Supplemental Information. Mitophagy Controls the Activities. of Tumor Suppressor p53. to Regulate Hepatic Cancer Stem Cells
|
|
|
- Crystal Chase
- 6 years ago
- Views:
Transcription
1 Molecular Cell, Volume 68 Supplemental Information Mitophagy Controls the Activities of Tumor Suppressor to Regulate Hepatic Cancer Stem Cells Kai Liu, Jiyoung Lee, Ja Yeon Kim, Linya Wang, Yongjun Tian, Stephanie T. Chan, Cecilia Cho, Keigo Machida, Dexi Chen, and JingHsiung James Ou
2 SUPPLEMENTAL INFORMATION Supplemental Figure Legends Figure S1. Representative flow cytometry and effect of ATG7 on HepG2 cells. Related to Figure 1. (A) Flow cytometry analysis of CD133 cells of HepG2 with various treatments or with stable expression of the control shrna or the Atg5 shrna. () Flow cytometry analysis of CD133 CD49f cells in liver tumors of control mice and ATG5knockout mice that had been treated with DEN. (C)(E) HepG2 cells without treatment (None) or with the stable expression of control shrna (shctrl) or ATG7 shrna (shatg7) were subjected to immunoblot analysis (C), flow cytometry analysis of CD133 cells (D) or sphereformation assay (E). Figure S2. Analysis of the effect of PFTα on. Related to Figure 2. (A) Analysis of the effect of PFTα on and its subcellular localization. Cytoplasmic and nuclear lysates of HepG2 cells that had been treated with PFTα or DMSO for one day were lysed and separated into cytoplasmic and nuclear fractions for immunoblot analysis. () Analysis on the effect of PFTα on cell viability. The viability of HepG2 cells treated with DMSO or PFTα were analyzed by the MTT assay. Figure S3. Effects of ATG5 on. Related to Figure 3. (A) Immunoblot analysis of liver tumors of control mice and ATG5KO mice that had been treated with DEN. () The level of mrna in HepG2 cells with various treatments for 24 hours or with stable ATG5 knockdown were quantified by realtime RTPCR. No significant difference of mrna levels was observed. Figure S4. Analysis of the effect of on the NANOG promoter. Related to Figure 4. (A) The nucleotide sequence of the Nanog promoter that contains the putative response element (highlighted in yellow color) and the OCT4SOX2 binding site (highlighted in blue color). The consensus sequence of the binding site is shown below the NANOG promoter sequence. R, A and G; W, T and A; and Y, T and C. () The supershift assay using the anti antibody that recognized phosphoserine392. The supershift assay was conducted as shown in Figure 4D, with the exception that an increasing amount of the antibody was used for the supershift assay. (C) Effects of various constructs on the NANOG promoter in Huh7 cells. Huh7 cells were cotransfected with the Nanogluc2 reporter and the control vector or the expression plasmid for (WT), (S392A) or (S392D). The reporter psvrl, which
3 expressed renilla luciferase, was included in the transfection to monitor the transfection efficiency. The luciferase activity expressed from the parental vector pgl3basic was arbitrarily defined as 1. The results represent the mean ± SEM of three independent experiments Figure S5. Effects of Mdivi1, CCCP and DFP on mitophagy. Related to Figure 5. HepG2 cells were treated with DMSO, Mdivi1, CCCP or DFP for 24 hours. (A) Cells were lysed for the preparation of total cell lysates or for the isolation of mitochondria for immunoblot analysis. () Mitochondrial DNA (mtdna) was quantified by qpcr and normalized against GAPDH DNA. The mtdna level of control cells was arbitrarily defined as 1. The results represent the mean of at least three different experiments. (C) Analysis of mitophagy using the mkeimared reporter. Red color indicated mitophagy. Scale bar, 1 µm. (D) Immunoblot analysis of cells without (Ctrl) and with treatment with DFP. Figure S6. and the phosphorylation of S392 of. Related to Figure 6. (A) HepG2 cells without treatment or with transfection of a control vector or the expressing plasmid were lysed 48 hours after transfection and analyzed by immunoblot for phosphorylated ubiquitin, total ubiquitin, phosphorylated PARKIN, total PARKIN and. Actin served as the loading control. () Analysis of the effect of silencing of various kinases on the phosphorylation of S392 of. HepG2 cells transfected with various sirnas for two days were lysed for immunoblot analysis. (C) Huh7 cells, which expressed (Y22C), were treated with the control sirna (si Ctrl) or si and lysed for immunoblot analysis. (D) Stable HepG2 cells that expressed the control shrna (shctrl) or shrna (sh) were lysed for immunoblot analysis. (E) Stable HepG2 cells with knockdown were transfected with a control vector or a plasmid that expressed the shrnaresistant wildtype (left panel), or with a control vector or a plasmid that expressed the kinase dead mutant (mt) (right panel) followed by immunoblot analysis. Figure S7. Role of in phosphorylation and tumorigenesis. Related to Figure 6. (A) GSTPARKIN was mixed with GST or GST and incubated in the presence of ATP. The phosphorylation of PARKIN at S65 was then analyzed with the antibody that recognized phosphorylated S65. GSTPARKIN, GST and GST used for the reaction was also analyzed by antiparkin, anti and antigst antibodies, respectively (bottom three panels). () The phosphorylation of ubiquitin by GST was conducted the same way as in (A), with the exception that GSTPARKIN was replaced with tetraubiquitin. (C) The
4 phosphorylation analysis of GST was conducted the same way as described in the legend to Figure 6E, with the exception that γ 32 PATP was used for the labeling reaction and the phosphorylated protein was analyzed by autoradiography. (D) The representative SiMPull images are shown on the top. The chart demonstrated a dosedependent binding of GST by. The Kd was estimated to be ~5 nm. (E) HepG2 cells without or with treatment of DMSO or 3MA as well as stable HepG2 cells that expressed shctrl, shatg5 or shatg7 were lysed for immunoblot analysis. (F) Stable HepG2 cells that expressed shctrl, shatg5 or sh Atg5 and sh were lysed for immunoblot analysis. (G) CD133 cells of the cell lines shown in (F) were subcutaneously injected into nude mice for tumorigenesis studies. Tumor imaging was conducted 3 months later. Representative tumor images were shown to the left and the quantification of photons, which reflected tumor sizes, were shown in the chart to the right.
5 Figure S1 A None DMSO 3MA afa1 Rapa Serum free shctrl shatg5 Counts 5.6% 5.5% 1.5% 1.9% 13.7% 11.4% 5.6% 1.9% CD133 CD49f CD133 Control.85% ATG5KO.14% C LC3I LC3II p62 Atg7 CD133 NANOG OCT4 None shctrl shatg7 D % CD133 % CD None p<.1 p<.1 shctrl shatg7 E Number of Spheres shctrl shatg7 CD133 CD133 SOX2 βactin
6 None PFT Á DMSO No treatment PFTα DMSO No treatment PFTα DMSO Figure S2 A (ps392) Lamin 1 βactin nucleus cytoplasm None PFTα DMSO Cell Viability viability(%)
7 None shctrl shatg5 3MA afa1 Rapamycin Serum free Figure S Relative mrna levels A Ctrl Atg5 KO CD133 Nanog (ps392) βactin
8 Figure S4 A Motif 1 Motif 2 OCT4SOX2 site NANOG promoter Supershift shift Free probe probe Nuclear extracts C Relative luciferase activity pgl3basic Nanogluc2 (WT) (S392A) (S392D) Anti(pS392) (μg)
9 Figure S5 A Mitochondria Total lysates Treatment: Tom2 Tim23 βactin LC3I LC3II TOM2 Relative mtdna levels Ctrl DFP CCCP Mdivi D CD133 Nanog (ps392) actin Ctrl DFP C Control DFP CCCP Mdivi
10 Figure S6 A sirna: (ps392 ) CK2α C Ubiquitin(pS65) Ubiquitin p38(mapk) PKR Cdk9 βactin (ps392 ) βactin PARKIN(pS65) PARKIN βactin D (ps392) βactin None shctrl sh E (ps392) βactin mt (ps392) βactin
11 Figure S7 A C GSTPARKIN GST GST PARKIN(pS65) TetraUbiquitin GST GST GST GST GST D GST GST PARKIN GST Ubiquitin(pS65) TetraUbiquitin GST ( 32 ps392) GST Fraction ound Kd 4.98 nm Concentration (nm) y =.1775ln(x).2147 R² =.9988 GST E βactin F Atg5 (ps392) NANOG βactin None shctrl shatg5 shatg5/ G ATG5 KD: Control ATG5 Photons/sec/cm 2 (x1 5 ) KD: Ctrl ATG5 ATG5
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1.
