From Disease Genes to Cellular Pathways: A Progress Report
|
|
|
- Kelly Baldwin
- 6 years ago
- Views:
Transcription
1 From Disease Genes to Cellular Pathways: A Progress Report J. Yu 1,2, A.J. Mears 1, S. Yoshida 1, R. Farjo 1, T.A. Carter 3, D. Ghosh 4, A. Hero 2,5, C. Barlow 3, A. 1, 6, # Swaroop Departments of 1 Ophthalmology and Visual Sciences, 2 Biomedical Engineering, 4 Biostatistics, 5 Statistics and 6 Human Genetics, University of Michigan, Ann Arbor, MI, USA. 3 The Salk Institute for Biological Studies, Laboratory of Genetics, North Torrey Pines Road, La Jolla, CA 92037, USA. Short title: Gene Profiling of retinal development and degeneration Keywords: Nrl, transcription factors, signaling pathways, gene expression, microarray, GeneChips, retinal dystrophy, knockout # To whom correspondence should be addressed: Anand Swaroop, Ph.D., University of Michigan, W.K. Kellogg Eye Center; 1000 Wall Street, Ann Arbor, MI Phone: (734) ; Fax: (734) ; swaroop@umich.edu - 1 -
2 Abstract Mutations in a large number of retinal and retinal pigment epithelium (RPE) expressed genes can lead to the degeneration of photoreceptors and consequently the loss of vision. The genetic and phenotypic heterogeneity of retinal dystrophies poses a complex problem with respect to rational development of therapeutic strategies. Delineation of physiological functions of disease genes and identification of pathways that lead to disease pathogenesis represent essential goals towards developing a systematic and global approach to gene-based treatments. We are interested in identifying cellular pathways that are involved in photoreceptor differentiation, function and degeneration. We are, therefore, generating comprehensive gene expression profiles of retina and RPE of humans and mice using both cdna- and oligonucleotide-based (Affymetrix) microarrays. Because of the under-representation of retinal/ RPE genes in the public databases, we have constructed several unamplified cdna libraries and produced almost twenty thousand expressed sequence tags (ESTs) that are being printed onto glass slides ( I-Gene microarrays). In this presentation, we will report the microarray analysis of the rod-less (and cone-enhanced) retina from the Nrl-knockout mouse as a paradigm to initiate the identification of cellular pathways involved in photoreceptor differentiation and function
3 Background and Basic Concepts Retinal dystrophies (RD) comprise a group of clinically and genetically heterogeneous retinal disorders, which typically result in the degeneration of photoreceptors followed by the impairment or loss of vision. To date, the online retinal information network (RetNet, has listed over 130 loci associated with retinal dystrophies. RD is a major cause of blindness in the industrialized world and is, for the most part, currently untreatable. Retinitis pigmentosa (RP) primarily causes rod photoreceptor degeneration and early symptoms include night blindness and loss of peripheral vision. The prevalence of RP is approximately 1/3000, with a total of over 1.5 million people affected world-wide (Saleem & Walter 2002). In contrast, cone dysfunction occurs early during the progression of cone or conerod dystrophies (CRD), thereby affecting visual acuity and color vision. Leber congenital amaurosis (LCA) is the most common cause of congenital visual impairment with age of onset in infants or children. LCA accounts for 5-10% of all retinal dystrophies and is perhaps the most severe RD. Age-related Macular degeneration (AMD) is highly prevalent in the elderly population, accounting for 22% of monocular blindness and 75% of legal blindness in adults over age 50 in the Unites States (Klein et al 1995). It preferentially affects the macular region, leading to loss of central vision and visual acuity. Unlike other forms of RD, AMD is the culmination of a complex interplay of genetic and non-genetic components. The complexity afforded by the considerable genetic heterogeneity in RD has greatly hindered the application of gene-based therapies; nonetheless, all of these diseases result in the same fate, i.e., the death of the photoreceptors
4 A number of innovative strategies have been employed with the objectives of slowing down, preventing, or even reversing photoreceptor cell death in RD. One approach of circumventing the heterogeneity of RD is symptom-based disease treatments without correcting underlying genetic defect. To restore sight in highly visually handicapped individuals, several research groups are working on the development of electronic photoreceptor prosthesis (Zrenner et al 2001; Hammerle et al 2002) and cell/tissue transplantations (Otani et al 2002; Radner et al 2002; Semkova et al 2002). However, these strategies are currently limited due to issues regarding biocompatibility, stability and longevity of transplants. Another generic approach involves the use of growth or survival factors (LaVail et al 1998). In any event, the need for understanding both the physiological function of disease genes and the cellular processes leading to photoreceptor degeneration is inescapable. Gene-based therapy seeks to rescue retinal diseases by correcting the underlying genetic defect or a consequent physiological deficiency. Over 80 genes have been associated with retinal dystrophies (Bessant et al 2001; Saleem & Walter 2002), including the neural retinal leucine zipper (NRL) gene, NR2E3 (nuclear receptor subfamily 2, group E, member 3), PDE6B (phosphodiesterase 6B, cgmp-specific, rod, beta), CRX (cone-rod homeobox ) and RHO (Rhodopsin). To rescue RD, numerous researchers have attempted to deliver a functional copy of the mutant gene into photoreceptor cells using viral-based vectors (Bennett et al 1998; Cheng et al 2002). However, gene transfer technology faces a number of hurdles, including the sheer number of distinct targets that need to be addressed due to the heterogeneity of RD, and issues regarding the safety and efficacy of such vectors
5 An alternative approach that we advocate is a therapeutic design based on the understanding of the cellular pathways leading to photoreceptor cell death (Figure 1). Although a large number of retinal and RPE expressed genes can lead to RD, studies have shown that only a few common cellular pathways are involved in disease progression and the photoreceptor cells in many, if not all, forms of RD die via apoptosis (Travis 1998). Pharmacological approaches have been advanced to slow photoreceptor degeneration through the introduction of growth and survival factors (LaVail et al 1998; Liang et al 2001; Tao et al 2002). Unfortunately, most experiments were only able to slow cell death for a week to a month, possibly due to the irreversible stage of disease by the time apoptotic pathways are induced. In order to devise a therapeutic strategy that targets multiple forms of RD prior to the induction of massive photoreceptor cell death, we are elucidating the common pathways of photoreceptor degeneration at a pre-apoptotic stage of disease. As illustrated in Figure 1, pathways of disease pathogenesis initiated by different mutant gene products (or the lack thereof) must converge over time and follow limited routes to cell death. Therefore, temporal profiling of gene expression in normal developing, mature and aging retinas and in retinal degeneration mouse models should lead to the identification of common pre-apoptotic signals (PAS) that can be targeted for drug discovery. A crucial aspect of this approach is the understanding of normal differentiation and function of rods and cones since it serves as the baseline against which abnormal changes may be recognized. We propose that the adaptive response of the retinal neurons or RPE to disease or aging is reflected by modulation of specific cellular pathways and, consequently, changes in gene expression. Profiling of diseased or aging retina or RPE from humans and mice will facilitate in - 5 -
6 the identification of these pathways. In this manuscript, we will primarily focus on the regulatory networks of photoreceptor development and function in the context of the transcription factor Nrl, using the Nrl / mouse as a paradigm. Nrl, an essential transcription factor for rod development and function The Nrl gene, encoding a basic motif leucine zipper protein of Maf-subfamily, was initially identified from a subtracted retinal library (Swaroop et al 1992). It showed a highly restricted pattern of expression, primarily in rod photoreceptors (Farjo et al 1993; Swain et al 2001). Six phosphorylated isoforms of Nrl have been identified in rod but not cone photoreceptor nuclei (Swain et al 2001). The Nrl protein can positively regulate rhodopsin gene expression by binding to an extended AP-1 like sequence element (called NRE) in the upstream promoter region (Kumar et al 1996; Rehemtulla et al 1996). Further studies indicated that Nrl regulates several other rod genes, and can interact with other transcriptional factors, such as Crx, in the regulation of retinal expressed genes ( Chen et al 1997; Mitton et al 2000; Lerner et al 2001). Mutations in the human NRL gene have been associated with autosomal dominant RP (Bessant et al 1999; Bessant et al 2000; Martinez-Gimeno et al 2001; DeAngelis et al 2002). Interestingly, 5 of the 6 currently identified mutations alter the residues S50 and P51, resulting in possibly hypermorphic alleles of NRL and suggesting their functional importance. To define the role of Nrl in photoreceptor development and function, the Nrl gene was deleted in mice by homologous recombination (Mears et al 2001). Since Nrl plays a key role in - 6 -
