|
|
|
- Shannon Richards
- 6 years ago
- Views:
Transcription
1
2
3
4
5
6
7
8
9
10 Supplemental Table 1. The progeny of crosses between MPK4/mpk4-2 and ANQ/anq-2. Genotype Number Number Number observed expected-1 expected-2 (1) MPK4/MPK4 ANQ/ANQ (2) MPK4/MPK4 ANQ/anq (3) MPK4/MPK4 anq-2/anq (4) MPK4/mpk4-2 ANQ/ANQ (5) MPK4/mpk4-2 ANQ/anq (6) MPK4/mpk4-2 anq-2/anq (7) mpk4-2/mpk4-2 ANQ/ANQ (8) mpk4-2/mpk4-2 ANQ/anq (9) mpk4-2/mpk4-2 anq-2/anq Total number P-value 1.5E-12<0.05* 0.46>0.05 The genotypes of 197 progeny plants with the respective indicated genotypes were determined by PCR. Expected values were calculated using Mendel s law (expected-1). Expected values were also calculated by reference to the viability of the female gametophyte and/or the embryo, as summarized in Table 2 (expected-2). P-values were calculated by the χ 2 -test from observed and expected values. A P- value indicative of a significant difference is
11 indicated by an asterisk.
12 Supplemental Table 2. The frequency of cells with an incomplete cell plate. Genotype Number of Number of The frequency of cells with an cells in cells with an incomplete cell one incomplete cell plate plate in one section (b) (%) section (a) Col-0 # Col-0 # Col-0 # Col-0 # Col-0 # Col-0 # Average frequency 0 ± 0 mpk13 # mpk13 # mpk13 # mpk13 # mpk13 # mpk13 # Average frequency 0 ± 0 mpk4-2 #
13 mpk4-2 # mpk4-2 # mpk4-2 # mpk4-2 # mpk4-2 # Average frequency 2.5 ± 0.62 P-value * 2.0E-06<0.05 mpk4-2 mpk13 # mpk4-2 mpk13 # mpk4-2 mpk13 # mpk4-2 mpk13 # mpk4-2 mpk13 # mpk4-2 mpk13 # Average frequency 2.1 ± 0.61 P-value * 7.0E-06<0.05 P-value ** 0.37>0.05 (a-b) Transverse sections (5 µm) were prepared from cotyledons of 9-day-old Col-0, mpk13, mpk4-2, and mpk4-2 mpk13 seedlings. The number of cells and the number of cells with incomplete cell plates were counted in a typical single section from cotyledons of six homozygous siblings (#1 to #6). Average frequencies are presented as means ± SD (n = 6). P-values were calculated by Student s t-test. A single asterisk indicates comparison with Col-0. Two asterisks indicate comparison with mpk4-2.
14 Supplemental Table 3. Primers used in this study. Application Sequence (5-3 ) (forward/reverse) Genotyping of MPK4 ATTAAGCTTCTCAAGCATATGGATCATGA/ TCCTCCTTGAAATATCTACAGAGTTGGTGT Genotyping of mpk4-1 mutant CCGGATCGTATCGGTTTTCG /CTTGAGTCATCAGGAGATCC Genotyping of mpk4-2 mutant GCGTGGACCGCTTGCTGCAACT/ TCCTCCTTGAAATATCTACAGAGTTGGTGT Genotyping of ANQ/MKK6 CTACGCGTCTTGAGATCTTG/ GGATTGAGAGAAAGCGTTAAGATCAGAG Genotyping of anq-2 mutant GCGTGGACCGCTTGCTGCAACT/ GGATTGAGAGAAAGCGTTAAGATCAGAG Genotyping of MPK13 GAAGATGGAGGGATCTTGAC/ CAGACAAGCTCCTTGATGTC Genotyping of mpk13 mutant GAAGATGGAGGGATCTTGAC/ CATTTTATAATAACGCTGCGGACATCTAC
15 Genotyping of MPK11 CACACTAAGAATCTTAAGGGTTAAACTTGACT G/ATGTCAATAGAGAAACCATTCTTCGGTG Genotyping of mpk11 mutant GCGTGGACCGCTTGCTGCAACT /ATGTCAATAGAGAAACCATTCTTCGGTG Genotyping of NACK1 GCGGCCATATACCTTACAGA/ GCTCTCCGATTTCCATTTCCA Genotyping of nack1-3 mutant GCGGCCATATACCTTACAGA/ GCGTGGACCGCTTGCTGCAACT RT-PCR analysis of MPK4 TGCTCTGAATACACAGCAGC/ CACACTGAGTCTTGAGGATTG Quantitative RT-PCR analysis β-6-tubulin ACCACTCCTAGCTTTGGTGATCTG/ AGGTTCACTGCGAGCTTCCTCA Quantitative RT-PCR analysis EF1-a TGAGCACGCTCTTCTTGCTTTCA/ GGTGGTGGCATCCATCTTGTTACA Quantitative RT-PCR analysis of MPK4 CACAGTTGATGAGGCGTTGTG/ CACATACCGGTTCTTCGTTGATAT Quantitative RT-PCR analysis of MPK5 ACGGTTGAGGAAGCACTGTGT/ GAACAAACAGGCTCGTCGTTAA
16 Quantitative RT-PCR analysis of MPK11 TTTGACCCAAACAGACGCATT/ TCGTGTAGCGGTGCTAAGTAAGG Quantitative RT-PCR analysis of MPK12 CCTAACCGGCGCATCTCA/ TCATGGTGCGGTGATAGGTAAG Quantitative RT-PCR analysis of MPK13 K4_5p_F GCATTGAAGCAGCCGTACTTG/ CAAAGCTGAAAGGAGTAGGACATG GGTACCGACTTGTTTGTGAATATAGAGGAAA CATG (Underline = Kpn1 site) K4_5p_R GGATCCCACTGAGTCTTGAGGATTGAACTTGA C (Underline = BamH1 site) K4_3p_F GGATCCGAGAAAGAGAGAGAGATATATATCC (Underline = BamH1 site) K4_3p_R GCGGCCGCGATAATTAGTGGATGTAATTAGA GTTAAGAC YFP_F GGATCCATGGTGAGCAAGGGCGAGGAGCTG (Underline = BamH1 site) K4WT_S K4WT_R LB6316 RT-PCR analysis of MPK11 GTGACAATGCAAGAAGATACGTTAGACAGC CTTGAAATATCTACAGAGTTGGTGTG TCAAACAGGATTTTCGCCTGCT AACCGAGTAACTTGCTCCTG CACAAGGCTTCATCGACTGT
17 KK 93 GCGGCCGCGACACTGAGTCTTGAGGATTGAA KK 138 GTCGACGATCTCGAGGCTGTTGAAAC (Underline = Sal1 site) Cloning of MPK4 CCATGGCGGCGGAGAGTTGTTTCG GCGGCCGCCACTGAGTCTTGAGGATTGAA Cloning of MPK5 CCATGGCGAAGGAAATTGAATCA GCGGCCGCAATGCTCGGCAGAGGATTGAA Cloning of MPK11 CCATGGCAATAGAGAAACCATTC GCGGCCGCAGGGTTAAACTTGACTGATTC Cloning of MPK12 CCATGGCTGGAGAATCAAGCTCT GCGGCCGCGTGGTCAGGATTGAATTTGACAGA Cloning of MPK13 CCATGGAGAAAAGGGAAGATGGA GCGGCCGCCATATTCTTGAAGTGTAAAGACTCT
18 Cloning of MPK1 CCATGGCGACTTTGGTTGATCC GCGGCCGCGAGCTCAGTGTTTAAGGTTG Cloning of MPK6 CCATGGACGGTGGTTCAGGTC GCGGCCGCTTGCTGATATTCTGGATTGAAA Cloning of MKK6/ANQ CCATGGTGAAGATCAAATCGAA GCGGCCGCTCTAAGGTAGTTAACAGGTG Cloning of MPK2 CCATGGCGACTCCTGTTGATCC GCGGCCGCAAACTCAGAGACCTCATTGT Cloning of MPK3 CCATGGACACCGGCGGTGGCCA GCGGCCGCACCGTATGTTGGATTGAGTG Cloning of MPK7 CCATGGCGATGTTAGTTGAGCC GCGGCCGCGGCATTTGAGATTTCAGCTT
19 Cloning of MPK8 CCATGGGTGGTGGTGGGAATCT GCGGCCGCAGAATTGTGAAGAGAAGCAA Cloning of MPK14 CCATGGCGATGCTAGTTGATCC GCGGCCGCAGCTCGGGGGAGGTAATGAA Cloning of MPK15 CCATGGGTGGTGGTGGCAATCT GCGGCCGCAGAATTGTGTAGAGATGCAA Cloning of MPK16 CCATGGAGCCTGATCACCGCAA GCGGCCGCATACCAGCGACTCATTGCAG Cloning of MPK17 CCATGGTGGAGAAAGAGTTTTT GCGGCCGCTGACACTGCAGAGGAGACAC Cloning of MPK18 CCATGGAACAAAATCAAGTGAA GCGGCCGCTGATGCTGCGCTGTAACTAA
20 Cloning of MPK19 CCATGGAAAAAACTCAGGAGAA GCGGCCGCAGACATGCCATACCCAACAG Supplemental Methods RT-PCR We used 15-day-old seedlings for analyses by RT-PCR, which was performed as described in Methods. Primers used for the experiments are listed in Supplemental Table 3. Plant materials and growth conditions Tobacco suspension-cultured BY-2 cells were maintained as described elsewhere (Banno et al., 1993; Nishihama et al., 2001). Transformed BY-2 cells were generated by Agrobacterium-mediated transformation (An. 1985). Fluorescence microscopy To generate the MPK4-GFP fusions, MPK4-coding regions without a termination codon were cloned into the pgwb5 vector. To generate the GFP-MAP65-3 fusions, MAP65-3-coding regions were cloned into the pgwb6 vector (Nakagawa et al. 2001). Living BY-2 cells that expressed MPK4-GFP were stained with 0.5 µg/ml Hoechst (Dojin, Kumamoto, Japan). Aniline blue (0.1% in 0.1 M K 3 PO 4, ph 11) was added to roots of 4-day-old plants on a Gellan gum plate and imaged after a 5-min incubation.
