0.5% Sucrose. Supplemental Data. Chao et al. (2011). Plant Cell /tpc Col-0 tsc10a-2. Root length (mm)
|
|
|
- Chad Pierce
- 5 years ago
- Views:
Transcription
1 A 0% 0.5% 1% Col-0 tsc10a-2 B Root length (mm) % 0.50% 1% 2% 4% Sucrose concentration 2% 4% Sucrose Col-0 tsc10a-2 Supplemental Figure 1. The effect of sucrose on root growth of tsc10a.col-0 and tsc10a-2 plants were germinated on ½ MS media with sucrose at the indicated concentration and plants photographed (A) and root length quantified after 1-week (B). Error bars represent standard deviations.
2 A Col-0 tsc10a-2 Col-0 tsc10a-1 Percentage of tricotyledon plants 60% 40% 20% 0% B Supplemental Figure 2. Developmental changes in tsc10a. (A) A significant proportion of the plants of the TSC10A loss of function mutants show abnormal cotyledon development. (B) Percentage of Col-0 wild-type tsc10a-1 and tsc10a-2 plants with tricotyledons. Error bars represent standard deviations.
3 A Col-0 WT tsc10a-1 B Col-0 WT tsc10a-1 tsc10a-2 tsc10b-1 Supplemental Figure 3. Flowering and hypocotyl elongation phenotypes in tsc10a. (A) Abnormal flowering pattern of tsc10a-1. (B) Hypocotyl elongation of TSC10A loss of function mutants in response to etiolation. Wild type and mutants were placed in darkness, and grown vertically for 5 days. Error bars represent standard deviations.
4 Frequency (number of plants) tsc10a-1 F2 tsc10a-1 x Ler-0 Col-0 Ler Leaf Mg + Ca (percentage of Col-0) Supplemental Figure 4. Segregation of the leaf ionomic phenotype in five week old F2 plants from a Ler-0 x tsc10a-1 outcross. Data represents the percentage difference from the mean of the wild-type (Col-0) control plants (n = 48). Raw data can be viewed and obtained from in trays 890, 891, 892 and 893.
5 SPHD
6 Supplemental Figure 5. Map-based cloning of tsc10a-1. (A) Bulk Segregant analysis of tsc10a- 1 using F2 plants with low Mg and Ca from a Ler-0 x tsc10a-1 outcross. Data are presented as a scaled pool hybridization difference (SPHD), representing the difference between the hybridization of the two pools at the SFPs, scaled so that a pure Col-0 pool would be at 1 and a pure Ler-0 pool would be at -1. Pools were prepared from F2 plants with low leaf Ca and Mg content (n = 31) and F2 plants with a Col-0-like Ca and Mg content (n = 31). SFPs were scored after hybridization of genomic DNA from these pools to Affymetrix ATH1 DNA microarrays. Green lines denote likely location of the causal loci. (B) The mutated gene was mapped between SSLP makers T1540K and T1.9M using 341 F2 plants. (C) Fine mapping narrowed tsc10a-1 down to a 30 kb region between markers T1818K and T1848K using 1279 F2 plants. Numbers above the horizontal line in (B) and (C) are the number of recombinants between the indicated marker and tsc10a-1. (D) Sequenced region of tsc10a-1mutant and different tsc10a alleles. The white box represents the sequenced region between markers T1818K and T1848K. Black triangles indicate T-DNA insertion sites for two tsc10a alleles. Grey arrows represent genes in the sequenced region. Dashed line shows the mutated site of tsc10a-1. Start, start codon; Stop, stop codon.
7 A TSC10B TSC10A FVT1 Tsc10p TSC10B TSC10A FVT1 Tsc10p TSC10B TSC10A FVT1 Tsc10p TSC10B TSC10A FVT1 Tsc10p TSC10B TSC10A FVT1 Tsc10p B GABI_524F10 (tsc10b-2) SAIL_659_C12 (tsc10b-1) Supplemental Figure 6. Relationship between A. thaliana TSC10A and its various homologs. (A) Alignment of the predicted amino acid seqeunce of TSC10A and its homologs in A. thaliana (TSC10B), yeast (Tsc10p) and humans (FVT1). The N-terminal extensions of the two A. thaliana and the human 3-KDS reductases (blue), the Rossmann folds (red), conserved catalytic triad residues (orange), the C-terminal hydrophobic domains (purple) and the dilysine motif presumed to be important for ER retention in Tsc10p (pink) are indicated. (B) Structure of TSC10B and its loss of function alleles tsc10b-1 and tsc10b-2. The position of the T-DNA insertions are indicated by triangles. Exons are indicated by boxes (pink, 5' UTR; gray, coding region; blue, 3'UTR ), introns are shown as black lines and dashed lines represent promoter region. Note that TSC10B genomic sequence in tsc10b-2 was rearrangement by the T-DNA insertion.
8 Supplemental Data. Chao et al. (2011). Plant Cell /tpc A C B D Col-0 tsc10a-2 Col-0 tsc10a-1 Supplemental Figure 7. Wilting resistance of tsc10a mutants. Col-0 and tsc10a plants germinated and grew on soil for 3 weeks (A & C), and then water was withheld for 1 week (B & D).
9 Supplemental Table 1. Predicted transmembrane domains in the yeast, A. thaliana and human 3-KDS Reductases Tsc10p TSC10A TSC10B FVT1 1 HMMTop TMPred TopPred MEMSAT TMHMM PhDhtm ConPred II Tusnady, G. E. and Simon, I. (1998) Principles governing amino acid composition of integral membrane proteins: application to topology prediction. J Mol Biol. 283, Hofmann, K. and Stoffel, W. (1993) TMbase - A database of membrane spanning proteins segments. Biol. Chem. Hoppe-Seyler. 374, von Heijne, G. (1992) Membrane protein structure prediction hydrophobicity analysis and the positive-inside rule. J. Mol. Biol. 225, Jones, D. T., Taylor, W. R. and Thornton, J. M. (1994) A model recognition approach to the prediction of all-helical membrane protein structure and topology. Biochemistry. 33, Krogh, A., Larsson, B., von Heijne, G. and Sonnhammer, E. L. (2001) Predicting transmembrane protein topology with a hidden Markov model: application to complete genomes. J Mol Biol. 305, Rost, B., Yachdav, G. and Liu, J. (2004) The PredictProtein server. Nucleic acids research. 32, W Arai, M., Mitsuke, H., Ikeda, M., Xia, J. X., Kikuchi, T., Satake, M. and Shimizu, T. (2004) ConPred II: a consensus prediction method for obtaining transmembrane topology models with high reliability. Nucleic acids research. 32, W
10 Supplemental Table 2 Name Sequence Introduced restriction Purpose sites At3g06060 sense GGATCCGTCGAC GGCGGCCATTTCTCCT Sal I Expression vector in Yeast At3g06060 antisense GGATCCGTCGAC TCATTTGGTTTTGGT Sal I Expression vector in Yeast At5g19200 sense GGATCCCTCGAG GGCGGCAATTTTTTCT Xho I Expression vector in Yeast At5g19200 antisense GGATCCCTCGAG CTAAGCTAACTTACT Xho I Expression vector in Yeast AtTSC10ACS ATGCggcgcgcc ATAGCTCAACATAACCAACTTGACCGGTGTGTGAC Asc I Expression vector for complementing in Arabidopsis AtTSC10ACA ATGCggcgcgcc TTAGCTTTAGAATTGACATCGCCATCG Asc I Expression vector for complementing in Arabidopsis AtTSC10A_PmL ggatccaagctctcggcgatgtgaag BamH I Expression vector for promoter activity AtTSC10A_PmU CTGCAGcaacaaccaagaaagcatcaCC Pst I Expression vector for promoter activity AtTSC10A_GFP_S gtctctagagcttcacatcgccgagag Xba I Expression vector for subcelluar localization of AtTSC10A AtTSC10A_GFP_A aacggatccgttcaaagaccggaaacatga BamH I Expression vector for subcelluar localization of AtTSC10A AtTSC10A-QRS accgcagggcctataagatt - qrt-pcr AtTSC10A-QRA GGCTGtcatttggttttgg - qrt-pcr AtTSC10B-QF ggtttgaacaagaactgaagaag - qrt-pcr AtTSC10B-QR cttcattggttttcattgagc - qrt-pcr IRT1-QF ACCCGGGTTTGAACAAGAACT - qrt-pcr IRT1-QR CATATTTTGGCTACTTCATTGGTTTTC - qrt-pcr GABI_524F10_LP ctcaagctggtcaggttcgta - T-DNA insertion checking GABI_524F10_RP tttcattgagccagatgatgc; - T-DNA insertion checking GABI TDNA CCCATTTGGACGTGAATGTAGACAC - T-DNA insertion checking SALK_149589_LP accagctccttcgccgtaaac - T-DNA insertion checking SALK_149589_RP aagactaaccacccatatgtgaaa - T-DNA insertion checking LBa1 TGGTTCACGTAGTGGGCCATCG - T-DNA insertion checking SAIL_131A01_LP TGTGAGGGTAGCTAGGATTAGG - T-DNA insertion checking SAIL_131A01_RP AGACCGGAAACATGAATCTTG - T-DNA insertion checking LBb1 GCCTTTTCAGAAATGGATAAATAGCCTTGCTTCC - T-DNA insertion checking TSC10BGFTF GGGGACAAGTTTGTACAAAAAAGCAGGCTtcAGAAGAGTTTCAatggcggcaa- Localization TSC10BGFTR GGGGACcactttgtacaagaaagctgggtTTCCAAAGAGATAATGGTTCATACC - Localization
