SUPPLEMENTARY INFORMATION
|
|
|
- Elizabeth Wilkins
- 6 years ago
- Views:
Transcription
1 doi: /nature09861 & &' -(' ()*+ ')(+,,(','-*+,&,,+ ',+' ' 23,45/0*6787*9:./09 &' )*+*(,-* -(' ()*+ ')(+,,(','-*+,&,,+ ',+'./)*+*(,-*..)*+*(,-*./)*+*(,-*.0)*+*(,-*..)*+*(,-* )*+*(,-* )*+1*(,-*,C3(7*D,C3(7*E,C3(7*2 & '( Supplementary Figure 1. ONSEN retrotransposon family in Arabidopsis Columbia accession. a, Chromosomal location of eight ONSEN copies (open triangles with gene IDs according to TAIR8, chromosome numbers are given on the left, centromeres are marked by open circles. b, Structures of eight ONSEN copies; predicted ORFs larger than 500 base pairs (bp) marked as grey arrows, LTRs as blue arrows a noncoding regions are marked in white. Four ONSEN copies encode a putative polyprotein (AT1G11265, AT1G48710, AT3G61330, AT3G13205). The predicted reverse-transcriptase domain ( PF07727 domain) is iicated above as RVT_2. The scale below is in kilobase pairs (kb). PsiI iicated was used to digest DNA for Southern blots on Figure 2a a Supplementary Figure 3b. The location of ONSEN specific probes (A, B a C) used for hybridizations are iicated. Note that probe B was designed from the AT1G11265 locus. 1
2 '()*+,-./ 012( 01'( 01 & '()*& +,-'.& +,-'&,',& & & & &' 3456 Supplementary Figure 2. Northern blot of RNA isolated in wild type plants a mutants 10 days after HS. WT CS a WT HS are shown as negative a positive controls, respectively. RNA size correspoing to the full-length transposon transcript is marked by the arrow on the right. EtBr-stained gel is shown below as a loading control. ONSEN specific probe A (Supplementary Fig. 1b) was used for hybridization. 2
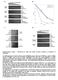
3 1& /, & /, & &/, )*/, )* 0 '( )* +, )* /, -. +,. doi: /nature /, & &+,-. &/,-. '()*(&9:;<-=>(? Supplementary Figure 3. ONSEN transposition assays. a, Transposon display performed on DNA extracted from wild-type a nrpd1 mutant plants 40 days after HS. nrprd1 HS 1 to 3 represent three iepeent HS-treated nrpd1 plants. ONSEN specific primer Copia78 3 LTR was used (see Supplementary Table 2). b, Southern blot of DNA extracted from the progeny (seco generation, 2 ) of CS- or HS-treated plants. DNA was digested with PsiI a hybridized with probe C (Supplementary Fig. 1). Six nrpd1 HS 2 plants are progeny siblings of a single nrpd1 plant which was HS-treated as a 7-day-old seedling. EtBr-stained gel is shown below as a loading control.c, Quantification of ONSEN copy number increase in five nrpd1 HS 2 Five nrpd1 CS 2 plants. Error bars correspo to technical repetitions (mean ± s.e.m., n=3). plants are shown as controls. 3
4 doi: /nature09861 &' (' *+, &'() /0 -. Supplementary Figure 4. Transposon display assays with DNA isolated from progeny siblings (seco generation, 2 ) of CS- a HS-treated mutant plants. For each mutant five iividual plants (marked above), derived from a CS- or HS-treated mutant ancestor were used for DNA isolation (mutant identities are given on the right). ONSEN specific primer Copia78 3 LTR was used (see Supplementary Table 2). 4
5 &'&& &'& ''()& '',-(,'&,& (*& ',*+)&,'*+-& (,(& )+& ((,,&& Supplementary Figure 5. Chromosomal distribution of new ONSEN insertions. Black triangles iicate the new ONSEN insertion sites in the progeny of HS-treated nrpd1 plants. See also Supplementary Table 1. White circles on the chromosomes specify the location of the centromeres. 5
6 Supplementary Table 1. Sequences of new ONSEN insertion sites. Black triangles iicate the position of new ONSEN insertions in the progeny of HS-treated nrpd1 plants, as shown in Supplementary Figure 5. Flanking sequences were determined by transposon display followed by sequencing (see Supplementary Table 2 for primer information) a validated by PCR (data not shown). 6
7 Experiment Primer Sequence (5-3 ) At1g11265 full-f TAATGTTCCCTTCCAAGTCCC ONSEN probe A At1g11265 full-r TTTCAACACCCCCTCTTAAAC ONSEN LTR F GGTTGAAGGGTCAAAGAGTAAAT ONSEN probe B ONSEN LTR R CCTCCAAACTACAAAATATCTAAAA ONSEN probe C ONSEN RT-PCR ACTIN2 Copia1-F Copia mix-r Copia1-F Copia1-R qact2_f1 qact2_r1 ACT2_probe 18S F TCTCTGTCTAATCTATCAAG TGATCTCCATTCTTCAATCG TCTCTGTCTAATCTATCAAG AAGTTGCTCTATTGTCATAGC TGCCAATCTACGAGGGTTTC TTACAATTTCCCGCTCTGCT TCCGTCTTGACCTTGCTGGACG CGTCCCTGCCCTTTGTACAC 18S ONSEN qpcr Transposon display 18S R CGAACACTTCACCGGATCATT 18S probe CCGCCCGTCGCTCCTACCGAT COPIA F CCACAAGAGGAACCAACGAA COPIA R TTCGATCATGGAAGACCGG COPIA T (probe) AAGTCGGCAATAGCTTTGGCGAAGA GenWalkAdaptator 1 GTAATACGACTCACTATAGGGCACGCGTGGTC GACGGCCCGGGCTGGT GenWalkAdaptator 2 ACCAGCCC (AMINO 3 ) GenWalk_AP1 GTAATACGACTCACTATAGGGC Copia78 3 LTR AACACTTAAACACTTTCTCCA ONS_312_R GCCTCCAAACTACAAAATATCTAAA Supplementary Table 2. Primer sequences. 7
8 Plant Material Additional Arabidopsis mutant used in this study: kyp-7 1 Supplementary Reference 1. Mathieu, O., Probst, A.V. & Paszkowski, J. Distinct regulation of histone H3 methylation at lysines 27 a 9 by CpG methylation in Arabidopsis. EMBO J 24, (2005). 8
Construction of plant complementation vector and generation of transgenic plants
 MATERIAL S AND METHODS Plant materials and growth conditions Arabidopsis ecotype Columbia (Col0) was used for this study. SALK_072009, SALK_076309, and SALK_027645 were obtained from the Arabidopsis Biological
MATERIAL S AND METHODS Plant materials and growth conditions Arabidopsis ecotype Columbia (Col0) was used for this study. SALK_072009, SALK_076309, and SALK_027645 were obtained from the Arabidopsis Biological
Using mutants to clone genes
 Using mutants to clone genes Objectives 1. What is positional cloning? 2. What is insertional tagging? 3. How can one confirm that the gene cloned is the same one that is mutated to give the phenotype
Using mutants to clone genes Objectives 1. What is positional cloning? 2. What is insertional tagging? 3. How can one confirm that the gene cloned is the same one that is mutated to give the phenotype
Supplementary Figure 1 An overview of pirna biogenesis during fetal mouse reprogramming. (a) (b)
 Supplementary Figure 1 An overview of pirna biogenesis during fetal mouse reprogramming. (a) A schematic overview of the production and amplification of a single pirna from a transposon transcript. The
Supplementary Figure 1 An overview of pirna biogenesis during fetal mouse reprogramming. (a) A schematic overview of the production and amplification of a single pirna from a transposon transcript. The
Genomes summary. Bacterial genome sizes
 Genomes summary 1. >930 bacterial genomes sequenced. 2. Circular. Genes densely packed. 3. 2-10 Mbases, 470-7,000 genes 4. Genomes of >200 eukaryotes (45 higher ) sequenced. 5. Linear chromosomes 6. On
Genomes summary 1. >930 bacterial genomes sequenced. 2. Circular. Genes densely packed. 3. 2-10 Mbases, 470-7,000 genes 4. Genomes of >200 eukaryotes (45 higher ) sequenced. 5. Linear chromosomes 6. On
Supplementary Methods
 Supplementary Methods Reverse transcribed Quantitative PCR. Total RNA was isolated from bone marrow derived macrophages using RNeasy Mini Kit (Qiagen), DNase-treated (Promega RQ1), and reverse transcribed
Supplementary Methods Reverse transcribed Quantitative PCR. Total RNA was isolated from bone marrow derived macrophages using RNeasy Mini Kit (Qiagen), DNase-treated (Promega RQ1), and reverse transcribed
GENETICS EXAM 3 FALL a) is a technique that allows you to separate nucleic acids (DNA or RNA) by size.
