Computational Biology I LSM5191
|
|
|
- Sydney Stone
- 6 years ago
- Views:
Transcription
1 Computational Biology I LSM5191 Aylwin Ng, D.Phil Lecture 4 Notes: Gene Regulation & Control
2 Do all cells in an individual have the same DNA content?
3 PART I: CONTROL OF GENE EXPRESSION
4 REGULATION OF GENE EXPRESSION Any of these stages could be used to regulate expression of specific genes in particular tissues. But in general, the primary control of gene expression is at the level of transcription. DNA Start Exon1 Intron Exon2 Termination *Transcription m 7 Gppp Addition of 5 cap Cleavage & addn of polya tail at 3 end m 7 Gppp A (A) 200 RNA splicing m 7 Gppp A (A) 200 Transport to cytoplasm Translation Protein
5 TRANSCRIPTIONAL CONTROL Regulatory Sequence Elements Short regulatory elements Enhancers or Enhancer Elements Locus control regions Transcriptional activators
6 Short regulatory elements Elements commonly found in many genes e.g. TATA, CCAAT and Sp1 boxes. Elements found only in specific genes, e.g. heat-shock element found only in genes whose transcription is increased in response to elevated temperature. Heat inducible Hsp 70 Non-heat inducible tk Heat-shock element Chimeric gene tk Pelham, 1982, Cell 30: Heat-inducible transcription
7 Sequences present in the upstream region of hsp70 gene also found in other genes Element Name Consensus sequence Other genes containing sequences TATA box TATA A/T A A/T Very many genes. CCAAT box TGTGGCTNNNAGC CAA α- and β-globin, albumin, HSV tk, cellular oncogenes: c-ras, c-myc, etc. Sp1 box GGGCGG Metallothionein IIA, type II procollagen, dihydrofolate reductase, etc. CRE T/G T/A CGTCA Somatostatin, fibronectin, α-gonadotrophin, c-fos, etc. AP2 box CCCCAGGC Collagenase, MHC class 1 antigen H-2K b, metallothionein IIA. Heat-shock consensus CTNGAATNTTCTAG A Heat-inducible genes hsp83, hsp27, etc. Adapted from Latchman, D., 1998, Gene regulation, Stanley Thornes Publ.
8 Enhancer or Enhancer Elements These elements can activate a promoter when placed: up to several kb from promoter, in either orientation relative to promoter, upstream or downstream of the transcribed region, or within introns. Genes exhibiting tissue-specific expression found to contain enhancers. A tissue-specific enhancer can activate the promoter of its own or another gene only in one particular tissue and not others: Enhancer is transferred to unrelated gene Gene active in cell type A, not in cell type B Promoter X transcription unit X enhancer Cell type A Promoter Y Promoter Y transcription unit Y Cell type B Promoter Y enhancer + transcription unit Y enhancer transcription unit Y Enhancer is active; Activates promoter High level transcription Enhancer inactive Low level transcription
9 Enhancer or Enhancer Elements Case example: Tissue-specific expression of Insulin gene in vivo. Insulin gene enhancer element linked to gene encoding large T antigen (Ag) of SV40 virus. This construct introduced into a fertilized mouse egg. Egg returned to oviduct of mouse. Expression of large T Ag analyzed in all tissues of the transgenic mouse (using specific antibody). Expression of large T was detectable only in the pancreas (specifically in the ß cells of the pancreatic islets which produce insulin). Enhancer is therefore capable of conferring the specific pattern of insulin gene expression on an unrelated gene in vivo. Hanahan, D., 1985, Nature 315:
10 Enhancer or Enhancer Elements Enhancer elements often contain sequences (motifs) similar to sequences found adjacent to promoters. Octamer motif (ATGCAAAT) of heavy chain Immunoglobulin (Ig) enhancers is also found in Ig promoters. Possible mechanisms of action: By changing chromatin structure leading to nucleosome displacement, By direct interaction with the proteins of the transcriptional apparatus: (a) (b) (c) Enhancer Enhancer Protein factor binds to Enhancer element & slides along DNA Protein factor binds & contacts apparatus by looping out of intervening DNA Bound protein Enhancer Transcription start site Protein factor binds to Enhancer element & contacts transcriptional apparatus via other proteins
11 Models (a) and (b) cannot explain the following finding: Immunoglobulin enhancer activates equally well 2 promoters located 1.7kb and 7.7kb away on the same DNA molecule Models (a) & (b) would postulate that the sliding of factors or the assembly of connecting molecules would stop at the 1 st promoter. E 1.7kb P1 6kb P Atchison & Perry, 1986, Cell 46: Model (c) readily explains the observation showing the critical importance of DNA structure on the action of enhancers. Region between SV40 enhancer & promoter: Removal of multiples of 10 bases (1 helical turn) Removal of bases corresponding to half a helical turn Activity. Activity disrupted. Takahashi et al., 1986, Nature 319:
12 Enhancer-binding proteins bend DNA Enhancer-binding proteins actually bend the DNA so that interactions can occur between regulatory proteins bound at distant sites on DNA. e.g. T-cell receptor α chain gene enhancer. LEF-1 factor binds to a site at the centre of this enhancer, bends DNA, brings other regulatory factors into close proximity: X X Y Y X LEF-1 X Y Y LEF-1
13 LEF-1 LEF-1 Werner and Burley, 1997, Cell 88:
14 LOCUS CONTROL REGIONS (LCRs) LCRs are sequences (additional to promoters & enhancers) that are necessary for high-level gene expression. Influence expression of adjacent genes in a position-independent manner, i.e. regardless of the position of the genes in the genome. Act in a tissue-specific manner. LCR elements have been identified in α- and β- globin gene clusters, the major histocompatibility (MHC) locus, CD2 and lysozyme genes.
15 LOCUS CONTROL REGIONS (LCRs) In the β-globin gene cluster, LCR is located 10-20kb upstream of the β-globin genes. Its deletion leads to a lack of expression of any genes in the cluster. In Humans, this leads to a lethal disease (Hispanic thalassaemia), in which no functional haemoglobin is produced DNase I sensitivity ε Gγ Aγ ψβ δ β LCR
16 LOCUS CONTROL REGIONS (LCRs) LCRs are rich in sequence motifs also found in promoters & enhancers. LCRs function by affecting chromatin structure: When gene is introduced transiently into cells (I.e. exogenous DNA is not packaged into chromatin) LCR has no effect on gene activity. When gene is an integral part of the chromosome LCR affects gene activity. LCR induces DNase I hypersensitivity in adjacent regions (e.g. β-globin cluster). DNase I hypersensitivity is characteristic of active or potentially active genes. Gene lacking LCR will be subject to the influence of adjacent regulatory elements which might repress its expression by directing its organization into a closed chromatin conformation.
17 TRANSCRIPTIONAL CONTROL Transcription Factors (Transcriptional Activators) Proteins that bind to DNA in a sequence-specific manner and regulate the level of transcription. transcriptional activators have characteristic structural features: Helix-turn-helix motif Zinc finger motif The Leucine zipper References: Travers A., 1993, DNA-protein interactions, Chapman & Hall Ptashne, M. & Gann, A., 2002, Genes & Signals, CSH Lab Press. Ptashne, M , Genes & Regulation, CSH Lab Press. Branden, C & Tooze, J., 2000, Introduction to protein structure. 2 nd Ed, Garland Pub.
18 Transcriptional Activators Genome-wide comparison of transcriptional activator families across Eukaryotes No. of members Transcriptional activator f Adapted from Tupler et al., 2001, Nature 409:832 amilies
19 Helix-turn-Helix motif Many transcriptional activators with this type of DNA-binding domain are called homeodomain proteins. Name derived from a group of Drosophila genes (homeotic genes) in which the conserved sequence encoding this structural motif was 1 st observed. Mutations in these homeotic genes transformation of one body part into another during fly s development. These genes encode regulatory proteins activate or repress activity of other genes encoding proteins req.d for development of certain structures. e.g. The Engrailed (Eng) protein binds the identical sequence recognized by Ftz (another homeodomain protein) and blocks gene induction by Ftz. Eng α-helix α-helix Turn DNA binding Adapted from Harrison, 1991, Nature 353:715
20 Zinc finger motif This motif is common in eukaryotic proteins. Est.d 1% of all mammalian genes code for zinc finger proteins. At least 6 different versions of this motif. The first identified was the Cys 2 His 2 finger. Consensus sequence: Tyr/Phe-X-Cys-X 2-4 -Cys-X 3 -Phe/Tyr-X 5 -Leu-X 2 -His-X 3-4 -His This structure binds one Zn 2+ ion through the 2 Cys and 2 His side chains. The transcriptional activators, TFIIIA (for the gene encoding 5S RNA of the ribosome) was the 1 st to be identified bearing this motif. Cys 2 His 2 Cys Cys Zn His His
21 Cys 4 Zinc finger motif The second type is the Cys 4 zinc finger. Found in more than 100 transcriptional activators. Steroid receptor superfamily or now known as nuclear receptors. 2 groups of 4 critical Cys bind a Zn 2+ ion. Cys 2 His 2 proteins generally contain 3 or 4 repeating finger units and bind to DNA as monomers. Cys 4 proteins generally contain only 2-finger units and bind to DNA as homodimers or heterodimers. The yeast Gal4 protein exhibits the Cys 6 zinc finger motif.