 Supplementary Figure 1. Characterization and high expression of Lnc-β-Catm in liver CSCs. (a) Heatmap of differently expressed lncrnas in Liver CSCs (CD13 + CD133 + ) and non-cscs (CD13 - CD133 - ) according
Supplementary Figure 1. Characterization and high expression of Lnc-β-Catm in liver CSCs. (a) Heatmap of differently expressed lncrnas in Liver CSCs (CD13 + CD133 + ) and non-cscs (CD13 - CD133 - ) according
Supplementary Table 1. The Q-PCR primer sequence is summarized in the following table.
 Supplementary Table 1. The Q-PCR primer sequence is summarized in the following table. Name Sequence (5-3 ) Application Flag-u ggactacaaggacgacgatgac Shared upstream primer for all the amplifications of
Supplementary Table 1. The Q-PCR primer sequence is summarized in the following table. Name Sequence (5-3 ) Application Flag-u ggactacaaggacgacgatgac Shared upstream primer for all the amplifications of
ASPP1 Fw GGTTGGGAATCCACGTGTTG ASPP1 Rv GCCATATCTTGGAGCTCTGAGAG
 Supplemental Materials and Methods Plasmids: the following plasmids were used in the supplementary data: pwzl-myc- Lats2 (Aylon et al, 2006), pretrosuper-vector and pretrosuper-shp53 (generous gift of
Supplemental Materials and Methods Plasmids: the following plasmids were used in the supplementary data: pwzl-myc- Lats2 (Aylon et al, 2006), pretrosuper-vector and pretrosuper-shp53 (generous gift of
Journal of Cell Science Supplementary Material
 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 SUPPLEMENTARY FIGURE LEGENDS Figure S1: Eps8 is localized at focal adhesions and binds directly to FAK (A) Focal
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 SUPPLEMENTARY FIGURE LEGENDS Figure S1: Eps8 is localized at focal adhesions and binds directly to FAK (A) Focal
Supporting Information
 Supporting Information Su et al. 10.1073/pnas.1211604110 SI Materials and Methods Cell Culture and Plasmids. Tera-1 and Tera-2 cells (ATCC: HTB- 105/106) were maintained in McCoy s 5A medium with 15% FBS
Supporting Information Su et al. 10.1073/pnas.1211604110 SI Materials and Methods Cell Culture and Plasmids. Tera-1 and Tera-2 cells (ATCC: HTB- 105/106) were maintained in McCoy s 5A medium with 15% FBS
Supplementary Figure 1. Lnc-β-Catm characterization. (A, B) LncBRM silenced
 Supplementary Figure 1. Lnc-β-Catm characterization. (A, B) LncBRM silenced Hep3B (A) and Huh7 (B) cells were established using psicor lentivirus, followed by sphere formation assays. (C, D) LncBRM deleted
Supplementary Figure 1. Lnc-β-Catm characterization. (A, B) LncBRM silenced Hep3B (A) and Huh7 (B) cells were established using psicor lentivirus, followed by sphere formation assays. (C, D) LncBRM deleted
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION Dynamic Phosphorylation of HP1 Regulates Mitotic Progression in Human Cells Supplementary Figures Supplementary Figure 1. NDR1 interacts with HP1. (a) Immunoprecipitation using
SUPPLEMENTARY INFORMATION Dynamic Phosphorylation of HP1 Regulates Mitotic Progression in Human Cells Supplementary Figures Supplementary Figure 1. NDR1 interacts with HP1. (a) Immunoprecipitation using
Supplementary Fig. 1 related to Fig. 1 Clinical relevance of lncrna candidate
 Supplementary Figure Legends Supplementary Fig. 1 related to Fig. 1 Clinical relevance of lncrna candidate BC041951 in gastric cancer. (A) The flow chart for selected candidate lncrnas in 660 up-regulated
Supplementary Figure Legends Supplementary Fig. 1 related to Fig. 1 Clinical relevance of lncrna candidate BC041951 in gastric cancer. (A) The flow chart for selected candidate lncrnas in 660 up-regulated
HCT116 SW48 Nutlin: p53
 Figure S HCT6 SW8 Nutlin: - + - + p GAPDH Figure S. Nutlin- treatment induces p protein. HCT6 and SW8 cells were left untreated or treated for 8 hr with Nutlin- ( µm) to up-regulate p. Whole cell lysates
Figure S HCT6 SW8 Nutlin: - + - + p GAPDH Figure S. Nutlin- treatment induces p protein. HCT6 and SW8 cells were left untreated or treated for 8 hr with Nutlin- ( µm) to up-regulate p. Whole cell lysates
Figure S1. Figure S2. Figure S3 HB Anti-FSP27 (COOH-terminal peptide) Ab. Anti-GST-FSP27(45-127) Ab.
 / 36B4 mrna ratio Figure S1 * 2. 1.6 1.2.8 *.4 control TNFα BRL49653 Figure S2 Su bw AT p iw Anti- (COOH-terminal peptide) Ab Blot : Anti-GST-(45-127) Ab β-actin Figure S3 HB2 HW AT BA T Figure S4 A TAG
/ 36B4 mrna ratio Figure S1 * 2. 1.6 1.2.8 *.4 control TNFα BRL49653 Figure S2 Su bw AT p iw Anti- (COOH-terminal peptide) Ab Blot : Anti-GST-(45-127) Ab β-actin Figure S3 HB2 HW AT BA T Figure S4 A TAG
Supplementary Figure Legend
 Supplementary Figure Legend Supplementary Figure S1. Effects of MMP-1 silencing on HEp3-hi/diss cell proliferation in 2D and 3D culture conditions. (A) Downregulation of MMP-1 expression in HEp3-hi/diss
Supplementary Figure Legend Supplementary Figure S1. Effects of MMP-1 silencing on HEp3-hi/diss cell proliferation in 2D and 3D culture conditions. (A) Downregulation of MMP-1 expression in HEp3-hi/diss
Figure S2. Response of mouse ES cells to GSK3 inhibition. Mentioned in discussion
 Stem Cell Reports, Volume 1 Supplemental Information Robust Self-Renewal of Rat Embryonic Stem Cells Requires Fine-Tuning of Glycogen Synthase Kinase-3 Inhibition Yaoyao Chen, Kathryn Blair, and Austin
Stem Cell Reports, Volume 1 Supplemental Information Robust Self-Renewal of Rat Embryonic Stem Cells Requires Fine-Tuning of Glycogen Synthase Kinase-3 Inhibition Yaoyao Chen, Kathryn Blair, and Austin
Coleman et al., Supplementary Figure 1
 Coleman et al., Supplementary Figure 1 BrdU Merge G1 Early S Mid S Supplementary Figure 1. Sequential destruction of CRL4 Cdt2 targets during the G1/S transition. HCT116 cells were synchronized by sequential
Coleman et al., Supplementary Figure 1 BrdU Merge G1 Early S Mid S Supplementary Figure 1. Sequential destruction of CRL4 Cdt2 targets during the G1/S transition. HCT116 cells were synchronized by sequential
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb2774 Figure S1 TRF2 dosage modulates the tumorigenicity of mouse and human tumor cells. (a) Left: immunoblotting with antibodies directed against the Myc tag of the transduced TRF2 forms
DOI: 10.1038/ncb2774 Figure S1 TRF2 dosage modulates the tumorigenicity of mouse and human tumor cells. (a) Left: immunoblotting with antibodies directed against the Myc tag of the transduced TRF2 forms
Short hairpin RNA (shrna) against MMP14. Lentiviral plasmids containing shrna
 Supplemental Materials and Methods Short hairpin RNA (shrna) against MMP14. Lentiviral plasmids containing shrna (Mission shrna, Sigma) against mouse MMP14 were transfected into HEK293 cells using FuGene6
Supplemental Materials and Methods Short hairpin RNA (shrna) against MMP14. Lentiviral plasmids containing shrna (Mission shrna, Sigma) against mouse MMP14 were transfected into HEK293 cells using FuGene6
Supplemental Table 1 Gene Symbol FDR corrected p-value PLOD1 CSRP2 PFKP ADFP ADM C10orf10 GPI LOX PLEKHA2 WIPF1
 Supplemental Table 1 Gene Symbol FDR corrected p-value PLOD1 4.52E-18 PDK1 6.77E-18 CSRP2 4.42E-17 PFKP 1.23E-14 MSH2 3.79E-13 NARF_A 5.56E-13 ADFP 5.56E-13 FAM13A1 1.56E-12 FAM29A_A 1.22E-11 CA9 1.54E-11
Supplemental Table 1 Gene Symbol FDR corrected p-value PLOD1 4.52E-18 PDK1 6.77E-18 CSRP2 4.42E-17 PFKP 1.23E-14 MSH2 3.79E-13 NARF_A 5.56E-13 ADFP 5.56E-13 FAM13A1 1.56E-12 FAM29A_A 1.22E-11 CA9 1.54E-11
Supplemental Information. The TRAIL-Induced Cancer Secretome. Promotes a Tumor-Supportive Immune. Microenvironment via CCR2
 Molecular Cell, Volume 65 Supplemental Information The TRAIL-Induced Cancer Secretome Promotes a Tumor-Supportive Immune Microenvironment via CCR2 Torsten Hartwig, Antonella Montinaro, Silvia von Karstedt,
Molecular Cell, Volume 65 Supplemental Information The TRAIL-Induced Cancer Secretome Promotes a Tumor-Supportive Immune Microenvironment via CCR2 Torsten Hartwig, Antonella Montinaro, Silvia von Karstedt,
Confocal immunofluorescence microscopy
 Confocal immunofluorescence microscopy HL-6 and cells were cultured and cytospun onto glass slides. The cells were double immunofluorescence stained for Mt NPM1 and fibrillarin (nucleolar marker). Briefly,
Confocal immunofluorescence microscopy HL-6 and cells were cultured and cytospun onto glass slides. The cells were double immunofluorescence stained for Mt NPM1 and fibrillarin (nucleolar marker). Briefly,
SANTA CRUZ BIOTECHNOLOGY, INC.
