Floral stem cell termination involves the direct regulation of AGAMOUS by PERIANTHIA
|
|
|
- Cori Alexander
- 6 years ago
- Views:
Transcription
1 RESEARCH REPORT 165 Development 136, (29) doi:1.1242/dev Floral stem cell termination involves the direct regulation of AGAMOUS by PERIANTHIA Pradeep Das 1,2, *, Toshiro Ito 1,3, Frank Wellmer 1,4, Teva Vernoux 2, Annick Dedieu 2, Jan Traas 2 and Elliot M. Meyerowitz 1, * In Arabidopsis, the population of stem cells present in young flower buds is lost after the production of a fixed number of floral organs. The precisely timed repression of the stem cell identity gene WUSCHEL (WUS) by the floral homeotic protein AGAMOUS (AG) is a key part of this process. In this study, we report on the identification of a novel input into the process of floral stem cell regulation. We use genetics and chromatin immunoprecipitation assays to demonstrate that the bzip transcription factor PERIANTHIA (PAN) plays a role in regulating stem cell fate by directly controlling AG expression and suggest that this activity is spatially restricted to the centermost region of the AG expression domain. These results suggest that the termination of floral stem cell fate is a multiply redundant process involving loci with unrelated floral patterning functions. KEY WORDS: AGAMOUS, Flower development, Stem cells, Arabidopsis INTRODUCTION In Arabidopsis, the indeterminate shoot apical meristem (SAM) produces organs such as leaves and flowers throughout the life of the plant. By contrast, the determinate floral meristem (FM), from which flowers are derived, produces a stereotypical number of floral organs: four sepals, four petals, six stamens and two carpels. Underlying the different behaviors of these two meristematic tissues are the different properties of their respective stem cell populations. In the SAM, as well as in the FM, the expression of the homeodomain gene WUSCHEL (WUS) in a small group of cells at the center of the structures, the so-called stem cell organizing center, is essential for maintaining the pool of stem cells. In the FM, once the correct numbers of floral organs have formed, WUS is quickly downregulated and the stem cells lost (Laux et al., 1996; Mayer et al., 1998). The floral identity regulator LEAFY (LFY) (Schultz and Haughn, 1991; Weigel et al., 1992) activates the expression of the homeotic gene AGAMOUS (AG) in the center of young flower buds, and the AG gene product then acts to downregulate WUS, leading to a loss of stem cell activity (Busch et al., 1999; Lenhard et al., 21; Lohmann et al., 21; Parcy et al., 1998). However, loss of LFY function only leads to a delay in the onset of AG expression, and not to its absence (Weigel and Meyerowitz, 1993), suggesting that other factors also play a role in the early activation of AG. One of these factors was recently shown to be WUS itself, which directly binds AG regulatory sequences in combination with LFY (Lohmann et al., 21). Flowers mutant for AG display stem cell maintenance phenotypes, resulting in the formation of flowers within flowers, and also show homeotic transformations of stamens to petals (Bowman et al., 1989). It has been suggested that these are functionally distinct activities of AG (Mizukami and Ma, 1995; Sieburth et al., 1995), yet not much is 1 Division of Biology , California Institute of Technology, Pasadena, California 91125, USA. 2 Laboratoire RDP, Ecole Normale Supérieure de Lyon, 46 allée d Italie, 697 Lyon, France. 3 Temasek Life Sciences Laboratory, National University of Singapore, Singapore 11764, Singapore. 4 Smurfit Institute of Genetics, Trinity College Dublin, College Green, Dublin 2, Ireland. *Authors for correspondence ( s: pradeep.das@ens-lyon.fr; meyerow@its.caltech.edu) Accepted 16 March 29 known about how this is regulated: whether AG is regulated at the RNA level, for example, via the regulation of AG expression in specific floral domains, or at the protein level, through interactions between AG and other spatially restricted molecules. Furthermore, it is unclear how AG shuts down WUS expression and thus the floral stem cell population (Laux et al., 1996; Mayer et al., 1998). In this study, we report on the identification of a novel input into the process of floral stem cell arrest and suggest that this activity is spatially restricted to the centermost region of the AG expression domain. MATERIALS AND METHODS Mutagenesis pan-3 seeds (1,-15,; in the L-er accession) were treated with a.3% (v/v) aqueous solution of ethyl methanesulfonate (Sigma) in a volume of 15 ml for 1 hours, then washed with water for 8 hours (with hourly changes) before being resuspended in a.15% (v/v) agar solution and sowed on soil 1 cm apart. Seeds from M1 plants were harvested individually and 2-3 M2 plants per M1 line (~1) were screened for altered floral phenotypes, which were reconfirmed in the M3 generation. Ten putative modifiers were retained after re-screening. To identify the mutation in the novel lfy allele, the genomic coding region was amplified in two fragments of 1.3 kb and 1.4 kb by PCR (using Ex Taq, Takara) and sequenced. The mutation was found to be a nonsense mutation (Q162Stop), similar to all published null alleles. Plasmid constructs and sequences Details of primers available upon request. All PCR amplifications were carried out using the Phusion high fidelity polymerase (Finnzymes). All constructs were sequenced. To construct the PAN repressor domain chimeric fusion, we first annealed complementary oligonucleotides carrying the enhanced SUPERMAN repressor domain motif flanked by BamHI and BglII sites, and ligated this to the pgem-t EZ cloning vector (Promega), to yield pgem- SRDX. The PAN cdna was PCR-amplified, digested with KpnI and BglII and cloned into the KpnI and BamHI sites of pgem-srdx to yield ppd64.1. The PAN-RD fragment was extracted with BamHI and BglII, and cloned into pbj36 (Gleave, 1992) carrying either the p35s, ppan or pap1(1.7-kb) promoters to yield ppd66.1, ppd199.2 or ppd143.1 respectively. The PAN promoter was PCR-amplified from the L-er accession and cloned into pbj36 using the SalI and KpnI sites. pap1(1.7-kb) was a kind gift of Dr Marty Yanofsky (University of California, San Diego, CA, USA). The promoter- PAN-RD fragments were then extracted from pbj36 using NotI and ligated to the plant transformation vector pml BART (Eshed et al., 21) yielding ppd74.1, ppd218.1 or ppd171.2, respectively.
2 166 RESEARCH REPORT Development 136 (1) For the ethanol-inducible version of PAN-RD under the control of the PAN promoter, we used the MultiSite Gateway Three-Fragment Vector Construction Kit (Invitrogen) to generate a single plasmid harboring the two components. We PCR-amplified a fragment of the LFY::alcR--alcA::ER- GFP pgreen binary vector (gift of Patrick Laufs; INRA, Versailles, France); the alcr gene harboring a 3 nos terminator, followed by a 35S terminator in an inverted orientation. This fragment was recombined with the pdonr 221 vector to generate pentr-alcr-2xt. We also modified a destination vector (pgreen 229; gift of Philip Benfey; Duke University, Durham, NC, USA) by inserting the chimeric promoter alca immediately after, and oriented towards, the attr3 recombination sequence. To do so, the alca promoter was PCR-amplified from the LFY::alcR--alcA::ER-GFP pgreen vector, digested with SpeI and HinDIII and ligated to the pgreen 229 binary vector, to yield the dpgreenbar-alca binary vector. The PAN promoter was PCR-amplified from Col- and recombined with pdonr P4-P1R to yield ppd277. The PAN-RD fragment was amplified from ppd269 and recombined with pdonr P2R-P3 to yield ppd317. Finally, the three pentr vectors (ppd277, pentr-alcr-2xt and ppd317) were recombined into dpgreenbar-alca, to yield the PAN::alcR--alcA::PAN-RD binary vector. The putative bzip binding sites in the second AG intron were identified based on the presence of ACGT core sequences (Jakoby et al., 22). The six observed sites are (5-3 ): ACTTATACGTACATGT, AGTCCC - ACGTGATTAC, TTGATCACGTCATCAC, TGTAATACGT ATTTGT, TATGGAACGTTGTGAT and TCCATCACGTTTAAAT. For the p35s::pan-vp16 construct, VP16 was PCR-amplified, digested with BamHI and BglII, and ligated to pbj36. The PAN-VP16 fragment was then PCR-amplified and cloned into the pdonr 221 vector (Invitrogen). The triple gateway system was then used to generate the final p35s::pan- VP16 plasmid in pdpgreen-bart. The reporter construct used in the particle bombardment assays, pagi-3, is published as KB31 (Busch et al., 1999). Plant lines, transgenics and plant growth conditions All plants were grown at either 16 C or 22 C with continuous white light, except the pan-3 and L-er plants shown in Fig. S6, which were grown at long days (22 C) under a combination of white and gro-lux light. Photos of flowers were taken using either a Zeiss Stemi SV 11 stereomicroscope fitted with a Zeiss Axiocam or a Leica MZ12 stereomicroscope with a Leica DFC32 camera. Some images were adjusted for clarity by altering the brightness or contrast but any changes were applied evenly, across the entirety of the picture, without exception. In situ hybridizations were performed according to published protocols. The WUS antisense RNA probe corresponds to the entire cdna. Photos were taken on a Nikon Optiphot-2 equipped with a Zeiss Axiocam. Transgenic plants were generated using standard floral dipping methods. Transformant lines were selected on soil for resistance to the herbicide Basta. To determine the copy number of the PAN-RD transgene, T2 seeds were plated on petri dishes containing 1 μg/ml ammonium glufosinate (Basta) and the ratio of resistant to sensitive seedlings determined (~75% resistant seedlings indicates the parent had a single insertion). The sterile 2 PAN-RD and 1 PAN-RD pan-2 plants were used as pollen donors to fertilize emasculated wild type flowers, and the F1 seeds were tested as above. pap1(1.7-kb)-driven expression patterns were assayed in inflorescences of ethanol-induced pap1(1.7-kb)::alcr--alca::er-gfp lines treated with the water soluble lipophilic dye FM4-64 and imaged on a Zeiss 51 confocal microscope. Chromatin immunoprecipitation Chromatin immunoprecipitation (ChIP) experiments were performed according to published protocols (Ito et al., 1997). Inflorescences from plants mutant for the redundant APETALA1 and CAULIFLOWER genes, and expressing the dexamethasone-inducible 35S::APETALA-GR transgene (p35s::ap1-gr ap1-1 cal-1), were used. Plants were induced as described (Wellmer et al., 26) and tissue were collected 5-7 days later for a synchronized population of flowers. Inflorescences were ground in liquid nitrogen, resuspended in buffer M1 (1 mm phosphate buffer.1 M NaCl, 1 mm β-mercaptoethanol, 1 M hexylene glycol), fixed with 1% formaldehyde for 1 minutes, washed in buffers M2 [buffer M1 containing 1 mm MgCl 2,.5% (v/v) Triton X-1] and M3 (1 mm phosphate buffer,.1 M NaCl, 1 mm β-mercaptoethanol) and centrifuged to collect nuclei. Chromatin was isolated by resuspending the pellet in lysis buffer [1% SDS (w/v), 1 mm EDTA, 5 mm Tris-HCl ph 8.1] containing protease inhibitors (1 μg/ml Leupeptin, 15 μg/ml Aprotinin), incubating on ice for 1 minutes before adding ChIP dilution buffer (Upstate) supplemented with protease inhibitors. This mixture was sonicated and then pre-cleared with salmon sperm DNAtreated Protein A beads (Upstate) at 4 C for 1-3 hours. The solution was centrifuged, an aliquot of the supernatant kept for use as control for total input DNA (I) and the rest was incubated overnight with anti-pan antiserum (Chuang et al., 1999) that had been pre-cleared at 4 C for 2 days against leaves and inflorescences from pan mutant plants. Chromatin-antibody complexes were captured by incubating with a Protein A slurry at 4 C for 1 hour, centrifuging to remove unbound chromatin and eluting off the beads with elution buffer [1% SDS (w/v),.1 M NaHCO 3 ]. Histone-DNA crosslinks were reversed with.2 M NaCl at 65 C for 4 hours, RNA and protein were degraded with 4 μg/ml RNase and 4 μg/ml Proteinase K, respectively, and DNA was purified on a standard PCR purification column (Qiagen). This purified, antibody-bound DNA (B) was used to determine enrichment using quantitative real-time PCR on an ABI 79HT system (Applied Biosystems) with a MUTATOR-LIKE (MU) locus (At4g387) serving as control. Relative enrichment levels were calculated by determining the ratio of the mean quantity (calculated by the ABI software) of antibody-bound DNA (B) to total input DNA (I) for the control primers (B CTRL /I CTRL ) as well as for each experimental primer pair (B EXP /I EXP ), and then normalizing the ratio of the experimental value to the control value [(B EXP /I EXP )/(B CTRL /I CTRL )]. Particle bombardments Three milligrams of one micron gold microcarrier particles were coated with 2.5 μg of the p3 AGi reporter construct alone (for the control) or with an additional 2.5 μg of the p35s::pan-vp16 (for the co-bombardments). DNA was premixed prior to each coating. Equal aliquots of the coated particles were then placed onto macrocarriers and bombarded onto onion epidermal cells using a PDS-1/He Biolistic Particle Delivery System (Bio-Rad) and 11-psi rupture discs. The onion cells were incubated at 25 C for 2-3 days and then visualized for GUS staining using standard protocols. RESULTS AND DISCUSSION A genetic screen uncovers a combined role for LEAFY and PERIANTHIA in floral stem cell regulation In a mutagenesis experiment designed to identify modifiers of the floral organ number regulator PERIANTHIA (PAN) (Chuang et al., 1999; Running and Meyerowitz, 1996), we isolated a new lfy allele (see Materials and methods), which we named lfy-31. When compared with wild-type flowers (Fig. 1A), lfy single mutant flowers (of either lfy-31 or the well-described lfy-6 allele) bear organs that resemble leaf-like or sepal-like structures instead of sepals, petals or stamens; and sepalloid organs with stigmatic papillae instead of carpels (Fig. 1B,C). However, flowers of pan-3 mutants, which we used in the mutagenesis experiment, show no defects in floral identity, but rather bear increased numbers of sepals and petals, and reduced numbers of stamens (Fig. 1D). Flowers of the pan-3 lfy-31 double mutant line bear several notable differences from flowers of either single mutant. First, the overall number of primary organs is slightly increased with respect to lfy flowers (Table 1). Second, whereas the carpelloid structures of lfy mutant flowers are fully or partially fused (Fig. 1B,C), those of pan lfy flowers remain unfused (Fig. 1E,F). Third, ovule-like structures are often visible within carpels of lfy mutants (Fig. 1G) but only very rarely in the pan lfy double mutant (Fig. 1F). Finally, whereas all lfy flowers produce a determinate number of organs (Weigel et al., 1992), 89% (n=38) of pan lfy flowers are indeterminate, such that ectopic floral structures continue to develop interior to the fourth whorl organs (Table 1; Fig. 1F). As a further test of this interaction, we generated lines doubly mutant for pan-3 and the weak lfy-5 allele