Supplemental Data. Cui et al. (2012). Plant Cell /tpc a b c d. Stem UBC32 ACTIN
 A Root Stem Leaf Flower Silique Senescence leaf B a b c d UBC32 ACTIN C * Supplemental Figure 1. Expression Pattern and Protein Sequence of UBC32 Homologues in Yeast, Human, and Arabidopsis. (A) Expression
A Root Stem Leaf Flower Silique Senescence leaf B a b c d UBC32 ACTIN C * Supplemental Figure 1. Expression Pattern and Protein Sequence of UBC32 Homologues in Yeast, Human, and Arabidopsis. (A) Expression
SUPPLEMENTARY INFORMATION
 AS-NMD modulates FLM-dependent thermosensory flowering response in Arabidopsis NATURE PLANTS www.nature.com/natureplants 1 Supplementary Figure 1. Genomic sequence of FLM along with the splice sites. Sequencing
AS-NMD modulates FLM-dependent thermosensory flowering response in Arabidopsis NATURE PLANTS www.nature.com/natureplants 1 Supplementary Figure 1. Genomic sequence of FLM along with the splice sites. Sequencing
Supplemental Data. Guo et al. (2015). Plant Cell /tpc
 Supplemental Figure 1. The Mutant exb1-d Displayed Pleiotropic Phenotypes and Produced Branches in the Axils of Cotyledons. (A) Branches were developed in exb1-d but not in wild-type plants. (B) and (C)
Supplemental Figure 1. The Mutant exb1-d Displayed Pleiotropic Phenotypes and Produced Branches in the Axils of Cotyledons. (A) Branches were developed in exb1-d but not in wild-type plants. (B) and (C)
scrm-d/+ scrm-d Supplemental Fig. 1.
 Kanaoka, Pillitteri, Fujii, Yoshida, Bogenschutz, Takabayashi, Zhu, and Torii (2008) SCRM/ICE1 and SCRM2 specify three cell-state transitional steps leading to Arabidopsis stomatal differentiation WT scrm-d/+
Kanaoka, Pillitteri, Fujii, Yoshida, Bogenschutz, Takabayashi, Zhu, and Torii (2008) SCRM/ICE1 and SCRM2 specify three cell-state transitional steps leading to Arabidopsis stomatal differentiation WT scrm-d/+
- 1 - Supplemental Data
 - 1-1 Supplemental Data 2 3 4 5 6 7 8 9 Supplemental Figure S1. Differential expression of AtPIP Genes in DC3000-inoculated plants. Gene expression in leaves was analyzed by real-time RT-PCR and expression
- 1-1 Supplemental Data 2 3 4 5 6 7 8 9 Supplemental Figure S1. Differential expression of AtPIP Genes in DC3000-inoculated plants. Gene expression in leaves was analyzed by real-time RT-PCR and expression
Construction of plant complementation vector and generation of transgenic plants
 MATERIAL S AND METHODS Plant materials and growth conditions Arabidopsis ecotype Columbia (Col0) was used for this study. SALK_072009, SALK_076309, and SALK_027645 were obtained from the Arabidopsis Biological
MATERIAL S AND METHODS Plant materials and growth conditions Arabidopsis ecotype Columbia (Col0) was used for this study. SALK_072009, SALK_076309, and SALK_027645 were obtained from the Arabidopsis Biological
Supplemental Data. Na Xu et al. (2016). Plant Cell /tpc
 Supplemental Figure 1. The weak fluorescence phenotype is not caused by the mutation in At3g60240. (A) A mutation mapped to the gene At3g60240. Map-based cloning strategy was used to map the mutated site
Supplemental Figure 1. The weak fluorescence phenotype is not caused by the mutation in At3g60240. (A) A mutation mapped to the gene At3g60240. Map-based cloning strategy was used to map the mutated site
Supplemental Data. Zhang et al. (2010). Plant Cell /tpc
 Supplemental Figure 1. uvs90 gene cloning The T-DNA insertion in uvs90 was identified using thermal asymmetric interlaced (TAIL)-PCR. Three rounds of amplification were performed; the second (2 nd ) and
Supplemental Figure 1. uvs90 gene cloning The T-DNA insertion in uvs90 was identified using thermal asymmetric interlaced (TAIL)-PCR. Three rounds of amplification were performed; the second (2 nd ) and
S156AT168AY175A (AAA) were purified as GST-fusion proteins and incubated with GSTfused
 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 Supplemental Materials Supplemental Figure S1 (a) Phenotype of the wild type and grik1-2 grik2-1 plants after 8 days in darkness.
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 Supplemental Materials Supplemental Figure S1 (a) Phenotype of the wild type and grik1-2 grik2-1 plants after 8 days in darkness.
Supplemental Data. Borg et al. Plant Cell (2014) /tpc
 Supplementary Figure 1 - Alignment of selected angiosperm DAZ1 and DAZ2 homologs Multiple sequence alignment of selected DAZ1 and DAZ2 homologs. A consensus sequence built using default parameters is shown
Supplementary Figure 1 - Alignment of selected angiosperm DAZ1 and DAZ2 homologs Multiple sequence alignment of selected DAZ1 and DAZ2 homologs. A consensus sequence built using default parameters is shown
Abcam.com. hutton.ac.uk. Ipmdss.dk. Bo Gong and Eva Chou
 Abcam.com Bo Gong and Eva Chou Ipmdss.dk hutton.ac.uk What is a homeotic gene? A gene which regulates the developmental fate of anatomical structures in an organism Why study them? Understand the underlying
Abcam.com Bo Gong and Eva Chou Ipmdss.dk hutton.ac.uk What is a homeotic gene? A gene which regulates the developmental fate of anatomical structures in an organism Why study them? Understand the underlying
AD-FIL AD-YAB2 -2 BD-JAZ3 AD-YAB3 AD-YAB5 AD-FIL -4 3AT BD-JAZ3 AD-YAB3
 3 4 9 10 11 12 AD-FIL AD-YAB5 2 B AD-YAB3 1 BD-JAZ 5 6 7 8 AD-FIL A AD-YAB2 Supplemental Data. Boter et al. (2015). Plant Cell 10.1105/tpc.15.00220 BD AD-YAB2-2 -2 BD-JAZ3 AD-YAB3 BD AD-YAB5-4 AD-FIL -4
3 4 9 10 11 12 AD-FIL AD-YAB5 2 B AD-YAB3 1 BD-JAZ 5 6 7 8 AD-FIL A AD-YAB2 Supplemental Data. Boter et al. (2015). Plant Cell 10.1105/tpc.15.00220 BD AD-YAB2-2 -2 BD-JAZ3 AD-YAB3 BD AD-YAB5-4 AD-FIL -4
Map-Based Cloning of Qualitative Plant Genes
 Map-Based Cloning of Qualitative Plant Genes Map-based cloning using the genetic relationship between a gene and a marker as the basis for beginning a search for a gene Chromosome walking moving toward
Map-Based Cloning of Qualitative Plant Genes Map-based cloning using the genetic relationship between a gene and a marker as the basis for beginning a search for a gene Chromosome walking moving toward
A Nucleus-Encoded Chloroplast Protein YL1 Is Involved in Chloroplast. Development and Efficient Biogenesis of Chloroplast ATP Synthase in Rice
 A Nucleus-Encoded Chloroplast Protein YL1 Is Involved in Chloroplast Development and Efficient Biogenesis of Chloroplast ATP Synthase in Rice Fei Chen 1,*, Guojun Dong 2,*, Limin Wu 1, Fang Wang 3, Xingzheng
A Nucleus-Encoded Chloroplast Protein YL1 Is Involved in Chloroplast Development and Efficient Biogenesis of Chloroplast ATP Synthase in Rice Fei Chen 1,*, Guojun Dong 2,*, Limin Wu 1, Fang Wang 3, Xingzheng
Supplemental Data. Zhou et al. (2016). Plant Cell /tpc
 Supplemental Figure 1. Confirmation of mutant mapping results. (A) Complementation assay with stably transformed genomic fragments (ComN-N) (2 kb upstream of TSS and 1.5 kb downstream of TES) and CaMV
Supplemental Figure 1. Confirmation of mutant mapping results. (A) Complementation assay with stably transformed genomic fragments (ComN-N) (2 kb upstream of TSS and 1.5 kb downstream of TES) and CaMV
A Repressor Complex Governs the Integration of
 Developmental Cell 15 Supplemental Data A Repressor Complex Governs the Integration of Flowering Signals in Arabidopsis Dan Li, Chang Liu, Lisha Shen, Yang Wu, Hongyan Chen, Masumi Robertson, Chris A.