 TECHNICAL SERVICE GUIDE: Western Blotting 2. What size bands were expected and what size bands were detected? 3. Was the blot blank or was a dark background or non-specific bands seen? 4. Did this same
TECHNICAL SERVICE GUIDE: Western Blotting 2. What size bands were expected and what size bands were detected? 3. Was the blot blank or was a dark background or non-specific bands seen? 4. Did this same
Tumor Growth Suppression Through the Activation of p21, a Cyclin-Dependent Kinase Inhibitor
 Tumor Growth Suppression Through the Activation of p21, a Cyclin-Dependent Kinase Inhibitor Nicholas Love 11/28/01 A. What is p21? Introduction - p21 is a gene found on chromosome 6 at 6p21.2 - this gene
Tumor Growth Suppression Through the Activation of p21, a Cyclin-Dependent Kinase Inhibitor Nicholas Love 11/28/01 A. What is p21? Introduction - p21 is a gene found on chromosome 6 at 6p21.2 - this gene
Int. J. Mol. Sci. 2016, 17, 1259; doi: /ijms
 S1 of S5 Supplementary Materials: Fibroblast-Derived Extracellular Matrix Induces Chondrogenic Differentiation in Human Adipose-Derived Mesenchymal Stromal/Stem Cells in Vitro Kevin Dzobo, Taegyn Turnley,
S1 of S5 Supplementary Materials: Fibroblast-Derived Extracellular Matrix Induces Chondrogenic Differentiation in Human Adipose-Derived Mesenchymal Stromal/Stem Cells in Vitro Kevin Dzobo, Taegyn Turnley,
Hossain_Supplemental Figure 1
 Hossain_Supplemental Figure 1 GFP-PACT GFP-PACT Motif I GFP-PACT Motif II A. MG132 (1µM) GFP Tubulin GFP-PACT Pericentrin GFP-PACT GFP-PACT Pericentrin Fig. S1. Expression and localization of Orc1 PACT
Hossain_Supplemental Figure 1 GFP-PACT GFP-PACT Motif I GFP-PACT Motif II A. MG132 (1µM) GFP Tubulin GFP-PACT Pericentrin GFP-PACT GFP-PACT Pericentrin Fig. S1. Expression and localization of Orc1 PACT
Regulation of Synaptic Structure and Function by FMRP- Associated MicroRNAs mir-125b and mir-132
 Neuron, Volume 65 Regulation of Synaptic Structure and Function by FMRP- Associated MicroRNAs mir-125b and mir-132 Dieter Edbauer, Joel R. Neilson, Kelly A. Foster, Chi-Fong Wang, Daniel P. Seeburg, Matthew
Neuron, Volume 65 Regulation of Synaptic Structure and Function by FMRP- Associated MicroRNAs mir-125b and mir-132 Dieter Edbauer, Joel R. Neilson, Kelly A. Foster, Chi-Fong Wang, Daniel P. Seeburg, Matthew
RNA oligonucleotides and 2 -O-methylated oligonucleotides were synthesized by. 5 AGACACAAACACCAUUGUCACACUCCACAGC; Rand-2 OMe,
 Materials and methods Oligonucleotides and DNA constructs RNA oligonucleotides and 2 -O-methylated oligonucleotides were synthesized by Dharmacon Inc. (Lafayette, CO). The sequences were: 122-2 OMe, 5
Materials and methods Oligonucleotides and DNA constructs RNA oligonucleotides and 2 -O-methylated oligonucleotides were synthesized by Dharmacon Inc. (Lafayette, CO). The sequences were: 122-2 OMe, 5
The microtubule-associated tau protein has intrinsic acetyltransferase activity. Todd J. Cohen, Dave Friedmann, Andrew W. Hwang, Ronen Marmorstein and
 SUPPLEMENTARY INFORMATION: The microtubule-associated tau protein has intrinsic acetyltransferase activity Todd J. Cohen, Dave Friedmann, Andrew W. Hwang, Ronen Marmorstein and Virginia M.Y. Lee Cohen
SUPPLEMENTARY INFORMATION: The microtubule-associated tau protein has intrinsic acetyltransferase activity Todd J. Cohen, Dave Friedmann, Andrew W. Hwang, Ronen Marmorstein and Virginia M.Y. Lee Cohen
Supplementary Information
 Supplementary Information Live imaging reveals the dynamics and regulation of mitochondrial nucleoids during the cell cycle in Fucci2-HeLa cells Taeko Sasaki 1, Yoshikatsu Sato 2, Tetsuya Higashiyama 1,2,
Supplementary Information Live imaging reveals the dynamics and regulation of mitochondrial nucleoids during the cell cycle in Fucci2-HeLa cells Taeko Sasaki 1, Yoshikatsu Sato 2, Tetsuya Higashiyama 1,2,
GFP CCD2 GFP IP:GFP
 D1 D2 1 75 95 148 178 492 GFP CCD1 CCD2 CCD2 GFP D1 D2 GFP D1 D2 Beclin 1 IB:GFP IP:GFP Supplementary Figure 1: Mapping domains required for binding to HEK293T cells are transfected with EGFP-tagged mutant
D1 D2 1 75 95 148 178 492 GFP CCD1 CCD2 CCD2 GFP D1 D2 GFP D1 D2 Beclin 1 IB:GFP IP:GFP Supplementary Figure 1: Mapping domains required for binding to HEK293T cells are transfected with EGFP-tagged mutant
Supporting Online Material for
 www.sciencemag.org/cgi/content/full/1137999/dc1 Supporting Online Material for Disrupting the Pairing Between let-7 and Enhances Oncogenic Transformation Christine Mayr, Michael T. Hemann, David P. Bartel*
www.sciencemag.org/cgi/content/full/1137999/dc1 Supporting Online Material for Disrupting the Pairing Between let-7 and Enhances Oncogenic Transformation Christine Mayr, Michael T. Hemann, David P. Bartel*
Gaussia Luciferase-a Novel Bioluminescent Reporter for Tracking Stem Cells Survival, Proliferation and Differentiation in Vivo
 Gaussia Luciferase-a Novel Bioluminescent Reporter for Tracking Stem Cells Survival, Proliferation and Differentiation in Vivo Rampyari Raja Walia and Bakhos A. Tannous 1 2 1 Pluristem Innovations, 1453
Gaussia Luciferase-a Novel Bioluminescent Reporter for Tracking Stem Cells Survival, Proliferation and Differentiation in Vivo Rampyari Raja Walia and Bakhos A. Tannous 1 2 1 Pluristem Innovations, 1453
Supplementary Information Design of small molecule-responsive micrornas based on structural requirements for Drosha processing
 Supplementary Information Design of small molecule-responsive micrornas based on structural requirements for Drosha processing Chase L. Beisel, Yvonne Y. Chen, Stephanie J. Culler, Kevin G. Hoff, & Christina
Supplementary Information Design of small molecule-responsive micrornas based on structural requirements for Drosha processing Chase L. Beisel, Yvonne Y. Chen, Stephanie J. Culler, Kevin G. Hoff, & Christina
Supplementary Figure 1: MYCER protein expressed from the transgene can enhance
 Relative luciferase activity Relative luciferase activity MYC is a critical target FBXW7 MYC Supplementary is a critical Figures target 1-7. FBXW7 Supplementary Material A E-box sequences 1 2 3 4 5 6 HSV-TK
Relative luciferase activity Relative luciferase activity MYC is a critical target FBXW7 MYC Supplementary is a critical Figures target 1-7. FBXW7 Supplementary Material A E-box sequences 1 2 3 4 5 6 HSV-TK
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION Legends for Supplementary Tables. Supplementary Table 1. An excel file containing primary screen data. Worksheet 1, Normalized quantification data from a duplicated screen: valid
SUPPLEMENTARY INFORMATION Legends for Supplementary Tables. Supplementary Table 1. An excel file containing primary screen data. Worksheet 1, Normalized quantification data from a duplicated screen: valid
Comparison of Commercial Transfection Reagents: Cell line optimized transfection kits for in vitro cancer research.