21 All techniques for microscopic visualization and analysis were described previously (Nishihama et al., 2001; Tanaka et al., 2001). Production and purification of recombinant proteins His-T7-tagged-MPK4, GST-fused MKK6/ANQ, and GST-fused-MAP65-3/PLE proteins were produced in E. coli BL21 using pet28 (Novagen) and pgex (GE Healthcare) vectors. His-T7-MPK4 was purified with Ni Sepharose TM 6 Fast Flow (GE Healthcare). GST-NQK1 and GST-MAP65-3 proteins were purified with Glutathione-Sepharose TM 4B (GE Healthcare). Phosphorylation assays in vitro His-T7-MPK4 and GST-MKK6/ANQ were incubated in 200 µl of kinase buffer (as described in Methods) that contained 500 µm ATP at 25 C for 120 min. Reaction products, which included His-T7-MPK4 (150 ng) and GST-MKK6/ANQ (225 ng), were incubated with GST-MAP65-3-PLE (1 mg) in 20 µl of kinase buffer that contained 50 µm ATP and 10 µci of [γ- 32 P] ATP (GE Healthcare) at 25 C for 60 min. Reactions were terminated by the addition of 10 ml of 3 SDS sample buffer and the mixtures were boiled. The samples were subjected to SDS-PAGE and radioactivity was visualized with an imaging analyzer (BAS-1800; Fujifilm, Tokyo, Japan).
22 Supplemental References An, G. (1985). High Efficiency Transformation of Cultured Tobacco Cells. Plant Physiol. 79: Banno, H., Hirano, K., Nakamura, T., Irie, K., Nomoto, S., Matsumoto, K., and Machida, Y. (1993). NPK1, a tobacco gene that encodes a protein with a domain homologous to yeast BCK1, STE11, and Byr2 protein kinases. Mol. Cell. Biol. 13: Nakagawa, T., Kurose, T., Hino, T., Tanaka, K., Kawamukai, M., Niwa, Y., Toyooka, K., Matsuoka, K., Jinbo, T., and Kimura, T. (2007). Development of series of gateway binary vectors, pgwbs, for realizing efficient construction of fusion genes for plant transformation. J. Biosci. Bioeng. 104: Tanaka, H., Onouchi, H., Kondo, M., Hara-Nishimura, I., Nishimura, M., Machida, C., and Machida, Y. (2001). A subtilisin-like serine protease is required for epidermal surface formation in Arabidopsis embryos and juvenile plants. Development 128: Nishihama, R., Ishikawa, M., Araki, S., Soyano, T., Asada, T., and Machida, Y. (2001). The NPK1 mitogen-activated protein kinase kinase kinase is a regulator of cell-plate formation in plant cytokinesis. Genes Dev. 15:
At E17.5, the embryos were rinsed in phosphate-buffered saline (PBS) and immersed in
 Supplementary Materials and Methods Barrier function assays At E17.5, the embryos were rinsed in phosphate-buffered saline (PBS) and immersed in acidic X-gal mix (100 mm phosphate buffer at ph4.3, 3 mm
Supplementary Materials and Methods Barrier function assays At E17.5, the embryos were rinsed in phosphate-buffered saline (PBS) and immersed in acidic X-gal mix (100 mm phosphate buffer at ph4.3, 3 mm
Supplemental Data. Lee et al. Plant Cell. (2010) /tpc Supplemental Figure 1. Protein and Gene Structures of DWA1 and DWA2.
 Supplemental Figure 1. Protein and Gene Structures of DWA1 and DWA2. (A) Protein structures of DWA1 and DWA2. WD40 region was determined based on the NCBI conserved domain databases (B, C) Schematic representation
Supplemental Figure 1. Protein and Gene Structures of DWA1 and DWA2. (A) Protein structures of DWA1 and DWA2. WD40 region was determined based on the NCBI conserved domain databases (B, C) Schematic representation
Fertilization Recovery after Defective. Sperm Cell Release in Arabidopsis. Current Biology, Volume 22 Supplemental Information
 Current Biology, Volume 22 Supplemental Information Fertilization Recovery after Defective Sperm Cell Release in Arabidopsis Ryushiro D. Kasahara, Daisuke Maruyama, Yuki Hamamura, Takashi Sakakibara, David
Current Biology, Volume 22 Supplemental Information Fertilization Recovery after Defective Sperm Cell Release in Arabidopsis Ryushiro D. Kasahara, Daisuke Maruyama, Yuki Hamamura, Takashi Sakakibara, David
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/5/244/ra72/dc1 Supplementary Materials for An Interaction Between BZR1 and DELLAs Mediates Direct Signaling Crosstalk Between Brassinosteroids and Gibberellins
www.sciencesignaling.org/cgi/content/full/5/244/ra72/dc1 Supplementary Materials for An Interaction Between BZR1 and DELLAs Mediates Direct Signaling Crosstalk Between Brassinosteroids and Gibberellins
Supplemental Data. Liu et al. (2013). Plant Cell /tpc
 Supplemental Figure 1. The GFP Tag Does Not Disturb the Physiological Functions of WDL3. (A) RT-PCR analysis of WDL3 expression in wild-type, WDL3-GFP, and WDL3 (without the GFP tag) transgenic seedlings.
Supplemental Figure 1. The GFP Tag Does Not Disturb the Physiological Functions of WDL3. (A) RT-PCR analysis of WDL3 expression in wild-type, WDL3-GFP, and WDL3 (without the GFP tag) transgenic seedlings.
Supplemental Data. Furlan et al. Plant Cell (2017) /tpc
 Supplemental Data. Furlan et al. Plant Cell (0) 0.0/tpc..00. Supplemental Data. Furlan et al. Plant Cell (0) 0.0/tpc..00. Supplemental Data. Furlan et al. Plant Cell (0) 0.0/tpc..00. Supplemental Figure.
Supplemental Data. Furlan et al. Plant Cell (0) 0.0/tpc..00. Supplemental Data. Furlan et al. Plant Cell (0) 0.0/tpc..00. Supplemental Data. Furlan et al. Plant Cell (0) 0.0/tpc..00. Supplemental Figure.
Supplementary Fig 1. The responses of ERF109 to different hormones and stresses. (a to k) The induced expression of ERF109 in 7-day-old Arabidopsis
 Supplementary Fig 1. The responses of ERF109 to different hormones and stresses. (a to k) The induced expression of ERF109 in 7-day-old Arabidopsis seedlings expressing ERF109pro-GUS. The GUS staining
Supplementary Fig 1. The responses of ERF109 to different hormones and stresses. (a to k) The induced expression of ERF109 in 7-day-old Arabidopsis seedlings expressing ERF109pro-GUS. The GUS staining
Supplementary information
 Supplementary information The E3 ligase RNF8 regulates KU80 removal and NHEJ repair Lin Feng 1, Junjie Chen 1 1 Department of Experimental Radiation Oncology, The University of Texas M. D. Anderson Cancer
Supplementary information The E3 ligase RNF8 regulates KU80 removal and NHEJ repair Lin Feng 1, Junjie Chen 1 1 Department of Experimental Radiation Oncology, The University of Texas M. D. Anderson Cancer
Supplemental Figure 1. Conserved regions of the kinase domain of PEPR1, PEPR2, CLV1 and BRI1.