22 Leucine zipper Motif present in many transcriptional activators. Contains the hydrophobic leucine at every 7 th position in the C-terminal portion of their DNA-binding domains. These proteins bind to DNA as dimers. Dimers form via hydrophobic interactions between the C-terminal regions of the α-helices, forming a coiled-coil structure. Hydrophobic side chains form a stripe down one side of the α-helix. Hydrophobic stripes make up the interacting surfaces between the helices in the coiled-coil dimer. L L L L L L L L Basic DNA-binding domain
23 Fos and Jun proteins: Examples of transcriptional activators bearing the leucine zipper motif: Jun can bind as a homodimer to the AP1 recognition sequence, TGAGTCAG, transcriptional induction of phorbol esters. Fos cannot bind to DNA alone, but can form a heterodimer with Jun. Jun-Fos heterodimer binds AP1 with 30-fold greater affinity than Jun homodimer.
24 Modular nature of Transcriptional activators transcriptional activators have modular structures (e.g. with a DNA-binding and activation domains). Classic domain-type structure seen in yeast transcriptional activators GCN4. GCN4 induces genes encoding enzymes of amino-acid (a.a.) biosynthesis in response to a.a. starvation. Expt: 60a.a. region (containing DNA-binding site) introduced into cells binds GCN4-responsive genes but fails to activate transcription. This only confirms the DNA-binding domain. A functional test is needed to identify the activation domain.
25 Identify activation domain: Perform Domain-swap experiment to locate activation domain of FactorA: Link various regions of FactorA to DNA-binding domain of FactorB FactorA FactorB Where is activation domain? DNA-binding domain DNA-binding domain Reporter gene activation Response element & binding site for FactorB Reporter gene
26 An extreme example of ( parasitic ) modularity: Herpes simplex virus (HSV) VP16 protein activates the transcription of viral immediate-early genes during lytic infection. VP16 contains a potent activation region. VP16 contains no DNA-binding domain & therefore cannot bind DNA itself. But following infection, VP16 complexes with cellular Oct-1 protein, which binds the sequence, TAATGARAT, in the viral promoters, and Activation is achieved. VP16 ACTIVATION Oct-1 DNA-binding domain Binding site
27 General features of Activating Domains: Do not show strong a.a. sequence similarity amongst transcriptional activators. But in many cases, activating domains contain a very high proportion of acidic a.a., a region of strong negative charge. E.g. 17 acidic a.a. residues were found in the 82-a.a. N-terminal activating domain of the glucocorticoid receptor. E.g. 17 acidic a.a. residues were found in the 60-a.a. activating domain of GCN4. Hence activating domains also known as Acidic Blobs or Negative Noodles. It has been suggested that activating domains adopt an α-helical or an antiparallel β-sheet conformation.
28 TRANSLATIONAL CONTROL Regulation at the level of translation. Significance of Translational Control: Translational control tends to occur in situations where very rapid responses are required. Translational control viewed as supplementing the regulation of transcription, to meet the requirements of particular specialized cases. E.g. following heat shock, it is necessary to: Shut down rapidly enzyme and structural protein synthesis, Rapidly synthesize heat-shock proteins.
29 TRANSLATIONAL CONTROL Some interesting examples of how control is mediated by untranslated region of mrna: Ferritin expression: Control is mediated by sequences in the 5 untranslated region of ferritin mrna. Sequences in this region can fold into a stem-loop structure. Stem-loop structure is stabilized by the Iron-response-element binding protein (IRE-BP) interacting with this structure. Presence of Iron: IRE-BP binds iron and dissociates from the stem-loop in the process, stem-loop structure unfolds, enhanced translation of gene encoding ferritin. IRE-BP Stem loop + Fe Nascent polypeptide IRE-BP Fe ribosome Start of translation Start of translation
30 TRANSLATIONAL CONTROL Transferin receptor expression: Control is mediated by sequences in the 3 untranslated region (important for the stability) of the transferin receptor mrna. Sequences in this region can also fold into a stem-loop structure. Stem-loop structure is stabilized by the Iron-response-element binding protein (IRE-BP) interacting with this structure. Presence of Iron: IRE-BP binds iron and dissociates from the stem-loop in the process, stem-loop structure unfolds, RNA becomes susceptible to nuclease degradation at a rapid rate. Nascent polypeptide IRE-BP Stem loop 5 3 ribosome + Fe IRE-BP Fe Stem loop unfolds 5 3 RNA is rapidly degraded
31 PART II: CONTROL SYSTEMS in Gene Expression Putting it all together: coordinated control of transcriptional regulatory molecules
32 Simple Control: Lactose (lac) Operon in bacteria E. coli can use glucose or lactose as source of carbon and energy. In glucose-containing medium repression of lactose-metabolizing enzymes syn. In lactose-containing medium induction of lactose-metabolizing enzymes syn. Enzymes induced in the presence of lactose are encoded by the lac operon. lacy gene encodes lactose permease (pumps lactose into cell), lacz gene encodes β-galactosidase (splits lactose glucose + galactose), laca gene encodes thiogalactoside transacetylase (function not well understood). These genes (in the operon) are coordinately regulated. I P O Z Y A Another gene (adjacent to lac control region), laci, encodes the lactose repressor. In the absence of lactose: the repressor binds to the (O)perator, blocking the binding of RNA polymerase to (P)romoter. In the presence of lactose/allolactose: The repressor binds to allolactose, structural change to HLH motif can t bind operator, RNA pol can now gain access to the Promoter syn of lacz,y,a-encoded products.
33 Simple Control: Control by Lipid-soluble Steroid Hormones Steroid hormones are signaling compounds that coordinate a range of physiological activities in eukaryotic cells. Lipid-soluble steroids (e.g. cortisol) can directly penetrate the cell membrane. Once inside cell, steroid hormone interacts with specific steroid receptor (also known as nuclear receptor) protein (e.g. Glucocorticoid receptor) in the cytoplasm. DNA binding Hormone-binding Glucocorticoid receptor
34 In the absence of cortisol, Glucocorticoid receptor (GR) is bound in a complex with Hsp90 (heat-shock chaperon ) in the cytoplasm. Upon interaction with its ligand (cortisol), GR undergoes a conformational change: Hsp90 is released. Receptor with bound cortisol translocates into the nucleus. Receptor functions as a transcription activator. DNA-binding domain (Zn finger motif) binds to the glucocorticoid response element (GRE). Activation Domain stimulates transcription of genes. Glucocorticoid Response Element LBD: Ligand-Binding Domain DBD: DNA-Binding Domain AD: Activation Domain
35 Signaling mediated by cell surface receptors Many other extracellular signaling compounds: Cannot penetrate cell membrane, or Lack specific transport mechanism for their uptake. Signaling is transmitted by binding to specific receptor proteins that span across the cell membrane: Binding conformation change in the receptor, Inducing a series of biochem. events within the cell, e.g. phosphorylation of an intracellular protein. This constitutes the 1 st step in the intracellular stage of Signal Transduction. Receptor OUT Cell membrane IN Signaling cpd. P- phosphorylated protein Signal Transduction
36 Direct Signal Transduction Stimulation of cell surface receptor direct activation of a protein that influences transcription activity. Direct system used by many cytokines (e.g. interleukins and interferons). Binding of cytokines to their cognate receptors: activation of transcriptional activator called STAT (signal transducer & activator of transcription). If receptor is a member of the tyrosine kinase family activate STAT directly i.e. phosphorylation of single tyrosine residue (near C-term) of STAT. If receptor is a tyrosine-kinase-associated receptor activation via JAKs (Janus kinases), which auto-phosphorylate activate STAT. Receptor (Tyr kinase family) Receptor (Tyr kinase-associated) STAT dimerizes activation of JAK involves Dimerization
37 Signal Transduction Pathways activated by Interferon (IFN-γ) IFN-γ is secreted by antigen-activated T- helper lymphocytes. Binding of IFN-γ to its receptor induces oligomerization of the IFN-γ -receptor subunits IFNGR1 and IFNGR2, Phosphorylation and activation of Jak1, Jak2, IFNGR1 and Stat1. Stat1 homodimers translocate to the nucleus, bind to γ-activated sequence (GAS) elements and, regulate gene expression with other transcriptional activators (e.g. BRCA1 and MCM5). Several other signal-transduction pathways are activated also in parallel with the Jak Stat1 pathway in response to IFN-γ (shown in the small box). Concensus for DNA-binding 5 -TTN 5-6 AA-3 Adapted from Ramana et. al., 2002, Trends Immunol, 23:96
38 Numerous genes regulated by IFN-γ in macrophages Adapted from Ramana et. al., 2002, Trends Immunol, 23:96
39 Complex Signal Transduction Cascades Activation of receptor represents just the 1 st in a series of steps that eventually lead to one or more transcriptional activators or repressors being switched on or off. MAP (mitogen activated protein) kinase system A minimal signaling module consists of: a MAP kinase (MAPK), a MAP kinase kinase (MKK or MEK), & a MAP kinase kinase kinase (MKKK or MEKK). Signals are transmitted through the module by sequential phosphorylation and activation of these components arranged in a signaling cascade of ser/threonine kinases. Different groups of MAP kinases are activated by different signaling modules that are composed of distinct protein kinases. Three major groups of MAP kinases have been identified by molecular cloning: the extracellular signal regulated kinases (ERKs), the p38 MAP kinases, & the c-jun amino-terminal kinases (JNKs).