Developmental Cell 15 Supplemental Data A Repressor Complex Governs the Integration of Flowering Signals in Arabidopsis Dan Li, Chang Liu, Lisha Shen, Yang Wu, Hongyan Chen, Masumi Robertson, Chris A.
Supplemental Data. Jing et al. (2013). Plant Cell /tpc
 Supplemental Figure 1. Characterization of epp1 Mutants. (A) Cotyledon angles of 5-d-old Col wild-type (gray bars) and epp1-1 (black bars) seedlings under red (R), far-red (FR) and blue (BL) light conditions,
Supplemental Figure 1. Characterization of epp1 Mutants. (A) Cotyledon angles of 5-d-old Col wild-type (gray bars) and epp1-1 (black bars) seedlings under red (R), far-red (FR) and blue (BL) light conditions,
Supplemental Data. Huo et al. (2013). Plant Cell /tpc
 Supplemental Data. Huo et al. (2013). Plant Cell 10.1105/tpc.112.108902 1 Supplemental Figure 1. Alignment of NCED4 Amino Acid Sequences from Different Lettuce Varieties. NCED4 amino acid sequences are
Supplemental Data. Huo et al. (2013). Plant Cell 10.1105/tpc.112.108902 1 Supplemental Figure 1. Alignment of NCED4 Amino Acid Sequences from Different Lettuce Varieties. NCED4 amino acid sequences are
Supplemental Figure 1. Rosette Leaf Morphology of Single, Double and Triple Mutants of eid3, phya-201 and phyb-5. Photographs of Ler wild type,
 Supplemental Figure 1. Rosette Leaf Morphology of Single, Double and Triple Mutants of eid3, phya-201 and phyb-5. Photographs of Ler wild type, phya-201, phyb-5, phya-201 phyb-5, eid3, phya-201 eid3, phyb-5
Supplemental Figure 1. Rosette Leaf Morphology of Single, Double and Triple Mutants of eid3, phya-201 and phyb-5. Photographs of Ler wild type, phya-201, phyb-5, phya-201 phyb-5, eid3, phya-201 eid3, phyb-5
Supplemental Materials
 Supplemental Materials Supplemental Figure S. Phenotypic assessment of alb4 mutant plants under different stress conditions. (A) High-light stress and drought stress. Wild-type (WT) and alb4 mutant plants
Supplemental Materials Supplemental Figure S. Phenotypic assessment of alb4 mutant plants under different stress conditions. (A) High-light stress and drought stress. Wild-type (WT) and alb4 mutant plants
Supplemental Figure 1
 Supplemental Figure 1 A gta2-1 gta2-2 1kb AT4G08350 B Col-0 gta2-1 LP+RP+LBb1.3 C Col-0 gta2-2 LP+RP+LB1 D Col-0 gta2-2 gta2-1 GTA2 TUBLIN E Col0 gta2-1 gta2-2 Supplemental Figure 1. Phenotypic analysis
Supplemental Figure 1 A gta2-1 gta2-2 1kb AT4G08350 B Col-0 gta2-1 LP+RP+LBb1.3 C Col-0 gta2-2 LP+RP+LB1 D Col-0 gta2-2 gta2-1 GTA2 TUBLIN E Col0 gta2-1 gta2-2 Supplemental Figure 1. Phenotypic analysis
Supplemental Figure 1.
 Supplemental Data. Charron et al. Dynamic landscapes of four histone modifications during de-etiolation in Arabidopsis. Plant Cell (2009). 10.1105/tpc.109.066845 Supplemental Figure 1. Immunodetection
Supplemental Data. Charron et al. Dynamic landscapes of four histone modifications during de-etiolation in Arabidopsis. Plant Cell (2009). 10.1105/tpc.109.066845 Supplemental Figure 1. Immunodetection
Supplemental Data. Wang et al. Plant Cell. (2013) /tpc
 SNL1 SNL Supplemental Figure 1. The Expression Patterns of SNL1 and SNL in Different Tissues of Arabidopsis from Genevestigator Web Site (https://www.genevestigator.com/gv/index.jsp). 1 A 1. 1..8.6.4..
SNL1 SNL Supplemental Figure 1. The Expression Patterns of SNL1 and SNL in Different Tissues of Arabidopsis from Genevestigator Web Site (https://www.genevestigator.com/gv/index.jsp). 1 A 1. 1..8.6.4..
Supplemental Figure 1. Floral commitment in Arabidopsis WT and mutants.
 Percentage of induction Supplemental Data. Torti et al. (2012). Plant Cell 10.1105/tpc.111.092791 A +0 +0 LD LDs +1 LDs +3 +3 LDs +5 +5 LDs LD 50 μm AP1 B C 140 120 100 80 60 40 20 0-20 -40 wt Col soc1-2
Percentage of induction Supplemental Data. Torti et al. (2012). Plant Cell 10.1105/tpc.111.092791 A +0 +0 LD LDs +1 LDs +3 +3 LDs +5 +5 LDs LD 50 μm AP1 B C 140 120 100 80 60 40 20 0-20 -40 wt Col soc1-2
Using mutants to clone genes
 Using mutants to clone genes Objectives: 1. What is positional cloning? 2. What is insertional tagging? 3. How can one confirm that the gene cloned is the same one that is mutated to give the phenotype
Using mutants to clone genes Objectives: 1. What is positional cloning? 2. What is insertional tagging? 3. How can one confirm that the gene cloned is the same one that is mutated to give the phenotype
IDN1 and IDN2: two proteins required for de novo DNA methylation in Arabidopsis thaliana.
 IDN1 and IDN2: two proteins required for de novo DNA methylation in Arabidopsis thaliana. Israel Ausin, Todd C. Mockler, Joanne Chory, and Steven E. Jacobsen Col Ler nrpd1a rdr2 dcl3-1 ago4-1 idn1-1 idn2-1
IDN1 and IDN2: two proteins required for de novo DNA methylation in Arabidopsis thaliana. Israel Ausin, Todd C. Mockler, Joanne Chory, and Steven E. Jacobsen Col Ler nrpd1a rdr2 dcl3-1 ago4-1 idn1-1 idn2-1
Supplemental Data. Farmer et al. (2010) Plant Cell /tpc
 Supplemental Figure 1. Amino acid sequence comparison of RAD23 proteins. Identical and similar residues are shown in the black and gray boxes, respectively. Dots denote gaps. The sequence of plant Ub is
Supplemental Figure 1. Amino acid sequence comparison of RAD23 proteins. Identical and similar residues are shown in the black and gray boxes, respectively. Dots denote gaps. The sequence of plant Ub is
Nature Genetics: doi: /ng Supplementary Figure 1. ChIP-seq genome browser views of BRM occupancy at previously identified BRM targets.