 Supplemental Figure 1. Conserved regions of the kinase domain of PEPR1, PEPR2, CLV1 and BRI1. The asterisks and colons indicate the important residues for ATP-binding pocket and substrate binding pocket,
Supplemental Figure 1. Conserved regions of the kinase domain of PEPR1, PEPR2, CLV1 and BRI1. The asterisks and colons indicate the important residues for ATP-binding pocket and substrate binding pocket,
Supplemental Data. Dai et al. (2013). Plant Cell /tpc Absolute FyPP3. Absolute
 A FyPP1 Absolute B FyPP3 Absolute Dry seeds Imbibed 24 hours Dry seeds Imbibed 24 hours C ABI5 Absolute Dry seeds Imbibed 24 hours Supplemental Figure 1. Expression of FyPP1, FyPP3 and ABI5 during seed
A FyPP1 Absolute B FyPP3 Absolute Dry seeds Imbibed 24 hours Dry seeds Imbibed 24 hours C ABI5 Absolute Dry seeds Imbibed 24 hours Supplemental Figure 1. Expression of FyPP1, FyPP3 and ABI5 during seed
Supplemental Data. Sethi et al. (2014). Plant Cell /tpc
 Supplemental Data Supplemental Figure 1. MYC2 Binds to the E-box but not the E1-box of the MPK6 Promoter. (A) E1-box and E-box (wild type) containing MPK6 promoter fragment. The region shown in red denotes
Supplemental Data Supplemental Figure 1. MYC2 Binds to the E-box but not the E1-box of the MPK6 Promoter. (A) E1-box and E-box (wild type) containing MPK6 promoter fragment. The region shown in red denotes
S156AT168AY175A (AAA) were purified as GST-fusion proteins and incubated with GSTfused
 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 Supplemental Materials Supplemental Figure S1 (a) Phenotype of the wild type and grik1-2 grik2-1 plants after 8 days in darkness.
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 Supplemental Materials Supplemental Figure S1 (a) Phenotype of the wild type and grik1-2 grik2-1 plants after 8 days in darkness.
Supplementary Figures 1-12
 Supplementary Figures 1-12 Supplementary Figure 1. The specificity of anti-abi1 antibody. Total Proteins extracted from the wild type seedlings or abi1-3 null mutant seedlings were used for immunoblotting
Supplementary Figures 1-12 Supplementary Figure 1. The specificity of anti-abi1 antibody. Total Proteins extracted from the wild type seedlings or abi1-3 null mutant seedlings were used for immunoblotting
PCR Cloning II fusion domains for purification His 6 GST chitin binding protein MBP
 PCR cloning : why secondary amplification? May 26, 2009 secondary amplification amplification of the coding sequence alone or of defined fragments ligation into expression vector insertion of fusion domains
PCR cloning : why secondary amplification? May 26, 2009 secondary amplification amplification of the coding sequence alone or of defined fragments ligation into expression vector insertion of fusion domains
Supplementary Figure S1. Immunodetection of full-length XA21 and the XA21 C-terminal cleavage product.
 Supplementary Information Supplementary Figure S1. Immunodetection of full-length XA21 and the XA21 C-terminal cleavage product. Total protein extracted from Kitaake wild type and rice plants carrying
Supplementary Information Supplementary Figure S1. Immunodetection of full-length XA21 and the XA21 C-terminal cleavage product. Total protein extracted from Kitaake wild type and rice plants carrying
Supplemental Data. Cui et al. (2012). Plant Cell /tpc a b c d. Stem UBC32 ACTIN
 A Root Stem Leaf Flower Silique Senescence leaf B a b c d UBC32 ACTIN C * Supplemental Figure 1. Expression Pattern and Protein Sequence of UBC32 Homologues in Yeast, Human, and Arabidopsis. (A) Expression
A Root Stem Leaf Flower Silique Senescence leaf B a b c d UBC32 ACTIN C * Supplemental Figure 1. Expression Pattern and Protein Sequence of UBC32 Homologues in Yeast, Human, and Arabidopsis. (A) Expression
SUPPLEMENTARY INFORMATION
 R2A MASNSEKNPLL-SDEKPKSTEENKSS-KPESASGSSTSSAMP---GLNFNAFDFSNMASIL 56 R2B MASSSEKTPLIPSDEKNDTKEESKSTTKPESGSGAPPSPS-PTDPGLDFNAFDFSGMAGIL 60 R2A NDPSIREMAEQIAKDPAFNQLAEQLQRSIPNAGQEGGFPNFDPQQYVNTMQQVMHNPEFK
R2A MASNSEKNPLL-SDEKPKSTEENKSS-KPESASGSSTSSAMP---GLNFNAFDFSNMASIL 56 R2B MASSSEKTPLIPSDEKNDTKEESKSTTKPESGSGAPPSPS-PTDPGLDFNAFDFSGMAGIL 60 R2A NDPSIREMAEQIAKDPAFNQLAEQLQRSIPNAGQEGGFPNFDPQQYVNTMQQVMHNPEFK
Figure S1. DELLA Proteins Act as Positive Regulators to Mediate GA-Regulated Anthocyanin
 Supplemental Information Figure S1. DELLA Proteins Act as Positive Regulators to Mediate GA-Regulated Anthocyanin Biosynthesis. (A) Effect of GA on anthocyanin content in WT and ga1-3 seedlings. Mock,
Supplemental Information Figure S1. DELLA Proteins Act as Positive Regulators to Mediate GA-Regulated Anthocyanin Biosynthesis. (A) Effect of GA on anthocyanin content in WT and ga1-3 seedlings. Mock,
- 1 - Supplemental Data
 - 1-1 Supplemental Data 2 3 4 5 6 7 8 9 Supplemental Figure S1. Differential expression of AtPIP Genes in DC3000-inoculated plants. Gene expression in leaves was analyzed by real-time RT-PCR and expression
- 1-1 Supplemental Data 2 3 4 5 6 7 8 9 Supplemental Figure S1. Differential expression of AtPIP Genes in DC3000-inoculated plants. Gene expression in leaves was analyzed by real-time RT-PCR and expression
Data Sheet Quick PCR Cloning Kit
 Data Sheet Quick PCR Cloning Kit 6044 Cornerstone Ct. West, Ste. E DESCRIPTION: The Quick PCR Cloning Kit is a simple and highly efficient method to insert any gene or DNA fragment into a vector, without
Data Sheet Quick PCR Cloning Kit 6044 Cornerstone Ct. West, Ste. E DESCRIPTION: The Quick PCR Cloning Kit is a simple and highly efficient method to insert any gene or DNA fragment into a vector, without
PIE1 ARP6 SWC6 KU70 ARP6 PIE1. HSA SNF2_N HELICc SANT. pie1-3 A1,A2 K1,K2 K1,K3 K3,LB2 A3, A4 A3,LB1 A1,A2 K1,K2 K1,K3. swc6-1 A3,A4.