40 Mitogen receptor Signal transduction from a mitogen receptor Activated receptor recruits Raf (a protein kinase), Initiates a cascade of phosphorylations: MEK MAPK Rsk Activated MAPK translocates into nucleus and switches on (by phosphorylating) ELK-1 and c-myc MAPK Rsk activates SRF (serum response factor) by phosphorylation. ELK-1, c-myc & SRF are transcriptional activators. MAPK
41 Mammalian MAP kinase signaling pathways Adapted from Dong et al., 2002, Annu. Rev. Immunol 20:55
42 MAP kinase signaling cascades
43 Complex Signal Transduction Cascades The Ras System Ras family of proteins are important intermediates in signal transduction pathways initiating from activated receptor tyrosine kinases (RTKs). Ligands for RTKs include NGF (nerve growth factor), PDGF (platelet-derived growth factor), FGF (fibroblast growth factor), EGF (epidermal growth factor) and Insulin. Ras is a GTP-binding switch that alternates between active state (with bound GTP) and an inactive state (with bound GDP). [1 & 2] Ras activation is facilitated by guanine nucleotide exchange factor (GEF). GEF facilitates dissoc. of GDP from Ras. [3 & 4] GAP (GTPase-activating protein) accelerates the hydrolysis of bound GTP regenerate inactive Ras.GDP
44 Adapter protein & GEF establish link between RTKs and Ras GEF activity of Sos Second messenger pathways Raf MAP kinase pathway
45 Complex Signal Transduction via second messengers Some signal transduction pathways transfer an external signal to the nucleus using an indirect mechanism via second messengers. Second messengers transduce signals from cell surface receptors in several directions, so that a variety of cellular activities respond to one signal. Second messengers: camp (cyclic AMP), cgmp, IP 3, DAG, calcium ions. Levels of camp or cgmp (regulated by cylase and decylase activities), control the activities of various target enzymes. e.g. camp activates protein kinase A phosphorylates CREB (camp ) interacts with p300/cbp modify histones / nucleosome positioning / affect chromatin structure. e.g. Activation of phospholipases cleave phosphatidylinositol-4,5-bisphosphate (PIP 2 ) to give inositol-1,4,5-trisphosphate (IP 3 ) and 1,2-diacylglycerol (DAG). IP 3 increases intracellular [Ca 2+ ] activates p300/cbp IP 3 increases intracellular [Ca 2+ ] activates calmodulin
46 Complex Signal Transduction via second messengers Phospholipase C cleaves PIP 2 to give IP 3 and DAG. IP 3 increases intracellular [Ca 2+ ], which recruits Protein kinase C (PKC) from cytosol to membrane. At membrane, PKC is activated by DAG. Activated PKC then phosphorylates several cellular enzymes. Animation clip: 2 nd messengers Activation via calmodulin
47 Control of immune cell repression & activation Interplay of the various signal transduction pathways to bring about control Stimulating CD4 + Th1 cells with peptide antigen on MHC class II molecules, but in the absence of co-stimulatory signal via CD28, does not activate Th1 cells, but instead, drive them towards an unresponsive state (anergy). Interleukin-2 (IL-2) production is severely impaired (20-fold decrease). The cause: a block in p21 ras activation (due to either inhibited msos activity or altered Ras-GAP function). decrease in signaling through ERK and JNK pathways, decrease in c-fos and JunB induction, transactivation by AP-1 is diminished, transcription activation of the IL-2 gene impaired.
48 Cross-talk between classical & alternative MAPK pathways thought to occur between GTP-p21ras and GTP-Rac Adapted from Schwartz, 1997, Curr Opin Immunol 9:351
49 Adapted from Gerondakis et al., 1998, Curr Opin Immunol 10:353
50 A model gene expression regulatory network Colored circles represent distinct transcriptional activators. Rectangular ovals represent potential target genes in the genome. The color of the rectangular oval indicates which transcriptional activator is regulating its expression in response to the environmental stimulus; in addition, arrows point from each transcriptional activator to its regulated genes. Note that this model can be thought of as an individual regulatory network or as a collection of regulatory networks. Wyrick JJ & Young RA, Curr Opin Genet Dev 2002, 12(2):130-6
DNA Binding Domains: Structural Motifs. Effector Domain. Zinc Fingers. Zinc Fingers, continued. Zif268
 DNA Binding Domains: Structural Motifs Studies of known transcription factors have found several motifs of protein design to allow sequence-specific binding of DNA. We will cover only three of these motifs:
DNA Binding Domains: Structural Motifs Studies of known transcription factors have found several motifs of protein design to allow sequence-specific binding of DNA. We will cover only three of these motifs:
Regulation of gene expression. (Lehninger pg )
 Regulation of gene expression (Lehninger pg. 1072-1085) Today s lecture Gene expression Constitutive, inducible, repressible genes Specificity factors, activators, repressors Negative and positive gene
Regulation of gene expression (Lehninger pg. 1072-1085) Today s lecture Gene expression Constitutive, inducible, repressible genes Specificity factors, activators, repressors Negative and positive gene
Computational Biology I LSM5191 (2003/4)
 Computational Biology I LSM5191 (2003/4) Aylwin Ng, D.Phil Lecture Notes: Transcriptome: Molecular Biology of Gene Expression I Flow of information: DNA to polypeptide DNA Start Exon1 Intron Exon2 Termination
Computational Biology I LSM5191 (2003/4) Aylwin Ng, D.Phil Lecture Notes: Transcriptome: Molecular Biology of Gene Expression I Flow of information: DNA to polypeptide DNA Start Exon1 Intron Exon2 Termination
Chapter 18: Regulation of Gene Expression. 1. Gene Regulation in Bacteria 2. Gene Regulation in Eukaryotes 3. Gene Regulation & Cancer
 Chapter 18: Regulation of Gene Expression 1. Gene Regulation in Bacteria 2. Gene Regulation in Eukaryotes 3. Gene Regulation & Cancer Gene Regulation Gene regulation refers to all aspects of controlling
Chapter 18: Regulation of Gene Expression 1. Gene Regulation in Bacteria 2. Gene Regulation in Eukaryotes 3. Gene Regulation & Cancer Gene Regulation Gene regulation refers to all aspects of controlling
Chapter 25: Regulating Eukaryotic Transcription The Ligand Responsive Activators
 Chapter 25: Regulating Eukaryotic Transcription The Ligand Responsive Activators At least 5 potential gene expression control points Superfamily of Gene Regulators Activation of gene structure Initiation
Chapter 25: Regulating Eukaryotic Transcription The Ligand Responsive Activators At least 5 potential gene expression control points Superfamily of Gene Regulators Activation of gene structure Initiation
BIOLOGY. Chapter 16 GenesExpression
 BIOLOGY Chapter 16 GenesExpression CAMPBELL BIOLOGY TENTH EDITION Reece Urry Cain Wasserman Minorsky Jackson 18 Gene Expression 2014 Pearson Education, Inc. Figure 16.1 Differential Gene Expression results
BIOLOGY Chapter 16 GenesExpression CAMPBELL BIOLOGY TENTH EDITION Reece Urry Cain Wasserman Minorsky Jackson 18 Gene Expression 2014 Pearson Education, Inc. Figure 16.1 Differential Gene Expression results
Regulation of Gene Expression
 Slide 1 Chapter 18 Regulation of Gene Expression PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley with contributions
Slide 1 Chapter 18 Regulation of Gene Expression PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley with contributions
CHAPTER 18 LECTURE NOTES: CONTROL OF GENE EXPRESSION PART B: CONTROL IN EUKARYOTES
 CHAPTER 18 LECTURE NOTES: CONTROL OF GENE EXPRESSION PART B: CONTROL IN EUKARYOTES I. Introduction A. No operon structures in eukaryotes B. Regulation of gene expression is frequently tissue specific.
