Activation of a Floral Homeotic Gene in Arabidopsis
|
|
|
- Amanda Craig
- 6 years ago
- Views:
Transcription
1 Activation of a Floral Homeotic Gene in Arabidopsis Busch, M.A., Bomblies, K., Weigel, D. kuleuven-kortrijk.be Simon Olthof Louie Christodoulidis 1
2 In Arabidopsis flowers, appearance is based, in part, upon the expression of floral meristemindentity genes and their interaction with homeotic genes. 2
3 What s a meristem-identity gene? - A gene which codes for a transcription factor that regulates other genes that contribute to phenotype. What s a homeotic gene? - A gene that controls the pattern of body formation during development, contributing to the overall phenotype. 3
4 What they knew at the time Floral meristem-identity genes which contribute to the appearance of Arabidopsis include APETALA1 (AP1), CAULIFLOWER and LEAFY (LFY). These genes are expressed early to control flower initiation throughout the floral primordium. 4
5 Growth Phases of Arabidopsis - Vegetative phase = plants grow leaves and shoots. - Reproductive phase = flowers produced. 5
6 Inactivation and overexpression experiments had been done which demonstrated the control these genes had on flower initiation. For example, loss-of-function experiments which used lfy mutants saw leaves with associated shoots instead of early flowers. wild type lfy mutant 6
7 What more they knew at the time Homeotic genes are expressed in distinct domains of the flower and specify the identity of the floral organs which include sepals, petals, stamens and carpels. Section C is specified by the AGAMOUS (AG) gene. 7
8 Loss-of-function experiments demonstrated how the genes of the ABC model specify floral organ identity. For example, when A function is disrupted (which is specified by the gene APETALA2), mutant phenotype plants lack sepals and petals. Wild Type ap2 mutant 8
9 9
10 10
11 How are LFY and AG related? Besides playing a part in initiating floral development, a similar study conducted by Parcy et al. (1999) found that LFY plays a role in the activation of AG. In order to demonstrate this, they developed a version of LFY with increased transcriptional-activation ability by fusing it with a strong activation domain taken from the viral transcription factor VP16 to make LFY:VP16. Question: Why use a viral transcription factor? 11
12 Answer: because viral-proten 16 (VP16) is a strong activator and can be used to amplify the effects of LFY on AGAMOUS (AG). 12
13 What they found in LFY:VP16 plants with respect to AG RNA: - detected earlier - found ectopically, meaning gene was expressed in cell types in which it is normally inactive. - produced more RNA all in comparison with wild type. - This suggests not only that AG is activated in some way by LFY, but also that AG expression is normally region-specific. - Parcy et al. (1999) state that coregulators interact with LFY protein to restrict AG expression. 13
14 So if they knew all this, what was left to prove? 14
15 What they didn t know at the time was the directness of interaction which in the case of this study, was between LFY and the AGAMOUS (AG) gene of the ABC model. 15
16 In order to better understand the interaction between LFY and AG, they first identified enhancers responsible for AG activation. why? 16
17 They isolated enhancers to see if and how they interact with the LFY protein produced by the LFY gene. Does LFY protein bind to the AG gene or does it bind elsewhere? 17
18 To illustrate the LFY AG interaction They created a translation fusion of AGAMOUS (AG) to β-glucuronidase (GUS). This is where a gene is constructed to produce a protein that can be tagged in order to locate where in the cell it is produced and at what levels. It was found (regarding AG) that the second intron (of AG) was required for expression of this chimeric (artificial) gene because it contained enhancers necessary and sufficient for wild type AG expression. It was also found that this intron was spaced mostly within a 3kb Hind-III restriction fragment. 18
19 This was demonstrated using the KB9 fragment, which includes the whole Hind-III fragment: When arranged in the forward position ( 3 closest to promoter), transformants showed intermediate-to-zero staining. Oppositely, when arranged in the reverse position (5 closest to promoter) transformants showed weak-to-strong staining. weak intermediate strong 19
20 This was demonstrated using the KB9 fragment, which includes the whole Hind-III fragment: When arranged in the forward position ( 3 closest to promoter), transformants showed intermediate-to-zero staining. Oppositely, when arranged in the reverse position (5 closest to promoter) transformants showed weak-to-strong staining. 20
21 Oppositely, when the Hind-III fragment was removed, there was no response of the artificial gene. This was with the use of LFY:VP16 to amplify the interaction. 21
22 At the same time, the KB9 reporter also illustrated how AG responds to LFY, suggesting AG contains LFYresponsive enhancer sequences. In lfy mutants, GUS expression was delayed (and reduced) as indicated by the stain: 22
23 In LFY:VP16 subjects, GUS was expressed earlier (than normal) and ectopically, illustrated by the stain: 23
24 Ag-Enhancers Deletion analysis of 3-kb fragment revealed two enhancers that drive the GUS expression in Arabidopsis Flowers Combined these two enhancers could drive GUS expression in young wild type flowers in a similar way to those under the control of the 3kb fragment
25 Ag-Enhancers The enhancers KB14 and KB31 both are expressed in the centre of young flowers. These enhancers are in the 5' end and 3' of the fragment respectively This is very similar to expected endogenous AG expression in the early stages of flower development KB14-wild type KB9-wild type KB31-wild type
26 Two main findings from studying LFY mutants Ag-Enhancers One, the appearance of what appears to be enhancers that help to mediate AG activation independent of LFY As evident from how expression of smaller reporters is reduced more than full length reporters. What does this say about LFY mediated activation of AG?
27 Two main findings from studying LFY mutants Ag-Enhancers One, the appearance of what appears to be enhancers that help to mediate AG activation independent of LFY As evident form how expression of smaller reporters is reduced more than full length reporters. What does this say about LFY mediated activation of AG? - That other factors play a role in AG activation as well.
28 Ag-Enhancers Second, that the reporters appear to have a regulatory role in AG expression. Several reporters are only expressed in certain regions even when exposed to the activated protein LFY:VP16 Particularly evident in KB18's and KB14's Outer whorls, Floral stem, and Pedical tissues KB14 KB18
29 The 3' Enhancer was chosen for as a main focus, Why? Ag-Enhancers Analysis
30 Ag-Enhancers Analysis The 3' Enhancer was chosen for as a main focus, Why? - Size, the 3' enhancers were consideraly smaller than those of the 5' end ones - Their GUS activity was reduced in older flowers which would make them easier to interpret their expression
31 Ag-Enhancers Analysis Using LFY:VP16 activated protein, certain reporters which normally either have very low expression or no expression in wild type can be detected in the LFY:VP16 As these reporters were expressed in the presence of LFY:VP16 it must mean that there is an interaction between the reporter and the protein Using KB30 as a guide the responsive element for LFY:VP16 was mapped.
32 Finding the Binding Sites The binding sites for LFY were found by performing a immunoprecipitation of the protein-dna complex This done by using an antibody which binds to the transcription factor protein LFY and allows for the isolation of the DNA that it's bound to which can then be sequenced.
33 Finding the Binding Sites From a electromoblity assay it was found that using a 160 bp fragment from the 3' enhancer that the protein bound to the DNA at two binding sites As the DNA isolated from the immunoprecipitation only 31 bp long looking for common motifs which resemble LFY binding sites in AP1 it was possible to isolated the two different binding sites in AG's 3' enhancer
34 LFY binding sites Using this method, it was found that LFY did bind to AG at two sites named AG I AND AGII LFY binds to both sites with to an equal levels. To test whether these binding sites are needed to express the 3' enhancer these binding sites were deleted, to see it's effect on GUS expression.