 Student Name: All questions are worth 5 pts. each. GENETICS EXAM 3 FALL 2004 1. a) is a technique that allows you to separate nucleic acids (DNA or RNA) by size. b) Name one of the materials (of the two
Student Name: All questions are worth 5 pts. each. GENETICS EXAM 3 FALL 2004 1. a) is a technique that allows you to separate nucleic acids (DNA or RNA) by size. b) Name one of the materials (of the two
Supplementary Figure 1 qrt-pcr expression analysis of NLP8 with and without KNO 3 during germination.
 Supplementary Figure 1 qrt-pcr expression analysis of NLP8 with and without KNO 3 during germination. Seeds of Col-0 were harvested from plants grown at 16 C, stored for 2 months, imbibed for indicated
Supplementary Figure 1 qrt-pcr expression analysis of NLP8 with and without KNO 3 during germination. Seeds of Col-0 were harvested from plants grown at 16 C, stored for 2 months, imbibed for indicated
WiscDsLox485 ATG < > //----- E1 E2 E3 E4 E bp. Col-0 arr7 ARR7 ACTIN7. s of mrna/ng total RNA (x10 3 ) ARR7.
 A WiscDsLox8 ATG < > -9 +0 > ---------//----- E E E E E UTR +9 UTR +0 > 00bp B S D ol-0 arr7 ol-0 arr7 ARR7 ATIN7 D s of mrna/ng total RNA opie 0. ol-0 (x0 ) arr7 ARR7 Supplemental Fig.. Genotyping and
A WiscDsLox8 ATG < > -9 +0 > ---------//----- E E E E E UTR +9 UTR +0 > 00bp B S D ol-0 arr7 ol-0 arr7 ARR7 ATIN7 D s of mrna/ng total RNA opie 0. ol-0 (x0 ) arr7 ARR7 Supplemental Fig.. Genotyping and
Lecture 3 Mutagens and Mutagenesis. 1. Mutagens A. Physical and Chemical mutagens B. Transposons and retrotransposons C. T-DNA
 Lecture 3 Mutagens and Mutagenesis 1. Mutagens A. Physical and Chemical mutagens B. Transposons and retrotransposons C. T-DNA 2. Mutagenesis A. Screen B. Selection C. Lethal mutations Read: 508-514 Figs:
Lecture 3 Mutagens and Mutagenesis 1. Mutagens A. Physical and Chemical mutagens B. Transposons and retrotransposons C. T-DNA 2. Mutagenesis A. Screen B. Selection C. Lethal mutations Read: 508-514 Figs:
R1 12 kb R1 4 kb R1. R1 10 kb R1 2 kb R1 4 kb R1
 Bcor101 Sample questions Midterm 3 1. The maps of the sites for restriction enzyme EcoR1 (R1) in the wild type and mutated cystic fibrosis genes are shown below: Wild Type R1 12 kb R1 4 kb R1 _ _ CF probe
Bcor101 Sample questions Midterm 3 1. The maps of the sites for restriction enzyme EcoR1 (R1) in the wild type and mutated cystic fibrosis genes are shown below: Wild Type R1 12 kb R1 4 kb R1 _ _ CF probe
Supplementary Figures Montero et al._supplementary Figure 1
 Montero et al_suppl. Info 1 Supplementary Figures Montero et al._supplementary Figure 1 Montero et al_suppl. Info 2 Supplementary Figure 1. Transcripts arising from the structurally conserved subtelomeres
Montero et al_suppl. Info 1 Supplementary Figures Montero et al._supplementary Figure 1 Montero et al_suppl. Info 2 Supplementary Figure 1. Transcripts arising from the structurally conserved subtelomeres
% Viability. isw2 ino isw2 ino isw2 ino isw2 ino mM HU 4-NQO CPT
 a Drug concentration b 1.3% MMS nhp1 nhp1 8 nhp1 mag1.5% MMS.3% MMS nhp1 nhp1 ino8 9 ino8 9 % Viability 4.5% MMS ino8 9 ino8 9 2.5.1.15 % MMS c d nhp1 nhp1 nhp1 nhp1 nhp1 nhp1 Control (YPD) γ IR (1 gy)
a Drug concentration b 1.3% MMS nhp1 nhp1 8 nhp1 mag1.5% MMS.3% MMS nhp1 nhp1 ino8 9 ino8 9 % Viability 4.5% MMS ino8 9 ino8 9 2.5.1.15 % MMS c d nhp1 nhp1 nhp1 nhp1 nhp1 nhp1 Control (YPD) γ IR (1 gy)
SUPPLEMENTARY INFORMATION
 doi:.38/nature899 Supplementary Figure Suzuki et al. a c p7 -/- / WT ratio (+)/(-) p7 -/- / WT ratio Log X 3. Fold change by treatment ( (+)/(-)) Log X.5 3-3. -. b Fold change by treatment ( (+)/(-)) 8
doi:.38/nature899 Supplementary Figure Suzuki et al. a c p7 -/- / WT ratio (+)/(-) p7 -/- / WT ratio Log X 3. Fold change by treatment ( (+)/(-)) Log X.5 3-3. -. b Fold change by treatment ( (+)/(-)) 8
Enzyme that uses RNA as a template to synthesize a complementary DNA
 Biology 105: Introduction to Genetics PRACTICE FINAL EXAM 2006 Part I: Definitions Homology: Comparison of two or more protein or DNA sequence to ascertain similarities in sequences. If two genes have
Biology 105: Introduction to Genetics PRACTICE FINAL EXAM 2006 Part I: Definitions Homology: Comparison of two or more protein or DNA sequence to ascertain similarities in sequences. If two genes have
Jung-Nam Cho, Jee-Youn Ryu, Young-Min Jeong, Jihye Park, Ji-Joon Song, Richard M. Amasino, Bosl Noh, and Yoo-Sun Noh
 Developmental Cell, Volume 22 Supplemental Information Control of Seed Germination by Light-Induced Histone Arginine Demethylation Activity Jung-Nam Cho, Jee-Youn Ryu, Young-Min Jeong, Jihye Park, Ji-Joon
Developmental Cell, Volume 22 Supplemental Information Control of Seed Germination by Light-Induced Histone Arginine Demethylation Activity Jung-Nam Cho, Jee-Youn Ryu, Young-Min Jeong, Jihye Park, Ji-Joon
Genetics - Problem Drill 19: Dissection of Gene Function: Mutational Analysis of Model Organisms
 Genetics - Problem Drill 19: Dissection of Gene Function: Mutational Analysis of Model Organisms No. 1 of 10 1. The mouse gene knockout is based on. (A) Homologous recombination (B) Site-specific recombination
Genetics - Problem Drill 19: Dissection of Gene Function: Mutational Analysis of Model Organisms No. 1 of 10 1. The mouse gene knockout is based on. (A) Homologous recombination (B) Site-specific recombination
Alternative Cleavage and Polyadenylation of RNA
 Developmental Cell 18 Supplemental Information The Spen Family Protein FPA Controls Alternative Cleavage and Polyadenylation of RNA Csaba Hornyik, Lionel C. Terzi, and Gordon G. Simpson Figure S1, related