 Supplementary Figure 1 ChIP-seq genome browser views of BRM occupancy at previously identified BRM targets. Gene structures are shown underneath each panel. Supplementary Figure 2 pref6::ref6-gfp complements
Supplementary Figure 1 ChIP-seq genome browser views of BRM occupancy at previously identified BRM targets. Gene structures are shown underneath each panel. Supplementary Figure 2 pref6::ref6-gfp complements
Supplementary Information
 Supplementary Information Supplementary Fig. 1. Seed dormancy and germination responses of RVE1 and PIF1. (a) Diagram of RVE1 and the T-DNA insertion of the rve1-2 mutant (SAIL_326_A01). Black boxes represent
Supplementary Information Supplementary Fig. 1. Seed dormancy and germination responses of RVE1 and PIF1. (a) Diagram of RVE1 and the T-DNA insertion of the rve1-2 mutant (SAIL_326_A01). Black boxes represent
Proposed experiments to study SAR
 Proposed experiments to study SAR Question or hypothesis Proposed experiments Expectation 1. Arabidopsis can be used to study SAR Test SAR response in Arabidopsis Yes, SAR on Arabidopsis 2. What are contributing
Proposed experiments to study SAR Question or hypothesis Proposed experiments Expectation 1. Arabidopsis can be used to study SAR Test SAR response in Arabidopsis Yes, SAR on Arabidopsis 2. What are contributing
Supplemental Figure 1. Transcript profiles of Arabidopsis IPMS1 and IPMS2 in different tissues and developmental stages.
 Transcript level (intensity) Supplemental Data. Xing et al. Plant Cell (2017) 10.1105/tpc.17.00186 1600 1400 1200 1000 800 600 400 200 0 IPMS1 IPMS2 Supplemental Figure 1. Transcript profiles of Arabidopsis
Transcript level (intensity) Supplemental Data. Xing et al. Plant Cell (2017) 10.1105/tpc.17.00186 1600 1400 1200 1000 800 600 400 200 0 IPMS1 IPMS2 Supplemental Figure 1. Transcript profiles of Arabidopsis
Supplemental Data. Lee et al. Plant Cell. (2010) /tpc Supplemental Figure 1. Protein and Gene Structures of DWA1 and DWA2.
 Supplemental Figure 1. Protein and Gene Structures of DWA1 and DWA2. (A) Protein structures of DWA1 and DWA2. WD40 region was determined based on the NCBI conserved domain databases (B, C) Schematic representation
Supplemental Figure 1. Protein and Gene Structures of DWA1 and DWA2. (A) Protein structures of DWA1 and DWA2. WD40 region was determined based on the NCBI conserved domain databases (B, C) Schematic representation
Fig. S1. Molecular phylogenetic analysis of AtHD-ZIP IV family. A phylogenetic tree was constructed using Bayesian analysis with Markov Chain Monte
 Fig. S1. Molecular phylogenetic analysis of AtHD-ZIP IV family. A phylogenetic tree was constructed using Bayesian analysis with Markov Chain Monte Carlo algorithm for one million generations to obtain
Fig. S1. Molecular phylogenetic analysis of AtHD-ZIP IV family. A phylogenetic tree was constructed using Bayesian analysis with Markov Chain Monte Carlo algorithm for one million generations to obtain
Supplemental Data. Bai et al. Plant Cell. (2012) /tpc A
 A B Unconserved Conserved AIF4 AIF2 AIF3 AIF1 UPB1 IBH1 Supplemental Figure 1. IBH1, UPB1 and AIFs belong to the same HLH family. (A), Part of a phylogenetic tree constructed using conserved domains shows
A B Unconserved Conserved AIF4 AIF2 AIF3 AIF1 UPB1 IBH1 Supplemental Figure 1. IBH1, UPB1 and AIFs belong to the same HLH family. (A), Part of a phylogenetic tree constructed using conserved domains shows
Supplemental Figure 1. VLN5 retains conserved residues at both type 1 and type 2 Ca 2+ -binding
 Supplemental Figure 1. VLN5 retains conserved residues at both type 1 and type 2 Ca 2+ -binding sites in the G1 domain. Multiple sequence alignment was performed with DNAMAN6.0.40. Secondary structural
Supplemental Figure 1. VLN5 retains conserved residues at both type 1 and type 2 Ca 2+ -binding sites in the G1 domain. Multiple sequence alignment was performed with DNAMAN6.0.40. Secondary structural
Supplemental Data. Li et al. (2015). Plant Cell /tpc
 Supplemental Data Supplemental Figure 1: Characterization of asr3 T-DNA knockout lines and complementation transgenic lines. (A) The scheme of At2G33550 (ASR3) with gray boxes indicating exons and dash
Supplemental Data Supplemental Figure 1: Characterization of asr3 T-DNA knockout lines and complementation transgenic lines. (A) The scheme of At2G33550 (ASR3) with gray boxes indicating exons and dash
Functional conservation and diversification of the soybean maturity gene E1 and its homologs in legumes
 Functional conservation and diversification of the soybean maturity gene E1 and its homologs in legumes Xingzheng Zhang 1,2#, Hong Zhai 1#, Yaying Wang 1,2, Xiaojie Tian 1,2, Yupeng Zhang 1,2, Hongyan
Functional conservation and diversification of the soybean maturity gene E1 and its homologs in legumes Xingzheng Zhang 1,2#, Hong Zhai 1#, Yaying Wang 1,2, Xiaojie Tian 1,2, Yupeng Zhang 1,2, Hongyan
Supplemental Data. Seo et al. (2014). Plant Cell /tpc
 Supplemental Figure 1. Protein alignment of ABD1 from other model organisms. The alignment was performed with H. sapiens DCAF8, M. musculus DCAF8 and O. sativa Os10g0544500. The WD40 domains are underlined.
Supplemental Figure 1. Protein alignment of ABD1 from other model organisms. The alignment was performed with H. sapiens DCAF8, M. musculus DCAF8 and O. sativa Os10g0544500. The WD40 domains are underlined.
PIE1 ARP6 SWC6 KU70 ARP6 PIE1. HSA SNF2_N HELICc SANT. pie1-3 A1,A2 K1,K2 K1,K3 K3,LB2 A3, A4 A3,LB1 A1,A2 K1,K2 K1,K3. swc6-1 A3,A4.
 A B N-terminal SWC2 H2A.Z SWC6 ARP6 PIE1 HSA SNF2_N HELICc SANT C pie1-3 D PIE1 ARP6 5 Kb A1 200 bp A3 A2 LB1 arp6-3 A4 E A1,A2 A3, A4 A3,LB1 K1,K2 K1,K3 K3,LB2 SWC6 swc6-1 A1,A2 A3,A4 K1,K2 K1,K3 100
A B N-terminal SWC2 H2A.Z SWC6 ARP6 PIE1 HSA SNF2_N HELICc SANT C pie1-3 D PIE1 ARP6 5 Kb A1 200 bp A3 A2 LB1 arp6-3 A4 E A1,A2 A3, A4 A3,LB1 K1,K2 K1,K3 K3,LB2 SWC6 swc6-1 A1,A2 A3,A4 K1,K2 K1,K3 100
Supplementary Information
 Supplementary Information MED18 interaction with distinct transcription factors regulates plant immunity, flowering time and responses to hormones Supplementary Figure 1. Diagram showing T-DNA insertion
Supplementary Information MED18 interaction with distinct transcription factors regulates plant immunity, flowering time and responses to hormones Supplementary Figure 1. Diagram showing T-DNA insertion
Supplemental Figure 1. Immunoblot analysis of wild-type and mutant forms of JAZ3
 Supplemental Data. Chung and Howe (2009). A Critical Role for the TIFY Motif in Repression of Jasmonate Signaling by a Stabilized Splice Variant of the JASMONATE ZIM-domain protein JAZ10 in Arabidopsis.
Supplemental Data. Chung and Howe (2009). A Critical Role for the TIFY Motif in Repression of Jasmonate Signaling by a Stabilized Splice Variant of the JASMONATE ZIM-domain protein JAZ10 in Arabidopsis.
