Protein-protein interaction analysis
|
|
|
- Audra Harvey
- 6 years ago
- Views:
Transcription
1 Protein-protein interaction analysis Protein-protein interactions Golemis & Adams CSHL Press, 2005 Review and research papers (referenced on slides) doc. Jan Paleček - Matrix/beads-based: pull-down (in vitro), coip - Hybrid-based: Y2H (yeast 2-hybrid), BiFC - Proximity-based: PLA, BioID - MS-based: crosslink, D/H-exchange - Quantitative methods: SPR, ITC - Structural methods: co-crystalization, NMR - Genetic methods: synthetic lethality - Bioinformatics methods: databases, docking
2 Protein-protein interaction analysis - Matrix/beads-based: - pull-down assay - co-purification gel filtration - co-immunoprecipitation - Analysis of protein domains - Analysis of interaction surfaces - Peptide libraries - Hybrid-based: Y2H (yeast 2-hybrid), BiFC - Proximity-based: PLA, BioID - MS-based: crosslink, D/H-exchange - Quantitative methods: SPR, ITC - Structural methods: co-crystalization, NMR - Genetic methods: synthetic lethality - Bioinformatics methods: databases, docking
3 strong = nm-pm range weak = mm-µm range Palecek et al, JBC, 2006 strong interaction both proteins can be at equal concentrations (expressed/purified from bacteria or expressed/labelled in TnT in vitro expression system) weak interaction bait overexpressed vs prey from TnT Pull-down 1. tagged (e.g. GST) protein (bait) is bound to (glutathione) beads/particles (GP) 2. Partner protein (prey) is added - if the bait and prey interact then prey will be pulled down (together with bait protein) on the beads strong Nse4-SMC5 interaction GST
4 Pull-down Common tags for pull-down assay: GST (glutathione) Streptactin (biotinstreptavidin) MBP (maltose) S-tag (protein S) + tags recognized by antibodies (see co-immunoprecipitation) Weak Nse3-SMC6 interaction Palecek et al, JBC, GST-Nse3 expressed in bacteria (pre)purified on glutathione particles detected with anti-gst - Smc6 expressed in TnT (radiolabeled) radiogram detection (neither tag nor antibody needed) - control nonspecific binding of prey (Smc6 does not bind to GST-bound beads)
5 co-purification Strong interactions (protein complexes) can be recognized during the purification of the proteins (similar approach to pulldown assay) proteins can be co-purified through different tags and using gel filtration Nse1 6xHis Nse3 Strep Strep Nse4 Total extract Soluble fraction FT 1. His-tag purification 2. Strep-tag purification Elution fractions 10ml Input FT Elution fractions 1,5ml Strep- Nse4 Nse3 His- Nse Zabrady et al, NAR, 2016 Nse1-Nse3-Nse4 co-purify (interact strongly)
6 co-purification Strong interactions (protein complexes) can be recognized during the purification of the proteins (similar approach to pulldown assay) proteins can be co-purified through different tags and using gel filtration Gel filtration Nse1-Nse3 co-purify (interact strongly) Interaction strength/stability can be compared by gel filtration (NSE3-L264F mutation affects the structure and interaction of NSE3 with NSE1 resulting in broader elution peak in gel filtration) Van Crabben et al, JCI, 2016
7 Co-immunoprecipitation Similar to pull-down assay, beads/matrix/particles are used to precipitate bait protein with its bound partners Protein A-antibody Protein Tag Common tags for coimmunoprecipitation assay: GFP, FLAG, myc, HA Y These tags are recognized by specific antibodies (commercially available) TAP-tag (can be used as well): Tandem-affinity purification = immunoglobulin tag + calmodulin tag (usually used to purify complexes)
8 Co-immunoprecipitation Similar to pull-down assay, beads/matrix/particles are used to precipitate bait protein with its partners bound However, whole cell extracts are used (instead of purified proteins) co-ip Common tags for coimmunoprecipitation assay: GFP, FLAG, myc, HA FLAG-MAGEA1 NSE4 These tags are recognized by specific antibodies (commercially available) NSE1 Hudson et al, PLoS One, 2011 precipitated proteins may be associated indirectly (NSE1 is bound via NSE4 linker protein to MAGEA1) with the bait fusion protein (pulldown with pre-purified proteins is more reliable)
9 Protein-protein interaction analysis - Matrix/beads-based: - pull-down assay - co-purification gel filtration - co-immunoprecipitation - Analysis of protein domains - Analysis of interaction surfaces - Peptide libraries - Hybrid-based: Y2H (yeast 2-hybrid), BiFC - Proximity-based: FRET, PLA - MS-based: crosslink, D/H-exchange - Quantitative methods: SPR, ITC - Structural methods: co-crystalization, NMR - Genetic methods: synthetic lethality - Bioinformatics methods: databases, docking
10 Characterization of binding domain Proteins interact via their domains (motifs) analyze domain composition of your protein prepare fragments of your protein defined by domain boundaries test them in pull-down, co- Doyle et al, Mol Cell, 2010 Truncated versions of TRIM28 were used to determine MAGEC2- binding domain (only fragments A, C, D overlapping in coiled-coil domain do interact)
11 A. B. Reichman, Diploma thesis, 2009 Characterization of binding regions Proteins interact via their domains (motifs) (sometimes) only fragments of the domain can interact (can be precipitated) A. Peptide coverage of the protein B. Peptide enrichment after coimmunoprecipitation with the bait protein (red arrow points to enriched/bound peptide in MS spectra)
12 ELISA binding partner Peptide libraries region definition Proteins interact via their domains (motifs) (sometimes) only fragments of the domain can interact (can be precipitated) - peptide library can be synthetized (with conjugated biotin tag) and used in pull-down or ELISA assays (similar to antigen-epitope mapping) wells are coated with streptavidin which anchors biotinylated peptides binding partner interacts with peptide antibody against the partner with conjugated enzyme (or 2 nd antibody-enzyme) is applied - luminescence or colour detection
13 ELISA Guerineau, PhD thesis, 2013 Peptide libraries region definition Proteins interact via their domains (motifs) (sometimes) only fragments of the domain can interact (can be precipitated) - peptide library can be synthetized (with conjugated biotin tag) and used in pull-down or ELISA assays (similar to antigen-epitope mapping) wells are coated with streptavidin which anchors biotinylated peptides binding partner interacts with peptide antibody against the partner with conjugated enzyme (or 2 nd antibody-enzyme) is applied - luminescence or colour detection
14 Guerineau, PhD thesis, 2013 ELISA Peptide libraries 25 amino acids long (18) peptides library with 4 amino acids overlap (covering 90 amino acids region of Nse4 protein) peptides #6-8 bind with highest affinity, suggesting the core of the binding region
15 Peptide libraries surface mapping Proteins interact via their domains (motifs) amino acids essential for the interaction can be identified (via mutational analysis e.g. alanine substitutions = alanine scan ) - peptide library or yeast two-hybrid system (see below) can be used 21 amino acids long (20) peptides library with single amino acid alanine substitution (covering every non-ala amino acid) Guerineau, PLoS One, 2012
16 Peptide libraries alanine scan ELISA mapping Guerineau, PLoS One, 2012
17 ELISA Peptide libraries surface mapping Helical peptide is sitting in the pocket of the partner protein most peptide residues are in contact (red labeled) with the pocket (so, their mutations reduced the mutant peptide affinity), while the D204 (black labeled) residue is exposed to solvent Guerineau, PLoS One, 2012 Guerineau, PLoS One, 2012
18 Protein-protein interaction analysis - matrix/beads-based: pull-down (in vitro), coip - hybrid-based: - classical systems- domain - transcription 2-hybrid systems - reverse systems - multi-hybrid systems - alternative (membrane) systems - complementation systems fold - BiFC, DHFR - proximity/transfer system - FRET - proximity-based: PLA, BioID - MS-based: crosslink, D/H-exchange - Quantitative methods: SPR, ITC - Structural methods: co-crystalization, NMR - Genetic methods: synthetic lethality ) - Bioinformatics methods: databases, docking
19 Principal differences in hybrid systems hybrid protein hybrid protein hybrid protein hybrid protein complete domain A. In classical systems, PPI reconnects two separated domains (normally present in one protein) back to one tight complex B. In complementation systems, PPI reconnects fragments of one domain and reconstitutes its fold In FRET system, PPI enables energy transfer (see below) Stynen et al, MMBR, 2012