3 Floral stem cell regulation RESEARCH REPORT 167 Fig. 1. pan lfy double mutant flowers are indeterminate and fail to downregulate WUS. (A-G) Phenotypes of pan and lfy single and double mutant flowers. (A) Wild-type flower bearing four sepals, four petals, six stamens and two fused carpels in whorls one to four, respectively. (B) lfy-31 flower with sepal-like organs in whorls one to three and fused sepalloid/carpelloid organs (arrow) in whorl four. (C) Flower of the well-characterized strong lfy-6 allele displaying similar phenotypes to lfy-31. (D) pan-3 flower bears extra sepals and petals but no carpel defects. (E) pan-3 lfy-31 double mutant flower with sepallike organs in whorls one to three and unfused sepalloid/carpelloid organs (arrow) in whorl four. (F) The same pan lfy flower as in E, with several organs removed to expose the ectopic floral structures growing within (arrowhead). The approximate position of the removed fourth-whorl organs is indicated (dotted line). (G) The same lfy flower as in B, dissected to reveal a relatively normal gynoecium. (H-M) In situ localization of WUS transcript in stage 2 flowers (H-J), or in stage 7 or older flowers (K-M), of pan-3 (H,K); lfy-6 (I,L); or pan-3 lfy-31 (J,M) plants. (H-J) Early WUS expression (arrows) is similar in all three genotypes. (K-M) In older flowers, no WUS expression is observed at the base of the gynoecium (arrowheads) of pan (K) and lfy (L) single mutants, but remains strong in pan lfy flowers (M) of a similar stage. Scale bars: 5 μm. The role of LFY in the center of the floral meristem, via the activation of AG expression, has been well studied. In addition, three lines of evidence suggest that PAN might also be active in this region. First, when certain alleles of pan (such as pan-3) are grown under specific culture conditions (see Materials and methods), some flowers (13%, n=84) show slight indeterminacy (see Fig. S5 in the supplementary material). Second, flowers from plants doubly mutant for pan and crabs claw (the carpel patterning gene) are indeterminate, with a reiteration of carpel structures in internal whorls (see Fig. S6C-F in the supplementary material; Y. Eshed and J. Bowman, personal communication). Third, pan mutations restore fourth-whorl carpels to flowers of the superman-1 (sup-1) single mutant that normally either lack carpels or have staminoid carpels, also suggesting a role in this domain (see Fig. S6G-I in the supplementary material) (Running and Meyerowitz, 1996). Because these data reveal a function for PAN in the presumptive fourth whorl, we hypothesized that the determinacy defects apparent in lfy pan double mutant flowers were due to a hidden role for PAN in regulating the floral stem cell population, which lies within the fourth whorl. A dominant-negative pan allele induces floral indeterminacy by suppressing AG expression One explanation for the absence of floral indeterminacy phenotypes in most pan mutant flowers is that this activity of PAN might be masked by functional redundancy with other factors. To overcome Table 1. Identities and numbers of organs in lfy and pan mutant flowers* Mutant pan-3 lfy-6 lfy-31 pan-3 Sepals/sepal-like Petals Stamens Carpels/sepalloid-carpels Flowers with interior organs (%) n 5.1±.7 9.8± ± ±.6 5.3± ±.5 6.9± *lfy-6 and lfy-31 pan-3 flowers were later-arising structures with floral and secondary inflorescence characteristics. (Weigel et al., 1992), and observed that these flowers also bear unfused carpels and are indeterminate (see Fig. S1 in the supplementary material). Thus, in plants carrying mutations in both LFY and PAN, there is an apparent loss of floral determinacy that is not observed in the single mutants alone. To test the molecular basis of the fourth-whorl phenotypes in pan lfy double mutant flowers, we performed in situ hybridization experiments to determine the expression dynamics of the WUS transcript, as the downregulation of WUS in the stem cell organizing center is essential for floral determinacy (Mayer et al., 1998). As in the case of wild-type flowers (Mayer et al., 1998), WUS mrna is clearly detectable in the center of very early flower buds of pan-3 and lfy-6 single mutants, as well as in pan-3 lfy-31 double mutants (Fig. 1H-J). In the wild type, WUS mrna becomes undetectable by approximately stage 7, when the carpel primordia first appear (Mayer et al., 1998). Similarly, in pan-3 and lfy-6 single mutant flowers, WUS expression is absent in later-stage flowers (Fig. 1K,L; see Figs S2 and S3 in the supplementary material). By contrast, in older pan lfy double mutant flowers, WUS expression is clearly detectable within the broad expanse of tissue at the center of the flower from where the organs of the interior floral structures will arise (Fig. 1M; see Fig. S4 in the supplementary material). These results indicate that the indeterminacy phenotypes of pan lfy flowers are associated with the persistence of the stem cell pool, as revealed by the continued expression of WUS in the stem cell organizing center.
4 168 RESEARCH REPORT Development 136 (1) Fig. 2. A dominant-negative PAN chimera induces floral determinacy phenotypes. (A) pap1(1.7-kb)::alcr--alca::er-gfp expression in an inflorescence meristem. GFP signal (green) is observed in the central domes (arrowheads) of all visible flowers after induction with ethanol vapors. Autofluorescence from the shoot apical meristem is visible (red). (B-H) Floral phenotypes of PAN wild-type or mutant plants harboring one or more copies of PAN-RD under the control of the AP1(1.7-kb) promoter. Red and green bars indicate the number of copies of PAN-RD or wild-type PAN, respectively. Open bar indicates no copies, half-filled indicates one copy and filled indicates two copies (see key, bottom right). (B) pan-2 mutant flower. (C) pan flower harboring one copy of PAN-RD displaying amplified pan-like phenotypes, as well as additional phenotypes such as extra carpels (arrow) and severe floral indeterminacy. (D) Side view of the flower in C with some organs removed to reveal ectopic floral structures (arrow) developing interior to the fourth whorl (demarcated by a dashed line). (E) A pan-like phenotype is observed in flowers arising from a cross of genotype in C to wild type. This flower harbors one copy of PAN-RD and is heterozygous at the PAN locus. (F) Wild-type flower. (G) Wild-type flower harboring one copy of PAN-RD displaying a pan-like phenotype. (H) Flower from progeny of the plant in G harboring two copies of PAN-RD presenting strong indeterminacy defects, similar to PAN-RD/+ pan plants (C). (I-K) In situ localizations of AG transcript in stage 5-6 flowers of pan-2 (I) or PAN-RD/+ pan-2 (J,K) plants. AG localization appears unperturbed in pan flowers (I), but in PAN-RD/+ pan-2 flowers a central region within the AG domain shows diminished signal intensity. Also marked are sepals (dashed arrow) that do not express AG and early stamen primordia (arrowhead) that do. (K) Magnified view of the dashed box in J. Scale bars: 1 μm. such redundancy, we generated a constitutively repressing form of PAN by fusing it to the SUPERMAN repressor domain motif (Hiratsu et al., 23). Such constructions have been used to study bzip factors in several model systems (Fukazawa et al., 2; Rieping et al., 1994) and, in Arabidopsis, SUPERMAN repressor domain (RD) fusions of several transcription factors have been shown to phenocopy their corresponding loss-of-function mutants (Baudry et al., 26; Hiratsu et al., 23; Xu et al., 26). The expression of such a PAN fusion, either ubiquitously or from the endogenous promoter, yielded plants with severe growth defects, thereby masking any floral phenotypes (data not shown). In order to assay its effects on floral patterning, we thus used the flower-specific APETALA1 (AP1) promoter (Hempel et al., 1997). A 1.7 kb upstream fragment of the AP1 locus drives expression throughout the early flower; in addition, and at variance with the normal AP1 expression pattern, it remains active in the entire flower, presumably due to the absence of additional regulatory elements (Fig. 2A; see Fig. S7 in the supplementary material; M. Yanofsky, personal communication). In order to remove the effects of competition between PAN-RD and endogenous PAN, we introduced the pap1(1.7-kb)::pan-rd construct (hereafter referred to as PAN-RD) into pan-2 mutant plants (Fig. 2B). The majority (82%) of these PAN-RD/+; pan/pan primary transformants bore flowers with increased numbers of petals and stamens (6.3±.6 and 5.8±.4 respectively, n=2), secondary flowers in the axils of first-whorl organs (2.8±.6 versus in the wild type and pan), extra carpels in the fourth whorl (3.5±.5) and severe indeterminacy (Fig. 2C,D). The other 18% (n=85) of transformants showed a slight enhancement of the pan organ number phenotypes without any determinacy defects (data not shown). To eliminate the possibility that the indeterminacy phenotypes were caused by the use of the AP1(1.7-kb) promoter fragment, we used an ethanol-inducible two component system (Deveaux et al., 23; Maizel and Weigel, 24) to drive PAN-RD expression under the endogenous PAN promoter. In the absence of induction, pan-2 plants bearing this construct show no fourth-whorl phenotypes (see Fig. S8A in the supplementary material). However, after induction with ethanol, we observe flowers bearing unfused gynoecia and ectopic internal carpel structures (see Fig. S8B in the supplementary material). Thus, expression of a PAN-repressor domain fusion protein in the flower leads to the loss of floral determinacy, a phenotype observed at low frequency in pan mutant plants grown in specific culture conditions (see above). A caveat to the use of dominant-negative alleles is that they may act as neomorphs, altering the expression of ectopic, rather than genuine, downstream targets of the protein under study. We decided to study this by varying the ratio of chimeric to wild-type protein. If PAN-RD behaves as a true dominant-negative allele, increasing doses of it should yield increasingly stronger phenotypes, whereas increasing doses of unmodified PAN should attenuate the phenotypes. We first