35 LFY binding sites The first experiment was to the delete the AG1 site. This resulted in the KB45 mutant, this mutant has no detectable GUS expression in the Wild type and has a reduced level of expression in the LFY:VP16 Wild type KB30-LFY:VP16 LFY:VP16
36 This indicates that both AG sites are needed for normal levels of in vivo 3' AG enhancer expression LFY binding sites A further double deletion was also made which both of the AG sites deleted, called KB46 In KB46 expression in both the wild type and those exposed to LFY:VP16 showed no expression of the AG enhancer. This appears to show that with out the binding sites LFY can't activate the 3' enhancer. KB30- LFY:VP16 KB46-wild type KB46-LFY:VP16
37 To prove that the lost of enhancer activity was indeed due to loss of function of the binding site, a small 2 bp mutation was induced a the beginning of the conserved region of the binding site. A similar mutation rends the binding site of AP1 non functional LFY binding sites Another mutation was also induced this time a single bp in the center of the binding site. These mutation were named m1 and m2 respectivily and in a electomobility assay, the m1 mutation did indeed appear to prevent binding of LFY, while m2 mutation had no effect.
38 To show this the case in actually plants these mutation were introduced into two plants One MX68 had the m1 mutation in both binding sites LFY binding sites MX68-Wild type KB30-FLY:VP16 MX68-FLY:VP16
39 LFY binding sites While the other mutant was given the m2 mutation in both binding sites This mutation however as expected has no effect on expression as MX 100 as a very similar level of expression as the KB 31 reporter which contains the 3' enhancer. MX100-wild type KB31-wild type
40 LFY Gene Unlike ABC and other transrciption factors LFY is not part of a gene family Only one homolog of LFY is present in the hapoloid genome What this means is that the binding sites on the AG enchancer is unlikily to bind any other protein than LFY.
41 So What did they prove? Conclusions
42 Conclusions So What did they prove? Hypothesis-That Binding of LFY is critical for Transcriptional Activation mediated by the 3' AG enhancer
43 Conclusions So What did they prove? Hypothesis-That Binding of LFY is critical for Transcriptional Activation mediated by the 3' AG enhancer How did they prove this? -First they isolated the 5' and 3' enhancers in the 3 kb fragment -Then show that the LFY made a difference to the level of expression of the the reporter, and there fore AG - Then they isolated the binding sites of the 3' enhancer - Using these they deleted them to prove that with the ability to bind to AG the gene could not be expressed. - They also proved how specific and conserved the LFY binding motif is.
44 Conclusions So What did they prove? Hypothesis-That Binding of LFY is critical for Transcriptional Activation mediated by the 3' AG enhancer How did they prove this? -First they isolated the 5' and 3' enhancers in the 3 kb fragment -Then show that the LFY made a difference to the level of expression of the the reporter, and there fore AG - Then they isolated the binding sites of the 3' enhancer - Using these they deleted them to prove that with the ability to bind to AG the gene could not be expressed. - They also proved how specific and conserved the LFY binding motif is.
45 Conclusions So What did they prove? Hypothesis-That Binding of LFY is critical for Transcriptional Activation mediated by the 3' AG enhancer How did they prove this? -First they isolated the 5' and 3' enhancers in the 3 kb fragment -Then show that the LFY made a difference to the level of expression of the the reporter, and there fore AG - Then they isolated the binding sites of the 3' enhancer - Using these they deleted them to prove that with the ability to bind to AG the gene could not be expressed. - They also proved how specific and conserved the LFY binding motif is. This makes LFY a recognized AG activator
46 Conclusions So What did they prove? Hypothesis-That Binding of LFY is critical for Transcriptional Activation mediated by the 3' AG enhancer How did they prove this? -First they isolated the 5' and 3' enhancers in the 3 kb fragment -Then show that the LFY made a difference to the level of expression of the the reporter, and there fore AG - Then they isolated the binding sites of the 3' enhancer - Using these they deleted them to prove that with the ability to bind to AG the gene could not be expressed. - They also proved how specific and conserved the IFY binding motif is. This makes LFY a recognized AG activator
47 So what Now? Conclusions
48 Conclusions So what Now? As you may remember even with out LFY some expression of AG was occurring As well IFY in certain enhancer elements was only expressed in certain regions So now that the main regulator for AG is known, it is important to find the CO-regulators that help govern the expression of AG in flowers KB9-LFY mt We now know of two possible co-regulators of LFY the region specific genes UFO(Unsual Floral Organ) and WUS(Wuschel) KB18-LFY:VP16
49 Why is this paper so important? Conclusions
50 Conclusions Why is this paper so important? -As it proved that the AG an ABC was directly mediated by LFY -Prior this it was unclear if it was a direct link or that it went through intermediates IFY ABC With the work also outline in this paper it proved that IFY bound to and regulated the expression of AG and AP1, and would likily regulated expression of all ABC genes. Nature Oct 8;395(6702): A genetic framework for floral patterning. Parcy F, Nilsson O, Busch MA, Lee I, Weigel D. The Salk Institute for Biological Studies, La Jolla, California 92037, USA.
51 References A molecular link between stem cell regulation and floral patterning in Arabidopsis. Lohmann JU, Hong RL, Hobe M, Busch MA, Parcy F, Simon R, Weigel D Plant Biology Laboratory, The Salk Institute for Biological Studies, La Jolla, CA 92037, USA. A molecular link between stem cell regulation and floral patterning in Arabidopsis. Lohmann JU, Hong RL, Hobe M, Busch MA, Parcy F, Simon R, Weigel D Plant Biology Laboratory, The Salk Institute for Biological Studies, La Jolla, CA 92037, USA.
Supplemental Data. mir156-regulated SPL Transcription. Factors Define an Endogenous Flowering. Pathway in Arabidopsis thaliana
 Cell, Volume 138 Supplemental Data mir156-regulated SPL Transcription Factors Define an Endogenous Flowering Pathway in Arabidopsis thaliana Jia-Wei Wang, Benjamin Czech, and Detlef Weigel Table S1. Interaction
Cell, Volume 138 Supplemental Data mir156-regulated SPL Transcription Factors Define an Endogenous Flowering Pathway in Arabidopsis thaliana Jia-Wei Wang, Benjamin Czech, and Detlef Weigel Table S1. Interaction
Genetics - Problem Drill 19: Dissection of Gene Function: Mutational Analysis of Model Organisms
 Genetics - Problem Drill 19: Dissection of Gene Function: Mutational Analysis of Model Organisms No. 1 of 10 1. The mouse gene knockout is based on. (A) Homologous recombination (B) Site-specific recombination
Genetics - Problem Drill 19: Dissection of Gene Function: Mutational Analysis of Model Organisms No. 1 of 10 1. The mouse gene knockout is based on. (A) Homologous recombination (B) Site-specific recombination
Floral stem cell termination involves the direct regulation of AGAMOUS by PERIANTHIA
 RESEARCH REPORT 165 Development 136, 165-1611 (29) doi:1.1242/dev.35436 Floral stem cell termination involves the direct regulation of AGAMOUS by PERIANTHIA Pradeep Das 1,2, *, Toshiro Ito 1,3, Frank Wellmer
RESEARCH REPORT 165 Development 136, 165-1611 (29) doi:1.1242/dev.35436 Floral stem cell termination involves the direct regulation of AGAMOUS by PERIANTHIA Pradeep Das 1,2, *, Toshiro Ito 1,3, Frank Wellmer
Reconstitution of floral quartets in vitro involving class B and class E floral homeotic proteins
 Published online 1 March 9 Nucleic Acids Research, 9, Vol. 37, No. 8 73 736 doi:1.193/nar/gkp19 Reconstitution of floral quartets in vitro involving class B and class E floral homeotic proteins Rainer
Published online 1 March 9 Nucleic Acids Research, 9, Vol. 37, No. 8 73 736 doi:1.193/nar/gkp19 Reconstitution of floral quartets in vitro involving class B and class E floral homeotic proteins Rainer
Lecture 2-3: Using Mutants to study Biological processes
 Lecture 2-3: Using Mutants to study Biological processes Objectives: 1. Why use mutants? 2. How are mutants isolated? 3. What important genetic analyses must be done immediately after a genetic screen
Lecture 2-3: Using Mutants to study Biological processes Objectives: 1. Why use mutants? 2. How are mutants isolated? 3. What important genetic analyses must be done immediately after a genetic screen
Genome research in eukaryotes
 Functional Genomics Genome and EST sequencing can tell us how many POTENTIAL genes are present in the genome Proteomics can tell us about proteins and their interactions The goal of functional genomics
Functional Genomics Genome and EST sequencing can tell us how many POTENTIAL genes are present in the genome Proteomics can tell us about proteins and their interactions The goal of functional genomics
Highly efficient genome engineering in flowering plants ~ Development of a rapid method to knockout genes in Arabidopsis thaliana ~
 Highly efficient genome engineering in flowering plants ~ Development of a rapid method to knockout genes in Arabidopsis thaliana ~ December 5, 2016 Plant biologists at ITbM, Nagoya University have developed
Highly efficient genome engineering in flowering plants ~ Development of a rapid method to knockout genes in Arabidopsis thaliana ~ December 5, 2016 Plant biologists at ITbM, Nagoya University have developed
An Arabidopsis F-box protein acts as a transcriptional co-factor to regulate floral development
 RESEARCH ARTICLE 1235 Development 135, 1235-1245 (2008) doi:10.1242/dev.015842 An Arabidopsis F-box protein acts as a transcriptional co-factor to regulate floral development Eunyoung Chae 1, Queenie K.-G.