CHAPTER 18 LECTURE NOTES: CONTROL OF GENE EXPRESSION PART B: CONTROL IN EUKARYOTES I. Introduction A. No operon structures in eukaryotes B. Regulation of gene expression is frequently tissue specific.
CHAPTER 13 LECTURE SLIDES
 CHAPTER 13 LECTURE SLIDES Prepared by Brenda Leady University of Toledo To run the animations you must be in Slideshow View. Use the buttons on the animation to play, pause, and turn audio/text on or off.
CHAPTER 13 LECTURE SLIDES Prepared by Brenda Leady University of Toledo To run the animations you must be in Slideshow View. Use the buttons on the animation to play, pause, and turn audio/text on or off.
Structure/function relationship in DNA-binding proteins
 PHRM 836 September 22, 2015 Structure/function relationship in DNA-binding proteins Devlin Chapter 8.8-9 u General description of transcription factors (TFs) u Sequence-specific interactions between DNA
PHRM 836 September 22, 2015 Structure/function relationship in DNA-binding proteins Devlin Chapter 8.8-9 u General description of transcription factors (TFs) u Sequence-specific interactions between DNA
7.1 The lac Operon 7-1
 7.1 The lac Operon The lac operon was the first operon discovered It contains 3 genes coding for E. coli proteins that permit the bacteria to use the sugar lactose Galactoside permease (lacy) which transports
7.1 The lac Operon The lac operon was the first operon discovered It contains 3 genes coding for E. coli proteins that permit the bacteria to use the sugar lactose Galactoside permease (lacy) which transports
GENE REGULATION IN PROKARYOTES
 GENE REGULATION IN PROKARYOTES Prepared by Brenda Leady, University of Toledo Copyright (c) The McGraw-Hill Companies, Inc. Permission required for reproduction or display. 1 Gene regulation refers to
GENE REGULATION IN PROKARYOTES Prepared by Brenda Leady, University of Toledo Copyright (c) The McGraw-Hill Companies, Inc. Permission required for reproduction or display. 1 Gene regulation refers to
Gene Regulation in Eukaryotes
 Gene Regulation in Eukaryotes The latest estimates are that a human cell, a eukaryotic cell, contains 20,000 25,000 genes. Some of these are expressed in all cells all the time. These so-called housekeeping
Gene Regulation in Eukaryotes The latest estimates are that a human cell, a eukaryotic cell, contains 20,000 25,000 genes. Some of these are expressed in all cells all the time. These so-called housekeeping
Chem 465 Biochemistry II
 Chem 465 Biochemistry II Name: 2 points Multiple choice (4 points apiece): 1. Which of the following is not true of trna molecules? A) The 3'-terminal sequence is -CCA. B) Their anticodons are complementary
Chem 465 Biochemistry II Name: 2 points Multiple choice (4 points apiece): 1. Which of the following is not true of trna molecules? A) The 3'-terminal sequence is -CCA. B) Their anticodons are complementary
Chapter 8 DNA Recognition in Prokaryotes by Helix-Turn-Helix Motifs
 Chapter 8 DNA Recognition in Prokaryotes by Helix-Turn-Helix Motifs 1. Helix-turn-helix proteins 2. Zinc finger proteins 3. Leucine zipper proteins 4. Beta-scaffold factors 5. Others λ-repressor AND CRO
Chapter 8 DNA Recognition in Prokaryotes by Helix-Turn-Helix Motifs 1. Helix-turn-helix proteins 2. Zinc finger proteins 3. Leucine zipper proteins 4. Beta-scaffold factors 5. Others λ-repressor AND CRO
Differential Gene Expression
 Biology 4361 Developmental Biology Differential Gene Expression September 28, 2006 Chromatin Structure ~140 bp ~60 bp Transcriptional Regulation: 1. Packing prevents access CH 3 2. Acetylation ( C O )
Biology 4361 Developmental Biology Differential Gene Expression September 28, 2006 Chromatin Structure ~140 bp ~60 bp Transcriptional Regulation: 1. Packing prevents access CH 3 2. Acetylation ( C O )
Transcription in Prokaryotes. Jörg Bungert, PhD Phone:
 Transcription in Prokaryotes Jörg Bungert, PhD Phone: 352-273-8098 Email: jbungert@ufl.edu Objectives Understand the basic mechanism of transcription. Know the function of promoter elements and associating
Transcription in Prokaryotes Jörg Bungert, PhD Phone: 352-273-8098 Email: jbungert@ufl.edu Objectives Understand the basic mechanism of transcription. Know the function of promoter elements and associating
Differential Gene Expression
 Biology 4361 Developmental Biology Differential Gene Expression June 19, 2008 Differential Gene Expression Overview Chromatin structure Gene anatomy RNA processing and protein production Initiating transcription:
Biology 4361 Developmental Biology Differential Gene Expression June 19, 2008 Differential Gene Expression Overview Chromatin structure Gene anatomy RNA processing and protein production Initiating transcription:
DNA Transcription. Dr Aliwaini
 DNA Transcription 1 DNA Transcription-Introduction The synthesis of an RNA molecule from DNA is called Transcription. All eukaryotic cells have five major classes of RNA: ribosomal RNA (rrna), messenger
DNA Transcription 1 DNA Transcription-Introduction The synthesis of an RNA molecule from DNA is called Transcription. All eukaryotic cells have five major classes of RNA: ribosomal RNA (rrna), messenger
Chapter 11: Regulation of Gene Expression
 Chapter Review 1. It has long been known that there is probably a genetic link for alcoholism. Researchers studying rats have begun to elucidate this link. Briefly describe the genetic mechanism found
Chapter Review 1. It has long been known that there is probably a genetic link for alcoholism. Researchers studying rats have begun to elucidate this link. Briefly describe the genetic mechanism found
Mechanism of action-1
 Mechanism of action-1 receptors: mediators of hormone action, membrane associated vs. intracellular receptors: measurements of receptor - ligand interactions, regulation mechanisms surface - receptors:
Mechanism of action-1 receptors: mediators of hormone action, membrane associated vs. intracellular receptors: measurements of receptor - ligand interactions, regulation mechanisms surface - receptors:
Lecture 9 Controlling gene expression
 Lecture 9 Controlling gene expression BIOLOGY Campbell, Reece and Mitchell Chapter 18 334- (352-356) Every cell in your body contains the same number of genes approximately 35, 000 DNA is wound around
Lecture 9 Controlling gene expression BIOLOGY Campbell, Reece and Mitchell Chapter 18 334- (352-356) Every cell in your body contains the same number of genes approximately 35, 000 DNA is wound around
Transcription in Eukaryotes
 Transcription in Eukaryotes Biology I Hayder A Giha Transcription Transcription is a DNA-directed synthesis of RNA, which is the first step in gene expression. Gene expression, is transformation of the
Transcription in Eukaryotes Biology I Hayder A Giha Transcription Transcription is a DNA-directed synthesis of RNA, which is the first step in gene expression. Gene expression, is transformation of the
Bi 8 Lecture 7. Ellen Rothenberg 26 January Reading: Ch. 3, pp ; panel 3-1
 Bi 8 Lecture 7 PROTEIN STRUCTURE, Functional analysis, and evolution Ellen Rothenberg 26 January 2016 Reading: Ch. 3, pp. 109-134; panel 3-1 (end with free amine) aromatic, hydrophobic small, hydrophilic
Bi 8 Lecture 7 PROTEIN STRUCTURE, Functional analysis, and evolution Ellen Rothenberg 26 January 2016 Reading: Ch. 3, pp. 109-134; panel 3-1 (end with free amine) aromatic, hydrophobic small, hydrophilic
Protein Synthesis Notes
 Protein Synthesis Notes Protein Synthesis: Overview Transcription: synthesis of mrna under the direction of DNA. Translation: actual synthesis of a polypeptide under the direction of mrna. Transcription
Protein Synthesis Notes Protein Synthesis: Overview Transcription: synthesis of mrna under the direction of DNA. Translation: actual synthesis of a polypeptide under the direction of mrna. Transcription
Year III Pharm.D Dr. V. Chitra
 Year III Pharm.D Dr. V. Chitra 1 Genome entire genetic material of an individual Transcriptome set of transcribed sequences Proteome set of proteins encoded by the genome 2 Only one strand of DNA serves
Year III Pharm.D Dr. V. Chitra 1 Genome entire genetic material of an individual Transcriptome set of transcribed sequences Proteome set of proteins encoded by the genome 2 Only one strand of DNA serves
GENE REGULATION slide shows by Kim Foglia modified Slides with blue edges are Kim s
 GENE REGULATION slide shows by Kim Foglia modified Slides with blue edges are Kim s 2007-2008 Bacterial metabolism Bacteria need to respond quickly to changes in their environment STOP GO if they have
GENE REGULATION slide shows by Kim Foglia modified Slides with blue edges are Kim s 2007-2008 Bacterial metabolism Bacteria need to respond quickly to changes in their environment STOP GO if they have
Fig Ch 17: From Gene to Protein
 Fig. 17-1 Ch 17: From Gene to Protein Basic Principles of Transcription and Translation RNA is the intermediate between genes and the proteins for which they code Transcription is the synthesis of RNA
Fig. 17-1 Ch 17: From Gene to Protein Basic Principles of Transcription and Translation RNA is the intermediate between genes and the proteins for which they code Transcription is the synthesis of RNA
Review of Protein (one or more polypeptide) A polypeptide is a long chain of..
