Introduction to Protein Structure
|
|
|
- Avice Blankenship
- 6 years ago
- Views:
Transcription
1 Introduction to Protein Structure Second Edition Carl Branden Microbiology and Tumor Biology Center Karolinska Institute Stockholm Sweden John Tooze Imperial Cancer Research Fund Laboratories Lincolns Inn Fields London UK
2 Contents Part I Basic Structural Principles 1. The Building Blocks Proteins are polypeptide chains The genetic code specifies 20 different amino acid side chains Cysteines can form disulfide bridges Peptide units are building blocks of protein structures Glycine residues can adopt many different conformations Certain side-chain conformations are energetically favorable Many proteins contain intrinsic metal atoms 2. Motifs of Protein Structure The interior of proteins is hydrophobic The alpha (a) helix is an important element of secondary structure The a helix has a dipole moment Some amino acids are preferred in a helices Beta (P) sheets usually have their p strands either parallel or antiparallel Loop regions are at the surface of protein molecules Schematic pictures of proteins highlight secondary structure Topology diagrams are useful for classification of protein structures Secondary structure elements are connected to form simple motifs The hairpin p motif occurs frequently in protein structures The Greek key motif is found in antiparallel P sheets The P-a-P motif contains two parallel p strands Protein molecules are organized in a structural hierarchy Large polypeptide chains fold into several domains Domains are built from structural motifs 30 Simple motifs combine to form complex motifs 30 Protein structures can be divided into three main classes g 3. Alpha-Domain Structures 35 Coiled-coil a helices contain a repetitive 9 heptad amino acid sequence pattern 35 The four-helix bundle is a common domain 20 structure in a proteins 37 \\ Alpha-helical domains are sometimes large \2 and complex 39 The globin fold is present in myoglobin and hemoglobin Geometric considerations determine 14 a-helix packing 40 Ridges of one a helix fit into grooves of an 14 adjacent helix The globin fold has been preserved during evolution The hydrophobic interior is preserved 42 Helix movements accommodate interior 19 side-chain mutations 43 Sickle-cell hemoglobin confers resistance 21 to malaria Alpha/Beta Structures 47 Parallel p strands are arranged in barrels 24 or sheets 47 Alpha/beta barrels occur in many different 26 enzymes 48 Branched hydrophobic side chains dominate 27 the core of a/p barrels 49 Pyruvate kinase contains several domains, 27 one of which is an a/p barrel 51 Double barrels have occurred by gene 28 fusion 52 The active site is formed by loops at one 29 end of the a/p barrel 53 ix
3 Alpha/beta barrels provide examples of evolution of new enzyme activities 54 Leucine-rich motifs form an a/p-horseshoe 55 fold Alpha/beta twisted open-sheet structures contain a helices on both sides of the P sheet 56 Open p-sheet structures have a variety of topologies 57 The positions of active sites can be predicted in a/p structures 57 Tyrosyl-tRNA synthetase has two different domains (a/p + a) 59 Carboxypeptidase is an a/p protein with a mixed P sheet 60 Arabinose-binding protein has two similar a/p domains Beta Structures 67 Up-and-down barrels have a simple topology 68 The retinol-binding protein binds retinol inside an up-and-down p barrel 68 Amino acid sequence reflects P structure 69 The retinol-binding protein belongs to a superfamily of protein structures 70 Neuraminidase folds into up-and-down P sheets 70 Folding motifs form a propeller-like structure in neuraminidase 71 The active site is in the middle of one side of the propeller 72 Greek key motifs occur frequently in antiparallel p structures 72 The y-crystallin molecule has two domains 74 The domain structure has a simple topology 74 Two Greek key motifs form the domain 74 The two domains have identical topology 75 The two domains have similar structures 76 The Greek key motifs in y crystallin are evolutionary related 76 The Greek key motifs can form jelly roll barrels 77 The jelly roll motif is wrapped around a barrel 77 The jelly roll barrel is usually divided into two sheets 78 The functional hemagglutinin subunit has two polypeptide chains 79 The subunit structure is divided into a stem and a tip 79 The receptor binding site is formed by the jelly roll domain 80 Hemagglutinin acts as a membrane fusogen 80 The structure of hemagglutinin is affected by ph changes 81 Parallel p-helix domains have a novel fold Folding and Flexibility 89 Globular proteins are only marginally stable 90 Kinetic factors are important for folding 91 Molten globules are intermediates in folding 92 Burying hydrophobic side chains is a key event 93 Both single and multiple folding pathways have been observed Enzymes assist formation of proper disulfide bonds during folding Isomerization of proline residues can be a rate-limiting step in protein folding Proteins can fold or unfold inside chaperonins GroEL is a cylindrical structure with a central channel in which newly synthesized polypeptides bind GroES closes off one end of the GroEL cylinder The GroEL-GroES complex binds and releases newly synthesized polypeptides in an ATP-dependent cycle The folded state has a flexible structure Conformational changes in a protein kinase are important for cell cycle regulation Peptide binding to calmodulin induces a large interdomain movement Serpins inhibit serine proteinases with a spring-loaded safety catch mechanism Effector molecules switch allosteric proteins between R and T states X-ray structures explain the allosteric properties of phosphofructokinase 7. DNA Structures The DNA double helix is different in A- and B-DNA The DNA helix has major and minor grooves Z-DNA forms a zigzag pattern B-DNA is the preferred conformation in vivo Specific base sequences can be recognized in B-DNA Part 2 Structure, Function, and Engineering 8. DNA Recognition in Procaryotes by Helix-Turn-Helix Motifs A molecular mechanism for gene control Repressor and Cro proteins operate a procaryotic genetic switch region The x-ray structure of the complete lambda Cro protein is known The x-ray structure of the DNA-binding domain of the lambda repressor is known Both lambda Cro and repressor proteins have a specific DNA-binding motif Model building predicts Cro-DNA interactions Genetic studies agree with the structural model The x-ray structure of DNA complexes with 434 Cro and repressor revealed novel features of protein-dna interactions The structures of 434 Cro and the 434 repressor DNA-binding domain are very similar The proteins impose precise distortions on the B-DNA in the complexes Sequence-specific protein-dna interactions recognize operator regions
4 Protein-DNA backbone interactions determine DNA conformation Conformational changes of DNA are important for differential binding of repressor and Cro to different operator sites The essence of phage repressor and Cro DNA binding is regulated by allosteric control The trp repressor forms a helix-turn-helix motif A conformational change operates a functional switch Lac repressor binds to both the major and minor grooves inducing a sharp bend in the DNA CAP-induced DNA bending could activate transcription 9. DNA Recognition by Eucaryotic Transcription Factors Transcription is activated by protein-protein interactions The TATA box-binding protein is ubiquitous The three-dimensional structures of TBP-TATA box complexes are known A p sheet in TBP forms the DNA-binding site TBP binds in the minor groove and induces large structural changes in DNA The interaction area between TBP and the TATA box is mainly hydrophobic Functional implications of the distortion of DNA by TBP TFIIA and TFIIB bind to both TBP and DNA Homeodomain proteins are involved in the development of many eucaryotic organisms Monomers of homeodomain proteins bind to DNA through a helix-turn-helix motif In vivo specificity of homeodomain transcription factors depends on interactions with other proteins POU regions bind to DNA by two tandemly oriented helix-turn-helix motifs Much remains to be learnt about the function of homeodomains in vivo Understanding tumorigenic mutations The monomeric p53 polypeptide chain is divided in three domains The oligomerization domain forms tetramers The DNA-binding domain of p53 is an antiparallel p barrel Two loop regions and one a helix of p53 bind to DNA Tumorigenic mutations occur mainly in three regions involved in DNA binding s 10. Specific Transcription Factors Belong to a Few Families Several different groups of zinc-containing motifs have been observed The classic zinc fingers bind to DNA in tandem along the major groove The finger region of the classic zinc finger motif interacts with DNA 178 Two zinc-containing motifs in the glucocorticoid receptor form one DNA-binding domain 181 A dimer of the glucocorticoid receptor binds to DNA 183 An a helix in the first zinc motif provides the specific protein-dna interactions 184 Three residues in the recognition helix provide the sequence-specific interactions with DNA 184 The