7 the regulation of rod specific genes, it was anticipated that the deletion of Nrl would affect rod photoreceptors. Surprisingly, the Nrl / mouse retina is functionally rod-less. The knockout retina has abnormal histology, with rosettes and whorls within the outer nuclear layer. Only 20% of photoreceptors elaborate outer segments, most of which have abnormal disk morphology. Electroretinogram (ERG) recording revealed no scotopic response and detected a light-adapted b-wave of two to three times larger amplitude in knockout than that of wild type retina, demonstrating the absence of rod function and an enhanced cone function. Using monochromatic stimuli of 400 nm or 530nm, this large b-wave amplitude is explained by increased S-cone activity. Preliminary gene expression analysis revealed an absence of rodspecific transcripts, and an increase in the expression of cone-specific genes (Mears et al 2001). Dramatic retinal changes observed in this mouse establish it as an excellent model for expression profiling corresponding to different pathways associated with rod and cone development and function. We propose genes with reduced expression at the Nrl / retina relative to normal would be associated with rod signaling pathways, while those with augmented expression relate to cone function. Microarray Analysis High-throughput technologies, including cdna microarrays and Affymetrix GeneChips have made large-scale gene expression studies of retinal tissues readily achievable (Farjo et al 2002; Yoshida et al 2002; Swaroop & Zack 2002). Microarrays allow us to investigate changes in expression at a genome scale in a single experiment. This approach is limited only by the number and types of genes represented on the arrays. In addition to being a powerful gene
8 discovery tool in the identification of candidate genes, microarrays may shed considerable light on the cellular pathways of the tissue under study (Livesey et al 2000; Livesey 2002). A schema of microarray analysis is presented in Figure 2. Although Affymetrix technology is relatively well developed, with appropriate quality controls, standard data preprocessing and ready-to-use data analysis software, its application to our studies is limited by the under-representation of retinal expressed genes on their GeneChips. For comprehensive profiling, customized I-Gene cdna microarrays were also utilized. These arrays were generated by printing retina / eye expressed genes and ESTs obtained from a variety of cdna libraries onto glass slides using a robotic micro-arrayer (Farjo et al 2002; Yu et al 2002). For these high-throughput studies, total RNA was isolated from either control (normal) or experimental (diseased or aging) retinas, labeled with fluorescent dyes and hybridized to either Affymetrix GeneChips or I-Gene microarrays (Figure 2). Image analysis and statistical modeling were employed to identify differentially expressed genes between control and experimental samples. Clustering algorithms were used to group co-expressed genes under different experimental conditions, which might lead to the identification of functional / regulatory networks and pathways (Figure 2). We have used gene profiling of retinas from the normal and Nrl-knockout mice as a paradigm and to establish the proof of principle. Affymetrix GeneChip Study Gene profiling of postnatal day 2 (PN2), PN10 and 2 month-old retinas from the control and Nrl-knockout mice using mouse GeneChips showed approximately equal number of up- or down-regulated genes at each time point (data not shown). At PN2, only 6 genes are found to be - 8 -
9 differentially expressed, compared with 74 at PN10 and 136 at 2-months. As predicted, several rod photoreceptor-specific genes, including rhodopsin (Rho) and rod transducin alpha (Gnat1), were found to be greatly under-expressed in the knockout mouse, while cone genes, such as S- opsin (Opn1sw) and cone transducin alpha (Gnat2), are up-regulated. Quantitative real-time PCR (qrt-pcr) analyses of almost 50 genes have validated gene expression changes revealed by GeneChips; qrt-pcr profiles of 4 genes are shown in Figure 3. More than 20% of differentially expressed transcripts were unknown ESTs. These are of considerable interest, as they may represent novel retinal dystrophy candidate genes or lead to the elucidation of specific cellular pathways associated with photoreceptor differentiation and function. Clusters of differentially expressed genes may also provide insights into pathways and functional networks (Figure 4). I-Gene Micoarray Study Gene expression of wild type and Nrl / mice retinas were compared at 5 developmental time points: PN0, PN2, PN6, PN10, and PN21. Custom I-Gene microarrays containing over 6500 eye/retina expressed genes and ESTs printed in duplicate were generated for hybridization (Figure 5A, B). Five replicates were performed for each stage utilizing labeled targets from different mice to reduce individual variance. Density plots of the log-ratios of gene expression in PN21 Nrl +/+ and Nrl / mice retinas detected by five independent replicated experiments showed similar patterns of distribution. Log-ratios of all replicates are centered at 0, with most genes lying within 1 and +1 (Figure 5C), suggesting that the expression of a majority of genes is unaltered or minimally altered between the control and Nrl-knockout retinas. Microarrays tend - 9 -
10 to underestimate the true biological change and perhaps a lower threshold instead of 1 needs to be established. Statistical analysis of PN21 expression data identified 52 cdnas, representing 39 unique genes, with highest possibility of differential expression. Over 30% of these genes are known to play important roles in the retina; these include Rho, Opn1sw, Gnat1, Gnat2, Nr2e3, Retinoschisis 1 homolog (Rs1h), myosin 5a (Myo5a), and Recoverin (Rcvrn). qrt-pcr analyses validated these expression alterations (Figure 6). Further examination of these differentially expressed genes suggests a bias in the utilization of the Bone Morphogenetic Protein (Bmp) signaling pathway, Wnt/Ca 2+ signaling pathway and the retinoid acid pathway between rods and cones (Yu, Mears and Swaroop, unpublished data). Pathway Consolidation Affymetrix GeneChip studies, presented here, showed differential gene expression from PN2, PN10 to 2-month old retinas, whereas I-Gene cdna microarray data indicated alterations of signaling pathways in the PN21 knockout mice retinas. Systematic examination of gene expression levels at PN0, PN2, PN6, PN10 and PN21 followed by statistical analysis should further assist in the identification of genes that are downstream of Nrl in regulatory hierarchy and play key roles in photoreceptor differentiation and/or function. Clustering based on temporal expression profiles may identify coordinately regulated genes involved in rod and cone photoreceptor development. Since hypermorphic alleles of Nrl are predicted to cause retinal degeneration, the signaling pathways downstream of Nrl may also be studied in the context of other retinal degenerative mouse models
11 Conclusions Delineation of cellular pathways involved in photoreceptor differentiation and disease pathogenesis presents an attractive approach to identify targets for treatment of RD. In this presentation, we have used a single paradigm to illustrate our research approach and the focus on cellular pathways downstream of an important retinal gene. Nrl is a rod-specific transcription factor that is required for rod differentiation and regulation of rod-specific gene expression. Mutations in the human NRL gene have been identified in patients with autosomal dominant RP. The Nrl / mouse retina is rod-less, with an increased number of functional S-cones. Using Affymetrix GeneChips and custom I-Gene cdna microarrays, we have so far identified over 150 genes that are differentially expressed in the Nrl-knockout mouse retina as compared to controls. Several of these cdnas represent novel genes that are attractive candidates for RD. Further characterization of differentially-expressed cdnas should reveal direct or indirect targets of Nrl and assist in developing transcriptional regulatory hierarchy downstream of Nrl. Initial studies also suggest differential utilization of signaling pathways in rods and cones. Our investigations provide an initial framework for establishing pathway-based treatment strategies for retinal and macular diseases
12 Acknowledgements We thank Mohammad Othman and Sean MacNee for their advice and assistance. The research in our laboratory is supported by grants from the National Institutes of Health (EY11115 including administrative supplements, EY07961, and EY07003), The Foundation Fighting Blindness (Owings Mills, MD), Macula Vision Research Foundation (West Conshohocken, PA), Research to Prevent Blindness (New York, NY), Elmer and Sylvia Sramek Charitable Foundation (Chicago, IL), and Juvenile Diabetes Research Foundation (New York, NY)