Using mutants to clone genes
 Using mutants to clone genes Objectives 1. What is positional cloning? 2. What is insertional tagging? 3. How can one confirm that the gene cloned is the same one that is mutated to give the phenotype
Using mutants to clone genes Objectives 1. What is positional cloning? 2. What is insertional tagging? 3. How can one confirm that the gene cloned is the same one that is mutated to give the phenotype
Fig. S1. Clustering analysis of expression array and ChIP-PCR assay in the ARF3 locus. (A) Typical examples of the transgenic plants used for
 Fig. S1. Clustering analysis of expression array and ChIP-PCR assay in the ARF3 locus. (A) Typical examples of the transgenic plants used for ChIP-chip and ChIP-PCR assays. The presence of pas1:t7:as1
Fig. S1. Clustering analysis of expression array and ChIP-PCR assay in the ARF3 locus. (A) Typical examples of the transgenic plants used for ChIP-chip and ChIP-PCR assays. The presence of pas1:t7:as1
Supplemental Figure 1
 Supplemental Figure 1 A LK sls1 lks1-2 F 1 sls1 B LK sls1 lks1-2 F 1 lks1-2 sls1 F 1 lks1-2 sls1 F 2 Col lks1-2 Col+LKS1 Col Col+LKS1 Supplemental Figure 1. Genetic analysis of sls1 mutant. (A) and (B)
Supplemental Figure 1 A LK sls1 lks1-2 F 1 sls1 B LK sls1 lks1-2 F 1 lks1-2 sls1 F 1 lks1-2 sls1 F 2 Col lks1-2 Col+LKS1 Col Col+LKS1 Supplemental Figure 1. Genetic analysis of sls1 mutant. (A) and (B)
Supplemental materials
 Supplemental materials Materials and methods for supplemental figures Yeast two-hybrid assays TAP46-PP2Ac interactions I. The TAP46 was used as the bait and the full-length cdnas of the five C subunits
Supplemental materials Materials and methods for supplemental figures Yeast two-hybrid assays TAP46-PP2Ac interactions I. The TAP46 was used as the bait and the full-length cdnas of the five C subunits
Supplemental Figure legends Figure S1. (A) (B) (C) (D) Figure S2. Figure S3. (A-E) Figure S4. Figure S5. (A, C, E, G, I) (B, D, F, H, Figure S6.
 Supplemental Figure legends Figure S1. Map-based cloning and complementation testing for ZOP1. (A) ZOP1 was mapped to a ~273-kb interval on Chromosome 1. In the interval, a single-nucleotide G to A substitution
Supplemental Figure legends Figure S1. Map-based cloning and complementation testing for ZOP1. (A) ZOP1 was mapped to a ~273-kb interval on Chromosome 1. In the interval, a single-nucleotide G to A substitution
Next Generation Genetics: Using deep sequencing to connect phenotype to genotype
 Next Generation Genetics: Using deep sequencing to connect phenotype to genotype http://1001genomes.org Korbinian Schneeberger Connecting Genotype and Phenotype Genotyping SNPs small Resequencing SVs*
Next Generation Genetics: Using deep sequencing to connect phenotype to genotype http://1001genomes.org Korbinian Schneeberger Connecting Genotype and Phenotype Genotyping SNPs small Resequencing SVs*
Supplemental Figure 1 A
 Supplemental Figure 1 A Supplemental Data. Han et al. (2016). Plant Cell 10.1105/tpc.15.00997 BamHI -1616 bp SINE repeats SalI ATG SacI +1653 bp HindIII LB HYG p35s pfwa LUC Tnos RB pfwa-bamhi-f: CGGGATCCCGCCTTTCTCTTCCTCATCTGC
Supplemental Figure 1 A Supplemental Data. Han et al. (2016). Plant Cell 10.1105/tpc.15.00997 BamHI -1616 bp SINE repeats SalI ATG SacI +1653 bp HindIII LB HYG p35s pfwa LUC Tnos RB pfwa-bamhi-f: CGGGATCCCGCCTTTCTCTTCCTCATCTGC
Supplemental Data. Benstein et al. (2013). Plant Cell /tpc
 Supplemental Figure 1. Purification of the heterologously expressed PGDH1, PGDH2 and PGDH3 enzymes by Ni-NTA affinity chromatography. Protein extracts (2 µl) of different fractions (lane 1 = total extract,
Supplemental Figure 1. Purification of the heterologously expressed PGDH1, PGDH2 and PGDH3 enzymes by Ni-NTA affinity chromatography. Protein extracts (2 µl) of different fractions (lane 1 = total extract,
Supplementary Figures 1-12
 Supplementary Figures 1-12 Supplementary Figure 1. The specificity of anti-abi1 antibody. Total Proteins extracted from the wild type seedlings or abi1-3 null mutant seedlings were used for immunoblotting
Supplementary Figures 1-12 Supplementary Figure 1. The specificity of anti-abi1 antibody. Total Proteins extracted from the wild type seedlings or abi1-3 null mutant seedlings were used for immunoblotting
Supplemental Data. Sethi et al. (2014). Plant Cell /tpc
 Supplemental Data Supplemental Figure 1. MYC2 Binds to the E-box but not the E1-box of the MPK6 Promoter. (A) E1-box and E-box (wild type) containing MPK6 promoter fragment. The region shown in red denotes
Supplemental Data Supplemental Figure 1. MYC2 Binds to the E-box but not the E1-box of the MPK6 Promoter. (A) E1-box and E-box (wild type) containing MPK6 promoter fragment. The region shown in red denotes
7.012 Problem Set 5. Question 1
 Name Section 7.012 Problem Set 5 Question 1 While studying the problem of infertility, you attempt to isolate a hypothetical rabbit gene that accounts for the prolific reproduction of rabbits. After much
Name Section 7.012 Problem Set 5 Question 1 While studying the problem of infertility, you attempt to isolate a hypothetical rabbit gene that accounts for the prolific reproduction of rabbits. After much
WiscDsLox485 ATG < > //----- E1 E2 E3 E4 E bp. Col-0 arr7 ARR7 ACTIN7. s of mrna/ng total RNA (x10 3 ) ARR7.
 A WiscDsLox8 ATG < > -9 +0 > ---------//----- E E E E E UTR +9 UTR +0 > 00bp B S D ol-0 arr7 ol-0 arr7 ARR7 ATIN7 D s of mrna/ng total RNA opie 0. ol-0 (x0 ) arr7 ARR7 Supplemental Fig.. Genotyping and
A WiscDsLox8 ATG < > -9 +0 > ---------//----- E E E E E UTR +9 UTR +0 > 00bp B S D ol-0 arr7 ol-0 arr7 ARR7 ATIN7 D s of mrna/ng total RNA opie 0. ol-0 (x0 ) arr7 ARR7 Supplemental Fig.. Genotyping and
Supplementary Fig 1. The responses of ERF109 to different hormones and stresses. (a to k) The induced expression of ERF109 in 7-day-old Arabidopsis
 Supplementary Fig 1. The responses of ERF109 to different hormones and stresses. (a to k) The induced expression of ERF109 in 7-day-old Arabidopsis seedlings expressing ERF109pro-GUS. The GUS staining
Supplementary Fig 1. The responses of ERF109 to different hormones and stresses. (a to k) The induced expression of ERF109 in 7-day-old Arabidopsis seedlings expressing ERF109pro-GUS. The GUS staining
SUPPLEMENTARY INFORMATION
 Ca 2+ /calmodulin Regulates Salicylic Acid-mediated Plant Immunity Liqun Du, Gul S. Ali, Kayla A. Simons, Jingguo Hou, Tianbao Yang, A.S.N. Reddy and B. W. Poovaiah * *To whom correspondence should be
Ca 2+ /calmodulin Regulates Salicylic Acid-mediated Plant Immunity Liqun Du, Gul S. Ali, Kayla A. Simons, Jingguo Hou, Tianbao Yang, A.S.N. Reddy and B. W. Poovaiah * *To whom correspondence should be
Supplementary Figure 1. jmj30-2 and jmj32-1 produce null mutants. (a) Schematic drawing of JMJ30 and JMJ32 genome structure showing regions amplified
 Supplementary Figure 1. jmj30-2 and jmj32-1 produce null mutants. (a) Schematic drawing of JMJ30 and JMJ32 genome structure showing regions amplified by primers used for mrna expression analysis. Gray
Supplementary Figure 1. jmj30-2 and jmj32-1 produce null mutants. (a) Schematic drawing of JMJ30 and JMJ32 genome structure showing regions amplified by primers used for mrna expression analysis. Gray
Nature Methods: doi: /nmeth Supplementary Figure 1. Construction of a sensitive TetR mediated auxotrophic off-switch.