 Comparison of Commercial Transfection Reagents: Cell line optimized transfection kits for in vitro cancer research. by Altogen Labs, 11200 Manchaca Road, Suite 203 Austin TX 78748 USA Tel. (512) 433-6177
Comparison of Commercial Transfection Reagents: Cell line optimized transfection kits for in vitro cancer research. by Altogen Labs, 11200 Manchaca Road, Suite 203 Austin TX 78748 USA Tel. (512) 433-6177
ingenio electroporation kits & solution
 ingenio electroporation kits & solution Electroporation x DEFINITION and OPTIMIZATION What is ELECTROPORATION? Electroporation is a physical method of nucleic acid transfer wherein the cells and nucleic
ingenio electroporation kits & solution Electroporation x DEFINITION and OPTIMIZATION What is ELECTROPORATION? Electroporation is a physical method of nucleic acid transfer wherein the cells and nucleic
Supplementary Figures
 Supplementary Figures Supplementary Figure 1. Description of the observed lymphatic metastases in two different SIX1-induced MCF7 metastasis models (Nude and NOD/SCID). Supplementary Figure 2. MCF7-SIX1
Supplementary Figures Supplementary Figure 1. Description of the observed lymphatic metastases in two different SIX1-induced MCF7 metastasis models (Nude and NOD/SCID). Supplementary Figure 2. MCF7-SIX1
FBH1 Catalyzes Regression of Stalled Replication Forks
 Cell Reports Supplemental Information FBH1 Catalyzes Regression of Stalled Replication Forks Kasper Fugger, Martin Mistrik, Kai J. Neelsen, Qi Yao, Ralph Zellweger, Arne Nedergaard Kousholt, Peter Haahr,
Cell Reports Supplemental Information FBH1 Catalyzes Regression of Stalled Replication Forks Kasper Fugger, Martin Mistrik, Kai J. Neelsen, Qi Yao, Ralph Zellweger, Arne Nedergaard Kousholt, Peter Haahr,
Nature Immunology: doi: /ni Supplementary Figure 1. Zranb1 gene targeting.
 Supplementary Figure 1 Zranb1 gene targeting. (a) Schematic picture of Zranb1 gene targeting using an FRT-LoxP vector, showing the first 6 exons of Zranb1 gene (exons 7-9 are not shown). Targeted mice
Supplementary Figure 1 Zranb1 gene targeting. (a) Schematic picture of Zranb1 gene targeting using an FRT-LoxP vector, showing the first 6 exons of Zranb1 gene (exons 7-9 are not shown). Targeted mice
Supplementary information to accompany: A novel role for the DNA repair gene Rad51 in Netrin-1 signalling
 Supplementary information to accompany: A novel role for the DNA repair gene Rad51 in Netrin-1 signalling Glendining KA 1, Markie D 2, Gardner RJM 4, Franz EA 3, Robertson SP 4, Jasoni CL 1 Supplementary
Supplementary information to accompany: A novel role for the DNA repair gene Rad51 in Netrin-1 signalling Glendining KA 1, Markie D 2, Gardner RJM 4, Franz EA 3, Robertson SP 4, Jasoni CL 1 Supplementary
* ** ** * IB: p-p90rsk. p90rsk (Ser380) (arbitrary units) (Ser380) p90rsk. IB: p90rsk. Tubulin. IB: Tubulin. Ang II (200 nm) Ang II (200 nm)
 I: p-p9rsk I: p9rsk I: C I: p-p9rsk I: p9rsk 5 (ka) 5 5 (min) Ang II ( nm) p-p9rsk (Ser8) p9rsk p-p9rsk (Ser8) p9rsk (h) Mannitol 5 mm -Glucose 5 mm p9rsk (Ser8) (arbitrary units) p-p9rsk (Ser8) (arbitrary
I: p-p9rsk I: p9rsk I: C I: p-p9rsk I: p9rsk 5 (ka) 5 5 (min) Ang II ( nm) p-p9rsk (Ser8) p9rsk p-p9rsk (Ser8) p9rsk (h) Mannitol 5 mm -Glucose 5 mm p9rsk (Ser8) (arbitrary units) p-p9rsk (Ser8) (arbitrary
supplementary information
 DOI: 1.138/ncb1839 a b Control 1 2 3 Control 1 2 3 Fbw7 Smad3 1 2 3 4 1 2 3 4 c d IGF-1 IGF-1Rβ IGF-1Rβ-P Control / 1 2 3 4 Real-time RT-PCR Relative quantity (IGF-1/ mrna) 2 1 IGF-1 1 2 3 4 Control /
DOI: 1.138/ncb1839 a b Control 1 2 3 Control 1 2 3 Fbw7 Smad3 1 2 3 4 1 2 3 4 c d IGF-1 IGF-1Rβ IGF-1Rβ-P Control / 1 2 3 4 Real-time RT-PCR Relative quantity (IGF-1/ mrna) 2 1 IGF-1 1 2 3 4 Control /
Toll Receptor-Mediated Hippo Signaling Controls Innate Immunity in Drosophila
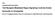 Cell Supplemental Information Toll Receptor-Mediated Hippo Signaling Controls Innate Immunity in Drosophila Bo Liu, Yonggang Zheng, Feng Yin, Jianzhong Yu, Neal Silverman, and Duojia Pan Supplemental Experimental
Cell Supplemental Information Toll Receptor-Mediated Hippo Signaling Controls Innate Immunity in Drosophila Bo Liu, Yonggang Zheng, Feng Yin, Jianzhong Yu, Neal Silverman, and Duojia Pan Supplemental Experimental
To isolate single GNS 144 cell clones, cells were plated at a density of 1cell/well
 Supplemental Information: Supplemental Methods: Cell culture To isolate single GNS 144 cell clones, cells were plated at a density of 1cell/well in 96 well Primaria plates in GNS media and incubated at
Supplemental Information: Supplemental Methods: Cell culture To isolate single GNS 144 cell clones, cells were plated at a density of 1cell/well in 96 well Primaria plates in GNS media and incubated at
Supplementary Information
 Supplementary Information MLL histone methylases regulate expression of HDLR- in presence of estrogen and control plasma cholesterol in vivo Khairul I. Ansari 1, Sahba Kasiri 1, Imran Hussain 1, Samara
Supplementary Information MLL histone methylases regulate expression of HDLR- in presence of estrogen and control plasma cholesterol in vivo Khairul I. Ansari 1, Sahba Kasiri 1, Imran Hussain 1, Samara
X2-C/X1-Y X2-C/VCAM-Y. FRET efficiency. Ratio YFP/CFP
 FRET efficiency.7.6..4.3.2 X2-C/X1-Y X2-C/VCAM-Y.1 1 2 3 Ratio YFP/CFP Supplemental Data 1. Analysis of / heterodimers in live cells using FRET. FRET saturation curves were obtained using cells transiently
FRET efficiency.7.6..4.3.2 X2-C/X1-Y X2-C/VCAM-Y.1 1 2 3 Ratio YFP/CFP Supplemental Data 1. Analysis of / heterodimers in live cells using FRET. FRET saturation curves were obtained using cells transiently
Nature Methods: doi: /nmeth Supplementary Figure 1. Validation of RaPID with EDEN15
 Supplementary Figure 1 Validation of RaPID with EDEN15 (a) Full Western Blot of conventional biotinylated RNA pulldown with EDEN15 and scrambled control (n=3 biologically independent experiments, representative
Supplementary Figure 1 Validation of RaPID with EDEN15 (a) Full Western Blot of conventional biotinylated RNA pulldown with EDEN15 and scrambled control (n=3 biologically independent experiments, representative
The Human Protein PRR14 Tethers Heterochromatin to the Nuclear Lamina During Interphase and Mitotic Exit
 Cell Reports, Volume 5 Supplemental Information The Human Protein PRR14 Tethers Heterochromatin to the Nuclear Lamina During Interphase and Mitotic Exit Andrey Poleshko, Katelyn M. Mansfield, Caroline
Cell Reports, Volume 5 Supplemental Information The Human Protein PRR14 Tethers Heterochromatin to the Nuclear Lamina During Interphase and Mitotic Exit Andrey Poleshko, Katelyn M. Mansfield, Caroline
sirna Transfection Reagent
 Description RiboJuice 0.3 ml 71115-3 1.0 ml 71115-4 Description RiboJuice efficiently delivers small interfering RNA (sirna) into a wide range of mammalian cell lines for targeted gene suppression (1).