RESEARCH ARTICLE 1235 Development 135, 1235-1245 (2008) doi:10.1242/dev.015842 An Arabidopsis F-box protein acts as a transcriptional co-factor to regulate floral development Eunyoung Chae 1, Queenie K.-G.
Construction of plant complementation vector and generation of transgenic plants
 MATERIAL S AND METHODS Plant materials and growth conditions Arabidopsis ecotype Columbia (Col0) was used for this study. SALK_072009, SALK_076309, and SALK_027645 were obtained from the Arabidopsis Biological
MATERIAL S AND METHODS Plant materials and growth conditions Arabidopsis ecotype Columbia (Col0) was used for this study. SALK_072009, SALK_076309, and SALK_027645 were obtained from the Arabidopsis Biological
The Two-Hybrid System
 Encyclopedic Reference of Genomics and Proteomics in Molecular Medicine The Two-Hybrid System Carolina Vollert & Peter Uetz Institut für Genetik Forschungszentrum Karlsruhe PO Box 3640 D-76021 Karlsruhe
Encyclopedic Reference of Genomics and Proteomics in Molecular Medicine The Two-Hybrid System Carolina Vollert & Peter Uetz Institut für Genetik Forschungszentrum Karlsruhe PO Box 3640 D-76021 Karlsruhe
CRISPR/Cas9 Genome Editing: Transfection Methods
 CRISPR/ Genome Editing: Transfection Methods For over 20 years Mirus Bio has developed and manufactured high performance transfection products and technologies. That expertise is now being applied to the
CRISPR/ Genome Editing: Transfection Methods For over 20 years Mirus Bio has developed and manufactured high performance transfection products and technologies. That expertise is now being applied to the
7 Gene Isolation and Analysis of Multiple
 Genetic Techniques for Biological Research Corinne A. Michels Copyright q 2002 John Wiley & Sons, Ltd ISBNs: 0-471-89921-6 (Hardback); 0-470-84662-3 (Electronic) 7 Gene Isolation and Analysis of Multiple
Genetic Techniques for Biological Research Corinne A. Michels Copyright q 2002 John Wiley & Sons, Ltd ISBNs: 0-471-89921-6 (Hardback); 0-470-84662-3 (Electronic) 7 Gene Isolation and Analysis of Multiple
The Genetic Basis of Development
 Chapter 21 The Genetic Basis of Development Overview: From Single Cell to Multicellular Organism The application of genetic analysis and DNA technology Has revolutionized the study of development PowerPoint
Chapter 21 The Genetic Basis of Development Overview: From Single Cell to Multicellular Organism The application of genetic analysis and DNA technology Has revolutionized the study of development PowerPoint
Introducing new DNA into the genome requires cloning the donor sequence, delivery of the cloned DNA into the cell, and integration into the genome.
 Key Terms Chapter 32: Genetic Engineering Cloning describes propagation of a DNA sequence by incorporating it into a hybrid construct that can be replicated in a host cell. A cloning vector is a plasmid
Key Terms Chapter 32: Genetic Engineering Cloning describes propagation of a DNA sequence by incorporating it into a hybrid construct that can be replicated in a host cell. A cloning vector is a plasmid
A subclass of HSP70s regulate development and abiotic stress responses in Arabidopsis thaliana
 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 Journal of Plant Research A subclass of HSP70s regulate development and abiotic stress responses in Arabidopsis thaliana Linna Leng 1 Qianqian Liang
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 Journal of Plant Research A subclass of HSP70s regulate development and abiotic stress responses in Arabidopsis thaliana Linna Leng 1 Qianqian Liang
Trasposable elements: Uses of P elements Problem set B at the end
 Trasposable elements: Uses of P elements Problem set B at the end P-elements have revolutionized the way Drosophila geneticists conduct their research. Here, we will discuss just a few of the approaches
Trasposable elements: Uses of P elements Problem set B at the end P-elements have revolutionized the way Drosophila geneticists conduct their research. Here, we will discuss just a few of the approaches
Genomes summary. Bacterial genome sizes
 Genomes summary 1. >930 bacterial genomes sequenced. 2. Circular. Genes densely packed. 3. 2-10 Mbases, 470-7,000 genes 4. Genomes of >200 eukaryotes (45 higher ) sequenced. 5. Linear chromosomes 6. On
Genomes summary 1. >930 bacterial genomes sequenced. 2. Circular. Genes densely packed. 3. 2-10 Mbases, 470-7,000 genes 4. Genomes of >200 eukaryotes (45 higher ) sequenced. 5. Linear chromosomes 6. On
Learning Objectives. Define RNA interference. Define basic terminology. Describe molecular mechanism. Define VSP and relevance
 Learning Objectives Define RNA interference Define basic terminology Describe molecular mechanism Define VSP and relevance Describe role of RNAi in antigenic variation A Nobel Way to Regulate Gene Expression
Learning Objectives Define RNA interference Define basic terminology Describe molecular mechanism Define VSP and relevance Describe role of RNAi in antigenic variation A Nobel Way to Regulate Gene Expression
