SUPPLEMENTARY INFORMATION
|
|
|
- Brendan Barton
- 6 years ago
- Views:
Transcription
1 doi: /nature11726 Supplementary Figure 1 SRP RNA constructs used in single molecule experiments. a, Secondary structure of wildtype E. coli SRP RNA, which forms an elongated hairpin structure capped by a conserved GGAA tetraloop. Pink region highlights the distal docking site for the SRP-FtsY GTPase complex observed in the crystal structure (PBD code: 2xxa, from Ataide et al, Science, 2011). b, Wildtype, 99A, and 82mer smrna constructs used in this work. c, Analytical RNA gel verified the generation and purification of smrna. smrnas (lanes 4, 6, 7) were purified from in vitro transcription mixtures (lanes 2, 5) by a preparative RNA gel, which removed the HDV ribozyme (90 nt, lane 3) and uncleaved precursor (210 ntd, lane 5). d, GTPase reactions for SRPs assembled with DNA-smRNA hybrid and with labeled FtsY. 1
2 RESEARCH SUPPLEMENTARY INFORMATION Supplementary Figure 2 smfret of the SRP-FtsY complex with Cy3-labeled Ffh. a, Schematics of the immobilized SRP-FtsY complex on the coverslip with Cy3-labeled Ffh. b, Fluorescent signals (upper) and FRET trajectory (lower) of the SRP-FtsY complex. Color coding is the same as in Figure 1b. The arrow denotes bleaching of Cy3. c, Zoom in of the grey box in b to show the four FRET states resolved by HMM. d, smfret histogram. e, HMM analysis of a sample FRET trajectory using different numbers of states. As the number of FRET states increases, better fits to experimental data were obtained. f, Probability score of each model in the HMM analysis in e. The probability maxed out at 4 states, which was chosen as the optimal model to describe experimental data. g, The FRET values of each state from the HMM analyses in e. Bold indicates the converged FRET values of the identified states. h, TDP of the GTPase rearrangement. i-n, Analysis of the transition kinetics between states L and H (i), L and M1 (j), L and 2 W W W. N A T U R E. C O M / N A T U R E
3 RESEARCH in the HMM analysis in e. The probability maxed out at 4 states, which was chosen as the optimal model to describe experimental data. g, The FRET values of each state from the HMM analyses in e. Bold indicates the converged FRET values of the identified states. h, TDP of the GTPase rearrangement. i-n, Analysis of the transition kinetics between states L and H (i), L and M1 (j), L and M2 (k), M1 and M2 (l), H and M1 (m), H and M2 (n). o, Summary of transition rates between different FRET states. p-r, Scatter plots of the transition dwell times of individual molecules. Each cross represents an individual molecule. 3
4 RESEARCH SUPPLEMENTARY INFORMATION Supplementary Figure 3 smfret analysis of the SRP-FtsY complex with labeled FtsY. a, HMM analysis of the FRET trajectory in Figure 1b using different numbers of FRET states. As the number of FRET states increases, better fits to experimental data were obtained. b, Probability score of each model in the HMM analysis in a. The probability maxed out at 4 states, which was chosen as the optimal model to describe experimental data. c, The FRET values of each state from the HMM analyses in a. Bold indicates the converged FRET values of the identified states. d-h, Analysis of the transition kinetics between states L and M1 (d), L and M2 (e), H and M1 (f), H and M2 (g), M1 and M2 (h). i-k, Scatter plots of the transition dwell times of individual molecules. Each cross represents an individual molecule. 4
5 RESEARCH Supplementary Figure 4 Activity of SRP RNA distal end mutants in GTPase reactions and in co-translational protein targeting and translocation. a, Representative data for stimulated GTPase reactions upon assembly of the SRP-FtsY complex with wildtype (red), 82mer (blue), and 99A (orange) SRP RNA. The data in the absence of SRP RNA are in black. Summary of the rate constants, with n = 5, is provided in Figure 2a. b, Schematics of the co-translational protein targeting and translocation assay. c & e, Representative data for co-translational targeting and translocation of ppl (c) and ppl signal sequence variants (e) by the wildtype and mutant SRPs. Color coding is the same as in a. Quantification of the data from three independent experiments is provided in Figures 2b and c. d, Signal sequence variants of ppl used in this study. Bold highlights the hydrophobic core of the signal sequence. 5
6 RESEARCH SUPPLEMENTARY INFORMATION Supplementary Figure 5 smfret data and analysis for SRP-FtsY complexes assembled with the 82mer SRP RNA. a & c, Fluorescent signals (upper) and FRET trajectory (lower) of the SRP(82mer)-FtsY complex with Cy3-labeled Ffh (a) or FtsY (c). Color coding is the same as in Figure 1b. b & d, smfret histograms of the GTPase complex with Cy3-labeled Ffh (b) or FtsY (d). 6
7 RESEARCH Supplementary Figure 6 smfret data and analyses for complexes assembled with mutant 99A RNA. a & f, Fluorescent signals (upper) and FRET trajectory (lower) of the SRP(99A)-FtsY complex with Cy3-labeled Ffh (a) or FtsY (f). Color coding is the same as in Figure 1b. b & g, smfret histograms of the GTPase complex with Cy3-labeled Ffh (b) or FtsY (g). c & h, Summary of the FRET distributions of SRP(99A)-FtsY and comparison with the wildtype complex for Cy3-labeled Ffh (c) or FtsY (h). d & i, TDP of the SRP(99A)-FtsY complex with Cy3-labeled Ffh (d) or FtsY (i). e & j, Summary of transition kinetics of the SRP(99A)- FtsY complex with Cy3-labeled Ffh (e) or FtsY (j). 7
8 RESEARCH SUPPLEMENTARY INFORMATION Supplementary Figure 7 smfret traces of SRP at different stages of its GTPase cycle, including free SRP (a) and the SRP- FtsY complex in the early (b) and closed (c) states. Ffh was labeled with Cy3. Color coding is the same as in Figure 1b. d, TDP of the SRP-FtsY complex in the closed state. 8
9 RESEARCH Supplementary Figure 8 Dwell time analysis of real-time GTP hydrolysis. a, Exponential fit of Δτ gave a lower limit for the GTPase rate. b, Exponential fit of the low FRET state between high FRET burst clusters. 9
10 RESEARCH SUPPLEMENTARY INFORMATION Supplementary Figure 9 RNC carrying a correct SRP substrate abolishes GTPase movement along the SRP RNA. a, RNCbound SRP-FtsY complex on the coverslip surface. b & f, Fluorescent signals (upper) and FRET trajectory (lower) of the SRP-FtsY complex bound to RNC FtsQ (b) or RNC luciferase (f). Color coding is the same as in Figure 1b. c-e, smfret histograms of the RNC FtsQ - SRP complex (c) and the RNC FtsQ -SRP-FtsY complex in the early (d) and closed (e) states. 10
11 RESEARCH Supplementary Figure 10 RNC s pausing effect is independent of the SRP RNA distal end. a & c, Fluorescent (upper) and FRET (lower) trajectory of the RNC FtsQ -SRP(82mer)-FtsY complex in the absence (a) and presence (c) of SecYEG. Color coding is the same as in Fig. 1b. b, smfret histogram of the RNC FtsQ -SRP(82mer)-FtsY complex. d, Representative data for GTPase assay of mutant FtsY T307A in the absence (green) and presence (grey) of RNC FtsQ. e, Summary of the k cat values from (d). Data represent mean±s.d. (n=3). 11