20 Classical yeast two-hybrid system Classical (first) yeast two-hybrid system is based on transcription factor Gal4 function Gal4 binds promotor regions (sequences) of GAL genes and activates their transcription Uetz and Finley, FEBS lett., 2005
21 Gal4-based two-hybrid system Gal4 transcription factor binds specific DNA sequence through its DNA-binding domain (DBD) - Gal4 transcription activation domain (AD) binds to general TFII factors/rna polymerase II and activates transcription machinery TFII
22 Gal4-based 2-hybrid system A. Gal4 (DBD-AD) protein activates reporter gene (lacz) B. When DNA-binding domain (DBD) and activation domain (AD) are separated, they are not able to activate transcription machinery C. When DBD and AD are fused in frame to interacting proteins (X and Y), then PPI reconnects DBD- AD and enables transcription TFII
23 Other transcription factors have been employed in two-hybrid variants: Stynen et al, MMBR, 2012
24 To detect/score transcription activation (i.e. see interaction of partner proteins), different reporter genes are used BD-Snf1/- BD-Snf1/AD-Snf4 -/AD-Snf4 Only yeast cells expressing binding partners will turn blue (as the lacz reporter will be transcribed/expressed and will convert transparent X-gal substrate to blue product) lacz enzymatic activity can be measured (thus, the strength of the PPI can be quantified)
25 Reporter genes quantitative His3 enzyme activity can be titrated by its 3- aminotriazol inhibitor auxotrophy (selective) FACSorting antibiotic resistance Stynen et al, Microbiol Mol Biol Rev, 2012
26 Yeast 2-hybrid strain example AH109 (and other strains) contains His3 and lacz reporter genes (integrated in LYS2 and URA3 genes, respectively) under different Gal4-binding promotors (GAL1 and MEL1, respectively) genotype Trp1 and Leu2 genes must be mutated to enable (auxotrophy) selection of plasmids (bearing hybrid genes) - many yeast strains exist; systems adopted to bacterial and mammalian cells exist as well
27 Yeast 2-hybrid plasmid example pgbkt7 and pgadt7 plasmids contain Gal4 BD and AD elements (to make hybrid proteins) as well as selective markers (Trp1 and Leu2 for yeast selection) myc Tag HA Tag T7 promoters in front of myc and HA tag, respectively, are suitable for additional pull-down experiments (see previous slides)
28 Yeast 2-hybrid α cells screens a cells X DBD x Y AD High-throughput screens can be done as 1. simple study: one bait is screened against AD-library (e.g. of all human hybrid proteins) - or 2. interactom study: collection of all BD-proteins is screened against ADlibrary (e.g. 6000x6000 yeast proteins = yeast interactom) Uetz et al, Nature, 2000
29 Reverse systems Vidal & Endoh, T in Biotech, 1999 For detail PPI analysis (e.g. binding surface mapping), mutation (drug) will disturb interaction - it (loss of interaction) is detected by the loss of growth of the yeast cells on selective plate (or inability to turn on the blue colour) reverse systems were developed to visualize loss of interaction
30 Reverse systems in reverse systems, PPI results in lethal phenotype yeast cells will not grow until PPI is disturbed (by mutation or drug) for example, cells expressing URA3 reporter gene will grow on plates without uracil, but these cells will be killed by 5-flouro-orothic acid (Ura3 enzyme converts FOA to toxic compound); in contrast, when PPI is disturbed, yeast cells will not express URA3 reporter gene (will not grow on plates without uracil), but these cells will not convert 5-flouro-orothic acid and therefore they will be able to grow on plates with FOA Stynen et al, MMBR, 2012
31 Reverse systems new reverse system (also called split system) is based on two transcription regulation steps: PPI activates transcription of repressor which blocks transcription of reporter gene (only when PPI is disturbed, the His3 reporter gene is transcribed) Stynen et al, MMBR, 2012
32 (multi) three-hybrid systems First three-hybrid system was developed to study RNAbinding proteins DBD-hybrid protein (1) binds one RNA motif (MS2) within the RNA-hybrid molecule (2), while the other part of the RNA-hybrid molecule (X) is recognized by AD-hybrid protein (3) this RNA-protein complex will switch on lacz reporter gene transcription in this way, you can screen ADhybrid library for RNA-X binding proteins 1. hybrid protein 2. hybrid RNA molecule 3. hybrid protein SenGupta et al, PNAS, 1996 Stynen et al, MMBR, 2012
33 Three-component 2-hybrid system DBD-hybrid protein binds one part of bridging protein, while the other part of the bridging (non-hybrid) protein is bound by AD-hybrid protein (several bridging proteins can be used) bridging protein 1. hybrid protein 2. hybrid protein DBD AD Nse1-Nse3-Nse4 complex Bednarova, Diploma thesis, 2009 Stynen et al, MMBR, 2012
34 Alternative membrane systems - Ras Number of proteins can t be used in transcription-based hybrid systems (e.g protein can t be localized to the yeast cell nucleus) CytoTrap (Ras recruitment) system is based on membrane-anchored Ras pathway reactivation A. RAS protein is activated only when human hsos-hybrid, otholog of yeast cdc25 (guanine exchange factor; cdc25-2 mutant cells are used), is anchored at the cytoplasmic membrane via interaction of myristylated hybrid-protein partner B. RAS-hybrid protein is activated when it binds to myristylated hybrid-protein partner myristyl anchor myristyl anchor 1. hybrid protein 1. hybrid protein 2. hybrid protein 2. hybrid protein Stynen et al, Microbiol Mol Biol Rev, 2012
35 Alternative membrane systems - Aga Yeast surface display system Aga2-hybrid protein is localized at the yeast surface tagged-partner interaction anchors it at the yeast surface anti-tag antibody recognizes the tagged protein fluorescence of the antibody (primary or secondary antibody) is detected and can be used for yeast strain selection (by FACS) Stynen et al, MMBR, 2012
36 Complementation systems PPI reconnects fragments of one domain and reconstitutes its fold original (A) assay based on reconstitution of ubiquitin (western blot analysis of protein degradation) new alternative versions use different detection approaches for example (B), in transcription-based approach, reporter gene is transcribed only when LexA-VP16 transcription factor is released from membrane localization Johnsson et al, PNAS, 1994 Stynen et al, MMBR, 2012
37 Complementation systems PPI reconnects fragments of ubiquitin molecule ubiquitin attracts Dub (de-ubiquitination) enzyme, which releases LexA- VP16 transcription factor from membrane - LexA-VP16 transcription factor goes to the cell nucleus and activates transcription of reporter genes Ivanusic et al, BioTech, 2015
38 Complementation systems Several systems based on complementation of different protein folds have been developed Shekhat & Ghosh, CO in ChB, 2011 Bimolecular fluorescence complementation (BiFC) PPI reconnects GFP Kodama & Hu, Biotechniques, 2012
39 Bimolecular fluorescence complementation (BiFC) binding nonbinding Pekarova et al, Plant J., 2011 Bimolecular fluorescence complementation (BiFC) PPI reconnects GFP and its fluorescence is detected Kodama & Hu, Biotechniques, 2012
40 Kodama & Hu, Biotechniques, 2012
41 Overview of yeast 2-hybrid systems Bruckner et al, IJMS, 2009
42 FRET (Forster/fluorescence resonance energy transfer) - CFP-hybrid protein emits nm light when excited (by 458nm light) when CFP-hybrid protein binds partner YFPhybrid protein, the nm emitted light excites YFP which then emits nm light (detected in the fluorescence microscope) Proximity based
43 Protein-protein interaction analysis - matrix/beads-based: pull-down (in vitro), coip - Hybrid-based: Y2H (yeast 2-hybrid), BiFC - Proximity-based: - PLA - BioID - MS-based: crosslink, D/H-exchange - Quantitative methods: SPR, ITC - Structural methods: co-crystalization, NMR - Genetic methods: synthetic lethality - Bioinformatics methods: databases, docking
44 Proximity ligation assay - PLA Weibrecht et al, ER Prot, Specific antibodies conjugated with oligonucleotides, which are complementary to circular DNA if the antibodies come close (<16nm) via PPI of they target proteins then polymerase synthesis reaction can run