retinoid X receptor forms heterodimers that recognize tandem repeats with variable spacings 185 Yeast transcription factor GAL4 contains a binuclear zinc cluster in its DNA-binding domain 187 The zinc cluster regions of GAL4 bind at the two ends of the enhancer element 188 The linker region also contributes to DNA binding 189 DNA-binding site specificity among the C6-zinc cluster family of transcription factors is achieved by the linker regions 190 Families of zinc-containing transcription factors bind to DNA in several different ways 191 Leucine zippers provide dimerization interactions for some eucaryotic transcription factors 191 The GCN4 basic region leucine zipper binds DNA as a dimer of two uninterrupted a helices 193 GCN4 binds to DNA with both specific and nonspecific contacts 194 The HLH motif is involved in homodimer and heterodimer associations 196 The a-helical basic region of the b/hlh motif binds in the major groove of DNA 198 The b/hlh/zip family of transcription factors have both HLH and leucine zipper dimerization motifs 199 Max and MyoD recognize the DNA HLH consensus sequence by different specific protein-dna interactions An Example of Enzyme Catalysis: Serine Proteinases 205 Proteinases form four functional families 205 The catalytic properties of enzymes are reflected in K m and k ca t values 206 Enzymes decrease the activation energy of chemical reactions 206 Serine proteinases cleave peptide bonds by forming tetrahedral transition states 208 Four important structural features are required for the catalytic action of serine proteinases 209 Convergent evolution has produced two different serine proteinases with similar catalytic mechanisms 210 The chymotrypsin structure has two antiparallel P-barrel domains 210 XI
5 The active site is formed by two loop regions from each domain Did the chymotrypsin molecule evolve by gene duplication? Different side chains in the substrate specificity pocket confer preferential cleavage Engineered mutations in the substrate specificity pocket change the rate of catalysis The Asp 189-Lys mutation in trypsin causes unexpected changes in substrate specificity The structure of the serine proteinase subtilisin is of the a/p type The active sites of subtilisin and chymotrypsin are similar A structural anomaly in subtilisin has functional consequences Transition-state stabilization in subtilisin is dissected by protein engineering Catalysis occurs without a catalytic triad Substrate molecules provide catalytic groups in substrate-assisted catalysis 12. Membrane Proteins Membrane proteins are difficult to crystallize Novel crystallization methods are being developed Two-dimensional crystals of membrane proteins can be studied by electron microscopy Bacteriorhodopsin contains seven transmembrane a helices Bacteriorhodopsin is a light-driven proton pump Porins form transmembrane channels by P strands Porin channels are made by up and down P barrels Each porin molecule has three channels Ion channels combine ion selectivity with high levels of ion conductance The K + channel is a tetrameric molecule with one ion pore in the interface between the four subunits The ion pore has a narrow ion selectivity filter The bacterial photosynthetic reaction center is built up from four different polypeptide chains and many pigments The L, M, and H subunits have transmembrane a helices The photosynthetic pigments are bound to the L and M subunits Reaction centers convert light energy into electrical energy by electron flow through the membrane Antenna pigment proteins assemble into multimeric light-harvesting particles Chlorophyll molecules form circular rings in the light-harvesting complex LH2 The reaction center is surrounded by a ring of 16 antenna proteins of the light-harvesting complex LH Transmembrane a helices can be predicted from amino acid sequences 244 Hydrophobicity scales measure the degree of hydrophobicity of different amino acid side chains 245 Hydropathy plots identify transmembrane helices 245 Reaction center hydropathy plots agree with crystal structural data 246 Membrane lipids have no specific interaction with protein transmembrane a helices Signal Transduction 251 G proteins are molecular amplifiers 252 Ras proteins and the catalytic domain of G a have similar three-dimensional 254 structures G a is activated by conformational changes of three switch regions 257 GTPases hydrolyze GTP through nucleophilic attack by a water molecule 259 The Gp subunit has a seven-blade propeller fold, built up from seven WD repeat units 261 The GTPase domain of G a binds to Gp in the heterotrimeric G a p Y complex 263 Phosducin regulates light adaptation in retinal rods 265 Phosducin binding to Gp Y blocks binding of G a 265 The human growth hormone induces dimerization of its cognate receptor 267 Dimerization of the growth hormone receptor is a sequential process 268 The growth hormone also binds to the prolactin receptor 269 Tyrosine kinase receptors are important enzyme-linked receptors 270 Small protein modules form adaptors for a signaling network 272 SH2 domains bind to phosphotyrosinecontaining regions of target molecules 273 SH3 domains bind to proline-rich regions of target molecules 274 Src tyrosine kinases comprise SH2 and SH3 domains in addition to a tyrosine kinase 275 The two domains of the kinase in the inactive state are held in a closed conformation by assembly of the regulatory domains Fibrous Proteins 283 Collagen is a superhelix formed by three parallel, very extended left-handed helices 284 Coiled coils are frequently used to form oligomers of fibrous and globular proteins 286 Amyloid fibrils are suggested to be built up from continuous P sheet helices 288 Spider silk is nature's high-performance fiber 289 Muscle fibers contain myosin and actin which slide against each other during muscle contraction 290 XII
6 Myosin heads form cross-bridges between the actin and myosin filaments 291 Time-resolved x-ray diffraction of frog muscle confirmed movement of the cross-bridges 292 Structures of actin and myosin have been determined 293 The structure of myosin supports the swinging cross-bridge hypothesis 295 The role of ATP in muscular contraction has parallels to the role of GTP in G-protein activation Recognition of Foreign Molecules by the Immune System 299 The polypeptide chains of antibodies are divided into domains 300 Antibody diversity is generated by several different mechanisms 302 All immunoglobulin domains have similar three-dimensional structures 303 The immunoglobulin fold is best described as two antiparallel p sheets packed tightly against each other 304 The hypervariable regions are clustered in loop regions at one end of the variable domain 305 The antigen-binding site is formed by close association of the hypervariable regions from both heavy and light chains 306 The antigen-binding site binds haptens in crevices and protein antigens on large flat surfaces 308 The CDR loops assume only a limited range of conformations, except for the heavy chain CDR3 311 An IgG molecule has several degrees of conformational flexibility 312 Structures of MHC molecules have provided insights into the molecular mechanisms of T-cell activation 312 MHC molecules are composed of antigenbinding and immunoglobulin-like domains 313 Recognition of antigen is different in MHC molecules compared with immunoglobulins 314 Peptides are bound differently by class I and class II MHC molecules 315 T-cell receptors have variable and constant immunoglobulin domains and hypervariable regions 316 MHC-peptide complexes are the ligands for T-cell receptors 318 Many cell-surface receptors contain immunoglobulin-like domains The Structure of Spherical Viruses 325 The protein shells of spherical viruses have icosahedral symmetry 327 The icosahedron has high symmetry 327 The simplest virus has a shell of 60 protein subunits 328 Complex spherical viruses have more than one polypeptide chain in the asymmetric unit 329 Structural versatility gives quasi-equivalent packing in T = 3 plant viruses 331 The protein subunits recognize specific parts of the RNA inside the shell 332 The protein capsid of picornaviruses contains four polypeptide chains 333 There are four different structural proteins in picornaviruses 334 The arrangement of subunits in the shell of picornaviruses is similar to that of T = 3 plant viruses 334 The coat proteins of many different spherical plant and animal viruses have similar jelly roll barrel structures, indicating an evolutionary relationship 335 Drugs against the common cold may be designed from the structure of rhinovirus 337 Bacteriophage MS2 has a different subunit structure 339 A dimer of MS2 subunits recognizes an RNA packaging signal 339 The core protein of alphavirus has a chymotrypsin-like fold 340 SV40 and polyomavirus shells are constructed from pentamers of the major coat protein with nonequivalent packing but largely equivalent interactions Prediction, Engineering, and Design of Protein Structures 347 Homologous proteins have similar structure and function 348 Homologous proteins have conserved structural cores and variable loop regions 349 Knowledge of secondary structure is necessary for prediction of tertiary structure 350 Prediction methods for secondary structure benefit from multiple alignment of homologous proteins 351 Many different amino acid sequences give similar three-dimensional structures 352 Prediction of protein structure from sequence is an unsolved problem 352 Threading methods can assign amino acid sequences to known three-dimensional folds 353 Proteins can be made more stable by engineering 354 Disulfide bridges increase protein stability 355 Glycine and proline have opposite effects on stability 356 Stabilizing the dipoles of a helices increases stability 357 Mutants that fill cavities in hydrophobic cores do not stabilize T4 lysozyme 358 Proteins can be engineered by combinatorial methods 358 Phage display links the protein library to DNA 359 Affinity and specificity of proteinase inhibitors can be optimized by phage display 361 xiii
