrations. Cells are ordered by increasing HBEGF expression, from left to right, as indicated by the red wedge on top.
6 Picco et al., Supplementary Figure 3. DPH2 silencing by DT R does not affect basal growth rate of cancer cell lines. Histograms representing the growth of OVCAR4 (ovary), SNB19 (glioblastoma), HCT116 (colon) and A549 (lung) cancer cell lines transduced with DT R or scramble construct. The growth rates on the y- axes were calculated by comparing ATP-based viability measurements at each time point vs. the measurements at plating.
7 Picco et al., Supplementary Figure 4. DT R is an efficient selectable marker in vitro. HCT116 cells were infected with DT R at low MOI (~0.02), to ensure transduction of a low fraction of cells with a single copy of the vector. Three days after infection, only ~2% of the cells expressed GFP. Cells were then incubated with puromycin (2 ng/ml) or DT (10 ng/ml) for two weeks. Subsequently, selection was released for four weeks to assess stability of the GFP+ fraction. Histograms represent the distribution of GFP signal: (i) in unselected transduced cells, (ii) after puromycin/dt selection, and (iii) after release from puromycin/dt selection, as indicated.
8 Picco et al., Supplementary Figure 5. Stability of DT resistance and GFP expression after tumor propagation. (a) Scramble and DT R transduced/selected HCT116 xenografts (n = 2 per group) were re-implanted in the right flank of CD1-nude mice. When tumors reached approximately a volume of 250 mm 3, tumour growth was monitored for three weeks. At day 24, DT (5 µg/kg) was administered to mice. DT R tumors (green line) continued growing in presence of DT, while scramble tumors (black line) rapidly reduced their volume. Flow cytometry analysis (b) and fluorescence micrograph (c) of the DT R xenograft explanted at day 24 (before the new DT selection) revealed a very high fraction of GFP+ cells.
9 a b GFP Signal (RE) c Days d GFP Signal (RE) Volume (mm 3 ) e Days f Days g GFP signal (RE) after explant h Tumor weight (mg) i Tumor weight (mg) GFP signal (RE) before explant GFP signal (RE) before explant GFP signal (RE) after explant Picco et al., Supplementary Figure 6. Live imaging of DT R -transduced HCT116 xenografts growing subcutaneously and intraperitoneally. (a) HCT116 xenografts transduced in vivo with DT R - (GFP+) were propagated subcutaneously in CD1 nude mice. (b) Scatter plot representing the correlation between caliper measurements and GFP fluorescence signal (Radiant efficiency, RE) detected by live imaging in subcutaneous xenografts. (c) Growth of DT R -transduced HCT116 xenografts (GFP+) propagated in the intraperitoneal (IP) cavity of CD1 nude mice. (d) Line chart reporting the GFP signal measured by live imaging (IVIS-Caliper) as a surrogate maker of tumor growth. (e) GFP signal of four DT R IP xenografts. (f) GFP expression of the tumours after explant, imaged by IVIS. (g-h) Scatter plots comparing GFP signal before explant with GFP signal after explant or with tumor weight after explant. (i) Comparison of GFP signal and tumor weight, both measured after explant.
10 a Tumor volume (mm 3 ) b Days Tumor volume (mm 3 ) c Days Tumor volume (mm 3 ) Days Picco et al., Supplementary Figure 7. Colorectal cancer PDXs are sensitive to DT. Patient-derived xenografts from three metastatic CRC specimens, with different KRAS/BRAF genetic backgound were grown in NOD/SCID mice. The three graphs represent tumor growth inhibition of xenografts derived from: (a) KRAS mutated (G13D), (b) BRAF mutated (V600E) or (c) KRAS/BRAF WT CRC tumors after administration of DT (10 µg/kg) for three weeks.
11 a Parental PDX DTR-transduced and selected PDX b Parental PDX DTR-transduced and selected PDX Picco et al., Supplementary Figure 8. DTR-transduced PDXs retain histopathologic and functional characteristics of the original sample. (a) Haematoxylin and eosin staining of parental and DTR-transduced PDX. (b) Ki67 expression in parental and DTR-transduced PDX.
12 CDX2 Parental PDX DTR-transduced and DT-selected PDX CK20 β-catenin Picco et al., Supplementary Figure 9. DTR transduction and DT selection do not significantly alter the PDX phenotype. CDX2, CK20 and β-catenin expression in a CRC PDX before (left panels) and after (right panels) DTR transduction and DT selection.
13 a b 92% HLA-APC c GFP Picco et al., Supplementary Figure 10. Stability of GFP expression after tumor propagation. After DT R transduction and DT selection, a CRC PDX was propagated for four passages in NOD-SCID mice. (a) Live imaging highlighting the GFP-positive tumor mass (white arrow). (b) Flow cytometry analysis of cells from the P4 tumor explant, displaying GFP signal on the x-axis and the HLA-APC human marker on the y-axis. (c) fluorescence microscopy displaying the GFP signal (green) in cancer cells vs. the nuclear DAPI signal (blue) in all cells.
14 a b c Pearson DPH2 expression (percent) Parental vs DT R s DT R s vs DT R s Parental P1 P2 P4 P0 DT R Picco et al., Supplementary Figure 11. DT R trasduction and DT selection do not alter global gene expression of CRC PDX. (a) Downregulation, respect to the parental PDX, of the DPH2 transcript in all the DT R derivatives passaged in the absence of DT. (b) Pearson correlation values based on global expression profiles, comparing parental PDX with DT R derivatives (blue diamonds) or DT R derivatives with each other (red diamonds). (c) hierarchical clustering based on global gene expression profiling of a parental PDX and its DT R derivatives at different passages in mice.
Supplementary Fig. 1 related to Fig. 1 Clinical relevance of lncrna candidate
 Supplementary Figure Legends Supplementary Fig. 1 related to Fig. 1 Clinical relevance of lncrna candidate BC041951 in gastric cancer. (A) The flow chart for selected candidate lncrnas in 660 up-regulated
Supplementary Figure Legends Supplementary Fig. 1 related to Fig. 1 Clinical relevance of lncrna candidate BC041951 in gastric cancer. (A) The flow chart for selected candidate lncrnas in 660 up-regulated
Supplementary Figure 1. Isolation of GFPHigh cells.
 Supplementary Figure 1. Isolation of GFP High cells. (A) Schematic diagram of cell isolation based on Wnt signaling activity. Colorectal cancer (CRC) cell lines were stably transduced with lentivirus encoding
Supplementary Figure 1. Isolation of GFP High cells. (A) Schematic diagram of cell isolation based on Wnt signaling activity. Colorectal cancer (CRC) cell lines were stably transduced with lentivirus encoding
Supplementary Figure 1
 Supplementary Figure 1 Supplementary Figure 1: Vector maps of TRMPV and TRMPVIR variants. Many derivatives of TRMPV have been generated and tested. Unless otherwise noted, experiments in this paper use
Supplementary Figure 1 Supplementary Figure 1: Vector maps of TRMPV and TRMPVIR variants. Many derivatives of TRMPV have been generated and tested. Unless otherwise noted, experiments in this paper use
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1.