5 Floral stem cell regulation RESEARCH REPORT 169 used pollen from the PAN-RD/+; pan/pan flowers described above in a cross to wild type and examined the phenotypes of F1 plants selected for the presence of the transgene. We found that these flowers (PAN- RD/+; pan/+) had phenotypes similar to the pan mutant (compare Fig. 2E with 2B). Thus the same PAN-RD insertion that confers strong indeterminacy phenotypes on pan flowers does not do so on pan/+ heterozygotes (instead yielding only the more sensitive organ number defect), showing that PAN-RD expression does not ectopically induce floral indeterminacy. Next we examined primary transformants in wild-type plants, thus with two additional wild-type copies of PAN (Fig. 2F). Approximately 2% (n=6) of these plants (genotypically PAN-RD/+; PAN/PAN) phenocopied the pan mutant (Fig. 2G; 4.3±.5 sepals, 4.9±.3 petals and 4.7±.5 stamens; n=2), whereas the rest showed no discernable phenotypes. We then examined the progeny of these plants, to determine the phenotypes of plants harboring two copies of the transgene (see Materials and methods). These flowers (PAN-RD/PAN-RD; PAN/PAN) displayed strong phenotypes, including extra carpels and floral indeterminacy (Fig. 2H), and closely resembled PAN-RD/+; pan/pan flowers (Fig. 2C,D). Taken together, these data show that the effects of the PAN-RD fusion protein are modified by wild-type PAN in a dosage sensitive manner. This suggests that PAN-RD and endogenous PAN compete for the same targets and that the PAN-RD phenotypes are due to the repression of genuine PAN targets. To characterize the molecular basis of the PAN-RD phenotype, we performed in situ hybridizations to determine whether the PAN-RD transgene induced changes in AG expression patterns. We observed that, as in the wild type, AG is expressed uniformly throughout the third and fourth whorls of stage 5-6 pan-2 flowers (Fig. 2I). However, in similarly staged PAN-RD/+; pan/pan flowers, AG expression is patchy, with the central region of the floral meristem showing reduced expression (Fig. 2J,K). Since pap1(1.7-kb) drives expression throughout the central dome of the flower during these stages, these results indicate that the PAN-RD chimera might exert an unequal influence on different cells within the AG-expressing region. Thus our results show that expression of a PAN-RD fusion protein in the flower is sufficient to mimic pan loss-of-function phenotypes and to produce floral indeterminacy. Since the indeterminacy phenotype occurs more stably in PAN-RD plants than in pan simple mutants, this role possibly requires other spatially or temporally restricted factors. Taken together with the phenotypes of the double mutant flowers described above, this suggests that PAN plays a role in the development of the fourth whorl, specifically in the proper regulation of the floral stem cell population, and that this role is achieved through the regulation of AG. Furthermore, as the effects of PAN-RD were similar to the loss-of-function phenotypes of pan mutant alleles, the role of PAN in floral determinacy is likely to be that of an activator of a gene involved in the process. PAN binds AG regulatory sequences in vivo We next sought to determine the precise mechanism by which PAN affects floral stem cell fate. As discussed above, AG is a key regulator of this process, by itself acting to repress WUS expression. Because AG expression is perturbed in PAN-RD flowers and because PAN is a predicted transcriptional activator, we reasoned that its role might be to positively regulate AG expression. To test whether PAN directly associates with the AG promoter, we used chromatin immunoprecipitation (ChIP) assays, which examine the in vivo binding of transcription factors to DNA. We maximized the sensitivity of our assays by using a synchronized population of flowers at stages 5-7 (Wellmer et al., 26) (see Materials and methods), as the stem cell organizing activity is known to be Fig. 3. PAN directly binds AGAMOUS regulatory sequences. (A) Structure of the 5.7 kb AG locus. Exons and introns (top) are represented in bold and dashed lines, respectively. The second intron (asterisk) contains all known AG regulatory elements. The detailed view of the 3 kb intron shows the fragments used to determine enrichment in the chromatin immunoprecipitation assays (blue bars), the 3 fragment used in co-bombardment experiments (red line), and evolutionarily conserved elements and known or predicted transcription factor binding motifs (Davies et al., 1999; Hong et al., 23; Lohmann et al., 21; Parcy et al., 1998). Black triangles, LFY/WUS binding sites; white triangle, predicted LFY binding site; stars, CArG boxes; circles, CCAAT boxes; diamond, AAGAAT motif; green squares, predicted core bzip binding sites; vertical tick marks: 2 bp intervals. (B) Results of chromatin immunoprecipitation experiments from stage 5-7 flowers. Anti-PAN antiserum was used to isolate protein-dna complexes and DNA enrichment levels were measured by quantitative real-time RT-PCR (see Materials and methods). Enrichment was calculated for antibodybound DNA relative to total input DNA and was then normalized against an internal genomic control. Vertical bars show mean enrichment levels from duplicate experiments for amplicons distributed along the intron (shown in 3A). The scale for amplicon H is different and is shown in red. Results are shown only for amplicons with a coefficient of correlation (r 2 )>.98. (C) Results of particle cobombardment experiments in onion epidermal cells. Vertical bars represent mean values from duplicate experiments for numbers of cells stained for GUS enzymatic activity. The pagi-3 ::GUS reporter, which recapitulates most facets of AG expression in vivo, shows some background activity (left bar) in onion epidermal cells. When cobombarded with p35s::pan-vp16 (right bar), the numbers of cells stained for GUS activity is 4- to 5-fold greater. Error bars represent standard deviation from the mean. terminated during this time. In addition, we used a characterized PAN-specific polyclonal antibody that, when used in immunohistochemical analyses, detects protein signals closely resembling PAN mrna expression patterns, but showing no signal in mutant flowers (Chuang et al., 1999). We further pre-cleared this antibody against tissue from pan-2 plants prior to immunoprecipitating intact protein-dna complexes. We then performed quantitative real-time RT-PCR to assay for the enrichment of sequences within the 3 kb second intron of the AG locus with respect to input DNA, where important cis regulatory sequences are located (Fig. 3A) (Busch et al., 1999; Deyholos and Sieburth, 2; Hong et al., 23; Lohmann et al., 21; Parcy et al., 1998; Sieburth and Meyerowitz, 1997). We observed that five amplicons out of a total of eight tested within this region are significantly enriched (Fig. 3B; amplicons B=9.2-fold±2., C=9.±.6, F=6.6±.7, G=4.9±.4 and H=4.5±9.2) when
6 161 RESEARCH REPORT Development 136 (1) normalized relative to internal genomic controls. These data demonstrate that PAN, either alone or in a complex, binds AG regulatory sequences in vivo. Because of the nature of ChIP assays, not every enriched fragment necessarily contains a PAN binding site. Four of the five ChIPenriched amplicons (B, C, F and H) map to regions previously described as being very highly conserved (Hong et al., 23). Of these, amplicon C contains binding sites for LFY and WUS (Lohmann et al., 21; Parcy et al., 1998), whereas predicted bzip core binding sites (Jakoby et al., 22) are located within or close to two others (F and H). Hong et al. (Hong et al., 23) have shown that an AG reporter containing a deletion within amplicon H loses expression at later floral stages, suggesting that it plays a role in the maintenance of AG expression. In an accompanying article (Maier et al., 29), the authors use one-hybrid assays to show that amplicon F contains PAN binding sites. Furthermore, they show that mutations in this bzip motif disrupt binding in vitro and eliminate in vivo expression in the context of an AG reporter. Given the very high enrichment of amplicon H in our ChIP assays, we were interested in determining whether this region might also contain PAN binding sites. To this end, we made use of a published 3 AG reporter construct that reproduces the normal AG expression pattern in vivo (Busch et al., 1999). In this reporter, an 8 bp 3 pag fragment (Fig. 3A) drives expression of the uida gene, which encodes the β-glucuronidase (GUS) enzyme (Busch et al., 1999). We performed particle co-bombardment experiments in onion epidermal cells with a putative constitutively activated form of PAN (p35s::pan-vp16). Although the reporter alone presents basal reporter activity in onion cells (44.5±34.6 cells; Fig. 3C; see Fig. S9A,C in the supplementary material), perhaps due to the presence of minimal p35s sequences, 4- to 5-fold greater numbers of cells (157±11.3; Fig. 3C; see Fig. S9B,D in the supplementary material) show GUS activity when co-bombarded with p35s::pan-vp16. It is thus possible that PAN has two or more target sites within the AG promoter, to which it might bind with different affinities or different partners. Flowers mutant for AG display stem cell overproliferation phenotypes and, in addition, also show homeotic transformations of stamens to petals (Bowman et al., 1989). Since PAN-RD disrupts floral stem cell regulation without inducing the homeotic transformations associated with the loss of AG function, we asked whether PAN might function as a general regulator of AG, or only to modify its activity in the fourth whorl. To this end, we used the weak ag-4 allele, which produces flowers with reduced numbers of stamens and with fourth whorl organs replaced by another flower (Fig. 4A) (Sieburth et al., 1995). Mutations in several loci, including HUA1 and HUA2, REBELOTE (RBL) or ULTRAPETALA1 (ULT1) enhance the ag-4 allele by fully or partially converting stamens to petals (Chen and Meyerowitz, 1999; Fletcher, 21; Prunet et al., 28). We reasoned that if PAN plays a general role in regulating AG expression, a pan allele might similarly enhance the weak ag phenotype. Conversely, if PAN participates primarily in the floral determinacy aspects of AG, the double mutant might not be significantly enhanced. In fact, we observe that pan-2 ag-4 double mutant flowers (Fig. 4B) differ from ag-4 single mutants only in that they present an additional pan-like phenotype: extra perianth organs (4.8±.5 sepals and 4.8±.5 petals compared with four each in ag- 4; n=2 for both genotypes), as is the case for double mutants of pan and the strong ag-1 allele (Running and Meyerowitz, 1996). The third-whorl stamens of pan-2 ag-4 flowers appear morphologically normal, although they are reduced in number (5.5±.5 compared with 5.8±.4 in ag-4). This suggests either that the stamen identity Fig. 4. pan mutations do not enhance the third-whorl phenotypes of a weak ag allele. (A) Flower of the weak ag-4 allele in a mixed background of wild-type accessions (L-er/Ws). Such flowers have normal outer organs but fourth-whorl carpels are replaced with a new flower (arrow). (B)A pan-2 ag-4 flower in the same genetic background has the altered floral organ numbers of the pan mutant and the new fourth-whorl flower phenotype of ag-4 (arrow). functions of the mutant protein encoded by the ag-4 allele are robust and are not perturbed by the absence of PAN, or that PAN regulates AG expression only in the fourth whorl. Prunet et al. (Prunet et al., 28) have recently proposed the existence of a distinct subdomain within the floral fourth whorl, in which a decrease in AG expression is sufficient to disrupt the specification of floral determinacy. Two observations indicate that the effects of PAN on the AG promoter might be spatially restricted, perhaps to this subdomain. First, in PAN-RD flowers, there is a marked reduction in the accumulation of AG transcript at the very center of the dome of the floral meristem. Second, unlike many other AG interactors (Chen and Meyerowitz, 1999; Fletcher, 21; Prunet et al., 28), pan mutations have no effect on the third whorl phenotypes of a weak ag allele. A restricted effect of PAN on AG expression might explain why mutations in PAN attenuate the fourth whorl phenotype of sup mutants (Running and Meyerowitz, 1996), which produce reduced or masculinized carpels (see Fig. S6G,H in the supplementary material) (Bowman et al., 1992). One model is that in the center of the fourth whorl of pan sup double mutant flowers (see Fig. S6I in the supplementary material), even the mild reduction in AG expression caused by the absence of PAN could lead to increased or prolonged WUS expression, and thus to an increase in the size of this region. Since the stamen identity genes AP3 and PI are excluded from the very center of sup flowers (Bowman et al., 1992; Goto and Meyerowitz, 1994), the corresponding, but enlarged, region in pan sup flowers could now become specified into a functional carpel. Like PAN, the broadly expressed APETALA2 (AP2) gene, the absence of which causes only specific floral phenotypes (a change in sepal and petal identities), was recently shown, through the characterization of a semi-dominant allele, to play a role in the control of stem cells in the shoot (Würschum et al., 26). Thus both PAN and AP2 have primary roles in specific aspects of floral patterning, but also have masked functions in stem cell regulation that are likely to require interactions with other domain- and/or stage-specific factors. Identifying these interactors through genetic screens or other methods could prove invaluable in gaining a full understanding of the complex regulatory mechanisms that control stem cell fate. As the main function of AG in floral determinacy appears to be to downregulate WUS expression in a narrow temporal window, AG expression must be quickly upregulated in order to ensure the complete arrest of stem cell fate, and thereby ensure proper floral