7 Structural scaffolds can be reduced in size while function is retained Phage display of random peptide libraries identified agonists of erythropoetin receptor DNA shuffling allows accelerated evolution of genes Protein structures can be designed from first principles A P structure has been converted to an a structure by changing only half of the sequence 18. Determination of Protein Structures Several different techniques are used to study the structure of protein molecules Protein crystals are difficult to grow X-ray sources are either monochromatic or polychromatic X-ray data are recorded either on image plates or by electronic detectors The rules for diffraction are given by Bragg's law Phase determination is the major crystallographic problem Phase information can also be obtained by Multiwavelength Anomalous Diffraction experiments Building a model involves subjective 363 interpretation of the data 381 Errors in the initial model are removed 364 by refinement 383 Recent technological advances have greatly 365 influenced protein crystallography 383 X-ray diffraction can be used to study the 367 structure of fibers as well as crystals 384 The structure of biopolymers can be studied 368 using fiber diffraction NMR methods use the magnetic properties of 371 atomic nuclei 387 Two-dimensional NMR spectra of proteins are 373 interpreted by the method of sequential assignment Distance constraints are used to derive possible 374 structures of a protein molecule 390 Biochemical studies and molecular 376 structure give complementary functional information Protein Structure on the World Wide Web xiv
Introduction to Protein Structure
 Introduction to Protein Structure Carl Branden John Tooze Protein Structure Protein molecules are organized in a structural Part 1 Basic Structural Principles 1 hierarchy 2 6 1. The Building Blocks 3
Introduction to Protein Structure Carl Branden John Tooze Protein Structure Protein molecules are organized in a structural Part 1 Basic Structural Principles 1 hierarchy 2 6 1. The Building Blocks 3
Chapter 8 DNA Recognition in Prokaryotes by Helix-Turn-Helix Motifs
 Chapter 8 DNA Recognition in Prokaryotes by Helix-Turn-Helix Motifs 1. Helix-turn-helix proteins 2. Zinc finger proteins 3. Leucine zipper proteins 4. Beta-scaffold factors 5. Others λ-repressor AND CRO
Chapter 8 DNA Recognition in Prokaryotes by Helix-Turn-Helix Motifs 1. Helix-turn-helix proteins 2. Zinc finger proteins 3. Leucine zipper proteins 4. Beta-scaffold factors 5. Others λ-repressor AND CRO
STRUCTURAL BIOLOGY. α/β structures Closed barrels Open twisted sheets Horseshoe folds
 STRUCTURAL BIOLOGY α/β structures Closed barrels Open twisted sheets Horseshoe folds The α/β domains Most frequent domain structures are α/β domains: A central parallel or mixed β sheet Surrounded by α
STRUCTURAL BIOLOGY α/β structures Closed barrels Open twisted sheets Horseshoe folds The α/β domains Most frequent domain structures are α/β domains: A central parallel or mixed β sheet Surrounded by α
Chapter 17. Prediction, Engineering, and Design of Protein Structures
 Chapter 17. Prediction, Engineering, and Design of Protein Structures Homologous proteins have similar structure and function Branden & Tooze (1998), Introduction to protein structure, 2 nd ed., p.349.
Chapter 17. Prediction, Engineering, and Design of Protein Structures Homologous proteins have similar structure and function Branden & Tooze (1998), Introduction to protein structure, 2 nd ed., p.349.
RNA does not adopt the classic B-DNA helix conformation when it forms a self-complementary double helix
 Reason: RNA has ribose sugar ring, with a hydroxyl group (OH) If RNA in B-from conformation there would be unfavorable steric contact between the hydroxyl group, base, and phosphate backbone. RNA structure
Reason: RNA has ribose sugar ring, with a hydroxyl group (OH) If RNA in B-from conformation there would be unfavorable steric contact between the hydroxyl group, base, and phosphate backbone. RNA structure
Chapter 14 Regulation of Transcription
 Chapter 14 Regulation of Transcription Cis-acting sequences Distance-independent cis-acting elements Dissecting regulatory elements Transcription factors Overview transcriptional regulation Transcription
Chapter 14 Regulation of Transcription Cis-acting sequences Distance-independent cis-acting elements Dissecting regulatory elements Transcription factors Overview transcriptional regulation Transcription
BIRKBECK COLLEGE (University of London)
 BIRKBECK COLLEGE (University of London) SCHOOL OF BIOLOGICAL SCIENCES M.Sc. EXAMINATION FOR INTERNAL STUDENTS ON: Postgraduate Certificate in Principles of Protein Structure MSc Structural Molecular Biology
BIRKBECK COLLEGE (University of London) SCHOOL OF BIOLOGICAL SCIENCES M.Sc. EXAMINATION FOR INTERNAL STUDENTS ON: Postgraduate Certificate in Principles of Protein Structure MSc Structural Molecular Biology
Transcription factors
 Atlas of Genetics and Cytogenetics in Oncology and Haematology Transcription factors I Introduction * II Initiation of transcription III Transcription factors family pdf version I Introduction III.1 Helix-Turn-Helix
Atlas of Genetics and Cytogenetics in Oncology and Haematology Transcription factors I Introduction * II Initiation of transcription III Transcription factors family pdf version I Introduction III.1 Helix-Turn-Helix
Protein Structure and Function: From Sequence to Consequence 1. From Sequence to Structure
 Protein Structure and Function: From Sequence to Consequence Gregory A. Petsko and Dagmar Ringe, Brandeis University Published by New Science Press Ltd and distributed in the United States and Canada by
Protein Structure and Function: From Sequence to Consequence Gregory A. Petsko and Dagmar Ringe, Brandeis University Published by New Science Press Ltd and distributed in the United States and Canada by
Protein Structure/Function Relationships
 Protein Structure/Function Relationships W. M. Grogan, Ph.D. OBJECTIVES 1. Describe and cite examples of fibrous and globular proteins. 2. Describe typical tertiary structural motifs found in proteins.
Protein Structure/Function Relationships W. M. Grogan, Ph.D. OBJECTIVES 1. Describe and cite examples of fibrous and globular proteins. 2. Describe typical tertiary structural motifs found in proteins.
Protein Structure. Protein Structure Tertiary & Quaternary
 Lecture 4 Protein Structure Protein Structure Tertiary & Quaternary Dr. Sameh Sarray Hlaoui Primary structure: The linear sequence of amino acids held together by peptide bonds. Secondary structure: The
Lecture 4 Protein Structure Protein Structure Tertiary & Quaternary Dr. Sameh Sarray Hlaoui Primary structure: The linear sequence of amino acids held together by peptide bonds. Secondary structure: The
DNA Binding Domains: Structural Motifs. Effector Domain. Zinc Fingers. Zinc Fingers, continued. Zif268
 DNA Binding Domains: Structural Motifs Studies of known transcription factors have found several motifs of protein design to allow sequence-specific binding of DNA. We will cover only three of these motifs:
DNA Binding Domains: Structural Motifs Studies of known transcription factors have found several motifs of protein design to allow sequence-specific binding of DNA. We will cover only three of these motifs:
Examination I PHRM 836 Biochemistry for Pharmaceutical Sciences II September 29, 2015
 Examination I PHRM 836 Biochemistry for Pharmaceutical Sciences II September 29, 2015 PHRM 836 Exam I - 1 Name: Instructions 1. Check your exam to make certain that it has 9 pages including this cover
Examination I PHRM 836 Biochemistry for Pharmaceutical Sciences II September 29, 2015 PHRM 836 Exam I - 1 Name: Instructions 1. Check your exam to make certain that it has 9 pages including this cover
Introduction to Proteins
 Introduction to Proteins Lecture 4 Module I: Molecular Structure & Metabolism Molecular Cell Biology Core Course (GSND5200) Matthew Neiditch - Room E450U ICPH matthew.neiditch@umdnj.edu What is a protein?
Introduction to Proteins Lecture 4 Module I: Molecular Structure & Metabolism Molecular Cell Biology Core Course (GSND5200) Matthew Neiditch - Room E450U ICPH matthew.neiditch@umdnj.edu What is a protein?