 Supplementary Figure 1. Characterization and high expression of Lnc-β-Catm in liver CSCs. (a) Heatmap of differently expressed lncrnas in Liver CSCs (CD13 + CD133 + ) and non-cscs (CD13 - CD133 - ) according
Supplementary Figure 1. Characterization and high expression of Lnc-β-Catm in liver CSCs. (a) Heatmap of differently expressed lncrnas in Liver CSCs (CD13 + CD133 + ) and non-cscs (CD13 - CD133 - ) according
TF-1a lymphoblastic leukemia cell line: marking with GFP, phenotyping and sorting
 Supplemental Material Supplemental Methods TF-1a lymphoblastic leukemia cell line: marking with GFP, phenotyping and sorting In order to determine if the multi-parameter FACS approach would be successful
Supplemental Material Supplemental Methods TF-1a lymphoblastic leukemia cell line: marking with GFP, phenotyping and sorting In order to determine if the multi-parameter FACS approach would be successful
Nature Methods doi: /nmeth Supplementary Figure 1. Screening for cancer stem cell marker(s)
 Supplementary Figure 1 Screening for cancer stem cell marker(s) (a) Flow cytometry analysis of CD133 at increasing times to analysis. Cells were trypsinized, stained, transferred to RPMI medium and then
Supplementary Figure 1 Screening for cancer stem cell marker(s) (a) Flow cytometry analysis of CD133 at increasing times to analysis. Cells were trypsinized, stained, transferred to RPMI medium and then
-Immune phenotype -Cancer xenografts -Immune system humanization -Services
 -Immune phenotype -Cancer xenografts -Immune system humanization -Services services@herabiolabs.com 859-414-0648 About Hera BioLabs Precision Toxicology & Efficacy: utilizing precisely gene-edited models
-Immune phenotype -Cancer xenografts -Immune system humanization -Services services@herabiolabs.com 859-414-0648 About Hera BioLabs Precision Toxicology & Efficacy: utilizing precisely gene-edited models
Supplementary Figure 1
 number of cells, normalized number of cells, normalized number of cells, normalized Supplementary Figure CD CD53 Cd3e fluorescence intensity fluorescence intensity fluorescence intensity Supplementary
number of cells, normalized number of cells, normalized number of cells, normalized Supplementary Figure CD CD53 Cd3e fluorescence intensity fluorescence intensity fluorescence intensity Supplementary
Supporting Information for. Bongseo Choi, 1, Hyojin Moon, 1, Sung Joon Hong, 1 Changsik Shin, 1 Yoonkyung Do, 1 Seongho Ryu, 2,* Sebyung Kang 1,*
 Supporting Information for Effective Delivery of Antigen-Encapsulin Nanoparticle Fusions to Dendritic Cells Leads to Antigen-Specific Cytotoxic T Cell Activation and Tumor Rejection Bongseo Choi, 1, Hyojin
Supporting Information for Effective Delivery of Antigen-Encapsulin Nanoparticle Fusions to Dendritic Cells Leads to Antigen-Specific Cytotoxic T Cell Activation and Tumor Rejection Bongseo Choi, 1, Hyojin
Gene-Level Analysis of Exon Array Data using Partek Genomics Suite 6.6
 Gene-Level Analysis of Exon Array Data using Partek Genomics Suite 6.6 Overview This tutorial will demonstrate how to: Summarize core exon-level data to produce gene-level data Perform exploratory analysis
Gene-Level Analysis of Exon Array Data using Partek Genomics Suite 6.6 Overview This tutorial will demonstrate how to: Summarize core exon-level data to produce gene-level data Perform exploratory analysis
SUPPLEMENTARY INFORMATION
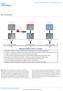 DOI: 10.1038/ncb2239 Hepatocytes (Endoderm) Foreskin fibroblasts (Mesoderm) Melanocytes (Ectoderm) +Dox rtta TetO CMV O,S,M,K & N H1 hes H7 hes H9 hes H1 hes-derived EBs H7 hes-derived EBs H9 hes-derived
DOI: 10.1038/ncb2239 Hepatocytes (Endoderm) Foreskin fibroblasts (Mesoderm) Melanocytes (Ectoderm) +Dox rtta TetO CMV O,S,M,K & N H1 hes H7 hes H9 hes H1 hes-derived EBs H7 hes-derived EBs H9 hes-derived
Supplementary Figures
 Supplementary Figures Supplementary Figure 1. Description of the observed lymphatic metastases in two different SIX1-induced MCF7 metastasis models (Nude and NOD/SCID). Supplementary Figure 2. MCF7-SIX1
Supplementary Figures Supplementary Figure 1. Description of the observed lymphatic metastases in two different SIX1-induced MCF7 metastasis models (Nude and NOD/SCID). Supplementary Figure 2. MCF7-SIX1
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb2774 Figure S1 TRF2 dosage modulates the tumorigenicity of mouse and human tumor cells. (a) Left: immunoblotting with antibodies directed against the Myc tag of the transduced TRF2 forms
DOI: 10.1038/ncb2774 Figure S1 TRF2 dosage modulates the tumorigenicity of mouse and human tumor cells. (a) Left: immunoblotting with antibodies directed against the Myc tag of the transduced TRF2 forms
Bioware Brite Cell Line HepG2-Red-FLuc
 TECHNICAL DATA SHEET Bioware Brite Cell Line Research Use Only. Not for use in diagnostic procedures. Bioware Brite Cell Line HepG2-Red-FLuc Product No.: BW134280 Material Provided Cells: Format: 2 x 1
TECHNICAL DATA SHEET Bioware Brite Cell Line Research Use Only. Not for use in diagnostic procedures. Bioware Brite Cell Line HepG2-Red-FLuc Product No.: BW134280 Material Provided Cells: Format: 2 x 1
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION Legends for Supplementary Tables. Supplementary Table 1. An excel file containing primary screen data. Worksheet 1, Normalized quantification data from a duplicated screen: valid
SUPPLEMENTARY INFORMATION Legends for Supplementary Tables. Supplementary Table 1. An excel file containing primary screen data. Worksheet 1, Normalized quantification data from a duplicated screen: valid
Multiplex Fluorescence Assays for Adherence Cells without Trypsinization
 Multiplex Fluorescence Assays for Adherence Cells without Trypsinization The combination of a bright field and three fluorescent channels allows the Celigo to perform many multiplexed assays. A gating
Multiplex Fluorescence Assays for Adherence Cells without Trypsinization The combination of a bright field and three fluorescent channels allows the Celigo to perform many multiplexed assays. A gating
Nature Genetics: doi: /ng Supplementary Figure 1
 Supplementary Figure 1 Processing of mutations and generation of simulated controls. On the left, a diagram illustrates the manner in which covariate-matched simulated mutations were obtained, filtered
Supplementary Figure 1 Processing of mutations and generation of simulated controls. On the left, a diagram illustrates the manner in which covariate-matched simulated mutations were obtained, filtered
DEPArray Technology. Sorting and Recovery of Rare Cells
 DEPArray Technology Sorting and Recovery of Rare Cells Delivering pure, single, viable cells The DEPArray system from Silicon Biosystems is the only automated instrument that can identify, quantify, and
DEPArray Technology Sorting and Recovery of Rare Cells Delivering pure, single, viable cells The DEPArray system from Silicon Biosystems is the only automated instrument that can identify, quantify, and
Nature Immunology: doi: /ni Supplementary Figure 1
 Supplementary Figure 1 BALB/c LYVE1-deficient mice exhibited reduced lymphatic trafficking of all DC subsets after oxazolone-induced sensitization. (a) Schematic overview of the mouse skin oxazolone contact
Supplementary Figure 1 BALB/c LYVE1-deficient mice exhibited reduced lymphatic trafficking of all DC subsets after oxazolone-induced sensitization. (a) Schematic overview of the mouse skin oxazolone contact
Gaussia Luciferase-a Novel Bioluminescent Reporter for Tracking Stem Cells Survival, Proliferation and Differentiation in Vivo
 Gaussia Luciferase-a Novel Bioluminescent Reporter for Tracking Stem Cells Survival, Proliferation and Differentiation in Vivo Rampyari Raja Walia and Bakhos A. Tannous 1 2 1 Pluristem Innovations, 1453
Gaussia Luciferase-a Novel Bioluminescent Reporter for Tracking Stem Cells Survival, Proliferation and Differentiation in Vivo Rampyari Raja Walia and Bakhos A. Tannous 1 2 1 Pluristem Innovations, 1453
supplementary information
 DOI: 1.138/ncb1839 a b Control 1 2 3 Control 1 2 3 Fbw7 Smad3 1 2 3 4 1 2 3 4 c d IGF-1 IGF-1Rβ IGF-1Rβ-P Control / 1 2 3 4 Real-time RT-PCR Relative quantity (IGF-1/ mrna) 2 1 IGF-1 1 2 3 4 Control /
DOI: 1.138/ncb1839 a b Control 1 2 3 Control 1 2 3 Fbw7 Smad3 1 2 3 4 1 2 3 4 c d IGF-1 IGF-1Rβ IGF-1Rβ-P Control / 1 2 3 4 Real-time RT-PCR Relative quantity (IGF-1/ mrna) 2 1 IGF-1 1 2 3 4 Control /
a Award Number: W81XWH TITLE:
 AD Award Number: W81XWH-04-1-0138 TITLE: Imaging Metastatic Prostate Cancer After Genetic Manipulation of Transcriptional Memory Regulators EZH2 and EED PRINCIPAL INVESTIGATOR: Lily Wu, M.D., Ph.D. CONTRACTING
AD Award Number: W81XWH-04-1-0138 TITLE: Imaging Metastatic Prostate Cancer After Genetic Manipulation of Transcriptional Memory Regulators EZH2 and EED PRINCIPAL INVESTIGATOR: Lily Wu, M.D., Ph.D. CONTRACTING
Figure S1. Mechanism of ON induced MM cell death.