2. Outline the levels of DNA packing in the eukaryotic nucleus below next to the diagram provided.
 AP Biology Reading Packet 6- Molecular Genetics Part 2 Name Chapter 19: Eukaryotic Genomes 1. Define the following terms: a. Euchromatin b. Heterochromatin c. Nucleosome 2. Outline the levels of DNA packing
AP Biology Reading Packet 6- Molecular Genetics Part 2 Name Chapter 19: Eukaryotic Genomes 1. Define the following terms: a. Euchromatin b. Heterochromatin c. Nucleosome 2. Outline the levels of DNA packing
Lecture 3 Mutagens and Mutagenesis. 1. Mutagens A. Physical and Chemical mutagens B. Transposons and retrotransposons C. T-DNA
 Lecture 3 Mutagens and Mutagenesis 1. Mutagens A. Physical and Chemical mutagens B. Transposons and retrotransposons C. T-DNA 2. Mutagenesis A. Screen B. Selection C. Lethal mutations Read: 508-514 Figs:
Lecture 3 Mutagens and Mutagenesis 1. Mutagens A. Physical and Chemical mutagens B. Transposons and retrotransposons C. T-DNA 2. Mutagenesis A. Screen B. Selection C. Lethal mutations Read: 508-514 Figs:
M I C R O B I O L O G Y WITH DISEASES BY TAXONOMY, THIRD EDITION
 M I C R O B I O L O G Y WITH DISEASES BY TAXONOMY, THIRD EDITION Chapter 7 Microbial Genetics Lecture prepared by Mindy Miller-Kittrell, University of Tennessee, Knoxville The Structure and Replication
M I C R O B I O L O G Y WITH DISEASES BY TAXONOMY, THIRD EDITION Chapter 7 Microbial Genetics Lecture prepared by Mindy Miller-Kittrell, University of Tennessee, Knoxville The Structure and Replication
CHAPTER 9 DNA Technologies
 CHAPTER 9 DNA Technologies Recombinant DNA Artificially created DNA that combines sequences that do not occur together in the nature Basis of much of the modern molecular biology Molecular cloning of genes
CHAPTER 9 DNA Technologies Recombinant DNA Artificially created DNA that combines sequences that do not occur together in the nature Basis of much of the modern molecular biology Molecular cloning of genes
Year III Pharm.D Dr. V. Chitra
 Year III Pharm.D Dr. V. Chitra 1 Genome entire genetic material of an individual Transcriptome set of transcribed sequences Proteome set of proteins encoded by the genome 2 Only one strand of DNA serves
Year III Pharm.D Dr. V. Chitra 1 Genome entire genetic material of an individual Transcriptome set of transcribed sequences Proteome set of proteins encoded by the genome 2 Only one strand of DNA serves
Multiple choice questions (numbers in brackets indicate the number of correct answers)
 1 Multiple choice questions (numbers in brackets indicate the number of correct answers) February 1, 2013 1. Ribose is found in Nucleic acids Proteins Lipids RNA DNA (2) 2. Most RNA in cells is transfer
1 Multiple choice questions (numbers in brackets indicate the number of correct answers) February 1, 2013 1. Ribose is found in Nucleic acids Proteins Lipids RNA DNA (2) 2. Most RNA in cells is transfer
Functional Genomics in Plants
 Functional Genomics in Plants Jeffrey L Bennetzen, Purdue University, West Lafayette, Indiana, USA Functional genomics refers to a suite of genetic technologies that will contribute to a comprehensive
Functional Genomics in Plants Jeffrey L Bennetzen, Purdue University, West Lafayette, Indiana, USA Functional genomics refers to a suite of genetic technologies that will contribute to a comprehensive
Theoretical cloning project
 Theoretical cloning project Needed to get credits Make it up yourself, don't copy Possible to do in groups of 2-4 students If you need help or an idea, ask! If you have no idea what to clone, I can give
Theoretical cloning project Needed to get credits Make it up yourself, don't copy Possible to do in groups of 2-4 students If you need help or an idea, ask! If you have no idea what to clone, I can give
CHAPTER 21 LECTURE SLIDES
 CHAPTER 21 LECTURE SLIDES Prepared by Brenda Leady University of Toledo To run the animations you must be in Slideshow View. Use the buttons on the animation to play, pause, and turn audio/text on or off.
CHAPTER 21 LECTURE SLIDES Prepared by Brenda Leady University of Toledo To run the animations you must be in Slideshow View. Use the buttons on the animation to play, pause, and turn audio/text on or off.
Lecture 25 (11/15/17)
 Lecture 25 (11/15/17) Reading: Ch9; 328-332 Ch25; 990-995, 1005-1012 Problems: Ch9 (study-guide: applying); 1,2 Ch9 (study-guide: facts); 7,8 Ch25 (text); 1-3,5-7,9,10,13-15 Ch25 (study-guide: applying);
Lecture 25 (11/15/17) Reading: Ch9; 328-332 Ch25; 990-995, 1005-1012 Problems: Ch9 (study-guide: applying); 1,2 Ch9 (study-guide: facts); 7,8 Ch25 (text); 1-3,5-7,9,10,13-15 Ch25 (study-guide: applying);
Supplementary information, Figure S1
 Supplementary information, Figure S1 (A) Schematic diagram of the sgrna and hspcas9 expression cassettes in a single binary vector designed for Agrobacterium-mediated stable transformation of Arabidopsis
Supplementary information, Figure S1 (A) Schematic diagram of the sgrna and hspcas9 expression cassettes in a single binary vector designed for Agrobacterium-mediated stable transformation of Arabidopsis
Title: Functional analysis of the novel TBX5 c.1333delc mutation resulting in an extended TBX5 protein
 Author's response to reviews Title: Functional analysis of the novel TBX5 c.1333delc mutation resulting in an extended TBX5 protein Authors: Johann Bohm (Johann.Boehm@titus.u-strasbg.fr) Wolfram Heinritz
Author's response to reviews Title: Functional analysis of the novel TBX5 c.1333delc mutation resulting in an extended TBX5 protein Authors: Johann Bohm (Johann.Boehm@titus.u-strasbg.fr) Wolfram Heinritz
MIT Department of Biology 7.013: Introductory Biology - Spring 2005 Instructors: Professor Hazel Sive, Professor Tyler Jacks, Dr.
 MIT Department of Biology 7.01: Introductory Biology - Spring 2005 Instructors: Professor Hazel Sive, Professor Tyler Jacks, Dr. Claudette Gardel iv) Would Xba I be useful for cloning? Why or why not?
MIT Department of Biology 7.01: Introductory Biology - Spring 2005 Instructors: Professor Hazel Sive, Professor Tyler Jacks, Dr. Claudette Gardel iv) Would Xba I be useful for cloning? Why or why not?
Groups of new plant breeding techniques
 WORKSHOP COMPERATIVE SITUATION OF NEW PLANT BREEDING TECHNIQUES 12-13 SEPTEMBER 2011 SEVILLE, SPAIN Groups of new plant breeding techniques Maria Lusser Joint Research Centre, European Commission Workshop
WORKSHOP COMPERATIVE SITUATION OF NEW PLANT BREEDING TECHNIQUES 12-13 SEPTEMBER 2011 SEVILLE, SPAIN Groups of new plant breeding techniques Maria Lusser Joint Research Centre, European Commission Workshop
Methods. An efficient reverse genetics platform in the model legume Medicago truncatula. Research
 Research Methods An efficient reverse genetics platform in the model legume Medicago truncatula Xiaofei Cheng 1, Mingyi Wang 1, Hee-Kyung Lee 1, Million Tadege 1, Pascal Ratet 2, Michael Udvardi 1, Kirankumar
Research Methods An efficient reverse genetics platform in the model legume Medicago truncatula Xiaofei Cheng 1, Mingyi Wang 1, Hee-Kyung Lee 1, Million Tadege 1, Pascal Ratet 2, Michael Udvardi 1, Kirankumar
HiPer RT-PCR Teaching Kit
 HiPer RT-PCR Teaching Kit Product Code: HTBM024 Number of experiments that can be performed: 5 Duration of Experiment: Protocol: 4 hours Agarose Gel Electrophoresis: 45 minutes Storage Instructions: The
HiPer RT-PCR Teaching Kit Product Code: HTBM024 Number of experiments that can be performed: 5 Duration of Experiment: Protocol: 4 hours Agarose Gel Electrophoresis: 45 minutes Storage Instructions: The
Introduction to C. elegans and RNA interference
 Introduction to C. elegans and RNA interference Why study model organisms? The problem: In order to understand biology, we need to learn about the function of the underlying genes How can we find out what
Introduction to C. elegans and RNA interference Why study model organisms? The problem: In order to understand biology, we need to learn about the function of the underlying genes How can we find out what
F 11/23 Happy Thanksgiving! 8 M 11/26 Gene identification in the genomic era Bamshad et al. Nature Reviews Genetics 12: , 2011
 3 rd Edition 4 th Edition Lecture Day Date Topic Reading Problems Reading Problems 1 M 11/5 Complementation testing reveals that genes are distinct entities Ch. 7 224-232 2 W 11/7 One gene makes one protein
3 rd Edition 4 th Edition Lecture Day Date Topic Reading Problems Reading Problems 1 M 11/5 Complementation testing reveals that genes are distinct entities Ch. 7 224-232 2 W 11/7 One gene makes one protein