13 Figure Legends Figure 1 The progression of disease in retinal dystrophies: from genes to pathways. This schematic representation shows our approach for gene-based therapy that focuses on the convergence of different pre-apoptotic cellular pathways in time, in order to develop novel therapeutic targets for several forms of RD. In a majority of retinal dystrophies, the photoreceptors die by apoptosis. Mutations in hundreds of genes may disrupt the cellular homeostasis and selected signaling pathways. M1, M2 represent different mutations in the same gene (rd), and the blue squares indicate various "disease" genes. The response of photoreceptors to the presence of a mutation is predicted to converge on a few pre-apoptotic signaling pathways (PAS1 indicates pre-apoptic signals) that lead eventually to photoreceptor cell death via apoptotic pathways (apo1-3). In this model, various pre-apoptotic signals (PAS) would be ideal targets for drug discovery. Figure 2 Comprehensive gene profiling of control and mutant retinas using Affymetrix GeneChips and custom I-Gene microarrays. Temporal expression profiling followed by statistical modeling and cluster analysis can lead to the identification of pathways and molecular targets. Figure
14 qrt-pcr analyses of four differentially-expressed genes identified by Affymetrix GeneChip analysis. Total RNA from wild type (wt) and Nrl -/- (ko) mice retinas were first reverse transcribed either with or without (-rt) reverse-transcriptase, and then subjected to Realtime PCR. The negative control (-rt) experiments were utilized to demonstrate that RNA samples are free from genomic contamination. qrt-pcr profiles of wt, ko and -rt were shown for 4 genes, Gnat1, Rho, Gnat2 and op1sw. The fold difference was calculated as 2^ difference in threshold cycles (Ct) between wt and ko samples. Affymetrix chips and qrt-pcr showed high concordance for all genes, with qrt-pcr generally being more sensitive. Figure 4 Cluster analysis of the temporal expression profiles generated from Affymetrix GeneChips. (A) Representation of clustering analysis of differentially expressed genes. The data matrix was first standardized to z-score and hierarchical clustering analysis performed using the Euclidean distance method. Color-coding indicates relative expression: green being low, red high. Eight genes shown are clustered based on their similarity of expression profile, which is also graphically represented in (B), where z-scores (Y-axis) are plotted against time-point (X-axis). Figure 5 I-Gene microarray and density plots of log-ratios in five replicate experiments. (A) A TIFF image of the Cy3 channel of an I-Gene microarrays containing over 6500 genes or ESTs printed in duplicate. False color has been applied to indicate the intensity of hybridization, with black
15 having no signal, blue low, red high, and white saturated. (B) Enlargement of the left lower corner grid of the array, showing uniform spot diameter, clear hybridization and low background signal. (C) Ratios of gene expression indicate the abundance of each gene in Nrl -/- mice retinas relative to Nrl +/+ retinas. Smooth density plots of log-ratios shows that, in all replicates (expt1- expt5), log-ratios are centered at 0, with majority of spots lying within 1 and +1. Figure 6 qrt-pcr validation of I-Gene microarray results: Analysis of Nr2e3, Rs1h, Myo5a, and Rcvrn expression in retinas of wild-type (wt) and Nrl-knockout (ko) mice. Total RNA from wt and ko mice retinas were first reverse transcribed either with or without (-rt) reversetranscriptase, and then subjected to real-time PCR. The negative control (-rt) experiments were utilized to demonstrate that RNA samples are free from genomic DNA contamination. qrt- PCR tends to be more sensitive than the hybridization-based microarray experiments
16 Figure
17 Figure
18 Figure
19 Figure
20 Figure
21 Figure
22 REFERENCES Bennett J, Zeng Y, Bajwa R, Klatt L, Li Y, Maguire AM 1998 Adenovirus-mediated delivery of rhodopsin-promoted bcl-2 results in a delay in photoreceptor cell death in the rd/rd mouse. Gene Ther 5: Bessant DA, Ali RR, Bhattacharya SS 2001 Molecular genetics and prospects for therapy of the inherited retinal dystrophies. Curr Opin Genet Dev 11: Bessant DA, Payne AM, Mitton KP, et al 1999 A mutation in NRL is associated with autosomal dominant retinitis pigmentosa. Nat Genet 21: Bessant DA, Payne AM, Plant C, Bird AC, Swaroop A, Bhattacharya SS 2000 NRL S50T mutation and the importance of 'founder effects' in inherited retinal dystrophies. Eur J Hum Genet 8: Chen S, Wang QL, Nie Z, et al 1997 Crx, a novel Otx-like paired-homeodomain protein, binds to and transactivates photoreceptor cell-specific genes. Neuron 19: Cheng L, Chaidhawangul S, Wong-Staal F, et al 2002 Human immunodeficiency virus type 2 (HIV-2) vector-mediated in vivo gene transfer into adult rabbit retina. Curr Eye Res 24: DeAngelis MM, Grimsby JL, Sandberg MA, Berson EL, Dryja TP 2002 Novel mutations in the NRL gene and associated clinical findings in patients with dominant retinitis pigmentosa. Arch Ophthalmol 120: Farjo Q, Jackson AU, Xu J, et al 1993 Molecular characterization of the murine neural retina leucine zipper gene, Nrl. Genomics 18: Farjo R, Yu J, Othman MI, et al 2002 Mouse eye gene microarrays for investigating ocular development and disease. Vision Res 42: Hammerle H, Kobuch K, Kohler K, Nisch W, Sachs H, Stelzle M 2002 Biostability of microphotodiode arrays for subretinal implantation. Biomaterials 23: Klein R, Wang Q, Klein BE, Moss SE, Meuer SM 1995 The relationship of age-related maculopathy, cataract, and glaucoma to visual acuity. Invest Ophthalmol Vis Sci 36: Kumar R, Chen S, Scheurer D, et al 1996 The bzip transcription factor Nrl stimulates rhodopsin promoter activity in primary retinal cell cultures. J Biol Chem 271: LaVail MM, Yasumura D, Matthes MT, et al 1998 Protection of mouse photoreceptors by survival factors in retinal degenerations. Invest Ophthalmol Vis Sci 39: Lerner LE, Gribanova YE, Ji M, Knox BE, Farber DB 2001 Nrl and Sp nuclear proteins mediate transcription of rod-specific cgmp-phosphodiesterase beta-subunit gene: involvement of multiple response elements. J Biol Chem 276: Liang FQ, Dejneka NS, Cohen DR, et al 2001 AAV-mediated delivery of ciliary neurotrophic factor prolongs photoreceptor survival in the rhodopsin knockout mouse. Mol Ther 3: Livesey FJ, Furukawa T, Steffen MA, Church GM, Cepko CL 2000 Microarray analysis of the transcriptional network controlled by the photoreceptor homeobox gene Crx. Curr Biol 10: Livesey R 2002 Have microarrays failed to deliver for developmental biology? Genome Biol 3: COMMENT
23 Martinez-Gimeno M, Maseras M, Baiget M, et al 2001 Mutations P51U and G122E in retinal transcription factor NRL associated with autosomal dominant and sporadic retinitis pigmentosa. Hum Mutat 17: 520 Mears AJ, Kondo M, Swain PK, et al 2001 Nrl is required for rod photoreceptor development. Nat Genet 29: Mitton KP, Swain PK, Chen S, Xu S, Zack DJ, Swaroop A 2000 The leucine zipper of NRL interacts with the CRX homeodomain. A possible mechanism of transcriptional synergy in rhodopsin regulation. J Biol Chem 275: Otani A, Kinder K, Ewalt K, Otero FJ, Schimmel P, Friedlander M 2002 Bone marrow-derived stem cells target retinal astrocytes and can promote or inhibit retinal angiogenesis. Nat Med 8: Radner W, Sadda SR, Humayun MS, Suzuki S, de Juan E, Jr Increased spontaneous retinal ganglion cell activity in rd mice after neural retinal transplantation. Invest Ophthalmol Vis Sci 43: Rehemtulla A, Warwar R, Kumar R, Ji X, Zack DJ, Swaroop A 1996 The basic motif-leucine zipper transcription factor Nrl can positively regulate rhodopsin gene expression. Proc Natl Acad Sci U S A 93: Saleem RA, Walter MA 2002 The complexities of ocular genetics. Clin Genet 61: Semkova I, Kreppel F, Welsandt G, et al 2002 Autologous transplantation of genetically modified iris pigment epithelial cells: A promising concept for the treatment of agerelated macular degeneration and other disorders of the eye. Proc Natl Acad Sci U S A 99: Swain PK, Hicks D, Mears AJ, et al 2001 Multiple phosphorylated isoforms of NRL are expressed in rod photoreceptors. J Biol Chem 276: Swaroop A, Xu JZ, Pawar H, Jackson A, Skolnick C, Agarwal N 1992 A conserved retinaspecific gene encodes a basic motif/leucine zipper domain. Proc Natl Acad Sci U S A 89: Swaroop A, Zack DJ 2002 Transcriptome analysis of the retina. Genome Biol 3: REVIEWS1022 Tao W, Wen R, Goddard MB, et al 2002 Encapsulated cell-based delivery of CNTF reduces photoreceptor degeneration in animal models of retinitis pigmentosa. Invest Ophthalmol Vis Sci 43: Travis GH 1998 Mechanisms of cell death in the inherited retinal degenerations. Am J Hum Genet 62: Yoshida S, Yashar BM, Hiriyanna S, Swaroop A 2002 Microarray analysis of gene expression in the aging human retina. Invest Ophthalmol Vis Sci 43: Yu J, Othman MI, Farjo R, et al 2002 Evaluation and optimization of procedures for target labeling and hybridization of cdna microarrays. Mol Vis 8: Zrenner E, Gekeler F, Gabel VP, et al 2001 Subretinal microphotodiode array as replacement for degenerated photoreceptors? Ophthalmologe 98:
ALLERGAN ENTERS STRATEGIC R&D ALLIANCE WITH EDITAS MEDICINE TO DISCOVER AND DEVELOP CRISPR GENE EDITING PROGRAMS FOR OCULAR DISEASES
 ALLERGAN ENTERS STRATEGIC R&D ALLIANCE WITH EDITAS MEDICINE TO DISCOVER AND DEVELOP CRISPR GENE EDITING PROGRAMS FOR OCULAR DISEASES Novel Development Programs for Potential Treatments of Serious Ocular
ALLERGAN ENTERS STRATEGIC R&D ALLIANCE WITH EDITAS MEDICINE TO DISCOVER AND DEVELOP CRISPR GENE EDITING PROGRAMS FOR OCULAR DISEASES Novel Development Programs for Potential Treatments of Serious Ocular
Recent technology allow production of microarrays composed of 70-mers (essentially a hybrid of the two techniques)
 Microarrays and Transcript Profiling Gene expression patterns are traditionally studied using Northern blots (DNA-RNA hybridization assays). This approach involves separation of total or polya + RNA on