 Figure S1. Mechanism of ON123300 induced MM cell death. To elucidate whether ON123300 induced decrease of viability of MM cells was associated with induction of apoptosis, the number of apoptotic cells
Figure S1. Mechanism of ON123300 induced MM cell death. To elucidate whether ON123300 induced decrease of viability of MM cells was associated with induction of apoptosis, the number of apoptotic cells
Supplementary Figure 1: sgrna library generation and the length of sgrnas for the functional screen. (a) A diagram of the retroviral vector for sgrna
 Supplementary Figure 1: sgrna library generation and the length of sgrnas for the functional screen. (a) A diagram of the retroviral vector for sgrna expression. It contains a U6-promoter-driven sgrna
Supplementary Figure 1: sgrna library generation and the length of sgrnas for the functional screen. (a) A diagram of the retroviral vector for sgrna expression. It contains a U6-promoter-driven sgrna
Figure S1. Figure S2. Figure S3 HB Anti-FSP27 (COOH-terminal peptide) Ab. Anti-GST-FSP27(45-127) Ab.
 / 36B4 mrna ratio Figure S1 * 2. 1.6 1.2.8 *.4 control TNFα BRL49653 Figure S2 Su bw AT p iw Anti- (COOH-terminal peptide) Ab Blot : Anti-GST-(45-127) Ab β-actin Figure S3 HB2 HW AT BA T Figure S4 A TAG
/ 36B4 mrna ratio Figure S1 * 2. 1.6 1.2.8 *.4 control TNFα BRL49653 Figure S2 Su bw AT p iw Anti- (COOH-terminal peptide) Ab Blot : Anti-GST-(45-127) Ab β-actin Figure S3 HB2 HW AT BA T Figure S4 A TAG
Development of a Tissue Slice Culture Model for Prostate Cancer
 Development of a Tissue Slice Culture Model for Prostate Cancer Hanneke van Zoggel, PhD Experimental Urology Erasmus MC ; JNI, Be 330 j.vanzoggel@erasmusmc.nl 3D workshop 23/24-09-2013 Create novel models
Development of a Tissue Slice Culture Model for Prostate Cancer Hanneke van Zoggel, PhD Experimental Urology Erasmus MC ; JNI, Be 330 j.vanzoggel@erasmusmc.nl 3D workshop 23/24-09-2013 Create novel models
Supplemental Information. The TRAIL-Induced Cancer Secretome. Promotes a Tumor-Supportive Immune. Microenvironment via CCR2
 Molecular Cell, Volume 65 Supplemental Information The TRAIL-Induced Cancer Secretome Promotes a Tumor-Supportive Immune Microenvironment via CCR2 Torsten Hartwig, Antonella Montinaro, Silvia von Karstedt,
Molecular Cell, Volume 65 Supplemental Information The TRAIL-Induced Cancer Secretome Promotes a Tumor-Supportive Immune Microenvironment via CCR2 Torsten Hartwig, Antonella Montinaro, Silvia von Karstedt,
A Level. A Level Biology. DNA Technology Questions. AQA, OCR, Edexcel. Name: Total Marks: Page 1
 AQA, OCR, Edexcel A Level A Level Biology DNA Technology Questions Name: Total Marks: Page 1 Q1.(a) (i) A mutation of a tumour suppressor gene can result in the formation of a tumour. Explain how.........(2)
AQA, OCR, Edexcel A Level A Level Biology DNA Technology Questions Name: Total Marks: Page 1 Q1.(a) (i) A mutation of a tumour suppressor gene can result in the formation of a tumour. Explain how.........(2)
SUPPLEMENTAL FIGURE LEGENDS. Figure S1: Homology alignment of DDR2 amino acid sequence. Shown are
 SUPPLEMENTAL FIGURE LEGENDS Figure S1: Homology alignment of DDR2 amino acid sequence. Shown are the amino acid sequences of human DDR2, mouse DDR2 and the closest homologs in zebrafish and C. Elegans.
SUPPLEMENTAL FIGURE LEGENDS Figure S1: Homology alignment of DDR2 amino acid sequence. Shown are the amino acid sequences of human DDR2, mouse DDR2 and the closest homologs in zebrafish and C. Elegans.