Description RiboJuice 0.3 ml 71115-3 1.0 ml 71115-4 Description RiboJuice efficiently delivers small interfering RNA (sirna) into a wide range of mammalian cell lines for targeted gene suppression (1).
mir-24-mediated down-regulation of H2AX suppresses DNA repair
 Supplemental Online Material mir-24-mediated down-regulation of H2AX suppresses DNA repair in terminally differentiated blood cells Ashish Lal 1,4, Yunfeng Pan 2,4, Francisco Navarro 1,4, Derek M. Dykxhoorn
Supplemental Online Material mir-24-mediated down-regulation of H2AX suppresses DNA repair in terminally differentiated blood cells Ashish Lal 1,4, Yunfeng Pan 2,4, Francisco Navarro 1,4, Derek M. Dykxhoorn
Supplementary information
 Supplementary information Inhibition of mitochondrial dysfunction and neuronal cell death in models of Huntington s disease and in HD patients-derived cells Xing Guo 1,#, Marie-Helene Disatnik 3,#, Marie
Supplementary information Inhibition of mitochondrial dysfunction and neuronal cell death in models of Huntington s disease and in HD patients-derived cells Xing Guo 1,#, Marie-Helene Disatnik 3,#, Marie
Supporting Information: Core-Shell Nanoparticle-Based Peptide Therapeutics and Combined. Hyperthermia for Enhanced Cancer Cell Apoptosis
 Supporting Information: Core-Shell Nanoparticle-Based Peptide Therapeutics and Combined Hyperthermia for Enhanced Cancer Cell Apoptosis Birju P. Shah a, Nicholas Pasquale a, Gejing De b, Tao Tan b, Jianjie
Supporting Information: Core-Shell Nanoparticle-Based Peptide Therapeutics and Combined Hyperthermia for Enhanced Cancer Cell Apoptosis Birju P. Shah a, Nicholas Pasquale a, Gejing De b, Tao Tan b, Jianjie
Supplementary Figure 1 Characterization of sirna-onv stability. (a) Fluorescence recovery curves of SQ-siRNA-ONV and SQ-ds-siRNA in 1 TAMg buffer
 Supplementary Figure 1 Characterization of sirna-onv stability. (a) Fluorescence recovery curves of SQ-siRNA-ONV and SQ-ds-siRNA in 1 TAMg buffer containing 10% serum The data error bars indicate means
Supplementary Figure 1 Characterization of sirna-onv stability. (a) Fluorescence recovery curves of SQ-siRNA-ONV and SQ-ds-siRNA in 1 TAMg buffer containing 10% serum The data error bars indicate means
Multiplex Fluorescence Assays for Adherence Cells without Trypsinization
 Multiplex Fluorescence Assays for Adherence Cells without Trypsinization The combination of a bright field and three fluorescent channels allows the Celigo to perform many multiplexed assays. A gating
Multiplex Fluorescence Assays for Adherence Cells without Trypsinization The combination of a bright field and three fluorescent channels allows the Celigo to perform many multiplexed assays. A gating
The MAP Kinase Family
 The MAP Kinase Family Extracellular stimuli Classical MAP kinases Atypical MAP kinases MAPKKK MLK1/2/3/7; LZK RAF-1/A/B TAK1; TPL2 c-mos MEKK1-4; DLK ASK1/2; MLTK TAO1/2 ASK1 TAK1 MEKK1-4 MEKK2/3 TPL2???
The MAP Kinase Family Extracellular stimuli Classical MAP kinases Atypical MAP kinases MAPKKK MLK1/2/3/7; LZK RAF-1/A/B TAK1; TPL2 c-mos MEKK1-4; DLK ASK1/2; MLTK TAO1/2 ASK1 TAK1 MEKK1-4 MEKK2/3 TPL2???
Real-time PCR. TaqMan Protein Assays. Unlock the power of real-time PCR for protein analysis
 Real-time PCR TaqMan Protein Assays Unlock the power of real-time PCR for protein analysis I can use my real-time PCR instrument to quantitate protein? Do protein levels correlate with related mrna levels
Real-time PCR TaqMan Protein Assays Unlock the power of real-time PCR for protein analysis I can use my real-time PCR instrument to quantitate protein? Do protein levels correlate with related mrna levels
Plasmid DNA transfection of SW480 human colorectal cancer cells with the Biontex K2 Transfection System
 Plasmid DNA transfection of human colorectal cancer cells with the Biontex K2 Transfection System Stephanie Hehlgans and Franz Rödel, Department of Radiotherapy and Oncology, Goethe- University Frankfurt,
Plasmid DNA transfection of human colorectal cancer cells with the Biontex K2 Transfection System Stephanie Hehlgans and Franz Rödel, Department of Radiotherapy and Oncology, Goethe- University Frankfurt,
Supplementary Figure 1. Isolation of GFPHigh cells.
 Supplementary Figure 1. Isolation of GFP High cells. (A) Schematic diagram of cell isolation based on Wnt signaling activity. Colorectal cancer (CRC) cell lines were stably transduced with lentivirus encoding
Supplementary Figure 1. Isolation of GFP High cells. (A) Schematic diagram of cell isolation based on Wnt signaling activity. Colorectal cancer (CRC) cell lines were stably transduced with lentivirus encoding
SureSilencing sirna Array Technology Overview
 SureSilencing sirna Array Technology Overview Pathway-Focused sirna-based RNA Interference Topics to be Covered Who is SuperArray? Brief Introduction to RNA Interference Challenges Facing RNA Interference
SureSilencing sirna Array Technology Overview Pathway-Focused sirna-based RNA Interference Topics to be Covered Who is SuperArray? Brief Introduction to RNA Interference Challenges Facing RNA Interference
Supplementary Fig. S1. SAMHD1c has a more potent dntpase activity than. SAMHD1c. Purified recombinant SAMHD1c and SAMHD1c proteins (with
 Supplementary Fig. S1. SAMHD1c has a more potent dntpase activity than SAMHD1c. Purified recombinant SAMHD1c and SAMHD1c proteins (with concentration of 800nM) were incubated with 1mM dgtp for the indicated
Supplementary Fig. S1. SAMHD1c has a more potent dntpase activity than SAMHD1c. Purified recombinant SAMHD1c and SAMHD1c proteins (with concentration of 800nM) were incubated with 1mM dgtp for the indicated
Supplementary Figure 1. Characterization of EVs (a) Phase-contrast electron microscopy was used to visualize resuspended EV pellets.