Microarrays and Transcript Profiling Gene expression patterns are traditionally studied using Northern blots (DNA-RNA hybridization assays). This approach involves separation of total or polya + RNA on
Impact of Retinoic acid induced-1 (Rai1) on Regulators of Metabolism and Adipogenesis
 Impact of Retinoic acid induced-1 (Rai1) on Regulators of Metabolism and Adipogenesis The mammalian system undergoes ~24 hour cycles known as circadian rhythms that temporally orchestrate metabolism, behavior,
Impact of Retinoic acid induced-1 (Rai1) on Regulators of Metabolism and Adipogenesis The mammalian system undergoes ~24 hour cycles known as circadian rhythms that temporally orchestrate metabolism, behavior,
Gene Expression Technology
 Gene Expression Technology Bing Zhang Department of Biomedical Informatics Vanderbilt University bing.zhang@vanderbilt.edu Gene expression Gene expression is the process by which information from a gene
Gene Expression Technology Bing Zhang Department of Biomedical Informatics Vanderbilt University bing.zhang@vanderbilt.edu Gene expression Gene expression is the process by which information from a gene
The Effects of Scaffold Rigidity on Retinal Pigment Epithelial Cells. Corina White Symposium on Biomaterials Science 24 October 2016
 The Effects of Scaffold Rigidity on Retinal Pigment Epithelial Cells Corina White Symposium on Biomaterials Science 24 October 2016 Background Physiology The retina is the light-responsive tissue layer
The Effects of Scaffold Rigidity on Retinal Pigment Epithelial Cells Corina White Symposium on Biomaterials Science 24 October 2016 Background Physiology The retina is the light-responsive tissue layer
Data Mining For Genomics
 Data Mining For Genomics Alfred O. Hero III The University of Michigan, Ann Arbor, MI ISTeC Seminar, CSU Feb. 22, 23 1. Biotechnology Overview 2. Gene Microarray Technology 3. Mining the genomic database
Data Mining For Genomics Alfred O. Hero III The University of Michigan, Ann Arbor, MI ISTeC Seminar, CSU Feb. 22, 23 1. Biotechnology Overview 2. Gene Microarray Technology 3. Mining the genomic database
Lecture #1. Introduction to microarray technology
 Lecture #1 Introduction to microarray technology Outline General purpose Microarray assay concept Basic microarray experimental process cdna/two channel arrays Oligonucleotide arrays Exon arrays Comparing
Lecture #1 Introduction to microarray technology Outline General purpose Microarray assay concept Basic microarray experimental process cdna/two channel arrays Oligonucleotide arrays Exon arrays Comparing
Microarray Technique. Some background. M. Nath
 Microarray Technique Some background M. Nath Outline Introduction Spotting Array Technique GeneChip Technique Data analysis Applications Conclusion Now Blind Guess? Functional Pathway Microarray Technique
Microarray Technique Some background M. Nath Outline Introduction Spotting Array Technique GeneChip Technique Data analysis Applications Conclusion Now Blind Guess? Functional Pathway Microarray Technique
Genetics - Problem Drill 19: Dissection of Gene Function: Mutational Analysis of Model Organisms
 Genetics - Problem Drill 19: Dissection of Gene Function: Mutational Analysis of Model Organisms No. 1 of 10 1. The mouse gene knockout is based on. (A) Homologous recombination (B) Site-specific recombination
Genetics - Problem Drill 19: Dissection of Gene Function: Mutational Analysis of Model Organisms No. 1 of 10 1. The mouse gene knockout is based on. (A) Homologous recombination (B) Site-specific recombination
Severe Stargardt disease with peripapillary sparing
 www.edoriumjournals.com CLINICAL IMAGES PEER REVIEWED OPEN ACCESS Severe Stargardt disease with peripapillary sparing Heather Leisy, Meleha Ahmad, Nathaniel Tracer, R. Theodore Smith ABSTRACT Abstract
www.edoriumjournals.com CLINICAL IMAGES PEER REVIEWED OPEN ACCESS Severe Stargardt disease with peripapillary sparing Heather Leisy, Meleha Ahmad, Nathaniel Tracer, R. Theodore Smith ABSTRACT Abstract
Feature Selection of Gene Expression Data for Cancer Classification: A Review
 Available online at www.sciencedirect.com ScienceDirect Procedia Computer Science 50 (2015 ) 52 57 2nd International Symposium on Big Data and Cloud Computing (ISBCC 15) Feature Selection of Gene Expression
Available online at www.sciencedirect.com ScienceDirect Procedia Computer Science 50 (2015 ) 52 57 2nd International Symposium on Big Data and Cloud Computing (ISBCC 15) Feature Selection of Gene Expression
sirna Overview and Technical Tips
 1 sirna Overview and Technical Tips 2 CONTENTS 3 4 5 7 8 10 11 13 14 18 19 20 21 Introduction Applications How Does It Work? Handy Tips Troubleshooting Conclusions Further References Contact Us 3 INTRODUCTION
1 sirna Overview and Technical Tips 2 CONTENTS 3 4 5 7 8 10 11 13 14 18 19 20 21 Introduction Applications How Does It Work? Handy Tips Troubleshooting Conclusions Further References Contact Us 3 INTRODUCTION
SIMS2003. Instructors:Rus Yukhananov, Alex Loguinov BWH, Harvard Medical School. Introduction to Microarray Technology.
 SIMS2003 Instructors:Rus Yukhananov, Alex Loguinov BWH, Harvard Medical School Introduction to Microarray Technology. Lecture 1 I. EXPERIMENTAL DETAILS II. ARRAY CONSTRUCTION III. IMAGE ANALYSIS Lecture
SIMS2003 Instructors:Rus Yukhananov, Alex Loguinov BWH, Harvard Medical School Introduction to Microarray Technology. Lecture 1 I. EXPERIMENTAL DETAILS II. ARRAY CONSTRUCTION III. IMAGE ANALYSIS Lecture
Functional Assessment and Clinical Outcome
 Tissue Engineering Functional Assessment and Clinical Outcome STEVEN A. GOLDSTEIN Orthopaedic Research Laboratories, Department of Orthopaedic Surgery, University of Michigan, Ann Arbor, Michigan 48109,
Tissue Engineering Functional Assessment and Clinical Outcome STEVEN A. GOLDSTEIN Orthopaedic Research Laboratories, Department of Orthopaedic Surgery, University of Michigan, Ann Arbor, Michigan 48109,
Serial Analysis of Gene Expression
 Serial Analysis of Gene Expression Cloning of Tissue-Specific Genes Using SAGE and a Novel Computational Substraction Approach. Genomic (2001) Hung-Jui Shih Outline of Presentation SAGE EST Article TPE
Serial Analysis of Gene Expression Cloning of Tissue-Specific Genes Using SAGE and a Novel Computational Substraction Approach. Genomic (2001) Hung-Jui Shih Outline of Presentation SAGE EST Article TPE
Methods of Biomaterials Testing Lesson 3-5. Biochemical Methods - Molecular Biology -
 Methods of Biomaterials Testing Lesson 3-5 Biochemical Methods - Molecular Biology - Chromosomes in the Cell Nucleus DNA in the Chromosome Deoxyribonucleic Acid (DNA) DNA has double-helix structure The
Methods of Biomaterials Testing Lesson 3-5 Biochemical Methods - Molecular Biology - Chromosomes in the Cell Nucleus DNA in the Chromosome Deoxyribonucleic Acid (DNA) DNA has double-helix structure The
ARTIFICAL VISION. Regina Leung, Michelle Ngai
 ARTIFICAL VISION http://www.sfn.org/skins/main/images/brainbriefings/big.nov.jpg http://www.ubergizmo.com/photos/2006/4/wireless-ocular-implant.jpg http://www.crystalinks.com/eyeblind.gif Regina Leung,
ARTIFICAL VISION http://www.sfn.org/skins/main/images/brainbriefings/big.nov.jpg http://www.ubergizmo.com/photos/2006/4/wireless-ocular-implant.jpg http://www.crystalinks.com/eyeblind.gif Regina Leung,
Outline. Analysis of Microarray Data. Most important design question. General experimental issues
 Outline Analysis of Microarray Data Lecture 1: Experimental Design and Data Normalization Introduction to microarrays Experimental design Data normalization Other data transformation Exercises George Bell,
Outline Analysis of Microarray Data Lecture 1: Experimental Design and Data Normalization Introduction to microarrays Experimental design Data normalization Other data transformation Exercises George Bell,
SureSilencing sirna Array Technology Overview
 SureSilencing sirna Array Technology Overview Pathway-Focused sirna-based RNA Interference Topics to be Covered Who is SuperArray? Brief Introduction to RNA Interference Challenges Facing RNA Interference
SureSilencing sirna Array Technology Overview Pathway-Focused sirna-based RNA Interference Topics to be Covered Who is SuperArray? Brief Introduction to RNA Interference Challenges Facing RNA Interference
Trasposable elements: Uses of P elements Problem set B at the end
 Trasposable elements: Uses of P elements Problem set B at the end P-elements have revolutionized the way Drosophila geneticists conduct their research. Here, we will discuss just a few of the approaches
Trasposable elements: Uses of P elements Problem set B at the end P-elements have revolutionized the way Drosophila geneticists conduct their research. Here, we will discuss just a few of the approaches
Enhancer genetics Problem set E
 Enhancer genetics Problem set E How are specific cell types generated during development? There are thought to be 10,000 different types of neurons in the human nervous system. How are all of these neural
Enhancer genetics Problem set E How are specific cell types generated during development? There are thought to be 10,000 different types of neurons in the human nervous system. How are all of these neural
Newborn screening for spinal muscular atrophy
 Newborn screening for spinal muscular atrophy Kristina Mercer, MPH, PhD ORISE Fellow Newborn Screening Translational Research Initiative Newborn Screening and Molecular Biology Branch APHL Newborn Screening
Newborn screening for spinal muscular atrophy Kristina Mercer, MPH, PhD ORISE Fellow Newborn Screening Translational Research Initiative Newborn Screening and Molecular Biology Branch APHL Newborn Screening
Where are we with gene therapy?