Label-free, real-time live-cell assays for spheroids: IncuCyte bright-field analysis
 Introduction APPLICATION NOTE IncuCyte Live-Cell Analysis System Label-free, real-time live-cell assays for spheroids: IncuCyte bright-field analysis Susana L. Alcantara, Miniver Oliver, Kalpana Patel,
Introduction APPLICATION NOTE IncuCyte Live-Cell Analysis System Label-free, real-time live-cell assays for spheroids: IncuCyte bright-field analysis Susana L. Alcantara, Miniver Oliver, Kalpana Patel,
Figure S2. Response of mouse ES cells to GSK3 inhibition. Mentioned in discussion
 Stem Cell Reports, Volume 1 Supplemental Information Robust Self-Renewal of Rat Embryonic Stem Cells Requires Fine-Tuning of Glycogen Synthase Kinase-3 Inhibition Yaoyao Chen, Kathryn Blair, and Austin
Stem Cell Reports, Volume 1 Supplemental Information Robust Self-Renewal of Rat Embryonic Stem Cells Requires Fine-Tuning of Glycogen Synthase Kinase-3 Inhibition Yaoyao Chen, Kathryn Blair, and Austin
Pre-made Lentiviral Particles for Nuclear Permeant CRE Recombinase Expression
 Pre-made Lentiviral Particles for Nuclear Permeant CRE Recombinase Expression LVP336 LVP336-PBS LVP339 LVP339-PBS LVP297 LVP297-PBS LVP013 LVP013-PBS LVP338 LVP338-PBS LVP027 LVP027-PBS LVP337 LVP337-PBS
Pre-made Lentiviral Particles for Nuclear Permeant CRE Recombinase Expression LVP336 LVP336-PBS LVP339 LVP339-PBS LVP297 LVP297-PBS LVP013 LVP013-PBS LVP338 LVP338-PBS LVP027 LVP027-PBS LVP337 LVP337-PBS
Tumor Growth Suppression Through the Activation of p21, a Cyclin-Dependent Kinase Inhibitor
 Tumor Growth Suppression Through the Activation of p21, a Cyclin-Dependent Kinase Inhibitor Nicholas Love 11/28/01 A. What is p21? Introduction - p21 is a gene found on chromosome 6 at 6p21.2 - this gene
Tumor Growth Suppression Through the Activation of p21, a Cyclin-Dependent Kinase Inhibitor Nicholas Love 11/28/01 A. What is p21? Introduction - p21 is a gene found on chromosome 6 at 6p21.2 - this gene
Wnt16 smact merge VK/AB
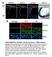 A WT Wnt6 smact merge VK/A KO ctrl IgG WT KO Wnt6 smact DAPI SUPPLEMENTAL FIGURE I: Wnt6 expression in MGP-deficient aortae. Immunostaining for Wnt6 and smooth muscle actin (smact) in aortae from 7 day
A WT Wnt6 smact merge VK/A KO ctrl IgG WT KO Wnt6 smact DAPI SUPPLEMENTAL FIGURE I: Wnt6 expression in MGP-deficient aortae. Immunostaining for Wnt6 and smooth muscle actin (smact) in aortae from 7 day
To isolate single GNS 144 cell clones, cells were plated at a density of 1cell/well
 Supplemental Information: Supplemental Methods: Cell culture To isolate single GNS 144 cell clones, cells were plated at a density of 1cell/well in 96 well Primaria plates in GNS media and incubated at
Supplemental Information: Supplemental Methods: Cell culture To isolate single GNS 144 cell clones, cells were plated at a density of 1cell/well in 96 well Primaria plates in GNS media and incubated at
Supplementary Materials and Methods:
 Supplementary Materials and Methods: Preclinical chemoprevention experimental design Mice were weighed and each mammary tumor was manually palpated and measured with digital calipers once a week from 4
Supplementary Materials and Methods: Preclinical chemoprevention experimental design Mice were weighed and each mammary tumor was manually palpated and measured with digital calipers once a week from 4
CQAs for C> Products to Enable Comparability Assessment. Ben Thompson Snr Director, Biopharmaceutical CMC RA GlaxoSmithKline
 CQAs for C> Products to Enable Comparability Assessment Ben Thompson Snr Director, Biopharmaceutical CMC RA GlaxoSmithKline Overview Demonstrate the value of defining CQAs early in product development
CQAs for C> Products to Enable Comparability Assessment Ben Thompson Snr Director, Biopharmaceutical CMC RA GlaxoSmithKline Overview Demonstrate the value of defining CQAs early in product development
Supplementary Fig. S1. SAMHD1c has a more potent dntpase activity than. SAMHD1c. Purified recombinant SAMHD1c and SAMHD1c proteins (with
 Supplementary Fig. S1. SAMHD1c has a more potent dntpase activity than SAMHD1c. Purified recombinant SAMHD1c and SAMHD1c proteins (with concentration of 800nM) were incubated with 1mM dgtp for the indicated
Supplementary Fig. S1. SAMHD1c has a more potent dntpase activity than SAMHD1c. Purified recombinant SAMHD1c and SAMHD1c proteins (with concentration of 800nM) were incubated with 1mM dgtp for the indicated
Supplemental Information Inventory
 Cell Stem Cell, Volume 6 Supplemental Information Distinct Hematopoietic Stem Cell Subtypes Are Differentially Regulated by TGF-β1 Grant A. Challen, Nathan C. Boles, Stuart M. Chambers, and Margaret A.
Cell Stem Cell, Volume 6 Supplemental Information Distinct Hematopoietic Stem Cell Subtypes Are Differentially Regulated by TGF-β1 Grant A. Challen, Nathan C. Boles, Stuart M. Chambers, and Margaret A.
Supporting Information
 Supporting Information Chakrabarty et al. 10.1073/pnas.1018001108 SI Materials and Methods Cell Lines. All cell lines were purchased from the American Type Culture Collection. Media and FBS were purchased
Supporting Information Chakrabarty et al. 10.1073/pnas.1018001108 SI Materials and Methods Cell Lines. All cell lines were purchased from the American Type Culture Collection. Media and FBS were purchased
Supplemental Table 1 Gene Symbol FDR corrected p-value PLOD1 CSRP2 PFKP ADFP ADM C10orf10 GPI LOX PLEKHA2 WIPF1
 Supplemental Table 1 Gene Symbol FDR corrected p-value PLOD1 4.52E-18 PDK1 6.77E-18 CSRP2 4.42E-17 PFKP 1.23E-14 MSH2 3.79E-13 NARF_A 5.56E-13 ADFP 5.56E-13 FAM13A1 1.56E-12 FAM29A_A 1.22E-11 CA9 1.54E-11
Supplemental Table 1 Gene Symbol FDR corrected p-value PLOD1 4.52E-18 PDK1 6.77E-18 CSRP2 4.42E-17 PFKP 1.23E-14 MSH2 3.79E-13 NARF_A 5.56E-13 ADFP 5.56E-13 FAM13A1 1.56E-12 FAM29A_A 1.22E-11 CA9 1.54E-11
Supplementary Information Design of small molecule-responsive micrornas based on structural requirements for Drosha processing
 Supplementary Information Design of small molecule-responsive micrornas based on structural requirements for Drosha processing Chase L. Beisel, Yvonne Y. Chen, Stephanie J. Culler, Kevin G. Hoff, & Christina
Supplementary Information Design of small molecule-responsive micrornas based on structural requirements for Drosha processing Chase L. Beisel, Yvonne Y. Chen, Stephanie J. Culler, Kevin G. Hoff, & Christina
TOOLS sirna and mirna. User guide
 TOOLS sirna and mirna User guide Introduction RNA interference (RNAi) is a powerful tool for suppression gene expression by causing the destruction of specific mrna molecules. Small Interfering RNAs (sirnas)
TOOLS sirna and mirna User guide Introduction RNA interference (RNAi) is a powerful tool for suppression gene expression by causing the destruction of specific mrna molecules. Small Interfering RNAs (sirnas)
Supplementary Figure S1. Immunodetection of full-length XA21 and the XA21 C-terminal cleavage product.