7 Floral stem cell regulation RESEARCH REPORT 1611 patterning. We suggest that plants have evolved a complex, multiply redundant system to ensure the proper regulation of AG, and furthermore, that many of the factors involved in the regulation of floral stem cells probably also perform unrelated patterning functions. We thank J. Lohmann for sharing unpublished results; Arnavaz Garda, Alexis Lacroix, Claudia Bardoux and Hervé Leyral for technical assistance; Ioan Negrutiu, Christophe Trehin, Nathanaël Prunet and Olivier Hamant for useful discussions and comments on the manuscript; Ioan Negrutiu for sharing unpublished results and seeds; and John Bowman and Marty Yanofsky for communicating unpublished results. This work was supported by a Caltech Division of Biology postdoctoral fellowship (to P.D.), a Marie Curie Incoming International Fellowship IIF-222 (to P.D.), the EU Marie Curie SY-STEM Network (to J.T.) and a NSF Grant IOS (to E.M.M.). Supplementary material Supplementary material available online at References Baudry, A., Caboche, M. and Lepiniec, L. (26). TT8 controls its own expression in a feedback regulation involving TTG1 and homologous MYB and bhlh factors, allowing a strong and cell-specific accumulation of flavonoids in Arabidopsis thaliana. Plant J. 46, Bowman, J. L., Smyth, D. R. and Meyerowitz, E. M. (1989). Genes directing flower development in Arabidopsis. Plant Cell 1, Bowman, J. L., Sakai, H., Jack, T., Weigel, D., Mayer, U. and Meyerowitz, E. M. (1992). SUPERMAN, a regulator of floral homeotic genes in Arabidopsis. Development 114, Busch, M. A., Bomblies, K. and Weigel, D. (1999). Activation of a floral homeotic gene in Arabidopsis. Science 285, Chen, X. and Meyerowitz, E. M. (1999). HUA1 and HUA2 are two members of the floral homeotic AGAMOUS pathway. Mol. Cell 3, Chuang, C. F., Running, M. P., Williams, R. W. and Meyerowitz, E. M. (1999). The PERIANTHIA gene encodes a bzip protein involved in the determination of floral organ number in Arabidopsis thaliana. Genes Dev. 13, Davies, B., Motte, P., Keck, E., Saedler, H., Sommer, H. and Schwarz- Sommer, Z. (1999). PLENA and FARINELLI: redundancy and regulatory interactions between two Antirrhinum MADS-box factors controlling flower development. EMBO J. 18, Deveaux, Y., Peaucelle, A., Roberts, G. R., Coen, E., Simon, R., Mizukami, Y., Traas, J., Murray, J. A., Doonan, J. H. and Laufs, P. (23). The ethanol switch: a tool for tissue-specific gene induction during plant development. Plant J. 36, Deyholos, M. K. and Sieburth, L. E. (2). Separable whorl-specific expression and negative regulation by enhancer elements within the AGAMOUS second intron. Plant Cell 12, Eshed, Y., Baum, S. F., Perea, J. V. and Bowman, J. L. (21). Establishment of polarity in lateral organs of plants. Curr. Biol. 11, Fletcher, J. C. (21). The ULTRAPETALA gene controls shoot and floral meristem size in Arabidopsis. Development 128, Fukazawa, J., Sakai, T., Ishida, S., Yamaguchi, I., Kamiya, Y. and Takahashi, Y. (2). REPRESSION OF SHOOT GROWTH, a bzip transcriptional activator, regulates cell elongation by controlling the level of gibberellins. Plant Cell 12, Gleave, A. P. (1992). A versatile binary vector system with a T-DNA organisational structure conducive to efficient integration of cloned DNA into the plant genome. Plant Mol. Biol. 2, Goto, K. and Meyerowitz, E. M. (1994). Function and regulation of the Arabidopsis floral homeotic gene PISTILLATA. Genes Dev. 8, Hempel, F. D., Weigel, D., Mandel, M. A., Ditta, G., Zambryski, P. C., Feldman, L. J. and Yanofsky, M. F. (1997). Floral determination and expression of floral regulatory genes in Arabidopsis. Development 124, Hiratsu, K., Matsui, K., Koyama, T. and Ohme-Takagi, M. (23). Dominant repression of target genes by chimeric repressors that include the EAR motif, a repression domain, in Arabidopsis. Plant J. 34, Hong, R. L., Hamaguchi, L., Busch, M. A. and Weigel, D. (23). Regulatory elements of the floral homeotic gene AGAMOUS identified by phylogenetic footprinting and shadowing. Plant Cell 15, Ito, T., Takahashi, N., Shimura, Y. and Okada, K. (1997). A serine/threonine protein kinase gene isolated by an in vivo binding procedure using the Arabidopsis floral homeotic gene product, AGAMOUS. Plant Cell Physiol. 38, Jakoby, M., Weisshaar, B., Droge-Laser, W., Vicente-Carbajosa, J., Tiedemann, J., Kroj, T. and Parcy, F. (22). bzip transcription factors in Arabidopsis. Trends Plant Sci. 7, Laux, T., Mayer, K. F., Berger, J. and Jurgens, G. (1996). The WUSCHEL gene is required for shoot and floral meristem integrity in Arabidopsis. Development 122, Lenhard, M., Bohnert, A., Jurgens, G. and Laux, T. (21). Termination of stem cell maintenance in Arabidopsis floral meristems by interactions between WUSCHEL and AGAMOUS. Cell 15, Lohmann, J. U., Hong, R. L., Hobe, M., Busch, M. A., Parcy, F., Simon, R. and Weigel, D. (21). A molecular link between stem cell regulation and floral patterning in Arabidopsis. Cell 15, Maier, A. T., Stehling-Sun, S., Wollmann, H., Demar, M., Hong, R. L., Haubeiß, S., Weigel, D. and Lohmann, J. U. (29). Dual roles of the bzip transcription factor PERIANTHIA in the control of floral architecture and homeotic gene expression. Development 136, Maizel, A. and Weigel, D. (24). Temporally and spatially controlled induction of gene expression in Arabidopsis thaliana. Plant J. 38, Mayer, K. F., Schoof, H., Haecker, A., Lenhard, M., Jurgens, G. and Laux, T. (1998). Role of WUSCHEL in regulating stem cell fate in the Arabidopsis shoot meristem. Cell 95, Mizukami, Y. and Ma, H. (1995). Separation of AG function in floral meristem determinacy from that in reproductive organ identity by expressing antisense AG RNA. Plant Mol. Biol. 28, Parcy, F., Nilsson, O., Busch, M. A., Lee, I. and Weigel, D. (1998). A genetic framework for floral patterning. Nature 395, Prunet, N., Morel, P., Thierry, A. M., Eshed, Y., Bowman, J. L., Negrutiu, I. and Trehin, C. (28). REBELOTE, SQUINT, and ULTRAPETALA1 function redundantly in the temporal regulation of floral meristem termination in Arabidopsis thaliana. Plant Cell 2, Rieping, M., Fritz, M., Prat, S. and Gatz, C. (1994). A dominant negative mutant of PG13 suppresses transcription from a cauliflower mosaic virus 35S truncated promoter in transgenic tobacco plants. Plant Cell 6, Running, M. P. and Meyerowitz, E. M. (1996). Mutations in the PERIANTHIA gene of Arabidopsis specifically alter floral organ number and initiation pattern. Development 122, Schultz, E. A. and Haughn, G. W. (1991). LEAFY, a homeotic gene that regulates inflorescence development in Arabidopsis. Plant Cell 3, Sieburth, L. E. and Meyerowitz, E. M. (1997). Molecular dissection of the AGAMOUS control region shows that cis elements for spatial regulation are located intragenically. Plant Cell 9, Sieburth, L. E., Running, M. P. and Meyerowitz, E. M. (1995). Genetic separation of third and fourth whorl functions of AGAMOUS. Plant Cell 7, Weigel, D. and Meyerowitz, E. M. (1993). Activation of Floral Homeotic Genes in Arabidopsis. Science 261, Weigel, D., Alvarez, J., Smyth, D. R., Yanofsky, M. F. and Meyerowitz, E. M. (1992). LEAFY controls floral meristem identity in Arabidopsis. Cell 69, Wellmer, F., Alves-Ferreira, M., Dubois, A., Riechmann, J. L. and Meyerowitz, E. M. (26). Genome-wide analysis of gene expression during early Arabidopsis flower development. PLoS Genet. 2, e117. Würschum, T., Gross-Hardt, R. and Laux, T. (26). APETALA2 regulates the stem cell niche in the Arabidopsis shoot meristem. Plant Cell 18, Xu, Y., Teo, L. L., Zhou, J., Kumar, P. P. and Yu, H. (26). Floral organ identity genes in the orchid Dendrobium crumenatum. Plant J. 46,
Activation of a Floral Homeotic Gene in Arabidopsis
 Activation of a Floral Homeotic Gene in Arabidopsis Busch, M.A., Bomblies, K., Weigel, D. kuleuven-kortrijk.be Simon Olthof Louie Christodoulidis 1 In Arabidopsis flowers, appearance is based, in part,
Activation of a Floral Homeotic Gene in Arabidopsis Busch, M.A., Bomblies, K., Weigel, D. kuleuven-kortrijk.be Simon Olthof Louie Christodoulidis 1 In Arabidopsis flowers, appearance is based, in part,
Fig. S1. TPL and TPL N176H protein interactions. (A) Semi-in vivo pull-down assays using recombinant GST N-TPL and GST N-TPL N176H fusions and
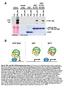 Fig. S1. TPL and TPL N176H protein interactions. (A) Semi-in vivo pull-down assays using recombinant GST N-TPL and GST N-TPL N176H fusions and transgenic Arabidopsis TPL-HA lysates. Immunoblotting of input
Fig. S1. TPL and TPL N176H protein interactions. (A) Semi-in vivo pull-down assays using recombinant GST N-TPL and GST N-TPL N176H fusions and transgenic Arabidopsis TPL-HA lysates. Immunoblotting of input
A Repressor Complex Governs the Integration of
 Developmental Cell 15 Supplemental Data A Repressor Complex Governs the Integration of Flowering Signals in Arabidopsis Dan Li, Chang Liu, Lisha Shen, Yang Wu, Hongyan Chen, Masumi Robertson, Chris A.
Developmental Cell 15 Supplemental Data A Repressor Complex Governs the Integration of Flowering Signals in Arabidopsis Dan Li, Chang Liu, Lisha Shen, Yang Wu, Hongyan Chen, Masumi Robertson, Chris A.
Activation of a Floral Homeotic Gene in Arabidopsis
 Activation of a Floral Homeotic Gene in Arabidopsis By Maximiliam A. Busch, Kirsten Bomblies, and Detlef Weigel Presentation by Lis Garrett and Andrea Stevenson http://ucsdnews.ucsd.edu/archive/graphics/images/image5.jpg
Activation of a Floral Homeotic Gene in Arabidopsis By Maximiliam A. Busch, Kirsten Bomblies, and Detlef Weigel Presentation by Lis Garrett and Andrea Stevenson http://ucsdnews.ucsd.edu/archive/graphics/images/image5.jpg
Abcam.com. hutton.ac.uk. Ipmdss.dk. Bo Gong and Eva Chou
 Abcam.com Bo Gong and Eva Chou Ipmdss.dk hutton.ac.uk What is a homeotic gene? A gene which regulates the developmental fate of anatomical structures in an organism Why study them? Understand the underlying
Abcam.com Bo Gong and Eva Chou Ipmdss.dk hutton.ac.uk What is a homeotic gene? A gene which regulates the developmental fate of anatomical structures in an organism Why study them? Understand the underlying
The homeo'c gene AGAMOUS (AG) in Arabidopsis. Presenta'on by: Yang Liu and Zhonghang Zhang (Daisy) February 27th, 2018
 The homeo'c gene AGAMOUS (AG) in Arabidopsis Presenta'on by: Yang Liu and Zhonghang Zhang (Daisy) February 27th, 2018 Arabidopsis Flower George Haughn, UBC Genes Direc'ng Flower Development in Arabidopsis
The homeo'c gene AGAMOUS (AG) in Arabidopsis Presenta'on by: Yang Liu and Zhonghang Zhang (Daisy) February 27th, 2018 Arabidopsis Flower George Haughn, UBC Genes Direc'ng Flower Development in Arabidopsis
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:10.1038/nature10928 Materials and Methods 1. Plant material and growth conditions. All plant lines used were in Col-0 background unless otherwise specified. pif4-101 mutant
SUPPLEMENTARY INFORMATION doi:10.1038/nature10928 Materials and Methods 1. Plant material and growth conditions. All plant lines used were in Col-0 background unless otherwise specified. pif4-101 mutant
Supplemental Data. Noncoding Transcription by RNA Polymerase Pol IVb/Pol V Mediates Transcriptional Silencing of Overlapping and Adjacent Genes
 Cell, Volume 135 Supplemental Data Noncoding Transcription by RNA Polymerase Pol IVb/Pol V Mediates Transcriptional Silencing of Overlapping and Adjacent Genes Andrzej T. Wierzbicki, Jeremy R. Haag, and
Cell, Volume 135 Supplemental Data Noncoding Transcription by RNA Polymerase Pol IVb/Pol V Mediates Transcriptional Silencing of Overlapping and Adjacent Genes Andrzej T. Wierzbicki, Jeremy R. Haag, and
Sperm cells are passive cargo of the pollen tube in plant fertilization
 In the format provided by the authors and unedited. SUPPLEMENTARY INFORMATION VOLUME: 3 ARTICLE NUMBER: 17079 Sperm cells are passive cargo of the pollen tube in plant fertilization Jun Zhang 1, Qingpei
In the format provided by the authors and unedited. SUPPLEMENTARY INFORMATION VOLUME: 3 ARTICLE NUMBER: 17079 Sperm cells are passive cargo of the pollen tube in plant fertilization Jun Zhang 1, Qingpei
Schematic representation of the endogenous PALB2 locus and gene-disruption constructs
 Supplementary Figures Supplementary Figure 1. Generation of PALB2 -/- and BRCA2 -/- /PALB2 -/- DT40 cells. (A) Schematic representation of the endogenous PALB2 locus and gene-disruption constructs carrying
Supplementary Figures Supplementary Figure 1. Generation of PALB2 -/- and BRCA2 -/- /PALB2 -/- DT40 cells. (A) Schematic representation of the endogenous PALB2 locus and gene-disruption constructs carrying
AGAMOUS Terminates Floral Stem Cell Maintenance in Arabidopsis by Directly Repressing WUSCHEL through Recruitment of Polycomb Group Proteins W
 The Plant Cell, Vol. 23: 3654 3670, October 2011, www.plantcell.org ã 2011 American Society of Plant Biologists. All rights reserved. AGAMOUS Terminates Floral Stem Cell Maintenance in Arabidopsis by Directly
The Plant Cell, Vol. 23: 3654 3670, October 2011, www.plantcell.org ã 2011 American Society of Plant Biologists. All rights reserved. AGAMOUS Terminates Floral Stem Cell Maintenance in Arabidopsis by Directly
1. Cross-linking and cell harvesting
 ChIP is a powerful tool that allows the specific matching of proteins or histone modifications to regions of the genome. Chromatin is isolated and antibodies to the antigen of interest are used to determine
ChIP is a powerful tool that allows the specific matching of proteins or histone modifications to regions of the genome. Chromatin is isolated and antibodies to the antigen of interest are used to determine
This contribution is part of the special series of Inaugural Articles by members of the National Academy of Sciences elected on April 25, 1995.