 Gene expression Review of Protein (one or more polypeptide) A polypeptide is a long chain of.. In a protein, the sequence of amino acid determines its which determines the protein s A protein with an enzymatic
Gene expression Review of Protein (one or more polypeptide) A polypeptide is a long chain of.. In a protein, the sequence of amino acid determines its which determines the protein s A protein with an enzymatic
M I C R O B I O L O G Y WITH DISEASES BY TAXONOMY, THIRD EDITION
 M I C R O B I O L O G Y WITH DISEASES BY TAXONOMY, THIRD EDITION Chapter 7 Microbial Genetics Lecture prepared by Mindy Miller-Kittrell, University of Tennessee, Knoxville The Structure and Replication
M I C R O B I O L O G Y WITH DISEASES BY TAXONOMY, THIRD EDITION Chapter 7 Microbial Genetics Lecture prepared by Mindy Miller-Kittrell, University of Tennessee, Knoxville The Structure and Replication
Gene Expression: Transcription
 Gene Expression: Transcription The majority of genes are expressed as the proteins they encode. The process occurs in two steps: Transcription = DNA RNA Translation = RNA protein Taken together, they make
Gene Expression: Transcription The majority of genes are expressed as the proteins they encode. The process occurs in two steps: Transcription = DNA RNA Translation = RNA protein Taken together, they make
REGULATION OF GENE EXPRESSION
 REGULATION OF GENE EXPRESSION Each cell of a living organism contains thousands of genes. But all genes do not function at a time. Genes function according to requirements of the cell. Genes control the
REGULATION OF GENE EXPRESSION Each cell of a living organism contains thousands of genes. But all genes do not function at a time. Genes function according to requirements of the cell. Genes control the
Genetics Lecture Notes Lectures 17 19
 Genetics Lecture Notes 7.03 2005 Lectures 17 19 Lecture 17 Gene Regulation We are now going to look at ways that genetics can be used to study gene regulation. The issue is how cells adjust the expression
Genetics Lecture Notes 7.03 2005 Lectures 17 19 Lecture 17 Gene Regulation We are now going to look at ways that genetics can be used to study gene regulation. The issue is how cells adjust the expression
TRANSCRIPTION COMPARISON OF DNA & RNA TRANSCRIPTION. Umm AL Qura University. Sugar Ribose Deoxyribose. Bases AUCG ATCG. Strand length Short Long
 Umm AL Qura University TRANSCRIPTION Dr Neda Bogari TRANSCRIPTION COMPARISON OF DNA & RNA RNA DNA Sugar Ribose Deoxyribose Bases AUCG ATCG Strand length Short Long No. strands One Two Helix Single Double
Umm AL Qura University TRANSCRIPTION Dr Neda Bogari TRANSCRIPTION COMPARISON OF DNA & RNA RNA DNA Sugar Ribose Deoxyribose Bases AUCG ATCG Strand length Short Long No. strands One Two Helix Single Double
Nucleic acids deoxyribonucleic acid (DNA) ribonucleic acid (RNA) nucleotide
 Nucleic Acids Nucleic acids are molecules that store information for cellular growth and reproduction There are two types of nucleic acids: - deoxyribonucleic acid (DNA) and ribonucleic acid (RNA) These
Nucleic Acids Nucleic acids are molecules that store information for cellular growth and reproduction There are two types of nucleic acids: - deoxyribonucleic acid (DNA) and ribonucleic acid (RNA) These
TRANSCRIPTION AND PROCESSING OF RNA
 TRANSCRIPTION AND PROCESSING OF RNA 1. The steps of gene expression. 2. General characterization of transcription: steps, components of transcription apparatus. 3. Transcription of eukaryotic structural
TRANSCRIPTION AND PROCESSING OF RNA 1. The steps of gene expression. 2. General characterization of transcription: steps, components of transcription apparatus. 3. Transcription of eukaryotic structural
DNA Transcription. Visualizing Transcription. The Transcription Process
 DNA Transcription By: Suzanne Clancy, Ph.D. 2008 Nature Education Citation: Clancy, S. (2008) DNA transcription. Nature Education 1(1) If DNA is a book, then how is it read? Learn more about the DNA transcription
DNA Transcription By: Suzanne Clancy, Ph.D. 2008 Nature Education Citation: Clancy, S. (2008) DNA transcription. Nature Education 1(1) If DNA is a book, then how is it read? Learn more about the DNA transcription
Transcription Eukaryotic Cells
 Transcription Eukaryotic Cells Packet #20 1 Introduction Transcription is the process in which genetic information, stored in a strand of DNA (gene), is copied into a strand of RNA. Protein-encoding genes
Transcription Eukaryotic Cells Packet #20 1 Introduction Transcription is the process in which genetic information, stored in a strand of DNA (gene), is copied into a strand of RNA. Protein-encoding genes
Prokaryotic Transcription
 Prokaryotic Transcription Transcription Basics DNA is the genetic material Nucleic acid Capable of self-replication and synthesis of RNA RNA is the middle man Nucleic acid Structure and base sequence are
Prokaryotic Transcription Transcription Basics DNA is the genetic material Nucleic acid Capable of self-replication and synthesis of RNA RNA is the middle man Nucleic acid Structure and base sequence are
From Gene to Protein transcription, messenger RNA (mrna) translation, RNA processing triplet code, template strand, codons,
 From Gene to Protein I. Transcription and translation are the two main processes linking gene to protein. A. RNA is chemically similar to DNA, except that it contains ribose as its sugar and substitutes
From Gene to Protein I. Transcription and translation are the two main processes linking gene to protein. A. RNA is chemically similar to DNA, except that it contains ribose as its sugar and substitutes
CHAPTERS , 17: Eukaryotic Genetics
 CHAPTERS 14.1 14.6, 17: Eukaryotic Genetics 1. Review the levels of DNA packing within the eukaryote nucleus. Label each level. (A similar diagram is on pg 188 of your textbook.) 2. How do the coding regions
CHAPTERS 14.1 14.6, 17: Eukaryotic Genetics 1. Review the levels of DNA packing within the eukaryote nucleus. Label each level. (A similar diagram is on pg 188 of your textbook.) 2. How do the coding regions
Chapter 4: How Cells Work
 Chapter 4: How Cells Work David Shonnard Department of Chemical Engineering 1 Presentation Outline: l l l l l Introduction : Central Dogma DNA Replication: Preserving and Propagating DNA Transcription:
Chapter 4: How Cells Work David Shonnard Department of Chemical Engineering 1 Presentation Outline: l l l l l Introduction : Central Dogma DNA Replication: Preserving and Propagating DNA Transcription:
Molecular Genetics Student Objectives
 Molecular Genetics Student Objectives Exam 1: Enduring understanding 3.A: Heritable information provides for continuity of life. Essential knowledge 3.A.1: DNA, and in some cases RNA, is the primary source
Molecular Genetics Student Objectives Exam 1: Enduring understanding 3.A: Heritable information provides for continuity of life. Essential knowledge 3.A.1: DNA, and in some cases RNA, is the primary source
Einführung in die Genetik
 Einführung in die Genetik Prof. Dr. Kay Schneitz (EBio Pflanzen) http://plantdev.bio.wzw.tum.de schneitz@wzw.tum.de Prof. Dr. Claus Schwechheimer (PlaSysBiol) http://wzw.tum.de/sysbiol claus.schwechheimer@wzw.tum.de
Einführung in die Genetik Prof. Dr. Kay Schneitz (EBio Pflanzen) http://plantdev.bio.wzw.tum.de schneitz@wzw.tum.de Prof. Dr. Claus Schwechheimer (PlaSysBiol) http://wzw.tum.de/sysbiol claus.schwechheimer@wzw.tum.de
I. Prokaryotic Gene Regulation. Figure 1: Operon. Operon:
 I. Prokaryotic Gene Regulation Figure 1: Operon Operon: a) Regulatory Elements consist of an Operator that serves as the on-off switch for the genes of the operon. Also contains a promoter for the Structural
I. Prokaryotic Gene Regulation Figure 1: Operon Operon: a) Regulatory Elements consist of an Operator that serves as the on-off switch for the genes of the operon. Also contains a promoter for the Structural
Eukaryotic & Prokaryotic Transcription. RNA polymerases
 Eukaryotic & Prokaryotic Transcription RNA polymerases RNA Polymerases A. E. coli RNA polymerase 1. core enzyme = ββ'(α)2 has catalytic activity but cannot recognize start site of transcription ~500,000
Eukaryotic & Prokaryotic Transcription RNA polymerases RNA Polymerases A. E. coli RNA polymerase 1. core enzyme = ββ'(α)2 has catalytic activity but cannot recognize start site of transcription ~500,000
AP Biology Gene Expression/Biotechnology REVIEW
 AP Biology Gene Expression/Biotechnology REVIEW Multiple Choice Identify the choice that best completes the statement or answers the question. 1. Gene expression can be a. regulated before transcription.