Developmental Cell 18 Supplemental Information The Spen Family Protein FPA Controls Alternative Cleavage and Polyadenylation of RNA Csaba Hornyik, Lionel C. Terzi, and Gordon G. Simpson Figure S1, related
Supplementary Information
 Supplementary Information Supplementary Figure 1. ZBTB20 expression in the developing DRG. ZBTB20 expression in the developing DRG was detected by immunohistochemistry using anti-zbtb20 antibody 9A10 on
Supplementary Information Supplementary Figure 1. ZBTB20 expression in the developing DRG. ZBTB20 expression in the developing DRG was detected by immunohistochemistry using anti-zbtb20 antibody 9A10 on
GENES AND CHROMOSOMES II
 1 GENES AND CHROMOSOMES II Lecture 4 BIOL 266/2 2014-15 Dr. S. Azam Biology Department Concordia University 2 GENE AND THE GENOME The Structure of the Genome DNA fingerprinting 3 DNA fingerprinting: DNA-based
1 GENES AND CHROMOSOMES II Lecture 4 BIOL 266/2 2014-15 Dr. S. Azam Biology Department Concordia University 2 GENE AND THE GENOME The Structure of the Genome DNA fingerprinting 3 DNA fingerprinting: DNA-based
Candidate region (0.74 Mb) ATC TCT GGG ACT CAT GAG CAG GAG GCT AGC ATC TCT GGG ACT CAT TAG CAG GAG GCT AGC
 A idm-3 idm-3 B Physical distance (Mb) 4.6 4.86.6 8.4 C Chr.3 Recom. Rate (%) ATG 3.9.9.9 9.74 Candidate region (.74 Mb) n=4 TAA D idm-3 G3T(E4) G4A(W988) WT idm-3 ATC TCT GGG ACT CAT GAG CAG GAG GCT AGC
A idm-3 idm-3 B Physical distance (Mb) 4.6 4.86.6 8.4 C Chr.3 Recom. Rate (%) ATG 3.9.9.9 9.74 Candidate region (.74 Mb) n=4 TAA D idm-3 G3T(E4) G4A(W988) WT idm-3 ATC TCT GGG ACT CAT GAG CAG GAG GCT AGC
Exploring Chromodomain Genes in the Fungus Fusarium graminearum Through Targeted Genetic Manipulation
 Exploring Chromodomain Genes in the Fungus Fusarium graminearum Through Targeted Genetic Manipulation Alec Peters Dr. Michael Freitag Laboratory Dept of Biochemistry/Biophysics Oregon State University
Exploring Chromodomain Genes in the Fungus Fusarium graminearum Through Targeted Genetic Manipulation Alec Peters Dr. Michael Freitag Laboratory Dept of Biochemistry/Biophysics Oregon State University
TRANSPOSON INSERTION SITE VERIFICATION
 TRANSPOSON INSERTION SITE VERIFICATION Transposon and T-DNA insertion in Arabidopsis genes can be identified using the Arabidopsis thaliana Insertion Database (ATIdb) (http://atidb.org/cgi-perl/gbrowse/atibrowse).
TRANSPOSON INSERTION SITE VERIFICATION Transposon and T-DNA insertion in Arabidopsis genes can be identified using the Arabidopsis thaliana Insertion Database (ATIdb) (http://atidb.org/cgi-perl/gbrowse/atibrowse).
PrimePCR Assay Validation Report
 Gene Information Gene Name Gene Symbol Organism Gene Summary Gene Aliases RefSeq Accession No. UniGene ID Ensembl Gene ID sema domain, immunoglobulin domain (Ig), short basic domain, secreted, (semaphorin)
Gene Information Gene Name Gene Symbol Organism Gene Summary Gene Aliases RefSeq Accession No. UniGene ID Ensembl Gene ID sema domain, immunoglobulin domain (Ig), short basic domain, secreted, (semaphorin)
Supplementary Material
 Supplementary Material The Cerato-Platanin protein Epl-1 from Trichoderma harzianum is involved in mycoparasitism, plant resistance induction and self cell wall protection Eriston Vieira Gomes 1, Mariana
Supplementary Material The Cerato-Platanin protein Epl-1 from Trichoderma harzianum is involved in mycoparasitism, plant resistance induction and self cell wall protection Eriston Vieira Gomes 1, Mariana
Molecular Cell Biology - Problem Drill 11: Recombinant DNA
 Molecular Cell Biology - Problem Drill 11: Recombinant DNA Question No. 1 of 10 1. Which of the following statements about the sources of DNA used for molecular cloning is correct? Question #1 (A) cdna
Molecular Cell Biology - Problem Drill 11: Recombinant DNA Question No. 1 of 10 1. Which of the following statements about the sources of DNA used for molecular cloning is correct? Question #1 (A) cdna
Percent survival. Supplementary fig. S3 A.
 Supplementary fig. S3 A. B. 100 Percent survival 80 60 40 20 Ml 0 0 100 C. Fig. S3 Comparison of leukaemia incidence rate in the triple targeted chimaeric mice and germline-transmission translocator mice
Supplementary fig. S3 A. B. 100 Percent survival 80 60 40 20 Ml 0 0 100 C. Fig. S3 Comparison of leukaemia incidence rate in the triple targeted chimaeric mice and germline-transmission translocator mice
Supplemental Figure 1. Immunoblot analysis of wild-type and mutant forms of JAZ3
 Supplemental Data. Chung and Howe (2009). A Critical Role for the TIFY Motif in Repression of Jasmonate Signaling by a Stabilized Splice Variant of the JASMONATE ZIM-domain protein JAZ10 in Arabidopsis.
Supplemental Data. Chung and Howe (2009). A Critical Role for the TIFY Motif in Repression of Jasmonate Signaling by a Stabilized Splice Variant of the JASMONATE ZIM-domain protein JAZ10 in Arabidopsis.
3I03 - Eukaryotic Genetics Repetitive DNA
 Repetitive DNA Satellite DNA Minisatellite DNA Microsatellite DNA Transposable elements LINES, SINES and other retrosequences High copy number genes (e.g. ribosomal genes, histone genes) Multifamily member
Repetitive DNA Satellite DNA Minisatellite DNA Microsatellite DNA Transposable elements LINES, SINES and other retrosequences High copy number genes (e.g. ribosomal genes, histone genes) Multifamily member
Concepts: What are RFLPs and how do they act like genetic marker loci?