 Supplementary Information Supplementary Figure S1. Immunodetection of full-length XA21 and the XA21 C-terminal cleavage product. Total protein extracted from Kitaake wild type and rice plants carrying
Supplementary Information Supplementary Figure S1. Immunodetection of full-length XA21 and the XA21 C-terminal cleavage product. Total protein extracted from Kitaake wild type and rice plants carrying
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:10.1038/nature09937 a Name Position Primersets 1a 1b 2 3 4 b2 Phenotype Genotype b Primerset 1a D T C R I E 10000 8000 6000 5000 4000 3000 2500 2000 1500 1000 800 Donor (D)
SUPPLEMENTARY INFORMATION doi:10.1038/nature09937 a Name Position Primersets 1a 1b 2 3 4 b2 Phenotype Genotype b Primerset 1a D T C R I E 10000 8000 6000 5000 4000 3000 2500 2000 1500 1000 800 Donor (D)
SUPPLEMENTARY INFORMATION
 a before amputation regeneration regenerated limb DERMIS SKELETON MUSCLE SCHWANN CELLS EPIDERMIS DERMIS SKELETON MUSCLE SCHWANN CELLS EPIDERMIS developmental origin: lateral plate mesoderm presomitic mesoderm
a before amputation regeneration regenerated limb DERMIS SKELETON MUSCLE SCHWANN CELLS EPIDERMIS DERMIS SKELETON MUSCLE SCHWANN CELLS EPIDERMIS developmental origin: lateral plate mesoderm presomitic mesoderm
Nature Immunology: doi: /ni Supplementary Figure 1
 Supplementary Figure 1 Validation of the monoclonal antibody to mouse ACKR1 and expression of ACKR1 by BM hematopoietic cells. (a to d) Comparison of immunostaining of BM cells by anti-mouse ACKR1 antibodies:
Supplementary Figure 1 Validation of the monoclonal antibody to mouse ACKR1 and expression of ACKR1 by BM hematopoietic cells. (a to d) Comparison of immunostaining of BM cells by anti-mouse ACKR1 antibodies:
Supporting Information: Core-Shell Nanoparticle-Based Peptide Therapeutics and Combined. Hyperthermia for Enhanced Cancer Cell Apoptosis
 Supporting Information: Core-Shell Nanoparticle-Based Peptide Therapeutics and Combined Hyperthermia for Enhanced Cancer Cell Apoptosis Birju P. Shah a, Nicholas Pasquale a, Gejing De b, Tao Tan b, Jianjie
Supporting Information: Core-Shell Nanoparticle-Based Peptide Therapeutics and Combined Hyperthermia for Enhanced Cancer Cell Apoptosis Birju P. Shah a, Nicholas Pasquale a, Gejing De b, Tao Tan b, Jianjie
Supporting Information
 Supporting Information Table S1. Overview of samples used for sequencing, and the number of sequences obtained from each sample. Visit 1 is day 0, Visit 2 is day 7, Visit 3 is day 28, and Visit 4 is day
Supporting Information Table S1. Overview of samples used for sequencing, and the number of sequences obtained from each sample. Visit 1 is day 0, Visit 2 is day 7, Visit 3 is day 28, and Visit 4 is day
IKK is a therapeutic target in KRAS-induced lung cancer with disrupted p53 activity
 IKK is a therapeutic target in KRAS-induced lung cancer with disrupted p5 activity H6 5 5 H58 A59 H6 H58 A59 anti-ikkα anti-ikkβ anti-panras anti-gapdh anti-ikkα anti-ikkβ anti-panras anti-gapdh anti-ikkα
IKK is a therapeutic target in KRAS-induced lung cancer with disrupted p5 activity H6 5 5 H58 A59 H6 H58 A59 anti-ikkα anti-ikkβ anti-panras anti-gapdh anti-ikkα anti-ikkβ anti-panras anti-gapdh anti-ikkα
Supplementary Figure legends
 Supplementary Figure legends Supplementary Figure S1. Genome wide shrna screen. A, Screen schematic. The OpenBiosystems GIPZ lentiviral shrna library was divided into six pools, each encompassing 9600
Supplementary Figure legends Supplementary Figure S1. Genome wide shrna screen. A, Screen schematic. The OpenBiosystems GIPZ lentiviral shrna library was divided into six pools, each encompassing 9600
Genetics - Problem Drill 19: Dissection of Gene Function: Mutational Analysis of Model Organisms
 Genetics - Problem Drill 19: Dissection of Gene Function: Mutational Analysis of Model Organisms No. 1 of 10 1. The mouse gene knockout is based on. (A) Homologous recombination (B) Site-specific recombination
Genetics - Problem Drill 19: Dissection of Gene Function: Mutational Analysis of Model Organisms No. 1 of 10 1. The mouse gene knockout is based on. (A) Homologous recombination (B) Site-specific recombination
efluor Organic Dyes 450/50 BP Fluorescence Intensity Wavelength (nm)
 efluor Organic Dyes efluor Organic Dyes Catalog No. Description Clone Application Anti-Mouse efluor 450 Products 48-0042 Anti-mouse CD4 RM4-5 Flow Cytometry 48-0081 Anti-mouse CD8 53-6.7 Flow Cytometry
efluor Organic Dyes efluor Organic Dyes Catalog No. Description Clone Application Anti-Mouse efluor 450 Products 48-0042 Anti-mouse CD4 RM4-5 Flow Cytometry 48-0081 Anti-mouse CD8 53-6.7 Flow Cytometry
SUPPLEMENTARY INFORMATION
 doi:.38/nature899 Supplementary Figure Suzuki et al. a c p7 -/- / WT ratio (+)/(-) p7 -/- / WT ratio Log X 3. Fold change by treatment ( (+)/(-)) Log X.5 3-3. -. b Fold change by treatment ( (+)/(-)) 8
doi:.38/nature899 Supplementary Figure Suzuki et al. a c p7 -/- / WT ratio (+)/(-) p7 -/- / WT ratio Log X 3. Fold change by treatment ( (+)/(-)) Log X.5 3-3. -. b Fold change by treatment ( (+)/(-)) 8
Supporting information. Single-cell and subcellular pharmacokinetic imaging allows insight into drug action in vivo
 Supporting information Single-cell and subcellular pharmacokinetic imaging allows insight into drug action in vivo Greg Thurber 1, Katy Yang 1, Thomas Reiner 1, Rainer Kohler 1, Peter Sorger 2, Tim Mitchison
Supporting information Single-cell and subcellular pharmacokinetic imaging allows insight into drug action in vivo Greg Thurber 1, Katy Yang 1, Thomas Reiner 1, Rainer Kohler 1, Peter Sorger 2, Tim Mitchison
FIG S1: Calibration curves of standards for HPLC detection of Dopachrome (Dopac), Dopamine (DA) and Homovanillic acid (HVA), showing area of peak vs
 FIG S1: Calibration curves of standards for HPLC detection of Dopachrome (Dopac), Dopamine (DA) and Homovanillic acid (HVA), showing area of peak vs amount of standard analysed. Each point represents the
FIG S1: Calibration curves of standards for HPLC detection of Dopachrome (Dopac), Dopamine (DA) and Homovanillic acid (HVA), showing area of peak vs amount of standard analysed. Each point represents the
Supplemental Data. ALDH1 Is a Marker of Normal and Malignant. Human Mammary Stem Cells. and a Predictor of Poor Clinical Outcome
 Cell Stem Cell, Volume 1 Supplemental Data ALDH1 Is a Marker of Normal and Malignant Human Mammary Stem Cells and a Predictor of Poor Clinical Outcome Christophe Ginestier, Min Hee Hur, Emmanuelle Charafe-Jauffret,
Cell Stem Cell, Volume 1 Supplemental Data ALDH1 Is a Marker of Normal and Malignant Human Mammary Stem Cells and a Predictor of Poor Clinical Outcome Christophe Ginestier, Min Hee Hur, Emmanuelle Charafe-Jauffret,
Nature Immunology: doi: /ni Supplementary Figure 1
 Supplementary Figure 1 PPAR-γ is dispensable for the development of tissue macrophages in the heart, kidneys, lamina propria and white adipose tissue. Plots show the expression of F4/80 and CD11b (a) or
Supplementary Figure 1 PPAR-γ is dispensable for the development of tissue macrophages in the heart, kidneys, lamina propria and white adipose tissue. Plots show the expression of F4/80 and CD11b (a) or
Supplementary Figure Legend
 Supplementary Figure Legend Supplementary Figure S1. Effects of MMP-1 silencing on HEp3-hi/diss cell proliferation in 2D and 3D culture conditions. (A) Downregulation of MMP-1 expression in HEp3-hi/diss
Supplementary Figure Legend Supplementary Figure S1. Effects of MMP-1 silencing on HEp3-hi/diss cell proliferation in 2D and 3D culture conditions. (A) Downregulation of MMP-1 expression in HEp3-hi/diss
Supplementary Figure and Table Legends
 1 Supplementary Figure and Table Legends Figure S1: Whole-animal metabolic analysis. 12 week old WT and Dvl1 / were singly housed in CLAMS cages (Comprehensive Laboratory Animals Monitoring System) for
1 Supplementary Figure and Table Legends Figure S1: Whole-animal metabolic analysis. 12 week old WT and Dvl1 / were singly housed in CLAMS cages (Comprehensive Laboratory Animals Monitoring System) for
Different Potential of Extracellular Vesicles to Support Thrombin Generation: Contributions of Phosphatidylserine, Tissue Factor, and Cellular Origin
 Different Potential of Extracellular Vesicles to Support Thrombin Generation: Contributions of Phosphatidylserine, Tissue Factor, and Cellular Origin Carla Tripisciano 1, René Weiss 1, Tanja Eichhorn 1,
Different Potential of Extracellular Vesicles to Support Thrombin Generation: Contributions of Phosphatidylserine, Tissue Factor, and Cellular Origin Carla Tripisciano 1, René Weiss 1, Tanja Eichhorn 1,
Genome Editing: Cas9 Stable Cell Lines for CRISPR sgrna Validation, Library Screening, and More. Ed Davis, Ph.D.