 A B N-terminal SWC2 H2A.Z SWC6 ARP6 PIE1 HSA SNF2_N HELICc SANT C pie1-3 D PIE1 ARP6 5 Kb A1 200 bp A3 A2 LB1 arp6-3 A4 E A1,A2 A3, A4 A3,LB1 K1,K2 K1,K3 K3,LB2 SWC6 swc6-1 A1,A2 A3,A4 K1,K2 K1,K3 100
A B N-terminal SWC2 H2A.Z SWC6 ARP6 PIE1 HSA SNF2_N HELICc SANT C pie1-3 D PIE1 ARP6 5 Kb A1 200 bp A3 A2 LB1 arp6-3 A4 E A1,A2 A3, A4 A3,LB1 K1,K2 K1,K3 K3,LB2 SWC6 swc6-1 A1,A2 A3,A4 K1,K2 K1,K3 100
(phosphatase tensin) domain is shown in dark gray, the FH1 domain in black, and the
 Supplemental Figure 1. Predicted Domain Organization of the AFH14 Protein. (A) Schematic representation of the predicted domain organization of AFH14. The PTEN (phosphatase tensin) domain is shown in dark
Supplemental Figure 1. Predicted Domain Organization of the AFH14 Protein. (A) Schematic representation of the predicted domain organization of AFH14. The PTEN (phosphatase tensin) domain is shown in dark
Supplemental Information
 Supplemental Information ATP-dependent unwinding of U4/U6 snrnas by the Brr2 helicase requires the C-terminus of Prp8 Corina Maeder 1,3, Alan K. Kutach 1,2,3, and Christine Guthrie 1 1 Department of Biochemistry
Supplemental Information ATP-dependent unwinding of U4/U6 snrnas by the Brr2 helicase requires the C-terminus of Prp8 Corina Maeder 1,3, Alan K. Kutach 1,2,3, and Christine Guthrie 1 1 Department of Biochemistry
pt7ht vector and over-expressed in E. coli as inclusion bodies. Cells were lysed in 6 M
 Supplementary Methods MIG6 production, purification, inhibition, and kinase assays MIG6 segment 1 (30mer, residues 334 364) peptide was synthesized using standard solid-phase peptide synthesis as described
Supplementary Methods MIG6 production, purification, inhibition, and kinase assays MIG6 segment 1 (30mer, residues 334 364) peptide was synthesized using standard solid-phase peptide synthesis as described
Supplemental data. Zhao et al. (2009). The Wuschel-related homeobox gene WOX11 is required to activate shoot-borne crown root development in rice.
 Supplemental data. Zhao et al. (2009). The Wuschel-related homeobox gene WOX11 is required to activate shoot-borne crown root development in rice. A B Supplemental Figure 1. Expression of WOX11p-GUS WOX11-GFP
Supplemental data. Zhao et al. (2009). The Wuschel-related homeobox gene WOX11 is required to activate shoot-borne crown root development in rice. A B Supplemental Figure 1. Expression of WOX11p-GUS WOX11-GFP
Supplemental Information
 FOOT-AND-MOUTH DISEASE VIRUS PROTEIN 2C IS A HEXAMERIC AAA+ PROTEIN WITH A COORDINATED ATP HYDROLYSIS MECHANISM. Trevor R. Sweeney 1, Valentina Cisnetto 1, Daniel Bose 2, Matthew Bailey 3, Jon R. Wilson
FOOT-AND-MOUTH DISEASE VIRUS PROTEIN 2C IS A HEXAMERIC AAA+ PROTEIN WITH A COORDINATED ATP HYDROLYSIS MECHANISM. Trevor R. Sweeney 1, Valentina Cisnetto 1, Daniel Bose 2, Matthew Bailey 3, Jon R. Wilson
Supplementary Information
 Supplementary Information ER-localized auxin transporter PIN8 regulates auxin homeostasis and male gametophyte development in Arabidopsis Supplemental Figures a 8 6 4 2 Absolute Expression b Absolute Expression
Supplementary Information ER-localized auxin transporter PIN8 regulates auxin homeostasis and male gametophyte development in Arabidopsis Supplemental Figures a 8 6 4 2 Absolute Expression b Absolute Expression
Experimental Tools and Resources Available in Arabidopsis. Manish Raizada, University of Guelph, Canada
 Experimental Tools and Resources Available in Arabidopsis Manish Raizada, University of Guelph, Canada Community website: The Arabidopsis Information Resource (TAIR) at http://www.arabidopsis.org Can order
Experimental Tools and Resources Available in Arabidopsis Manish Raizada, University of Guelph, Canada Community website: The Arabidopsis Information Resource (TAIR) at http://www.arabidopsis.org Can order
Figure legends for supplement
 Figure legends for supplement Supplemental Figure 1 Characterization of purified and recombinant proteins Relevant fractions related the final stage of the purification protocol(bingham et al., 1998; Toba
Figure legends for supplement Supplemental Figure 1 Characterization of purified and recombinant proteins Relevant fractions related the final stage of the purification protocol(bingham et al., 1998; Toba
Figure S1. Verification of ihog Mutation by Protein Immunoblotting Figure S2. Verification of ihog and boi
 Figure S1. Verification of ihog Mutation by Protein Immunoblotting Extracts from S2R+ cells, embryos, and adults were analyzed by immunoprecipitation and immunoblotting with anti-ihog antibody. The Ihog
Figure S1. Verification of ihog Mutation by Protein Immunoblotting Extracts from S2R+ cells, embryos, and adults were analyzed by immunoprecipitation and immunoblotting with anti-ihog antibody. The Ihog
Supplemental Materials
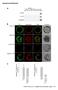 Supplemental Materials Flores-Pérez et al., Supplemental Materials, page 1 of 5 Supplemental Figure S1. Pull-down and BiFC controls, and quantitative analyses associated with the BiFC studies. (A) Controls
Supplemental Materials Flores-Pérez et al., Supplemental Materials, page 1 of 5 Supplemental Figure S1. Pull-down and BiFC controls, and quantitative analyses associated with the BiFC studies. (A) Controls
Supplemental Figure 1. Immunoblot analysis of wild-type and mutant forms of JAZ3
 Supplemental Data. Chung and Howe (2009). A Critical Role for the TIFY Motif in Repression of Jasmonate Signaling by a Stabilized Splice Variant of the JASMONATE ZIM-domain protein JAZ10 in Arabidopsis.
Supplemental Data. Chung and Howe (2009). A Critical Role for the TIFY Motif in Repression of Jasmonate Signaling by a Stabilized Splice Variant of the JASMONATE ZIM-domain protein JAZ10 in Arabidopsis.
Supplemental Data. Wu et al. (2). Plant Cell..5/tpc RGLG Hormonal treatment H2O B RGLG µm ABA µm ACC µm GA Time (hours) µm µm MJ µm IA
 Supplemental Data. Wu et al. (2). Plant Cell..5/tpc..4. A B Supplemental Figure. Immunoblot analysis verifies the expression of the AD-PP2C and BD-RGLG proteins in the Y2H assay. Total proteins were extracted
Supplemental Data. Wu et al. (2). Plant Cell..5/tpc..4. A B Supplemental Figure. Immunoblot analysis verifies the expression of the AD-PP2C and BD-RGLG proteins in the Y2H assay. Total proteins were extracted
Supplemental Data. Li et al. (2015). Plant Cell /tpc
 Supplemental Data Supplemental Figure 1: Characterization of asr3 T-DNA knockout lines and complementation transgenic lines. (A) The scheme of At2G33550 (ASR3) with gray boxes indicating exons and dash
Supplemental Data Supplemental Figure 1: Characterization of asr3 T-DNA knockout lines and complementation transgenic lines. (A) The scheme of At2G33550 (ASR3) with gray boxes indicating exons and dash
Supplemental Experimental Procedures
 Supplemental Experimental Procedures Cloning, expression and purification of untagged P C, Q D -N 0123, Q D- N 012 and Q D -N 01 - The DNA sequence encoding non tagged P C (starting at the Arginine 57
Supplemental Experimental Procedures Cloning, expression and purification of untagged P C, Q D -N 0123, Q D- N 012 and Q D -N 01 - The DNA sequence encoding non tagged P C (starting at the Arginine 57
pgbkt7 Anti- Myc AH109 strain (KDa) 50
 pgbkt7 (KDa) 50 37 Anti- Myc AH109 strain Supplementary Figure 1. Protein expression of CRN and TDR in yeast. To analyse the protein expression of CRNKD and TDRKD, total proteins extracted from yeast culture
pgbkt7 (KDa) 50 37 Anti- Myc AH109 strain Supplementary Figure 1. Protein expression of CRN and TDR in yeast. To analyse the protein expression of CRNKD and TDRKD, total proteins extracted from yeast culture
Supplemental Data. Na Xu et al. (2016). Plant Cell /tpc
 Supplemental Figure 1. The weak fluorescence phenotype is not caused by the mutation in At3g60240. (A) A mutation mapped to the gene At3g60240. Map-based cloning strategy was used to map the mutated site
Supplemental Figure 1. The weak fluorescence phenotype is not caused by the mutation in At3g60240. (A) A mutation mapped to the gene At3g60240. Map-based cloning strategy was used to map the mutated site
Supplemental Data. Steiner et al. Plant Cell. (2012) /tpc
 Supplemental Figure 1. SPY does not interact with free GST. Invitro pull-down assay using E. coli-expressed MBP-SPY and GST, GST-TCP14 and GST-TCP15. MBP-SPY was used as bait and incubated with equal amount
Supplemental Figure 1. SPY does not interact with free GST. Invitro pull-down assay using E. coli-expressed MBP-SPY and GST, GST-TCP14 and GST-TCP15. MBP-SPY was used as bait and incubated with equal amount
Supplementary information
 Supplementary information Supplementary figures Figure S1 Level of mycdet1 protein in DET1 OE-1, OE-2 and OE-3 transgenic lines. Total protein extract from wild type Col0, det1-1 mutant and DET1 OE lines
Supplementary information Supplementary figures Figure S1 Level of mycdet1 protein in DET1 OE-1, OE-2 and OE-3 transgenic lines. Total protein extract from wild type Col0, det1-1 mutant and DET1 OE lines
SUPPLEMENTAL DATA. Supplementary data
 1 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 34 35 SUPPLEMENTAL DATA Supplementary data Figure S1: Verification of pla-iα1 knockdown lines a: Semiquantitative
1 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 34 35 SUPPLEMENTAL DATA Supplementary data Figure S1: Verification of pla-iα1 knockdown lines a: Semiquantitative
Figure S1. Figure S2. Figure S3 HB Anti-FSP27 (COOH-terminal peptide) Ab. Anti-GST-FSP27(45-127) Ab.