12 RESEARCH SUPPLEMENTARY INFORMATION Supplementary Figure 11 SecYEG restores GTPase movement to the SRP RNA distal site. a, SecYEG-bound RNC FtsQ -SRP- FtsY complex on the surface of the coverslip. b, smfret histogram of the RNC FtsQ -SRP-FtsY complex in the presence of 0.1% DDM. For comparison, the DDM concentration in reactions containing SecYEG is 0.07%. c, smfret histogram of the RNC FtsQ - SRP-FtsY(A335W) closed complex in the presence of SecYEG. d & e, FRET trajectory (d) and smfret histogram (e) of the SRP- FtsY complex in the presence of SecYEG but absence of RNC. Cy3-labeled Ffh was used. 12
13 RESEARCH Supplementary Figure 12 Co-localization assay of SRP and RNC. Cy3-labeled SRP (a) and Alexa647-labeled RNC FtsQ (b) colocalize before (c) and after (d) incubation with SecYEG. 13
Masayoshi Honda, Jeehae Park, Robert A. Pugh, Taekjip Ha, and Maria Spies
 Molecular Cell, Volume 35 Supplemental Data Single-Molecule Analysis Reveals Differential Effect of ssdna-binding Proteins on DNA Translocation by XPD Helicase Masayoshi Honda, Jeehae Park, Robert A. Pugh,
Molecular Cell, Volume 35 Supplemental Data Single-Molecule Analysis Reveals Differential Effect of ssdna-binding Proteins on DNA Translocation by XPD Helicase Masayoshi Honda, Jeehae Park, Robert A. Pugh,
Supplementary Figure 1: Two modes of low concentration of BsSMC on a DNA (a) Protein staining (left) and fluorescent imaging of Cy3 (right) confirm
 Supplementary Figure 1: Two modes of low concentration of BsSMC on a DNA (a) Protein staining (left) and fluorescent imaging of Cy3 (right) confirm that BsSMC was labeled with Cy3 NHS-Ester. In each panel,
Supplementary Figure 1: Two modes of low concentration of BsSMC on a DNA (a) Protein staining (left) and fluorescent imaging of Cy3 (right) confirm that BsSMC was labeled with Cy3 NHS-Ester. In each panel,
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION Conserved arginines on the rim of Hfq catalyze base pair formation and exchange Subrata Panja and Sarah A. Woodson T.C. Jenkins Department of Biophysics, Johns Hopkins University,
SUPPLEMENTARY INFORMATION Conserved arginines on the rim of Hfq catalyze base pair formation and exchange Subrata Panja and Sarah A. Woodson T.C. Jenkins Department of Biophysics, Johns Hopkins University,
Technical Review. Real time PCR
 Technical Review Real time PCR Normal PCR: Analyze with agarose gel Normal PCR vs Real time PCR Real-time PCR, also known as quantitative PCR (qpcr) or kinetic PCR Key feature: Used to amplify and simultaneously
Technical Review Real time PCR Normal PCR: Analyze with agarose gel Normal PCR vs Real time PCR Real-time PCR, also known as quantitative PCR (qpcr) or kinetic PCR Key feature: Used to amplify and simultaneously
Enhancers mutations that make the original mutant phenotype more extreme. Suppressors mutations that make the original mutant phenotype less extreme
 Interactomics and Proteomics 1. Interactomics The field of interactomics is concerned with interactions between genes or proteins. They can be genetic interactions, in which two genes are involved in the
Interactomics and Proteomics 1. Interactomics The field of interactomics is concerned with interactions between genes or proteins. They can be genetic interactions, in which two genes are involved in the
Nature Structural & Molecular Biology: doi: /nsmb.3175
 Supplementary Figure 1 Medium- and low-fret groups probably representing varying positions on the DNA show similar changes between H3 and CENP-A nucleosomes with and without CENP-C CD, as does the high-fret
Supplementary Figure 1 Medium- and low-fret groups probably representing varying positions on the DNA show similar changes between H3 and CENP-A nucleosomes with and without CENP-C CD, as does the high-fret
Contents... vii. List of Figures... xii. List of Tables... xiv. Abbreviatons... xv. Summary... xvii. 1. Introduction In vitro evolution...
 vii Contents Contents... vii List of Figures... xii List of Tables... xiv Abbreviatons... xv Summary... xvii 1. Introduction...1 1.1 In vitro evolution... 1 1.2 Phage Display Technology... 3 1.3 Cell surface
vii Contents Contents... vii List of Figures... xii List of Tables... xiv Abbreviatons... xv Summary... xvii 1. Introduction...1 1.1 In vitro evolution... 1 1.2 Phage Display Technology... 3 1.3 Cell surface
RNA polymerase pausing and nascent-rna structure formation are linked through clamp-domain movement
 a r t i c l e s RNA polymerase pausing and nascent-rna structure formation are linked through clamp-domain movement Pyae P Hein 1, Kellie E Kolb 1, Tricia Windgassen 1, Michael J Bellecourt 1, Seth A Darst
a r t i c l e s RNA polymerase pausing and nascent-rna structure formation are linked through clamp-domain movement Pyae P Hein 1, Kellie E Kolb 1, Tricia Windgassen 1, Michael J Bellecourt 1, Seth A Darst
Supplementary Figure 1 Two distinct conformational states of the HNH domain in crystal structures. a
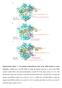 Supplementary Figure 1 Two distinct conformational states of the HNH domain in crystal structures. a HNH-state 1 in PDB 4OO8, in which the distance from the C atom of the HNH catalytic residue 840 to the
Supplementary Figure 1 Two distinct conformational states of the HNH domain in crystal structures. a HNH-state 1 in PDB 4OO8, in which the distance from the C atom of the HNH catalytic residue 840 to the
Applications and Uses. (adapted from Roche RealTime PCR Application Manual)
 What Can You Do With qpcr? Applications and Uses (adapted from Roche RealTime PCR Application Manual) What is qpcr? Real time PCR also known as quantitative PCR (qpcr) measures PCR amplification as it
What Can You Do With qpcr? Applications and Uses (adapted from Roche RealTime PCR Application Manual) What is qpcr? Real time PCR also known as quantitative PCR (qpcr) measures PCR amplification as it
Roche Molecular Biochemicals Technical Note No. LC 10/2000
 Roche Molecular Biochemicals Technical Note No. LC 10/2000 LightCycler Overview of LightCycler Quantification Methods 1. General Introduction Introduction Content Definitions This Technical Note will introduce
Roche Molecular Biochemicals Technical Note No. LC 10/2000 LightCycler Overview of LightCycler Quantification Methods 1. General Introduction Introduction Content Definitions This Technical Note will introduce
Supplemental Information. Natural RNA Polymerase Aptamers. Regulate Transcription in E. coli
 Molecular Cell, Volume 67 Supplemental Information Natural RNA Polymerase Aptamers Regulate Transcription in E. coli Nadezda Sedlyarova, Philipp Rescheneder, Andrés Magán, Niko Popitsch, Natascha Rziha,
Molecular Cell, Volume 67 Supplemental Information Natural RNA Polymerase Aptamers Regulate Transcription in E. coli Nadezda Sedlyarova, Philipp Rescheneder, Andrés Magán, Niko Popitsch, Natascha Rziha,
Product Specifications & Manual
 Product Specifications & Manual Custom Oligo Synthesis, antisense oligos, RNA oligos, chimeric oligos, Fluorescent dye labeled oligos, Molecular Beacons, sirna, phosphonates Affinity Ligands, 2-5 linked
Product Specifications & Manual Custom Oligo Synthesis, antisense oligos, RNA oligos, chimeric oligos, Fluorescent dye labeled oligos, Molecular Beacons, sirna, phosphonates Affinity Ligands, 2-5 linked
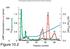
Supplementary Fig. 1. Characteristics of transcription elongation by YonO. a. YonO forms a saltstable EC. Immobilized ECs were washed with