 TECHNICAL NOTE Genome Editing: Cas9 Stable Cell Lines for CRISPR sgrna Validation, Library Screening, and More Introduction Ed Davis, Ph.D. The CRISPR-Cas9 system has become greatly popular for genome
TECHNICAL NOTE Genome Editing: Cas9 Stable Cell Lines for CRISPR sgrna Validation, Library Screening, and More Introduction Ed Davis, Ph.D. The CRISPR-Cas9 system has become greatly popular for genome
Initial genotyping of all new litters was performed by Transnetyx (Memphis,
 SUPPLEMENTAL INFORMATION MURILLO ET AL. SUPPLEMENTAL EXPERIMENTAL PROCEDURES Mouse Genotyping Initial genotyping of all new litters was performed by Transnetyx (Memphis, USA). Assessment of LoxPpα recombination
SUPPLEMENTAL INFORMATION MURILLO ET AL. SUPPLEMENTAL EXPERIMENTAL PROCEDURES Mouse Genotyping Initial genotyping of all new litters was performed by Transnetyx (Memphis, USA). Assessment of LoxPpα recombination
Introduction to Microarray Analysis
 Introduction to Microarray Analysis Methods Course: Gene Expression Data Analysis -Day One Rainer Spang Microarrays Highly parallel measurement devices for gene expression levels 1. How does the microarray
Introduction to Microarray Analysis Methods Course: Gene Expression Data Analysis -Day One Rainer Spang Microarrays Highly parallel measurement devices for gene expression levels 1. How does the microarray
Beta3 integrin promotes long-lasting activation and polarization of Vascular Endothelial Growth Factor Receptor 2 by immobilized ligand
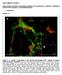 SUPPLEMENTAL FIGURES Beta3 integrin promotes long-lasting activation and polarization of Vascular Endothelial Growth Factor Receptor 2 by immobilized ligand C. Ravelli et al. FIGURE S. I Figure S. I: Gremlin
SUPPLEMENTAL FIGURES Beta3 integrin promotes long-lasting activation and polarization of Vascular Endothelial Growth Factor Receptor 2 by immobilized ligand C. Ravelli et al. FIGURE S. I Figure S. I: Gremlin
Rejuvenation of the muscle stem cell population restores strength to injured aged muscles
 Rejuvenation of the muscle stem cell population restores strength to injured aged muscles Benjamin D Cosgrove, Penney M Gilbert, Ermelinda Porpiglia, Foteini Mourkioti, Steven P Lee, Stephane Y Corbel,
Rejuvenation of the muscle stem cell population restores strength to injured aged muscles Benjamin D Cosgrove, Penney M Gilbert, Ermelinda Porpiglia, Foteini Mourkioti, Steven P Lee, Stephane Y Corbel,
Automated Method for Determination of Infectious Dose (TCID 50 ) using Celigo Imaging Cytometer
 Automated Method for Determination of Infectious Dose (TCID 50 ) using Celigo Imaging Cytometer Nexcelom Bioscience LLC. 360 Merrimack Street, Building 9 Lawrence, MA 01843 T: 978.327.5340 F: 978.327.5341
Automated Method for Determination of Infectious Dose (TCID 50 ) using Celigo Imaging Cytometer Nexcelom Bioscience LLC. 360 Merrimack Street, Building 9 Lawrence, MA 01843 T: 978.327.5340 F: 978.327.5341
ab CFSE Fluorescent Cell Labeling Kit
 ab113853 CFSE Fluorescent Cell Labeling Kit Instructions for Use For the durable fluorescent labeling of live cells for fluorescent microscopy and flow cytometry, population growth studies and within sample
ab113853 CFSE Fluorescent Cell Labeling Kit Instructions for Use For the durable fluorescent labeling of live cells for fluorescent microscopy and flow cytometry, population growth studies and within sample
File name: Supplementary Information Description: Supplementary figures and supplementary tables. File name: Peer review file Description:
 File name: Supplementary Information Description: Supplementary figures and supplementary tables. File name: Peer review file Description: Supplementary figure 1 Flow diagram summarizing the overall experimental
File name: Supplementary Information Description: Supplementary figures and supplementary tables. File name: Peer review file Description: Supplementary figure 1 Flow diagram summarizing the overall experimental
Supplementary Figure 1: MYCER protein expressed from the transgene can enhance
 Relative luciferase activity Relative luciferase activity MYC is a critical target FBXW7 MYC Supplementary is a critical Figures target 1-7. FBXW7 Supplementary Material A E-box sequences 1 2 3 4 5 6 HSV-TK
Relative luciferase activity Relative luciferase activity MYC is a critical target FBXW7 MYC Supplementary is a critical Figures target 1-7. FBXW7 Supplementary Material A E-box sequences 1 2 3 4 5 6 HSV-TK
Intestinal Epithelial Cell-Specific Deletion of PLD2 Alleviates DSS-Induced Colitis by. Regulating Occludin
 Intestinal Epithelial Cell-Specific Deletion of PLD2 Alleviates DSS-Induced Colitis by Regulating Occludin Chaithanya Chelakkot 1,ǂ, Jaewang Ghim 2,3,ǂ, Nirmal Rajasekaran 4, Jong-Sun Choi 5, Jung-Hwan
Intestinal Epithelial Cell-Specific Deletion of PLD2 Alleviates DSS-Induced Colitis by Regulating Occludin Chaithanya Chelakkot 1,ǂ, Jaewang Ghim 2,3,ǂ, Nirmal Rajasekaran 4, Jong-Sun Choi 5, Jung-Hwan
Cell autonomous vs. cell non-autonomous gene function
 Cell autonomous vs. cell non-autonomous gene function In multicellular organisms, it is important to know in what cell(s) the activity of a gene is required. - while RNA expression can be highly informative,
Cell autonomous vs. cell non-autonomous gene function In multicellular organisms, it is important to know in what cell(s) the activity of a gene is required. - while RNA expression can be highly informative,