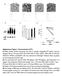 Supplementary Figure 1. Characterization of EVs (a) Phase-contrast electron microscopy was used to visualize resuspended EV pellets. Scale bar represent 100 nm. The sizes of EVs from MDA-MB-231-D3H1 (D3H1),
Supplementary Figure 1. Characterization of EVs (a) Phase-contrast electron microscopy was used to visualize resuspended EV pellets. Scale bar represent 100 nm. The sizes of EVs from MDA-MB-231-D3H1 (D3H1),
Ingenio Electroporation Kits and Solution
 Protocol for MIR 08, 09, 01, 0111, 0112, 0113, 0114, 011, 0116, 0117, 0118, 0119 Quick Reference Protocol, SDS and Certificate of Analysis available at mirusbio.com/0111 INTRODUCTION Ingenio Electroporation
Protocol for MIR 08, 09, 01, 0111, 0112, 0113, 0114, 011, 0116, 0117, 0118, 0119 Quick Reference Protocol, SDS and Certificate of Analysis available at mirusbio.com/0111 INTRODUCTION Ingenio Electroporation
Isolation, culture, and transfection of primary mammary epithelial organoids
 Supplementary Experimental Procedures Isolation, culture, and transfection of primary mammary epithelial organoids Primary mammary epithelial organoids were prepared from 8-week-old CD1 mice (Charles River)
Supplementary Experimental Procedures Isolation, culture, and transfection of primary mammary epithelial organoids Primary mammary epithelial organoids were prepared from 8-week-old CD1 mice (Charles River)
Supplementary Figure 1. (a) The qrt-pcr for lnc-2, lnc-6 and lnc-7 RNA level in DU145, 22Rv1, wild type HCT116 and HCT116 Dicer ex5 cells transfected
 Supplementary Figure 1. (a) The qrt-pcr for lnc-2, lnc-6 and lnc-7 RNA level in DU145, 22Rv1, wild type HCT116 and HCT116 Dicer ex5 cells transfected with the sirna against lnc-2, lnc-6, lnc-7, and the
Supplementary Figure 1. (a) The qrt-pcr for lnc-2, lnc-6 and lnc-7 RNA level in DU145, 22Rv1, wild type HCT116 and HCT116 Dicer ex5 cells transfected with the sirna against lnc-2, lnc-6, lnc-7, and the
TOOLS sirna and mirna. User guide
 TOOLS sirna and mirna User guide Introduction RNA interference (RNAi) is a powerful tool for suppression gene expression by causing the destruction of specific mrna molecules. Small Interfering RNAs (sirnas)
TOOLS sirna and mirna User guide Introduction RNA interference (RNAi) is a powerful tool for suppression gene expression by causing the destruction of specific mrna molecules. Small Interfering RNAs (sirnas)
Thyroid peroxidase gene expression is induced by lipopolysaccharide involving Nuclear Factor (NF)-κB p65 subunit phosphorylation
 1 2 3 4 5 SUPPLEMENTAL DATA Thyroid peroxidase gene expression is induced by lipopolysaccharide involving Nuclear Factor (NF)-κB p65 subunit phosphorylation Magalí Nazar, Juan Pablo Nicola, María Laura
1 2 3 4 5 SUPPLEMENTAL DATA Thyroid peroxidase gene expression is induced by lipopolysaccharide involving Nuclear Factor (NF)-κB p65 subunit phosphorylation Magalí Nazar, Juan Pablo Nicola, María Laura
Forced Expression of Heat Shock Protein 27 (Hsp27) Reverses P-Glycoprotein (ABCB1)- mediated Drug Efflux and MDR1 Gene Expression in
 Forced Expression of Heat Shock Protein 27 (Hsp27) Reverses P-Glycoprotein (ABCB1)- mediated Drug Efflux and MDR1 Gene Expression in Adriamycin-resistant Human Breast Cancer Cells Authors: Ragu Kanagasabai,
Forced Expression of Heat Shock Protein 27 (Hsp27) Reverses P-Glycoprotein (ABCB1)- mediated Drug Efflux and MDR1 Gene Expression in Adriamycin-resistant Human Breast Cancer Cells Authors: Ragu Kanagasabai,
Supplemental Information. HEXIM1 and NEAT1 Long Non-coding RNA Form. a Multi-subunit Complex that Regulates. DNA-Mediated Innate Immune Response
 Molecular Cell, Volume 67 Supplemental Information HEXIM1 and NEAT1 Long Non-coding RNA Form a Multi-subunit Complex that Regulates DNA-Mediated Innate Immune Response Mehdi Morchikh, Alexandra Cribier,
Molecular Cell, Volume 67 Supplemental Information HEXIM1 and NEAT1 Long Non-coding RNA Form a Multi-subunit Complex that Regulates DNA-Mediated Innate Immune Response Mehdi Morchikh, Alexandra Cribier,
Supporting Online Material for
 www.sciencemag.org/cgi/content/full/323/5910/124/dc1 Supporting Online Material for Regulation of Neuronal Survival Factor MEF2D by Chaperone-Mediated Autophagy Qian Yang, Hua She, Marla Gearing, Emanuela
www.sciencemag.org/cgi/content/full/323/5910/124/dc1 Supporting Online Material for Regulation of Neuronal Survival Factor MEF2D by Chaperone-Mediated Autophagy Qian Yang, Hua She, Marla Gearing, Emanuela
Product: Arrest-In TM Transfection Reagent for RNAi
 Product: Arrest-In TM Transfection Reagent for RNAi Catalog #: ATR1740, ATR1741, ATR1742, ATR1743 Product Description Arrest-In transfection reagent is a proprietary polymeric formulation, developed and
Product: Arrest-In TM Transfection Reagent for RNAi Catalog #: ATR1740, ATR1741, ATR1742, ATR1743 Product Description Arrest-In transfection reagent is a proprietary polymeric formulation, developed and
Tandem E2F Binding Sites in the Promoter of the p107 Cell Cycle Regulator Control p107 Expression and Its Cellular Functions
 Tandem E2F Binding Sites in the Promoter of the p107 Cell Cycle Regulator Control p107 Expression and Its Cellular Functions Deborah L. Burkhart 1,2, Stacey E. Wirt 1,2, Anne-Flore Zmoos 1, Michael S.
Tandem E2F Binding Sites in the Promoter of the p107 Cell Cycle Regulator Control p107 Expression and Its Cellular Functions Deborah L. Burkhart 1,2, Stacey E. Wirt 1,2, Anne-Flore Zmoos 1, Michael S.
NTM486-04, NTM174-04,
 Transfection of transformed human trabecular meshwork TM5, and primary human NTM210-05, NTM486-04, NTM174-04, and NTM153-00 cells with Metafectene Easy Adnan Dibas1A,C, Ming Jiang1A,C, Thomas Yorio1A,C.