45 BioID assay BirA biotin ligase (hybrid protein) biotin ligase domain biotinylates interacting (or close proximity <20nm) partner Roux, CMLS, 2013 (highly sensitive method covalently bound biotin persists even after transient interaction dissociates)
46 Protein-protein interaction analysis - beads-based: pull-down (in vitro), coip - hybridní: Y2H (kvasinkový 2-hybridní), BiFC - proximity-based: FRET, PLA - MS-based: - crosslink - D/H-exchange - Quantitative methods: SPR, ITC - Structural methods: co-crystalization, NMR - Genetic methods: synthetic lethality - Bioinformatics methods: databases, docking
47 A. ester Protein cross-linking ε-amine lysine groups react with cross linking reagent and form covalent ester bonds - MS analysis of dipeptides can show partner peptides in close proximity Sinz, MS Reviews, 2006 Bian, AJBE, 2014
48 Hydrogen/deuterium exchange single protein deuteriated peptide profile is compared to profile of the partner-bound protein (deuteriated after partner s interaction) - peptides buried inside the contact zones are not available for H/D exchange) Trcka et al, JBC, 2014
49 Protein-protein interaction analysis - overview Stynen et al, MMBR, 2012
The Two-Hybrid System
 Encyclopedic Reference of Genomics and Proteomics in Molecular Medicine The Two-Hybrid System Carolina Vollert & Peter Uetz Institut für Genetik Forschungszentrum Karlsruhe PO Box 3640 D-76021 Karlsruhe
Encyclopedic Reference of Genomics and Proteomics in Molecular Medicine The Two-Hybrid System Carolina Vollert & Peter Uetz Institut für Genetik Forschungszentrum Karlsruhe PO Box 3640 D-76021 Karlsruhe
The yeast two-hybrid assay
 Master program Molecular and Cellular Life Sciences Block 1: Intracellular membrane processes 11th October 2011 The yeast two-hybrid assay Fulvio Reggiori Department of Cell Biology, UMC Utrecht For what
Master program Molecular and Cellular Life Sciences Block 1: Intracellular membrane processes 11th October 2011 The yeast two-hybrid assay Fulvio Reggiori Department of Cell Biology, UMC Utrecht For what
Enhancers mutations that make the original mutant phenotype more extreme. Suppressors mutations that make the original mutant phenotype less extreme
 Interactomics and Proteomics 1. Interactomics The field of interactomics is concerned with interactions between genes or proteins. They can be genetic interactions, in which two genes are involved in the
Interactomics and Proteomics 1. Interactomics The field of interactomics is concerned with interactions between genes or proteins. They can be genetic interactions, in which two genes are involved in the
27041, Week 02. Review of Week 01
 27041, Week 02 Review of Week 01 The human genome sequencing project (HGP) 2 CBS, Department of Systems Biology Systems Biology and emergent properties 3 CBS, Department of Systems Biology Different model
27041, Week 02 Review of Week 01 The human genome sequencing project (HGP) 2 CBS, Department of Systems Biology Systems Biology and emergent properties 3 CBS, Department of Systems Biology Different model
Yeast 2-Hybrid Kayla Nygaard
 Yeast 2-Hybrid 2.26.18 Kayla Nygaard Y2H - What is it? A method to screen for protein-protein interactions in yeast Capitalizes on GAL4 system in yeast. GAL4 has 2 domains DNA-Binding Domain (DB) Transcriptional
Yeast 2-Hybrid 2.26.18 Kayla Nygaard Y2H - What is it? A method to screen for protein-protein interactions in yeast Capitalizes on GAL4 system in yeast. GAL4 has 2 domains DNA-Binding Domain (DB) Transcriptional
LARGE-SCALE PROTEIN INTERACTOMICS. Karl Frontzek Institute of Neuropathology
 LARGE-SCALE PROTEIN INTERACTOMICS Karl Frontzek Institute of Neuropathology STUDYING THE INTERACTOME A yeast 2 hybrid (DB: DNA binding domain, AD: activation domain) B tandem affinity purification (dashed
LARGE-SCALE PROTEIN INTERACTOMICS Karl Frontzek Institute of Neuropathology STUDYING THE INTERACTOME A yeast 2 hybrid (DB: DNA binding domain, AD: activation domain) B tandem affinity purification (dashed
20.320, notes for 9/13
 20.320 Notes Page 1 20.320, notes for 9/13 Thursday, September 13, 2012 9:36 AM Coming Assignments 1st part of design project coming up, assigned 9/14 (tomorrow) and due 9/21. Last time We covered ITC
20.320 Notes Page 1 20.320, notes for 9/13 Thursday, September 13, 2012 9:36 AM Coming Assignments 1st part of design project coming up, assigned 9/14 (tomorrow) and due 9/21. Last time We covered ITC
FRAUNHOFER IME SCREENINGPORT
 FRAUNHOFER IME SCREENINGPORT Detection technologies used in drug discovery Introduction Detection technologies in drug discovery is unlimited only a biased snapshot can be presented Differences can presented
FRAUNHOFER IME SCREENINGPORT Detection technologies used in drug discovery Introduction Detection technologies in drug discovery is unlimited only a biased snapshot can be presented Differences can presented
Technical tips Session 4
 Technical tips Session 4 Biotinylation assay: Streptavidin is a small bacterial protein that binds with high affinity to the vitamin biotin. This streptavidin-biotin combination can be used to link molecules
Technical tips Session 4 Biotinylation assay: Streptavidin is a small bacterial protein that binds with high affinity to the vitamin biotin. This streptavidin-biotin combination can be used to link molecules
Supplementary Figure 1 Collision-induced dissociation (CID) mass spectra of peptides from PPK1, PPK2, PPK3 and PPK4 respectively.
 Supplementary Figure 1 lision-induced dissociation (CID) mass spectra of peptides from PPK1, PPK, PPK3 and PPK respectively. % of nuclei with signal / field a 5 c ppif3:gus pppk1:gus 0 35 30 5 0 15 10
Supplementary Figure 1 lision-induced dissociation (CID) mass spectra of peptides from PPK1, PPK, PPK3 and PPK respectively. % of nuclei with signal / field a 5 c ppif3:gus pppk1:gus 0 35 30 5 0 15 10
Experimental Bioinformatics Lab Institute for Research in Biomedicine Barcelona Barcelona Supercomputing Center
 EMBO Practical Course on Modern Biophysical Methods for protein-ligand interactions Montse Soler-López Experimental Bioinformatics Lab Institute for Research in Biomedicine Barcelona Barcelona Supercomputing
EMBO Practical Course on Modern Biophysical Methods for protein-ligand interactions Montse Soler-López Experimental Bioinformatics Lab Institute for Research in Biomedicine Barcelona Barcelona Supercomputing
Lecture 25 (11/15/17)
 Lecture 25 (11/15/17) Reading: Ch9; 328-332 Ch25; 990-995, 1005-1012 Problems: Ch9 (study-guide: applying); 1,2 Ch9 (study-guide: facts); 7,8 Ch25 (text); 1-3,5-7,9,10,13-15 Ch25 (study-guide: applying);
Lecture 25 (11/15/17) Reading: Ch9; 328-332 Ch25; 990-995, 1005-1012 Problems: Ch9 (study-guide: applying); 1,2 Ch9 (study-guide: facts); 7,8 Ch25 (text); 1-3,5-7,9,10,13-15 Ch25 (study-guide: applying);
Supporting Information
 Supporting Information Üstün and Börnke, 2015 Figure S1: XopJ does not display acetyltransferase activity. Autoacetylation activity in vitro. Acetylation reactions using MBP, MBP-XopJ, GST and GST-HopZ1a
Supporting Information Üstün and Börnke, 2015 Figure S1: XopJ does not display acetyltransferase activity. Autoacetylation activity in vitro. Acetylation reactions using MBP, MBP-XopJ, GST and GST-HopZ1a
Contents... vii. List of Figures... xii. List of Tables... xiv. Abbreviatons... xv. Summary... xvii. 1. Introduction In vitro evolution...
 vii Contents Contents... vii List of Figures... xii List of Tables... xiv Abbreviatons... xv Summary... xvii 1. Introduction...1 1.1 In vitro evolution... 1 1.2 Phage Display Technology... 3 1.3 Cell surface
vii Contents Contents... vii List of Figures... xii List of Tables... xiv Abbreviatons... xv Summary... xvii 1. Introduction...1 1.1 In vitro evolution... 1 1.2 Phage Display Technology... 3 1.3 Cell surface
Solutions to 7.02 Quiz II 10/27/05
 Solutions to 7.02 Quiz II 10/27/05 Class Average = 83 Standard Deviation = 9 Range Grade % 87-100 A 43 74-86 B 39 55-73 C 17 > 54 D 1 Question 1 (56 points) While studying deep sea bacteria, you discover
Solutions to 7.02 Quiz II 10/27/05 Class Average = 83 Standard Deviation = 9 Range Grade % 87-100 A 43 74-86 B 39 55-73 C 17 > 54 D 1 Question 1 (56 points) While studying deep sea bacteria, you discover
Lecture 22 Eukaryotic Genes and Genomes III
 Lecture 22 Eukaryotic Genes and Genomes III In the last three lectures we have thought a lot about analyzing a regulatory system in S. cerevisiae, namely Gal regulation that involved a hand full of genes.