 Supplementary Fig. 1. Characteristics of transcription elongation by YonO. a. YonO forms a saltstable EC. Immobilized ECs were washed with transcription buffer with or without a high salt concentration
Supplementary Fig. 1. Characteristics of transcription elongation by YonO. a. YonO forms a saltstable EC. Immobilized ECs were washed with transcription buffer with or without a high salt concentration
High Throughput Sequencing Technologies. UCD Genome Center Bioinformatics Core Monday 15 June 2015
 High Throughput Sequencing Technologies UCD Genome Center Bioinformatics Core Monday 15 June 2015 Sequencing Explosion www.genome.gov/sequencingcosts http://t.co/ka5cvghdqo Sequencing Explosion 2011 PacBio
High Throughput Sequencing Technologies UCD Genome Center Bioinformatics Core Monday 15 June 2015 Sequencing Explosion www.genome.gov/sequencingcosts http://t.co/ka5cvghdqo Sequencing Explosion 2011 PacBio
Quantitative Real time PCR. Only for teaching purposes - not for reproduction or sale
 Quantitative Real time PCR PCR reaction conventional versus real time PCR real time PCR principles threshold cycle C T efficiency relative quantification reference genes primers detection chemistry GLP
Quantitative Real time PCR PCR reaction conventional versus real time PCR real time PCR principles threshold cycle C T efficiency relative quantification reference genes primers detection chemistry GLP
Supporting Information
 Copyright WILEY-VCH Verlag GmbH & Co. KGaA, 69469 Weinheim, Germany, 2014. Supporting Information for Small, DOI: 10.1002/smll.201303558 Ultraspecific and Highly Sensitive Nucleic Acid Detection by Integrating
Copyright WILEY-VCH Verlag GmbH & Co. KGaA, 69469 Weinheim, Germany, 2014. Supporting Information for Small, DOI: 10.1002/smll.201303558 Ultraspecific and Highly Sensitive Nucleic Acid Detection by Integrating
Current ( pa) Current (pa) Voltage (mv) Voltage ( mv)
 Current ( pa) 3000 2000 1000 0-1000 -2000-3000 a -400-200 0 200 400 Voltage (mv) P1 P2 P3 P4 P5 P6 P7 P8 P9 P10 P11 P12 P13 P14 P15 P16 P17 P18 Average Current (pa) 4500 3000 1500 0-1500 -3000-4500 b -400-200
Current ( pa) 3000 2000 1000 0-1000 -2000-3000 a -400-200 0 200 400 Voltage (mv) P1 P2 P3 P4 P5 P6 P7 P8 P9 P10 P11 P12 P13 P14 P15 P16 P17 P18 Average Current (pa) 4500 3000 1500 0-1500 -3000-4500 b -400-200
Biology 201 (Genetics) Exam #3 120 points 20 November Read the question carefully before answering. Think before you write.
 Name KEY Section Biology 201 (Genetics) Exam #3 120 points 20 November 2006 Read the question carefully before answering. Think before you write. You will have up to 50 minutes to take this exam. After
Name KEY Section Biology 201 (Genetics) Exam #3 120 points 20 November 2006 Read the question carefully before answering. Think before you write. You will have up to 50 minutes to take this exam. After
Recent technology allow production of microarrays composed of 70-mers (essentially a hybrid of the two techniques)
 Microarrays and Transcript Profiling Gene expression patterns are traditionally studied using Northern blots (DNA-RNA hybridization assays). This approach involves separation of total or polya + RNA on
Microarrays and Transcript Profiling Gene expression patterns are traditionally studied using Northern blots (DNA-RNA hybridization assays). This approach involves separation of total or polya + RNA on
Supplementary Fig. S1. SAMHD1c has a more potent dntpase activity than. SAMHD1c. Purified recombinant SAMHD1c and SAMHD1c proteins (with
 Supplementary Fig. S1. SAMHD1c has a more potent dntpase activity than SAMHD1c. Purified recombinant SAMHD1c and SAMHD1c proteins (with concentration of 800nM) were incubated with 1mM dgtp for the indicated
Supplementary Fig. S1. SAMHD1c has a more potent dntpase activity than SAMHD1c. Purified recombinant SAMHD1c and SAMHD1c proteins (with concentration of 800nM) were incubated with 1mM dgtp for the indicated
Supplemental Figure 1. Ribose-614 cleavage is unaffected by the addition of 10%
 Supplemental Figure 1. Ribose-614 cleavage is unaffected by the addition of 1% complementary strand. Addition of equimolar amounts of the complement greatly lowers cleavage rate and plateau. (A) Cleavage
Supplemental Figure 1. Ribose-614 cleavage is unaffected by the addition of 1% complementary strand. Addition of equimolar amounts of the complement greatly lowers cleavage rate and plateau. (A) Cleavage
Chem 465 Biochemistry II
 Chem 465 Biochemistry II Name: 2 points Multiple choice (4 points apiece): 1. Which of the following is not true of trna molecules? A) The 3'-terminal sequence is -CCA. B) Their anticodons are complementary
Chem 465 Biochemistry II Name: 2 points Multiple choice (4 points apiece): 1. Which of the following is not true of trna molecules? A) The 3'-terminal sequence is -CCA. B) Their anticodons are complementary
Gene Expression Technology
 Gene Expression Technology Bing Zhang Department of Biomedical Informatics Vanderbilt University bing.zhang@vanderbilt.edu Gene expression Gene expression is the process by which information from a gene
Gene Expression Technology Bing Zhang Department of Biomedical Informatics Vanderbilt University bing.zhang@vanderbilt.edu Gene expression Gene expression is the process by which information from a gene
XactEdit Cas9 Nuclease with NLS User Manual
 XactEdit Cas9 Nuclease with NLS User Manual An RNA-guided recombinant endonuclease for efficient targeted DNA cleavage Catalog Numbers CE1000-50K, CE1000-50, CE1000-250, CE1001-250, CE1001-1000 Table of
XactEdit Cas9 Nuclease with NLS User Manual An RNA-guided recombinant endonuclease for efficient targeted DNA cleavage Catalog Numbers CE1000-50K, CE1000-50, CE1000-250, CE1001-250, CE1001-1000 Table of
Supporting Information
 Supporting Information Velocity of DNA during translocation through a solid state nanopore Calin Plesa, Nick van Loo, Philip Ketterer, Hendrik Dietz, Cees Dekker Department of Bionanoscience, Kavli Institute
Supporting Information Velocity of DNA during translocation through a solid state nanopore Calin Plesa, Nick van Loo, Philip Ketterer, Hendrik Dietz, Cees Dekker Department of Bionanoscience, Kavli Institute
DNA Transcription. Visualizing Transcription. The Transcription Process
 DNA Transcription By: Suzanne Clancy, Ph.D. 2008 Nature Education Citation: Clancy, S. (2008) DNA transcription. Nature Education 1(1) If DNA is a book, then how is it read? Learn more about the DNA transcription
DNA Transcription By: Suzanne Clancy, Ph.D. 2008 Nature Education Citation: Clancy, S. (2008) DNA transcription. Nature Education 1(1) If DNA is a book, then how is it read? Learn more about the DNA transcription
Non-Organic-Based Isolation of Mammalian microrna using Norgen s microrna Purification Kit
 Application Note 13 RNA Sample Preparation Non-Organic-Based Isolation of Mammalian microrna using Norgen s microrna Purification Kit B. Lam, PhD 1, P. Roberts, MSc 1 Y. Haj-Ahmad, M.Sc., Ph.D 1,2 1 Norgen
Application Note 13 RNA Sample Preparation Non-Organic-Based Isolation of Mammalian microrna using Norgen s microrna Purification Kit B. Lam, PhD 1, P. Roberts, MSc 1 Y. Haj-Ahmad, M.Sc., Ph.D 1,2 1 Norgen
Chromatographic Separation of the three forms of RNA Polymerase II.