 / 36B4 mrna ratio Figure S1 * 2. 1.6 1.2.8 *.4 control TNFα BRL49653 Figure S2 Su bw AT p iw Anti- (COOH-terminal peptide) Ab Blot : Anti-GST-(45-127) Ab β-actin Figure S3 HB2 HW AT BA T Figure S4 A TAG
/ 36B4 mrna ratio Figure S1 * 2. 1.6 1.2.8 *.4 control TNFα BRL49653 Figure S2 Su bw AT p iw Anti- (COOH-terminal peptide) Ab Blot : Anti-GST-(45-127) Ab β-actin Figure S3 HB2 HW AT BA T Figure S4 A TAG
Construction of plant complementation vector and generation of transgenic plants
 MATERIAL S AND METHODS Plant materials and growth conditions Arabidopsis ecotype Columbia (Col0) was used for this study. SALK_072009, SALK_076309, and SALK_027645 were obtained from the Arabidopsis Biological
MATERIAL S AND METHODS Plant materials and growth conditions Arabidopsis ecotype Columbia (Col0) was used for this study. SALK_072009, SALK_076309, and SALK_027645 were obtained from the Arabidopsis Biological
TECHNICAL BULLETIN. In Vitro Bacterial Split Fluorescent Protein Fold n Glow Solubility Assay Kits
 In Vitro Bacterial Split Fluorescent Protein Fold n Glow Solubility Assay Kits Catalog Numbers APPA001 In Vitro Bacterial Split GFP "Fold 'n' Glow" Solubility Assay Kit (Green) APPA008 In Vitro Bacterial
In Vitro Bacterial Split Fluorescent Protein Fold n Glow Solubility Assay Kits Catalog Numbers APPA001 In Vitro Bacterial Split GFP "Fold 'n' Glow" Solubility Assay Kit (Green) APPA008 In Vitro Bacterial
PRODUCT INFORMATION. Composition of SOC medium supplied :
 Product Name : Competent Cell BL21(DE3)pLysS Code No. : DS260 Size : 100 μl 10 Competency : > 5 10 7 cfu/μg (puc19) Supplied product : SOC medium, 1 ml 10 This product is for research use only Description
Product Name : Competent Cell BL21(DE3)pLysS Code No. : DS260 Size : 100 μl 10 Competency : > 5 10 7 cfu/μg (puc19) Supplied product : SOC medium, 1 ml 10 This product is for research use only Description
Supplemental Data. Shimoda et al. (2011). Plant Cell /tpc B C D E
 (kda) 80 58 46 30 25 B C D E (0/15) (29/29) (0/29) (0/32) Supplemental Figure 1. Immunodetection of Myc-tagged CCaMK Protein Variants on ccamk-3 Roots. () Immunodetection of myc-tag fusion CCaMK WT, 1-340
(kda) 80 58 46 30 25 B C D E (0/15) (29/29) (0/29) (0/32) Supplemental Figure 1. Immunodetection of Myc-tagged CCaMK Protein Variants on ccamk-3 Roots. () Immunodetection of myc-tag fusion CCaMK WT, 1-340
Generation of gene knockout vectors for Dictyostelium discoideum
 Generation of gene knockout vectors for Dictyostelium discoideum Instruction manual Last date of revision April 2012 2015 Version PR29-0001 PR29-0003 www.stargate.com This manual can be downloaded under
Generation of gene knockout vectors for Dictyostelium discoideum Instruction manual Last date of revision April 2012 2015 Version PR29-0001 PR29-0003 www.stargate.com This manual can be downloaded under
Supplemental Figure 1
 Supplemental Figure 1 A gta2-1 gta2-2 1kb AT4G08350 B Col-0 gta2-1 LP+RP+LBb1.3 C Col-0 gta2-2 LP+RP+LB1 D Col-0 gta2-2 gta2-1 GTA2 TUBLIN E Col0 gta2-1 gta2-2 Supplemental Figure 1. Phenotypic analysis
Supplemental Figure 1 A gta2-1 gta2-2 1kb AT4G08350 B Col-0 gta2-1 LP+RP+LBb1.3 C Col-0 gta2-2 LP+RP+LB1 D Col-0 gta2-2 gta2-1 GTA2 TUBLIN E Col0 gta2-1 gta2-2 Supplemental Figure 1. Phenotypic analysis
Supplementary data. sienigma. F-Enigma F-EnigmaSM. a-p53
 Supplementary data Supplemental Figure 1 A sienigma #2 sienigma sicontrol a-enigma - + ++ - - - - - - + ++ - - - - - - ++ B sienigma F-Enigma F-EnigmaSM a-flag HLK3 cells - - - + ++ + ++ - + - + + - -
Supplementary data Supplemental Figure 1 A sienigma #2 sienigma sicontrol a-enigma - + ++ - - - - - - + ++ - - - - - - ++ B sienigma F-Enigma F-EnigmaSM a-flag HLK3 cells - - - + ++ + ++ - + - + + - -
Supplemental Figure 1 HDA18 has an HDAC domain and therefore has concentration dependent and TSA inhibited histone deacetylase activity.
 Supplemental Figure 1 HDA18 has an HDAC domain and therefore has concentration dependent and TSA inhibited histone deacetylase activity. (A) Amino acid alignment of HDA5, HDA15 and HDA18. The blue line
Supplemental Figure 1 HDA18 has an HDAC domain and therefore has concentration dependent and TSA inhibited histone deacetylase activity. (A) Amino acid alignment of HDA5, HDA15 and HDA18. The blue line
Supplemental Figure 1. Alignment of the NbGAPC amino acid sequences with their Arabidopsis homologues.
 Supplemental Figure 1. Alignment of the NbGAPC amino acid sequences with their Arabidopsis homologues. Homologs from N. benthamiana (NbGAPC1, NbGAPC2, NbGAPC3), Arabidopsis (AtGAPC1, AT3G04120; AtGAPC2,
Supplemental Figure 1. Alignment of the NbGAPC amino acid sequences with their Arabidopsis homologues. Homologs from N. benthamiana (NbGAPC1, NbGAPC2, NbGAPC3), Arabidopsis (AtGAPC1, AT3G04120; AtGAPC2,
Supporting Information
 Supporting Information Üstün and Börnke, 2015 Figure S1: XopJ does not display acetyltransferase activity. Autoacetylation activity in vitro. Acetylation reactions using MBP, MBP-XopJ, GST and GST-HopZ1a
Supporting Information Üstün and Börnke, 2015 Figure S1: XopJ does not display acetyltransferase activity. Autoacetylation activity in vitro. Acetylation reactions using MBP, MBP-XopJ, GST and GST-HopZ1a
SUPPLEMENTARY INFORMATION
 Local auxin metabolism regulates environment-induced hypocotyl elongation Zuyu Zheng 1,2, Yongxia Guo 3, Ondřej Novák 4,5, William Chen 2, Karin Ljung 4, Joseph P. Noel 1,3, *, and Joanne Chory 1,2, *
Local auxin metabolism regulates environment-induced hypocotyl elongation Zuyu Zheng 1,2, Yongxia Guo 3, Ondřej Novák 4,5, William Chen 2, Karin Ljung 4, Joseph P. Noel 1,3, *, and Joanne Chory 1,2, *
indicated numbers of pups at day of life (DOL) 10, or embryonic day (ED) B. Male mice of
 SUPPLEMENTRY FIGURE LEGENDS Figure S1. USP44 loss leads to chromosome missegregation.. Genotypes obtained from the indicated numbers of pups at day of life (DOL) 10, or embryonic day (ED) 13.5.. Male mice
SUPPLEMENTRY FIGURE LEGENDS Figure S1. USP44 loss leads to chromosome missegregation.. Genotypes obtained from the indicated numbers of pups at day of life (DOL) 10, or embryonic day (ED) 13.5.. Male mice
Supplemental Data. Aung et al. (2011). Plant Cell /tpc
 35S pro:pmd1-yfp 10 µm Supplemental Figure 1. -terminal YFP fusion of PMD1 (PMD1-YFP, in green) is localized to the cytosol. grobacterium cells harboring 35S pro :PMD1-YFP were infiltrated into tobacco
35S pro:pmd1-yfp 10 µm Supplemental Figure 1. -terminal YFP fusion of PMD1 (PMD1-YFP, in green) is localized to the cytosol. grobacterium cells harboring 35S pro :PMD1-YFP were infiltrated into tobacco
Supplemental Data. Seo et al. (2014). Plant Cell /tpc
 Supplemental Figure 1. Protein alignment of ABD1 from other model organisms. The alignment was performed with H. sapiens DCAF8, M. musculus DCAF8 and O. sativa Os10g0544500. The WD40 domains are underlined.