DNA Binding Domains: Structural Motifs. Effector Domain. Zinc Fingers. Zinc Fingers, continued. Zif268
 DNA Binding Domains: Structural Motifs Studies of known transcription factors have found several motifs of protein design to allow sequence-specific binding of DNA. We will cover only three of these motifs:
DNA Binding Domains: Structural Motifs Studies of known transcription factors have found several motifs of protein design to allow sequence-specific binding of DNA. We will cover only three of these motifs:
Learning Objectives :
 Learning Objectives : Understand the basic differences between genomic and cdna libraries Understand how genomic libraries are constructed Understand the purpose for having overlapping DNA fragments in
Learning Objectives : Understand the basic differences between genomic and cdna libraries Understand how genomic libraries are constructed Understand the purpose for having overlapping DNA fragments in
Genomics and Gene Recognition Genes and Blue Genes
 Genomics and Gene Recognition Genes and Blue Genes November 1, 2004 Prokaryotic Gene Structure prokaryotes are simplest free-living organisms studying prokaryotes can give us a sense what is the minimum
Genomics and Gene Recognition Genes and Blue Genes November 1, 2004 Prokaryotic Gene Structure prokaryotes are simplest free-living organisms studying prokaryotes can give us a sense what is the minimum
Molecular Cloning and Expression Analysis of a Zinc Finger Protein Gene in Apple
 Acta Botanica Sinica 2004, 46 (9): 1091 1099 http://www.chineseplantscience.com Molecular Cloning and Expression Analysis of a Zinc Finger Protein Gene in Apple CAO Qiu-Fen 1*, Masato WADA 2, MENG Yu-Ping
Acta Botanica Sinica 2004, 46 (9): 1091 1099 http://www.chineseplantscience.com Molecular Cloning and Expression Analysis of a Zinc Finger Protein Gene in Apple CAO Qiu-Fen 1*, Masato WADA 2, MENG Yu-Ping
Bi 8 Lecture 7. Ellen Rothenberg 26 January Reading: Ch. 3, pp ; panel 3-1
 Bi 8 Lecture 7 PROTEIN STRUCTURE, Functional analysis, and evolution Ellen Rothenberg 26 January 2016 Reading: Ch. 3, pp. 109-134; panel 3-1 (end with free amine) aromatic, hydrophobic small, hydrophilic
Bi 8 Lecture 7 PROTEIN STRUCTURE, Functional analysis, and evolution Ellen Rothenberg 26 January 2016 Reading: Ch. 3, pp. 109-134; panel 3-1 (end with free amine) aromatic, hydrophobic small, hydrophilic
3. human genomics clone genes associated with genetic disorders. 4. many projects generate ordered clones that cover genome
 Lectures 30 and 31 Genome analysis I. Genome analysis A. two general areas 1. structural 2. functional B. genome projects a status report 1. 1 st sequenced: several viral genomes 2. mitochondria and chloroplasts
Lectures 30 and 31 Genome analysis I. Genome analysis A. two general areas 1. structural 2. functional B. genome projects a status report 1. 1 st sequenced: several viral genomes 2. mitochondria and chloroplasts
Supplementary Figure 1 qrt-pcr expression analysis of NLP8 with and without KNO 3 during germination.
 Supplementary Figure 1 qrt-pcr expression analysis of NLP8 with and without KNO 3 during germination. Seeds of Col-0 were harvested from plants grown at 16 C, stored for 2 months, imbibed for indicated
Supplementary Figure 1 qrt-pcr expression analysis of NLP8 with and without KNO 3 during germination. Seeds of Col-0 were harvested from plants grown at 16 C, stored for 2 months, imbibed for indicated
Chapter 24: Promoters and Enhancers
 Chapter 24: Promoters and Enhancers A typical gene transcribed by RNA polymerase II has a promoter that usually extends upstream from the site where transcription is initiated the (#1) of transcription
Chapter 24: Promoters and Enhancers A typical gene transcribed by RNA polymerase II has a promoter that usually extends upstream from the site where transcription is initiated the (#1) of transcription
Some types of Mutagenesis
 Mutagenesis What Is a Mutation? Genetic information is encoded by the sequence of the nucleotide bases in DNA of the gene. The four nucleotides are: adenine (A), thymine (T), guanine (G), and cytosine
Mutagenesis What Is a Mutation? Genetic information is encoded by the sequence of the nucleotide bases in DNA of the gene. The four nucleotides are: adenine (A), thymine (T), guanine (G), and cytosine
AP Biology Gene Expression/Biotechnology REVIEW
 AP Biology Gene Expression/Biotechnology REVIEW Multiple Choice Identify the choice that best completes the statement or answers the question. 1. Gene expression can be a. regulated before transcription.
AP Biology Gene Expression/Biotechnology REVIEW Multiple Choice Identify the choice that best completes the statement or answers the question. 1. Gene expression can be a. regulated before transcription.
BIOLOGICAL DIMENSIONS OF THE GMO ISSUE
 Carlson, J. 2005. Biological dimensions of the GMO issue. In, proc. of conf. on restoration of American chestnut to forest lands, Steiner, K.C. and J.E. Carlson (eds.). BIOLOGICAL DIMENSIONS OF THE GMO
Carlson, J. 2005. Biological dimensions of the GMO issue. In, proc. of conf. on restoration of American chestnut to forest lands, Steiner, K.C. and J.E. Carlson (eds.). BIOLOGICAL DIMENSIONS OF THE GMO
UNIVERSITY OF YORK BA, BSc, and MSc Degree Examinations
 Examination Candidate Number: Desk Number: UNIVERSITY OF YORK BA, BSc, and MSc Degree Examinations 2017-8 Department : BIOLOGY Title of Exam: Bacterial pathogenesis Time Allowed: 2 hours Marking Scheme:
Examination Candidate Number: Desk Number: UNIVERSITY OF YORK BA, BSc, and MSc Degree Examinations 2017-8 Department : BIOLOGY Title of Exam: Bacterial pathogenesis Time Allowed: 2 hours Marking Scheme:
DNA is the genetic material. DNA structure. Chapter 7: DNA Replication, Transcription & Translation; Mutations & Ames test
 DNA is the genetic material Chapter 7: DNA Replication, Transcription & Translation; Mutations & Ames test Dr. Amy Rogers Bio 139 General Microbiology Hereditary information is carried by DNA Griffith/Avery
DNA is the genetic material Chapter 7: DNA Replication, Transcription & Translation; Mutations & Ames test Dr. Amy Rogers Bio 139 General Microbiology Hereditary information is carried by DNA Griffith/Avery
The plant-specific LEAFY (LFY) protein is necessary and
 Genomic identification of direct target genes of LEAFY Dilusha A. William*, Yanhui Su*, Michael R. Smith*, Meina Lu*, Don A. Baldwin, and Doris Wagner* *Department of Biology and Penn Microarray Facility,
Genomic identification of direct target genes of LEAFY Dilusha A. William*, Yanhui Su*, Michael R. Smith*, Meina Lu*, Don A. Baldwin, and Doris Wagner* *Department of Biology and Penn Microarray Facility,
Non-Organic-Based Isolation of Mammalian microrna using Norgen s microrna Purification Kit
 Application Note 13 RNA Sample Preparation Non-Organic-Based Isolation of Mammalian microrna using Norgen s microrna Purification Kit B. Lam, PhD 1, P. Roberts, MSc 1 Y. Haj-Ahmad, M.Sc., Ph.D 1,2 1 Norgen
Application Note 13 RNA Sample Preparation Non-Organic-Based Isolation of Mammalian microrna using Norgen s microrna Purification Kit B. Lam, PhD 1, P. Roberts, MSc 1 Y. Haj-Ahmad, M.Sc., Ph.D 1,2 1 Norgen
New Tool for Late Stage Visualization of Oskar Protein in Drosophila melanogaster. Oocytes. Kathleen Harnois. Supervised by Paul Macdonald
 New Tool for Late Stage Visualization of Oskar Protein in Drosophila melanogaster Oocytes Kathleen Harnois Supervised by Paul Macdonald Institute of Cellular and Molecular Biology, The University of Texas
New Tool for Late Stage Visualization of Oskar Protein in Drosophila melanogaster Oocytes Kathleen Harnois Supervised by Paul Macdonald Institute of Cellular and Molecular Biology, The University of Texas
Le proteine regolative variano nei vari tipi cellulari e in funzione degli stimoli ambientali
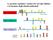 Le proteine regolative variano nei vari tipi cellulari e in funzione degli stimoli ambientali Tipo cellulare 1 Tipo cellulare 2 Tipo cellulare 3 DNA-protein Crosslink Lisi Frammentazione Immunopurificazione
Le proteine regolative variano nei vari tipi cellulari e in funzione degli stimoli ambientali Tipo cellulare 1 Tipo cellulare 2 Tipo cellulare 3 DNA-protein Crosslink Lisi Frammentazione Immunopurificazione
Gene Expression and Heritable Phenotype. CBS520 Eric Nabity
 Gene Expression and Heritable Phenotype CBS520 Eric Nabity DNA is Just the Beginning DNA was determined to be the genetic material, and the structure was identified as a (double stranded) double helix.