Translation. Protein Synthesis
 Protein Structure Translation Protein Synthesis Size and Shape Comparison of Proteins Levels of Protein Structure 1 o 2 o 3 o 4 o Amino Acids Peptide Bonds Proteins are formed by creating peptide bonds
Protein Structure Translation Protein Synthesis Size and Shape Comparison of Proteins Levels of Protein Structure 1 o 2 o 3 o 4 o Amino Acids Peptide Bonds Proteins are formed by creating peptide bonds
Protein Folding Problem I400: Introduction to Bioinformatics
 Protein Folding Problem I400: Introduction to Bioinformatics November 29, 2004 Protein biomolecule, macromolecule more than 50% of the dry weight of cells is proteins polymer of amino acids connected into
Protein Folding Problem I400: Introduction to Bioinformatics November 29, 2004 Protein biomolecule, macromolecule more than 50% of the dry weight of cells is proteins polymer of amino acids connected into
Structure/function relationship in DNA-binding proteins
 PHRM 836 September 22, 2015 Structure/function relationship in DNA-binding proteins Devlin Chapter 8.8-9 u General description of transcription factors (TFs) u Sequence-specific interactions between DNA
PHRM 836 September 22, 2015 Structure/function relationship in DNA-binding proteins Devlin Chapter 8.8-9 u General description of transcription factors (TFs) u Sequence-specific interactions between DNA
Bi 8 Lecture 7. Ellen Rothenberg 26 January Reading: Ch. 3, pp ; panel 3-1
 Bi 8 Lecture 7 PROTEIN STRUCTURE, Functional analysis, and evolution Ellen Rothenberg 26 January 2016 Reading: Ch. 3, pp. 109-134; panel 3-1 (end with free amine) aromatic, hydrophobic small, hydrophilic
Bi 8 Lecture 7 PROTEIN STRUCTURE, Functional analysis, and evolution Ellen Rothenberg 26 January 2016 Reading: Ch. 3, pp. 109-134; panel 3-1 (end with free amine) aromatic, hydrophobic small, hydrophilic
Examination I PHRM 836 Biochemistry for Pharmaceutical Sciences II September 29, 2015
 Examination I PHRM 836 Biochemistry for Pharmaceutical Sciences II September 29, 2015 PHRM 836 Exam I - 1 Name: Instructions 1. Check your exam to make certain that it has 9 pages including this cover
Examination I PHRM 836 Biochemistry for Pharmaceutical Sciences II September 29, 2015 PHRM 836 Exam I - 1 Name: Instructions 1. Check your exam to make certain that it has 9 pages including this cover
Chapter 25: Regulating Eukaryotic Transcription The Ligand Responsive Activators
 Chapter 25: Regulating Eukaryotic Transcription The Ligand Responsive Activators At least 5 potential gene expression control points Superfamily of Gene Regulators Activation of gene structure Initiation
Chapter 25: Regulating Eukaryotic Transcription The Ligand Responsive Activators At least 5 potential gene expression control points Superfamily of Gene Regulators Activation of gene structure Initiation
Proteins Higher Order Structures
 Proteins Higher Order Structures Dr. Mohammad Alsenaidy Department of Pharmaceutics College of Pharmacy King Saud University Office: AA 101 msenaidy@ksu.edu.sa Previously on PHT 426!! Protein Structures
Proteins Higher Order Structures Dr. Mohammad Alsenaidy Department of Pharmaceutics College of Pharmacy King Saud University Office: AA 101 msenaidy@ksu.edu.sa Previously on PHT 426!! Protein Structures
BETA STRAND Prof. Alejandro Hochkoeppler Department of Pharmaceutical Sciences and Biotechnology University of Bologna
 Prof. Alejandro Hochkoeppler Department of Pharmaceutical Sciences and Biotechnology University of Bologna E-mail: a.hochkoeppler@unibo.it C-ter NH and CO groups: right, left, right (plane of the slide)
Prof. Alejandro Hochkoeppler Department of Pharmaceutical Sciences and Biotechnology University of Bologna E-mail: a.hochkoeppler@unibo.it C-ter NH and CO groups: right, left, right (plane of the slide)
Proteins the primary biological macromolecules of living organisms
 Proteins the primary biological macromolecules of living organisms Protein structure and folding Primary Secondary Tertiary Quaternary structure of proteins Structure of Proteins Protein molecules adopt
Proteins the primary biological macromolecules of living organisms Protein structure and folding Primary Secondary Tertiary Quaternary structure of proteins Structure of Proteins Protein molecules adopt
Nucleic Acids, Proteins, and Enzymes
 3 Nucleic Acids, Proteins, and Enzymes Chapter 3 Nucleic Acids, Proteins, and Enzymes Key Concepts 3.1 Nucleic Acids Are Informational Macromolecules 3.2 Proteins Are Polymers with Important Structural
3 Nucleic Acids, Proteins, and Enzymes Chapter 3 Nucleic Acids, Proteins, and Enzymes Key Concepts 3.1 Nucleic Acids Are Informational Macromolecules 3.2 Proteins Are Polymers with Important Structural
Enhancers. Activators and repressors of transcription
 Enhancers Can be >50 kb away from the gene they regulate. Can be upstream from a promoter, downstream from a promoter, within an intron, or even downstream of the final exon of a gene. Are often cell type
Enhancers Can be >50 kb away from the gene they regulate. Can be upstream from a promoter, downstream from a promoter, within an intron, or even downstream of the final exon of a gene. Are often cell type
CSE : Computational Issues in Molecular Biology. Lecture 19. Spring 2004
 CSE 397-497: Computational Issues in Molecular Biology Lecture 19 Spring 2004-1- Protein structure Primary structure of protein is determined by number and order of amino acids within polypeptide chain.
CSE 397-497: Computational Issues in Molecular Biology Lecture 19 Spring 2004-1- Protein structure Primary structure of protein is determined by number and order of amino acids within polypeptide chain.
PROTEINS & NUCLEIC ACIDS
 Chapter 3 Part 2 The Molecules of Cells PROTEINS & NUCLEIC ACIDS Lecture by Dr. Fernando Prince 3.11 Nucleic Acids are the blueprints of life Proteins are the machines of life We have already learned that
Chapter 3 Part 2 The Molecules of Cells PROTEINS & NUCLEIC ACIDS Lecture by Dr. Fernando Prince 3.11 Nucleic Acids are the blueprints of life Proteins are the machines of life We have already learned that
CS273: Algorithms for Structure Handout # 5 and Motion in Biology Stanford University Tuesday, 13 April 2004
 CS273: Algorithms for Structure Handout # 5 and Motion in Biology Stanford University Tuesday, 13 April 2004 Lecture #5: 13 April 2004 Topics: Sequence motif identification Scribe: Samantha Chui 1 Introduction
CS273: Algorithms for Structure Handout # 5 and Motion in Biology Stanford University Tuesday, 13 April 2004 Lecture #5: 13 April 2004 Topics: Sequence motif identification Scribe: Samantha Chui 1 Introduction
BIOLOGY 200 Molecular Biology Students registered for the 9:30AM lecture should NOT attend the 4:30PM lecture.