 Chromatographic Separation of the three forms of RNA Polymerase II. α-amanitin α-amanitin bound to Pol II Function of the three enzymes. Yeast Pol II. RNA Polymerase Subunit Structures 10-7 Subunit structure.
Chromatographic Separation of the three forms of RNA Polymerase II. α-amanitin α-amanitin bound to Pol II Function of the three enzymes. Yeast Pol II. RNA Polymerase Subunit Structures 10-7 Subunit structure.
Supporting Information
 Supporting Information Wiley-VCH 2006 69451 Weinheim, Germany RNA ligands that distinguish metabolite-induced conformations in the TPP riboswitch Günter Mayer, Marie-Sophie L. Raddatz, Julia D. Grunwald,
Supporting Information Wiley-VCH 2006 69451 Weinheim, Germany RNA ligands that distinguish metabolite-induced conformations in the TPP riboswitch Günter Mayer, Marie-Sophie L. Raddatz, Julia D. Grunwald,
Ensemble refinement shows conformational flexibility in crystal structures of human complement factor D
 Supplementary Information for Ensemble refinement shows conformational flexibility in crystal structures of human complement factor D Federico Forneris a,b, B. Tom Burnley a,b,c and Piet Gros a * a Crystal
Supplementary Information for Ensemble refinement shows conformational flexibility in crystal structures of human complement factor D Federico Forneris a,b, B. Tom Burnley a,b,c and Piet Gros a * a Crystal
Review of Protein (one or more polypeptide) A polypeptide is a long chain of..
 Gene expression Review of Protein (one or more polypeptide) A polypeptide is a long chain of.. In a protein, the sequence of amino acid determines its which determines the protein s A protein with an enzymatic
Gene expression Review of Protein (one or more polypeptide) A polypeptide is a long chain of.. In a protein, the sequence of amino acid determines its which determines the protein s A protein with an enzymatic
DNA Transcription. Dr Aliwaini
 DNA Transcription 1 DNA Transcription-Introduction The synthesis of an RNA molecule from DNA is called Transcription. All eukaryotic cells have five major classes of RNA: ribosomal RNA (rrna), messenger
DNA Transcription 1 DNA Transcription-Introduction The synthesis of an RNA molecule from DNA is called Transcription. All eukaryotic cells have five major classes of RNA: ribosomal RNA (rrna), messenger
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION Legends for Supplementary Tables. Supplementary Table 1. An excel file containing primary screen data. Worksheet 1, Normalized quantification data from a duplicated screen: valid
SUPPLEMENTARY INFORMATION Legends for Supplementary Tables. Supplementary Table 1. An excel file containing primary screen data. Worksheet 1, Normalized quantification data from a duplicated screen: valid
Figure 1: TDP-43 is subject to lysine acetylation within the RNA-binding domain a) QBI-293 cells were transfected with TDP-43 in the presence or
 Figure 1: TDP-43 is subject to lysine acetylation within the RNA-binding domain a) QBI-293 cells were transfected with TDP-43 in the presence or absence of the acetyltransferase CBP and acetylated TDP-43
Figure 1: TDP-43 is subject to lysine acetylation within the RNA-binding domain a) QBI-293 cells were transfected with TDP-43 in the presence or absence of the acetyltransferase CBP and acetylated TDP-43
The Two-Hybrid System
 Encyclopedic Reference of Genomics and Proteomics in Molecular Medicine The Two-Hybrid System Carolina Vollert & Peter Uetz Institut für Genetik Forschungszentrum Karlsruhe PO Box 3640 D-76021 Karlsruhe
Encyclopedic Reference of Genomics and Proteomics in Molecular Medicine The Two-Hybrid System Carolina Vollert & Peter Uetz Institut für Genetik Forschungszentrum Karlsruhe PO Box 3640 D-76021 Karlsruhe
% Viability. isw2 ino isw2 ino isw2 ino isw2 ino mM HU 4-NQO CPT
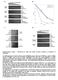 a Drug concentration b 1.3% MMS nhp1 nhp1 8 nhp1 mag1.5% MMS.3% MMS nhp1 nhp1 ino8 9 ino8 9 % Viability 4.5% MMS ino8 9 ino8 9 2.5.1.15 % MMS c d nhp1 nhp1 nhp1 nhp1 nhp1 nhp1 Control (YPD) γ IR (1 gy)
a Drug concentration b 1.3% MMS nhp1 nhp1 8 nhp1 mag1.5% MMS.3% MMS nhp1 nhp1 ino8 9 ino8 9 % Viability 4.5% MMS ino8 9 ino8 9 2.5.1.15 % MMS c d nhp1 nhp1 nhp1 nhp1 nhp1 nhp1 Control (YPD) γ IR (1 gy)
Mechanism of Transcription Termination by RNA Polymerase III Utilizes a Non-template Strand Sequence-Specific Signal Element
 Molecular Cell Supplemental Information Mechanism of ranscription ermination by RNA Polymerase III Utilizes a Non-template Strand Sequence-Specific Signal Element Aneeshkumar G. Arimbasseri and Richard
Molecular Cell Supplemental Information Mechanism of ranscription ermination by RNA Polymerase III Utilizes a Non-template Strand Sequence-Specific Signal Element Aneeshkumar G. Arimbasseri and Richard
Nature Methods: doi: /nmeth Supplementary Figure 1. Retention of RNA with LabelX.
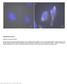 Supplementary Figure 1 Retention of RNA with LabelX. (a) Epi-fluorescence image of single molecule FISH (smfish) against GAPDH on HeLa cells expanded without LabelX treatment. (b) Epi-fluorescence image
Supplementary Figure 1 Retention of RNA with LabelX. (a) Epi-fluorescence image of single molecule FISH (smfish) against GAPDH on HeLa cells expanded without LabelX treatment. (b) Epi-fluorescence image
Deducing the kinetics of protein synthesis in vivo from the transition rates measured in vitro
 1 Deducing the kinetics of protein synthesis in vivo from the transition rates measured in vitro Supporting Information Sophia Rudorf 1, Michael Thommen 2, Marina V. Rodnina 2, and Reinhard Lipowsky 1
1 Deducing the kinetics of protein synthesis in vivo from the transition rates measured in vitro Supporting Information Sophia Rudorf 1, Michael Thommen 2, Marina V. Rodnina 2, and Reinhard Lipowsky 1
TransIT -mrna Transfection Kit
 INTRODUCTION TransIT -mrna Transfection Kit is designed to transfect RNA into a broad range of cell types with minimal cellular toxicity. RNA delivery avoids transcriptional regulation effects by directly
INTRODUCTION TransIT -mrna Transfection Kit is designed to transfect RNA into a broad range of cell types with minimal cellular toxicity. RNA delivery avoids transcriptional regulation effects by directly
Quantitation of mrna Using Real-Time Reverse Transcription PCR (RT-PCR)
 Quantitation of mrna Using Real-Time Reverse Transcription PCR (RT-PCR) Quantitative Real-Time RT-PCR Versus RT-PCR In Real-Time RT- PCR, DNA amplification monitored at each cycle but RT-PCR measures the
Quantitation of mrna Using Real-Time Reverse Transcription PCR (RT-PCR) Quantitative Real-Time RT-PCR Versus RT-PCR In Real-Time RT- PCR, DNA amplification monitored at each cycle but RT-PCR measures the
Polyclonal ARHGAP25 antibody was prepared from rabbit serum after intracutaneous
 Preparation and purification of polyclonal antibodies Polyclonal ARHGAP25 antibody was prepared from rabbit serum after intracutaneous injections of glutathione S-transferase-ARHGAP25-(509-619) (GST-coiled
Preparation and purification of polyclonal antibodies Polyclonal ARHGAP25 antibody was prepared from rabbit serum after intracutaneous injections of glutathione S-transferase-ARHGAP25-(509-619) (GST-coiled
T7-Based RNA Amplification Protocol (in progress)
 T7-Based RNA Amplification Protocol (in progress) Jacqueline Ann Lopez (modifications) Amy Cash & Justen Andrews INTRODUCTION T7 RNA Amplification, a technique originally developed in the laboratory of
T7-Based RNA Amplification Protocol (in progress) Jacqueline Ann Lopez (modifications) Amy Cash & Justen Andrews INTRODUCTION T7 RNA Amplification, a technique originally developed in the laboratory of
The GeneEditor TM in vitro Mutagenesis System: Site- Directed Mutagenesis Using Altered Beta-Lactamase Specificity
 Promega Notes Magazine Number 62, 1997, p. 02 The GeneEditor TM in vitro Mutagenesis System: Site- Directed Mutagenesis Using Altered Beta-Lactamase Specificity By Christine Andrews and Scott Lesley Promega
Promega Notes Magazine Number 62, 1997, p. 02 The GeneEditor TM in vitro Mutagenesis System: Site- Directed Mutagenesis Using Altered Beta-Lactamase Specificity By Christine Andrews and Scott Lesley Promega
The modified RNAfold program was used with molecule specific and molecule independent
 Supplemental Data The modified RNAfold program was used with molecule specific and molecule independent hairpin loop statistical potentials to predict the secondary structure for sixteen RNA molecular
Supplemental Data The modified RNAfold program was used with molecule specific and molecule independent hairpin loop statistical potentials to predict the secondary structure for sixteen RNA molecular
SUPPLEMENTARY INFORMATION. Transcriptional output transiently spikes upon mitotic exit
 SUPPLEMENTARY INFORMATION Transcriptional output transiently spikes upon mitotic exit Viola Vaňková Hausnerová 1, 2, Christian Lanctôt 1* 1 BIOCEV and Department of Cell Biology, Faculty of Science, Charles
SUPPLEMENTARY INFORMATION Transcriptional output transiently spikes upon mitotic exit Viola Vaňková Hausnerová 1, 2, Christian Lanctôt 1* 1 BIOCEV and Department of Cell Biology, Faculty of Science, Charles
Eukaryotic Transcription
 Eukaryotic Transcription I. Differences between eukaryotic versus prokaryotic transcription. II. (core vs holoenzyme): RNA polymerase II - Promotor elements. - General Pol II transcription factors (GTF).