Lecture 22 Eukaryotic Genes and Genomes III In the last three lectures we have thought a lot about analyzing a regulatory system in S. cerevisiae, namely Gal regulation that involved a hand full of genes.
7.03, 2006, Lecture 23 Eukaryotic Genes and Genomes IV
 1 Fall 2006 7.03 7.03, 2006, Lecture 23 Eukaryotic Genes and Genomes IV In the last three lectures we have thought a lot about analyzing a regulatory system in S. cerevisiae, namely Gal regulation that
1 Fall 2006 7.03 7.03, 2006, Lecture 23 Eukaryotic Genes and Genomes IV In the last three lectures we have thought a lot about analyzing a regulatory system in S. cerevisiae, namely Gal regulation that
7.05, 2005, Lecture 23 Eukaryotic Genes and Genomes IV
 7.05, 2005, Lecture 23 Eukaryotic Genes and Genomes IV In the last three lectures we have thought a lot about analyzing a regulatory system in S. cerevisiae, namely Gal regulation that involved a hand
7.05, 2005, Lecture 23 Eukaryotic Genes and Genomes IV In the last three lectures we have thought a lot about analyzing a regulatory system in S. cerevisiae, namely Gal regulation that involved a hand
Technical tips Session 5
 Technical tips Session 5 Chromatine Immunoprecipitation (ChIP): This is a powerful in vivo method to quantitate interaction of proteins associated with specific regions of the genome. It involves the immunoprecipitation
Technical tips Session 5 Chromatine Immunoprecipitation (ChIP): This is a powerful in vivo method to quantitate interaction of proteins associated with specific regions of the genome. It involves the immunoprecipitation
GFP as affinity tag for immunoprecipitation
 1/5 Preamble This document compares existing conventional antibodies against GFP in biochemical studies with the GFPTrap, a readytouse pulldown reagent derived from alpaca. What are the differences between
1/5 Preamble This document compares existing conventional antibodies against GFP in biochemical studies with the GFPTrap, a readytouse pulldown reagent derived from alpaca. What are the differences between
CHAPTER 9 DNA Technologies
 CHAPTER 9 DNA Technologies Recombinant DNA Artificially created DNA that combines sequences that do not occur together in the nature Basis of much of the modern molecular biology Molecular cloning of genes
CHAPTER 9 DNA Technologies Recombinant DNA Artificially created DNA that combines sequences that do not occur together in the nature Basis of much of the modern molecular biology Molecular cloning of genes
Fig. S1. CrgA intracellular levels in M. tuberculosis. Ten and twenty micrograms of
 Supplementary data Fig. S1. CrgA intracellular levels in M. tuberculosis. Ten and twenty micrograms of cell free protein lysates from WT M. tuberculosis (Rv) together with various known concentrations
Supplementary data Fig. S1. CrgA intracellular levels in M. tuberculosis. Ten and twenty micrograms of cell free protein lysates from WT M. tuberculosis (Rv) together with various known concentrations
Design. Construction. Characterization
 Design Construction Characterization DNA mrna (messenger) A C C transcription translation C A C protein His A T G C T A C G Plasmids replicon copy number incompatibility selection marker origin of replication
Design Construction Characterization DNA mrna (messenger) A C C transcription translation C A C protein His A T G C T A C G Plasmids replicon copy number incompatibility selection marker origin of replication
Experimental Techniques 2
 Experimental Techniques 2 High-throughput interaction detection Yeast two-hybrid - pairwise organisms as machines to learn about organisms yeast, worm, fly, human,... low intersection between repeated
Experimental Techniques 2 High-throughput interaction detection Yeast two-hybrid - pairwise organisms as machines to learn about organisms yeast, worm, fly, human,... low intersection between repeated
Molecular Cloning. Joseph Sambrook. David W. Russell A LABORATORY MANUAL COLD SPRING HARBOR LABORATORY PRESS VOLUME.
 VOLUME Molecular Cloning A LABORATORY MANUAL THIRD EDITION www.molecularcloning.com Joseph Sambrook PETER MACCALLUM CANCER INSTITUTE AND THE UNIVERSITY OF MELBOURNE, AUSTRALIA David W. Russell UNIVERSITY
VOLUME Molecular Cloning A LABORATORY MANUAL THIRD EDITION www.molecularcloning.com Joseph Sambrook PETER MACCALLUM CANCER INSTITUTE AND THE UNIVERSITY OF MELBOURNE, AUSTRALIA David W. Russell UNIVERSITY
Supplementary Results Supplementary Table 1. P1 and P2 enrichment scores for wild-type subtiligase.
 Supplementary Results Supplementary Table 1. P1 and P2 enrichment scores for wild-type subtiligase. Supplementary Table 2. Masses of subtiligase mutants measured by LC-MS. Supplementary Table 2 (cont d).
Supplementary Results Supplementary Table 1. P1 and P2 enrichment scores for wild-type subtiligase. Supplementary Table 2. Masses of subtiligase mutants measured by LC-MS. Supplementary Table 2 (cont d).
Selected Techniques Part I
 1 Selected Techniques Part I Gel Electrophoresis Can be both qualitative and quantitative Qualitative About what size is the fragment? How many fragments are present? Is there in insert or not? Quantitative
1 Selected Techniques Part I Gel Electrophoresis Can be both qualitative and quantitative Qualitative About what size is the fragment? How many fragments are present? Is there in insert or not? Quantitative
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION Supplementary figures Supplementary Figure 1: Suv39h1, but not Suv39h2, promotes HP1α sumoylation in vivo. In vivo HP1α sumoylation assay. Top: experimental scheme. Middle: we
SUPPLEMENTARY INFORMATION Supplementary figures Supplementary Figure 1: Suv39h1, but not Suv39h2, promotes HP1α sumoylation in vivo. In vivo HP1α sumoylation assay. Top: experimental scheme. Middle: we
Perform reproducible immunoprecipitation in less than 40 minutes
 Perform reproducible immunoprecipitation in less than 40 minutes Dynabeads products Immunoprecipitation made easy with low nonspecific binding, high yield, and reproducibility Magnetic beads have become
Perform reproducible immunoprecipitation in less than 40 minutes Dynabeads products Immunoprecipitation made easy with low nonspecific binding, high yield, and reproducibility Magnetic beads have become
CELL BIOLOGY - CLUTCH CH TECHNIQUES IN CELL BIOLOGY.
 !! www.clutchprep.com CONCEPT: LIGHT MICROSCOPE A light microscope uses light waves and magnification to view specimens Can be used to visualize transparent, and some cellular components - Can visualize
!! www.clutchprep.com CONCEPT: LIGHT MICROSCOPE A light microscope uses light waves and magnification to view specimens Can be used to visualize transparent, and some cellular components - Can visualize
Chapter 9 Proteomics. From genomics to proteomics
 Chapter 9 Proteomics From genomics to proteomics From structural genomics, functional genomics to proteomics http://www.nature.com/scitable/topicpage/the-proteome-discovering-the-structure-and-function-613
Chapter 9 Proteomics From genomics to proteomics From structural genomics, functional genomics to proteomics http://www.nature.com/scitable/topicpage/the-proteome-discovering-the-structure-and-function-613
Supplementary Fig. 1. Schematic structure of TRAIP and RAP80. The prey line below TRAIP indicates bait and the two lines above RAP80 highlight the
 Supplementary Fig. 1. Schematic structure of TRAIP and RAP80. The prey line below TRAIP indicates bait and the two lines above RAP80 highlight the prey clones identified in the yeast two hybrid screen.
Supplementary Fig. 1. Schematic structure of TRAIP and RAP80. The prey line below TRAIP indicates bait and the two lines above RAP80 highlight the prey clones identified in the yeast two hybrid screen.
Supplementary Fig. 1 Identification of Nedd4 as an IRS-2-associated protein in camp-treated FRTL-5 cells.