 BIOLOGY 200 Molecular Biology Students registered for the 9:30AM lecture should NOT attend the 4:30PM lecture. Midterm date change! The midterm will be held on October 19th (likely 6-8PM). Contact Kathy
BIOLOGY 200 Molecular Biology Students registered for the 9:30AM lecture should NOT attend the 4:30PM lecture. Midterm date change! The midterm will be held on October 19th (likely 6-8PM). Contact Kathy
Regulation of gene expression. (Lehninger pg )
 Regulation of gene expression (Lehninger pg. 1072-1085) Today s lecture Gene expression Constitutive, inducible, repressible genes Specificity factors, activators, repressors Negative and positive gene
Regulation of gene expression (Lehninger pg. 1072-1085) Today s lecture Gene expression Constitutive, inducible, repressible genes Specificity factors, activators, repressors Negative and positive gene
DNA & DNA : Protein Interactions BIBC 100
 DNA & DNA : Protein Interactions BIBC 100 Sequence = Information Alphabet = language L,I,F,E LIFE DNA = DNA code A, T, C, G CAC=Histidine CAG=Glutamine GGG=Glycine Protein = Protein code 20 a.a. LIVE EVIL
DNA & DNA : Protein Interactions BIBC 100 Sequence = Information Alphabet = language L,I,F,E LIFE DNA = DNA code A, T, C, G CAC=Histidine CAG=Glutamine GGG=Glycine Protein = Protein code 20 a.a. LIVE EVIL
Specificity: Induced Fit
 Specificity: Induced Fit Conformational changes may occur upon ligand binding (Daniel Koshland in 1958) This adaptation is called the induced fit Induced fit allows for tighter binding of the ligand Induced
Specificity: Induced Fit Conformational changes may occur upon ligand binding (Daniel Koshland in 1958) This adaptation is called the induced fit Induced fit allows for tighter binding of the ligand Induced
(Refer Slide Time: 00:15)
 (Refer Slide Time: 00:15) Proteins and Gel-Based Proteomics Professor Sanjeeva Srivastava Department of Biosciences and Bioengineering Indian Institute of Technology, Bombay Mod 02 Lecture Number 3 Let
(Refer Slide Time: 00:15) Proteins and Gel-Based Proteomics Professor Sanjeeva Srivastava Department of Biosciences and Bioengineering Indian Institute of Technology, Bombay Mod 02 Lecture Number 3 Let
Chem 465 Biochemistry II
 Chem 465 Biochemistry II Name: 2 points Multiple choice (4 points apiece): 1. Which of the following is not true of trna molecules? A) The 3'-terminal sequence is -CCA. B) Their anticodons are complementary
Chem 465 Biochemistry II Name: 2 points Multiple choice (4 points apiece): 1. Which of the following is not true of trna molecules? A) The 3'-terminal sequence is -CCA. B) Their anticodons are complementary
Antibodies (Recommended reading: Abbas et al., 4th edition, Chapter 3; Chapter 4; Janeway et al., 5th edition, Chapter 3)
 HST 175 Antibodies (Recommended reading: Abbas et al., 4th edition, Chapter 3; Chapter 4; Janeway et al., 5th edition, Chapter 3) Antibodies protect us from a vast variety of pathogens. Indeed the antibody
HST 175 Antibodies (Recommended reading: Abbas et al., 4th edition, Chapter 3; Chapter 4; Janeway et al., 5th edition, Chapter 3) Antibodies protect us from a vast variety of pathogens. Indeed the antibody
Chapter 4. Antigen Recognition by B-cell and T-cell Receptors
 Chapter 4 Antigen Recognition by B-cell and T-cell Receptors Antigen recognition by BCR and TCR B cells 2 separate functions of immunoglobulin (Ig) bind pathogen & induce immune responses recruit cells
Chapter 4 Antigen Recognition by B-cell and T-cell Receptors Antigen recognition by BCR and TCR B cells 2 separate functions of immunoglobulin (Ig) bind pathogen & induce immune responses recruit cells
Complex Transcription Machinery
 Complex Transcription Machinery Subunits of the Basal Txn Apparatus Stepwise Assembly of the Pre-initiation Complex Interplay of Activators, Co-regulators and RNA Polymerase at the Promoter Divide and
Complex Transcription Machinery Subunits of the Basal Txn Apparatus Stepwise Assembly of the Pre-initiation Complex Interplay of Activators, Co-regulators and RNA Polymerase at the Promoter Divide and
Gene expression DNA RNA. Protein DNA. Replication. Initiation Elongation Processing Export. DNA RNA Protein. Transcription. Degradation.
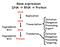 Gene expression DNA RNA Protein DNA DNA Degradation RNA Degradation Protein Replication Transcription Translation Initiation Elongation Processing Export Initiation Elongation Processing Targeting Chapter
Gene expression DNA RNA Protein DNA DNA Degradation RNA Degradation Protein Replication Transcription Translation Initiation Elongation Processing Export Initiation Elongation Processing Targeting Chapter
AMINO ACIDS, POLYPEPTIDES AND PROTEINS
 AMINO ACIDS, POLYPEPTIDES AND PROTEINS As mentioned above, some of the simplest and most stable forms of nucleic acid to be produced were the transfer RNA molecules - probably at random at first but later
AMINO ACIDS, POLYPEPTIDES AND PROTEINS As mentioned above, some of the simplest and most stable forms of nucleic acid to be produced were the transfer RNA molecules - probably at random at first but later
AP Biology Book Notes Chapter 3 v Nucleic acids Ø Polymers specialized for the storage transmission and use of genetic information Ø Two types DNA
 AP Biology Book Notes Chapter 3 v Nucleic acids Ø Polymers specialized for the storage transmission and use of genetic information Ø Two types DNA Encodes hereditary information Used to specify the amino
AP Biology Book Notes Chapter 3 v Nucleic acids Ø Polymers specialized for the storage transmission and use of genetic information Ø Two types DNA Encodes hereditary information Used to specify the amino
Protein Folding BIBC 100
 Protein Folding BIBC 100 The Folding Problem How Proteins Fold? Consider a protein with 100 a.a. 10 100 possible conformations (avg. of 10 conformations/a.a.) If it converts from one conformation to another
Protein Folding BIBC 100 The Folding Problem How Proteins Fold? Consider a protein with 100 a.a. 10 100 possible conformations (avg. of 10 conformations/a.a.) If it converts from one conformation to another
Ch Biophysical Chemistry
 Ch 247.53. Biophysical Chemistry Nina Rosario L. Rojas 2012-2013 sem 1 Review of Protein Structure Why structure? Primary, secondary, tertiary structure Disulfide bonds scheme 2 STRUCTURE- REGULAR STRUCTURE
Ch 247.53. Biophysical Chemistry Nina Rosario L. Rojas 2012-2013 sem 1 Review of Protein Structure Why structure? Primary, secondary, tertiary structure Disulfide bonds scheme 2 STRUCTURE- REGULAR STRUCTURE
BIO 311C Spring Lecture 16 Monday 1 March
 BIO 311C Spring 2010 Lecture 16 Monday 1 March Review Primary Structure of a portion of a polypeptide chain backbone of Polypeptide chain R-groups of amino acids Native conformation of a dimeric protein,
BIO 311C Spring 2010 Lecture 16 Monday 1 March Review Primary Structure of a portion of a polypeptide chain backbone of Polypeptide chain R-groups of amino acids Native conformation of a dimeric protein,
BASIC MOLECULAR GENETIC MECHANISMS Introduction:
 BASIC MOLECULAR GENETIC MECHANISMS Introduction: nucleic acids. (1) contain the information for determining the amino acid sequence & the structure and function of proteins (1) part of the cellular structures:
BASIC MOLECULAR GENETIC MECHANISMS Introduction: nucleic acids. (1) contain the information for determining the amino acid sequence & the structure and function of proteins (1) part of the cellular structures:
Plant Molecular and Cellular Biology Lecture 9: Nuclear Genome Organization: Chromosome Structure, Chromatin, DNA Packaging, Mitosis Gary Peter
 Plant Molecular and Cellular Biology Lecture 9: Nuclear Genome Organization: Chromosome Structure, Chromatin, DNA Packaging, Mitosis Gary Peter 9/16/2008 1 Learning Objectives 1. List and explain how DNA
Plant Molecular and Cellular Biology Lecture 9: Nuclear Genome Organization: Chromosome Structure, Chromatin, DNA Packaging, Mitosis Gary Peter 9/16/2008 1 Learning Objectives 1. List and explain how DNA
Comparative Modeling Part 1. Jaroslaw Pillardy Computational Biology Service Unit Cornell Theory Center
 Comparative Modeling Part 1 Jaroslaw Pillardy Computational Biology Service Unit Cornell Theory Center Function is the most important feature of a protein Function is related to structure Structure is
Comparative Modeling Part 1 Jaroslaw Pillardy Computational Biology Service Unit Cornell Theory Center Function is the most important feature of a protein Function is related to structure Structure is
Gene regulation V Biochemistry 302. March 6, 2006
 Gene regulation V Biochemistry 302 March 6, 2006 Common structural motifs associated with transcriptional regulatory proteins Helix-turn-helix Prokaryotic repressors and activators Eukaryotic homeodomain
Gene regulation V Biochemistry 302 March 6, 2006 Common structural motifs associated with transcriptional regulatory proteins Helix-turn-helix Prokaryotic repressors and activators Eukaryotic homeodomain
Molecular Biology (BIOL 4320) Exam #1 March 12, 2002
 Molecular Biology (BIOL 4320) Exam #1 March 12, 2002 Name KEY SS# This exam is worth a total of 100 points. The number of points each question is worth is shown in parentheses after the question number.
Molecular Biology (BIOL 4320) Exam #1 March 12, 2002 Name KEY SS# This exam is worth a total of 100 points. The number of points each question is worth is shown in parentheses after the question number.
Biochem Fall Sample Exam I Protein Structure. Vasopressin: CYFQNCPRG Oxytocin: CYIQNCPLG
 Biochem Fall 2011 1. Primary Structure and amino acid chemistry Sample Exam I Protein Structure The peptide hormones vasopressin (ADH) and oxytocin each contain only nine amino acids. Vasopressin is an
Biochem Fall 2011 1. Primary Structure and amino acid chemistry Sample Exam I Protein Structure The peptide hormones vasopressin (ADH) and oxytocin each contain only nine amino acids. Vasopressin is an
Protein Folding in vivo. Biochemistry 530 David Baker
 Protein Folding in vivo Biochemistry 530 David Baker I. Protein folding in vivo: Molecular Chaperones II. Protein unfolding in vivo: ATP driven unfolding in mitochondrial import and the proteasome. III.