Chicken EpithelialGut CellLines 1
 Chicken EpithelialGut CellLines 1 Content 01 Introduction p 3 02 Characterization p 5 03 Infection and inhibition p 6 04 Protein expression system p 8 05 NutriProof p 10 06 Contact p 12 01...which came
Chicken EpithelialGut CellLines 1 Content 01 Introduction p 3 02 Characterization p 5 03 Infection and inhibition p 6 04 Protein expression system p 8 05 NutriProof p 10 06 Contact p 12 01...which came
Supplementary Figures Montero et al._supplementary Figure 1
 Montero et al_suppl. Info 1 Supplementary Figures Montero et al._supplementary Figure 1 Montero et al_suppl. Info 2 Supplementary Figure 1. Transcripts arising from the structurally conserved subtelomeres
Montero et al_suppl. Info 1 Supplementary Figures Montero et al._supplementary Figure 1 Montero et al_suppl. Info 2 Supplementary Figure 1. Transcripts arising from the structurally conserved subtelomeres
Diagnostics in Oncology Mark Kockx MD, PhD
 HistoGeneX The Real World A Specialized of companion Biomarker & Integrated Pathology Laboratory Diagnostics in Oncology Mark Kockx MD, PhD 1 2 HistoGeneX located in Antwerp, Belgium and Chicago, Illinois
HistoGeneX The Real World A Specialized of companion Biomarker & Integrated Pathology Laboratory Diagnostics in Oncology Mark Kockx MD, PhD 1 2 HistoGeneX located in Antwerp, Belgium and Chicago, Illinois
Supplemental Information. Mitophagy Controls the Activities. of Tumor Suppressor p53. to Regulate Hepatic Cancer Stem Cells
 Molecular Cell, Volume 68 Supplemental Information Mitophagy Controls the Activities of Tumor Suppressor to Regulate Hepatic Cancer Stem Cells Kai Liu, Jiyoung Lee, Ja Yeon Kim, Linya Wang, Yongjun Tian,
Molecular Cell, Volume 68 Supplemental Information Mitophagy Controls the Activities of Tumor Suppressor to Regulate Hepatic Cancer Stem Cells Kai Liu, Jiyoung Lee, Ja Yeon Kim, Linya Wang, Yongjun Tian,
Supplementary Figure 1
 Supplementary Figure 1 Ex2 promotor region Cre IRES cherry pa Ex4 Ex5 Ex1 untranslated Ex3 Ex5 untranslated EYFP pa Rosa26 STOP loxp loxp Cre recombinase EYFP pa Rosa26 loxp 1 kb Interleukin-9 fate reporter
Supplementary Figure 1 Ex2 promotor region Cre IRES cherry pa Ex4 Ex5 Ex1 untranslated Ex3 Ex5 untranslated EYFP pa Rosa26 STOP loxp loxp Cre recombinase EYFP pa Rosa26 loxp 1 kb Interleukin-9 fate reporter
Isolation, culture, and transfection of primary mammary epithelial organoids
 Supplementary Experimental Procedures Isolation, culture, and transfection of primary mammary epithelial organoids Primary mammary epithelial organoids were prepared from 8-week-old CD1 mice (Charles River)
Supplementary Experimental Procedures Isolation, culture, and transfection of primary mammary epithelial organoids Primary mammary epithelial organoids were prepared from 8-week-old CD1 mice (Charles River)
SUPPLEMENTARY INFORMATION
 In the format provided by the authors and unedited. SUPPLEMENTARY INFORMATION ARTICLE NUMBER: 16206 DOI: 10.1038/NMICROBIOL.2016.206 Single cell RNA seq ties macrophage polarization to growth rate of intracellular
In the format provided by the authors and unedited. SUPPLEMENTARY INFORMATION ARTICLE NUMBER: 16206 DOI: 10.1038/NMICROBIOL.2016.206 Single cell RNA seq ties macrophage polarization to growth rate of intracellular
Supplementary Figure 1. Characterization of EVs (a) Phase-contrast electron microscopy was used to visualize resuspended EV pellets.
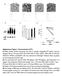 Supplementary Figure 1. Characterization of EVs (a) Phase-contrast electron microscopy was used to visualize resuspended EV pellets. Scale bar represent 100 nm. The sizes of EVs from MDA-MB-231-D3H1 (D3H1),
Supplementary Figure 1. Characterization of EVs (a) Phase-contrast electron microscopy was used to visualize resuspended EV pellets. Scale bar represent 100 nm. The sizes of EVs from MDA-MB-231-D3H1 (D3H1),
FRAUNHOFER IME SCREENINGPORT
 FRAUNHOFER IME SCREENINGPORT Detection technologies used in drug discovery Introduction Detection technologies in drug discovery is unlimited only a biased snapshot can be presented Differences can presented
FRAUNHOFER IME SCREENINGPORT Detection technologies used in drug discovery Introduction Detection technologies in drug discovery is unlimited only a biased snapshot can be presented Differences can presented
Supplementary Figure 1: Two modes of low concentration of BsSMC on a DNA (a) Protein staining (left) and fluorescent imaging of Cy3 (right) confirm
 Supplementary Figure 1: Two modes of low concentration of BsSMC on a DNA (a) Protein staining (left) and fluorescent imaging of Cy3 (right) confirm that BsSMC was labeled with Cy3 NHS-Ester. In each panel,
Supplementary Figure 1: Two modes of low concentration of BsSMC on a DNA (a) Protein staining (left) and fluorescent imaging of Cy3 (right) confirm that BsSMC was labeled with Cy3 NHS-Ester. In each panel,
Contents. The Right Surface for Every Cell Extracellular Matrices and Biologically Coated Surfaces ECM Mimetic and Advanced Surfaces...
 Contents The Right Surface for Every Cell... 1 Extracellular Matrices and Biologically Coated Surfaces... 2 Corning Matrigel Matrix... 2 Corning BioCoat Cultureware... 3 ECM Mimetic and Advanced Surfaces...
Contents The Right Surface for Every Cell... 1 Extracellular Matrices and Biologically Coated Surfaces... 2 Corning Matrigel Matrix... 2 Corning BioCoat Cultureware... 3 ECM Mimetic and Advanced Surfaces...