Supplemental Figure 1. Protein alignment of ABD1 from other model organisms. The alignment was performed with H. sapiens DCAF8, M. musculus DCAF8 and O. sativa Os10g0544500. The WD40 domains are underlined.
Supplemental Data. Guo et al. (2015). Plant Cell /tpc
 Supplemental Figure 1. The Mutant exb1-d Displayed Pleiotropic Phenotypes and Produced Branches in the Axils of Cotyledons. (A) Branches were developed in exb1-d but not in wild-type plants. (B) and (C)
Supplemental Figure 1. The Mutant exb1-d Displayed Pleiotropic Phenotypes and Produced Branches in the Axils of Cotyledons. (A) Branches were developed in exb1-d but not in wild-type plants. (B) and (C)
Expanded View Figures
 Expanded View Figures Figure EV1. AM3-CLE45 control experiments and bam3 alleles. A Relative primary root length of indicated genotypes at 9 dag, in response to increasing amounts of CLE45 in the media.
Expanded View Figures Figure EV1. AM3-CLE45 control experiments and bam3 alleles. A Relative primary root length of indicated genotypes at 9 dag, in response to increasing amounts of CLE45 in the media.
pri 201 DNA Series (High-Expression Vectors for Plant Transformation)
 For Research Use Cat #. 3264 3265 3266 3267 pri 201 DNA Series (High-Expression Vectors for Plant Transformation) Product Manual Table of Contents I. Description... 3 II. Product Information... 3 III.
For Research Use Cat #. 3264 3265 3266 3267 pri 201 DNA Series (High-Expression Vectors for Plant Transformation) Product Manual Table of Contents I. Description... 3 II. Product Information... 3 III.
SUPPLEMENTAL INFORMATION
 SUPPLEMENTAL INFORMATION SUPPLEMENTAL DATA Table S1: Related to Figure 4. DnaA screen for divalent cations and nucleotides. Concentration of reagents, T m and F fold30 75. Reagent Conc T m (±SEM) mm C
SUPPLEMENTAL INFORMATION SUPPLEMENTAL DATA Table S1: Related to Figure 4. DnaA screen for divalent cations and nucleotides. Concentration of reagents, T m and F fold30 75. Reagent Conc T m (±SEM) mm C
Supplemental Data. Farmer et al. (2010) Plant Cell /tpc
 Supplemental Figure 1. Amino acid sequence comparison of RAD23 proteins. Identical and similar residues are shown in the black and gray boxes, respectively. Dots denote gaps. The sequence of plant Ub is
Supplemental Figure 1. Amino acid sequence comparison of RAD23 proteins. Identical and similar residues are shown in the black and gray boxes, respectively. Dots denote gaps. The sequence of plant Ub is
GTTCGGGTTCC TTTTGAGCAG
 Supplementary Figures Splice variants of the SIP1 transcripts play a role in nodule organogenesis in Lotus japonicus. Wang C, Zhu H, Jin L, Chen T, Wang L, Kang H, Hong Z, Zhang Z. 5 UTR CDS 3 UTR TCTCAACCATCCTTTGTCTGCTTCCGCCGCATGGGTGAGGTCATTTTGTCTAGATGACGTGCAATTTACAATGA
Supplementary Figures Splice variants of the SIP1 transcripts play a role in nodule organogenesis in Lotus japonicus. Wang C, Zhu H, Jin L, Chen T, Wang L, Kang H, Hong Z, Zhang Z. 5 UTR CDS 3 UTR TCTCAACCATCCTTTGTCTGCTTCCGCCGCATGGGTGAGGTCATTTTGTCTAGATGACGTGCAATTTACAATGA
Supplementary Table 1. The Q-PCR primer sequence is summarized in the following table.
 Supplementary Table 1. The Q-PCR primer sequence is summarized in the following table. Name Sequence (5-3 ) Application Flag-u ggactacaaggacgacgatgac Shared upstream primer for all the amplifications of
Supplementary Table 1. The Q-PCR primer sequence is summarized in the following table. Name Sequence (5-3 ) Application Flag-u ggactacaaggacgacgatgac Shared upstream primer for all the amplifications of
Gateway Vectors for BiFC
 Gateway Vectors for BiFC 1. The enhanced YFP (EYFP) are used (Split EYFP). 2. The Fusion fusion gene is expressed by CaMV35S promoter. 3. The N- or C-terminal fragments of EYFP are fused subsequent to
Gateway Vectors for BiFC 1. The enhanced YFP (EYFP) are used (Split EYFP). 2. The Fusion fusion gene is expressed by CaMV35S promoter. 3. The N- or C-terminal fragments of EYFP are fused subsequent to
Supplemental Figure 1. Phos-Tag mobility shift detection of in vivo phosphorylated ERF6 in 35S:ERF6 WT plants after B. cinerea inoculation.
 Supplemental Figure 1. Phos-Tag mobility shift detection of in vivo phosphorylated ERF6 in 35S:ERF6 WT plants after B. cinerea inoculation. Protein extracts from 35S:ERF6 WT seedlings treated with B. cinerea
Supplemental Figure 1. Phos-Tag mobility shift detection of in vivo phosphorylated ERF6 in 35S:ERF6 WT plants after B. cinerea inoculation. Protein extracts from 35S:ERF6 WT seedlings treated with B. cinerea
BL21(DE3) expression competent cell pack DS265. Component Code No. Contents Competent cell BL21(DE3) DS250 5 tubes (100 μl/tube)
 Product Name : Code No. : Kit Component : BL21(DE3) expression competent cell pack DS265 Component Code No. Contents Competent cell BL21(DE3) DS250 5 tubes (100 μl/tube) transfromation efficiency: 5 10
Product Name : Code No. : Kit Component : BL21(DE3) expression competent cell pack DS265 Component Code No. Contents Competent cell BL21(DE3) DS250 5 tubes (100 μl/tube) transfromation efficiency: 5 10
Supplemental Figure 1. VLN5 retains conserved residues at both type 1 and type 2 Ca 2+ -binding
 Supplemental Figure 1. VLN5 retains conserved residues at both type 1 and type 2 Ca 2+ -binding sites in the G1 domain. Multiple sequence alignment was performed with DNAMAN6.0.40. Secondary structural
Supplemental Figure 1. VLN5 retains conserved residues at both type 1 and type 2 Ca 2+ -binding sites in the G1 domain. Multiple sequence alignment was performed with DNAMAN6.0.40. Secondary structural
oligonucleotide primers listed in Supplementary Table 2 and cloned into ppcr-script (Stratagene) before digestion
 Supplementary Methods. Construction of BiFC vectors The full-length ARC3 cdna, ARC3 1-1794 and ARC3 1-1083 were amplified from ppcr-script/arc3 using the oligonucleotide primers listed in Supplementary
Supplementary Methods. Construction of BiFC vectors The full-length ARC3 cdna, ARC3 1-1794 and ARC3 1-1083 were amplified from ppcr-script/arc3 using the oligonucleotide primers listed in Supplementary
Supplementary Figure S1. Growth patterns of WT and siz1-2 plants and the effect of different nitrogen sources on their growth. After germination on
