HBV RNA pre-genome encodes specific motifs that mediate interactions with the viral core protein that promote nucleocapsid assembly
|
|
|
- Stuart Sanders
- 6 years ago
- Views:
Transcription
1 In the format provided by the authors and unedited. SUPPLEMENTARY INFORMATION Nikesh Patel *, Simon J. White *, Rebecca F Thompson, Richard Bingham 1, Eva U. Weiß 1, Daniel P. Maskell, Adam Zlotnick 2, Eric Dykeman 1, Roman Tuma, Reidun Twarock 1², Neil A. Ranson ² & Peter G. Stockley ². Astbury Centre for Structural Molecular Biology, University of Leeds, Leeds, LS2 9JT, UK. VOLUME: 2 ARTICLE NUMBER: HBV RNA pre-genome encodes specific motifs that mediate interactions with the viral core protein that promote nucleocapsid assembly 1 Departments of Biology and Mathematics & York Centre for Complex Systems Analysis, University of York, York, YO10 5DD, UK 2 Department of Molecular & Cellular Biochemistry, Indiana University, Bloomington, IN 47405, USA. *These authors contributed equally to this work. ² Joint communicating authors Supplementary Figures NATURE MICROBIOLOGY DOI: /nmicrobiol Macmillan Publishers Limited, part of Springer Nature. All rights reserved.
2 Supplementary Figure 1. Characterising HBV VLPs from E.coli and SELEX protocol. (a) Hydrodynamic radial distribution and negative stain EM image of Alexa Fluor-488 labelled HBV VLPs purified from E.coli. Integration of peak yields suggests a roughly 2:1 ratio of T=4 (red circle, 63%) and T=3 (blue circle, 37%) VLPs. Scale bar represents 100 nm. (b) SELEX protocol showing selection for aptamers with high affinity to HBV 185 Cp. HBV VLPs were immobilised onto carboxylic magnetic beads (brown circles) and dissociated into Cp dimers (green and grey rectangles) using guanidinium chloride. An RNA pool encompassing a random region (40N) was enriched for sequences with affinity for Cp by repeated cycles of binding to these beads, partitioning and amplification. Negative selections at each round used carboxylic acid beads which had been treated with NHS-EDC and inactivated with Tris. Stringency was increased after 2
3 round 5 by decreasing the number of positive beads by half and increasing the number of washes from 8 to 10. The reverse transcriptase-pcr products at the end of each round were analyzed by native PAGE to confirm the isolation of products for the next round of selection. The 10 th round products were converted to DNA and sequenced. Supplementary Figure 2. PS oligo structures, example smfcs trace and EMs of PS containing VLPs. (a) PS1-3 and ε secondary structures, made using VARNA software 1, were predicted in Mfold. Preferred sites were taken from the HBV genome, NC_ , at positions: PS1 ( ), PS2 ( ) and PS3 ( ). In order to make them all the same length (47 nucleotides) to avoid effects of charge differences the following additions were made; PS1, 5ʹ - GGGUUUUGG and CCC - 3ʹ; PS2, 5ʹ - GGGUUUUGGGG and CCCC - 3ʹ; 3
4 PS3, 5ʹ- GGGUUUUGG and CCCC - 3ʹ. The consensus motif RGAG is highlighted in red in each of the loops. The stability of each RNA fold, as predicted by Mfold, is shown below each structure. (b) Example smfcs assay. R h values for, fluorescently labelled RNAs are determined before and after Cp is titrated in at fixed time points (vertical dashed lines), allowing the R h values to equilibrate after each step. The faint red trace represents real time, raw signal while the thick red line represents smoothed data. PS1 R h initially climbs slowly, until a threshold Cp concentration, which triggers rapid assembly into a T=3 or T=4 VLP (R h ~24-32 nm, orange dashed lines) as determined by measurements of Alexa-Flour 488 labelled HBV particles from E.coli (Supplementary Figure 1a). At the end of each titration, the complexes formed are challenged by addition of RNase A. An unchanged R h is assumed to mean that the test RNA has been encapsidated in a closed VLP. The time scale on which this occurs is indicated in the bottom right. (c) TEMs from assembly reactions of PS1, 2, 3 and Cp alone and unlabelled PS1 in Figure 3. Large white particles in Cp alone and Unlabelled PS1 TEMs are latex beads. Also present is TEM from empty particle assembly described in Sup Table 2. These empty HBV particles were assembled at much higher concentrations of Cp (1.5 µm) and in the absence of RNA. Scale bars represent 100 nm. 4