Gene Expression and Heritable Phenotype CBS520 Eric Nabity DNA is Just the Beginning DNA was determined to be the genetic material, and the structure was identified as a (double stranded) double helix.
Using mutants to clone genes
 Using mutants to clone genes Objectives 1. What is positional cloning? 2. What is insertional tagging? 3. How can one confirm that the gene cloned is the same one that is mutated to give the phenotype
Using mutants to clone genes Objectives 1. What is positional cloning? 2. What is insertional tagging? 3. How can one confirm that the gene cloned is the same one that is mutated to give the phenotype
Chapter 8: DNA and RNA
 Chapter 8: DNA and RNA Lecture Outline Enger, E. D., Ross, F. C., & Bailey, D. B. (2012). Concepts in biology (14th ed.). New York: McGraw- Hill. 1 8-1 DNA and the Importance of Proteins Proteins play
Chapter 8: DNA and RNA Lecture Outline Enger, E. D., Ross, F. C., & Bailey, D. B. (2012). Concepts in biology (14th ed.). New York: McGraw- Hill. 1 8-1 DNA and the Importance of Proteins Proteins play
Impact of Retinoic acid induced-1 (Rai1) on Regulators of Metabolism and Adipogenesis
 Impact of Retinoic acid induced-1 (Rai1) on Regulators of Metabolism and Adipogenesis The mammalian system undergoes ~24 hour cycles known as circadian rhythms that temporally orchestrate metabolism, behavior,
Impact of Retinoic acid induced-1 (Rai1) on Regulators of Metabolism and Adipogenesis The mammalian system undergoes ~24 hour cycles known as circadian rhythms that temporally orchestrate metabolism, behavior,
Methods of Biomaterials Testing Lesson 3-5. Biochemical Methods - Molecular Biology -
 Methods of Biomaterials Testing Lesson 3-5 Biochemical Methods - Molecular Biology - Chromosomes in the Cell Nucleus DNA in the Chromosome Deoxyribonucleic Acid (DNA) DNA has double-helix structure The
Methods of Biomaterials Testing Lesson 3-5 Biochemical Methods - Molecular Biology - Chromosomes in the Cell Nucleus DNA in the Chromosome Deoxyribonucleic Acid (DNA) DNA has double-helix structure The
Genetics Lecture Notes Lectures 17 19
 Genetics Lecture Notes 7.03 2005 Lectures 17 19 Lecture 17 Gene Regulation We are now going to look at ways that genetics can be used to study gene regulation. The issue is how cells adjust the expression
Genetics Lecture Notes 7.03 2005 Lectures 17 19 Lecture 17 Gene Regulation We are now going to look at ways that genetics can be used to study gene regulation. The issue is how cells adjust the expression
Epigenetics. Medical studies in English, Lecture # 12,
 Epigenetics Medical studies in English, 2018. Lecture # 12, Epigenetics Regulation of gene activity in eukaryotes Correlation of chromatin structure with transcription stably heritable phenotype resulting
Epigenetics Medical studies in English, 2018. Lecture # 12, Epigenetics Regulation of gene activity in eukaryotes Correlation of chromatin structure with transcription stably heritable phenotype resulting
LATE-PCR. Linear-After-The-Exponential
 LATE-PCR Linear-After-The-Exponential A Patented Invention of the Laboratory of Human Genetics and Reproductive Biology Lab. Director: Lawrence J. Wangh, Ph.D. Department of Biology, Brandeis University,
LATE-PCR Linear-After-The-Exponential A Patented Invention of the Laboratory of Human Genetics and Reproductive Biology Lab. Director: Lawrence J. Wangh, Ph.D. Department of Biology, Brandeis University,
Cloning of an ovule specific promoter from Arabidopsis thaliana and expression of β-glucuronidase
 Indian Journal of Experimental Biology Vol. 46, April 2008, pp. 207-211 Cloning of an ovule specific promoter from Arabidopsis thaliana and expression of β-glucuronidase Vikrant Nain, Anju Verma a, Neeraj
Indian Journal of Experimental Biology Vol. 46, April 2008, pp. 207-211 Cloning of an ovule specific promoter from Arabidopsis thaliana and expression of β-glucuronidase Vikrant Nain, Anju Verma a, Neeraj
7.06 Cell Biology EXAM #2 March 20, 2003
 7.06 Cell Biology EXAM #2 March 20, 2003 This is an open book exam, and you are allowed access to books, a calculator, and notes but not computers or any other types of electronic devices. Please write
7.06 Cell Biology EXAM #2 March 20, 2003 This is an open book exam, and you are allowed access to books, a calculator, and notes but not computers or any other types of electronic devices. Please write
Chapter 17. PCR the polymerase chain reaction and its many uses. Prepared by Woojoo Choi
 Chapter 17. PCR the polymerase chain reaction and its many uses Prepared by Woojoo Choi Polymerase chain reaction 1) Polymerase chain reaction (PCR): artificial amplification of a DNA sequence by repeated
Chapter 17. PCR the polymerase chain reaction and its many uses Prepared by Woojoo Choi Polymerase chain reaction 1) Polymerase chain reaction (PCR): artificial amplification of a DNA sequence by repeated
Higher Human Biology Unit 1: Human Cells Pupils Learning Outcomes
 Higher Human Biology Unit 1: Human Cells Pupils Learning Outcomes 1.1 Division and Differentiation in Human Cells I can state that cellular differentiation is the process by which a cell develops more
Higher Human Biology Unit 1: Human Cells Pupils Learning Outcomes 1.1 Division and Differentiation in Human Cells I can state that cellular differentiation is the process by which a cell develops more
The Use of Genetically-Modified Mouse Models to Study the Actin Cytoskeleton
 The Use of Genetically-Modified Mouse Models to Study the Actin Cytoskeleton Anthony Kee (PhD) Cellular and Genetic Medicine Unit School of Medical Sciences (a.kee@unsw.edu.au) 2017 Structure of the Prac
The Use of Genetically-Modified Mouse Models to Study the Actin Cytoskeleton Anthony Kee (PhD) Cellular and Genetic Medicine Unit School of Medical Sciences (a.kee@unsw.edu.au) 2017 Structure of the Prac
Today s lecture: Types of mutations and their impact on protein function
 Today s lecture: Types of mutations and their impact on protein function Mutations can be classified by their effect on the DNA sequence OR the encoded protein 1 From my Lecture 4 (10/1): Classification
Today s lecture: Types of mutations and their impact on protein function Mutations can be classified by their effect on the DNA sequence OR the encoded protein 1 From my Lecture 4 (10/1): Classification
If Dna Has The Instructions For Building Proteins Why Is Mrna Needed
 If Dna Has The Instructions For Building Proteins Why Is Mrna Needed if a strand of DNA has the sequence CGGTATATC, then the complementary each strand of DNA contains the info needed to produce the complementary
If Dna Has The Instructions For Building Proteins Why Is Mrna Needed if a strand of DNA has the sequence CGGTATATC, then the complementary each strand of DNA contains the info needed to produce the complementary
DICER-LIKE2 Plays a Primary Role in Transitive Silencing of Transgenes in Arabidopsis
 DICER-LIKE2 Plays a Primary Role in Transitive Silencing of Transgenes in Arabidopsis Sizolwenkosi Mlotshwa 1, Gail J. Pruss 1, Angela Peragine 2, Matthew W. Endres 1, Junjie Li 3, Xuemei Chen 3, R. Scott