AP Biology Gene Expression/Biotechnology REVIEW Multiple Choice Identify the choice that best completes the statement or answers the question. 1. Gene expression can be a. regulated before transcription.
Differential Gene Expression
 Biology 4361 - Developmental Biology Differential Gene Expression June 18, 2009 Differential Gene Expression Overview Chromatin structure Gene anatomy RNA processing and protein production Initiating transcription:
Biology 4361 - Developmental Biology Differential Gene Expression June 18, 2009 Differential Gene Expression Overview Chromatin structure Gene anatomy RNA processing and protein production Initiating transcription:
Regulation of Gene Expression in Eukaryotes
 12 Regulation of Gene Expression in Eukaryotes WORKING WITH THE FIGURES 1. In Figure 12-4, certain mutations decrease the relative transcription rate of the -globin gene. Where are these mutations located,
12 Regulation of Gene Expression in Eukaryotes WORKING WITH THE FIGURES 1. In Figure 12-4, certain mutations decrease the relative transcription rate of the -globin gene. Where are these mutations located,
Einführung in die Genetik
 Einführung in die Genetik Prof. Dr. Kay Schneitz (EBio Pflanzen) http://plantdev.bio.wzw.tum.de schneitz@wzw.tum.de Twitter: @PlantDevTUM, #genetiktum FB: Plant Development TUM Prof. Dr. Claus Schwechheimer
Einführung in die Genetik Prof. Dr. Kay Schneitz (EBio Pflanzen) http://plantdev.bio.wzw.tum.de schneitz@wzw.tum.de Twitter: @PlantDevTUM, #genetiktum FB: Plant Development TUM Prof. Dr. Claus Schwechheimer
Chapter 24: Promoters and Enhancers
 Chapter 24: Promoters and Enhancers A typical gene transcribed by RNA polymerase II has a promoter that usually extends upstream from the site where transcription is initiated the (#1) of transcription
Chapter 24: Promoters and Enhancers A typical gene transcribed by RNA polymerase II has a promoter that usually extends upstream from the site where transcription is initiated the (#1) of transcription
What is RNA? Another type of nucleic acid A working copy of DNA Does not matter if it is damaged or destroyed
 RNA Section 3.1 What is RNA? Another type of nucleic acid A working copy of DNA Does not matter if it is damaged or destroyed Used to direct the production of proteins that determines an organisms characteristics
RNA Section 3.1 What is RNA? Another type of nucleic acid A working copy of DNA Does not matter if it is damaged or destroyed Used to direct the production of proteins that determines an organisms characteristics
7.06 Cell Biology QUIZ #3
 Recitation Section: 7.06 Cell Biology QUIZ #3 This is an open book exam, and you are allowed access to books and notes, but not computers or any other types of electronic devices. Please write your answers
Recitation Section: 7.06 Cell Biology QUIZ #3 This is an open book exam, and you are allowed access to books and notes, but not computers or any other types of electronic devices. Please write your answers
Developmental Biology BY1101 P. Murphy
 Developmental Biology BY1101 P. Murphy Lecture 7 Cellular differentiation and the regulation of gene expression. In this lecture we looked at two main questions: How is gene expression regulated? (revision
Developmental Biology BY1101 P. Murphy Lecture 7 Cellular differentiation and the regulation of gene expression. In this lecture we looked at two main questions: How is gene expression regulated? (revision
Themes: RNA and RNA Processing. Messenger RNA (mrna) What is a gene? RNA is very versatile! RNA-RNA interactions are very important!
 Themes: RNA is very versatile! RNA and RNA Processing Chapter 14 RNA-RNA interactions are very important! Prokaryotes and Eukaryotes have many important differences. Messenger RNA (mrna) Carries genetic
Themes: RNA is very versatile! RNA and RNA Processing Chapter 14 RNA-RNA interactions are very important! Prokaryotes and Eukaryotes have many important differences. Messenger RNA (mrna) Carries genetic
Spring 2006 Biochemistry 302 Exam 2
 Name Spring 2006 Biochemistry 302 Exam 2 Directions: This exam has 45 questions/problems totaling 110 points. Check to make sure you have seven pages. Some questions have multiple parts so read each one
Name Spring 2006 Biochemistry 302 Exam 2 Directions: This exam has 45 questions/problems totaling 110 points. Check to make sure you have seven pages. Some questions have multiple parts so read each one
LECTURE: 22 IMMUNOGLOBULIN DIVERSITIES LEARNING OBJECTIVES: The student should be able to:
 LECTURE: 22 Title IMMUNOGLOBULIN DIVERSITIES LEARNING OBJECTIVES: The student should be able to: Identify the chromosome that contains the gene segments that encode the surface immunoglobulin heavy chain
LECTURE: 22 Title IMMUNOGLOBULIN DIVERSITIES LEARNING OBJECTIVES: The student should be able to: Identify the chromosome that contains the gene segments that encode the surface immunoglobulin heavy chain
Make the protein through the genetic dogma process.
 Make the protein through the genetic dogma process. Coding Strand 5 AGCAATCATGGATTGGGTACATTTGTAACTGT 3 Template Strand mrna Protein Complete the table. DNA strand DNA s strand G mrna A C U G T A T Amino
Make the protein through the genetic dogma process. Coding Strand 5 AGCAATCATGGATTGGGTACATTTGTAACTGT 3 Template Strand mrna Protein Complete the table. DNA strand DNA s strand G mrna A C U G T A T Amino
Genomics and Gene Recognition Genes and Blue Genes
 Genomics and Gene Recognition Genes and Blue Genes November 3, 2004 Eukaryotic Gene Structure eukaryotic genomes are considerably more complex than those of prokaryotes eukaryotic cells have organelles
Genomics and Gene Recognition Genes and Blue Genes November 3, 2004 Eukaryotic Gene Structure eukaryotic genomes are considerably more complex than those of prokaryotes eukaryotic cells have organelles
Chapter Fundamental Molecular Genetic Mechanisms
 Chapter 5-1 - Fundamental Molecular Genetic Mechanisms 5.1 Structure of Nucleic Acids 5.2 Transcription of Protein-Coding Genes and Formation of Functional mrna 5.3 The Decoding of mrna by trnas 5.4 Stepwise
Chapter 5-1 - Fundamental Molecular Genetic Mechanisms 5.1 Structure of Nucleic Acids 5.2 Transcription of Protein-Coding Genes and Formation of Functional mrna 5.3 The Decoding of mrna by trnas 5.4 Stepwise
Prokaryotic Transcription
 Prokaryotic Transcription Contents 1 The Lactose Intolerance of Bacteria 2 The Lac Operon 3 Lac Operon Simulation 4 LacZ as a reporter gene The Lactose Intolerance of Bacteria The standard growth kinetics
Prokaryotic Transcription Contents 1 The Lactose Intolerance of Bacteria 2 The Lac Operon 3 Lac Operon Simulation 4 LacZ as a reporter gene The Lactose Intolerance of Bacteria The standard growth kinetics
Ch. 10 Notes DNA: Transcription and Translation
 Ch. 10 Notes DNA: Transcription and Translation GOALS Compare the structure of RNA with that of DNA Summarize the process of transcription Relate the role of codons to the sequence of amino acids that
Ch. 10 Notes DNA: Transcription and Translation GOALS Compare the structure of RNA with that of DNA Summarize the process of transcription Relate the role of codons to the sequence of amino acids that
Regulatory Dynamics in Engineered Gene Networks
 Regulatory Dynamics in Engineered Gene Networks The Physico-chemical Foundation of Transcriptional Regulation with Applications to Systems Biology Mads Kærn Boston University Center for BioDynamics Center
Regulatory Dynamics in Engineered Gene Networks The Physico-chemical Foundation of Transcriptional Regulation with Applications to Systems Biology Mads Kærn Boston University Center for BioDynamics Center
Introduction to Proteins
 Introduction to Proteins Lecture 4 Module I: Molecular Structure & Metabolism Molecular Cell Biology Core Course (GSND5200) Matthew Neiditch - Room E450U ICPH matthew.neiditch@umdnj.edu What is a protein?
Introduction to Proteins Lecture 4 Module I: Molecular Structure & Metabolism Molecular Cell Biology Core Course (GSND5200) Matthew Neiditch - Room E450U ICPH matthew.neiditch@umdnj.edu What is a protein?