5 Supplementary Figure 3. smfcs assays of PS1 variants (a) smfcs assays of the PS1 variants (structures top left) and accompanying hydrodynamic radial distributions plotted in 2 nm bins and fitted with Gaussian peaks below, as colour coded in the key. 15 nm PS1 (black), PS1 loop mutant (dark orange) bulge mutant (dark blue) and epsilon (green) RNAs were tested for their ability to form VLPs under single molecule conditions. Vertical dotted lines indicate points of addition of Cp with the final concentrations shown in nm. Samples were allowed to equilibrate between additions. RNase A was added to check for correctly formed particles. Samples were taken prior to RNase A addition for analysis by TEM shown right, both here and throughout this figure. (b) - as (a) with RNA oligos PS1 (dashed black), L1 (green), L2 (red) and L3 (blue). (c) - as (a) with RNA oligos PS1 (dashed black), L4 (magenta), L5 (cyan) and B1 (light green). (d) as (a) with RNA oligos PS1 (dashed black) and DNA oligo PS1. Scale bars represent 100 nm. PS1 controls (dashed black) in each 5
6 panel were repeated for individual batches of purified Cp, accounting for the variations in assembly efficiency seen. smfcs and TEM were repeated in triplicate. 6
7 7
8 Supplementary Figure 4. Role(s) of ARD and its charge on assembly. (a) smfcs assays of 15 nm PS1 with Cp (grey) and Cp 149 (cyan) and accompanying hydrodynamic radial distributions plotted in 2 nm bins and fitted with Gaussian peaks below. EM images of particles are shown (right). (b) as (a) with PS1 and Cp (black), kinase and Cp pre equilibrated and added to PS1 (cyan) and PS1 and Cp with kinase added simultaneously (magenta). TEMs are shown right. Scale bars represent 100 nm. PS1 controls (black) in each panel were repeated for individual batches of purified Cp, accounting for the variations in assembly efficiency seen. smfcs and TEM were repeated in triplicate. Supplementary Figure 5. ARD structure in T=4 and T=3 VLPs. Slabs (~30 Å thick) through the structures of the icosahedrally-averaged T=4 particle at 4.7 Å (left), the same T=4 structure low pass filtered to 7 Å (middle), and the T=3 particle at 5.6 Å (right). A Cp dimer is fitted into each. Even at a slightly lower resolution than the T=3 VLP, there is no equivalent density for the ARD in the T=4 VLP, confirming that it has different conformations in each particle. Supplementary Tables Expected mass (Da) Observed mass (Da) SRPK ± 1.37 Phosphorylated Cp ± 0.71 Cp ± 0.86 Cp ± 0.06 Supplementary Table 1: Masses of the different forms of Cp and kinase (SRPK ) used, as determined by ESI-MS mass spectrometry 8
9 Fluorescence Polarisation Total Fluorescence Sample -RNase +RNase -RNase +RNase PS1 oligo PS1 VLP PS1 + empty VLP Supplementary Table 2: Association of Alexa-Fluor-488 labelled PS1 with Cp. Anisotropy was used to determine if 15 nm of Alexa-Fluor-488 labelled RNA PS oligos can bind to, or enter, 125 nm of preformed shells of Cp. The latter were formed by reassembly in the absence of RNA at high concentration 2 (Supplementary Fig 2c). Fluorescence polarisation values are influenced by the mass of the dye-labelled species 3. The polarisation value for PS1 oligo goes down following addition of RNase, as expected but remains unchanged when incorporated in VLPs assembled in the presence of the oligo. When labelled PS1 is added to the empty Cp VLP its fluorescence emission is unaffected, suggesting that it is not quenched, and it remains RNase sensitive confirming that it does not bind the outside of the protein shell or get internalised. RNA Oligo Loop Bulge PS1 GGGAGG GGG +++ Assembly behaviour Comment L1 UUUAUU GGG Loop G s are important L2 GUUAGG GGG Loop G s are important L3 UGGAUU GGG Loop G s are important L4 GGGUGG GGG Loop A is important L5 GGGGGG GGG Loop A is important B1 GGGAGG AAC Bulge sequence/structure is important Supplementary Table 3: Sequence changes and corresponding assembly behaviour of PS1 variant oligonucleotides, L1-5 and B1. Assembly behaviour is indicated as follows, the first + indicates RNA-Cp binding, the second signifies formation of T=3/ T=4 sized species, and the third indicates RNase protection. - indicates failure in that assay. Supplementary References 1. Darty, K., Denise, A. & Ponty, Y. VARNA: Interactive drawing and editing of the RNA secondary structure. Bioinformatics 25, (2009). 2. Porterfield, J. Z. et al. Full-length hepatitis B virus core protein packages viral and heterologous RNA with similarly high levels of cooperativity. J. Virol. 84, (2010). 3. Lakowicz, J. R. Principles of Fluorescence Spectroscopy. Ch. 1. page 12. (Springer US, 2006). 9
Masayoshi Honda, Jeehae Park, Robert A. Pugh, Taekjip Ha, and Maria Spies
 Molecular Cell, Volume 35 Supplemental Data Single-Molecule Analysis Reveals Differential Effect of ssdna-binding Proteins on DNA Translocation by XPD Helicase Masayoshi Honda, Jeehae Park, Robert A. Pugh,