Eukaryotic Transcription I. Differences between eukaryotic versus prokaryotic transcription. II. (core vs holoenzyme): RNA polymerase II - Promotor elements. - General Pol II transcription factors (GTF).
DNA polymerase proofreading: active site switching catalyzed by the bacteriophage T4 DNA polymerase
 5452 5463 Nucleic Acids Research, 2007, Vol. 35, No. 16 Published online 15 August 2007 doi:10.1093/nar/gkm591 DNA polymerase proofreading: active site switching catalyzed by the bacteriophage T4 DNA polymerase
5452 5463 Nucleic Acids Research, 2007, Vol. 35, No. 16 Published online 15 August 2007 doi:10.1093/nar/gkm591 DNA polymerase proofreading: active site switching catalyzed by the bacteriophage T4 DNA polymerase
Amino-allyl Dye Coupling Protocol
 Amino-allyl Dye Coupling Protocol Joseph DeRisi, June 2001 Typically, fluorescently labeled cdna is generated by incorporation of dyeconjugated nucleotide analogs during the reverse transcription process.
Amino-allyl Dye Coupling Protocol Joseph DeRisi, June 2001 Typically, fluorescently labeled cdna is generated by incorporation of dyeconjugated nucleotide analogs during the reverse transcription process.
Solutions to Quiz II
 MIT Department of Biology 7.014 Introductory Biology, Spring 2005 Solutions to 7.014 Quiz II Class Average = 79 Median = 82 Grade Range % A 90-100 27 B 75-89 37 C 59 74 25 D 41 58 7 F 0 40 2 Question 1
MIT Department of Biology 7.014 Introductory Biology, Spring 2005 Solutions to 7.014 Quiz II Class Average = 79 Median = 82 Grade Range % A 90-100 27 B 75-89 37 C 59 74 25 D 41 58 7 F 0 40 2 Question 1
Sequence-dependent Kinetic Model for Transcription Elongation by RNA Polymerase
 doi:10.1016/j.jmb.2004.08.107 J. Mol. Biol. (2004) 344, 335 349 Sequence-dependent Kinetic Model for Transcription Elongation by RNA Polymerase Lu Bai, Alla Shundrovsky and Michelle D. Wang* Department
doi:10.1016/j.jmb.2004.08.107 J. Mol. Biol. (2004) 344, 335 349 Sequence-dependent Kinetic Model for Transcription Elongation by RNA Polymerase Lu Bai, Alla Shundrovsky and Michelle D. Wang* Department
Combinatorial RNA libraries
 SELEX and Artificial Ribozymes Some definitions SELEX: Systematic Evolution of Ligands by EXponential Enrichment (alternatively: in vitro selection, in vitro evolution) Aptamer: nucleic acid ligand (from
SELEX and Artificial Ribozymes Some definitions SELEX: Systematic Evolution of Ligands by EXponential Enrichment (alternatively: in vitro selection, in vitro evolution) Aptamer: nucleic acid ligand (from
Supplementary Figure 1: sgrna library generation and the length of sgrnas for the functional screen. (a) A diagram of the retroviral vector for sgrna
 Supplementary Figure 1: sgrna library generation and the length of sgrnas for the functional screen. (a) A diagram of the retroviral vector for sgrna expression. It contains a U6-promoter-driven sgrna
Supplementary Figure 1: sgrna library generation and the length of sgrnas for the functional screen. (a) A diagram of the retroviral vector for sgrna expression. It contains a U6-promoter-driven sgrna
*To whom correspondence should be addressed. Tel/Fax: ;
 539 544 Nucleic Acids Research, 29, Vol. 37, No. 16 Published online 3 July 29 doi:1.193/nar/gkp56 DNA melting by RNA polymerase at the T7A1 promoter precedes the rate-limiting step at 378C and results
539 544 Nucleic Acids Research, 29, Vol. 37, No. 16 Published online 3 July 29 doi:1.193/nar/gkp56 DNA melting by RNA polymerase at the T7A1 promoter precedes the rate-limiting step at 378C and results
Learning Objectives. Define RNA interference. Define basic terminology. Describe molecular mechanism. Define VSP and relevance
 Learning Objectives Define RNA interference Define basic terminology Describe molecular mechanism Define VSP and relevance Describe role of RNAi in antigenic variation A Nobel Way to Regulate Gene Expression
Learning Objectives Define RNA interference Define basic terminology Describe molecular mechanism Define VSP and relevance Describe role of RNAi in antigenic variation A Nobel Way to Regulate Gene Expression
Answers to Module 1. An obligate aerobe is an organism that has an absolute requirement of oxygen for growth.