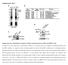 Supplementary Fig. 1 Supplementary Fig. 1 Identification of Nedd4 as an IRS-2-associated protein in camp-treated FRTL-5 cells. (a) FRTL-5 cells were treated with 1 mm dibutyryl camp for 24 h, and the lysates
Supplementary Fig. 1 Supplementary Fig. 1 Identification of Nedd4 as an IRS-2-associated protein in camp-treated FRTL-5 cells. (a) FRTL-5 cells were treated with 1 mm dibutyryl camp for 24 h, and the lysates
Gene Expression Technology
 Gene Expression Technology Bing Zhang Department of Biomedical Informatics Vanderbilt University bing.zhang@vanderbilt.edu Gene expression Gene expression is the process by which information from a gene
Gene Expression Technology Bing Zhang Department of Biomedical Informatics Vanderbilt University bing.zhang@vanderbilt.edu Gene expression Gene expression is the process by which information from a gene
Chapter One. Construction of a Fluorescent α5 Subunit. Elucidation of the unique contribution of the α5 subunit is complicated by several factors
 4 Chapter One Construction of a Fluorescent α5 Subunit The significance of the α5 containing nachr receptor (α5* receptor) has been a challenging question for researchers since its characterization by
4 Chapter One Construction of a Fluorescent α5 Subunit The significance of the α5 containing nachr receptor (α5* receptor) has been a challenging question for researchers since its characterization by
Practice Problems 5. Location of LSA-GFP fluorescence
 Life Sciences 1a Practice Problems 5 1. Soluble proteins that are normally secreted from the cell into the extracellular environment must go through a series of steps referred to as the secretory pathway.
Life Sciences 1a Practice Problems 5 1. Soluble proteins that are normally secreted from the cell into the extracellular environment must go through a series of steps referred to as the secretory pathway.
Supporting Information
 Supporting Information Wiley-VCH 2006 69451 Weinheim, Germany RNA ligands that distinguish metabolite-induced conformations in the TPP riboswitch Günter Mayer, Marie-Sophie L. Raddatz, Julia D. Grunwald,
Supporting Information Wiley-VCH 2006 69451 Weinheim, Germany RNA ligands that distinguish metabolite-induced conformations in the TPP riboswitch Günter Mayer, Marie-Sophie L. Raddatz, Julia D. Grunwald,
supplementary information
 supplementary information Figure S1 (a) Alignment of the Sgf11 amino acid sequences from S. cerevisiae, Yarrowia lipolytica and S. pombe with Sgf73 sequences from S. cerevisiae, S. pombe and Candida albicans.
supplementary information Figure S1 (a) Alignment of the Sgf11 amino acid sequences from S. cerevisiae, Yarrowia lipolytica and S. pombe with Sgf73 sequences from S. cerevisiae, S. pombe and Candida albicans.
Figure S1. Immunoblotting showing proper expression of tagged R and AVR proteins in N. benthamiana.
 SUPPLEMENTARY FIGURES Figure S1. Immunoblotting showing proper expression of tagged R and AVR proteins in N. benthamiana. YFP:, :CFP, HA: and (A) and HA, CFP and YFPtagged and AVR1-CO39 (B) were expressed
SUPPLEMENTARY FIGURES Figure S1. Immunoblotting showing proper expression of tagged R and AVR proteins in N. benthamiana. YFP:, :CFP, HA: and (A) and HA, CFP and YFPtagged and AVR1-CO39 (B) were expressed
pgbkt7 Anti- Myc AH109 strain (KDa) 50
 pgbkt7 (KDa) 50 37 Anti- Myc AH109 strain Supplementary Figure 1. Protein expression of CRN and TDR in yeast. To analyse the protein expression of CRNKD and TDRKD, total proteins extracted from yeast culture
pgbkt7 (KDa) 50 37 Anti- Myc AH109 strain Supplementary Figure 1. Protein expression of CRN and TDR in yeast. To analyse the protein expression of CRNKD and TDRKD, total proteins extracted from yeast culture
Newsletter Issue 7 One-STrEP Analysis of Protein:Protein-Interactions
 www.iba-biotagnology.com Newsletter Issue 7 One-STrEP Analysis of Protein:Protein-Interactions Strep-tag and One-STrEP-tag PPI Analysis with the co-precipitation/ mass spectrometry approach 3 Background
www.iba-biotagnology.com Newsletter Issue 7 One-STrEP Analysis of Protein:Protein-Interactions Strep-tag and One-STrEP-tag PPI Analysis with the co-precipitation/ mass spectrometry approach 3 Background
SUPPLEMENTAL MATERIAL. Supplemental material contains Supplemental Figure Legends and Supplemental Figures 1 to
 SUPPLEMENTAL MATERIAL Supplemental material contains Supplemental Figure Legends and Supplemental Figures 1 to 6. SUPPLEMENTAL FIGURE LEGENDS Supplemental Figure 1. Overview of the mechanisms by which
SUPPLEMENTAL MATERIAL Supplemental material contains Supplemental Figure Legends and Supplemental Figures 1 to 6. SUPPLEMENTAL FIGURE LEGENDS Supplemental Figure 1. Overview of the mechanisms by which
Phage Antibody Selection With Reichert SPR System
 Phage Antibody Selection With Reichert SPR System Reichert Technologies Webinar April 4, 2016 Mark A. Sullivan, Ph.D. Department of Microbiology and Immunology University of Rochester Presentation Outline
Phage Antibody Selection With Reichert SPR System Reichert Technologies Webinar April 4, 2016 Mark A. Sullivan, Ph.D. Department of Microbiology and Immunology University of Rochester Presentation Outline
This is the author's accepted version of the manuscript.
 This is the author's accepted version of the manuscript. The definitive version is published in Nature Communications Online Edition: 2015/4/16 (Japan time), doi:10.1038/ncomms7780. The final version published
This is the author's accepted version of the manuscript. The definitive version is published in Nature Communications Online Edition: 2015/4/16 (Japan time), doi:10.1038/ncomms7780. The final version published
transcription and the promoter occupancy of Smad proteins. (A) HepG2 cells were co-transfected with the wwp-luc reporter, and FLAG-tagged FHL1,
 Supplementary Data Supplementary Figure Legends Supplementary Figure 1 FHL-mediated TGFβ-responsive reporter transcription and the promoter occupancy of Smad proteins. (A) HepG2 cells were co-transfected
Supplementary Data Supplementary Figure Legends Supplementary Figure 1 FHL-mediated TGFβ-responsive reporter transcription and the promoter occupancy of Smad proteins. (A) HepG2 cells were co-transfected
Name with Last Name, First: BIOE111: Functional Biomaterial Development and Characterization MIDTERM EXAM (October 7, 2010) 93 TOTAL POINTS
 BIOE111: Functional Biomaterial Development and Characterization MIDTERM EXAM (October 7, 2010) 93 TOTAL POINTS Question 0: Fill in your name and student ID on each page. (1) Question 1: What is the role
BIOE111: Functional Biomaterial Development and Characterization MIDTERM EXAM (October 7, 2010) 93 TOTAL POINTS Question 0: Fill in your name and student ID on each page. (1) Question 1: What is the role
TECHNICAL BULLETIN. In Vitro Bacterial Split Fluorescent Protein Fold n Glow Solubility Assay Kits
 In Vitro Bacterial Split Fluorescent Protein Fold n Glow Solubility Assay Kits Catalog Numbers APPA001 In Vitro Bacterial Split GFP "Fold 'n' Glow" Solubility Assay Kit (Green) APPA008 In Vitro Bacterial
In Vitro Bacterial Split Fluorescent Protein Fold n Glow Solubility Assay Kits Catalog Numbers APPA001 In Vitro Bacterial Split GFP "Fold 'n' Glow" Solubility Assay Kit (Green) APPA008 In Vitro Bacterial
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/5/244/ra72/dc1 Supplementary Materials for An Interaction Between BZR1 and DELLAs Mediates Direct Signaling Crosstalk Between Brassinosteroids and Gibberellins
www.sciencesignaling.org/cgi/content/full/5/244/ra72/dc1 Supplementary Materials for An Interaction Between BZR1 and DELLAs Mediates Direct Signaling Crosstalk Between Brassinosteroids and Gibberellins
Chapter 20 DNA Technology & Genomics. If we can, should we?