 Restriction Fragment Length Polymorphisms (RFLPs) -1 Readings: Griffiths et al: 7th Edition: Ch. 12 pp. 384-386; Ch.13 pp404-407 8th Edition: pp. 364-366 Assigned Problems: 8th Ch. 11: 32, 34, 38-39 7th
Restriction Fragment Length Polymorphisms (RFLPs) -1 Readings: Griffiths et al: 7th Edition: Ch. 12 pp. 384-386; Ch.13 pp404-407 8th Edition: pp. 364-366 Assigned Problems: 8th Ch. 11: 32, 34, 38-39 7th
PrimePCR Assay Validation Report
 Gene Information Gene Name collagen, type IV, alpha 1 Gene Symbol Organism Gene Summary Gene Aliases RefSeq Accession No. UniGene ID Ensembl Gene ID COL4A1 Human This gene encodes the major type IV alpha
Gene Information Gene Name collagen, type IV, alpha 1 Gene Symbol Organism Gene Summary Gene Aliases RefSeq Accession No. UniGene ID Ensembl Gene ID COL4A1 Human This gene encodes the major type IV alpha
A CRISPR/Cas9 Vector System for Tissue-Specific Gene Disruption in Zebrafish
 Developmental Cell Supplemental Information A CRISPR/Cas9 Vector System for Tissue-Specific Gene Disruption in Zebrafish Julien Ablain, Ellen M. Durand, Song Yang, Yi Zhou, and Leonard I. Zon % larvae
Developmental Cell Supplemental Information A CRISPR/Cas9 Vector System for Tissue-Specific Gene Disruption in Zebrafish Julien Ablain, Ellen M. Durand, Song Yang, Yi Zhou, and Leonard I. Zon % larvae
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:10.1038/nature09937 a Name Position Primersets 1a 1b 2 3 4 b2 Phenotype Genotype b Primerset 1a D T C R I E 10000 8000 6000 5000 4000 3000 2500 2000 1500 1000 800 Donor (D)
SUPPLEMENTARY INFORMATION doi:10.1038/nature09937 a Name Position Primersets 1a 1b 2 3 4 b2 Phenotype Genotype b Primerset 1a D T C R I E 10000 8000 6000 5000 4000 3000 2500 2000 1500 1000 800 Donor (D)
Supplementary Figure 2
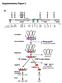 Supplementary Figure 2 a SBS-C1 SBS-C2 SBS-C3 SBS-C4 SBS-C5 SBS-C6 SBS-C7 SBS-C8 SBS-C9 LCR CNS-1 CNS-2 Il5 Rad50 Il13 Il4 Kif3a Sept8 0 50 100 150 200kb Sau3AI 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17
Supplementary Figure 2 a SBS-C1 SBS-C2 SBS-C3 SBS-C4 SBS-C5 SBS-C6 SBS-C7 SBS-C8 SBS-C9 LCR CNS-1 CNS-2 Il5 Rad50 Il13 Il4 Kif3a Sept8 0 50 100 150 200kb Sau3AI 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17
Site directed mutagenesis, Insertional and Deletion Mutagenesis. Mitesh Shrestha
 Site directed mutagenesis, Insertional and Deletion Mutagenesis Mitesh Shrestha Mutagenesis Mutagenesis (the creation or formation of a mutation) can be used as a powerful genetic tool. By inducing mutations
Site directed mutagenesis, Insertional and Deletion Mutagenesis Mitesh Shrestha Mutagenesis Mutagenesis (the creation or formation of a mutation) can be used as a powerful genetic tool. By inducing mutations
d. reading a DNA strand and making a complementary messenger RNA
 Biol/ MBios 301 (General Genetics) Spring 2003 Second Midterm Examination A (100 points possible) Key April 1, 2003 10 Multiple Choice Questions-4 pts. each (Choose the best answer) 1. Transcription involves:
Biol/ MBios 301 (General Genetics) Spring 2003 Second Midterm Examination A (100 points possible) Key April 1, 2003 10 Multiple Choice Questions-4 pts. each (Choose the best answer) 1. Transcription involves:
PrimePCR Assay Validation Report
 Gene Information Gene Name SRY (sex determining region Y)-box 6 Gene Symbol Organism Gene Summary Gene Aliases RefSeq Accession No. UniGene ID Ensembl Gene ID SOX6 Human This gene encodes a member of the
Gene Information Gene Name SRY (sex determining region Y)-box 6 Gene Symbol Organism Gene Summary Gene Aliases RefSeq Accession No. UniGene ID Ensembl Gene ID SOX6 Human This gene encodes a member of the
1a. What is the ratio of feathered to unfeathered shanks in the offspring of the above cross?
 Problem Set 5 answers 1. Whether or not the shanks of chickens contains feathers is due to two independently assorting genes. Individuals have unfeathered shanks when they are homozygous for recessive
Problem Set 5 answers 1. Whether or not the shanks of chickens contains feathers is due to two independently assorting genes. Individuals have unfeathered shanks when they are homozygous for recessive
SUPPLEMENTARY MATERIAL AND METHODS
 SUPPLEMENTARY MATERIAL AND METHODS Amplification of HEV ORF1, ORF2 and ORF3 genome regions Total RNA was extracted from 200 µl EDTA plasma using Cobas AmpliPrep total nucleic acid isolation kit (Roche,
SUPPLEMENTARY MATERIAL AND METHODS Amplification of HEV ORF1, ORF2 and ORF3 genome regions Total RNA was extracted from 200 µl EDTA plasma using Cobas AmpliPrep total nucleic acid isolation kit (Roche,
Your name: BSCI410-LIU/Spring 2007 Homework #2 Due March 27 (Tu), 07
 BSCI410-LIU/Spring 2007 Homework #2 Due March 27 (Tu), 07 KEY 1. What are each of the following molecular markers? (Indicate (a) what they stand for; (b) the nature of the molecular polymorphism and (c)
BSCI410-LIU/Spring 2007 Homework #2 Due March 27 (Tu), 07 KEY 1. What are each of the following molecular markers? (Indicate (a) what they stand for; (b) the nature of the molecular polymorphism and (c)
RNA oligonucleotides and 2 -O-methylated oligonucleotides were synthesized by. 5 AGACACAAACACCAUUGUCACACUCCACAGC; Rand-2 OMe,
 Materials and methods Oligonucleotides and DNA constructs RNA oligonucleotides and 2 -O-methylated oligonucleotides were synthesized by Dharmacon Inc. (Lafayette, CO). The sequences were: 122-2 OMe, 5
Materials and methods Oligonucleotides and DNA constructs RNA oligonucleotides and 2 -O-methylated oligonucleotides were synthesized by Dharmacon Inc. (Lafayette, CO). The sequences were: 122-2 OMe, 5
Multiple choice questions (numbers in brackets indicate the number of correct answers)
 1 Multiple choice questions (numbers in brackets indicate the number of correct answers) February 1, 2013 1. Ribose is found in Nucleic acids Proteins Lipids RNA DNA (2) 2. Most RNA in cells is transfer
1 Multiple choice questions (numbers in brackets indicate the number of correct answers) February 1, 2013 1. Ribose is found in Nucleic acids Proteins Lipids RNA DNA (2) 2. Most RNA in cells is transfer
Figure S1. Replication initiation sites at efficient ORIs do not coincide with nucleosomedepleted
 Figure S1. Replication initiation sites at efficient ORIs do not coincide with nucleosomedepleted regions in the mouse genome. Typical examples of SNS profiles (Cayrou et al., 11) and nucleosome occupancies