 Supplementary Figure 1 Construction of a sensitive TetR mediated auxotrophic off-switch. A Production of the Tet repressor in yeast when conjugated to either the LexA4 or LexA8 promoter DNA binding sequences.
Supplementary Figure 1 Construction of a sensitive TetR mediated auxotrophic off-switch. A Production of the Tet repressor in yeast when conjugated to either the LexA4 or LexA8 promoter DNA binding sequences.
SUPPLEMENTARY INFORMATION
 Gene replacements and insertions in rice by intron targeting using CRISPR Cas9 Table of Contents Supplementary Figure 1. sgrna-induced targeted mutations in the OsEPSPS gene in rice protoplasts. Supplementary
Gene replacements and insertions in rice by intron targeting using CRISPR Cas9 Table of Contents Supplementary Figure 1. sgrna-induced targeted mutations in the OsEPSPS gene in rice protoplasts. Supplementary
Wt (Col-0) pom2-1 Wt (Ler) pom2-3 pom2-5. pom2-4 (0%) pom2-1 (0%)
 Figure 1 Wt (Col-0) Wt (Ler) pom2-3 pom2-5 pom2-2 pom2-4 B Wt (0%) (0%) Wt (4.5%) (4.5%) Wt (0%) (0%) Wt (4.5%) (4.5%) (4.5%) Figure 1. Phenotypes of pom2 and wild type (Wt) seedlings after germination
Figure 1 Wt (Col-0) Wt (Ler) pom2-3 pom2-5 pom2-2 pom2-4 B Wt (0%) (0%) Wt (4.5%) (4.5%) Wt (0%) (0%) Wt (4.5%) (4.5%) (4.5%) Figure 1. Phenotypes of pom2 and wild type (Wt) seedlings after germination
Supplemental Data. Wu and Xue (2010). Plant Cell /tpc
 Supplemental Data. Wu and Xue (21). Plant Cell 1.115/tpc.11.75564 A P1-S P1-A P2-S P2-A P3-S P3-A P4-S P4-A B Relative expression Relative expression C 5. 4.5 4. 3.5 3. 2.5 2. 1.5 1..5 5. 4.5 4. 3.5 3.
Supplemental Data. Wu and Xue (21). Plant Cell 1.115/tpc.11.75564 A P1-S P1-A P2-S P2-A P3-S P3-A P4-S P4-A B Relative expression Relative expression C 5. 4.5 4. 3.5 3. 2.5 2. 1.5 1..5 5. 4.5 4. 3.5 3.
Supplemental Data. Ko et al. (2014). Plant Cell /tpc Supplemental Figure 1. Evidence of T-DNA Insertion in ms142 Mutant.
 Supplemental Figure 1. Evidence of T-DNA Insertion in ms142 Mutant. (A) DNA gel blotting to hptii probe confirmed single T-DNA insertion (marked with arrow) in ms142 mutant. (B) T-DNA tagged construction
Supplemental Figure 1. Evidence of T-DNA Insertion in ms142 Mutant. (A) DNA gel blotting to hptii probe confirmed single T-DNA insertion (marked with arrow) in ms142 mutant. (B) T-DNA tagged construction
The homeo'c gene AGAMOUS (AG) in Arabidopsis. Presenta'on by: Yang Liu and Zhonghang Zhang (Daisy) February 27th, 2018
 The homeo'c gene AGAMOUS (AG) in Arabidopsis Presenta'on by: Yang Liu and Zhonghang Zhang (Daisy) February 27th, 2018 Arabidopsis Flower George Haughn, UBC Genes Direc'ng Flower Development in Arabidopsis
The homeo'c gene AGAMOUS (AG) in Arabidopsis Presenta'on by: Yang Liu and Zhonghang Zhang (Daisy) February 27th, 2018 Arabidopsis Flower George Haughn, UBC Genes Direc'ng Flower Development in Arabidopsis
Supplementary information
 Supplementary information Supplementary figures Figure S1 Level of mycdet1 protein in DET1 OE-1, OE-2 and OE-3 transgenic lines. Total protein extract from wild type Col0, det1-1 mutant and DET1 OE lines
Supplementary information Supplementary figures Figure S1 Level of mycdet1 protein in DET1 OE-1, OE-2 and OE-3 transgenic lines. Total protein extract from wild type Col0, det1-1 mutant and DET1 OE lines
Table S1. Primer Sequences used in this work. Underlined bases indicate Shine Dalgarno
 Supplemental Tables Table S1. Primer Sequences used in this work. Underlined bases indicate Shine Dalgarno sequence. Bolded bases indicate restriction sites. Primer Name Purpose Sequence MJWtopo1 removes
Supplemental Tables Table S1. Primer Sequences used in this work. Underlined bases indicate Shine Dalgarno sequence. Bolded bases indicate restriction sites. Primer Name Purpose Sequence MJWtopo1 removes
Supplemental Figure 1. The phenotypes of flc-3, wild type (Col-0), skb1-1 flc-3 and skb1-1 grown in soil for 35 days under long day
 upplemental ata Zhang et al (2011) lant ell 101105/tpc110081356 upplemental igure 1 he phenotypes of flc-3, wild type (ol-0), skb1-1 flc-3 and skb1-1 grown in soil for 35 days under long day 1 upplemental
upplemental ata Zhang et al (2011) lant ell 101105/tpc110081356 upplemental igure 1 he phenotypes of flc-3, wild type (ol-0), skb1-1 flc-3 and skb1-1 grown in soil for 35 days under long day 1 upplemental
SUPPLEMENTARY INFORMATION
 doi: 10.1038/nature06147 SUPPLEMENTARY INFORMATION Figure S1 The genomic and domain structure of Dscam. The Dscam gene comprises 24 exons, encoding a signal peptide (SP), 10 IgSF domains, 6 fibronectin
doi: 10.1038/nature06147 SUPPLEMENTARY INFORMATION Figure S1 The genomic and domain structure of Dscam. The Dscam gene comprises 24 exons, encoding a signal peptide (SP), 10 IgSF domains, 6 fibronectin
Supplementary Materials
 Supplementary Materials Table S1. Oligonucleotide sequences and PCR conditions used to amplify the indicated genes. TA = annealing temperature; gdna = genomic DNA; cdna = complementary DNA; c = concentration.
Supplementary Materials Table S1. Oligonucleotide sequences and PCR conditions used to amplify the indicated genes. TA = annealing temperature; gdna = genomic DNA; cdna = complementary DNA; c = concentration.