 Chapter 20 DNA Technology & Genomics If we can, should we? Biotechnology Genetic manipulation of organisms or their components to make useful products Humans have been doing this for 1,000s of years plant
Chapter 20 DNA Technology & Genomics If we can, should we? Biotechnology Genetic manipulation of organisms or their components to make useful products Humans have been doing this for 1,000s of years plant
11/22/13. Proteomics, functional genomics, and systems biology. Biosciences 741: Genomics Fall, 2013 Week 11
 Proteomics, functional genomics, and systems biology Biosciences 741: Genomics Fall, 2013 Week 11 1 Figure 6.1 The future of genomics Functional Genomics The field of functional genomics represents the
Proteomics, functional genomics, and systems biology Biosciences 741: Genomics Fall, 2013 Week 11 1 Figure 6.1 The future of genomics Functional Genomics The field of functional genomics represents the
ALP (alkaline phosphatase) calibrators were analyzed manually in microtiter plates to find the linearity range by following this protocol:
 Exam Mol 3008 May 2009 Subject 1 (15p) ALP (alkaline phosphatase) calibrators were analyzed manually in microtiter plates to find the linearity range by following this protocol: Reaction solutions: 50
Exam Mol 3008 May 2009 Subject 1 (15p) ALP (alkaline phosphatase) calibrators were analyzed manually in microtiter plates to find the linearity range by following this protocol: Reaction solutions: 50
Polyclonal ARHGAP25 antibody was prepared from rabbit serum after intracutaneous
 Preparation and purification of polyclonal antibodies Polyclonal ARHGAP25 antibody was prepared from rabbit serum after intracutaneous injections of glutathione S-transferase-ARHGAP25-(509-619) (GST-coiled
Preparation and purification of polyclonal antibodies Polyclonal ARHGAP25 antibody was prepared from rabbit serum after intracutaneous injections of glutathione S-transferase-ARHGAP25-(509-619) (GST-coiled
Supporting Information
 Supporting Information Koh et al. 10.1073/pnas.1212917110 SI Materials and Methods Protein Purification. N-terminal His 6 -Dicer was purified as previously described with several modifications (1). After
Supporting Information Koh et al. 10.1073/pnas.1212917110 SI Materials and Methods Protein Purification. N-terminal His 6 -Dicer was purified as previously described with several modifications (1). After
Multiple choice questions (numbers in brackets indicate the number of correct answers)
 1 Multiple choice questions (numbers in brackets indicate the number of correct answers) February 1, 2013 1. Ribose is found in Nucleic acids Proteins Lipids RNA DNA (2) 2. Most RNA in cells is transfer
1 Multiple choice questions (numbers in brackets indicate the number of correct answers) February 1, 2013 1. Ribose is found in Nucleic acids Proteins Lipids RNA DNA (2) 2. Most RNA in cells is transfer
Chapter 14 Regulation of Transcription
 Chapter 14 Regulation of Transcription Cis-acting sequences Distance-independent cis-acting elements Dissecting regulatory elements Transcription factors Overview transcriptional regulation Transcription
Chapter 14 Regulation of Transcription Cis-acting sequences Distance-independent cis-acting elements Dissecting regulatory elements Transcription factors Overview transcriptional regulation Transcription
Supplementary Figure 1 Telomerase RNA fragments used in single-molecule FRET experiments. A pseudoknot fragment (nts ) labeled at position U42
 Supplementary Figure 1 Telomerase RNA fragments used in single-molecule FRET experiments. A pseudoknot fragment (nts 32-195) labeled at position U42 with Cy3 (green circle) was constructed by a two piece
Supplementary Figure 1 Telomerase RNA fragments used in single-molecule FRET experiments. A pseudoknot fragment (nts 32-195) labeled at position U42 with Cy3 (green circle) was constructed by a two piece
Supplemental Data. Wu et al. (2). Plant Cell..5/tpc RGLG Hormonal treatment H2O B RGLG µm ABA µm ACC µm GA Time (hours) µm µm MJ µm IA
 Supplemental Data. Wu et al. (2). Plant Cell..5/tpc..4. A B Supplemental Figure. Immunoblot analysis verifies the expression of the AD-PP2C and BD-RGLG proteins in the Y2H assay. Total proteins were extracted
Supplemental Data. Wu et al. (2). Plant Cell..5/tpc..4. A B Supplemental Figure. Immunoblot analysis verifies the expression of the AD-PP2C and BD-RGLG proteins in the Y2H assay. Total proteins were extracted
This Technical Note describes these characterization studies in more detail.
 Technical Note Characterization of SOMAmer Reagents Binding Specificity in the SOMAscan 1.3k Assay Summary Slow Off-Rate Modified Aptamer (SOMAmer) reagents are identified via the SELEX process against
Technical Note Characterization of SOMAmer Reagents Binding Specificity in the SOMAscan 1.3k Assay Summary Slow Off-Rate Modified Aptamer (SOMAmer) reagents are identified via the SELEX process against
Chapter 17: Immunization & Immune Testing. 1. Immunization 2. Diagnostic Immunology
 Chapter 17: Immunization & Immune Testing 1. Immunization 2. Diagnostic Immunology 1. Immunization Chapter Reading pp. 505-511 What is Immunization? A method of inducing artificial immunity by exposing
Chapter 17: Immunization & Immune Testing 1. Immunization 2. Diagnostic Immunology 1. Immunization Chapter Reading pp. 505-511 What is Immunization? A method of inducing artificial immunity by exposing
1. Immunization. What is Immunization? 12/9/2016. Chapter 17: Immunization & Immune Testing. 1. Immunization 2. Diagnostic Immunology
 Chapter 17: Immunization & Immune Testing 1. Immunization 2. Diagnostic Immunology 1. Immunization Chapter Reading pp. 505-511 What is Immunization? A method of inducing artificial immunity by exposing
Chapter 17: Immunization & Immune Testing 1. Immunization 2. Diagnostic Immunology 1. Immunization Chapter Reading pp. 505-511 What is Immunization? A method of inducing artificial immunity by exposing
Recitation CHAPTER 9 DNA Technologies
 Recitation CHAPTER 9 DNA Technologies DNA Cloning: General Scheme A cloning vector and eukaryotic chromosomes are separately cleaved with the same restriction endonuclease. (A single chromosome is shown
Recitation CHAPTER 9 DNA Technologies DNA Cloning: General Scheme A cloning vector and eukaryotic chromosomes are separately cleaved with the same restriction endonuclease. (A single chromosome is shown
Overview of Solulink Products. June 2011
 Overview of Solulink Products June 2011 1 Who we are Established in 2003, Solulink develops, patents, manufactures, and sells consumables to over 1,000 customers in life science, diagnostic, and pharmaceutical
Overview of Solulink Products June 2011 1 Who we are Established in 2003, Solulink develops, patents, manufactures, and sells consumables to over 1,000 customers in life science, diagnostic, and pharmaceutical
Challenges to measuring intracellular Ca 2+ Calmodulin: nature s Ca 2+ sensor
 Calcium Signals in Biological Systems Lecture 3 (2/9/0) Measuring intracellular Ca 2+ signals II: Genetically encoded Ca 2+ sensors Henry M. Colecraft, Ph.D. Challenges to measuring intracellular Ca 2+
Calcium Signals in Biological Systems Lecture 3 (2/9/0) Measuring intracellular Ca 2+ signals II: Genetically encoded Ca 2+ sensors Henry M. Colecraft, Ph.D. Challenges to measuring intracellular Ca 2+
The Biotechnology Toolbox
 Chapter 15 The Biotechnology Toolbox Cutting and Pasting DNA Cutting DNA Restriction endonuclease or restriction enzymes Cellular protection mechanism for infected foreign DNA Recognition and cutting specific
Chapter 15 The Biotechnology Toolbox Cutting and Pasting DNA Cutting DNA Restriction endonuclease or restriction enzymes Cellular protection mechanism for infected foreign DNA Recognition and cutting specific
NPTEL VIDEO COURSE PROTEOMICS PROF. SANJEEVA SRIVASTAVA
 LECTURE-26 Interactomics: Yeast two-hybrid immunoprecipitation protein microarrays TRANSCRIPT Welcome to the proteomics course. Today we will talk about interactomics, which is studying interactions of
LECTURE-26 Interactomics: Yeast two-hybrid immunoprecipitation protein microarrays TRANSCRIPT Welcome to the proteomics course. Today we will talk about interactomics, which is studying interactions of
Supplementary data. sienigma. F-Enigma F-EnigmaSM. a-p53
 Supplementary data Supplemental Figure 1 A sienigma #2 sienigma sicontrol a-enigma - + ++ - - - - - - + ++ - - - - - - ++ B sienigma F-Enigma F-EnigmaSM a-flag HLK3 cells - - - + ++ + ++ - + - + + - -
Supplementary data Supplemental Figure 1 A sienigma #2 sienigma sicontrol a-enigma - + ++ - - - - - - + ++ - - - - - - ++ B sienigma F-Enigma F-EnigmaSM a-flag HLK3 cells - - - + ++ + ++ - + - + + - -