Figure S1. Replication initiation sites at efficient ORIs do not coincide with nucleosomedepleted regions in the mouse genome. Typical examples of SNS profiles (Cayrou et al., 11) and nucleosome occupancies
Technical Review. Real time PCR
 Technical Review Real time PCR Normal PCR: Analyze with agarose gel Normal PCR vs Real time PCR Real-time PCR, also known as quantitative PCR (qpcr) or kinetic PCR Key feature: Used to amplify and simultaneously
Technical Review Real time PCR Normal PCR: Analyze with agarose gel Normal PCR vs Real time PCR Real-time PCR, also known as quantitative PCR (qpcr) or kinetic PCR Key feature: Used to amplify and simultaneously
Gene mutation and DNA polymorphism
 Gene mutation and DNA polymorphism Outline of this chapter Gene Mutation DNA Polymorphism Gene Mutation Definition Major Types Definition A gene mutation is a change in the nucleotide sequence that composes
Gene mutation and DNA polymorphism Outline of this chapter Gene Mutation DNA Polymorphism Gene Mutation Definition Major Types Definition A gene mutation is a change in the nucleotide sequence that composes
Reading Lecture 8: Lecture 9: Lecture 8. DNA Libraries. Definition Types Construction
 Lecture 8 Reading Lecture 8: 96-110 Lecture 9: 111-120 DNA Libraries Definition Types Construction 142 DNA Libraries A DNA library is a collection of clones of genomic fragments or cdnas from a certain
Lecture 8 Reading Lecture 8: 96-110 Lecture 9: 111-120 DNA Libraries Definition Types Construction 142 DNA Libraries A DNA library is a collection of clones of genomic fragments or cdnas from a certain
sides of the aleurone (Al) but it is excluded from the basal endosperm transfer layer
 Supplemental Data. Gómez et al. (2009). The maize transcription factor MRP-1 (Myb-Related-Protein-1) is a key regulator of the differentiation of transfer cells. Supplemental Figure 1. Expression analyses
Supplemental Data. Gómez et al. (2009). The maize transcription factor MRP-1 (Myb-Related-Protein-1) is a key regulator of the differentiation of transfer cells. Supplemental Figure 1. Expression analyses
Supplementary Materials
 Supplementary Materials Construction of Synthetic Nucleoli in Human Cells Reveals How a Major Functional Nuclear Domain is Formed and Propagated Through Cell Divisision Authors: Alice Grob, Christine Colleran
Supplementary Materials Construction of Synthetic Nucleoli in Human Cells Reveals How a Major Functional Nuclear Domain is Formed and Propagated Through Cell Divisision Authors: Alice Grob, Christine Colleran
Genomics and Gene Recognition Genes and Blue Genes
 Genomics and Gene Recognition Genes and Blue Genes November 3, 2004 Eukaryotic Gene Structure eukaryotic genomes are considerably more complex than those of prokaryotes eukaryotic cells have organelles
Genomics and Gene Recognition Genes and Blue Genes November 3, 2004 Eukaryotic Gene Structure eukaryotic genomes are considerably more complex than those of prokaryotes eukaryotic cells have organelles
3. human genomics clone genes associated with genetic disorders. 4. many projects generate ordered clones that cover genome
 Lectures 30 and 31 Genome analysis I. Genome analysis A. two general areas 1. structural 2. functional B. genome projects a status report 1. 1 st sequenced: several viral genomes 2. mitochondria and chloroplasts
Lectures 30 and 31 Genome analysis I. Genome analysis A. two general areas 1. structural 2. functional B. genome projects a status report 1. 1 st sequenced: several viral genomes 2. mitochondria and chloroplasts
Chapter Eleven: Chromosome Structure and Transposable Elements
 Chapter Eleven: Chromosome Structure and Transposable Elements COMPREHENSION QUESTIONS Section 11.1 *1. How does supercoiling arise? What is the difference between positive and negative supercoiling? Supercoiling
Chapter Eleven: Chromosome Structure and Transposable Elements COMPREHENSION QUESTIONS Section 11.1 *1. How does supercoiling arise? What is the difference between positive and negative supercoiling? Supercoiling
Supplemental Data. Lee et al. Plant Cell. (2010) /tpc Supplemental Figure 1. Protein and Gene Structures of DWA1 and DWA2.
 Supplemental Figure 1. Protein and Gene Structures of DWA1 and DWA2. (A) Protein structures of DWA1 and DWA2. WD40 region was determined based on the NCBI conserved domain databases (B, C) Schematic representation
Supplemental Figure 1. Protein and Gene Structures of DWA1 and DWA2. (A) Protein structures of DWA1 and DWA2. WD40 region was determined based on the NCBI conserved domain databases (B, C) Schematic representation
Supplementary Figures
 Supplementary Figures Supplementary Figure 1 Experimental schema for the identification of circular RNAs in six normal tissues and seven cancerous tissues. Supplementary Fiure 2 Comparison of human circrnas
Supplementary Figures Supplementary Figure 1 Experimental schema for the identification of circular RNAs in six normal tissues and seven cancerous tissues. Supplementary Fiure 2 Comparison of human circrnas
Figure S1. Figure S2. Figure S3 HB Anti-FSP27 (COOH-terminal peptide) Ab. Anti-GST-FSP27(45-127) Ab.
 / 36B4 mrna ratio Figure S1 * 2. 1.6 1.2.8 *.4 control TNFα BRL49653 Figure S2 Su bw AT p iw Anti- (COOH-terminal peptide) Ab Blot : Anti-GST-(45-127) Ab β-actin Figure S3 HB2 HW AT BA T Figure S4 A TAG
/ 36B4 mrna ratio Figure S1 * 2. 1.6 1.2.8 *.4 control TNFα BRL49653 Figure S2 Su bw AT p iw Anti- (COOH-terminal peptide) Ab Blot : Anti-GST-(45-127) Ab β-actin Figure S3 HB2 HW AT BA T Figure S4 A TAG
Figure S1. Characterization of the irx9l-1 mutant. (A) Diagram of the Arabidopsis IRX9L gene drawn based on information from TAIR (the Arabidopsis
 1 2 3 4 5 6 7 8 9 10 11 12 Figure S1. Characterization of the irx9l-1 mutant. (A) Diagram of the Arabidopsis IRX9L gene drawn based on information from TAIR (the Arabidopsis Information Research). Exons
1 2 3 4 5 6 7 8 9 10 11 12 Figure S1. Characterization of the irx9l-1 mutant. (A) Diagram of the Arabidopsis IRX9L gene drawn based on information from TAIR (the Arabidopsis Information Research). Exons
Genes found in the genome include protein-coding genes and non-coding RNA genes. Which nucleotide is not normally found in non-coding RNA genes?