DICER-LIKE2 Plays a Primary Role in Transitive Silencing of Transgenes in Arabidopsis Sizolwenkosi Mlotshwa 1, Gail J. Pruss 1, Angela Peragine 2, Matthew W. Endres 1, Junjie Li 3, Xuemei Chen 3, R. Scott
Lecture 20: Drosophila melanogaster
 Lecture 20: Drosophila melanogaster Model organisms Polytene chromosome Life cycle P elements and transformation Embryogenesis Read textbook: 732-744; Fig. 20.4; 20.10; 20.15-26 www.mhhe.com/hartwell3
Lecture 20: Drosophila melanogaster Model organisms Polytene chromosome Life cycle P elements and transformation Embryogenesis Read textbook: 732-744; Fig. 20.4; 20.10; 20.15-26 www.mhhe.com/hartwell3
Pre-Lab: Molecular Biology
 Pre-Lab: Molecular Biology Name 1. What are the three chemical parts of a nucleotide. Draw a simple sketch to show how the three parts are arranged. 2. What are the rules of base pairing? 3. In double
Pre-Lab: Molecular Biology Name 1. What are the three chemical parts of a nucleotide. Draw a simple sketch to show how the three parts are arranged. 2. What are the rules of base pairing? 3. In double
Name Class Date. Practice Test
 Name Class Date 12 DNA Practice Test Multiple Choice Write the letter that best answers the question or completes the statement on the line provided. 1. What do bacteriophages infect? a. mice. c. viruses.
Name Class Date 12 DNA Practice Test Multiple Choice Write the letter that best answers the question or completes the statement on the line provided. 1. What do bacteriophages infect? a. mice. c. viruses.
NAME TA SEC Problem Set 4 FRIDAY October 15, Answers to this problem set must be inserted into the box outside
 MIT Biology Department 7.012: Introductory Biology - Fall 2004 Instructors: Professor Eric Lander, Professor Robert A. Weinberg, Dr. Claudette Gardel NAME TA SEC 7.012 Problem Set 4 FRIDAY October 15,
MIT Biology Department 7.012: Introductory Biology - Fall 2004 Instructors: Professor Eric Lander, Professor Robert A. Weinberg, Dr. Claudette Gardel NAME TA SEC 7.012 Problem Set 4 FRIDAY October 15,
Differential Gene Expression
 Biology 4361 Developmental Biology Differential Gene Expression September 28, 2006 Chromatin Structure ~140 bp ~60 bp Transcriptional Regulation: 1. Packing prevents access CH 3 2. Acetylation ( C O )
Biology 4361 Developmental Biology Differential Gene Expression September 28, 2006 Chromatin Structure ~140 bp ~60 bp Transcriptional Regulation: 1. Packing prevents access CH 3 2. Acetylation ( C O )
Prions Other molecules besides organelle DNA are inherited in non-mendelian patterns.
 Prions Other molecules besides organelle DNA are inherited in non-mendelian patterns. Examples of non-mendelian patterns of inheritance extend beyond the inheritance of organelle DNA. Certain DNA and RNA
Prions Other molecules besides organelle DNA are inherited in non-mendelian patterns. Examples of non-mendelian patterns of inheritance extend beyond the inheritance of organelle DNA. Certain DNA and RNA
Make the protein through the genetic dogma process.
 Make the protein through the genetic dogma process. Coding Strand 5 AGCAATCATGGATTGGGTACATTTGTAACTGT 3 Template Strand mrna Protein Complete the table. DNA strand DNA s strand G mrna A C U G T A T Amino
Make the protein through the genetic dogma process. Coding Strand 5 AGCAATCATGGATTGGGTACATTTGTAACTGT 3 Template Strand mrna Protein Complete the table. DNA strand DNA s strand G mrna A C U G T A T Amino
SUPPLEMENTAL MATERIALS
 SUPPLEMENL MERILS Eh-seq: RISPR epitope tagging hip-seq of DN-binding proteins Daniel Savic, E. hristopher Partridge, Kimberly M. Newberry, Sophia. Smith, Sarah K. Meadows, rian S. Roberts, Mark Mackiewicz,
SUPPLEMENL MERILS Eh-seq: RISPR epitope tagging hip-seq of DN-binding proteins Daniel Savic, E. hristopher Partridge, Kimberly M. Newberry, Sophia. Smith, Sarah K. Meadows, rian S. Roberts, Mark Mackiewicz,
Supplementary Table 1. The Q-PCR primer sequence is summarized in the following table.
 Supplementary Table 1. The Q-PCR primer sequence is summarized in the following table. Name Sequence (5-3 ) Application Flag-u ggactacaaggacgacgatgac Shared upstream primer for all the amplifications of
Supplementary Table 1. The Q-PCR primer sequence is summarized in the following table. Name Sequence (5-3 ) Application Flag-u ggactacaaggacgacgatgac Shared upstream primer for all the amplifications of
DNA Structure and Properties Basic Properties Predicting Melting Temperature. Dinesh Yadav
 DNA Structure and Properties Basic Properties Predicting Melting Temperature Dinesh Yadav Nucleic Acid Structure Question: Is this RNA or DNA? Molecules of Life, pp. 15 2 Nucleic Acid Bases Molecules of
DNA Structure and Properties Basic Properties Predicting Melting Temperature Dinesh Yadav Nucleic Acid Structure Question: Is this RNA or DNA? Molecules of Life, pp. 15 2 Nucleic Acid Bases Molecules of
Structural variation. Marta Puig Institut de Biotecnologia i Biomedicina Universitat Autònoma de Barcelona
 Structural variation Marta Puig Institut de Biotecnologia i Biomedicina Universitat Autònoma de Barcelona Genetic variation How much genetic variation is there between individuals? What type of variants
Structural variation Marta Puig Institut de Biotecnologia i Biomedicina Universitat Autònoma de Barcelona Genetic variation How much genetic variation is there between individuals? What type of variants
REGULATION OF GENE EXPRESSION
 REGULATION OF GENE EXPRESSION Each cell of a living organism contains thousands of genes. But all genes do not function at a time. Genes function according to requirements of the cell. Genes control the
REGULATION OF GENE EXPRESSION Each cell of a living organism contains thousands of genes. But all genes do not function at a time. Genes function according to requirements of the cell. Genes control the
8/21/2014. From Gene to Protein
 From Gene to Protein Chapter 17 Objectives Describe the contributions made by Garrod, Beadle, and Tatum to our understanding of the relationship between genes and enzymes Briefly explain how information
From Gene to Protein Chapter 17 Objectives Describe the contributions made by Garrod, Beadle, and Tatum to our understanding of the relationship between genes and enzymes Briefly explain how information
Solutions to Quiz II
 MIT Department of Biology 7.014 Introductory Biology, Spring 2005 Solutions to 7.014 Quiz II Class Average = 79 Median = 82 Grade Range % A 90-100 27 B 75-89 37 C 59 74 25 D 41 58 7 F 0 40 2 Question 1
MIT Department of Biology 7.014 Introductory Biology, Spring 2005 Solutions to 7.014 Quiz II Class Average = 79 Median = 82 Grade Range % A 90-100 27 B 75-89 37 C 59 74 25 D 41 58 7 F 0 40 2 Question 1
Yeast Two Hybrid Assay: A Fishing Tale
 KEYWORDS: yeast two hybrid, molecular interactions, galactose metabolism Special section on techniques: Yeast Two Hybrid Assay: A Fishing Tale Solmaz Sobhanifar Pathology, University of British Columbia
KEYWORDS: yeast two hybrid, molecular interactions, galactose metabolism Special section on techniques: Yeast Two Hybrid Assay: A Fishing Tale Solmaz Sobhanifar Pathology, University of British Columbia