Chapter 31. Transcription and RNA processing
 Chapter 31 Transcription and RNA processing RNA polymerase (RNAP) E. coli promoters Components of E. coli RNA Polymerase Holoenzyme (α 2 ββ'ωσ) Structure of prokaryotic RNAP The closed and open state of
Chapter 31 Transcription and RNA processing RNA polymerase (RNAP) E. coli promoters Components of E. coli RNA Polymerase Holoenzyme (α 2 ββ'ωσ) Structure of prokaryotic RNAP The closed and open state of
RNA and Protein Synthesis
 Harriet Wilson, Lecture Notes Bio. Sci. 4 - Microbiology Sierra College RNA and Protein Synthesis Considerable evidence suggests that RNA molecules evolved prior to DNA molecules and proteins, and that
Harriet Wilson, Lecture Notes Bio. Sci. 4 - Microbiology Sierra College RNA and Protein Synthesis Considerable evidence suggests that RNA molecules evolved prior to DNA molecules and proteins, and that
The Two-Hybrid System
 Encyclopedic Reference of Genomics and Proteomics in Molecular Medicine The Two-Hybrid System Carolina Vollert & Peter Uetz Institut für Genetik Forschungszentrum Karlsruhe PO Box 3640 D-76021 Karlsruhe
Encyclopedic Reference of Genomics and Proteomics in Molecular Medicine The Two-Hybrid System Carolina Vollert & Peter Uetz Institut für Genetik Forschungszentrum Karlsruhe PO Box 3640 D-76021 Karlsruhe
Chapter 8 Lecture Outline. Transcription, Translation, and Bioinformatics
 Chapter 8 Lecture Outline Transcription, Translation, and Bioinformatics Replication, Transcription, Translation n Repetitive processes Build polymers of nucleotides or amino acids n All have 3 major steps
Chapter 8 Lecture Outline Transcription, Translation, and Bioinformatics Replication, Transcription, Translation n Repetitive processes Build polymers of nucleotides or amino acids n All have 3 major steps
CHAPTER 17 FROM GENE TO PROTEIN. Section C: The Synthesis of Protein
 CHAPTER 17 FROM GENE TO PROTEIN Section C: The Synthesis of Protein 1. Translation is the RNA-directed synthesis of a polypeptide: a closer look 2. Signal peptides target some eukaryotic polypeptides to
CHAPTER 17 FROM GENE TO PROTEIN Section C: The Synthesis of Protein 1. Translation is the RNA-directed synthesis of a polypeptide: a closer look 2. Signal peptides target some eukaryotic polypeptides to
Zool 3200: Cell Biology Exam 3 3/6/15
 Name: Trask Zool 3200: Cell Biology Exam 3 3/6/15 Answer each of the following questions in the space provided; circle the correct answer or answers for each multiple choice question and circle either
Name: Trask Zool 3200: Cell Biology Exam 3 3/6/15 Answer each of the following questions in the space provided; circle the correct answer or answers for each multiple choice question and circle either
Chapter 9-II - Transcriptional Control of Gene Expression
 Chapter 9-II - Transcriptional Control of Gene Expression Transcriptional Control of Gene Expression 9.3 RNA Polymerase II Promoters and General Transcription Factors Three types of promoter sequences
Chapter 9-II - Transcriptional Control of Gene Expression Transcriptional Control of Gene Expression 9.3 RNA Polymerase II Promoters and General Transcription Factors Three types of promoter sequences
Name: Class: Date: ID: A
 Class: _ Date: _ CH 12 Review Multiple Choice Identify the choice that best completes the statement or answers the question. 1. How many codons are needed to specify three amino acids? a. 6 c. 3 b. 12
Class: _ Date: _ CH 12 Review Multiple Choice Identify the choice that best completes the statement or answers the question. 1. How many codons are needed to specify three amino acids? a. 6 c. 3 b. 12
Summary 12 1 DNA RNA and Protein Synthesis Chromosomes and DNA Replication. Name Class Date
 Chapter 12 Summary DNA and RNA 12 1 DNA To understand genetics, biologists had to learn the chemical structure of the gene. Frederick Griffith first learned that some factor from dead, disease-causing
Chapter 12 Summary DNA and RNA 12 1 DNA To understand genetics, biologists had to learn the chemical structure of the gene. Frederick Griffith first learned that some factor from dead, disease-causing
8/21/2014. From Gene to Protein
 From Gene to Protein Chapter 17 Objectives Describe the contributions made by Garrod, Beadle, and Tatum to our understanding of the relationship between genes and enzymes Briefly explain how information
From Gene to Protein Chapter 17 Objectives Describe the contributions made by Garrod, Beadle, and Tatum to our understanding of the relationship between genes and enzymes Briefly explain how information
Bio11 Announcements. Ch 21: DNA Biology and Technology. DNA Functions. DNA and RNA Structure. How do DNA and RNA differ? What are genes?
 Bio11 Announcements TODAY Genetics (review) and quiz (CP #4) Structure and function of DNA Extra credit due today Next week in lab: Case study presentations Following week: Lab Quiz 2 Ch 21: DNA Biology
Bio11 Announcements TODAY Genetics (review) and quiz (CP #4) Structure and function of DNA Extra credit due today Next week in lab: Case study presentations Following week: Lab Quiz 2 Ch 21: DNA Biology
Gene Expression and Heritable Phenotype. CBS520 Eric Nabity
 Gene Expression and Heritable Phenotype CBS520 Eric Nabity DNA is Just the Beginning DNA was determined to be the genetic material, and the structure was identified as a (double stranded) double helix.
Gene Expression and Heritable Phenotype CBS520 Eric Nabity DNA is Just the Beginning DNA was determined to be the genetic material, and the structure was identified as a (double stranded) double helix.
BEADLE & TATUM EXPERIMENT
 FROM DNA TO PROTEINS: gene expression Chapter 14 LECTURE OBJECTIVES What Is the Evidence that Genes Code for Proteins? How Does Information Flow from Genes to Proteins? How Is the Information Content in
FROM DNA TO PROTEINS: gene expression Chapter 14 LECTURE OBJECTIVES What Is the Evidence that Genes Code for Proteins? How Does Information Flow from Genes to Proteins? How Is the Information Content in
Protein Synthesis. Presented by Dr. Mohammad Saadeh The requirements for the Pharmaceutical Biochemistry I Philadelphia University Faculty of pharmacy
 Protein Synthesis Presented by Dr. Mohammad Saadeh The requirements for the Pharmaceutical Biochemistry I Philadelphia University Faculty of pharmacy STRUCTURE OF RNA RNA, adenine forms a base pair with
Protein Synthesis Presented by Dr. Mohammad Saadeh The requirements for the Pharmaceutical Biochemistry I Philadelphia University Faculty of pharmacy STRUCTURE OF RNA RNA, adenine forms a base pair with
Transcription & post transcriptional modification
 Transcription & post transcriptional modification Transcription The synthesis of RNA molecules using DNA strands as the templates so that the genetic information can be transferred from DNA to RNA Similarity
Transcription & post transcriptional modification Transcription The synthesis of RNA molecules using DNA strands as the templates so that the genetic information can be transferred from DNA to RNA Similarity
Answers to Module 1. An obligate aerobe is an organism that has an absolute requirement of oxygen for growth.
 Answers to Module 1 Short Answers 1) What is an obligate aerobe? An obligate aerobe is an organism that has an absolute requirement of oxygen for growth. What about facultative anaerobe? 2) Distinguish
Answers to Module 1 Short Answers 1) What is an obligate aerobe? An obligate aerobe is an organism that has an absolute requirement of oxygen for growth. What about facultative anaerobe? 2) Distinguish
Transcriptional Regulation in Eukaryotes
 Transcriptional Regulation in Eukaryotes Concepts, Strategies, and Techniques Michael Carey Stephen T. Smale COLD SPRING HARBOR LABORATORY PRESS NEW YORK 2000 Cold Spring Harbor Laboratory Press, 0-87969-537-4
Transcriptional Regulation in Eukaryotes Concepts, Strategies, and Techniques Michael Carey Stephen T. Smale COLD SPRING HARBOR LABORATORY PRESS NEW YORK 2000 Cold Spring Harbor Laboratory Press, 0-87969-537-4
Read the question carefully before answering. Think before you write. If I can not read your handwriting, I will count the question wrong.
 Name KEY Note Total points added up to only 98 points so everyone received 2 free points to make total points 100. Biology 201 (Genetics) Exam #3 23 November 2004 Read the question carefully before answering.
Name KEY Note Total points added up to only 98 points so everyone received 2 free points to make total points 100. Biology 201 (Genetics) Exam #3 23 November 2004 Read the question carefully before answering.