 Answers to Module 1 Short Answers 1) What is an obligate aerobe? An obligate aerobe is an organism that has an absolute requirement of oxygen for growth. What about facultative anaerobe? 2) Distinguish
Answers to Module 1 Short Answers 1) What is an obligate aerobe? An obligate aerobe is an organism that has an absolute requirement of oxygen for growth. What about facultative anaerobe? 2) Distinguish
Kinetics Review. Tonight at 7 PM Phys 204 We will do two problems on the board (additional ones than in the problem sets)
 Quiz 1 Kinetics Review Tonight at 7 PM Phys 204 We will do two problems on the board (additional ones than in the problem sets) I will post the problems with solutions on Toolkit for those that can t make
Quiz 1 Kinetics Review Tonight at 7 PM Phys 204 We will do two problems on the board (additional ones than in the problem sets) I will post the problems with solutions on Toolkit for those that can t make
BIOLOGY. Chapter 15 Genes & Proteins
 BIOLOGY Chapter 15 Genes & Proteins CMPBELL BIOLOGY TENTH EDITION Reece Urry Cain Wasserman Minorsky Jackson 17 Protein Synthesis 2014 Pearson Education, Inc. Fig. 17-1 Figure 17.1a n albino racoon Condition
BIOLOGY Chapter 15 Genes & Proteins CMPBELL BIOLOGY TENTH EDITION Reece Urry Cain Wasserman Minorsky Jackson 17 Protein Synthesis 2014 Pearson Education, Inc. Fig. 17-1 Figure 17.1a n albino racoon Condition
qpcr Quantitative PCR or Real-time PCR Gives a measurement of PCR product at end of each cycle real time
 qpcr qpcr Quantitative PCR or Real-time PCR Gives a measurement of PCR product at end of each cycle real time Differs from endpoint PCR gel on last cycle Used to determines relative amount of template
qpcr qpcr Quantitative PCR or Real-time PCR Gives a measurement of PCR product at end of each cycle real time Differs from endpoint PCR gel on last cycle Used to determines relative amount of template
Molecular Cloning. Joseph Sambrook. David W. Russell A LABORATORY MANUAL COLD SPRING HARBOR LABORATORY PRESS VOLUME.
 VOLUME Molecular Cloning A LABORATORY MANUAL THIRD EDITION www.molecularcloning.com Joseph Sambrook PETER MACCALLUM CANCER INSTITUTE AND THE UNIVERSITY OF MELBOURNE, AUSTRALIA David W. Russell UNIVERSITY
VOLUME Molecular Cloning A LABORATORY MANUAL THIRD EDITION www.molecularcloning.com Joseph Sambrook PETER MACCALLUM CANCER INSTITUTE AND THE UNIVERSITY OF MELBOURNE, AUSTRALIA David W. Russell UNIVERSITY
Electrophoresis 101 Student Worksheet
 1 Electrophoresis 101 Student Worksheet Experiment Objective To develop an understanding of electrophoresis principles. To analyze results and to calculate the sizes of unknown charged molecules from given
1 Electrophoresis 101 Student Worksheet Experiment Objective To develop an understanding of electrophoresis principles. To analyze results and to calculate the sizes of unknown charged molecules from given
SUPPLEMENTARY MATERIAL
 SUPPLEMENTARY MATERIAL Materials and Methods Circular dichroism (CD) spectroscopy. Far ultraviolet (UV) CD spectra of apo- and holo- CaM and the CaM mutants were recorded on a Jasco J-715 spectropolarimeter
SUPPLEMENTARY MATERIAL Materials and Methods Circular dichroism (CD) spectroscopy. Far ultraviolet (UV) CD spectra of apo- and holo- CaM and the CaM mutants were recorded on a Jasco J-715 spectropolarimeter
Spontaneous Intersubunit Rotation in Single Ribosomes
 Article Spontaneous Intersubunit Rotation in Single Ribosomes Peter V. Cornish, 1,4 Dmitri N. Ermolenko, 3,4 Harry F. Noller, 3, * and Taekjip Ha 1,2, * 1 Department of Physics 2 Howard Hughes Medical
Article Spontaneous Intersubunit Rotation in Single Ribosomes Peter V. Cornish, 1,4 Dmitri N. Ermolenko, 3,4 Harry F. Noller, 3, * and Taekjip Ha 1,2, * 1 Department of Physics 2 Howard Hughes Medical
foodproof Sample Preparation Kit III
 For food testing purposes FOR IN VITRO USE ONLY Version 1, June 2015 For isolation of DNA from raw material and food products of plant and animal origin for PCR analysis Order No. S 400 06.1 Kit for 50
For food testing purposes FOR IN VITRO USE ONLY Version 1, June 2015 For isolation of DNA from raw material and food products of plant and animal origin for PCR analysis Order No. S 400 06.1 Kit for 50
INDEX KIT COMPONENTS 1 STORAGE AND STABILITY 1 INTRODUCTION 1 BENEFITS
 INDEX KIT COMPONENTS 1 STORAGE AND STABILITY 1 INTRODUCTION 1 BENEFITS ERRORE. IL SEGNALIBRO NON È DEFINITO. IMPORTANT NOTES 1 METHOD FOR DNA EXTRACTION FROM AGAROSE GELS 2 METHOD FOR DNA EXTRACTION FROM
INDEX KIT COMPONENTS 1 STORAGE AND STABILITY 1 INTRODUCTION 1 BENEFITS ERRORE. IL SEGNALIBRO NON È DEFINITO. IMPORTANT NOTES 1 METHOD FOR DNA EXTRACTION FROM AGAROSE GELS 2 METHOD FOR DNA EXTRACTION FROM
microrna-122 stimulates translation of Hepatitis C Virus RNA
 Supplementary information for: microrna-122 stimulates translation of Hepatitis C Virus RNA Jura Inga Henke 1,5, Dagmar Goergen 1,5, Junfeng Zheng 3, Yutong Song 4, Christian G. Schüttler 2, Carmen Fehr
Supplementary information for: microrna-122 stimulates translation of Hepatitis C Virus RNA Jura Inga Henke 1,5, Dagmar Goergen 1,5, Junfeng Zheng 3, Yutong Song 4, Christian G. Schüttler 2, Carmen Fehr
Prokaryotic Transcription
 Prokaryotic Transcription Contents 1 The Lactose Intolerance of Bacteria 2 The Lac Operon 3 Lac Operon Simulation 4 LacZ as a reporter gene The Lactose Intolerance of Bacteria The standard growth kinetics
Prokaryotic Transcription Contents 1 The Lactose Intolerance of Bacteria 2 The Lac Operon 3 Lac Operon Simulation 4 LacZ as a reporter gene The Lactose Intolerance of Bacteria The standard growth kinetics
TECHNICAL BULLETIN. in vitro Translation Labeling System. FluoroTect Green Lys. Instruc ons for Use of Product L5001. Revised 11/13 TB285
 TECHNICAL BULLETIN in vitro Translation Labeling System Instruc ons for Use of Product L5001 Revised 11/13 TB285 in vitro Translation Labeling System All technical literature is available at: www.promega.com/protocols/
TECHNICAL BULLETIN in vitro Translation Labeling System Instruc ons for Use of Product L5001 Revised 11/13 TB285 in vitro Translation Labeling System All technical literature is available at: www.promega.com/protocols/
Figure S1: NUN preparation yields nascent, unadenylated RNA with a different profile from Total RNA.