Coleman et al., Supplementary Figure 1
 Coleman et al., Supplementary Figure 1 BrdU Merge G1 Early S Mid S Supplementary Figure 1. Sequential destruction of CRL4 Cdt2 targets during the G1/S transition. HCT116 cells were synchronized by sequential
Coleman et al., Supplementary Figure 1 BrdU Merge G1 Early S Mid S Supplementary Figure 1. Sequential destruction of CRL4 Cdt2 targets during the G1/S transition. HCT116 cells were synchronized by sequential
Pre-made Lentiviral Particles for nuclear permeant CRE recombinase expression
 Pre-made Lentiviral Particles for nuclear permeant CRE recombinase expression Cat# Product Name Amounts* LVP336 NLS-CRE (Bsd) LVP336-PBS NLS-CRE (Bsd), in vivo ready LVP339 NLS-CRE (Puro) LVP339-PBS NLS-CRE
Pre-made Lentiviral Particles for nuclear permeant CRE recombinase expression Cat# Product Name Amounts* LVP336 NLS-CRE (Bsd) LVP336-PBS NLS-CRE (Bsd), in vivo ready LVP339 NLS-CRE (Puro) LVP339-PBS NLS-CRE
Supplementary Fig. S1. Schematic representation of mouse lines Pax6 fl/fl and mrx-cre used in this study. (A) To generate Pax6 fl/ fl
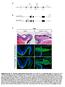 Supplementary Fig. S1. Schematic representation of mouse lines Pax6 fl/fl and mrx-cre used in this study. (A) To generate Pax6 fl/ fl, loxp sites flanking exons 3-6 (red arrowheads) were introduced into
Supplementary Fig. S1. Schematic representation of mouse lines Pax6 fl/fl and mrx-cre used in this study. (A) To generate Pax6 fl/ fl, loxp sites flanking exons 3-6 (red arrowheads) were introduced into
ab Hypoxic Response Human Flow Cytometry Kit
 ab126585 Hypoxic Response Human Flow Cytometry Kit Instructions for Use For measuring protein levels by flow cytometry: hypoxia-inducible factor 1-alpha (HIF1A) and BCL2/adenovirus E1B 19 kda proteininteracting
ab126585 Hypoxic Response Human Flow Cytometry Kit Instructions for Use For measuring protein levels by flow cytometry: hypoxia-inducible factor 1-alpha (HIF1A) and BCL2/adenovirus E1B 19 kda proteininteracting
Nature Methods: doi: /nmeth Supplementary Figure 1. Validation of RaPID with EDEN15
 Supplementary Figure 1 Validation of RaPID with EDEN15 (a) Full Western Blot of conventional biotinylated RNA pulldown with EDEN15 and scrambled control (n=3 biologically independent experiments, representative
Supplementary Figure 1 Validation of RaPID with EDEN15 (a) Full Western Blot of conventional biotinylated RNA pulldown with EDEN15 and scrambled control (n=3 biologically independent experiments, representative
Supplementary Material. TRIB3 inhibits proliferation and promotes osteogenesis in hbmscs by regulating the. ERK1/2 signaling pathway
 Supplementary Material TRIB3 inhibits proliferation and promotes osteogenesis in hbmscs by regulating the ERK1/2 signaling pathway Cui Zhang 1, Fan-Fan Hong 1, Cui-Cui Wang 1, Liang Li 1, Jian-Ling Chen
Supplementary Material TRIB3 inhibits proliferation and promotes osteogenesis in hbmscs by regulating the ERK1/2 signaling pathway Cui Zhang 1, Fan-Fan Hong 1, Cui-Cui Wang 1, Liang Li 1, Jian-Ling Chen
SureSilencing sirna Array Technology Overview
 SureSilencing sirna Array Technology Overview Pathway-Focused sirna-based RNA Interference Topics to be Covered Who is SuperArray? Brief Introduction to RNA Interference Challenges Facing RNA Interference
SureSilencing sirna Array Technology Overview Pathway-Focused sirna-based RNA Interference Topics to be Covered Who is SuperArray? Brief Introduction to RNA Interference Challenges Facing RNA Interference
Supplementary Figure 1: Derivation and characterization of RN ips cell lines. (a) RN ips cells maintain expression of pluripotency markers OCT4 and
 Supplementary Figure 1: Derivation and characterization of RN ips cell lines. (a) RN ips cells maintain expression of pluripotency markers OCT4 and SSEA4 after 10 passages in mtesr 1 medium. (b) Schematic
Supplementary Figure 1: Derivation and characterization of RN ips cell lines. (a) RN ips cells maintain expression of pluripotency markers OCT4 and SSEA4 after 10 passages in mtesr 1 medium. (b) Schematic
Xeno-Free Systems for hesc & hipsc. Facilitating the shift from Stem Cell Research to Clinical Applications
 Xeno-Free Systems for hesc & hipsc Facilitating the shift from Stem Cell Research to Clinical Applications NutriStem Defined, xeno-free (XF), serum-free media (SFM) specially formulated for growth and
Xeno-Free Systems for hesc & hipsc Facilitating the shift from Stem Cell Research to Clinical Applications NutriStem Defined, xeno-free (XF), serum-free media (SFM) specially formulated for growth and
ASPP1 Fw GGTTGGGAATCCACGTGTTG ASPP1 Rv GCCATATCTTGGAGCTCTGAGAG
 Supplemental Materials and Methods Plasmids: the following plasmids were used in the supplementary data: pwzl-myc- Lats2 (Aylon et al, 2006), pretrosuper-vector and pretrosuper-shp53 (generous gift of
Supplemental Materials and Methods Plasmids: the following plasmids were used in the supplementary data: pwzl-myc- Lats2 (Aylon et al, 2006), pretrosuper-vector and pretrosuper-shp53 (generous gift of
Supplementary Figure 1 qrt-pcr expression analysis of NLP8 with and without KNO 3 during germination.
 Supplementary Figure 1 qrt-pcr expression analysis of NLP8 with and without KNO 3 during germination. Seeds of Col-0 were harvested from plants grown at 16 C, stored for 2 months, imbibed for indicated
Supplementary Figure 1 qrt-pcr expression analysis of NLP8 with and without KNO 3 during germination. Seeds of Col-0 were harvested from plants grown at 16 C, stored for 2 months, imbibed for indicated
Center Drive, University of Michigan Health System, Ann Arbor, MI
 Leukotriene B 4 -induced reduction of SOCS1 is required for murine macrophage MyD88 expression and NFκB activation Carlos H. Serezani 1,3, Casey Lewis 1, Sonia Jancar 2 and Marc Peters-Golden 1,3 1 Division
Leukotriene B 4 -induced reduction of SOCS1 is required for murine macrophage MyD88 expression and NFκB activation Carlos H. Serezani 1,3, Casey Lewis 1, Sonia Jancar 2 and Marc Peters-Golden 1,3 1 Division
Real-time PCR. Total RNA was isolated from purified splenic or LP macrophages using
 Supplementary Methods Real-time PCR. Total RNA was isolated from purified splenic or LP macrophages using the Qiagen RNeasy Mini Kit, according to the manufacturer s protocol with on-column DNase digestion
Supplementary Methods Real-time PCR. Total RNA was isolated from purified splenic or LP macrophages using the Qiagen RNeasy Mini Kit, according to the manufacturer s protocol with on-column DNase digestion
% Viability. isw2 ino isw2 ino isw2 ino isw2 ino mM HU 4-NQO CPT
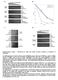 a Drug concentration b 1.3% MMS nhp1 nhp1 8 nhp1 mag1.5% MMS.3% MMS nhp1 nhp1 ino8 9 ino8 9 % Viability 4.5% MMS ino8 9 ino8 9 2.5.1.15 % MMS c d nhp1 nhp1 nhp1 nhp1 nhp1 nhp1 Control (YPD) γ IR (1 gy)
a Drug concentration b 1.3% MMS nhp1 nhp1 8 nhp1 mag1.5% MMS.3% MMS nhp1 nhp1 ino8 9 ino8 9 % Viability 4.5% MMS ino8 9 ino8 9 2.5.1.15 % MMS c d nhp1 nhp1 nhp1 nhp1 nhp1 nhp1 Control (YPD) γ IR (1 gy)
Supplementary Figure 1. (a) The qrt-pcr for lnc-2, lnc-6 and lnc-7 RNA level in DU145, 22Rv1, wild type HCT116 and HCT116 Dicer ex5 cells transfected
 Supplementary Figure 1. (a) The qrt-pcr for lnc-2, lnc-6 and lnc-7 RNA level in DU145, 22Rv1, wild type HCT116 and HCT116 Dicer ex5 cells transfected with the sirna against lnc-2, lnc-6, lnc-7, and the
Supplementary Figure 1. (a) The qrt-pcr for lnc-2, lnc-6 and lnc-7 RNA level in DU145, 22Rv1, wild type HCT116 and HCT116 Dicer ex5 cells transfected with the sirna against lnc-2, lnc-6, lnc-7, and the