Transfection of transformed human trabecular meshwork TM5, and primary human NTM210-05, NTM486-04, NTM174-04, and NTM153-00 cells with Metafectene Easy Adnan Dibas1A,C, Ming Jiang1A,C, Thomas Yorio1A,C.
from Dr. David Livingston. Rabbit anti-myc, V5, and BACH1 were raised by immunizing rabbits with peptides EQKLISEEDI, GKPIPNPLLGLDST, and
 upporting Material Experimental Procedures: Cell culture and antibodies All cell lines were maintained in RPMI 164 medium with 1% fetal calf serum at 37 C in 5% CO 2 (v/v). For HCC 1937-BRCA1 cells and
upporting Material Experimental Procedures: Cell culture and antibodies All cell lines were maintained in RPMI 164 medium with 1% fetal calf serum at 37 C in 5% CO 2 (v/v). For HCC 1937-BRCA1 cells and
Pre-made Lentiviral Particles for nuclear permeant CRE recombinase expression
 Pre-made Lentiviral Particles for nuclear permeant CRE recombinase expression Cat# Product Name Amounts* LVP336 NLS-CRE (Bsd) LVP336-PBS NLS-CRE (Bsd), in vivo ready LVP339 NLS-CRE (Puro) LVP339-PBS NLS-CRE
Pre-made Lentiviral Particles for nuclear permeant CRE recombinase expression Cat# Product Name Amounts* LVP336 NLS-CRE (Bsd) LVP336-PBS NLS-CRE (Bsd), in vivo ready LVP339 NLS-CRE (Puro) LVP339-PBS NLS-CRE
Intestinal Epithelial Cell-Specific Deletion of PLD2 Alleviates DSS-Induced Colitis by. Regulating Occludin
 Intestinal Epithelial Cell-Specific Deletion of PLD2 Alleviates DSS-Induced Colitis by Regulating Occludin Chaithanya Chelakkot 1,ǂ, Jaewang Ghim 2,3,ǂ, Nirmal Rajasekaran 4, Jong-Sun Choi 5, Jung-Hwan
Intestinal Epithelial Cell-Specific Deletion of PLD2 Alleviates DSS-Induced Colitis by Regulating Occludin Chaithanya Chelakkot 1,ǂ, Jaewang Ghim 2,3,ǂ, Nirmal Rajasekaran 4, Jong-Sun Choi 5, Jung-Hwan
TransIT-TKO Transfection Reagent
 Quick Reference Protocol, MSDS and Certificate of Analysis available at mirusbio.com/2150 INTRODUCTION TransIT-TKO is a broad spectrum sirna transfection reagent that enables high efficiency sirna delivery
Quick Reference Protocol, MSDS and Certificate of Analysis available at mirusbio.com/2150 INTRODUCTION TransIT-TKO is a broad spectrum sirna transfection reagent that enables high efficiency sirna delivery
Learning Objectives. Define RNA interference. Define basic terminology. Describe molecular mechanism. Define VSP and relevance
 Learning Objectives Define RNA interference Define basic terminology Describe molecular mechanism Define VSP and relevance Describe role of RNAi in antigenic variation A Nobel Way to Regulate Gene Expression
Learning Objectives Define RNA interference Define basic terminology Describe molecular mechanism Define VSP and relevance Describe role of RNAi in antigenic variation A Nobel Way to Regulate Gene Expression
Supplemental figures Supplemental Figure 1: Fluorescence recovery for FRAP experiments depicted in Figure 1.
 Supplemental figures Supplemental Figure 1: Fluorescence recovery for FRAP experiments depicted in Figure 1. Percent of original fluorescence was plotted as a function of time following photobleaching
Supplemental figures Supplemental Figure 1: Fluorescence recovery for FRAP experiments depicted in Figure 1. Percent of original fluorescence was plotted as a function of time following photobleaching
Supporting Information
 Supporting Information Horie et al. 10.1073/pnas.1008499107 SI Materials and Methods ell ulture and Reagents. THP-1 cells were obtained from the American Type ell ollection. THP-1 cells were transformed
Supporting Information Horie et al. 10.1073/pnas.1008499107 SI Materials and Methods ell ulture and Reagents. THP-1 cells were obtained from the American Type ell ollection. THP-1 cells were transformed
Wnt16 smact merge VK/AB
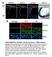 A WT Wnt6 smact merge VK/A KO ctrl IgG WT KO Wnt6 smact DAPI SUPPLEMENTAL FIGURE I: Wnt6 expression in MGP-deficient aortae. Immunostaining for Wnt6 and smooth muscle actin (smact) in aortae from 7 day
A WT Wnt6 smact merge VK/A KO ctrl IgG WT KO Wnt6 smact DAPI SUPPLEMENTAL FIGURE I: Wnt6 expression in MGP-deficient aortae. Immunostaining for Wnt6 and smooth muscle actin (smact) in aortae from 7 day
PROTEOMICS AND FUNCTIONAL GENOMICS PRESENTATION KIMBERLY DONG
 PROTEOMICS AND FUNCTIONAL GENOMICS PRESENTATION KIMBERLY DONG A Lentiviral RNAi Library for Human and Mouse Genes Applied to an Arrayed Viral High-Content Screen Jason Moffat,1,2,4,10 Dorre A. Grueneberg,1,10
PROTEOMICS AND FUNCTIONAL GENOMICS PRESENTATION KIMBERLY DONG A Lentiviral RNAi Library for Human and Mouse Genes Applied to an Arrayed Viral High-Content Screen Jason Moffat,1,2,4,10 Dorre A. Grueneberg,1,10
Chemically defined conditions for human ipsc derivation and culture
 Nature Methods Chemically defined conditions for human ipsc derivation and culture Guokai Chen, Daniel R Gulbranson, Zhonggang Hou, Jennifer M Bolin, Victor Ruotti, Mitchell D Probasco, Kimberly Smuga-Otto,
Nature Methods Chemically defined conditions for human ipsc derivation and culture Guokai Chen, Daniel R Gulbranson, Zhonggang Hou, Jennifer M Bolin, Victor Ruotti, Mitchell D Probasco, Kimberly Smuga-Otto,
Thermo Scientific Dharmacon SMARTvector 2.0 Lentiviral shrna Particles
 Thermo Scientific Dharmacon SMARTvector 2.0 Lentiviral shrna Particles Long-term gene silencing shrna-specific design algorithm High titer, purified particles Thermo Scientific Dharmacon SMARTvector shrna
Thermo Scientific Dharmacon SMARTvector 2.0 Lentiviral shrna Particles Long-term gene silencing shrna-specific design algorithm High titer, purified particles Thermo Scientific Dharmacon SMARTvector shrna
Supplementary Figure 1. RAD51 and RAD51 paralogs are enriched spontaneously onto
 Supplementary Figure legends Supplementary Figure 1. and paralogs are enriched spontaneously onto the S-phase chromatin during DN replication. () Chromatin fractionation was carried out as described in
Supplementary Figure legends Supplementary Figure 1. and paralogs are enriched spontaneously onto the S-phase chromatin during DN replication. () Chromatin fractionation was carried out as described in
SUPPLEMENTARY MATERIAL
 SUPPLEMENTARY MATERIAL Materials and Methods Circular dichroism (CD) spectroscopy. Far ultraviolet (UV) CD spectra of apo- and holo- CaM and the CaM mutants were recorded on a Jasco J-715 spectropolarimeter
SUPPLEMENTARY MATERIAL Materials and Methods Circular dichroism (CD) spectroscopy. Far ultraviolet (UV) CD spectra of apo- and holo- CaM and the CaM mutants were recorded on a Jasco J-715 spectropolarimeter
Supporting information. Single-cell and subcellular pharmacokinetic imaging allows insight into drug action in vivo
 Supporting information Single-cell and subcellular pharmacokinetic imaging allows insight into drug action in vivo Greg Thurber 1, Katy Yang 1, Thomas Reiner 1, Rainer Kohler 1, Peter Sorger 2, Tim Mitchison
Supporting information Single-cell and subcellular pharmacokinetic imaging allows insight into drug action in vivo Greg Thurber 1, Katy Yang 1, Thomas Reiner 1, Rainer Kohler 1, Peter Sorger 2, Tim Mitchison
Fig. S1. Nature Medicine: doi: /nm HoxA9 expression levels BM MOZ-TIF2 AML BM. Sca-1-H. c-kit-h CSF1R-H CD16/32-H. Mac1-H.