 Midterm Q Genes found in the genome include protein-coding genes and non-coding RNA genes Which nucleotide is not normally found in non-coding RNA genes? G T 3 A 4 C 5 U 00% Midterm Q Which of the following
Midterm Q Genes found in the genome include protein-coding genes and non-coding RNA genes Which nucleotide is not normally found in non-coding RNA genes? G T 3 A 4 C 5 U 00% Midterm Q Which of the following
Unit 6: Molecular Genetics & DNA Technology Guided Reading Questions (100 pts total)
 Name: AP Biology Biology, Campbell and Reece, 7th Edition Adapted from chapter reading guides originally created by Lynn Miriello Chapter 16 The Molecular Basis of Inheritance Unit 6: Molecular Genetics
Name: AP Biology Biology, Campbell and Reece, 7th Edition Adapted from chapter reading guides originally created by Lynn Miriello Chapter 16 The Molecular Basis of Inheritance Unit 6: Molecular Genetics
Genome research in eukaryotes
 Functional Genomics Genome and EST sequencing can tell us how many POTENTIAL genes are present in the genome Proteomics can tell us about proteins and their interactions The goal of functional genomics
Functional Genomics Genome and EST sequencing can tell us how many POTENTIAL genes are present in the genome Proteomics can tell us about proteins and their interactions The goal of functional genomics
Supplemental Materials. Matrix Proteases Contribute to Progression of Pelvic Organ Prolapse in Mice and Humans
 Supplemental Materials Matrix Proteases Contribute to Progression of Pelvic Organ Prolapse in Mice and Humans Madhusudhan Budatha, Shayzreen Roshanravan, Qian Zheng, Cecilia Weislander, Shelby L. Chapman,
Supplemental Materials Matrix Proteases Contribute to Progression of Pelvic Organ Prolapse in Mice and Humans Madhusudhan Budatha, Shayzreen Roshanravan, Qian Zheng, Cecilia Weislander, Shelby L. Chapman,
Chapter 6 - Molecular Genetic Techniques
 Chapter 6 - Molecular Genetic Techniques Two objects of molecular & genetic technologies For analysis For generation Molecular genetic technologies! For analysis DNA gel electrophoresis Southern blotting
Chapter 6 - Molecular Genetic Techniques Two objects of molecular & genetic technologies For analysis For generation Molecular genetic technologies! For analysis DNA gel electrophoresis Southern blotting
Supplemental Material Igreja and Izaurralde 1. CUP promotes deadenylation and inhibits decapping of mrna targets. Catia Igreja and Elisa Izaurralde
 Supplemental Material Igreja and Izaurralde 1 CUP promotes deadenylation and inhibits decapping of mrna targets Catia Igreja and Elisa Izaurralde Supplemental Materials and methods Functional assays and
Supplemental Material Igreja and Izaurralde 1 CUP promotes deadenylation and inhibits decapping of mrna targets Catia Igreja and Elisa Izaurralde Supplemental Materials and methods Functional assays and
Supplemental Data. Wu et al. (2). Plant Cell..5/tpc RGLG Hormonal treatment H2O B RGLG µm ABA µm ACC µm GA Time (hours) µm µm MJ µm IA
 Supplemental Data. Wu et al. (2). Plant Cell..5/tpc..4. A B Supplemental Figure. Immunoblot analysis verifies the expression of the AD-PP2C and BD-RGLG proteins in the Y2H assay. Total proteins were extracted
Supplemental Data. Wu et al. (2). Plant Cell..5/tpc..4. A B Supplemental Figure. Immunoblot analysis verifies the expression of the AD-PP2C and BD-RGLG proteins in the Y2H assay. Total proteins were extracted
Recombinant DNA Technology. The Role of Recombinant DNA Technology in Biotechnology. yeast. Biotechnology. Recombinant DNA technology.
 PowerPoint Lecture Presentations prepared by Mindy Miller-Kittrell, North Carolina State University C H A P T E R 8 Recombinant DNA Technology The Role of Recombinant DNA Technology in Biotechnology Biotechnology?
PowerPoint Lecture Presentations prepared by Mindy Miller-Kittrell, North Carolina State University C H A P T E R 8 Recombinant DNA Technology The Role of Recombinant DNA Technology in Biotechnology Biotechnology?
Reverse transcription-pcr (rt-pcr) Dr. Hani Alhadrami
 Reverse transcription-pcr (rt-pcr) Dr. Hani Alhadrami hanialhadrami@kau.edu.sa www.hanialhadrami.kau.edu.sa Overview Several techniques are available to detect and analyse RNA. Examples of these techniques
Reverse transcription-pcr (rt-pcr) Dr. Hani Alhadrami hanialhadrami@kau.edu.sa www.hanialhadrami.kau.edu.sa Overview Several techniques are available to detect and analyse RNA. Examples of these techniques
Roche Molecular Biochemicals Technical Note No. LC 9/2000
 Roche Molecular Biochemicals Technical Note No. LC 9/2000 LightCycler Optimization Strategy Introduction Purpose of this Note Table of Contents The LightCycler system provides different detection formats
Roche Molecular Biochemicals Technical Note No. LC 9/2000 LightCycler Optimization Strategy Introduction Purpose of this Note Table of Contents The LightCycler system provides different detection formats
A Low salt diet. C Low salt diet + mf4-31c1 3. D High salt diet + mf4-31c1 3. B High salt diet
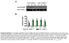 A Low salt diet GV [AU]:. [mmhg]: 0..... 09 9 7 B High salt diet GV [AU]:. [mmhg]: 7 8...0. 7.8 8.. 0 8 7 C Low salt diet + mf-c GV [AU]:. [mmhg]:.0.9 8.7.7. 7 8 0 D High salt diet + mf-c E Lymph capillary
A Low salt diet GV [AU]:. [mmhg]: 0..... 09 9 7 B High salt diet GV [AU]:. [mmhg]: 7 8...0. 7.8 8.. 0 8 7 C Low salt diet + mf-c GV [AU]:. [mmhg]:.0.9 8.7.7. 7 8 0 D High salt diet + mf-c E Lymph capillary
SUPPLEMENTARY INFORMATION
 Local auxin metabolism regulates environment-induced hypocotyl elongation Zuyu Zheng 1,2, Yongxia Guo 3, Ondřej Novák 4,5, William Chen 2, Karin Ljung 4, Joseph P. Noel 1,3, *, and Joanne Chory 1,2, *
Local auxin metabolism regulates environment-induced hypocotyl elongation Zuyu Zheng 1,2, Yongxia Guo 3, Ondřej Novák 4,5, William Chen 2, Karin Ljung 4, Joseph P. Noel 1,3, *, and Joanne Chory 1,2, *
Supplementary Figure S1. Immunodetection of full-length XA21 and the XA21 C-terminal cleavage product.
 Supplementary Information Supplementary Figure S1. Immunodetection of full-length XA21 and the XA21 C-terminal cleavage product. Total protein extracted from Kitaake wild type and rice plants carrying
Supplementary Information Supplementary Figure S1. Immunodetection of full-length XA21 and the XA21 C-terminal cleavage product. Total protein extracted from Kitaake wild type and rice plants carrying
Fig. A1 Characteristics of HepP and its homology to other proteins. Fig. A2 Amino acid sequence of the 78-kDa predicted protein encoded by zbdp
 Additional file 1 This file contains: Fig. A1 Characteristics of HepP and its homology to other proteins Fig. A2 Amino acid sequence of the 78-kDa predicted protein encoded by zbdp Fig. A3 Nucleotide sequences
Additional file 1 This file contains: Fig. A1 Characteristics of HepP and its homology to other proteins Fig. A2 Amino acid sequence of the 78-kDa predicted protein encoded by zbdp Fig. A3 Nucleotide sequences
PrimePCR Assay Validation Report
 Gene Information Gene Name laminin, beta 3 Gene Symbol Organism Gene Summary Gene Aliases RefSeq Accession No. UniGene ID Ensembl Gene ID LAMB3 Human The product encoded by this gene is a laminin that
Gene Information Gene Name laminin, beta 3 Gene Symbol Organism Gene Summary Gene Aliases RefSeq Accession No. UniGene ID Ensembl Gene ID LAMB3 Human The product encoded by this gene is a laminin that
Fatchiyah
 Fatchiyah Email: fatchiya@yahoo.co.id RNAs: mrna trna rrna RNAi DNAs: Protein: genome DNA cdna mikro-makro mono-poly single-multi Analysis: Identification human and animal disease Finger printing Sexing
Fatchiyah Email: fatchiya@yahoo.co.id RNAs: mrna trna rrna RNAi DNAs: Protein: genome DNA cdna mikro-makro mono-poly single-multi Analysis: Identification human and animal disease Finger printing Sexing
RNP purification, components and activity.