Supplemental Data. Wu et al. (2). Plant Cell..5/tpc RGLG Hormonal treatment H2O B RGLG µm ABA µm ACC µm GA Time (hours) µm µm MJ µm IA
 Supplemental Data. Wu et al. (2). Plant Cell..5/tpc..4. A B Supplemental Figure. Immunoblot analysis verifies the expression of the AD-PP2C and BD-RGLG proteins in the Y2H assay. Total proteins were extracted
Supplemental Data. Wu et al. (2). Plant Cell..5/tpc..4. A B Supplemental Figure. Immunoblot analysis verifies the expression of the AD-PP2C and BD-RGLG proteins in the Y2H assay. Total proteins were extracted
Sperm cells are passive cargo of the pollen tube in plant fertilization
 In the format provided by the authors and unedited. SUPPLEMENTARY INFORMATION VOLUME: 3 ARTICLE NUMBER: 17079 Sperm cells are passive cargo of the pollen tube in plant fertilization Jun Zhang 1, Qingpei
In the format provided by the authors and unedited. SUPPLEMENTARY INFORMATION VOLUME: 3 ARTICLE NUMBER: 17079 Sperm cells are passive cargo of the pollen tube in plant fertilization Jun Zhang 1, Qingpei
Supplemental Data. Steiner et al. Plant Cell. (2012) /tpc
 Supplemental Figure 1. SPY does not interact with free GST. Invitro pull-down assay using E. coli-expressed MBP-SPY and GST, GST-TCP14 and GST-TCP15. MBP-SPY was used as bait and incubated with equal amount
Supplemental Figure 1. SPY does not interact with free GST. Invitro pull-down assay using E. coli-expressed MBP-SPY and GST, GST-TCP14 and GST-TCP15. MBP-SPY was used as bait and incubated with equal amount
Schematic representation of the endogenous PALB2 locus and gene-disruption constructs
 Supplementary Figures Supplementary Figure 1. Generation of PALB2 -/- and BRCA2 -/- /PALB2 -/- DT40 cells. (A) Schematic representation of the endogenous PALB2 locus and gene-disruption constructs carrying
Supplementary Figures Supplementary Figure 1. Generation of PALB2 -/- and BRCA2 -/- /PALB2 -/- DT40 cells. (A) Schematic representation of the endogenous PALB2 locus and gene-disruption constructs carrying
Supplemental Information
 Supplemental Information Supplemental Figure 1. The Heterologous Yeast System for Screening of Arabidopsis JA Transporters. (A) Exogenous JA inhibited yeast cell growth. Yeast cells were diluted to various
Supplemental Information Supplemental Figure 1. The Heterologous Yeast System for Screening of Arabidopsis JA Transporters. (A) Exogenous JA inhibited yeast cell growth. Yeast cells were diluted to various
Biology 163 Laboratory in Genetics, Final Exam, Dec. 10, 2005
 1 Biology 163 Laboratory in Genetics, Final Exam, Dec. 10, 2005 Honor Pledge: I have neither given nor received any unauthorized help on this exam: Name Printed: Signature: 1. (2 pts) If you see the following
1 Biology 163 Laboratory in Genetics, Final Exam, Dec. 10, 2005 Honor Pledge: I have neither given nor received any unauthorized help on this exam: Name Printed: Signature: 1. (2 pts) If you see the following
Lecture 2: Biology Basics Continued. Fall 2018 August 23, 2018
 Lecture 2: Biology Basics Continued Fall 2018 August 23, 2018 Genetic Material for Life Central Dogma DNA: The Code of Life The structure and the four genomic letters code for all living organisms Adenine,
Lecture 2: Biology Basics Continued Fall 2018 August 23, 2018 Genetic Material for Life Central Dogma DNA: The Code of Life The structure and the four genomic letters code for all living organisms Adenine,
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/5/244/ra72/dc1 Supplementary Materials for An Interaction Between BZR1 and DELLAs Mediates Direct Signaling Crosstalk Between Brassinosteroids and Gibberellins
www.sciencesignaling.org/cgi/content/full/5/244/ra72/dc1 Supplementary Materials for An Interaction Between BZR1 and DELLAs Mediates Direct Signaling Crosstalk Between Brassinosteroids and Gibberellins
Supplemental Data. mir156-regulated SPL Transcription. Factors Define an Endogenous Flowering. Pathway in Arabidopsis thaliana
 Cell, Volume 138 Supplemental Data mir156-regulated SPL Transcription Factors Define an Endogenous Flowering Pathway in Arabidopsis thaliana Jia-Wei Wang, Benjamin Czech, and Detlef Weigel Table S1. Interaction
Cell, Volume 138 Supplemental Data mir156-regulated SPL Transcription Factors Define an Endogenous Flowering Pathway in Arabidopsis thaliana Jia-Wei Wang, Benjamin Czech, and Detlef Weigel Table S1. Interaction
3 -end. Sau3A. 3 -end TAA TAA. ~9.1kb SalI TAA. ~12.6kb. HindШ ATG 5,860bp TAA. ~12.7kb
 Supplemental Data. Chen et al. (2008). Badh2, encoding betaine aldehyde dehydrogenase, inhibits the biosynthesis of 2-acetyl-1-pyrroline, a major component in rice fragrance. A pcam-cah/cah Sau3A Promoter
Supplemental Data. Chen et al. (2008). Badh2, encoding betaine aldehyde dehydrogenase, inhibits the biosynthesis of 2-acetyl-1-pyrroline, a major component in rice fragrance. A pcam-cah/cah Sau3A Promoter
Identification and characterization of a pathogenicity-related gene VdCYP1 from Verticillium dahliae
 Identification and characterization of a pathogenicity-related gene VdCYP1 from Verticillium dahliae Dan-Dan Zhang*, Xin-Yan Wang*, Jie-Yin Chen*, Zhi-Qiang Kong, Yue-Jing Gui, Nan-Yang Li, Yu-Ming Bao,
Identification and characterization of a pathogenicity-related gene VdCYP1 from Verticillium dahliae Dan-Dan Zhang*, Xin-Yan Wang*, Jie-Yin Chen*, Zhi-Qiang Kong, Yue-Jing Gui, Nan-Yang Li, Yu-Ming Bao,
A subclass of HSP70s regulate development and abiotic stress responses in Arabidopsis thaliana
 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 Journal of Plant Research A subclass of HSP70s regulate development and abiotic stress responses in Arabidopsis thaliana Linna Leng 1 Qianqian Liang
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 Journal of Plant Research A subclass of HSP70s regulate development and abiotic stress responses in Arabidopsis thaliana Linna Leng 1 Qianqian Liang
DAY 1 DAY 2 DAY 3. GUS activity. Time h (ZT) Cluster 1. Cluster 2 Cluster 3. Cluster 4. Cluster 5. Cluster 6
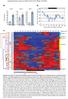 Supplemental Data. Giraud et al. (21). Plant Cell 1.115/tpc.11.74518 A) B) 6 5 4 3 2 1 DAY 1 2 15 1 5 IP SITE II DAY 2 7 6 5 4 3 2 1 IP SITE II DAY 3 IP SITE II ealtive transcript abundance 12 1 75 5 25
Supplemental Data. Giraud et al. (21). Plant Cell 1.115/tpc.11.74518 A) B) 6 5 4 3 2 1 DAY 1 2 15 1 5 IP SITE II DAY 2 7 6 5 4 3 2 1 IP SITE II DAY 3 IP SITE II ealtive transcript abundance 12 1 75 5 25
Nature Genetics: doi: /ng.3556 INTEGRATED SUPPLEMENTARY FIGURE TEMPLATE. Supplementary Figure 1
 INTEGRATED SUPPLEMENTARY FIGURE TEMPLATE Supplementary Figure 1 REF6 expression in transgenic lines. (a,b) Expression of REF6 in REF6-HA ref6 and REF6ΔZnF-HA ref6 plants detected by RT qpcr (a) and immunoblot
INTEGRATED SUPPLEMENTARY FIGURE TEMPLATE Supplementary Figure 1 REF6 expression in transgenic lines. (a,b) Expression of REF6 in REF6-HA ref6 and REF6ΔZnF-HA ref6 plants detected by RT qpcr (a) and immunoblot
Supporting Information
 Supporting Information Yuan et al. 10.1073/pnas.0906869106 Fig. S1. Heat map showing that Populus ICS is coregulated with orthologs of Arabidopsis genes involved in PhQ biosynthesis and PSI function, but
Supporting Information Yuan et al. 10.1073/pnas.0906869106 Fig. S1. Heat map showing that Populus ICS is coregulated with orthologs of Arabidopsis genes involved in PhQ biosynthesis and PSI function, but
Supplemental Data. Liu et al. (2013). Plant Cell /tpc
 Supplemental Figure 1. The GFP Tag Does Not Disturb the Physiological Functions of WDL3. (A) RT-PCR analysis of WDL3 expression in wild-type, WDL3-GFP, and WDL3 (without the GFP tag) transgenic seedlings.
Supplemental Figure 1. The GFP Tag Does Not Disturb the Physiological Functions of WDL3. (A) RT-PCR analysis of WDL3 expression in wild-type, WDL3-GFP, and WDL3 (without the GFP tag) transgenic seedlings.
Supporting Information
 Supporting Information Deng et al. 10.1073/pnas.1102117108 Fig. S1. Predicted structure of Arabidopsis bzip60 RNA. Lowest free energy form (ΔG = 309.72 (initially 343.10) of bzip60 mrna folded by M-Fold
Supporting Information Deng et al. 10.1073/pnas.1102117108 Fig. S1. Predicted structure of Arabidopsis bzip60 RNA. Lowest free energy form (ΔG = 309.72 (initially 343.10) of bzip60 mrna folded by M-Fold
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:10.1038/nature10928 Materials and Methods 1. Plant material and growth conditions. All plant lines used were in Col-0 background unless otherwise specified. pif4-101 mutant
SUPPLEMENTARY INFORMATION doi:10.1038/nature10928 Materials and Methods 1. Plant material and growth conditions. All plant lines used were in Col-0 background unless otherwise specified. pif4-101 mutant
previously. co and fca-9 alleles were isolated from the Col background. Plants were
 Supporting Online Material Materials and Methods Plant Growth Conditions The flc-3, FRI Sf2 -Col (S1), fve-4 (S2), ld-1 (S3) and fpa (S4) lines were described previously. co and fca-9 alleles were isolated
Supporting Online Material Materials and Methods Plant Growth Conditions The flc-3, FRI Sf2 -Col (S1), fve-4 (S2), ld-1 (S3) and fpa (S4) lines were described previously. co and fca-9 alleles were isolated
Supplementary Data 1.