 Proc. Natl. Acad. Sci. USA Vol. 93, pp. 4063 4070, April 1996 Plant Biology This contribution is part of the special series of Inaugural Articles by members of the National Academy of Sciences elected
Proc. Natl. Acad. Sci. USA Vol. 93, pp. 4063 4070, April 1996 Plant Biology This contribution is part of the special series of Inaugural Articles by members of the National Academy of Sciences elected
Supplementary Information: Materials and Methods. Immunoblot and immunoprecipitation. Cells were washed in phosphate buffered
 Supplementary Information: Materials and Methods Immunoblot and immunoprecipitation. Cells were washed in phosphate buffered saline (PBS) and lysed in TNN lysis buffer (50mM Tris at ph 8.0, 120mM NaCl
Supplementary Information: Materials and Methods Immunoblot and immunoprecipitation. Cells were washed in phosphate buffered saline (PBS) and lysed in TNN lysis buffer (50mM Tris at ph 8.0, 120mM NaCl
A Novel Single-Nucleotide Mutation in a CLAVATA3 Gene Homolog Controls a Multilocular Silique Trait in Brassica rapa L.
 Molecular Plant 7, 1788 1792, December 2014 LETTER TO THE EDITOR A Novel Single-Nucleotide Mutation in a CLAVATA3 Gene Homolog Controls a Multilocular Silique Trait in Brassica rapa L. Dear Editor, Brassica
Molecular Plant 7, 1788 1792, December 2014 LETTER TO THE EDITOR A Novel Single-Nucleotide Mutation in a CLAVATA3 Gene Homolog Controls a Multilocular Silique Trait in Brassica rapa L. Dear Editor, Brassica
Supplementary data. sienigma. F-Enigma F-EnigmaSM. a-p53
 Supplementary data Supplemental Figure 1 A sienigma #2 sienigma sicontrol a-enigma - + ++ - - - - - - + ++ - - - - - - ++ B sienigma F-Enigma F-EnigmaSM a-flag HLK3 cells - - - + ++ + ++ - + - + + - -
Supplementary data Supplemental Figure 1 A sienigma #2 sienigma sicontrol a-enigma - + ++ - - - - - - + ++ - - - - - - ++ B sienigma F-Enigma F-EnigmaSM a-flag HLK3 cells - - - + ++ + ++ - + - + + - -
Supplemental Data. Guo et al. (2015). Plant Cell /tpc
 Supplemental Figure 1. The Mutant exb1-d Displayed Pleiotropic Phenotypes and Produced Branches in the Axils of Cotyledons. (A) Branches were developed in exb1-d but not in wild-type plants. (B) and (C)
Supplemental Figure 1. The Mutant exb1-d Displayed Pleiotropic Phenotypes and Produced Branches in the Axils of Cotyledons. (A) Branches were developed in exb1-d but not in wild-type plants. (B) and (C)
Supplementary Figure 1. jmj30-2 and jmj32-1 produce null mutants. (a) Schematic drawing of JMJ30 and JMJ32 genome structure showing regions amplified
 Supplementary Figure 1. jmj30-2 and jmj32-1 produce null mutants. (a) Schematic drawing of JMJ30 and JMJ32 genome structure showing regions amplified by primers used for mrna expression analysis. Gray
Supplementary Figure 1. jmj30-2 and jmj32-1 produce null mutants. (a) Schematic drawing of JMJ30 and JMJ32 genome structure showing regions amplified by primers used for mrna expression analysis. Gray
Quantitative and non-quantitative RT-PCR. cdna was generated from 500ng RNA (iscript;
 Supplemental Methods Quantitative and non-quantitative RT-PCR. cdna was generated from 500ng RNA (iscript; Bio-Rad, Hercules, CA, USA) and standard RT-PCR experiments were carried out using the 2X GoTaq
Supplemental Methods Quantitative and non-quantitative RT-PCR. cdna was generated from 500ng RNA (iscript; Bio-Rad, Hercules, CA, USA) and standard RT-PCR experiments were carried out using the 2X GoTaq
Supplemental Data. Farmer et al. (2010) Plant Cell /tpc
 Supplemental Figure 1. Amino acid sequence comparison of RAD23 proteins. Identical and similar residues are shown in the black and gray boxes, respectively. Dots denote gaps. The sequence of plant Ub is
Supplemental Figure 1. Amino acid sequence comparison of RAD23 proteins. Identical and similar residues are shown in the black and gray boxes, respectively. Dots denote gaps. The sequence of plant Ub is
Supplemental Data. Polycomb Silencing of KNOX Genes Confines. Shoot Stem Cell Niches in Arabidopsis Current Biology, Volume 18
 Supplemental Data Polycomb Silencing of KNOX Genes Confines Shoot Stem Cell Niches in Arabidopsis Lin Xu and Wen-Hui Shen - 1 - Figure S1. Phylogram of RING1, BMI1 and RAD18 homologues in several organisms
Supplemental Data Polycomb Silencing of KNOX Genes Confines Shoot Stem Cell Niches in Arabidopsis Lin Xu and Wen-Hui Shen - 1 - Figure S1. Phylogram of RING1, BMI1 and RAD18 homologues in several organisms
Supplemental Data. Na Xu et al. (2016). Plant Cell /tpc
 Supplemental Figure 1. The weak fluorescence phenotype is not caused by the mutation in At3g60240. (A) A mutation mapped to the gene At3g60240. Map-based cloning strategy was used to map the mutated site
Supplemental Figure 1. The weak fluorescence phenotype is not caused by the mutation in At3g60240. (A) A mutation mapped to the gene At3g60240. Map-based cloning strategy was used to map the mutated site
supplementary information
 DOI: 1.138/ncb1839 a b Control 1 2 3 Control 1 2 3 Fbw7 Smad3 1 2 3 4 1 2 3 4 c d IGF-1 IGF-1Rβ IGF-1Rβ-P Control / 1 2 3 4 Real-time RT-PCR Relative quantity (IGF-1/ mrna) 2 1 IGF-1 1 2 3 4 Control /
DOI: 1.138/ncb1839 a b Control 1 2 3 Control 1 2 3 Fbw7 Smad3 1 2 3 4 1 2 3 4 c d IGF-1 IGF-1Rβ IGF-1Rβ-P Control / 1 2 3 4 Real-time RT-PCR Relative quantity (IGF-1/ mrna) 2 1 IGF-1 1 2 3 4 Control /
Supplemental Data. Cui et al. (2012). Plant Cell /tpc a b c d. Stem UBC32 ACTIN
 A Root Stem Leaf Flower Silique Senescence leaf B a b c d UBC32 ACTIN C * Supplemental Figure 1. Expression Pattern and Protein Sequence of UBC32 Homologues in Yeast, Human, and Arabidopsis. (A) Expression
A Root Stem Leaf Flower Silique Senescence leaf B a b c d UBC32 ACTIN C * Supplemental Figure 1. Expression Pattern and Protein Sequence of UBC32 Homologues in Yeast, Human, and Arabidopsis. (A) Expression
Use of the APETALA1 promoter to assay the in vivo function of chimeric MADS box genes
 Sex Plant Reprod (1999) 12:14 26 Springer-Verlag 1999 ORIGINAL PAPER selor&:beth Allyn Krizek José Luis Riechmann Elliot M. Meyerowitz Use of the APETALA1 promoter to assay the in vivo function of chimeric
Sex Plant Reprod (1999) 12:14 26 Springer-Verlag 1999 ORIGINAL PAPER selor&:beth Allyn Krizek José Luis Riechmann Elliot M. Meyerowitz Use of the APETALA1 promoter to assay the in vivo function of chimeric
Jung-Nam Cho, Jee-Youn Ryu, Young-Min Jeong, Jihye Park, Ji-Joon Song, Richard M. Amasino, Bosl Noh, and Yoo-Sun Noh
 Developmental Cell, Volume 22 Supplemental Information Control of Seed Germination by Light-Induced Histone Arginine Demethylation Activity Jung-Nam Cho, Jee-Youn Ryu, Young-Min Jeong, Jihye Park, Ji-Joon
Developmental Cell, Volume 22 Supplemental Information Control of Seed Germination by Light-Induced Histone Arginine Demethylation Activity Jung-Nam Cho, Jee-Youn Ryu, Young-Min Jeong, Jihye Park, Ji-Joon
Nature Genetics: doi: /ng.3556 INTEGRATED SUPPLEMENTARY FIGURE TEMPLATE. Supplementary Figure 1
 INTEGRATED SUPPLEMENTARY FIGURE TEMPLATE Supplementary Figure 1 REF6 expression in transgenic lines. (a,b) Expression of REF6 in REF6-HA ref6 and REF6ΔZnF-HA ref6 plants detected by RT qpcr (a) and immunoblot
INTEGRATED SUPPLEMENTARY FIGURE TEMPLATE Supplementary Figure 1 REF6 expression in transgenic lines. (a,b) Expression of REF6 in REF6-HA ref6 and REF6ΔZnF-HA ref6 plants detected by RT qpcr (a) and immunoblot
Using mutants to clone genes
 Using mutants to clone genes Objectives: 1. What is positional cloning? 2. What is insertional tagging? 3. How can one confirm that the gene cloned is the same one that is mutated to give the phenotype
Using mutants to clone genes Objectives: 1. What is positional cloning? 2. What is insertional tagging? 3. How can one confirm that the gene cloned is the same one that is mutated to give the phenotype
Experimental Tools and Resources Available in Arabidopsis. Manish Raizada, University of Guelph, Canada
 Experimental Tools and Resources Available in Arabidopsis Manish Raizada, University of Guelph, Canada Community website: The Arabidopsis Information Resource (TAIR) at http://www.arabidopsis.org Can order
Experimental Tools and Resources Available in Arabidopsis Manish Raizada, University of Guelph, Canada Community website: The Arabidopsis Information Resource (TAIR) at http://www.arabidopsis.org Can order
Supplementary information
 Supplementary information Supplementary figures Figure S1 Level of mycdet1 protein in DET1 OE-1, OE-2 and OE-3 transgenic lines. Total protein extract from wild type Col0, det1-1 mutant and DET1 OE lines
Supplementary information Supplementary figures Figure S1 Level of mycdet1 protein in DET1 OE-1, OE-2 and OE-3 transgenic lines. Total protein extract from wild type Col0, det1-1 mutant and DET1 OE lines
Determination of Floral Organ Identity by Arabidopsis
 Molecular Biology of the Cell Vol. 8, 1243-1259, July 1997 Determination of Floral Organ Identity by Arabidopsis MADS Domain Homeotic Proteins API, AP3, PI, and AG Is Independent of Their DNA-binding Specificity
Molecular Biology of the Cell Vol. 8, 1243-1259, July 1997 Determination of Floral Organ Identity by Arabidopsis MADS Domain Homeotic Proteins API, AP3, PI, and AG Is Independent of Their DNA-binding Specificity
Liu, Yang (2012) The characterization of a novel abscission-related gene in Arabidopsis thaliana. PhD thesis, University of Nottingham.