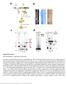 Supplementary Figure 1 RNP purification, components and activity. (a) Intein-mediated RNP production and purification. The Ll.LtrB intron RNA (red) (Exons E1 and E2 in green) and associated intron encoded
Supplementary Figure 1 RNP purification, components and activity. (a) Intein-mediated RNP production and purification. The Ll.LtrB intron RNA (red) (Exons E1 and E2 in green) and associated intron encoded
Supplementary Information
 Supplementary Information Deletion of the B-B and C-C regions of inverted terminal repeats reduces raav productivity but increases transgene expression Qingzhang Zhou 1, Wenhong Tian 2, Chunguo Liu 3,
Supplementary Information Deletion of the B-B and C-C regions of inverted terminal repeats reduces raav productivity but increases transgene expression Qingzhang Zhou 1, Wenhong Tian 2, Chunguo Liu 3,
Supplemental Data Supplemental Figure 1.
 Supplemental Data Supplemental Figure 1. Silique arrangement in the wild-type, jhs, and complemented lines. Wild-type (WT) (A), the jhs1 mutant (B,C), and the jhs1 mutant complemented with JHS1 (Com) (D)
Supplemental Data Supplemental Figure 1. Silique arrangement in the wild-type, jhs, and complemented lines. Wild-type (WT) (A), the jhs1 mutant (B,C), and the jhs1 mutant complemented with JHS1 (Com) (D)
PrimePCR Assay Validation Report
 Gene Information Gene Name transforming growth factor, beta 1 Gene Symbol Organism Gene Summary Gene Aliases RefSeq Accession No. UniGene ID Ensembl Gene ID TGFB1 Human This gene encodes a member of the
Gene Information Gene Name transforming growth factor, beta 1 Gene Symbol Organism Gene Summary Gene Aliases RefSeq Accession No. UniGene ID Ensembl Gene ID TGFB1 Human This gene encodes a member of the
User Manual. Catalog No.: DWSK-V101 (10 rxns), DWSK-V102 (25 rxns) For Research Use Only
 DNA Walking SpeedUp TM Kit SpeedUp Sequencing SpeedUp BAC Clone Sequencing SpeedUp Genome Walking SpeedUp Transgene Location Detection SpeedUp Deletion/ Insertion/ Isoform Detection User Manual Version
DNA Walking SpeedUp TM Kit SpeedUp Sequencing SpeedUp BAC Clone Sequencing SpeedUp Genome Walking SpeedUp Transgene Location Detection SpeedUp Deletion/ Insertion/ Isoform Detection User Manual Version
Supplementary Fig. S1. Schematic representation of mouse lines Pax6 fl/fl and mrx-cre used in this study. (A) To generate Pax6 fl/ fl
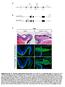 Supplementary Fig. S1. Schematic representation of mouse lines Pax6 fl/fl and mrx-cre used in this study. (A) To generate Pax6 fl/ fl, loxp sites flanking exons 3-6 (red arrowheads) were introduced into
Supplementary Fig. S1. Schematic representation of mouse lines Pax6 fl/fl and mrx-cre used in this study. (A) To generate Pax6 fl/ fl, loxp sites flanking exons 3-6 (red arrowheads) were introduced into
Supplemental Information
 Supplemental Information Itemized List Materials and Methods, Related to Supplemental Figures S5A-C and S6. Supplemental Figure S1, Related to Figures 1 and 2. Supplemental Figure S2, Related to Figure
Supplemental Information Itemized List Materials and Methods, Related to Supplemental Figures S5A-C and S6. Supplemental Figure S1, Related to Figures 1 and 2. Supplemental Figure S2, Related to Figure
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature11726 Supplementary Figure 1 SRP RNA constructs used in single molecule experiments. a, Secondary structure of wildtype E. coli SRP RNA, which forms an elongated hairpin structure capped
doi:10.1038/nature11726 Supplementary Figure 1 SRP RNA constructs used in single molecule experiments. a, Secondary structure of wildtype E. coli SRP RNA, which forms an elongated hairpin structure capped
Ramp1 EPD0843_4_B11. EUCOMM/KOMP-CSD Knockout-First Genotyping
 EUCOMM/KOMP-CSD Knockout-First Genotyping Introduction The majority of animals produced from the EUCOMM/KOMP-CSD ES cell resource contain the Knockout-First-Reporter Tagged Insertion allele. As well as
EUCOMM/KOMP-CSD Knockout-First Genotyping Introduction The majority of animals produced from the EUCOMM/KOMP-CSD ES cell resource contain the Knockout-First-Reporter Tagged Insertion allele. As well as
Usp14 EPD0582_2_G09. EUCOMM/KOMP-CSD Knockout-First Genotyping
 EUCOMM/KOMP-CSD Knockout-First Genotyping Introduction The majority of animals produced from the EUCOMM/KOMP-CSD ES cell resource contain the Knockout-First-Reporter Tagged Insertion allele. As well as
EUCOMM/KOMP-CSD Knockout-First Genotyping Introduction The majority of animals produced from the EUCOMM/KOMP-CSD ES cell resource contain the Knockout-First-Reporter Tagged Insertion allele. As well as
Meiotically Stable Natural Epialleles of Sadhu, a Novel Arabidopsis Retroposon
 Meiotically Stable Natural Epialleles of Sadhu, a Novel Arabidopsis Retroposon Sanjida H. Rangwala 1, Rangasamy Elumalai 2 a, Cheryl Vanier 3, Hakan Ozkan 2 b, David W. Galbraith 2, Eric J. Richards 1*
Meiotically Stable Natural Epialleles of Sadhu, a Novel Arabidopsis Retroposon Sanjida H. Rangwala 1, Rangasamy Elumalai 2 a, Cheryl Vanier 3, Hakan Ozkan 2 b, David W. Galbraith 2, Eric J. Richards 1*
Polymerase Chain Reaction
 Polymerase Chain Reaction Problem Suppose you have a patient with an infection or a heritable disease. You want to know which infection or disease it is and.. you want to know it fast and... from as little
Polymerase Chain Reaction Problem Suppose you have a patient with an infection or a heritable disease. You want to know which infection or disease it is and.. you want to know it fast and... from as little
Problem Set 8. Answer Key
 MCB 102 University of California, Berkeley August 11, 2009 Isabelle Philipp Online Document Problem Set 8 Answer Key 1. The Genetic Code (a) Are all amino acids encoded by the same number of codons? no
MCB 102 University of California, Berkeley August 11, 2009 Isabelle Philipp Online Document Problem Set 8 Answer Key 1. The Genetic Code (a) Are all amino acids encoded by the same number of codons? no
Molecular Analysis of Genes and Gene Products. BIT 220 Chapter 22
 Molecular Analysis of Genes and Gene Products BIT 220 Chapter 22 Credit: Courtesy Susan Lanzendorf, Ph.D., Jones Institute for Reproductive Medicine/Eastern Virginia Medical School 2003 John Wiley and
Molecular Analysis of Genes and Gene Products BIT 220 Chapter 22 Credit: Courtesy Susan Lanzendorf, Ph.D., Jones Institute for Reproductive Medicine/Eastern Virginia Medical School 2003 John Wiley and
Supplementary Material
 Fig S1. Interactions of OsDof24 with the OsC4PPDK promoter in rice protoplasts (a) Loss-of-function analysis of the OsC4PPDK promoter Effects of OsDof24 overexpression and the mapping of OsDof24-binding
Fig S1. Interactions of OsDof24 with the OsC4PPDK promoter in rice protoplasts (a) Loss-of-function analysis of the OsC4PPDK promoter Effects of OsDof24 overexpression and the mapping of OsDof24-binding
Sensitivity vs Specificity