 Where are we with gene therapy? Session 9: Gene Based Therapies Professor Alan Boyd PFPM 7 th February 2018 Faculty of Pharmaceutical Medicine, Royal Colleges of Physicians, UK Topics to be covered Gene
Where are we with gene therapy? Session 9: Gene Based Therapies Professor Alan Boyd PFPM 7 th February 2018 Faculty of Pharmaceutical Medicine, Royal Colleges of Physicians, UK Topics to be covered Gene
3.1.4 DNA Microarray Technology
 3.1.4 DNA Microarray Technology Scientists have discovered that one of the differences between healthy and cancer is which genes are turned on in each. Scientists can compare the gene expression patterns
3.1.4 DNA Microarray Technology Scientists have discovered that one of the differences between healthy and cancer is which genes are turned on in each. Scientists can compare the gene expression patterns
Introduction to BioMEMS & Medical Microdevices DNA Microarrays and Lab-on-a-Chip Methods
 Introduction to BioMEMS & Medical Microdevices DNA Microarrays and Lab-on-a-Chip Methods Companion lecture to the textbook: Fundamentals of BioMEMS and Medical Microdevices, by Prof., http://saliterman.umn.edu/
Introduction to BioMEMS & Medical Microdevices DNA Microarrays and Lab-on-a-Chip Methods Companion lecture to the textbook: Fundamentals of BioMEMS and Medical Microdevices, by Prof., http://saliterman.umn.edu/
Exam MOL3007 Functional Genomics
 Faculty of Medicine Department of Cancer Research and Molecular Medicine Exam MOL3007 Functional Genomics Tuesday May 29 th 9.00-13.00 ECTS credits: 7.5 Number of pages (included front-page): 5 Supporting
Faculty of Medicine Department of Cancer Research and Molecular Medicine Exam MOL3007 Functional Genomics Tuesday May 29 th 9.00-13.00 ECTS credits: 7.5 Number of pages (included front-page): 5 Supporting
Chapter 15 Gene Technologies and Human Applications
 Chapter Outline Chapter 15 Gene Technologies and Human Applications Section 1: The Human Genome KEY IDEAS > Why is the Human Genome Project so important? > How do genomics and gene technologies affect
Chapter Outline Chapter 15 Gene Technologies and Human Applications Section 1: The Human Genome KEY IDEAS > Why is the Human Genome Project so important? > How do genomics and gene technologies affect
Spark Therapeutics, Inc. (Exact Name of Registrant as Specified in its Charter)
 UNITED STATES SECURITIES AND EXCHANGE COMMISSION WASHINGTON, D.C. 20549 FORM 8-K CURRENT REPORT Pursuant to Section 13 or 15(d) of the Securities Exchange Act of 1934 Date of Report (Date of earliest event
UNITED STATES SECURITIES AND EXCHANGE COMMISSION WASHINGTON, D.C. 20549 FORM 8-K CURRENT REPORT Pursuant to Section 13 or 15(d) of the Securities Exchange Act of 1934 Date of Report (Date of earliest event
Introduction to Bioinformatics. Fabian Hoti 6.10.
 Introduction to Bioinformatics Fabian Hoti 6.10. Analysis of Microarray Data Introduction Different types of microarrays Experiment Design Data Normalization Feature selection/extraction Clustering Introduction
Introduction to Bioinformatics Fabian Hoti 6.10. Analysis of Microarray Data Introduction Different types of microarrays Experiment Design Data Normalization Feature selection/extraction Clustering Introduction
Altered Expression of Genes of the Bmp/Smad and Wnt/Calcium Signaling Pathways in the Cone-only Nrl
 THE JOURNAL OF BIOLOGICAL CHEMISTRY Vol. 279, No. 40, Issue of October 1, pp. 42211 42220, 2004 2004 by The American Society for Biochemistry and Molecular Biology, Inc. Printed in U.S.A. Altered Expression
THE JOURNAL OF BIOLOGICAL CHEMISTRY Vol. 279, No. 40, Issue of October 1, pp. 42211 42220, 2004 2004 by The American Society for Biochemistry and Molecular Biology, Inc. Printed in U.S.A. Altered Expression
Microarrays and Stem Cells
 Microarray Background Information Microarrays and Stem Cells Stem cells are the building blocks that allow the body to produce new cells and repair tissues. Scientists are actively investigating the potential
Microarray Background Information Microarrays and Stem Cells Stem cells are the building blocks that allow the body to produce new cells and repair tissues. Scientists are actively investigating the potential
Genetics Lecture 21 Recombinant DNA
 Genetics Lecture 21 Recombinant DNA Recombinant DNA In 1971, a paper published by Kathleen Danna and Daniel Nathans marked the beginning of the recombinant DNA era. The paper described the isolation of
Genetics Lecture 21 Recombinant DNA Recombinant DNA In 1971, a paper published by Kathleen Danna and Daniel Nathans marked the beginning of the recombinant DNA era. The paper described the isolation of
DNA Arrays Affymetrix GeneChip System
 DNA Arrays Affymetrix GeneChip System chip scanner Affymetrix Inc. hybridization Affymetrix Inc. data analysis Affymetrix Inc. mrna 5' 3' TGTGATGGTGGGAATTGGGTCAGAAGGACTGTGGGCGCTGCC... GGAATTGGGTCAGAAGGACTGTGGC
DNA Arrays Affymetrix GeneChip System chip scanner Affymetrix Inc. hybridization Affymetrix Inc. data analysis Affymetrix Inc. mrna 5' 3' TGTGATGGTGGGAATTGGGTCAGAAGGACTGTGGGCGCTGCC... GGAATTGGGTCAGAAGGACTGTGGC
AGRO/ANSC/BIO/GENE/HORT 305 Fall, 2016 Overview of Genetics Lecture outline (Chpt 1, Genetics by Brooker) #1
 AGRO/ANSC/BIO/GENE/HORT 305 Fall, 2016 Overview of Genetics Lecture outline (Chpt 1, Genetics by Brooker) #1 - Genetics: Progress from Mendel to DNA: Gregor Mendel, in the mid 19 th century provided the
AGRO/ANSC/BIO/GENE/HORT 305 Fall, 2016 Overview of Genetics Lecture outline (Chpt 1, Genetics by Brooker) #1 - Genetics: Progress from Mendel to DNA: Gregor Mendel, in the mid 19 th century provided the
The Two-Hybrid System
 Encyclopedic Reference of Genomics and Proteomics in Molecular Medicine The Two-Hybrid System Carolina Vollert & Peter Uetz Institut für Genetik Forschungszentrum Karlsruhe PO Box 3640 D-76021 Karlsruhe
Encyclopedic Reference of Genomics and Proteomics in Molecular Medicine The Two-Hybrid System Carolina Vollert & Peter Uetz Institut für Genetik Forschungszentrum Karlsruhe PO Box 3640 D-76021 Karlsruhe
Goals of pharmacogenomics
 Goals of pharmacogenomics Use drugs better and use better drugs! People inherit/exhibit differences in drug: Absorption Metabolism and degradation of the drug Transport of drug to the target molecule Excretion
Goals of pharmacogenomics Use drugs better and use better drugs! People inherit/exhibit differences in drug: Absorption Metabolism and degradation of the drug Transport of drug to the target molecule Excretion
Multiple choice questions (numbers in brackets indicate the number of correct answers)
 1 Multiple choice questions (numbers in brackets indicate the number of correct answers) February 1, 2013 1. Ribose is found in Nucleic acids Proteins Lipids RNA DNA (2) 2. Most RNA in cells is transfer
1 Multiple choice questions (numbers in brackets indicate the number of correct answers) February 1, 2013 1. Ribose is found in Nucleic acids Proteins Lipids RNA DNA (2) 2. Most RNA in cells is transfer
3. human genomics clone genes associated with genetic disorders. 4. many projects generate ordered clones that cover genome
 Lectures 30 and 31 Genome analysis I. Genome analysis A. two general areas 1. structural 2. functional B. genome projects a status report 1. 1 st sequenced: several viral genomes 2. mitochondria and chloroplasts
Lectures 30 and 31 Genome analysis I. Genome analysis A. two general areas 1. structural 2. functional B. genome projects a status report 1. 1 st sequenced: several viral genomes 2. mitochondria and chloroplasts
Gene replacement therapy in the retinal degeneration slow (rds) mouse: the effect on retinal degeneration following partial transduction of the retina
 2001 Oxford University Press Human Molecular Genetics, 2001, Vol. 10, No. 21 2353 2361 Gene replacement therapy in the retinal degeneration slow (rds) mouse: the effect on retinal degeneration following
2001 Oxford University Press Human Molecular Genetics, 2001, Vol. 10, No. 21 2353 2361 Gene replacement therapy in the retinal degeneration slow (rds) mouse: the effect on retinal degeneration following
EECS730: Introduction to Bioinformatics
 EECS730: Introduction to Bioinformatics Lecture 14: Microarray Some slides were adapted from Dr. Luke Huan (University of Kansas), Dr. Shaojie Zhang (University of Central Florida), and Dr. Dong Xu and
EECS730: Introduction to Bioinformatics Lecture 14: Microarray Some slides were adapted from Dr. Luke Huan (University of Kansas), Dr. Shaojie Zhang (University of Central Florida), and Dr. Dong Xu and
Chapter 20: Biotechnology
 Name Period The AP Biology exam has reached into this chapter for essay questions on a regular basis over the past 15 years. Student responses show that biotechnology is a difficult topic. This chapter
Name Period The AP Biology exam has reached into this chapter for essay questions on a regular basis over the past 15 years. Student responses show that biotechnology is a difficult topic. This chapter
Whole Transcriptome Analysis of Illumina RNA- Seq Data. Ryan Peters Field Application Specialist
 Whole Transcriptome Analysis of Illumina RNA- Seq Data Ryan Peters Field Application Specialist Partek GS in your NGS Pipeline Your Start-to-Finish Solution for Analysis of Next Generation Sequencing Data
Whole Transcriptome Analysis of Illumina RNA- Seq Data Ryan Peters Field Application Specialist Partek GS in your NGS Pipeline Your Start-to-Finish Solution for Analysis of Next Generation Sequencing Data
Analysis of Microarray Data
 Analysis of Microarray Data Lecture 1: Experimental Design and Data Normalization George Bell, Ph.D. Senior Bioinformatics Scientist Bioinformatics and Research Computing Whitehead Institute Outline Introduction
Analysis of Microarray Data Lecture 1: Experimental Design and Data Normalization George Bell, Ph.D. Senior Bioinformatics Scientist Bioinformatics and Research Computing Whitehead Institute Outline Introduction
Supporting Information
 Supporting Information Ho et al. 1.173/pnas.81288816 SI Methods Sequences of shrna hairpins: Brg shrna #1: ccggcggctcaagaaggaagttgaactcgagttcaacttccttcttgacgnttttg (TRCN71383; Open Biosystems). Brg shrna
Supporting Information Ho et al. 1.173/pnas.81288816 SI Methods Sequences of shrna hairpins: Brg shrna #1: ccggcggctcaagaaggaagttgaactcgagttcaacttccttcttgacgnttttg (TRCN71383; Open Biosystems). Brg shrna
Answers to additional linkage problems.