 Summary of Supplemental Information Figure S1: NUN preparation yields nascent, unadenylated RNA with a different profile from Total RNA. Figure S2: rrna removal procedure is effective for clearing out
Summary of Supplemental Information Figure S1: NUN preparation yields nascent, unadenylated RNA with a different profile from Total RNA. Figure S2: rrna removal procedure is effective for clearing out
HiPer Real-Time PCR Teaching Kit
 HiPer Real-Time PCR Teaching Kit Product Code: HTBM032 Number of experiments that can be performed: 10 Duration of Experiment Protocol: 1.5 hours Storage Instructions: The kit is stable for 12 months from
HiPer Real-Time PCR Teaching Kit Product Code: HTBM032 Number of experiments that can be performed: 10 Duration of Experiment Protocol: 1.5 hours Storage Instructions: The kit is stable for 12 months from
Genetics module. DNA Structure, Replication. The Genetic Code; Transcription and Translation. Principles of Heredity; Gene Mapping
 Genetics module Lectures DNA Structure, Replication The Genetic Code; Transcription and Translation Principles of Heredity; Gene Mapping Controlling Gene Expression Mutation and Cancer Textbook: Introduction
Genetics module Lectures DNA Structure, Replication The Genetic Code; Transcription and Translation Principles of Heredity; Gene Mapping Controlling Gene Expression Mutation and Cancer Textbook: Introduction
SUPPLEMENTARY INFORMATION
 a before amputation regeneration regenerated limb DERMIS SKELETON MUSCLE SCHWANN CELLS EPIDERMIS DERMIS SKELETON MUSCLE SCHWANN CELLS EPIDERMIS developmental origin: lateral plate mesoderm presomitic mesoderm
a before amputation regeneration regenerated limb DERMIS SKELETON MUSCLE SCHWANN CELLS EPIDERMIS DERMIS SKELETON MUSCLE SCHWANN CELLS EPIDERMIS developmental origin: lateral plate mesoderm presomitic mesoderm
Molecular Mechanisms of Transcription through Single-Molecule Experiments
 This is an open access article published under an ACS AuthorChoice License, which permits copying and redistribution of the article or any adaptations for non-commercial purposes. pubs.acs.org/cr Molecular
This is an open access article published under an ACS AuthorChoice License, which permits copying and redistribution of the article or any adaptations for non-commercial purposes. pubs.acs.org/cr Molecular
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:10.1038/nature09937 a Name Position Primersets 1a 1b 2 3 4 b2 Phenotype Genotype b Primerset 1a D T C R I E 10000 8000 6000 5000 4000 3000 2500 2000 1500 1000 800 Donor (D)
SUPPLEMENTARY INFORMATION doi:10.1038/nature09937 a Name Position Primersets 1a 1b 2 3 4 b2 Phenotype Genotype b Primerset 1a D T C R I E 10000 8000 6000 5000 4000 3000 2500 2000 1500 1000 800 Donor (D)
HiPer RT-PCR Teaching Kit
 HiPer RT-PCR Teaching Kit Product Code: HTBM024 Number of experiments that can be performed: 5 Duration of Experiment: Protocol: 4 hours Agarose Gel Electrophoresis: 45 minutes Storage Instructions: The
HiPer RT-PCR Teaching Kit Product Code: HTBM024 Number of experiments that can be performed: 5 Duration of Experiment: Protocol: 4 hours Agarose Gel Electrophoresis: 45 minutes Storage Instructions: The
Supplementary data Table 1
 Supplementary data Table 1 Amino acid sequences, antimicrobial effects, and hydrophobicity of peptides analysed. For determination of antimicrobial activities, S. aureus ATCC 29213, E. coli ATCC 25922
Supplementary data Table 1 Amino acid sequences, antimicrobial effects, and hydrophobicity of peptides analysed. For determination of antimicrobial activities, S. aureus ATCC 29213, E. coli ATCC 25922
RNA synthesis/transcription I Biochemistry 302. February 6, 2004 Bob Kelm
 RNA synthesis/transcription I Biochemistry 302 February 6, 2004 Bob Kelm Overview of RNA classes Messenger RNA (mrna) Encodes protein Relatively short half-life ( 3 min in E. coli, 30 min in eukaryotic
RNA synthesis/transcription I Biochemistry 302 February 6, 2004 Bob Kelm Overview of RNA classes Messenger RNA (mrna) Encodes protein Relatively short half-life ( 3 min in E. coli, 30 min in eukaryotic
Protein Synthesis Notes
 Protein Synthesis Notes Protein Synthesis: Overview Transcription: synthesis of mrna under the direction of DNA. Translation: actual synthesis of a polypeptide under the direction of mrna. Transcription
Protein Synthesis Notes Protein Synthesis: Overview Transcription: synthesis of mrna under the direction of DNA. Translation: actual synthesis of a polypeptide under the direction of mrna. Transcription
Coleman et al., Supplementary Figure 1
 Coleman et al., Supplementary Figure 1 BrdU Merge G1 Early S Mid S Supplementary Figure 1. Sequential destruction of CRL4 Cdt2 targets during the G1/S transition. HCT116 cells were synchronized by sequential
Coleman et al., Supplementary Figure 1 BrdU Merge G1 Early S Mid S Supplementary Figure 1. Sequential destruction of CRL4 Cdt2 targets during the G1/S transition. HCT116 cells were synchronized by sequential
MODULE 1: INTRODUCTION TO THE GENOME BROWSER: WHAT IS A GENE?
 MODULE 1: INTRODUCTION TO THE GENOME BROWSER: WHAT IS A GENE? Lesson Plan: Title Introduction to the Genome Browser: what is a gene? JOYCE STAMM Objectives Demonstrate basic skills in using the UCSC Genome
MODULE 1: INTRODUCTION TO THE GENOME BROWSER: WHAT IS A GENE? Lesson Plan: Title Introduction to the Genome Browser: what is a gene? JOYCE STAMM Objectives Demonstrate basic skills in using the UCSC Genome
AmpliScribe T7-Flash Transcription Kit
 AmpliScribe T7-Flash Transcription Kit Cat. Nos. ASF3257 and ASF3507 Available exclusively thru Lucigen. lucigen.com/epibio www.lucigen.com MA191E AmpliScribe T7-Flash Transcription Kit 12/2016 1 1. Introduction
AmpliScribe T7-Flash Transcription Kit Cat. Nos. ASF3257 and ASF3507 Available exclusively thru Lucigen. lucigen.com/epibio www.lucigen.com MA191E AmpliScribe T7-Flash Transcription Kit 12/2016 1 1. Introduction
Rat IGF-1 ELISA Kit (rigf-1-elisa)
 Rat IGF-1 ELISA Kit (rigf-1-elisa) Cat. No. EK0377 96 Tests in 8 x 12 divisible strips Background Insulin-like growth factor 1 (IGF-1), also known as somatomedin C, is a polypeptide protein hormone similar
Rat IGF-1 ELISA Kit (rigf-1-elisa) Cat. No. EK0377 96 Tests in 8 x 12 divisible strips Background Insulin-like growth factor 1 (IGF-1), also known as somatomedin C, is a polypeptide protein hormone similar
Fluorescence Imaging with One Nanometer Accuracy Lab
 I. Introduction. Fluorescence Imaging with One Nanometer Accuracy Lab Traditional light microscope is limited by the diffraction limit of light, typically around 250 nm. However, many biological processes
I. Introduction. Fluorescence Imaging with One Nanometer Accuracy Lab Traditional light microscope is limited by the diffraction limit of light, typically around 250 nm. However, many biological processes
Supplementary Figures
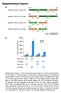 Supplementary Figures a HindIII XbaI pcdna3.1/v5-his A_Nfluc-VVD 1-398 37-186 Nfluc Linker 1 VVD HindIII XhoI pcdna3.1/v5-his A_VVD-Nfluc 37-186 1-398 VVD Linker 2 Nfluc HindIII XbaI pcdna3.1/v5-his A_Cfluc-VVD
Supplementary Figures a HindIII XbaI pcdna3.1/v5-his A_Nfluc-VVD 1-398 37-186 Nfluc Linker 1 VVD HindIII XhoI pcdna3.1/v5-his A_VVD-Nfluc 37-186 1-398 VVD Linker 2 Nfluc HindIII XbaI pcdna3.1/v5-his A_Cfluc-VVD