 Viral Detection Animal Inoculation Culturing the Virus Definitive Length of time Serology Detecting antibodies to the infectious agent Detecting Viral Proteins Western Blot ELISA Detecting the Viral Genome
Viral Detection Animal Inoculation Culturing the Virus Definitive Length of time Serology Detecting antibodies to the infectious agent Detecting Viral Proteins Western Blot ELISA Detecting the Viral Genome
NAME TA SEC Problem Set 4 FRIDAY October 15, Answers to this problem set must be inserted into the box outside
 MIT Biology Department 7.012: Introductory Biology - Fall 2004 Instructors: Professor Eric Lander, Professor Robert A. Weinberg, Dr. Claudette Gardel NAME TA SEC 7.012 Problem Set 4 FRIDAY October 15,
MIT Biology Department 7.012: Introductory Biology - Fall 2004 Instructors: Professor Eric Lander, Professor Robert A. Weinberg, Dr. Claudette Gardel NAME TA SEC 7.012 Problem Set 4 FRIDAY October 15,
Kerkel et al., Supplementary Tables and Figures
 Kerkel et al., Supplementary Tables and Figures Table S1. Summary of 16 SNP tagged loci from the combined 50K and 250K MSNP screens with independently validated ASM. PBL, total peripheral blood leukocytes;
Kerkel et al., Supplementary Tables and Figures Table S1. Summary of 16 SNP tagged loci from the combined 50K and 250K MSNP screens with independently validated ASM. PBL, total peripheral blood leukocytes;
Bacterial DNA replication
 Bacterial DNA replication Summary: What problems do these proteins solve? Tyr OH attacks PO4 and forms a covalent intermediate Structural changes in the protein open the gap by 20 Å! 1 Summary: What problems
Bacterial DNA replication Summary: What problems do these proteins solve? Tyr OH attacks PO4 and forms a covalent intermediate Structural changes in the protein open the gap by 20 Å! 1 Summary: What problems
2054, Chap. 14, page 1
 2054, Chap. 14, page 1 I. Recombinant DNA technology (Chapter 14) A. recombinant DNA technology = collection of methods used to perform genetic engineering 1. genetic engineering = deliberate modification
2054, Chap. 14, page 1 I. Recombinant DNA technology (Chapter 14) A. recombinant DNA technology = collection of methods used to perform genetic engineering 1. genetic engineering = deliberate modification
Nature Methods: doi: /nmeth Supplementary Figure 1
 Supplementary Figure 1 Workflow for multimodal analysis using sc-gem on a programmable microfluidic device (Fluidigm). 1) Cells are captured and lysed, 2) RNA from lysed single cell is reverse-transcribed
Supplementary Figure 1 Workflow for multimodal analysis using sc-gem on a programmable microfluidic device (Fluidigm). 1) Cells are captured and lysed, 2) RNA from lysed single cell is reverse-transcribed
Midterm 1 Results. Midterm 1 Akey/ Fields Median Number of Students. Exam Score
 Midterm 1 Results 10 Midterm 1 Akey/ Fields Median - 69 8 Number of Students 6 4 2 0 21 26 31 36 41 46 51 56 61 66 71 76 81 86 91 96 101 Exam Score Quick review of where we left off Parental type: the
Midterm 1 Results 10 Midterm 1 Akey/ Fields Median - 69 8 Number of Students 6 4 2 0 21 26 31 36 41 46 51 56 61 66 71 76 81 86 91 96 101 Exam Score Quick review of where we left off Parental type: the
CHAPTER 9 DNA Technologies
 CHAPTER 9 DNA Technologies Recombinant DNA Artificially created DNA that combines sequences that do not occur together in the nature Basis of much of the modern molecular biology Molecular cloning of genes
CHAPTER 9 DNA Technologies Recombinant DNA Artificially created DNA that combines sequences that do not occur together in the nature Basis of much of the modern molecular biology Molecular cloning of genes
Optimization of site-specific RNase H cutting of 1037 nt renilla luciferase mrna. The digestion
 SUPPLEMENTARY FIGURE 1. Optimization of site-specific RNase H cutting of 1037 nt renilla luciferase mrna. The digestion mixture was incubated at 37 C for 0 to 120 minutes. The cutting was nearly complete
SUPPLEMENTARY FIGURE 1. Optimization of site-specific RNase H cutting of 1037 nt renilla luciferase mrna. The digestion mixture was incubated at 37 C for 0 to 120 minutes. The cutting was nearly complete
Activation of a Floral Homeotic Gene in Arabidopsis
 Activation of a Floral Homeotic Gene in Arabidopsis By Maximiliam A. Busch, Kirsten Bomblies, and Detlef Weigel Presentation by Lis Garrett and Andrea Stevenson http://ucsdnews.ucsd.edu/archive/graphics/images/image5.jpg
Activation of a Floral Homeotic Gene in Arabidopsis By Maximiliam A. Busch, Kirsten Bomblies, and Detlef Weigel Presentation by Lis Garrett and Andrea Stevenson http://ucsdnews.ucsd.edu/archive/graphics/images/image5.jpg
Learning Objectives :
 Learning Objectives : Understand the basic differences between genomic and cdna libraries Understand how genomic libraries are constructed Understand the purpose for having overlapping DNA fragments in
Learning Objectives : Understand the basic differences between genomic and cdna libraries Understand how genomic libraries are constructed Understand the purpose for having overlapping DNA fragments in
ABSTRACT INTRODUCTION
 OMICS A Journal of Integrative Biology Volume 6, Number 2, 2002 Mary Ann Liebert, Inc. Characterization of T-DNA Insertion Sites in Arabidopsis thaliana and the Implications for Saturation Mutagenesis
OMICS A Journal of Integrative Biology Volume 6, Number 2, 2002 Mary Ann Liebert, Inc. Characterization of T-DNA Insertion Sites in Arabidopsis thaliana and the Implications for Saturation Mutagenesis
GTTCGGGTTCC TTTTGAGCAG
 Supplementary Figures Splice variants of the SIP1 transcripts play a role in nodule organogenesis in Lotus japonicus. Wang C, Zhu H, Jin L, Chen T, Wang L, Kang H, Hong Z, Zhang Z. 5 UTR CDS 3 UTR TCTCAACCATCCTTTGTCTGCTTCCGCCGCATGGGTGAGGTCATTTTGTCTAGATGACGTGCAATTTACAATGA
Supplementary Figures Splice variants of the SIP1 transcripts play a role in nodule organogenesis in Lotus japonicus. Wang C, Zhu H, Jin L, Chen T, Wang L, Kang H, Hong Z, Zhang Z. 5 UTR CDS 3 UTR TCTCAACCATCCTTTGTCTGCTTCCGCCGCATGGGTGAGGTCATTTTGTCTAGATGACGTGCAATTTACAATGA
MIT Department of Biology 7.013: Introductory Biology - Spring 2005 Instructors: Professor Hazel Sive, Professor Tyler Jacks, Dr.
 MIT Department of Biology 7.01: Introductory Biology - Spring 2005 Instructors: Professor Hazel Sive, Professor Tyler Jacks, Dr. Claudette Gardel iv) Would Xba I be useful for cloning? Why or why not?
MIT Department of Biology 7.01: Introductory Biology - Spring 2005 Instructors: Professor Hazel Sive, Professor Tyler Jacks, Dr. Claudette Gardel iv) Would Xba I be useful for cloning? Why or why not?
Quiz Submissions Quiz 4
 Quiz Submissions Quiz 4 Attempt 1 Written: Nov 1, 2015 17:35 Nov 1, 2015 22:19 Submission View Released: Nov 4, 2015 20:24 Question 1 0 / 1 point Three RNA polymerases synthesize most of the RNA present
Quiz Submissions Quiz 4 Attempt 1 Written: Nov 1, 2015 17:35 Nov 1, 2015 22:19 Submission View Released: Nov 4, 2015 20:24 Question 1 0 / 1 point Three RNA polymerases synthesize most of the RNA present