 Spring 2013 Biology 321 Answers to Assignment Set 8 Chapter 4 http://fire.biol.wwu.edu/trent/trent/iga_10e_sm_chapter_04.pdf Answers to additional linkage problems. Problem -1 In this cell, there two copies
Spring 2013 Biology 321 Answers to Assignment Set 8 Chapter 4 http://fire.biol.wwu.edu/trent/trent/iga_10e_sm_chapter_04.pdf Answers to additional linkage problems. Problem -1 In this cell, there two copies
Identifying Candidate Informative Genes for Biomarker Prediction of Liver Cancer
 Identifying Candidate Informative Genes for Biomarker Prediction of Liver Cancer Nagwan M. Abdel Samee 1, Nahed H. Solouma 2, Mahmoud Elhefnawy 3, Abdalla S. Ahmed 4, Yasser M. Kadah 5 1 Computer Engineering
Identifying Candidate Informative Genes for Biomarker Prediction of Liver Cancer Nagwan M. Abdel Samee 1, Nahed H. Solouma 2, Mahmoud Elhefnawy 3, Abdalla S. Ahmed 4, Yasser M. Kadah 5 1 Computer Engineering
Supplementary Figure 1
 Supplementary Figure 1 Supplementary Figure 1: Vector maps of TRMPV and TRMPVIR variants. Many derivatives of TRMPV have been generated and tested. Unless otherwise noted, experiments in this paper use
Supplementary Figure 1 Supplementary Figure 1: Vector maps of TRMPV and TRMPVIR variants. Many derivatives of TRMPV have been generated and tested. Unless otherwise noted, experiments in this paper use
M Keramatipour 2. M Keramatipour 1. M Keramatipour 4. M Keramatipour 3. M Keramatipour 5. M Keramatipour
 Molecular Cloning Methods Mohammad Keramatipour MD, PhD keramatipour@tums.ac.ir Outline DNA recombinant technology DNA cloning co Cell based PCR PCR-based Some application of DNA cloning Genomic libraries
Molecular Cloning Methods Mohammad Keramatipour MD, PhD keramatipour@tums.ac.ir Outline DNA recombinant technology DNA cloning co Cell based PCR PCR-based Some application of DNA cloning Genomic libraries
TRANSGENIC TECHNOLOGIES: Gene-targeting
 TRANSGENIC TECHNOLOGIES: Gene-targeting Reverse Genetics Wild-type Bmp7 -/- Forward Genetics Phenotype Gene or Mutations First Molecular Analysis Second Reverse Genetics Gene Phenotype or Molecular Analysis
TRANSGENIC TECHNOLOGIES: Gene-targeting Reverse Genetics Wild-type Bmp7 -/- Forward Genetics Phenotype Gene or Mutations First Molecular Analysis Second Reverse Genetics Gene Phenotype or Molecular Analysis
We are committed to translating ground-breaking science into genomic therapies that transform patients lives
 We are committed to translating ground-breaking science into genomic therapies that transform patients lives Liver-based expression of the human alpha-galactosidase A gene (GLA) in a murine Fabry model
We are committed to translating ground-breaking science into genomic therapies that transform patients lives Liver-based expression of the human alpha-galactosidase A gene (GLA) in a murine Fabry model
Construction of plant complementation vector and generation of transgenic plants
 MATERIAL S AND METHODS Plant materials and growth conditions Arabidopsis ecotype Columbia (Col0) was used for this study. SALK_072009, SALK_076309, and SALK_027645 were obtained from the Arabidopsis Biological
MATERIAL S AND METHODS Plant materials and growth conditions Arabidopsis ecotype Columbia (Col0) was used for this study. SALK_072009, SALK_076309, and SALK_027645 were obtained from the Arabidopsis Biological
PTH Effects Bone Healing and Vasculogenesis in Calvaria Bone Allograft Model
 PTH Effects Bone Healing and Vasculogenesis in Calvaria Bone Allograft Model Dmitriy Sheyn, PhD 1, Doron Cohn-Yakubovich, BSc 2, Ilan Kallai, MSc 2, Susan Su, MD 1, Xiaoyu Da, MSc 1, Gadi Pelled, DMD,
PTH Effects Bone Healing and Vasculogenesis in Calvaria Bone Allograft Model Dmitriy Sheyn, PhD 1, Doron Cohn-Yakubovich, BSc 2, Ilan Kallai, MSc 2, Susan Su, MD 1, Xiaoyu Da, MSc 1, Gadi Pelled, DMD,
Proteomics And Cancer Biomarker Discovery. Dr. Zahid Khan Institute of chemical Sciences (ICS) University of Peshawar. Overview. Cancer.
 Proteomics And Cancer Biomarker Discovery Dr. Zahid Khan Institute of chemical Sciences (ICS) University of Peshawar Overview Proteomics Cancer Aims Tools Data Base search Challenges Summary 1 Overview
Proteomics And Cancer Biomarker Discovery Dr. Zahid Khan Institute of chemical Sciences (ICS) University of Peshawar Overview Proteomics Cancer Aims Tools Data Base search Challenges Summary 1 Overview
American Society of Cytopathology Core Curriculum in Molecular Biology
 American Society of Cytopathology Core Curriculum in Molecular Biology American Society of Cytopathology Core Curriculum in Molecular Biology Chapter 3 Molecular Techniques Separation and Detection, Part
American Society of Cytopathology Core Curriculum in Molecular Biology American Society of Cytopathology Core Curriculum in Molecular Biology Chapter 3 Molecular Techniques Separation and Detection, Part
Functional Genomics in Plants
 Functional Genomics in Plants Jeffrey L Bennetzen, Purdue University, West Lafayette, Indiana, USA Functional genomics refers to a suite of genetic technologies that will contribute to a comprehensive
Functional Genomics in Plants Jeffrey L Bennetzen, Purdue University, West Lafayette, Indiana, USA Functional genomics refers to a suite of genetic technologies that will contribute to a comprehensive
DNA Transcription. Visualizing Transcription. The Transcription Process
 DNA Transcription By: Suzanne Clancy, Ph.D. 2008 Nature Education Citation: Clancy, S. (2008) DNA transcription. Nature Education 1(1) If DNA is a book, then how is it read? Learn more about the DNA transcription
DNA Transcription By: Suzanne Clancy, Ph.D. 2008 Nature Education Citation: Clancy, S. (2008) DNA transcription. Nature Education 1(1) If DNA is a book, then how is it read? Learn more about the DNA transcription
Learning Objectives :
 Learning Objectives : Understand the basic differences between genomic and cdna libraries Understand how genomic libraries are constructed Understand the purpose for having overlapping DNA fragments in
Learning Objectives : Understand the basic differences between genomic and cdna libraries Understand how genomic libraries are constructed Understand the purpose for having overlapping DNA fragments in
Protein-Protein-Interaction Networks. Ulf Leser, Samira Jaeger
 Protein-Protein-Interaction Networks Ulf Leser, Samira Jaeger SHK Stelle frei Ab 1.9.2015, 2 Jahre, 41h/Monat Verbundprojekt MaptTorNet: Pankreatische endokrine Tumore Insb. statistische Aufbereitung und
Protein-Protein-Interaction Networks Ulf Leser, Samira Jaeger SHK Stelle frei Ab 1.9.2015, 2 Jahre, 41h/Monat Verbundprojekt MaptTorNet: Pankreatische endokrine Tumore Insb. statistische Aufbereitung und
Lecture 20: Drosophila melanogaster
 Lecture 20: Drosophila melanogaster Model organisms Polytene chromosome Life cycle P elements and transformation Embryogenesis Read textbook: 732-744; Fig. 20.4; 20.10; 20.15-26 www.mhhe.com/hartwell3
Lecture 20: Drosophila melanogaster Model organisms Polytene chromosome Life cycle P elements and transformation Embryogenesis Read textbook: 732-744; Fig. 20.4; 20.10; 20.15-26 www.mhhe.com/hartwell3
ALSO: look at figure 5-11 showing exonintron structure of the beta globin gene
 S08 Biology 205 6/4/08 Reading Assignment Chapter 7: From DNA to Protein: How cells read the genome pg 237-243 on exons and introns (you are not responsible for the biochemistry of splicing: figures 7-15,16
S08 Biology 205 6/4/08 Reading Assignment Chapter 7: From DNA to Protein: How cells read the genome pg 237-243 on exons and introns (you are not responsible for the biochemistry of splicing: figures 7-15,16
The C-value paradox. PHAR2811: Genome Organisation. Is there too much DNA? C o t plots. What do these life forms have in common?