Nature Structural & Molecular Biology: doi: /nsmb.1583
 Acetylation by GCN5 regulates CDC6 phosphorylation in the S-phase of the cell cycle Roberta Paolinelli 1,2, Ramiro Mendoza-Maldonado 2, Anna Cereseto 1 and Mauro Giacca 2 1 Molecular Biology Laboratory,
Acetylation by GCN5 regulates CDC6 phosphorylation in the S-phase of the cell cycle Roberta Paolinelli 1,2, Ramiro Mendoza-Maldonado 2, Anna Cereseto 1 and Mauro Giacca 2 1 Molecular Biology Laboratory,
STUDYING THE INTERACTOME WITH THE YEAST TWO-HYBRID SYSTEM AND MASS SPECTROMETRY
 STUDYING THE INTERACTOME WITH THE YEAST TWO-HYBRID SYSTEM AND MASS SPECTROMETRY Barry Causier* School of Biology, University of Leeds, Leeds LS2 9JT, United Kingdom Received 24 July 2003; received (revised)
STUDYING THE INTERACTOME WITH THE YEAST TWO-HYBRID SYSTEM AND MASS SPECTROMETRY Barry Causier* School of Biology, University of Leeds, Leeds LS2 9JT, United Kingdom Received 24 July 2003; received (revised)
PHYS 498 HW3 Solutions: 1. We have two equations: (1) (2)
 PHYS 498 HW3 Solutions: 1. We have two equations: (1) (2) Where Conc is the initial concentration of [B] or [SA] Since [B] = [SA], the second equation simplifies to: (3) Using equation (1) and (3), we
PHYS 498 HW3 Solutions: 1. We have two equations: (1) (2) Where Conc is the initial concentration of [B] or [SA] Since [B] = [SA], the second equation simplifies to: (3) Using equation (1) and (3), we
MagSi Beads. Magnetic Silica Beads for Life Science and Biotechnology study
 MagSi Beads Magnetic Silica Beads for Life Science and Biotechnology study MagnaMedics Diagnostics B.V. / Rev. 9.2 / 2012 Wide range of products for numerous applications MagnaMedics separation solutions
MagSi Beads Magnetic Silica Beads for Life Science and Biotechnology study MagnaMedics Diagnostics B.V. / Rev. 9.2 / 2012 Wide range of products for numerous applications MagnaMedics separation solutions
Caspase-2 BiFC plasmids
 Caspase-2 BiFC plasmids Caspase-2 BiFC - background Bimolecular Fluorescence Complementation (BiFC) describes the use of split fluorescent proteins to measure protein-protein interactions in cells. The
Caspase-2 BiFC plasmids Caspase-2 BiFC - background Bimolecular Fluorescence Complementation (BiFC) describes the use of split fluorescent proteins to measure protein-protein interactions in cells. The
Fig. S1. Effect of p120-catenin overexpression on the interaction of SCUBE2 with E-cadherin. The expression plasmid encoding FLAG.
 Fig. S1. Effect of p120-catenin overexpression on the interaction of SCUBE2 with E-cadherin. The expression plasmid encoding FLAG.SCUBE2, E-cadherin.Myc, or HA.p120-catenin was transfected in a combination
Fig. S1. Effect of p120-catenin overexpression on the interaction of SCUBE2 with E-cadherin. The expression plasmid encoding FLAG.SCUBE2, E-cadherin.Myc, or HA.p120-catenin was transfected in a combination
Supplementary Information for. Regulation of Rev1 by the Fanconi Anemia Core Complex
 Supplementary Information for Regulation of Rev1 by the Fanconi Anemia Core Complex Hyungjin Kim, Kailin Yang, Donniphat Dejsuphong, Alan D. D Andrea* *Corresponding Author: Alan D. D Andrea, M.D. Alan_dandrea@dfci.harvard.edu
Supplementary Information for Regulation of Rev1 by the Fanconi Anemia Core Complex Hyungjin Kim, Kailin Yang, Donniphat Dejsuphong, Alan D. D Andrea* *Corresponding Author: Alan D. D Andrea, M.D. Alan_dandrea@dfci.harvard.edu
Figure S1. USP-46 is expressed in several tissues including the nervous system
 Supplemental Figure legends Figure S1. USP-46 is expressed in several tissues including the nervous system Transgenic animals expressing a transcriptional reporter (P::GFP) were imaged using epifluorescence
Supplemental Figure legends Figure S1. USP-46 is expressed in several tissues including the nervous system Transgenic animals expressing a transcriptional reporter (P::GFP) were imaged using epifluorescence
Review of previous work: Overview of previous work on DNA methylation and chromatin dynamics
 CONTENTS Preface Acknowledgements Lists of Commonly Used Abbreviations Review of previous work: Overview of previous work on DNA methylation and chromatin dynamics Introduction Enzymes involved in DNA
CONTENTS Preface Acknowledgements Lists of Commonly Used Abbreviations Review of previous work: Overview of previous work on DNA methylation and chromatin dynamics Introduction Enzymes involved in DNA
Post-translational modification
 Protein expression Western blotting, is a widely used and accepted technique to detect levels of protein expression in a cell or tissue extract. This technique measures protein levels in a biological sample
Protein expression Western blotting, is a widely used and accepted technique to detect levels of protein expression in a cell or tissue extract. This technique measures protein levels in a biological sample
PLA2 domain in parvoviruses The enzymatic features of viral PLA2
 PLA2 domain in parvoviruses Sequence analysis of 21 Pavovirinae members revealed that they contain an 80 amino acid conserved region in their VP1up. 20 amino acids out of the 80 were fully conserved and
PLA2 domain in parvoviruses Sequence analysis of 21 Pavovirinae members revealed that they contain an 80 amino acid conserved region in their VP1up. 20 amino acids out of the 80 were fully conserved and
Kinetics Review. Tonight at 7 PM Phys 204 We will do two problems on the board (additional ones than in the problem sets)
 Quiz 1 Kinetics Review Tonight at 7 PM Phys 204 We will do two problems on the board (additional ones than in the problem sets) I will post the problems with solutions on Toolkit for those that can t make
Quiz 1 Kinetics Review Tonight at 7 PM Phys 204 We will do two problems on the board (additional ones than in the problem sets) I will post the problems with solutions on Toolkit for those that can t make
Supplementary Figures 1-12
 Supplementary Figures 1-12 Supplementary Figure 1. The specificity of anti-abi1 antibody. Total Proteins extracted from the wild type seedlings or abi1-3 null mutant seedlings were used for immunoblotting
Supplementary Figures 1-12 Supplementary Figure 1. The specificity of anti-abi1 antibody. Total Proteins extracted from the wild type seedlings or abi1-3 null mutant seedlings were used for immunoblotting
Regulation of enzyme synthesis
 Regulation of enzyme synthesis The lac operon is an example of an inducible operon - it is normally off, but when a molecule called an inducer is present, the operon turns on. The trp operon is an example
Regulation of enzyme synthesis The lac operon is an example of an inducible operon - it is normally off, but when a molecule called an inducer is present, the operon turns on. The trp operon is an example
X ray structure data Molecular dynamics simulation Sequence homology Educated guessing. Rational. Protein design
 Biocatalyst Design Rational X ray structure data Molecular dynamics simulation Sequence homology Educated guessing Protein design Random Do enough mistakes, so many time you are bound to get it right some
Biocatalyst Design Rational X ray structure data Molecular dynamics simulation Sequence homology Educated guessing Protein design Random Do enough mistakes, so many time you are bound to get it right some
12/11/12. Dr. Sanjeeva Srivastava. IIT Bombay 2. Interactomics Yeast Two-Hybrid Immunoprecipitation Protein microarrays. Proteomics Course NPTEL
 Dr. Sanjeeva Srivastava IIT Bombay Interactomics Yeast Two-Hybrid Immunoprecipitation Protein microarrays IIT Bombay 2 1 IIT Bombay Tremendous effect of interactions IIT Bombay 4 2 Complex formation Protein
Dr. Sanjeeva Srivastava IIT Bombay Interactomics Yeast Two-Hybrid Immunoprecipitation Protein microarrays IIT Bombay 2 1 IIT Bombay Tremendous effect of interactions IIT Bombay 4 2 Complex formation Protein
Motivation From Protein to Gene
 MOLECULAR BIOLOGY 2003-4 Topic B Recombinant DNA -principles and tools Construct a library - what for, how Major techniques +principles Bioinformatics - in brief Chapter 7 (MCB) 1 Motivation From Protein
MOLECULAR BIOLOGY 2003-4 Topic B Recombinant DNA -principles and tools Construct a library - what for, how Major techniques +principles Bioinformatics - in brief Chapter 7 (MCB) 1 Motivation From Protein
AP Biology Gene Expression/Biotechnology REVIEW
 AP Biology Gene Expression/Biotechnology REVIEW Multiple Choice Identify the choice that best completes the statement or answers the question. 1. Gene expression can be a. regulated before transcription.