Protein Folding in vivo Biochemistry 530 David Baker I. Protein folding in vivo: Molecular Chaperones II. Protein unfolding in vivo: ATP driven unfolding in mitochondrial import and the proteasome. III.
Molecular recognition
 Lecture 9 Molecular recognition Antoine van Oijen BCMP201 Spring 2008 Structural principles of binding 4 fundamental functions of proteins: 1) Binding 2) Catalysis 3) Switching 4) Structural All involve
Lecture 9 Molecular recognition Antoine van Oijen BCMP201 Spring 2008 Structural principles of binding 4 fundamental functions of proteins: 1) Binding 2) Catalysis 3) Switching 4) Structural All involve
Chapter 3 Nucleic Acids, Proteins, and Enzymes
 3 Nucleic Acids, Proteins, and Enzymes Chapter 3 Nucleic Acids, Proteins, and Enzymes Key Concepts 3.1 Nucleic Acids Are Informational Macromolecules 3.2 Proteins Are Polymers with Important Structural
3 Nucleic Acids, Proteins, and Enzymes Chapter 3 Nucleic Acids, Proteins, and Enzymes Key Concepts 3.1 Nucleic Acids Are Informational Macromolecules 3.2 Proteins Are Polymers with Important Structural
Programme Good morning and summary of last week Levels of Protein Structure - I Levels of Protein Structure - II
 Programme 8.00-8.10 Good morning and summary of last week 8.10-8.30 Levels of Protein Structure - I 8.30-9.00 Levels of Protein Structure - II 9.00-9.15 Break 9.15-11.15 Exercise: Building a protein model
Programme 8.00-8.10 Good morning and summary of last week 8.10-8.30 Levels of Protein Structure - I 8.30-9.00 Levels of Protein Structure - II 9.00-9.15 Break 9.15-11.15 Exercise: Building a protein model
BC 367, Exam 2 November 13, Part I. Multiple Choice (3 pts each)- Please circle the single best answer.
 Name BC 367, Exam 2 November 13, 2008 Part I. Multiple Choice (3 pts each)- Please circle the single best answer. 1. The enzyme pyruvate dehydrogenase catalyzes the following reaction. What kind of enzyme
Name BC 367, Exam 2 November 13, 2008 Part I. Multiple Choice (3 pts each)- Please circle the single best answer. 1. The enzyme pyruvate dehydrogenase catalyzes the following reaction. What kind of enzyme
Protein Structure Function
 Protein Engineering 15hp Protein Structure Function christofer@xray.bmc.uu.se The Protein Domain 2 What is a domain? An evolutionary conserved, independently folding unit within a protein. A domain often,
Protein Engineering 15hp Protein Structure Function christofer@xray.bmc.uu.se The Protein Domain 2 What is a domain? An evolutionary conserved, independently folding unit within a protein. A domain often,
Bacteriophages get a foothold on their prey
 Bacteriophages get a foothold on their prey Long and thin, the receptor-binding needle of bacteriophage T4 Bacterial viruses, bacteriophages or phages, have served as a tool to decipher principles of molecular
Bacteriophages get a foothold on their prey Long and thin, the receptor-binding needle of bacteriophage T4 Bacterial viruses, bacteriophages or phages, have served as a tool to decipher principles of molecular
Solutions to Problem Set 1
 MIT Department of Biology 7.014 Introductory Biology, Spring 004 Question 1 Solutions to 7.014 Problem Set 1 a) Describe the conditions of the atmosphere on prebiotic earth and how these conditions differ
MIT Department of Biology 7.014 Introductory Biology, Spring 004 Question 1 Solutions to 7.014 Problem Set 1 a) Describe the conditions of the atmosphere on prebiotic earth and how these conditions differ
3.1 Storing Information. Polypeptides. Protein Structure. Helical Secondary Structure. Types of Protein Structure. Molecular Biology Fourth Edition
 Lecture PowerPoint to accompany Molecular Biology Fourth Edition Robert F. Weaver Chapter 3 An Introduction to Gene Function 3.1 Storing Information Producing a protein from DNA information involves both
Lecture PowerPoint to accompany Molecular Biology Fourth Edition Robert F. Weaver Chapter 3 An Introduction to Gene Function 3.1 Storing Information Producing a protein from DNA information involves both
Hmwk # 8 : DNA-Binding Proteins : Part II
 The purpose of this exercise is : Hmwk # 8 : DNA-Binding Proteins : Part II 1). to examine the case of a tandem head-to-tail homodimer binding to DNA 2). to view a Zn finger motif 3). to consider the case
The purpose of this exercise is : Hmwk # 8 : DNA-Binding Proteins : Part II 1). to examine the case of a tandem head-to-tail homodimer binding to DNA 2). to view a Zn finger motif 3). to consider the case
Τάσος Οικονόµου ιαλεξη 8. Kινηση, λειτουργια, ελεγχος.
 Τάσος Οικονόµου ιαλεξη 8 Kινηση, λειτουργια, ελεγχος http://ecoserver.imbb.forth.gr/bio321.htm εν ξεχνω. Cell The peptide bond Polypeptides are stabilized by: 1. Covalent bonds= amide bond 2. Noncovalent,
Τάσος Οικονόµου ιαλεξη 8 Kινηση, λειτουργια, ελεγχος http://ecoserver.imbb.forth.gr/bio321.htm εν ξεχνω. Cell The peptide bond Polypeptides are stabilized by: 1. Covalent bonds= amide bond 2. Noncovalent,
Differential Gene Expression
 Developmental Biology Biology 4361 Differential Gene Expression October 13, 2005 core transcription initiation site 5 promoter 3 TATAT +1 upstream downstream Basal transcription factors (eukaryotes) TFIID
Developmental Biology Biology 4361 Differential Gene Expression October 13, 2005 core transcription initiation site 5 promoter 3 TATAT +1 upstream downstream Basal transcription factors (eukaryotes) TFIID
Transcription in Prokaryotes. Jörg Bungert, PhD Phone:
 Transcription in Prokaryotes Jörg Bungert, PhD Phone: 352-273-8098 Email: jbungert@ufl.edu Objectives Understand the basic mechanism of transcription. Know the function of promoter elements and associating
Transcription in Prokaryotes Jörg Bungert, PhD Phone: 352-273-8098 Email: jbungert@ufl.edu Objectives Understand the basic mechanism of transcription. Know the function of promoter elements and associating
Current Status and Future Prospects p. 46 Acknowledgements p. 46 References p. 46 Hammerhead Ribozyme Crystal Structures and Catalysis
 Ribozymes and RNA Catalysis: Introduction and Primer What are Ribozymes? p. 1 What is the Role of Ribozymes in Cells? p. 1 Ribozymes Bring about Significant Rate Enhancements p. 4 Why Study Ribozymes?
Ribozymes and RNA Catalysis: Introduction and Primer What are Ribozymes? p. 1 What is the Role of Ribozymes in Cells? p. 1 Ribozymes Bring about Significant Rate Enhancements p. 4 Why Study Ribozymes?