 Supplementary Data 1. Evaluation of the effects of number of F2 progeny to be bulked (n) and average sequencing coverage (depth) of the genome (G) on the levels of false positive SNPs (SNP index = 1).
Supplementary Data 1. Evaluation of the effects of number of F2 progeny to be bulked (n) and average sequencing coverage (depth) of the genome (G) on the levels of false positive SNPs (SNP index = 1).
Supplemental Data. Hachez et al. Plant Cell (2014) /tpc Suppl. Figure 1A
 Suppl. Figure 1A Suppl. Figure 1B Supplemental Figure 1: Results of the commercial screening of interactants using split ubiquitin technique. (A) Isolated preys (192) using the bait construct pbt3-n- as
Suppl. Figure 1A Suppl. Figure 1B Supplemental Figure 1: Results of the commercial screening of interactants using split ubiquitin technique. (A) Isolated preys (192) using the bait construct pbt3-n- as
Chapter 1. from genomics to proteomics Ⅱ
 Proteomics Chapter 1. from genomics to proteomics Ⅱ 1 Functional genomics Functional genomics: study of relations of genomics to biological functions at systems level However, it cannot explain any more
Proteomics Chapter 1. from genomics to proteomics Ⅱ 1 Functional genomics Functional genomics: study of relations of genomics to biological functions at systems level However, it cannot explain any more
Supplemental Figure 1. Alignment of the NbGAPC amino acid sequences with their Arabidopsis homologues.
 Supplemental Figure 1. Alignment of the NbGAPC amino acid sequences with their Arabidopsis homologues. Homologs from N. benthamiana (NbGAPC1, NbGAPC2, NbGAPC3), Arabidopsis (AtGAPC1, AT3G04120; AtGAPC2,
Supplemental Figure 1. Alignment of the NbGAPC amino acid sequences with their Arabidopsis homologues. Homologs from N. benthamiana (NbGAPC1, NbGAPC2, NbGAPC3), Arabidopsis (AtGAPC1, AT3G04120; AtGAPC2,
TRANSGENIC ANIMALS. transient. stable. - Two methods to produce transgenic animals:
 Only for teaching purposes - not for reproduction or sale CELL TRANSFECTION transient stable TRANSGENIC ANIMALS - Two methods to produce transgenic animals: 1- DNA microinjection 2- embryonic stem cell-mediated
Only for teaching purposes - not for reproduction or sale CELL TRANSFECTION transient stable TRANSGENIC ANIMALS - Two methods to produce transgenic animals: 1- DNA microinjection 2- embryonic stem cell-mediated
Characterization of LysM-containing receptor-like kinases involved in plant innate immunity
 Characterization of LysM-containing receptor-like kinases involved in plant innate immunity Simone Ferrari Department of Biology and Biotechnology Charles Darwin Plant-pathogen interactions Plants are
Characterization of LysM-containing receptor-like kinases involved in plant innate immunity Simone Ferrari Department of Biology and Biotechnology Charles Darwin Plant-pathogen interactions Plants are
Nature Genetics: doi: /ng Supplementary Figure 1. Neighbor-joining tree of the 183 wild, cultivated, and weedy rice accessions.
 Supplementary Figure 1 Neighbor-joining tree of the 183 wild, cultivated, and weedy rice accessions. Relationships of cultivated and wild rice correspond to previously observed relationships 40. Wild rice
Supplementary Figure 1 Neighbor-joining tree of the 183 wild, cultivated, and weedy rice accessions. Relationships of cultivated and wild rice correspond to previously observed relationships 40. Wild rice
Fig. S1. TPL and TPL N176H protein interactions. (A) Semi-in vivo pull-down assays using recombinant GST N-TPL and GST N-TPL N176H fusions and
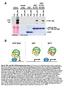 Fig. S1. TPL and TPL N176H protein interactions. (A) Semi-in vivo pull-down assays using recombinant GST N-TPL and GST N-TPL N176H fusions and transgenic Arabidopsis TPL-HA lysates. Immunoblotting of input
Fig. S1. TPL and TPL N176H protein interactions. (A) Semi-in vivo pull-down assays using recombinant GST N-TPL and GST N-TPL N176H fusions and transgenic Arabidopsis TPL-HA lysates. Immunoblotting of input
Nature Genetics: doi: /ng Supplementary Figure 1
 Supplementary Figure 1 Ihh interacts preferentially with its upstream neighboring gene Nhej1. Genes are indicated by gray lines, and Ihh and Nhej1 are highlighted in blue. 4C seq performed in E14.5 limbs
Supplementary Figure 1 Ihh interacts preferentially with its upstream neighboring gene Nhej1. Genes are indicated by gray lines, and Ihh and Nhej1 are highlighted in blue. 4C seq performed in E14.5 limbs
Candidate region (0.74 Mb) ATC TCT GGG ACT CAT GAG CAG GAG GCT AGC ATC TCT GGG ACT CAT TAG CAG GAG GCT AGC
 A idm-3 idm-3 B Physical distance (Mb) 4.6 4.86.6 8.4 C Chr.3 Recom. Rate (%) ATG 3.9.9.9 9.74 Candidate region (.74 Mb) n=4 TAA D idm-3 G3T(E4) G4A(W988) WT idm-3 ATC TCT GGG ACT CAT GAG CAG GAG GCT AGC
A idm-3 idm-3 B Physical distance (Mb) 4.6 4.86.6 8.4 C Chr.3 Recom. Rate (%) ATG 3.9.9.9 9.74 Candidate region (.74 Mb) n=4 TAA D idm-3 G3T(E4) G4A(W988) WT idm-3 ATC TCT GGG ACT CAT GAG CAG GAG GCT AGC
Intron distance to transcription start (nt)*
 Schwab et al. SOM: Enhanced microrna accumulation through stemloop-adjacent introns 1 Supplemental Table 1. Characteristics of introns used in this study Intron name Length intron (nt) Length cloned fragment
Schwab et al. SOM: Enhanced microrna accumulation through stemloop-adjacent introns 1 Supplemental Table 1. Characteristics of introns used in this study Intron name Length intron (nt) Length cloned fragment
Erhard et al. (2013). Plant Cell /tpc
 Supplemental Figure 1. c1-hbr allele structure. Diagram of the c1-hbr allele found in stocks segregating 1:1 for rpd1-1 and rpd1-2 homozygous mutants showing the presence of a 363 base pair (bp) Heartbreaker
Supplemental Figure 1. c1-hbr allele structure. Diagram of the c1-hbr allele found in stocks segregating 1:1 for rpd1-1 and rpd1-2 homozygous mutants showing the presence of a 363 base pair (bp) Heartbreaker
supplementary information
 DOI: 10.1038/ncb1979 Figure S1 Alignment and domain composition of HsTSN and PaTSN. Since both HsTSN (accession number AAA80488) and PaTSN (accession number CAL38976) have five staphylococcal nuclease
DOI: 10.1038/ncb1979 Figure S1 Alignment and domain composition of HsTSN and PaTSN. Since both HsTSN (accession number AAA80488) and PaTSN (accession number CAL38976) have five staphylococcal nuclease
Donor DNA Utilization during Gene Targeting with Zinc- finger Nucleases
 Donor DNA Utilization during Gene Targeting with Zinc- finger Nucleases Kelly J. Beumer, Jonathan K. Trautman, Kusumika Mukherjee and Dana Carroll Department of Biochemistry University of Utah School of
Donor DNA Utilization during Gene Targeting with Zinc- finger Nucleases Kelly J. Beumer, Jonathan K. Trautman, Kusumika Mukherjee and Dana Carroll Department of Biochemistry University of Utah School of