 A 1 4 1 4 1 4 CSF1RH 1 3 1 2 1 1 CSF1RH 1 3 1 2 1 1 CSF1RH 1 3 1 2 1 1 1 1 1 1 1 2 1 3 1 4 GFPH 1 1 1 1 1 2 1 3 1 4 Sca1H 1 1 1 1 1 2 1 3 1 4 ckith 1 4 1 4 1 4 CSF1RH 1 3 1 2 1 1 CSF1RH 1 3 1 2 1 1 CSF1RH
A 1 4 1 4 1 4 CSF1RH 1 3 1 2 1 1 CSF1RH 1 3 1 2 1 1 CSF1RH 1 3 1 2 1 1 1 1 1 1 1 2 1 3 1 4 GFPH 1 1 1 1 1 2 1 3 1 4 Sca1H 1 1 1 1 1 2 1 3 1 4 ckith 1 4 1 4 1 4 CSF1RH 1 3 1 2 1 1 CSF1RH 1 3 1 2 1 1 CSF1RH
Kinetics Review. Tonight at 7 PM Phys 204 We will do two problems on the board (additional ones than in the problem sets)
 Quiz 1 Kinetics Review Tonight at 7 PM Phys 204 We will do two problems on the board (additional ones than in the problem sets) I will post the problems with solutions on Toolkit for those that can t make
Quiz 1 Kinetics Review Tonight at 7 PM Phys 204 We will do two problems on the board (additional ones than in the problem sets) I will post the problems with solutions on Toolkit for those that can t make
Bronchial epithelium and its associated tissues act as a
 The Journal of Immunology A JNK-Independent Signaling Pathway Regulates TNF -Stimulated, c-jun-driven FRA-1 Protooncogene Transcription in Pulmonary Epithelial Cells 1 Pavan Adiseshaiah,* Dhananjaya V.
The Journal of Immunology A JNK-Independent Signaling Pathway Regulates TNF -Stimulated, c-jun-driven FRA-1 Protooncogene Transcription in Pulmonary Epithelial Cells 1 Pavan Adiseshaiah,* Dhananjaya V.
CRE/CREB Reporter Assay Kit camp/pka Cell Signaling Pathway Catalog #: 60611
 Data Sheet CRE/CREB Reporter Assay Kit camp/pka Cell Signaling Pathway Catalog #: 60611 Background The main role of the camp response element, or CRE, is mediating the effects of Protein Kinase A (PKA)
Data Sheet CRE/CREB Reporter Assay Kit camp/pka Cell Signaling Pathway Catalog #: 60611 Background The main role of the camp response element, or CRE, is mediating the effects of Protein Kinase A (PKA)
7.06 Problem Set #3, Spring 2005
 7.06 Problem Set #3, Spring 2005 1. The Drosophila compound eye is composed of about 800 units called ommatidia. Each ommatidium contains eight photoreceptor neurons (R1 through R8), which develop in a
7.06 Problem Set #3, Spring 2005 1. The Drosophila compound eye is composed of about 800 units called ommatidia. Each ommatidium contains eight photoreceptor neurons (R1 through R8), which develop in a
Small-Molecule Drug Target Identification/Deconvolution Technologies
 Small-Molecule Drug Target Identification/Deconvolution Technologies Case-Studies Shantani Target ID Technology Tool Box Target Deconvolution is not Trivial = A single Tool / Technology May Not necessarily
Small-Molecule Drug Target Identification/Deconvolution Technologies Case-Studies Shantani Target ID Technology Tool Box Target Deconvolution is not Trivial = A single Tool / Technology May Not necessarily
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/3/6/ra27/dc Supplementary Materials for AAA+ Proteins and Coordinate PIKK Activity and Function in Nonsense-Mediated mrna Decay Natsuko Izumi, Akio Yamashita,*
www.sciencesignaling.org/cgi/content/full/3/6/ra27/dc Supplementary Materials for AAA+ Proteins and Coordinate PIKK Activity and Function in Nonsense-Mediated mrna Decay Natsuko Izumi, Akio Yamashita,*
supplementary information
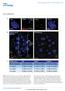 DOI: 10.1038/ncb2017 Figure S1 53BP1 sirna results in a heterochromatic DSB repair defect. Panel A: NIH3T3 cells were transfected with murine 53BP1 sirna. 48 hrs later, cells were irradiated with 3 Gy
DOI: 10.1038/ncb2017 Figure S1 53BP1 sirna results in a heterochromatic DSB repair defect. Panel A: NIH3T3 cells were transfected with murine 53BP1 sirna. 48 hrs later, cells were irradiated with 3 Gy
Supplementary Figure 1
 Supplementary Figure A Name Location Start position End position Oct4P Chr 3 29433673 29435 Annotation Mus musculus predicted gene 572 (Gm572), noncoding RNA RefSeq accession number NR_33594. Oct4P2 Chr
Supplementary Figure A Name Location Start position End position Oct4P Chr 3 29433673 29435 Annotation Mus musculus predicted gene 572 (Gm572), noncoding RNA RefSeq accession number NR_33594. Oct4P2 Chr
Supplemental Information
 Supplemental Information Genetic and Functional Studies Implicate HIF1α as a 14q Kidney Cancer Suppressor Gene Chuan Shen, Rameen Beroukhim, Steven E. Schumacher, Jing Zhou, Michelle Chang, Sabina Signoretti,
Supplemental Information Genetic and Functional Studies Implicate HIF1α as a 14q Kidney Cancer Suppressor Gene Chuan Shen, Rameen Beroukhim, Steven E. Schumacher, Jing Zhou, Michelle Chang, Sabina Signoretti,
Description: Nuclear morphology and dynamics in nontargeting sirna transfected cells. HeLa Kyoto
 Title of file for HTML: Supplementary Information Description: Supplementary Figures and Supplementary Tables Title of file for HTML: Supplementary Movie 1 Description: Nuclear morphology and dynamics
Title of file for HTML: Supplementary Information Description: Supplementary Figures and Supplementary Tables Title of file for HTML: Supplementary Movie 1 Description: Nuclear morphology and dynamics
Supporting Information
 Supporting Information Cieslewicz et al. 10.1073/pnas.1312197110 SI Results Human and mouse lesions of atherosclerosis contain both M1 and M2 macrophage phenotypes (1, 2). Previous work has suggested the
Supporting Information Cieslewicz et al. 10.1073/pnas.1312197110 SI Results Human and mouse lesions of atherosclerosis contain both M1 and M2 macrophage phenotypes (1, 2). Previous work has suggested the
sirna Overview and Technical Tips
 1 sirna Overview and Technical Tips 2 CONTENTS 3 4 5 7 8 10 11 13 14 18 19 20 21 Introduction Applications How Does It Work? Handy Tips Troubleshooting Conclusions Further References Contact Us 3 INTRODUCTION
1 sirna Overview and Technical Tips 2 CONTENTS 3 4 5 7 8 10 11 13 14 18 19 20 21 Introduction Applications How Does It Work? Handy Tips Troubleshooting Conclusions Further References Contact Us 3 INTRODUCTION
1,500 1,000. LPS + alum. * * Casp1 p10. Casp1 p45
 a NLRP3 Non-stimulation R46 BAY SI TAT LPS R46 BAY SI TAT 1,5 1, c 15 1 5 5 Pro-IL-18 Pro-IL-1 LPS + alum d e f IL-1 p17 Pro-IL-1 1 75 5 5 Casp1 p1 NLRP3 LPS + alum Supplementary Figure 1 Inhiition of
a NLRP3 Non-stimulation R46 BAY SI TAT LPS R46 BAY SI TAT 1,5 1, c 15 1 5 5 Pro-IL-18 Pro-IL-1 LPS + alum d e f IL-1 p17 Pro-IL-1 1 75 5 5 Casp1 p1 NLRP3 LPS + alum Supplementary Figure 1 Inhiition of
- NaCr. + NaCr. α H3K4me2 α H3K4me3 α H3K9me3 α H3K27me3 α H3K36me3 H3 H2A-2B H4 H3 H2A-2B H4 H3 H2A-2B H4. α Kcr. (rabbit) α Kac.
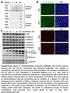 + NaCr NaCr + NaCr NaCr Peptides 10ng 50ng 250ng K α Pan (mouse) Pan (mouse) 10ng 50ng 250ng α Pan (rabbit) C 10ng 50ng 250ng α Pan (mouse) 0 1.25 2.5 5 10 20 40 (mm) NaCr 24h α Pan (rabbit) α K4me2 α
+ NaCr NaCr + NaCr NaCr Peptides 10ng 50ng 250ng K α Pan (mouse) Pan (mouse) 10ng 50ng 250ng α Pan (rabbit) C 10ng 50ng 250ng α Pan (mouse) 0 1.25 2.5 5 10 20 40 (mm) NaCr 24h α Pan (rabbit) α K4me2 α