CHAPTER 17 FROM GENE TO PROTEIN. Section C: The Synthesis of Protein
 CHAPTER 17 FROM GENE TO PROTEIN Section C: The Synthesis of Protein 1. Translation is the RNA-directed synthesis of a polypeptide: a closer look 2. Signal peptides target some eukaryotic polypeptides to
CHAPTER 17 FROM GENE TO PROTEIN Section C: The Synthesis of Protein 1. Translation is the RNA-directed synthesis of a polypeptide: a closer look 2. Signal peptides target some eukaryotic polypeptides to
Biochemistry Prof. S. DasGupta Department of Chemistry Indian Institute of Technology Kharagpur. Lecture - 5 Protein Structure - III
 Biochemistry Prof. S. DasGupta Department of Chemistry Indian Institute of Technology Kharagpur Lecture - 5 Protein Structure - III This is lecture number three on protein structure. (Refer Slide Time:
Biochemistry Prof. S. DasGupta Department of Chemistry Indian Institute of Technology Kharagpur Lecture - 5 Protein Structure - III This is lecture number three on protein structure. (Refer Slide Time:
Eukaryotic Gene Expression Prof. P. N. Rangarajan Department of Biochemistry Indian Institute of Science, Bangalore
 Eukaryotic Gene Expression Prof. P. N. Rangarajan Department of Biochemistry Indian Institute of Science, Bangalore Lecture No. # 06 Eukaryotic Transcription Factors: Transcription Activation Domains In
Eukaryotic Gene Expression Prof. P. N. Rangarajan Department of Biochemistry Indian Institute of Science, Bangalore Lecture No. # 06 Eukaryotic Transcription Factors: Transcription Activation Domains In
Structure formation and association of biomolecules. Prof. Dr. Martin Zacharias Lehrstuhl für Molekulardynamik (T38) Technische Universität München
 Structure formation and association of biomolecules Prof. Dr. Martin Zacharias Lehrstuhl für Molekulardynamik (T38) Technische Universität München Motivation Many biomolecules are chemically synthesized
Structure formation and association of biomolecules Prof. Dr. Martin Zacharias Lehrstuhl für Molekulardynamik (T38) Technische Universität München Motivation Many biomolecules are chemically synthesized
Structural Bioinformatics GENOME 541 Spring 2018
 Molecular composition of a rapidly dividing Escherichia coli cell Structural Bioinformatics GENOME 541 Spring 2018 Lecture 4: Nucleic Acids Frank DiMaio (dimaio@uw.edu) The major biopolymers DNA structure
Molecular composition of a rapidly dividing Escherichia coli cell Structural Bioinformatics GENOME 541 Spring 2018 Lecture 4: Nucleic Acids Frank DiMaio (dimaio@uw.edu) The major biopolymers DNA structure
A primer on the structure and function of genes
 A primer on the structure and function of genes What is the definition of a gene? GENE: the genetic element which is transmitted from parent to offspring during the process of reproduction that influences
A primer on the structure and function of genes What is the definition of a gene? GENE: the genetic element which is transmitted from parent to offspring during the process of reproduction that influences
Eukaryotic transcription (II)
 Eukaryotic transcription (II) Transcription factors Prokaryote: Sigma factors they are similar to general transcription factor of eukaryotes, they help bring RNA pol to the promoter Transcription factors
Eukaryotic transcription (II) Transcription factors Prokaryote: Sigma factors they are similar to general transcription factor of eukaryotes, they help bring RNA pol to the promoter Transcription factors
Chemistry 1050 Exam 4 Study Guide
 Chapter 19 Chemistry 1050 Exam 4 Study Guide 19.1 and 19.2 Know there are 20 common amino acids that can polymerize into proteins. Know why amino acids are called alpha amino acids. Identify the charges
Chapter 19 Chemistry 1050 Exam 4 Study Guide 19.1 and 19.2 Know there are 20 common amino acids that can polymerize into proteins. Know why amino acids are called alpha amino acids. Identify the charges
Chemistry 1120 Exam 3 Study Guide
 Chemistry 1120 Exam 3 Study Guide Chapter 9 9.1 and 9.2 Know there are 20 common amino acids that can polymerize into proteins. Know why amino acids are called alpha amino acids. Identify the charges of
Chemistry 1120 Exam 3 Study Guide Chapter 9 9.1 and 9.2 Know there are 20 common amino acids that can polymerize into proteins. Know why amino acids are called alpha amino acids. Identify the charges of
Gene Expression and Regulation - 1
 Gene Expression and Regulation - 1 We have been discussing the molecular structure of DNA and its function in DNA replication and in transcription. Earlier we discussed how genes interact in transmission
Gene Expression and Regulation - 1 We have been discussing the molecular structure of DNA and its function in DNA replication and in transcription. Earlier we discussed how genes interact in transmission
Giri Narasimhan. CAP 5510: Introduction to Bioinformatics. ECS 254; Phone: x3748
 CAP 5510: Introduction to Bioinformatics Giri Narasimhan ECS 254; Phone: x3748 giri@cis.fiu.edu www.cis.fiu.edu/~giri/teach/bioinfs07.html 2/19/07 CAP5510 1 HMM for Sequence Alignment 2/19/07 CAP5510 2
CAP 5510: Introduction to Bioinformatics Giri Narasimhan ECS 254; Phone: x3748 giri@cis.fiu.edu www.cis.fiu.edu/~giri/teach/bioinfs07.html 2/19/07 CAP5510 1 HMM for Sequence Alignment 2/19/07 CAP5510 2
Chapter 12: Molecular Biology of the Gene
 Biology Textbook Notes Chapter 12: Molecular Biology of the Gene p. 214-219 The Genetic Material (12.1) - Genetic Material must: 1. Be able to store information that pertains to the development, structure,
Biology Textbook Notes Chapter 12: Molecular Biology of the Gene p. 214-219 The Genetic Material (12.1) - Genetic Material must: 1. Be able to store information that pertains to the development, structure,
Gene Regulation in Eukaryotes
 Gene Regulation in Eukaryotes The latest estimates are that a human cell, a eukaryotic cell, contains 20,000 25,000 genes. Some of these are expressed in all cells all the time. These so-called housekeeping
Gene Regulation in Eukaryotes The latest estimates are that a human cell, a eukaryotic cell, contains 20,000 25,000 genes. Some of these are expressed in all cells all the time. These so-called housekeeping
STSs and ESTs. Sequence-Tagged Site: short, unique sequence Expressed Sequence Tag: short, unique sequence from a coding region
 STSs and ESTs Sequence-Tagged Site: short, unique sequence Expressed Sequence Tag: short, unique sequence from a coding region 1991: 609 ESTs [Adams et al.] June 2000: 4.6 million in dbest Genome sequencing
STSs and ESTs Sequence-Tagged Site: short, unique sequence Expressed Sequence Tag: short, unique sequence from a coding region 1991: 609 ESTs [Adams et al.] June 2000: 4.6 million in dbest Genome sequencing
Nucleic Acids and Proteins
 DNA Structure 7.1 Nucleic Acids and Proteins Topic 7 DNA is composed of subunits called nucleotides The four types of bases on nucleotides are: Adenine, Guanine, Cytosine, and Thyamine Purines: two ring
DNA Structure 7.1 Nucleic Acids and Proteins Topic 7 DNA is composed of subunits called nucleotides The four types of bases on nucleotides are: Adenine, Guanine, Cytosine, and Thyamine Purines: two ring
Pre-AP Biology DNA and Biotechnology Study Guide #1
 Last Name: First Name: Per. Pre-AP Biology DNA and Biotechnology Study Guide #1 Structure of DNA: Number of strands. Parallel or antiparallel?. Rosalind Franklin s x-ray crystallography image indicated
Last Name: First Name: Per. Pre-AP Biology DNA and Biotechnology Study Guide #1 Structure of DNA: Number of strands. Parallel or antiparallel?. Rosalind Franklin s x-ray crystallography image indicated
Molekulare Mechanismen der Signaltransduktion
 Molekulare Mechanismen der Signaltransduktion 12 Mechanism of auxin perception slides: http://tinyurl.com/modul-mms doi: 10.1038/nature05731 previous model http://www.plantcell.org/content/17/9/2425/f1.large.jpg
Molekulare Mechanismen der Signaltransduktion 12 Mechanism of auxin perception slides: http://tinyurl.com/modul-mms doi: 10.1038/nature05731 previous model http://www.plantcell.org/content/17/9/2425/f1.large.jpg
Figure A summary of spontaneous alterations likely to require DNA repair.
 DNA Damage Figure 5-46. A summary of spontaneous alterations likely to require DNA repair. The sites on each nucleotide that are known to be modified by spontaneous oxidative damage (red arrows), hydrolytic
DNA Damage Figure 5-46. A summary of spontaneous alterations likely to require DNA repair. The sites on each nucleotide that are known to be modified by spontaneous oxidative damage (red arrows), hydrolytic
Question Expected Answers Mark Additional Guidance 1 (a)
 Question Expected Answers Mark Additional Guidance (a) Mark the first answer for each letter. If the answer is correct and an additional answer is given that is incorrect or contradicts the correct answer
Question Expected Answers Mark Additional Guidance (a) Mark the first answer for each letter. If the answer is correct and an additional answer is given that is incorrect or contradicts the correct answer
Gene regulation II Biochemistry 302. Bob Kelm March 1, 2004
 Gene regulation II Biochemistry 302 Bob Kelm March 1, 2004 Lessons to learn from bacteriophage λ in terms of transcriptional regulation Similarities to E. coli Cis-elements (operator elements) are adjacent
Gene regulation II Biochemistry 302 Bob Kelm March 1, 2004 Lessons to learn from bacteriophage λ in terms of transcriptional regulation Similarities to E. coli Cis-elements (operator elements) are adjacent
Synthetic cells: do bacteria need all its genes? No.