 PHAR2811: enome Organisation Synopsis: -value paradox, different classes of DA, repetitive DA and disease. If protein-coding portions of the human genome make up only 1.5% what is the rest doing? The -value
PHAR2811: enome Organisation Synopsis: -value paradox, different classes of DA, repetitive DA and disease. If protein-coding portions of the human genome make up only 1.5% what is the rest doing? The -value
Use of Drosophila Melanogaster as a Model System in the Study of Human Sodium- Dependent Multivitamin Transporter. Michael Brinton BIOL 230W.
 Use of Drosophila Melanogaster as a Model System in the Study of Human Sodium- Dependent Multivitamin Transporter Michael Brinton BIOL 230W.001 28 October 2013 TA: Sashi Gollapudi Introduction Many human
Use of Drosophila Melanogaster as a Model System in the Study of Human Sodium- Dependent Multivitamin Transporter Michael Brinton BIOL 230W.001 28 October 2013 TA: Sashi Gollapudi Introduction Many human
7.06 Problem Set #3, Spring 2005
 7.06 Problem Set #3, Spring 2005 1. The Drosophila compound eye is composed of about 800 units called ommatidia. Each ommatidium contains eight photoreceptor neurons (R1 through R8), which develop in a
7.06 Problem Set #3, Spring 2005 1. The Drosophila compound eye is composed of about 800 units called ommatidia. Each ommatidium contains eight photoreceptor neurons (R1 through R8), which develop in a
CRX ChIP-seq reveals the cis-regulatory architecture of mouse photoreceptors
 Research CRX ChIP-seq reveals the cis-regulatory architecture of mouse photoreceptors Joseph C. Corbo, 1,6,7 Karen A. Lawrence, 1,6 Marcus Karlstetter, 2 Connie A. Myers, 1 Musa Abdelaziz, 1 William Dirkes,
Research CRX ChIP-seq reveals the cis-regulatory architecture of mouse photoreceptors Joseph C. Corbo, 1,6,7 Karen A. Lawrence, 1,6 Marcus Karlstetter, 2 Connie A. Myers, 1 Musa Abdelaziz, 1 William Dirkes,
Phase 1 SMA Type 2 Trial Initiation and Study Design. December 2017
 Phase 1 SMA Type 2 Trial Initiation and Study Design December 2017 Disclaimers This presentation contains forward-looking statements, including statements about: the timing, progress and results of preclinical
Phase 1 SMA Type 2 Trial Initiation and Study Design December 2017 Disclaimers This presentation contains forward-looking statements, including statements about: the timing, progress and results of preclinical
A Propagation-based Algorithm for Inferring Gene-Disease Associations
 A Propagation-based Algorithm for Inferring Gene-Disease Associations Oron Vanunu Roded Sharan Abstract: A fundamental challenge in human health is the identification of diseasecausing genes. Recently,
A Propagation-based Algorithm for Inferring Gene-Disease Associations Oron Vanunu Roded Sharan Abstract: A fundamental challenge in human health is the identification of diseasecausing genes. Recently,
INNOVATIVE TECHNOLOGY IN MEDICINE: EXPERIENCE OF POLAND AND UKRAINE
 LUBLIN SCIENCE AND TECHNOLOGY PARK S.A. International research and practice conference INNOVATIVE TECHNOLOGY IN MEDICINE: EXPERIENCE OF POLAND AND UKRAINE April 28 29, 2017 Lublin, Republic of Poland 2017
LUBLIN SCIENCE AND TECHNOLOGY PARK S.A. International research and practice conference INNOVATIVE TECHNOLOGY IN MEDICINE: EXPERIENCE OF POLAND AND UKRAINE April 28 29, 2017 Lublin, Republic of Poland 2017
1. Introduction. 1 Page 1
 1. INTRODUCTION Osteoblast differentiation is the key component of the bone formation, growth and remodeling. During this incompletely understood process a subpopulation of multipotential mesenchymal stem
1. INTRODUCTION Osteoblast differentiation is the key component of the bone formation, growth and remodeling. During this incompletely understood process a subpopulation of multipotential mesenchymal stem
Exploring the Genetic Basis of Congenital Heart Defects
 Exploring the Genetic Basis of Congenital Heart Defects Sanjay Siddhanti Jordan Hannel Vineeth Gangaram szsiddh@stanford.edu jfhannel@stanford.edu vineethg@stanford.edu 1 Introduction The Human Genome
Exploring the Genetic Basis of Congenital Heart Defects Sanjay Siddhanti Jordan Hannel Vineeth Gangaram szsiddh@stanford.edu jfhannel@stanford.edu vineethg@stanford.edu 1 Introduction The Human Genome
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature10163 Supplementary Table 1 Efficiency of vector construction. Process wells recovered efficiency (%) Recombineering* 480 461 96 Intermediate plasmids 461 381 83 Recombineering efficiency
doi:10.1038/nature10163 Supplementary Table 1 Efficiency of vector construction. Process wells recovered efficiency (%) Recombineering* 480 461 96 Intermediate plasmids 461 381 83 Recombineering efficiency
NAME TA SEC Problem Set 4 FRIDAY October 15, Answers to this problem set must be inserted into the box outside
 MIT Biology Department 7.012: Introductory Biology - Fall 2004 Instructors: Professor Eric Lander, Professor Robert A. Weinberg, Dr. Claudette Gardel NAME TA SEC 7.012 Problem Set 4 FRIDAY October 15,
MIT Biology Department 7.012: Introductory Biology - Fall 2004 Instructors: Professor Eric Lander, Professor Robert A. Weinberg, Dr. Claudette Gardel NAME TA SEC 7.012 Problem Set 4 FRIDAY October 15,
Microarray Analysis of Gene Expression in Huntington's Disease Peripheral Blood - a Platform Comparison. CodeLink compatible
 Microarray Analysis of Gene Expression in Huntington's Disease Peripheral Blood - a Platform Comparison CodeLink compatible Microarray Analysis of Gene Expression in Huntington's Disease Peripheral Blood
Microarray Analysis of Gene Expression in Huntington's Disease Peripheral Blood - a Platform Comparison CodeLink compatible Microarray Analysis of Gene Expression in Huntington's Disease Peripheral Blood
Le proteine regolative variano nei vari tipi cellulari e in funzione degli stimoli ambientali
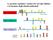 Le proteine regolative variano nei vari tipi cellulari e in funzione degli stimoli ambientali Tipo cellulare 1 Tipo cellulare 2 Tipo cellulare 3 DNA-protein Crosslink Lisi Frammentazione Immunopurificazione
Le proteine regolative variano nei vari tipi cellulari e in funzione degli stimoli ambientali Tipo cellulare 1 Tipo cellulare 2 Tipo cellulare 3 DNA-protein Crosslink Lisi Frammentazione Immunopurificazione
measuring gene expression December 5, 2017
 measuring gene expression December 5, 2017 transcription a usually short-lived RNA copy of the DNA is created through transcription RNA is exported to the cytoplasm to encode proteins some types of RNA
measuring gene expression December 5, 2017 transcription a usually short-lived RNA copy of the DNA is created through transcription RNA is exported to the cytoplasm to encode proteins some types of RNA
REAL TIME PCR USING SYBR GREEN
 REAL TIME PCR USING SYBR GREEN 1 THE PROBLEM NEED TO QUANTITATE DIFFERENCES IN mrna EXPRESSION SMALL AMOUNTS OF mrna LASER CAPTURE SMALL AMOUNTS OF TISSUE PRIMARY CELLS PRECIOUS REAGENTS 2 THE PROBLEM
REAL TIME PCR USING SYBR GREEN 1 THE PROBLEM NEED TO QUANTITATE DIFFERENCES IN mrna EXPRESSION SMALL AMOUNTS OF mrna LASER CAPTURE SMALL AMOUNTS OF TISSUE PRIMARY CELLS PRECIOUS REAGENTS 2 THE PROBLEM
Intravitreal and sub-retinal injections of plasmid DNA and electroporation in P0 pups
 Intravitreal and sub-retinal injections of plasmid DNA and electroporation in P0 pups Protocol modified from: Retinal Gene Delivery by raav and DNA Electroporation, Aditya Venkatesh et all, Current Protocols
Intravitreal and sub-retinal injections of plasmid DNA and electroporation in P0 pups Protocol modified from: Retinal Gene Delivery by raav and DNA Electroporation, Aditya Venkatesh et all, Current Protocols
7 Gene Isolation and Analysis of Multiple