AP Biology Gene Expression/Biotechnology REVIEW Multiple Choice Identify the choice that best completes the statement or answers the question. 1. Gene expression can be a. regulated before transcription.
Module 2 overview SPRING BREAK
 1 Module 2 overview lecture lab 1. Introduction to the module 1. Start-up protein eng. 2. Rational protein design 2. Site-directed mutagenesis 3. Fluorescence and sensors 3. DNA amplification 4. Protein
1 Module 2 overview lecture lab 1. Introduction to the module 1. Start-up protein eng. 2. Rational protein design 2. Site-directed mutagenesis 3. Fluorescence and sensors 3. DNA amplification 4. Protein
Protein isotopic enrichment for NMR studies
 Protein isotopic enrichment for NMR studies Protein NMR studies ARTGKYVDES sequence structure Structure of protein- ligand, protein-protein complexes Binding of molecules (perturbation mapping) Dynamic
Protein isotopic enrichment for NMR studies Protein NMR studies ARTGKYVDES sequence structure Structure of protein- ligand, protein-protein complexes Binding of molecules (perturbation mapping) Dynamic
DNA Cloning with Cloning Vectors
 Cloning Vectors A M I R A A. T. A L - H O S A R Y L E C T U R E R O F I N F E C T I O U S D I S E A S E S F A C U L T Y O F V E T. M E D I C I N E A S S I U T U N I V E R S I T Y - E G Y P T DNA Cloning
Cloning Vectors A M I R A A. T. A L - H O S A R Y L E C T U R E R O F I N F E C T I O U S D I S E A S E S F A C U L T Y O F V E T. M E D I C I N E A S S I U T U N I V E R S I T Y - E G Y P T DNA Cloning
Molecular characterization, detection & quantitation of biological products Purin Charoensuksai, PhD
 Molecular characterization, detection & quantitation of biological products Purin Charoensuksai, PhD Department of Biopharmacy, Faculty of Pharmacy, Silpakorn University Example of critical checkpoints
Molecular characterization, detection & quantitation of biological products Purin Charoensuksai, PhD Department of Biopharmacy, Faculty of Pharmacy, Silpakorn University Example of critical checkpoints
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION Contents: Supplementary Figure 1. Additional structural and binding data for designed tuim peptides. Supplementary Figure 2. Subcellular localization patterns of designed tuim
SUPPLEMENTARY INFORMATION Contents: Supplementary Figure 1. Additional structural and binding data for designed tuim peptides. Supplementary Figure 2. Subcellular localization patterns of designed tuim
Box 7905, Engineering Building I, North Carolina State University, Raleigh, NC 27695, Phone: Fax:
 Inefficient ribosomal skipping enables simultaneous secretion and display of proteins in Saccharomyces cerevisiae Carlos A. Cruz-Teran 1, Karthik Tiruthani 1, Adam Mischler, and Balaji M. Rao * Department
Inefficient ribosomal skipping enables simultaneous secretion and display of proteins in Saccharomyces cerevisiae Carlos A. Cruz-Teran 1, Karthik Tiruthani 1, Adam Mischler, and Balaji M. Rao * Department
T H E J O U R N A L O F C E L L B I O L O G Y
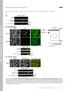 T H E J O U R N A L O F C E L L B I O L O G Y Supplemental material Nakajima and Tanoue, http://www.jcb.org/cgi/content/full/jcb.201104118/dc1 Figure S1. DLD-1 cells exhibit the characteristic morphology
T H E J O U R N A L O F C E L L B I O L O G Y Supplemental material Nakajima and Tanoue, http://www.jcb.org/cgi/content/full/jcb.201104118/dc1 Figure S1. DLD-1 cells exhibit the characteristic morphology
Fluorescent protein. Origin of fluorescent proteins. Structural and biophysical analysis of FPs. Application in molecular biology and biotechnology
 Origin of fluorescent proteins types of FPs mechanism of fluorescence Fluorescent protein Structural and biophysical analysis of FPs Application in molecular biology and biotechnology Variety of organisms
Origin of fluorescent proteins types of FPs mechanism of fluorescence Fluorescent protein Structural and biophysical analysis of FPs Application in molecular biology and biotechnology Variety of organisms
Lecture 5: 8/31. CHAPTER 5 Techniques in Protein Biochemistry
 Lecture 5: 8/31 CHAPTER 5 Techniques in Protein Biochemistry Chapter 5 Outline The proteome is the entire set of proteins expressed and modified by a cell under a particular set of biochemical conditions.
Lecture 5: 8/31 CHAPTER 5 Techniques in Protein Biochemistry Chapter 5 Outline The proteome is the entire set of proteins expressed and modified by a cell under a particular set of biochemical conditions.
INVESTIGATION OF THE BINDING SPECIFICITY OF IGF-IR USING MONOCLONAL ANTIBODIES
 INVESTIGATION OF THE BINDING SPECIFICITY OF IGF-IR USING MONOCLONAL ANTIBODIES By Mehrnaz Keyhanfar, Pharm.D. A thesis submitted to the University of Adelaide, South Australia in fulfilment of the requirements
INVESTIGATION OF THE BINDING SPECIFICITY OF IGF-IR USING MONOCLONAL ANTIBODIES By Mehrnaz Keyhanfar, Pharm.D. A thesis submitted to the University of Adelaide, South Australia in fulfilment of the requirements
BACTERIAL GENETICS. How does the DNA in the bacterial cell replicate
 BACTERIAL GENETICS Bacterial genetics is the study of gene structure and function in bacteria. Genetics itself is concerned with determining the number, location, and character of the genes of an organism.
BACTERIAL GENETICS Bacterial genetics is the study of gene structure and function in bacteria. Genetics itself is concerned with determining the number, location, and character of the genes of an organism.
Your Protein Manufacturer
 Expression of Toxic Proteins in E. coli Your Protein Manufacturer Contents 1. Definition of Toxic Protein 2. Mechanisms of Protein Toxicity 3. Percentage of Protein Toxicity 4. Phenotypes of Protein Toxicity
Expression of Toxic Proteins in E. coli Your Protein Manufacturer Contents 1. Definition of Toxic Protein 2. Mechanisms of Protein Toxicity 3. Percentage of Protein Toxicity 4. Phenotypes of Protein Toxicity
Really high sensitivity mass spectrometry and Discovery and analysis of protein complexes
 Really high sensitivity mass spectrometry and Discovery and analysis of protein complexes The PRIME lab and AMS Importance of protein complexes in biology Methods for isolation of protein complexes In
Really high sensitivity mass spectrometry and Discovery and analysis of protein complexes The PRIME lab and AMS Importance of protein complexes in biology Methods for isolation of protein complexes In
SPHERO TM Coated Particles
 SPHERO TM Coated Particles Manufactured by either passive adsorption or covalent coupling depending upon the intended application Stable for several years under proper storage condition Available in a
SPHERO TM Coated Particles Manufactured by either passive adsorption or covalent coupling depending upon the intended application Stable for several years under proper storage condition Available in a
Reading Lecture 3: 24-25, 45, Lecture 4: 66-71, Lecture 3. Vectors. Definition Properties Types. Transformation
 Lecture 3 Reading Lecture 3: 24-25, 45, 55-66 Lecture 4: 66-71, 75-79 Vectors Definition Properties Types Transformation 56 VECTORS- Definition Vectors are carriers of a DNA fragment of interest Insert
Lecture 3 Reading Lecture 3: 24-25, 45, 55-66 Lecture 4: 66-71, 75-79 Vectors Definition Properties Types Transformation 56 VECTORS- Definition Vectors are carriers of a DNA fragment of interest Insert
Topic 2: Proteins. 2-1 specific proteins can be purified from cell extracts. Molecular Biology and Public Health ( 分子生物学与公共卫生 )
 HAPTER20: Techniques of Molecular Biology Molecular Biology and Public Health ( 分子生物学与公共卫生 ) -------Protein manipulation ( 蛋白操作 ) Topic 2: Proteins 1. Protein purification ( 蛋白质纯化 ) 2. Affinity chromatography
HAPTER20: Techniques of Molecular Biology Molecular Biology and Public Health ( 分子生物学与公共卫生 ) -------Protein manipulation ( 蛋白操作 ) Topic 2: Proteins 1. Protein purification ( 蛋白质纯化 ) 2. Affinity chromatography