 Supplementary Figure S1. Growth patterns of WT and siz1-2 plants and the effect of different nitrogen sources on their growth. After germination on MS media, seedlings were transferred to soil and treated
Supplementary Figure S1. Growth patterns of WT and siz1-2 plants and the effect of different nitrogen sources on their growth. After germination on MS media, seedlings were transferred to soil and treated
Supplementary Information. c d e
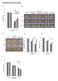 Supplementary Information a b c d e f Supplementary Figure 1. atabcg30, atabcg31, and atabcg40 mutant seeds germinate faster than the wild type on ½ MS medium supplemented with ABA (a and d-f) Germination
Supplementary Information a b c d e f Supplementary Figure 1. atabcg30, atabcg31, and atabcg40 mutant seeds germinate faster than the wild type on ½ MS medium supplemented with ABA (a and d-f) Germination
Rotation Report Sample Version 3. Due Date: August 9, Analysis of the Guanine Nucleotide Exchange Activity of the S. cerevisiae Ats1 Protein
 Rotation Report Sample Version 3 Due Date: August 9, 1998 Analysis of the Guanine Nucleotide Exchange Activity of the S. cerevisiae Ats1 Protein Anita H. Corbett Advisor: Amy Jones Rotation 1 Abstract:
Rotation Report Sample Version 3 Due Date: August 9, 1998 Analysis of the Guanine Nucleotide Exchange Activity of the S. cerevisiae Ats1 Protein Anita H. Corbett Advisor: Amy Jones Rotation 1 Abstract:
Enhancers mutations that make the original mutant phenotype more extreme. Suppressors mutations that make the original mutant phenotype less extreme
 Interactomics and Proteomics 1. Interactomics The field of interactomics is concerned with interactions between genes or proteins. They can be genetic interactions, in which two genes are involved in the
Interactomics and Proteomics 1. Interactomics The field of interactomics is concerned with interactions between genes or proteins. They can be genetic interactions, in which two genes are involved in the
Supporting information
 Supporting information Construction of strains and plasmids To create ptc67, a PCR product obtained with primers cc2570-162f (gcatgggcaagcttgaggacggcgtcatgt) and cc2570+512f (gaggccgtggtaccatagaggcgggcg),
Supporting information Construction of strains and plasmids To create ptc67, a PCR product obtained with primers cc2570-162f (gcatgggcaagcttgaggacggcgtcatgt) and cc2570+512f (gaggccgtggtaccatagaggcgggcg),
Supporting Information
 Electronic Supplementary Material (ESI) for ChemComm. This journal is The Royal Society of Chemistry 2016 Supporting Information An Ion Signal Responsive Dynamic Protein Nano-spring Constructed by High
Electronic Supplementary Material (ESI) for ChemComm. This journal is The Royal Society of Chemistry 2016 Supporting Information An Ion Signal Responsive Dynamic Protein Nano-spring Constructed by High
Supplementary Materials
 Supplementary Materials Supplementary Figure 1. PKM2 interacts with MLC2 in cytokinesis. a, U87, U87/EGFRvIII, and HeLa cells in cytokinesis were immunostained with DAPI and an anti-pkm2 antibody. Thirty
Supplementary Materials Supplementary Figure 1. PKM2 interacts with MLC2 in cytokinesis. a, U87, U87/EGFRvIII, and HeLa cells in cytokinesis were immunostained with DAPI and an anti-pkm2 antibody. Thirty
ABI3 Controls Embryo De-greening Through Mendel's I locus
 Supporting Online Material for ABI3 Controls Embryo De-greening Through Mendel's I locus Frédéric Delmas a,b,c, Subramanian Sankaranarayanan d, Srijani Deb d, Ellen Widdup d, Céline Bournonville b,c,norbert
Supporting Online Material for ABI3 Controls Embryo De-greening Through Mendel's I locus Frédéric Delmas a,b,c, Subramanian Sankaranarayanan d, Srijani Deb d, Ellen Widdup d, Céline Bournonville b,c,norbert
T H E J O U R N A L O F C E L L B I O L O G Y
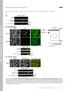 T H E J O U R N A L O F C E L L B I O L O G Y Supplemental material Nakajima and Tanoue, http://www.jcb.org/cgi/content/full/jcb.201104118/dc1 Figure S1. DLD-1 cells exhibit the characteristic morphology
T H E J O U R N A L O F C E L L B I O L O G Y Supplemental material Nakajima and Tanoue, http://www.jcb.org/cgi/content/full/jcb.201104118/dc1 Figure S1. DLD-1 cells exhibit the characteristic morphology
Li Wang, Fabio C. Rinaldi, Pallavi Singh, Erin L. Doyle, Zoe E. Dubrow, Tuan Tu Tran, Alvaro L. Pérez-Quintero, Boris Szurek, and Adam J.
 TAL effectors drive transcription bidirectionally in plants Li Wang, Fabio C. Rinaldi, Pallavi Singh, Erin L. Doyle, Zoe E. Dubrow, Tuan Tu Tran, Alvaro L. Pérez-Quintero, Boris Szurek, and Adam J. Bogdanove
TAL effectors drive transcription bidirectionally in plants Li Wang, Fabio C. Rinaldi, Pallavi Singh, Erin L. Doyle, Zoe E. Dubrow, Tuan Tu Tran, Alvaro L. Pérez-Quintero, Boris Szurek, and Adam J. Bogdanove
Supplemental Information. Pacer Mediates the Function of Class III PI3K. and HOPS Complexes in Autophagosome. Maturation by Engaging Stx17
 Molecular Cell, Volume 65 Supplemental Information Pacer Mediates the Function of Class III PI3K and HOPS Complexes in Autophagosome Maturation by Engaging Stx17 Xiawei Cheng, Xiuling Ma, Xianming Ding,
Molecular Cell, Volume 65 Supplemental Information Pacer Mediates the Function of Class III PI3K and HOPS Complexes in Autophagosome Maturation by Engaging Stx17 Xiawei Cheng, Xiuling Ma, Xianming Ding,
SUPPLEMENTARY INFORMATION
 (Supplementary Methods and Materials) GST pull-down assay GST-fusion proteins Fe65 365-533, and Fe65 538-700 were expressed in BL21 bacterial cells and purified with glutathione-agarose beads (Sigma).
(Supplementary Methods and Materials) GST pull-down assay GST-fusion proteins Fe65 365-533, and Fe65 538-700 were expressed in BL21 bacterial cells and purified with glutathione-agarose beads (Sigma).
Supplemental Data. Osakabe et al. (2013). Plant Cell /tpc
 Supplemental Figure 1. Phylogenetic analysis of KUP in various species. The amino acid sequences of KUPs from green algae and land plants were identified with a BLAST search and aligned using ClustalW
Supplemental Figure 1. Phylogenetic analysis of KUP in various species. The amino acid sequences of KUPs from green algae and land plants were identified with a BLAST search and aligned using ClustalW
Supplementary Figure 1. α-synuclein is truncated in PD and LBD brains. Nature Structural & Molecular Biology: doi: /nsmb.