 Liu, Yang (2012) The characterization of a novel abscission-related gene in Arabidopsis thaliana. PhD thesis, University of Nottingham. Access from the University of Nottingham repository: http://eprints.nottingham.ac.uk/12529/4/thesis_part_3_final.pdf
Liu, Yang (2012) The characterization of a novel abscission-related gene in Arabidopsis thaliana. PhD thesis, University of Nottingham. Access from the University of Nottingham repository: http://eprints.nottingham.ac.uk/12529/4/thesis_part_3_final.pdf
Supplemental Figure 1 HDA18 has an HDAC domain and therefore has concentration dependent and TSA inhibited histone deacetylase activity.
 Supplemental Figure 1 HDA18 has an HDAC domain and therefore has concentration dependent and TSA inhibited histone deacetylase activity. (A) Amino acid alignment of HDA5, HDA15 and HDA18. The blue line
Supplemental Figure 1 HDA18 has an HDAC domain and therefore has concentration dependent and TSA inhibited histone deacetylase activity. (A) Amino acid alignment of HDA5, HDA15 and HDA18. The blue line
SUPPLEMENTAL MATERIALS. Chromatin immunoprecipitation assays. Single-cell suspensions of pituitary
 SUPPLEMENTAL MATERIALS METHODS Chromatin immunoprecipitation assays. Single-cell suspensions of pituitary and liver cells were prepared from the indicated transgenic mice, as described (Ho et al, 2002).
SUPPLEMENTAL MATERIALS METHODS Chromatin immunoprecipitation assays. Single-cell suspensions of pituitary and liver cells were prepared from the indicated transgenic mice, as described (Ho et al, 2002).
SUPPLEMENTARY INFORMATION
 Ca 2+ /calmodulin Regulates Salicylic Acid-mediated Plant Immunity Liqun Du, Gul S. Ali, Kayla A. Simons, Jingguo Hou, Tianbao Yang, A.S.N. Reddy and B. W. Poovaiah * *To whom correspondence should be
Ca 2+ /calmodulin Regulates Salicylic Acid-mediated Plant Immunity Liqun Du, Gul S. Ali, Kayla A. Simons, Jingguo Hou, Tianbao Yang, A.S.N. Reddy and B. W. Poovaiah * *To whom correspondence should be
Supplementary Fig 1. The responses of ERF109 to different hormones and stresses. (a to k) The induced expression of ERF109 in 7-day-old Arabidopsis
 Supplementary Fig 1. The responses of ERF109 to different hormones and stresses. (a to k) The induced expression of ERF109 in 7-day-old Arabidopsis seedlings expressing ERF109pro-GUS. The GUS staining
Supplementary Fig 1. The responses of ERF109 to different hormones and stresses. (a to k) The induced expression of ERF109 in 7-day-old Arabidopsis seedlings expressing ERF109pro-GUS. The GUS staining
Supplementary Information. The flowering gene SINGLE FLOWER TRUSS drives heterosis for yield in tomato
 Supplementary Information The flowering gene SINGLE FLOWER TRUSS drives heterosis for yield in tomato Uri Krieger 1, Zachary B. Lippman 2 *, and Dani Zamir 1 * 1. The Hebrew University of Jerusalem Faculty
Supplementary Information The flowering gene SINGLE FLOWER TRUSS drives heterosis for yield in tomato Uri Krieger 1, Zachary B. Lippman 2 *, and Dani Zamir 1 * 1. The Hebrew University of Jerusalem Faculty
Supplemental Information. Autoregulatory Feedback Controls. Sequential Action of cis-regulatory Modules. at the brinker Locus
 Developmental Cell, Volume 26 Supplemental Information Autoregulatory Feedback Controls Sequential Action of cis-regulatory Modules at the brinker Locus Leslie Dunipace, Abbie Saunders, Hilary L. Ashe,
Developmental Cell, Volume 26 Supplemental Information Autoregulatory Feedback Controls Sequential Action of cis-regulatory Modules at the brinker Locus Leslie Dunipace, Abbie Saunders, Hilary L. Ashe,
ChIP DNA Clean & Concentrator Catalog Nos. D5201 & D5205
 Page 0 INSTRUCTION MANUAL ChIP DNA Clean & Concentrator Catalog Nos. D5201 & D5205 Highlights Quick (2 minute) recovery of ultra-pure DNA from chromatin immunoprecipitation (ChIP) assays, cell lysates,
Page 0 INSTRUCTION MANUAL ChIP DNA Clean & Concentrator Catalog Nos. D5201 & D5205 Highlights Quick (2 minute) recovery of ultra-pure DNA from chromatin immunoprecipitation (ChIP) assays, cell lysates,
Supplemental Figure 1. VLN5 retains conserved residues at both type 1 and type 2 Ca 2+ -binding
 Supplemental Figure 1. VLN5 retains conserved residues at both type 1 and type 2 Ca 2+ -binding sites in the G1 domain. Multiple sequence alignment was performed with DNAMAN6.0.40. Secondary structural
Supplemental Figure 1. VLN5 retains conserved residues at both type 1 and type 2 Ca 2+ -binding sites in the G1 domain. Multiple sequence alignment was performed with DNAMAN6.0.40. Secondary structural
Enhancers mutations that make the original mutant phenotype more extreme. Suppressors mutations that make the original mutant phenotype less extreme
 Interactomics and Proteomics 1. Interactomics The field of interactomics is concerned with interactions between genes or proteins. They can be genetic interactions, in which two genes are involved in the
Interactomics and Proteomics 1. Interactomics The field of interactomics is concerned with interactions between genes or proteins. They can be genetic interactions, in which two genes are involved in the
sides of the aleurone (Al) but it is excluded from the basal endosperm transfer layer
 Supplemental Data. Gómez et al. (2009). The maize transcription factor MRP-1 (Myb-Related-Protein-1) is a key regulator of the differentiation of transfer cells. Supplemental Figure 1. Expression analyses
Supplemental Data. Gómez et al. (2009). The maize transcription factor MRP-1 (Myb-Related-Protein-1) is a key regulator of the differentiation of transfer cells. Supplemental Figure 1. Expression analyses
Different Upstream Activators Stimulate Transcription Through Distinct Coactivator Complexes
 Supplemental Material for Different Upstream Activators Stimulate Transcription Through Distinct Coactivator Complexes Dong-ki Lee, Soyoun Kim, and John T. Lis Due to be published as a Research Communication
Supplemental Material for Different Upstream Activators Stimulate Transcription Through Distinct Coactivator Complexes Dong-ki Lee, Soyoun Kim, and John T. Lis Due to be published as a Research Communication
RNA was isolated using NucleoSpin RNA II (Macherey-Nagel, Bethlehem, PA) according to the
 Supplementary Methods RT-PCR and real-time PCR analysis RNA was isolated using NucleoSpin RNA II (Macherey-Nagel, Bethlehem, PA) according to the manufacturer s protocol and quantified by measuring the
Supplementary Methods RT-PCR and real-time PCR analysis RNA was isolated using NucleoSpin RNA II (Macherey-Nagel, Bethlehem, PA) according to the manufacturer s protocol and quantified by measuring the
SUPPLEMENTARY INFORMATION
 Materials and Methods Transgenic Plant Materials and DNA Constructs. VEX1::H2B-GFP, ACA3::H2B-GFP, KRP6::H2B-GFP, KRP6::mock21ts-GFP, KRP6::TE21ts- GFP and KRP6::miR161ts-GFP constructs were generated
Materials and Methods Transgenic Plant Materials and DNA Constructs. VEX1::H2B-GFP, ACA3::H2B-GFP, KRP6::H2B-GFP, KRP6::mock21ts-GFP, KRP6::TE21ts- GFP and KRP6::miR161ts-GFP constructs were generated
Supplementary Information. c d e
 Supplementary Information a b c d e f Supplementary Figure 1. atabcg30, atabcg31, and atabcg40 mutant seeds germinate faster than the wild type on ½ MS medium supplemented with ABA (a and d-f) Germination
Supplementary Information a b c d e f Supplementary Figure 1. atabcg30, atabcg31, and atabcg40 mutant seeds germinate faster than the wild type on ½ MS medium supplemented with ABA (a and d-f) Germination
Supplemental Data. Zhou et al. (2016). Plant Cell /tpc
 Supplemental Figure 1. Confirmation of mutant mapping results. (A) Complementation assay with stably transformed genomic fragments (ComN-N) (2 kb upstream of TSS and 1.5 kb downstream of TES) and CaMV
Supplemental Figure 1. Confirmation of mutant mapping results. (A) Complementation assay with stably transformed genomic fragments (ComN-N) (2 kb upstream of TSS and 1.5 kb downstream of TES) and CaMV
Supplementary Methods
 Supplementary Methods Reverse transcribed Quantitative PCR. Total RNA was isolated from bone marrow derived macrophages using RNeasy Mini Kit (Qiagen), DNase-treated (Promega RQ1), and reverse transcribed
Supplementary Methods Reverse transcribed Quantitative PCR. Total RNA was isolated from bone marrow derived macrophages using RNeasy Mini Kit (Qiagen), DNase-treated (Promega RQ1), and reverse transcribed
HiChIP Protocol Mumbach et al. (CHANG), p. 1 Chang Lab, Stanford University. HiChIP Protocol
 HiChIP Protocol Mumbach et al. (CHANG), p. 1 HiChIP Protocol Citation: Mumbach et. al., HiChIP: Efficient and sensitive analysis of protein-directed genome architecture. Nature Methods (2016). Cell Crosslinking
HiChIP Protocol Mumbach et al. (CHANG), p. 1 HiChIP Protocol Citation: Mumbach et. al., HiChIP: Efficient and sensitive analysis of protein-directed genome architecture. Nature Methods (2016). Cell Crosslinking
Intracellular receptors specify complex patterns of gene expression that are cell and gene
 SUPPLEMENTAL RESULTS AND DISCUSSION Some HPr-1AR ARE-containing Genes Are Unresponsive to Androgen Intracellular receptors specify complex patterns of gene expression that are cell and gene specific. For
SUPPLEMENTAL RESULTS AND DISCUSSION Some HPr-1AR ARE-containing Genes Are Unresponsive to Androgen Intracellular receptors specify complex patterns of gene expression that are cell and gene specific. For
Discrete spatial and temporal cis-acting elements regulate transcription of the Arabidopsis floral homeotic gene APETALA3
 Development 125, 1711-1721 (1998) Printed in Great Britain The Company of Biologists Limited 1998 DEV0158 1711 Discrete spatial and temporal cis-acting elements regulate transcription of the Arabidopsis
Development 125, 1711-1721 (1998) Printed in Great Britain The Company of Biologists Limited 1998 DEV0158 1711 Discrete spatial and temporal cis-acting elements regulate transcription of the Arabidopsis
SUPPLEMENTARY INFORMATION
 AS-NMD modulates FLM-dependent thermosensory flowering response in Arabidopsis NATURE PLANTS www.nature.com/natureplants 1 Supplementary Figure 1. Genomic sequence of FLM along with the splice sites. Sequencing
AS-NMD modulates FLM-dependent thermosensory flowering response in Arabidopsis NATURE PLANTS www.nature.com/natureplants 1 Supplementary Figure 1. Genomic sequence of FLM along with the splice sites. Sequencing
EZ-Zyme Chromatin Prep Kit
 Instruction Manual for EZ-Zyme Chromatin Prep Kit Catalog # 17 375 Sufficient reagents for 22 enzymatic chromatin preparations per kit. Contents Page I. INTRODUCTION 2 II. EZ-Zyme KIT COMPONENTS 3 A. Provided
Instruction Manual for EZ-Zyme Chromatin Prep Kit Catalog # 17 375 Sufficient reagents for 22 enzymatic chromatin preparations per kit. Contents Page I. INTRODUCTION 2 II. EZ-Zyme KIT COMPONENTS 3 A. Provided
To investigate the heredity of the WFP gene, we selected plants that were homozygous
 Supplementary information Supplementary Note ST-12 WFP allele is semi-dominant To investigate the heredity of the WFP gene, we selected plants that were homozygous for chromosome 1 of Nipponbare and heterozygous
Supplementary information Supplementary Note ST-12 WFP allele is semi-dominant To investigate the heredity of the WFP gene, we selected plants that were homozygous for chromosome 1 of Nipponbare and heterozygous
(phosphatase tensin) domain is shown in dark gray, the FH1 domain in black, and the
 Supplemental Figure 1. Predicted Domain Organization of the AFH14 Protein. (A) Schematic representation of the predicted domain organization of AFH14. The PTEN (phosphatase tensin) domain is shown in dark
Supplemental Figure 1. Predicted Domain Organization of the AFH14 Protein. (A) Schematic representation of the predicted domain organization of AFH14. The PTEN (phosphatase tensin) domain is shown in dark
Pattern Formation via Small RNA Mobility SUPPLEMENTAL FIGURE 1 SUPPLEMENTAL INFORMATION. Daniel H. Chitwood et al.