 NO NEED TO REFER TO THE SLIDES. بسم هللا الرحمن الرحيم Do we need all the non coding regions of the DNA? Two weeks ago, they discovered that the genome of a plant is very small (recall that plant genome
NO NEED TO REFER TO THE SLIDES. بسم هللا الرحمن الرحيم Do we need all the non coding regions of the DNA? Two weeks ago, they discovered that the genome of a plant is very small (recall that plant genome
Information specifying protein structure. Chapter 19 Nucleic Acids Nucleotides Are the Building Blocks of Nucleic Acids
 Chapter 19 Nucleic Acids Information specifying protein structure Nucleic acids represent the fourth major class of biomolecules (other major classes of biomolecules are proteins, carbohydrates, fats)
Chapter 19 Nucleic Acids Information specifying protein structure Nucleic acids represent the fourth major class of biomolecules (other major classes of biomolecules are proteins, carbohydrates, fats)
Proteins: Wide range of func2ons. Polypep2des. Amino Acid Monomers
 Proteins: Wide range of func2ons Proteins coded in DNA account for more than 50% of the dry mass of most cells Protein func9ons structural support storage transport cellular communica9ons movement defense
Proteins: Wide range of func2ons Proteins coded in DNA account for more than 50% of the dry mass of most cells Protein func9ons structural support storage transport cellular communica9ons movement defense
RNA Part I: Chemical Structure of RNA
 RA Part I: Chemical Structure of RA Structural differences between RA and DA Resistance of phosphate esters to basic hydrolysis The 2 - group of RA facilitates chemical cleavage in aqueous a by forming
RA Part I: Chemical Structure of RA Structural differences between RA and DA Resistance of phosphate esters to basic hydrolysis The 2 - group of RA facilitates chemical cleavage in aqueous a by forming
From DNA to Protein Structure and Function
 STO-106 From DNA to Protein Structure and Function The nucleus of every cell contains chromosomes. These chromosomes are made of DNA molecules. Each DNA molecule consists of many genes. Each gene carries
STO-106 From DNA to Protein Structure and Function The nucleus of every cell contains chromosomes. These chromosomes are made of DNA molecules. Each DNA molecule consists of many genes. Each gene carries
Introduction to Bioinformatics Online Course: IBT
 Introduction to Bioinformatics Online Course: IBT Multiple Sequence Alignment Building Multiple Sequence Alignment Lec6:Interpreting Your Multiple Sequence Alignment Interpreting Your Multiple Sequence
Introduction to Bioinformatics Online Course: IBT Multiple Sequence Alignment Building Multiple Sequence Alignment Lec6:Interpreting Your Multiple Sequence Alignment Interpreting Your Multiple Sequence
Biosensors. DNA Microarrays (for chemical analysis) Protein Sensors (for identifying viruses)
 Biosensors DNA Microarrays (for chemical analysis) Protein Sensors (for identifying viruses) DNA Microarrays 40 000 detectors in parallel, each detecting a specific DNA sequence. Combinatorial Chemistry
Biosensors DNA Microarrays (for chemical analysis) Protein Sensors (for identifying viruses) DNA Microarrays 40 000 detectors in parallel, each detecting a specific DNA sequence. Combinatorial Chemistry
CELL BIOLOGY - CLUTCH CH. 7 - GENE EXPRESSION.
 !! www.clutchprep.com CONCEPT: CONTROL OF GENE EXPRESSION BASICS Gene expression is the process through which cells selectively to express some genes and not others Every cell in an organism is a clone
!! www.clutchprep.com CONCEPT: CONTROL OF GENE EXPRESSION BASICS Gene expression is the process through which cells selectively to express some genes and not others Every cell in an organism is a clone
TALENs (Transcription Activator-Like Effector Nucleases)
 TALENs (Transcription Activator-Like Effector Nucleases) The fundamental rationale between TALENs and ZFNs is similar, namely, combine a sequencespecific DNA-binding peptide domain with a nuclease domain
TALENs (Transcription Activator-Like Effector Nucleases) The fundamental rationale between TALENs and ZFNs is similar, namely, combine a sequencespecific DNA-binding peptide domain with a nuclease domain
Comments. polyproline ( PXXP ) motif for SH3 binding RGD motif for integrin binding GXXXG motif within the TM domain of membrane protein
 Comments Structural motif v sequence motif polyproline ( PXXP ) motif for SH3 binding RGD motif for integrin binding GXXXG motif within the TM domain of membrane protein Most common type I beta turn sequences:
Comments Structural motif v sequence motif polyproline ( PXXP ) motif for SH3 binding RGD motif for integrin binding GXXXG motif within the TM domain of membrane protein Most common type I beta turn sequences:
1. Cell chemistry and biosynthesis (chapter 2) 3. DNA and chromosomes (chapter 4) 4. 추석 5. Control of gene expression (chapter 7)
 Week Topics 1. Cell chemistry and biosynthesis (chapter 2) 2. Proteins: structure and function (chapter 3) 3. DNA and chromosomes (chapter 4) 4. 추석 5. Control of gene expression (chapter 7) 6. Manipulating
Week Topics 1. Cell chemistry and biosynthesis (chapter 2) 2. Proteins: structure and function (chapter 3) 3. DNA and chromosomes (chapter 4) 4. 추석 5. Control of gene expression (chapter 7) 6. Manipulating
From mechanism to medicne
 From mechanism to medicne a look at proteins and drug design Chem 342 δ δ δ+ M 2009 δ+ δ+ δ M Drug Design - an Iterative Approach @ DSU Structural Analysis of Receptor Structural Analysis of Ligand-Receptor
From mechanism to medicne a look at proteins and drug design Chem 342 δ δ δ+ M 2009 δ+ δ+ δ M Drug Design - an Iterative Approach @ DSU Structural Analysis of Receptor Structural Analysis of Ligand-Receptor
Nucleic Acid Structure. Nucleic Acid Sequence Abbreviations. Sequence Abbreviations, con t.
 BC 4054 Spring 2001 Chapter 11 & 12 Review Lecture otes Slide 1 ucleic Acid Structure Linear polymer of nucleotides Phosphodiester linkage between 3 and 5 positions See Figure 11.17 Slide 2 ucleic Acid
BC 4054 Spring 2001 Chapter 11 & 12 Review Lecture otes Slide 1 ucleic Acid Structure Linear polymer of nucleotides Phosphodiester linkage between 3 and 5 positions See Figure 11.17 Slide 2 ucleic Acid
Proteins and folding
 Proteins and folding October 8th, 2013. Kinga Futó Biopolymers Mitotic spindle of a dividing cell Actin filament network in the epidermal cell of Tobacco leaf. DNA strand released from bacteriophage 1
Proteins and folding October 8th, 2013. Kinga Futó Biopolymers Mitotic spindle of a dividing cell Actin filament network in the epidermal cell of Tobacco leaf. DNA strand released from bacteriophage 1
Antibody-Antigen recognition. Structural Biology Weekend Seminar Annegret Kramer
 Antibody-Antigen recognition Structural Biology Weekend Seminar 10.07.2005 Annegret Kramer Contents Function and structure of antibodies Features of antibody-antigen interfaces Examples of antibody-antigen
Antibody-Antigen recognition Structural Biology Weekend Seminar 10.07.2005 Annegret Kramer Contents Function and structure of antibodies Features of antibody-antigen interfaces Examples of antibody-antigen
All Rights Reserved. U.S. Patents 6,471,520B1; 5,498,190; 5,916, North Market Street, Suite CC130A, Milwaukee, WI 53202
 Secondary Structure In the previous protein folding activity, you created a hypothetical 15-amino acid protein and learned that basic principles of chemistry determine how each protein spontaneously folds
Secondary Structure In the previous protein folding activity, you created a hypothetical 15-amino acid protein and learned that basic principles of chemistry determine how each protein spontaneously folds
CHAPTER 18 LECTURE NOTES: CONTROL OF GENE EXPRESSION PART B: CONTROL IN EUKARYOTES
 CHAPTER 18 LECTURE NOTES: CONTROL OF GENE EXPRESSION PART B: CONTROL IN EUKARYOTES I. Introduction A. No operon structures in eukaryotes B. Regulation of gene expression is frequently tissue specific.
CHAPTER 18 LECTURE NOTES: CONTROL OF GENE EXPRESSION PART B: CONTROL IN EUKARYOTES I. Introduction A. No operon structures in eukaryotes B. Regulation of gene expression is frequently tissue specific.