Division Ave. High School AP Biology
 Control of Eukaryotic Genes 2007-2008 The BIG Questions n How are genes turned on & off in eukaryotes? n How do cells with the same genes differentiate to perform completely different, specialized functions?
Control of Eukaryotic Genes 2007-2008 The BIG Questions n How are genes turned on & off in eukaryotes? n How do cells with the same genes differentiate to perform completely different, specialized functions?
Transcription. DNA to RNA
 Transcription from DNA to RNA The Central Dogma of Molecular Biology replication DNA RNA Protein transcription translation Why call it transcription and translation? transcription is such a direct copy
Transcription from DNA to RNA The Central Dogma of Molecular Biology replication DNA RNA Protein transcription translation Why call it transcription and translation? transcription is such a direct copy
RNA synthesis/transcription I Biochemistry 302. February 6, 2004 Bob Kelm
 RNA synthesis/transcription I Biochemistry 302 February 6, 2004 Bob Kelm Overview of RNA classes Messenger RNA (mrna) Encodes protein Relatively short half-life ( 3 min in E. coli, 30 min in eukaryotic
RNA synthesis/transcription I Biochemistry 302 February 6, 2004 Bob Kelm Overview of RNA classes Messenger RNA (mrna) Encodes protein Relatively short half-life ( 3 min in E. coli, 30 min in eukaryotic
Self-test Quiz for Chapter 12 (From DNA to Protein: Genotype to Phenotype)
 Self-test Quiz for Chapter 12 (From DNA to Protein: Genotype to Phenotype) Question#1: One-Gene, One-Polypeptide The figure below shows the results of feeding trials with one auxotroph strain of Neurospora
Self-test Quiz for Chapter 12 (From DNA to Protein: Genotype to Phenotype) Question#1: One-Gene, One-Polypeptide The figure below shows the results of feeding trials with one auxotroph strain of Neurospora
Name Class Date. Practice Test
 Name Class Date 12 DNA Practice Test Multiple Choice Write the letter that best answers the question or completes the statement on the line provided. 1. What do bacteriophages infect? a. mice. c. viruses.
Name Class Date 12 DNA Practice Test Multiple Choice Write the letter that best answers the question or completes the statement on the line provided. 1. What do bacteriophages infect? a. mice. c. viruses.
Matakuliah Genetika (BIO612206) Jurusan Biologi FMIPA Universitas Lampung. Priyambodo, M.Sc. staff.unila.ac.id/priyambodo
 Matakuliah Genetika (BIO612206) Jurusan Biologi FMIPA Universitas Lampung Priyambodo, M.Sc. staff.unila.ac.id/priyambodo Prokariotik Eukariotik staff.unila.ac.id/priyambodo Regulasi ekspresi gen pada
Matakuliah Genetika (BIO612206) Jurusan Biologi FMIPA Universitas Lampung Priyambodo, M.Sc. staff.unila.ac.id/priyambodo Prokariotik Eukariotik staff.unila.ac.id/priyambodo Regulasi ekspresi gen pada
Protein Synthesis & Gene Expression
 DNA provides the instructions for how to build proteins Each gene dictates how to build a single protein in prokaryotes The sequence of nucleotides (AGCT) in DNA dictates the order of amino acids that
DNA provides the instructions for how to build proteins Each gene dictates how to build a single protein in prokaryotes The sequence of nucleotides (AGCT) in DNA dictates the order of amino acids that
The Genetic Code and Transcription. Chapter 12 Honors Genetics Ms. Susan Chabot
 The Genetic Code and Transcription Chapter 12 Honors Genetics Ms. Susan Chabot TRANSCRIPTION Copy SAME language DNA to RNA Nucleic Acid to Nucleic Acid TRANSLATION Copy DIFFERENT language RNA to Amino
The Genetic Code and Transcription Chapter 12 Honors Genetics Ms. Susan Chabot TRANSCRIPTION Copy SAME language DNA to RNA Nucleic Acid to Nucleic Acid TRANSLATION Copy DIFFERENT language RNA to Amino
Protein Synthesis
 HEBISD Student Expectations: Identify that RNA Is a nucleic acid with a single strand of nucleotides Contains the 5-carbon sugar ribose Contains the nitrogen bases A, G, C and U instead of T. The U is
HEBISD Student Expectations: Identify that RNA Is a nucleic acid with a single strand of nucleotides Contains the 5-carbon sugar ribose Contains the nitrogen bases A, G, C and U instead of T. The U is
PROTEIN SYNTHESIS Flow of Genetic Information The flow of genetic information can be symbolized as: DNA RNA Protein
 PROTEIN SYNTHESIS Flow of Genetic Information The flow of genetic information can be symbolized as: DNA RNA Protein This is also known as: The central dogma of molecular biology Protein Proteins are made
PROTEIN SYNTHESIS Flow of Genetic Information The flow of genetic information can be symbolized as: DNA RNA Protein This is also known as: The central dogma of molecular biology Protein Proteins are made
Genome Architecture Structural Subdivisons
 Lecture 4 Hierarchical Organization of the Genome by John R. Finnerty Genome Architecture Structural Subdivisons 1. Nucleotide : monomer building block of DNA 2. DNA : polymer string of nucleotides 3.
Lecture 4 Hierarchical Organization of the Genome by John R. Finnerty Genome Architecture Structural Subdivisons 1. Nucleotide : monomer building block of DNA 2. DNA : polymer string of nucleotides 3.
Chapter 10: Gene Expression and Regulation
 Chapter 10: Gene Expression and Regulation Fact 1: DNA contains information but is unable to carry out actions Fact 2: Proteins are the workhorses but contain no information THUS Information in DNA must
Chapter 10: Gene Expression and Regulation Fact 1: DNA contains information but is unable to carry out actions Fact 2: Proteins are the workhorses but contain no information THUS Information in DNA must
Chapter 16: Gene Expression from Biology by OpenStax College is licensed under a Creative Commons Attribution 3.0 Unported license.
 Chapter 16: Gene Expression from Biology by OpenStax College is licensed under a Creative Commons Attribution 3.0 Unported license. 2013, Rice University. CHAPTER 16 GENE EXPRESSION 429 16 GENE EXPRESSION
Chapter 16: Gene Expression from Biology by OpenStax College is licensed under a Creative Commons Attribution 3.0 Unported license. 2013, Rice University. CHAPTER 16 GENE EXPRESSION 429 16 GENE EXPRESSION
DNA. Is a molecule that encodes the genetic instructions used in the development and functioning of all known living organisms and many viruses.
 Is a molecule that encodes the genetic instructions used in the development and functioning of all known living organisms and many viruses. Genetic information is encoded as a sequence of nucleotides (guanine,
Is a molecule that encodes the genetic instructions used in the development and functioning of all known living organisms and many viruses. Genetic information is encoded as a sequence of nucleotides (guanine,
Chromatographic Separation of the three forms of RNA Polymerase II.
 Chromatographic Separation of the three forms of RNA Polymerase II. α-amanitin α-amanitin bound to Pol II Function of the three enzymes. Yeast Pol II. RNA Polymerase Subunit Structures 10-7 Subunit structure.
Chromatographic Separation of the three forms of RNA Polymerase II. α-amanitin α-amanitin bound to Pol II Function of the three enzymes. Yeast Pol II. RNA Polymerase Subunit Structures 10-7 Subunit structure.
DNA makes RNA makes Proteins. The Central Dogma
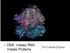 DNA makes RNA makes Proteins The Central Dogma TRANSCRIPTION DNA RNA transcript RNA polymerase RNA PROCESSING Exon RNA transcript (pre-mrna) Intron Aminoacyl-tRNA synthetase NUCLEUS CYTOPLASM FORMATION
DNA makes RNA makes Proteins The Central Dogma TRANSCRIPTION DNA RNA transcript RNA polymerase RNA PROCESSING Exon RNA transcript (pre-mrna) Intron Aminoacyl-tRNA synthetase NUCLEUS CYTOPLASM FORMATION
Scientific Method. Name: NetID: Exam 1 Version 1 September 12, 2017 Dr. A. Pimentel
 Name: NetID: Exam 1 Version 1 September 12, 2017 Dr. A. Pimentel Each question has a value of 4 points and there is a total of 156 points in the exam. However, the maximum score of this exam will be capped
Name: NetID: Exam 1 Version 1 September 12, 2017 Dr. A. Pimentel Each question has a value of 4 points and there is a total of 156 points in the exam. However, the maximum score of this exam will be capped
Biochemistry Eukaryotic Transcription
 1 Description of Module Subject Name Paper Name Module Name/Title Dr. Vijaya Khader Dr. MC Varadaraj 2 1. Objectives 1. Understand and have an overview of eucaryotic transcriptional regulation. 2. Explain
1 Description of Module Subject Name Paper Name Module Name/Title Dr. Vijaya Khader Dr. MC Varadaraj 2 1. Objectives 1. Understand and have an overview of eucaryotic transcriptional regulation. 2. Explain