M Keramatipour 2. M Keramatipour 1. M Keramatipour 4. M Keramatipour 3. M Keramatipour 5. M Keramatipour
 Molecular Cloning Methods Mohammad Keramatipour MD, PhD keramatipour@tums.ac.ir Outline DNA recombinant technology DNA cloning co Cell based PCR PCR-based Some application of DNA cloning Genomic libraries
Molecular Cloning Methods Mohammad Keramatipour MD, PhD keramatipour@tums.ac.ir Outline DNA recombinant technology DNA cloning co Cell based PCR PCR-based Some application of DNA cloning Genomic libraries
DNA Transcription. Visualizing Transcription. The Transcription Process
 DNA Transcription By: Suzanne Clancy, Ph.D. 2008 Nature Education Citation: Clancy, S. (2008) DNA transcription. Nature Education 1(1) If DNA is a book, then how is it read? Learn more about the DNA transcription
DNA Transcription By: Suzanne Clancy, Ph.D. 2008 Nature Education Citation: Clancy, S. (2008) DNA transcription. Nature Education 1(1) If DNA is a book, then how is it read? Learn more about the DNA transcription
Differential Gene Expression
 Biology 4361 Developmental Biology Differential Gene Expression June 19, 2008 Differential Gene Expression Overview Chromatin structure Gene anatomy RNA processing and protein production Initiating transcription:
Biology 4361 Developmental Biology Differential Gene Expression June 19, 2008 Differential Gene Expression Overview Chromatin structure Gene anatomy RNA processing and protein production Initiating transcription:
Gene Expression: Transcription
 Gene Expression: Transcription The majority of genes are expressed as the proteins they encode. The process occurs in two steps: Transcription = DNA RNA Translation = RNA protein Taken together, they make
Gene Expression: Transcription The majority of genes are expressed as the proteins they encode. The process occurs in two steps: Transcription = DNA RNA Translation = RNA protein Taken together, they make
Chapter 20: Biotechnology
 Name Period The AP Biology exam has reached into this chapter for essay questions on a regular basis over the past 15 years. Student responses show that biotechnology is a difficult topic. This chapter
Name Period The AP Biology exam has reached into this chapter for essay questions on a regular basis over the past 15 years. Student responses show that biotechnology is a difficult topic. This chapter
Wilkins Franklin s photo below proved model on left to be correct for DNA
 Watson Crick Franklin Wilkins Franklin s photo below proved model on left to be correct for DNA Pauling Most important scientific paper in Biology in last 100 years First time DNA double helix seen in
Watson Crick Franklin Wilkins Franklin s photo below proved model on left to be correct for DNA Pauling Most important scientific paper in Biology in last 100 years First time DNA double helix seen in
M P BHOJ (OPEN) UNIVERSITY, BHOPAL ASSIGNMENT QUESTION PAPER
 PAPER : I Plant Development and Reproduction Q.1 Explain use of mutants in understanding seeding development? OR Short note on :- (i) Tropisms (ii) Mobilization of food reserves Q.2 Describe the organization
PAPER : I Plant Development and Reproduction Q.1 Explain use of mutants in understanding seeding development? OR Short note on :- (i) Tropisms (ii) Mobilization of food reserves Q.2 Describe the organization
7.013 Practice Quiz
 MIT Department of Biology 7.013: Introductory Biology - Spring 2005 Instructors: Professor Hazel Sive, Professor Tyler Jacks, Dr. Claudette Gardel 7.013 Practice Quiz 2 2004 1 Question 1 A. The primer
MIT Department of Biology 7.013: Introductory Biology - Spring 2005 Instructors: Professor Hazel Sive, Professor Tyler Jacks, Dr. Claudette Gardel 7.013 Practice Quiz 2 2004 1 Question 1 A. The primer
Chapter 11: Regulation of Gene Expression
 Chapter Review 1. It has long been known that there is probably a genetic link for alcoholism. Researchers studying rats have begun to elucidate this link. Briefly describe the genetic mechanism found
Chapter Review 1. It has long been known that there is probably a genetic link for alcoholism. Researchers studying rats have begun to elucidate this link. Briefly describe the genetic mechanism found
Bioinformatics of Transcriptional Regulation
 Bioinformatics of Transcriptional Regulation Carl Herrmann IPMB & DKFZ c.herrmann@dkfz.de Wechselwirkung von Maßnahmen und Auswirkungen Einflussmöglichkeiten in einem Dialog From genes to active compounds
Bioinformatics of Transcriptional Regulation Carl Herrmann IPMB & DKFZ c.herrmann@dkfz.de Wechselwirkung von Maßnahmen und Auswirkungen Einflussmöglichkeiten in einem Dialog From genes to active compounds
DNA Structure and Analysis. Chapter 4: Background
 DNA Structure and Analysis Chapter 4: Background Molecular Biology Three main disciplines of biotechnology Biochemistry Genetics Molecular Biology # Biotechnology: A Laboratory Skills Course explorer.bio-rad.com
DNA Structure and Analysis Chapter 4: Background Molecular Biology Three main disciplines of biotechnology Biochemistry Genetics Molecular Biology # Biotechnology: A Laboratory Skills Course explorer.bio-rad.com
AP Biology. Scoring Guidelines
 2017 AP Biology Scoring Guidelines College Board, Advanced Placement Program, AP, AP Central, and the acorn logo are registered trademarks of the College Board. AP Central is the official online home for
2017 AP Biology Scoring Guidelines College Board, Advanced Placement Program, AP, AP Central, and the acorn logo are registered trademarks of the College Board. AP Central is the official online home for
Chapter 18: Regulation of Gene Expression. 1. Gene Regulation in Bacteria 2. Gene Regulation in Eukaryotes 3. Gene Regulation & Cancer
 Chapter 18: Regulation of Gene Expression 1. Gene Regulation in Bacteria 2. Gene Regulation in Eukaryotes 3. Gene Regulation & Cancer Gene Regulation Gene regulation refers to all aspects of controlling
Chapter 18: Regulation of Gene Expression 1. Gene Regulation in Bacteria 2. Gene Regulation in Eukaryotes 3. Gene Regulation & Cancer Gene Regulation Gene regulation refers to all aspects of controlling
Molecular Biology, Lecture 3 DNA Replication
 Molecular Biology, Lecture 3 DNA Replication We will continue talking about DNA replication. We have previously t discussed the structure of DNA. DNA replication is the copying of the whole DNA content
Molecular Biology, Lecture 3 DNA Replication We will continue talking about DNA replication. We have previously t discussed the structure of DNA. DNA replication is the copying of the whole DNA content
CRISPR Applications: Mouse
 CRISPR Applications: Mouse Lin He UC-Berkeley Advantages of mouse as a model organism similar to human Can be genetically manipulated Isogenic and congenic genetic background An accelerated lifespan. Well-characterized
CRISPR Applications: Mouse Lin He UC-Berkeley Advantages of mouse as a model organism similar to human Can be genetically manipulated Isogenic and congenic genetic background An accelerated lifespan. Well-characterized