 SUPPLEMENTAL INFORMATION Pattern Formation via Small RNA Mobility Daniel H. Chitwood et al. SUPPLEMENTAL FIGURE 1 Supplemental Figure 1. The ARF3 promoter drives expression throughout leaves. (A, B) Expression
SUPPLEMENTAL INFORMATION Pattern Formation via Small RNA Mobility Daniel H. Chitwood et al. SUPPLEMENTAL FIGURE 1 Supplemental Figure 1. The ARF3 promoter drives expression throughout leaves. (A, B) Expression
SUPPLEMENTARY INFORMATION. LIN-28 co-transcriptionally binds primary let-7 to regulate mirna maturation in C. elegans
 SUPPLEMENTARY INFORMATION LIN-28 co-transcriptionally binds primary let-7 to regulate mirna maturation in C. elegans Priscilla M. Van Wynsberghe 1, Zoya S. Kai 1, Katlin B. Massirer 2-4, Victoria H. Burton
SUPPLEMENTARY INFORMATION LIN-28 co-transcriptionally binds primary let-7 to regulate mirna maturation in C. elegans Priscilla M. Van Wynsberghe 1, Zoya S. Kai 1, Katlin B. Massirer 2-4, Victoria H. Burton
Solid Phase cdna Synthesis Kit
 #6123 v.02.09 Table of Contents I. Description... 2 II. Kit components... 2 III. Storage... 2 IV. Principle... 3 V. Protocol V-1. Preparation of immobilized mrna... 4 Protocol A: Starting from Tissue or
#6123 v.02.09 Table of Contents I. Description... 2 II. Kit components... 2 III. Storage... 2 IV. Principle... 3 V. Protocol V-1. Preparation of immobilized mrna... 4 Protocol A: Starting from Tissue or
Supplementary Table 1. The Q-PCR primer sequence is summarized in the following table.
 Supplementary Table 1. The Q-PCR primer sequence is summarized in the following table. Name Sequence (5-3 ) Application Flag-u ggactacaaggacgacgatgac Shared upstream primer for all the amplifications of
Supplementary Table 1. The Q-PCR primer sequence is summarized in the following table. Name Sequence (5-3 ) Application Flag-u ggactacaaggacgacgatgac Shared upstream primer for all the amplifications of
Supplemental Data. Lin et al. Plant Cell. (2014) /tpc
 Supplemental Figure 1. Scheme for extraction of polysome-mrna from in vitro germinated pollen (in vitro), pollinated (in vivo stage, ST14 and ST15a) and non-pollinated (Bud stage, ST10-12) floral buds.
Supplemental Figure 1. Scheme for extraction of polysome-mrna from in vitro germinated pollen (in vitro), pollinated (in vivo stage, ST14 and ST15a) and non-pollinated (Bud stage, ST10-12) floral buds.
Supplemental Data. LMO4 Controls the Balance between Excitatory. and Inhibitory Spinal V2 Interneurons
 Neuron, Volume 61 Supplemental Data LMO4 Controls the Balance between Excitatory and Inhibitory Spinal V2 Interneurons Kaumudi Joshi, Seunghee Lee, Bora Lee, Jae W. Lee, and Soo-Kyung Lee Supplemental
Neuron, Volume 61 Supplemental Data LMO4 Controls the Balance between Excitatory and Inhibitory Spinal V2 Interneurons Kaumudi Joshi, Seunghee Lee, Bora Lee, Jae W. Lee, and Soo-Kyung Lee Supplemental
Supplemental Data. mir156-regulated SPL Transcription. Factors Define an Endogenous Flowering. Pathway in Arabidopsis thaliana
 Cell, Volume 138 Supplemental Data mir156-regulated SPL Transcription Factors Define an Endogenous Flowering Pathway in Arabidopsis thaliana Jia-Wei Wang, Benjamin Czech, and Detlef Weigel Table S1. Interaction
Cell, Volume 138 Supplemental Data mir156-regulated SPL Transcription Factors Define an Endogenous Flowering Pathway in Arabidopsis thaliana Jia-Wei Wang, Benjamin Czech, and Detlef Weigel Table S1. Interaction
Genomic DNA was extracted from 3 to 5 ml of blood collected in EDTA blood collection tubes
 Supplementary information Methods DNA and RNA extraction Genomic DNA was extracted from to ml of blood collected in EDTA blood collection tubes using the Gentra Puregene Blood kit (Qiagen, California,
Supplementary information Methods DNA and RNA extraction Genomic DNA was extracted from to ml of blood collected in EDTA blood collection tubes using the Gentra Puregene Blood kit (Qiagen, California,
rev. 05/31/12 Box 2 contains the following buffers/reagents:
 Store at RT, 4 C, and 20 C #9002 SimpleChIP Enzymatic Chromatin IP Kit (Agarose Beads) 3 n 1 Kit (30 Immunoprecipitations) For Research Use Only. Not For Use In Diagnostic Procedures. rev. 05/31/12 Orders
Store at RT, 4 C, and 20 C #9002 SimpleChIP Enzymatic Chromatin IP Kit (Agarose Beads) 3 n 1 Kit (30 Immunoprecipitations) For Research Use Only. Not For Use In Diagnostic Procedures. rev. 05/31/12 Orders
Supplemental Figure legends Figure S1. (A) (B) (C) (D) Figure S2. Figure S3. (A-E) Figure S4. Figure S5. (A, C, E, G, I) (B, D, F, H, Figure S6.
 Supplemental Figure legends Figure S1. Map-based cloning and complementation testing for ZOP1. (A) ZOP1 was mapped to a ~273-kb interval on Chromosome 1. In the interval, a single-nucleotide G to A substitution
Supplemental Figure legends Figure S1. Map-based cloning and complementation testing for ZOP1. (A) ZOP1 was mapped to a ~273-kb interval on Chromosome 1. In the interval, a single-nucleotide G to A substitution
ChIP-Adembeads (cat #04242/04342)
 ChIP-Adem-Kit (cat # 04243/04343) ChIP-Adembeads (cat #04242/04342) Instruction manual for magnetic Chromatin Immunoprecipitation Ademtech SA Bioparc BioGalien 27, allée Charles Darwin 33600 PESSAC France
ChIP-Adem-Kit (cat # 04243/04343) ChIP-Adembeads (cat #04242/04342) Instruction manual for magnetic Chromatin Immunoprecipitation Ademtech SA Bioparc BioGalien 27, allée Charles Darwin 33600 PESSAC France
Enzyme that uses RNA as a template to synthesize a complementary DNA
 Biology 105: Introduction to Genetics PRACTICE FINAL EXAM 2006 Part I: Definitions Homology: Comparison of two or more protein or DNA sequence to ascertain similarities in sequences. If two genes have
Biology 105: Introduction to Genetics PRACTICE FINAL EXAM 2006 Part I: Definitions Homology: Comparison of two or more protein or DNA sequence to ascertain similarities in sequences. If two genes have
EPIGENTEK. EpiQuik Chromatin Immunoprecipitation Kit. Base Catalog # P-2002 PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE
 EpiQuik Chromatin Immunoprecipitation Kit Base Catalog # P-2002 PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE The EpiQuik Chromatin Immunoprecipitation Kit is suitable for combining the specificity of
EpiQuik Chromatin Immunoprecipitation Kit Base Catalog # P-2002 PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE The EpiQuik Chromatin Immunoprecipitation Kit is suitable for combining the specificity of
SUPPLEMENTARY INFORMATION
 Local auxin metabolism regulates environment-induced hypocotyl elongation Zuyu Zheng 1,2, Yongxia Guo 3, Ondřej Novák 4,5, William Chen 2, Karin Ljung 4, Joseph P. Noel 1,3, *, and Joanne Chory 1,2, *
Local auxin metabolism regulates environment-induced hypocotyl elongation Zuyu Zheng 1,2, Yongxia Guo 3, Ondřej Novák 4,5, William Chen 2, Karin Ljung 4, Joseph P. Noel 1,3, *, and Joanne Chory 1,2, *
Applicazioni biotecnologiche
 Applicazioni biotecnologiche Analisi forense Sintesi di proteine ricombinanti Restriction Fragment Length Polymorphism (RFLP) Polymorphism (more fully genetic polymorphism) refers to the simultaneous occurrence
Applicazioni biotecnologiche Analisi forense Sintesi di proteine ricombinanti Restriction Fragment Length Polymorphism (RFLP) Polymorphism (more fully genetic polymorphism) refers to the simultaneous occurrence
Supplementary Figure 1. BES1 specifically inhibits ABA responses in early seedling
 Supplementary Figure 1. BES1 specifically inhibits ABA responses in early seedling development. a. Exogenous BR application overcomes the hypersensitivity of bzr1-1d seedlings to ABA. Seed germination
Supplementary Figure 1. BES1 specifically inhibits ABA responses in early seedling development. a. Exogenous BR application overcomes the hypersensitivity of bzr1-1d seedlings to ABA. Seed germination
Site directed mutagenesis, Insertional and Deletion Mutagenesis. Mitesh Shrestha
 Site directed mutagenesis, Insertional and Deletion Mutagenesis Mitesh Shrestha Mutagenesis Mutagenesis (the creation or formation of a mutation) can be used as a powerful genetic tool. By inducing mutations
Site directed mutagenesis, Insertional and Deletion Mutagenesis Mitesh Shrestha Mutagenesis Mutagenesis (the creation or formation of a mutation) can be used as a powerful genetic tool. By inducing mutations
PIE1 ARP6 SWC6 KU70 ARP6 PIE1. HSA SNF2_N HELICc SANT. pie1-3 A1,A2 K1,K2 K1,K3 K3,LB2 A3, A4 A3,LB1 A1,A2 K1,K2 K1,K3. swc6-1 A3,A4.
 A B N-terminal SWC2 H2A.Z SWC6 ARP6 PIE1 HSA SNF2_N HELICc SANT C pie1-3 D PIE1 ARP6 5 Kb A1 200 bp A3 A2 LB1 arp6-3 A4 E A1,A2 A3, A4 A3,LB1 K1,K2 K1,K3 K3,LB2 SWC6 swc6-1 A1,A2 A3,A4 K1,K2 K1,K3 100
A B N-terminal SWC2 H2A.Z SWC6 ARP6 PIE1 HSA SNF2_N HELICc SANT C pie1-3 D PIE1 ARP6 5 Kb A1 200 bp A3 A2 LB1 arp6-3 A4 E A1,A2 A3, A4 A3,LB1 K1,K2 K1,K3 K3,LB2 SWC6 swc6-1 A1,A2 A3,A4 K1,K2 K1,K3 100
Ren Lab ENCODE in situ HiC Protocol for Tissue
 Ren Lab ENCODE in situ HiC Protocol for Tissue Pulverization, Crosslinking of Tissue Note: Ensure the samples are kept frozen on dry ice throughout pulverization. 1. Pour liquid nitrogen into a mortar
Ren Lab ENCODE in situ HiC Protocol for Tissue Pulverization, Crosslinking of Tissue Note: Ensure the samples are kept frozen on dry ice throughout pulverization. 1. Pour liquid nitrogen into a mortar
At E17.5, the embryos were rinsed in phosphate-buffered saline (PBS) and immersed in
 Supplementary Materials and Methods Barrier function assays At E17.5, the embryos were rinsed in phosphate-buffered saline (PBS) and immersed in acidic X-gal mix (100 mm phosphate buffer at ph4.3, 3 mm
Supplementary Materials and Methods Barrier function assays At E17.5, the embryos were rinsed in phosphate-buffered saline (PBS) and immersed in acidic X-gal mix (100 mm phosphate buffer at ph4.3, 3 mm
TRANSGENIC ANIMALS. -transient transfection of cells -stable transfection of cells. - Two methods to produce transgenic animals:
 TRANSGENIC ANIMALS -transient transfection of cells -stable transfection of cells - Two methods to produce transgenic animals: 1- DNA microinjection - random insertion 2- embryonic stem cell-mediated gene
TRANSGENIC ANIMALS -transient transfection of cells -stable transfection of cells - Two methods to produce transgenic animals: 1- DNA microinjection - random insertion 2- embryonic stem cell-mediated gene
S156AT168AY175A (AAA) were purified as GST-fusion proteins and incubated with GSTfused
 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 Supplemental Materials Supplemental Figure S1 (a) Phenotype of the wild type and grik1-2 grik2-1 plants after 8 days in darkness.