 Genetic Techniques for Biological Research Corinne A. Michels Copyright q 2002 John Wiley & Sons, Ltd ISBNs: 0-471-89921-6 (Hardback); 0-470-84662-3 (Electronic) 7 Gene Isolation and Analysis of Multiple
Genetic Techniques for Biological Research Corinne A. Michels Copyright q 2002 John Wiley & Sons, Ltd ISBNs: 0-471-89921-6 (Hardback); 0-470-84662-3 (Electronic) 7 Gene Isolation and Analysis of Multiple
Genetic testing of Tay-Sachs disease by PCR and Restriction Digest
 Genetic testing of Tay-Sachs disease by PCR and Restriction Digest ESC102-PRA0103 Submitted to: Elizabeth Berndl 18 February, 2009 Submitted by: Laila Hulbert (996625077) Maria Yancheva (996742173) Scientific
Genetic testing of Tay-Sachs disease by PCR and Restriction Digest ESC102-PRA0103 Submitted to: Elizabeth Berndl 18 February, 2009 Submitted by: Laila Hulbert (996625077) Maria Yancheva (996742173) Scientific
History of the CFTR chase
 Module II: Molecular and cellular phenotype Discuss the history of the gene. When was the gene discovered? How was the gene cloned? (Be brief.) Discuss the cellular phenotype. What cells or tissues are
Module II: Molecular and cellular phenotype Discuss the history of the gene. When was the gene discovered? How was the gene cloned? (Be brief.) Discuss the cellular phenotype. What cells or tissues are
University of Pennsylvania Perelman School of Medicine High Throughput Screening Core
 University of Pennsylvania Perelman School of Medicine High Throughput Screening Core Sara Cherry, Ph. D. David C. Schultz, Ph. D. dschultz@mail.med.upenn.edu (215) 573 9641 67 John Morgan Building Mission
University of Pennsylvania Perelman School of Medicine High Throughput Screening Core Sara Cherry, Ph. D. David C. Schultz, Ph. D. dschultz@mail.med.upenn.edu (215) 573 9641 67 John Morgan Building Mission
Functional Genomics Overview RORY STARK PRINCIPAL BIOINFORMATICS ANALYST CRUK CAMBRIDGE INSTITUTE 18 SEPTEMBER 2017
 Functional Genomics Overview RORY STARK PRINCIPAL BIOINFORMATICS ANALYST CRUK CAMBRIDGE INSTITUTE 18 SEPTEMBER 2017 Agenda What is Functional Genomics? RNA Transcription/Gene Expression Measuring Gene
Functional Genomics Overview RORY STARK PRINCIPAL BIOINFORMATICS ANALYST CRUK CAMBRIDGE INSTITUTE 18 SEPTEMBER 2017 Agenda What is Functional Genomics? RNA Transcription/Gene Expression Measuring Gene
A. AIPL1-Leber congenital amaurosis (LCA4). Accounting for approximately 5% of LCA cases, mutations
 IV. Online Supplement (Proof of concept) A. AIPL1-Leber congenital amaurosis (LCA4). Accounting for approximately 5% of LCA cases, mutations in aryl hydrocarbon receptor-interacting protein-like 1 (AIPL1)
IV. Online Supplement (Proof of concept) A. AIPL1-Leber congenital amaurosis (LCA4). Accounting for approximately 5% of LCA cases, mutations in aryl hydrocarbon receptor-interacting protein-like 1 (AIPL1)
Stem Cell Services. Driving Innovation for Stem Cell Researchers
 Driving Innovation for Stem Cell Researchers Stem Cell Services Partner with us and have access to the most advanced and comprehensive stem cell services available today. 675 W. Kendall St. Cambridge,
Driving Innovation for Stem Cell Researchers Stem Cell Services Partner with us and have access to the most advanced and comprehensive stem cell services available today. 675 W. Kendall St. Cambridge,
Stefano Monti. Workshop Format
 Gad Getz Stefano Monti Michael Reich {gadgetz,smonti,mreich}@broad.mit.edu http://www.broad.mit.edu/~smonti/aws Broad Institute of MIT & Harvard October 18-20, 2006 Cambridge, MA Workshop Format Morning
Gad Getz Stefano Monti Michael Reich {gadgetz,smonti,mreich}@broad.mit.edu http://www.broad.mit.edu/~smonti/aws Broad Institute of MIT & Harvard October 18-20, 2006 Cambridge, MA Workshop Format Morning
DNA Microarray Data Oligonucleotide Arrays
 DNA Microarray Data Oligonucleotide Arrays Sandrine Dudoit, Robert Gentleman, Rafael Irizarry, and Yee Hwa Yang Bioconductor Short Course 2003 Copyright 2002, all rights reserved Biological question Experimental
DNA Microarray Data Oligonucleotide Arrays Sandrine Dudoit, Robert Gentleman, Rafael Irizarry, and Yee Hwa Yang Bioconductor Short Course 2003 Copyright 2002, all rights reserved Biological question Experimental
Opportunities for industry/smes in EU-funded health research
 Opportunities for industry/smes in EU-funded health research Stéphane Hogan, M.Sc, MBA Head of Unit Applied Genomics and Biotechnology for Health DG Research - European Commission 1 Paris, 31 May 2005
Opportunities for industry/smes in EU-funded health research Stéphane Hogan, M.Sc, MBA Head of Unit Applied Genomics and Biotechnology for Health DG Research - European Commission 1 Paris, 31 May 2005
Calculation of Spot Reliability Evaluation Scores (SRED) for DNA Microarray Data
 Protocol Calculation of Spot Reliability Evaluation Scores (SRED) for DNA Microarray Data Kazuro Shimokawa, Rimantas Kodzius, Yonehiro Matsumura, and Yoshihide Hayashizaki This protocol was adapted from
Protocol Calculation of Spot Reliability Evaluation Scores (SRED) for DNA Microarray Data Kazuro Shimokawa, Rimantas Kodzius, Yonehiro Matsumura, and Yoshihide Hayashizaki This protocol was adapted from
Highly efficient genome engineering in flowering plants ~ Development of a rapid method to knockout genes in Arabidopsis thaliana ~
 Highly efficient genome engineering in flowering plants ~ Development of a rapid method to knockout genes in Arabidopsis thaliana ~ December 5, 2016 Plant biologists at ITbM, Nagoya University have developed
Highly efficient genome engineering in flowering plants ~ Development of a rapid method to knockout genes in Arabidopsis thaliana ~ December 5, 2016 Plant biologists at ITbM, Nagoya University have developed
Real-Time PCR Workshop Gene Expression. Applications Absolute and Relative Quantitation
 Real-Time PCR Workshop Gene Expression Applications Absolute and Relative Quantitation Absolute Quantitation Easy to understand the data, difficult to develop/qualify the standards Relative Quantitation
Real-Time PCR Workshop Gene Expression Applications Absolute and Relative Quantitation Absolute Quantitation Easy to understand the data, difficult to develop/qualify the standards Relative Quantitation
A Level. A Level Biology. DNA Technology Questions. AQA, OCR, Edexcel. Name: Total Marks: Page 1
 AQA, OCR, Edexcel A Level A Level Biology DNA Technology Questions Name: Total Marks: Page 1 Q1.(a) (i) A mutation of a tumour suppressor gene can result in the formation of a tumour. Explain how.........(2)
AQA, OCR, Edexcel A Level A Level Biology DNA Technology Questions Name: Total Marks: Page 1 Q1.(a) (i) A mutation of a tumour suppressor gene can result in the formation of a tumour. Explain how.........(2)
REGISTRATION DOCUMENT FOR RECOMBINANT DNA RESEARCH
 EHRS Date Received: Reg. Doc. No.: REGISTRATION DOCUMENT FOR RECOMBINANT DNA RESEARCH Principal Investigator: Penn ID#: Position Title: School: Department: Mailing Address: Mail Code: Telephone: FAX: E-mail:
EHRS Date Received: Reg. Doc. No.: REGISTRATION DOCUMENT FOR RECOMBINANT DNA RESEARCH Principal Investigator: Penn ID#: Position Title: School: Department: Mailing Address: Mail Code: Telephone: FAX: E-mail:
COMPASS. Cure SMA Awards 10 New Grants in Basic and Clinical Care Research. A Publication Dedicated to Research Updates SPRING 2016
 BASIC RESEARCH DRUG DISCOVERY CLINICAL TRIALS CLINICAL CARE RESEARCHER MEETING LATEST ADVANCES COMPASS A Publication Dedicated to Research Updates SPRING 2016 Cure SMA Awards 10 New Grants in Basic and
BASIC RESEARCH DRUG DISCOVERY CLINICAL TRIALS CLINICAL CARE RESEARCHER MEETING LATEST ADVANCES COMPASS A Publication Dedicated to Research Updates SPRING 2016 Cure SMA Awards 10 New Grants in Basic and