RayBio Human NF-κB p65 Transcription Factor Activity Assay Kit
 RayBio Human NF-κB p65 Transcription Factor Activity Assay Kit Catalog #: TFEH-p65 User Manual Mar 13, 2017 3607 Parkway Lane, Suite 200 Norcross, GA 30092 Tel: 1-888-494-8555 (Toll Free) or 770-729-2992,
RayBio Human NF-κB p65 Transcription Factor Activity Assay Kit Catalog #: TFEH-p65 User Manual Mar 13, 2017 3607 Parkway Lane, Suite 200 Norcross, GA 30092 Tel: 1-888-494-8555 (Toll Free) or 770-729-2992,
Computational Biology I LSM5191 (2003/4)
 Computational Biology I LSM5191 (2003/4) Aylwin Ng, D.Phil Lecture Notes: Transcriptome: Molecular Biology of Gene Expression I Flow of information: DNA to polypeptide DNA Start Exon1 Intron Exon2 Termination
Computational Biology I LSM5191 (2003/4) Aylwin Ng, D.Phil Lecture Notes: Transcriptome: Molecular Biology of Gene Expression I Flow of information: DNA to polypeptide DNA Start Exon1 Intron Exon2 Termination
DNA supercoiling, a critical signal regulating the basal expression of the lac operon in Escherichia coli
 Supplementary Information DNA supercoiling, a critical signal regulating the basal expression of the lac operon in Escherichia coli Geraldine Fulcrand 1,2, Samantha Dages 1,2, Xiaoduo Zhi 1,2, Prem Chapagain
Supplementary Information DNA supercoiling, a critical signal regulating the basal expression of the lac operon in Escherichia coli Geraldine Fulcrand 1,2, Samantha Dages 1,2, Xiaoduo Zhi 1,2, Prem Chapagain
phab Amine and Thiol Reactive Dyes for Antibody Internalization Studies Nidhi Nath, Ph.D. Group Leader, Protein Analysis Promega Corporation
 phab Amine and Thiol Reactive Dyes for Antibody Internalization Studies Nidhi Nath, Ph.D. Group Leader, Protein Analysis 1 Outline 1. phab Dyes 2. Protocols for conjugating phab Dyes to antibodies 3. Applications:
phab Amine and Thiol Reactive Dyes for Antibody Internalization Studies Nidhi Nath, Ph.D. Group Leader, Protein Analysis 1 Outline 1. phab Dyes 2. Protocols for conjugating phab Dyes to antibodies 3. Applications:
AmpliScribe T 7 Aminoallyl-RNA Transcription Kit
 Cat. No. AA50125 The AmpliScribe T7 Aminoallyl-RNA Transcription Kit enables high-yield production of aminoallyl-labeled RNA. The kit utilizes Epicentre s high yielding AmpliScribe T7-Flash in vitro transcription
Cat. No. AA50125 The AmpliScribe T7 Aminoallyl-RNA Transcription Kit enables high-yield production of aminoallyl-labeled RNA. The kit utilizes Epicentre s high yielding AmpliScribe T7-Flash in vitro transcription
Supporting Information
 Supporting Information Yuan et al. 10.1073/pnas.0906869106 Fig. S1. Heat map showing that Populus ICS is coregulated with orthologs of Arabidopsis genes involved in PhQ biosynthesis and PSI function, but
Supporting Information Yuan et al. 10.1073/pnas.0906869106 Fig. S1. Heat map showing that Populus ICS is coregulated with orthologs of Arabidopsis genes involved in PhQ biosynthesis and PSI function, but
42 fl organelles = 34.5 fl (1) 3.5X X 0.93 = 78,000 (2)
 SUPPLEMENTAL DATA Supplementary Experimental Procedures Fluorescence Microscopy - A Zeiss Axiovert 200M microscope equipped with a Zeiss 100x Plan- Apochromat (1.40 NA) DIC objective and Hamamatsu Orca
SUPPLEMENTAL DATA Supplementary Experimental Procedures Fluorescence Microscopy - A Zeiss Axiovert 200M microscope equipped with a Zeiss 100x Plan- Apochromat (1.40 NA) DIC objective and Hamamatsu Orca
interaction of MinE Supplementary Information
 Nature Structural & Molecular Biology doi:1.138/nsmb.237 Min protein patterns emerge from rapid rebinding and membrane interaction of MinE Supplementary Information Martin Loose, 1,2 Elisabeth Fischer-Friedrich,
Nature Structural & Molecular Biology doi:1.138/nsmb.237 Min protein patterns emerge from rapid rebinding and membrane interaction of MinE Supplementary Information Martin Loose, 1,2 Elisabeth Fischer-Friedrich,
SIBC504: TRANSCRIPTION & RNA PROCESSING Assistant Professor Dr. Chatchawan Srisawat
 SIBC504: TRANSCRIPTION & RNA PROCESSING Assistant Professor Dr. Chatchawan Srisawat TRANSCRIPTION: AN OVERVIEW Transcription: the synthesis of a single-stranded RNA from a doublestranded DNA template.
SIBC504: TRANSCRIPTION & RNA PROCESSING Assistant Professor Dr. Chatchawan Srisawat TRANSCRIPTION: AN OVERVIEW Transcription: the synthesis of a single-stranded RNA from a doublestranded DNA template.
RNA Clean-Up and Concentration Kit Product # 23600, 43200
 3430 Schmon Parkway Thorold, ON, Canada L2V 4Y6 Phone: 866-667-4362 (905) 227-8848 Fax: (905) 227-1061 Email: techsupport@norgenbiotek.com RNA Clean-Up and Concentration Kit Product # 23600, 43200 Product
3430 Schmon Parkway Thorold, ON, Canada L2V 4Y6 Phone: 866-667-4362 (905) 227-8848 Fax: (905) 227-1061 Email: techsupport@norgenbiotek.com RNA Clean-Up and Concentration Kit Product # 23600, 43200 Product
Dynamic protein assembly by programmable DNA strand displacement
 SUPPLEMENTARY INFORMATION Articles https://doi.org/10.1038/s41557-018-0016-9 In the format provided by the authors and unedited. Dynamic protein assembly by programmable DNA strand displacement Rebecca
SUPPLEMENTARY INFORMATION Articles https://doi.org/10.1038/s41557-018-0016-9 In the format provided by the authors and unedited. Dynamic protein assembly by programmable DNA strand displacement Rebecca
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Origin use and efficiency are similar among WT, rrm3, pif1-m2, and pif1-m2; rrm3 strains. A. Analysis of fork progression around confirmed and likely origins (from cerevisiae.oridb.org).
Supplementary Figure 1 Origin use and efficiency are similar among WT, rrm3, pif1-m2, and pif1-m2; rrm3 strains. A. Analysis of fork progression around confirmed and likely origins (from cerevisiae.oridb.org).
Zool 3200: Cell Biology Exam 3 3/6/15
 Name: Trask Zool 3200: Cell Biology Exam 3 3/6/15 Answer each of the following questions in the space provided; circle the correct answer or answers for each multiple choice question and circle either
Name: Trask Zool 3200: Cell Biology Exam 3 3/6/15 Answer each of the following questions in the space provided; circle the correct answer or answers for each multiple choice question and circle either
MIT Department of Biology 7.013: Introductory Biology - Spring 2005 Instructors: Professor Hazel Sive, Professor Tyler Jacks, Dr.
 MIT Department of Biology 7.01: Introductory Biology - Spring 2005 Instructors: Professor Hazel Sive, Professor Tyler Jacks, Dr. Claudette Gardel iv) Would Xba I be useful for cloning? Why or why not?
MIT Department of Biology 7.01: Introductory Biology - Spring 2005 Instructors: Professor Hazel Sive, Professor Tyler Jacks, Dr. Claudette Gardel iv) Would Xba I be useful for cloning? Why or why not?
Chapter 18: Regulation of Gene Expression. 1. Gene Regulation in Bacteria 2. Gene Regulation in Eukaryotes 3. Gene Regulation & Cancer
 Chapter 18: Regulation of Gene Expression 1. Gene Regulation in Bacteria 2. Gene Regulation in Eukaryotes 3. Gene Regulation & Cancer Gene Regulation Gene regulation refers to all aspects of controlling
Chapter 18: Regulation of Gene Expression 1. Gene Regulation in Bacteria 2. Gene Regulation in Eukaryotes 3. Gene Regulation & Cancer Gene Regulation Gene regulation refers to all aspects of controlling