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 Supplemental Materials Supplemental Figure S1 (a) Phenotype of the wild type and grik1-2 grik2-1 plants after 8 days in darkness.
Supplemental Data. Lee et al. Plant Cell. (2010) /tpc Supplemental Figure 1. Protein and Gene Structures of DWA1 and DWA2.
 Supplemental Figure 1. Protein and Gene Structures of DWA1 and DWA2. (A) Protein structures of DWA1 and DWA2. WD40 region was determined based on the NCBI conserved domain databases (B, C) Schematic representation
Supplemental Figure 1. Protein and Gene Structures of DWA1 and DWA2. (A) Protein structures of DWA1 and DWA2. WD40 region was determined based on the NCBI conserved domain databases (B, C) Schematic representation
Strep-Spin Protein Miniprep Kit Catalog No. P2004, P2005
 INSTRUCTION MANUAL Strep-Spin Protein Miniprep Kit Catalog No. P2004, P2005 Highlights Fast protocol to purify Strep-tagged proteins from cell-free extracts Screen your recombinant colonies directly for
INSTRUCTION MANUAL Strep-Spin Protein Miniprep Kit Catalog No. P2004, P2005 Highlights Fast protocol to purify Strep-tagged proteins from cell-free extracts Screen your recombinant colonies directly for
EPIGENTEK. EpiQuik Tissue Chromatin Immunoprecipitation Kit. Base Catalog # P-2003 PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE
 EpiQuik Tissue Chromatin Immunoprecipitation Kit Base Catalog # P-2003 PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE The EpiQuik Tissue Chromatin Immunoprecipitation Kit is suitable for combining the specificity
EpiQuik Tissue Chromatin Immunoprecipitation Kit Base Catalog # P-2003 PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE The EpiQuik Tissue Chromatin Immunoprecipitation Kit is suitable for combining the specificity
Transcriptional regulation of BRCA1 expression by a metabolic switch: Di, Fernandez, De Siervi, Longo, and Gardner. H3K4Me3
 ChIP H3K4Me3 enrichment.25.2.15.1.5 H3K4Me3 H3K4Me3 ctrl H3K4Me3 + E2 NS + E2 1. kb kb +82 kb Figure S1. Estrogen promotes entry of MCF-7 into the cell cycle but does not significantly change activation-associated
ChIP H3K4Me3 enrichment.25.2.15.1.5 H3K4Me3 H3K4Me3 ctrl H3K4Me3 + E2 NS + E2 1. kb kb +82 kb Figure S1. Estrogen promotes entry of MCF-7 into the cell cycle but does not significantly change activation-associated
Fig. S1. CrgA intracellular levels in M. tuberculosis. Ten and twenty micrograms of
 Supplementary data Fig. S1. CrgA intracellular levels in M. tuberculosis. Ten and twenty micrograms of cell free protein lysates from WT M. tuberculosis (Rv) together with various known concentrations
Supplementary data Fig. S1. CrgA intracellular levels in M. tuberculosis. Ten and twenty micrograms of cell free protein lysates from WT M. tuberculosis (Rv) together with various known concentrations
Construction of plant complementation vector and generation of transgenic plants
 MATERIAL S AND METHODS Plant materials and growth conditions Arabidopsis ecotype Columbia (Col0) was used for this study. SALK_072009, SALK_076309, and SALK_027645 were obtained from the Arabidopsis Biological
MATERIAL S AND METHODS Plant materials and growth conditions Arabidopsis ecotype Columbia (Col0) was used for this study. SALK_072009, SALK_076309, and SALK_027645 were obtained from the Arabidopsis Biological
Supplemental Data. Jing et al. (2013). Plant Cell /tpc
 Supplemental Figure 1. Characterization of epp1 Mutants. (A) Cotyledon angles of 5-d-old Col wild-type (gray bars) and epp1-1 (black bars) seedlings under red (R), far-red (FR) and blue (BL) light conditions,
Supplemental Figure 1. Characterization of epp1 Mutants. (A) Cotyledon angles of 5-d-old Col wild-type (gray bars) and epp1-1 (black bars) seedlings under red (R), far-red (FR) and blue (BL) light conditions,
Intron distance to transcription start (nt)*
 Schwab et al. SOM: Enhanced microrna accumulation through stemloop-adjacent introns 1 Supplemental Table 1. Characteristics of introns used in this study Intron name Length intron (nt) Length cloned fragment
Schwab et al. SOM: Enhanced microrna accumulation through stemloop-adjacent introns 1 Supplemental Table 1. Characteristics of introns used in this study Intron name Length intron (nt) Length cloned fragment
CHAPTER 4 Cloning, expression, purification and preparation of site-directed mutants of NDUFS3 and NDUFS7
 CHAPTER 4 Cloning, expression, purification and preparation of site-directed mutants of NDUFS3 and NDUFS7 subunits of human mitochondrial Complex-I Q module N DUFS2, 3, 7 and 8 form the core subunits of
CHAPTER 4 Cloning, expression, purification and preparation of site-directed mutants of NDUFS3 and NDUFS7 subunits of human mitochondrial Complex-I Q module N DUFS2, 3, 7 and 8 form the core subunits of
Nature Genetics: doi: /ng Supplementary Figure 1. ChIP-seq genome browser views of BRM occupancy at previously identified BRM targets.
 Supplementary Figure 1 ChIP-seq genome browser views of BRM occupancy at previously identified BRM targets. Gene structures are shown underneath each panel. Supplementary Figure 2 pref6::ref6-gfp complements
Supplementary Figure 1 ChIP-seq genome browser views of BRM occupancy at previously identified BRM targets. Gene structures are shown underneath each panel. Supplementary Figure 2 pref6::ref6-gfp complements
1. Introduction Drought stress and climate change Three strategies of plants in response to water stress 3
 Contents 1. Introduction 1 1.1 Drought stress and climate change 3 1.2 Three strategies of plants in response to water stress 3 1.3 Three closely related species of Linderniaceae family are experimental
Contents 1. Introduction 1 1.1 Drought stress and climate change 3 1.2 Three strategies of plants in response to water stress 3 1.3 Three closely related species of Linderniaceae family are experimental
The preparation of native chromatin from cultured human cells.
 Native chromatin immunoprecipitation protocol The preparation of native chromatin from cultured human cells. All solutions need to be ice cold. Sucrose containing solutions must be made up fresh on the
Native chromatin immunoprecipitation protocol The preparation of native chromatin from cultured human cells. All solutions need to be ice cold. Sucrose containing solutions must be made up fresh on the
WiscDsLox485 ATG < > //----- E1 E2 E3 E4 E bp. Col-0 arr7 ARR7 ACTIN7. s of mrna/ng total RNA (x10 3 ) ARR7.
 A WiscDsLox8 ATG < > -9 +0 > ---------//----- E E E E E UTR +9 UTR +0 > 00bp B S D ol-0 arr7 ol-0 arr7 ARR7 ATIN7 D s of mrna/ng total RNA opie 0. ol-0 (x0 ) arr7 ARR7 Supplemental Fig.. Genotyping and
A WiscDsLox8 ATG < > -9 +0 > ---------//----- E E E E E UTR +9 UTR +0 > 00bp B S D ol-0 arr7 ol-0 arr7 ARR7 ATIN7 D s of mrna/ng total RNA opie 0. ol-0 (x0 ) arr7 ARR7 Supplemental Fig.. Genotyping and
Fig. S1. Effect of p120-catenin overexpression on the interaction of SCUBE2 with E-cadherin. The expression plasmid encoding FLAG.
 Fig. S1. Effect of p120-catenin overexpression on the interaction of SCUBE2 with E-cadherin. The expression plasmid encoding FLAG.SCUBE2, E-cadherin.Myc, or HA.p120-catenin was transfected in a combination
Fig. S1. Effect of p120-catenin overexpression on the interaction of SCUBE2 with E-cadherin. The expression plasmid encoding FLAG.SCUBE2, E-cadherin.Myc, or HA.p120-catenin was transfected in a combination
Supporting Online Material for
 www.sciencemag.org/cgi/content/full/318/5851/794/dc1 Supporting Online Material for A Gene Regulatory Network Subcircuit Drives a Dynamic Pattern of Gene Expression Joel Smith, Christina Theodoris, Eric
www.sciencemag.org/cgi/content/full/318/5851/794/dc1 Supporting Online Material for A Gene Regulatory Network Subcircuit Drives a Dynamic Pattern of Gene Expression Joel Smith, Christina Theodoris, Eric
SUPPLEMENTARY INFORMATION
 Gene replacements and insertions in rice by intron targeting using CRISPR Cas9 Table of Contents Supplementary Figure 1. sgrna-induced targeted mutations in the OsEPSPS gene in rice protoplasts. Supplementary
Gene replacements and insertions in rice by intron targeting using CRISPR Cas9 Table of Contents Supplementary Figure 1. sgrna-induced targeted mutations in the OsEPSPS gene in rice protoplasts. Supplementary
scgem Workflow Experimental Design Single cell DNA methylation primer design
 scgem Workflow Experimental Design Single cell DNA methylation primer design The scgem DNA methylation assay uses qpcr to measure digestion of target loci by the methylation sensitive restriction endonuclease
scgem Workflow Experimental Design Single cell DNA methylation primer design The scgem DNA methylation assay uses qpcr to measure digestion of target loci by the methylation sensitive restriction endonuclease
Roche Molecular Biochemicals Technical Note No. LC 12/2000
 Roche Molecular Biochemicals Technical Note No. LC 12/2000 LightCycler Absolute Quantification with External Standards and an Internal Control 1. General Introduction Purpose of this Note Overview of Method
Roche Molecular Biochemicals Technical Note No. LC 12/2000 LightCycler Absolute Quantification with External Standards and an Internal Control 1. General Introduction Purpose of this Note Overview of Method
A subclass of HSP70s regulate development and abiotic stress responses in Arabidopsis thaliana
 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 Journal of Plant Research A subclass of HSP70s regulate development and abiotic stress responses in Arabidopsis thaliana Linna Leng 1 Qianqian Liang
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 Journal of Plant Research A subclass of HSP70s regulate development and abiotic stress responses in Arabidopsis thaliana Linna Leng 1 Qianqian Liang
ZR-96 Genomic DNA Clean & Concentrator -5 Catalog Nos. D4066 & D4067
 INSTRUCTION MANUAL ZR-96 Genomic DNA Clean & Concentrator -5 Catalog Nos. D4066 & D4067 Highlights 96-well plate recovery of large-sized DNA (e.g., genomic, mitochondrial, plasmid (BAC/PAC), viral, phage,
INSTRUCTION MANUAL ZR-96 Genomic DNA Clean & Concentrator -5 Catalog Nos. D4066 & D4067 Highlights 96-well plate recovery of large-sized DNA (e.g., genomic, mitochondrial, plasmid (BAC/PAC), viral, phage,
Figure S1. Verification of ihog Mutation by Protein Immunoblotting Figure S2. Verification of ihog and boi
 Figure S1. Verification of ihog Mutation by Protein Immunoblotting Extracts from S2R+ cells, embryos, and adults were analyzed by immunoprecipitation and immunoblotting with anti-ihog antibody. The Ihog
Figure S1. Verification of ihog Mutation by Protein Immunoblotting Extracts from S2R+ cells, embryos, and adults were analyzed by immunoprecipitation and immunoblotting with anti-ihog antibody. The Ihog
Supplemental data. Zhao et al. (2009). The Wuschel-related homeobox gene WOX11 is required to activate shoot-borne crown root development in rice.
 Supplemental data. Zhao et al. (2009). The Wuschel-related homeobox gene WOX11 is required to activate shoot-borne crown root development in rice. A B Supplemental Figure 1. Expression of WOX11p-GUS WOX11-GFP
Supplemental data. Zhao et al. (2009). The Wuschel-related homeobox gene WOX11 is required to activate shoot-borne crown root development in rice. A B Supplemental Figure 1. Expression of WOX11p-GUS WOX11-GFP
Supplementary Information
 Supplementary Information Table of contents: Pages: Supplementary Methods 2-3 Supplementary Figure 1 4 Supplementary Figure 2 5 Supplementary Figure 3 6 Supplementary Figure 4 7 Supplementary Figure 5
Supplementary Information Table of contents: Pages: Supplementary Methods 2-3 Supplementary Figure 1 4 Supplementary Figure 2 5 Supplementary Figure 3 6 Supplementary Figure 4 7 Supplementary Figure 5