 Supplementary Figure 1 α-synuclein is truncated in PD and LBD brains. (a) Specificity of anti-n103 antibody. Anti-N103 antibody was coated on an ELISA plate and different concentrations of full-length
Supplementary Figure 1 α-synuclein is truncated in PD and LBD brains. (a) Specificity of anti-n103 antibody. Anti-N103 antibody was coated on an ELISA plate and different concentrations of full-length
2.5. Equipment and materials supplied by user PCR based template preparation Influence of temperature on in vitro EGFP synthesis 11
 Manual 15 Reactions LEXSY in vitro Translation Cell-free protein expression kit based on Leishmania tarentolae for PCR-based template generation Cat. No. EGE-2010-15 FOR RESEARCH USE ONLY. NOT INTENDED
Manual 15 Reactions LEXSY in vitro Translation Cell-free protein expression kit based on Leishmania tarentolae for PCR-based template generation Cat. No. EGE-2010-15 FOR RESEARCH USE ONLY. NOT INTENDED
Supporting Online Material for
 www.sciencemag.org/cgi/content/full/319/5867/1241/dc1 Supporting Online Material for Local Positive Feedback Regulation Determines Cell Shape in Root Hair Cells Seiji Takeda, Catherine Gapper, Hidetaka
www.sciencemag.org/cgi/content/full/319/5867/1241/dc1 Supporting Online Material for Local Positive Feedback Regulation Determines Cell Shape in Root Hair Cells Seiji Takeda, Catherine Gapper, Hidetaka
Supplemental Data. Benstein et al. (2013). Plant Cell /tpc
 Supplemental Figure 1. Purification of the heterologously expressed PGDH1, PGDH2 and PGDH3 enzymes by Ni-NTA affinity chromatography. Protein extracts (2 µl) of different fractions (lane 1 = total extract,
Supplemental Figure 1. Purification of the heterologously expressed PGDH1, PGDH2 and PGDH3 enzymes by Ni-NTA affinity chromatography. Protein extracts (2 µl) of different fractions (lane 1 = total extract,
Supplementary Figure 1. BES1 specifically inhibits ABA responses in early seedling
 Supplementary Figure 1. BES1 specifically inhibits ABA responses in early seedling development. a. Exogenous BR application overcomes the hypersensitivity of bzr1-1d seedlings to ABA. Seed germination
Supplementary Figure 1. BES1 specifically inhibits ABA responses in early seedling development. a. Exogenous BR application overcomes the hypersensitivity of bzr1-1d seedlings to ABA. Seed germination
Nature Structural & Molecular Biology: doi: /nsmb.3308
 Supplementary Figure 1 Analysis of CED-3 autoactivation and CED-4-induced CED-3 activation. (a) Repeat experiments of Fig. 1a. (b) Repeat experiments of Fig. 1b. (c) Quantitative analysis of three independent
Supplementary Figure 1 Analysis of CED-3 autoactivation and CED-4-induced CED-3 activation. (a) Repeat experiments of Fig. 1a. (b) Repeat experiments of Fig. 1b. (c) Quantitative analysis of three independent
To investigate the heredity of the WFP gene, we selected plants that were homozygous
 Supplementary information Supplementary Note ST-12 WFP allele is semi-dominant To investigate the heredity of the WFP gene, we selected plants that were homozygous for chromosome 1 of Nipponbare and heterozygous
Supplementary information Supplementary Note ST-12 WFP allele is semi-dominant To investigate the heredity of the WFP gene, we selected plants that were homozygous for chromosome 1 of Nipponbare and heterozygous
(b) Genotyping of SALK_ (AT1g51690) and SALK_ (At1g17720) to find homozygous single mutant plants.
 Supplemental Table 1. Primers used for cloning and RT-PCR. The B-type subunits cdna CDS were amplified using high-fidelity PCR amplification with gene specific primers. The cdna CDS sequences were obtained
Supplemental Table 1. Primers used for cloning and RT-PCR. The B-type subunits cdna CDS were amplified using high-fidelity PCR amplification with gene specific primers. The cdna CDS sequences were obtained
Personal Exam Code: 1. 16p
 Exam, Methods in Molecular Biology, BIOR47, 2012-04-26 Please hand in answers to each question on its paper. Use additional papers if needed. Mark all papers with your code. Your answers should be logical
Exam, Methods in Molecular Biology, BIOR47, 2012-04-26 Please hand in answers to each question on its paper. Use additional papers if needed. Mark all papers with your code. Your answers should be logical
Supporting Information
 Supporting Information Deng et al. 10.1073/pnas.1102117108 Fig. S1. Predicted structure of Arabidopsis bzip60 RNA. Lowest free energy form (ΔG = 309.72 (initially 343.10) of bzip60 mrna folded by M-Fold
Supporting Information Deng et al. 10.1073/pnas.1102117108 Fig. S1. Predicted structure of Arabidopsis bzip60 RNA. Lowest free energy form (ΔG = 309.72 (initially 343.10) of bzip60 mrna folded by M-Fold
Product. Ni-NTA His Bind Resin. Ni-NTA His Bind Superflow. His Bind Resin. His Bind Magnetic Agarose Beads. His Bind Column. His Bind Quick Resin
 Novagen offers a large variety of affinity supports and kits for the purification of recombinant proteins containing popular peptide fusion tags, including His Tag, GST Tag, S Tag and T7 Tag sequences.
Novagen offers a large variety of affinity supports and kits for the purification of recombinant proteins containing popular peptide fusion tags, including His Tag, GST Tag, S Tag and T7 Tag sequences.
A subclass of HSP70s regulate development and abiotic stress responses in Arabidopsis thaliana
 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 Journal of Plant Research A subclass of HSP70s regulate development and abiotic stress responses in Arabidopsis thaliana Linna Leng 1 Qianqian Liang
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 Journal of Plant Research A subclass of HSP70s regulate development and abiotic stress responses in Arabidopsis thaliana Linna Leng 1 Qianqian Liang
Supplemental Information
 Supplemental Information Supplemental Figure 1. The Heterologous Yeast System for Screening of Arabidopsis JA Transporters. (A) Exogenous JA inhibited yeast cell growth. Yeast cells were diluted to various
Supplemental Information Supplemental Figure 1. The Heterologous Yeast System for Screening of Arabidopsis JA Transporters. (A) Exogenous JA inhibited yeast cell growth. Yeast cells were diluted to various
The poly(a) tail blocks RDR6 from converting self mrnas into substrates for gene silencing
 In the format provided by the authors and unedited. SUPPLEMENTARY INFORMATION VOLUME: 3 ARTICLE NUMBER: 17036 The poly(a) tail blocks RDR6 from converting self mrnas into substrates for gene silencing
In the format provided by the authors and unedited. SUPPLEMENTARY INFORMATION VOLUME: 3 ARTICLE NUMBER: 17036 The poly(a) tail blocks RDR6 from converting self mrnas into substrates for gene silencing
Supplementary Fig. 1. Schematic structure of TRAIP and RAP80. The prey line below TRAIP indicates bait and the two lines above RAP80 highlight the
 Supplementary Fig. 1. Schematic structure of TRAIP and RAP80. The prey line below TRAIP indicates bait and the two lines above RAP80 highlight the prey clones identified in the yeast two hybrid screen.
Supplementary Fig. 1. Schematic structure of TRAIP and RAP80. The prey line below TRAIP indicates bait and the two lines above RAP80 highlight the prey clones identified in the yeast two hybrid screen.
PHT1;2-CFP YFP-PHF + PHT1;2-CFP YFP-PHF
 YFP-PHF1 CFP-PHT1;2 PHT1;2-CFP YFP-PHF + PHT1;2-CFP YFP-PHF + CFP-PHT1;2 Negative control!-gfp Supplemental Figure 1: PHT1;2 accumulation is PHF1 dependent. Immunoblot analysis on total protein extract
YFP-PHF1 CFP-PHT1;2 PHT1;2-CFP YFP-PHF + PHT1;2-CFP YFP-PHF + CFP-PHT1;2 Negative control!-gfp Supplemental Figure 1: PHT1;2 accumulation is PHF1 dependent. Immunoblot analysis on total protein extract
Cat # Box1 Box2. DH5a Competent E. coli cells CCK-20 (20 rxns) 40 µl 40 µl 50 µl x 20 tubes. Choo-Choo Cloning TM Enzyme Mix
 Molecular Cloning Laboratories User Manual Version 3.3 Product name: Choo-Choo Cloning Kits Cat #: CCK-10, CCK-20, CCK-096, CCK-384 Description: Choo-Choo Cloning is a highly efficient directional PCR
Molecular Cloning Laboratories User Manual Version 3.3 Product name: Choo-Choo Cloning Kits Cat #: CCK-10, CCK-20, CCK-096, CCK-384 Description: Choo-Choo Cloning is a highly efficient directional PCR
Supplemental Experimental Procedures
 Supplemental Experimental Procedures Generation of BLM protein segments and mutants. For immunoprecipitation experiments with topoisomerase IIα, BLM N-terminal segments were generated by PCR amplification
Supplemental Experimental Procedures Generation of BLM protein segments and mutants. For immunoprecipitation experiments with topoisomerase IIα, BLM N-terminal segments were generated by PCR amplification
JCB. Supplemental material THE JOURNAL OF CELL BIOLOGY. Hong et al.,
 Supplemental material JCB Hong et al., http://www.jcb.org/cgi/content/full/jcb.201412127/dc1 THE JOURNAL OF CELL BIOLOGY Figure S1. Analysis of purified proteins by SDS-PAGE and pull-down assays. (A) Coomassie-stained
Supplemental material JCB Hong et al., http://www.jcb.org/cgi/content/full/jcb.201412127/dc1 THE JOURNAL OF CELL BIOLOGY Figure S1. Analysis of purified proteins by SDS-PAGE and pull-down assays. (A) Coomassie-stained
Supplemental materials
 Supplemental materials Materials and methods for supplemental figures Yeast two-hybrid assays TAP46-PP2Ac interactions I. The TAP46 was used as the bait and the full-length cdnas of the five C subunits
Supplemental materials Materials and methods for supplemental figures Yeast two-hybrid assays TAP46-PP2Ac interactions I. The TAP46 was used as the bait and the full-length cdnas of the five C subunits