Molecular Cell, Volume 35 Supplemental Data Single-Molecule Analysis Reveals Differential Effect of ssdna-binding Proteins on DNA Translocation by XPD Helicase Masayoshi Honda, Jeehae Park, Robert A. Pugh,
TECHNICAL NOTE Phosphorylation Site Specific Aptamers for Cancer Biomarker Peptide Enrichment and Detection in Mass Spectrometry Based Proteomics
 TECHNICAL NOTE Phosphorylation Site Specific Aptamers for Cancer Biomarker Peptide Enrichment and Detection in Mass Spectrometry Based Proteomics INTRODUCTION The development of renewable capture reagents
TECHNICAL NOTE Phosphorylation Site Specific Aptamers for Cancer Biomarker Peptide Enrichment and Detection in Mass Spectrometry Based Proteomics INTRODUCTION The development of renewable capture reagents
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION Conserved arginines on the rim of Hfq catalyze base pair formation and exchange Subrata Panja and Sarah A. Woodson T.C. Jenkins Department of Biophysics, Johns Hopkins University,
SUPPLEMENTARY INFORMATION Conserved arginines on the rim of Hfq catalyze base pair formation and exchange Subrata Panja and Sarah A. Woodson T.C. Jenkins Department of Biophysics, Johns Hopkins University,
Supporting Information
 Supporting Information Wiley-VCH 2006 69451 Weinheim, Germany RNA ligands that distinguish metabolite-induced conformations in the TPP riboswitch Günter Mayer, Marie-Sophie L. Raddatz, Julia D. Grunwald,
Supporting Information Wiley-VCH 2006 69451 Weinheim, Germany RNA ligands that distinguish metabolite-induced conformations in the TPP riboswitch Günter Mayer, Marie-Sophie L. Raddatz, Julia D. Grunwald,
Stabilization of a virus-like particle and its application as a nanoreactor at physiological conditions
 Supporting Information Stabilization of a virus-like particle and its application as a nanoreactor at physiological conditions Lise Schoonen, b Sjors Maassen, b Roeland J. M. Nolte b and Jan C. M. van
Supporting Information Stabilization of a virus-like particle and its application as a nanoreactor at physiological conditions Lise Schoonen, b Sjors Maassen, b Roeland J. M. Nolte b and Jan C. M. van
COMPAS for the Analysis of SELEX Experiments
 COMPAS for the Analysis of SELEX Experiments COMPAS (COMmon PAtternS) is a software tool that was especially developed to harness the technology of next generation sequencing (NGS) to bring light into
COMPAS for the Analysis of SELEX Experiments COMPAS (COMmon PAtternS) is a software tool that was especially developed to harness the technology of next generation sequencing (NGS) to bring light into
qpcr Quantitative PCR or Real-time PCR Gives a measurement of PCR product at end of each cycle real time
 qpcr qpcr Quantitative PCR or Real-time PCR Gives a measurement of PCR product at end of each cycle real time Differs from endpoint PCR gel on last cycle Used to determines relative amount of template
qpcr qpcr Quantitative PCR or Real-time PCR Gives a measurement of PCR product at end of each cycle real time Differs from endpoint PCR gel on last cycle Used to determines relative amount of template
A nucleic acid-based fluorescent sensor for expeditious detection of pyrophosphate anions at nanomolar concentrations
 Supporting Information for A nucleic acid-based fluorescent sensor for expeditious detection of pyrophosphate anions at nanomolar concentrations Xin Su, Chen Zhang, Xianjin Xiao, Anqin Xu, Zhendong Xu
Supporting Information for A nucleic acid-based fluorescent sensor for expeditious detection of pyrophosphate anions at nanomolar concentrations Xin Su, Chen Zhang, Xianjin Xiao, Anqin Xu, Zhendong Xu
Functional Genomics Research Stream. Research Meeting: June 19, 2012 SYBR Green qpcr, Research Update
 Functional Genomics Research Stream Research Meeting: June 19, 2012 SYBR Green qpcr, Research Update Updates Alternate Lab Meeting Fridays 11:30-1:00 WEL 4.224 Welcome to attend either one Lab Log thanks
Functional Genomics Research Stream Research Meeting: June 19, 2012 SYBR Green qpcr, Research Update Updates Alternate Lab Meeting Fridays 11:30-1:00 WEL 4.224 Welcome to attend either one Lab Log thanks
SUPPLEMENTARY MATERIAL AND METHODS
 SUPPLEMENTARY MATERIAL AND METHODS Amplification of HEV ORF1, ORF2 and ORF3 genome regions Total RNA was extracted from 200 µl EDTA plasma using Cobas AmpliPrep total nucleic acid isolation kit (Roche,
SUPPLEMENTARY MATERIAL AND METHODS Amplification of HEV ORF1, ORF2 and ORF3 genome regions Total RNA was extracted from 200 µl EDTA plasma using Cobas AmpliPrep total nucleic acid isolation kit (Roche,
American Society of Cytopathology Core Curriculum in Molecular Biology
 American Society of Cytopathology Core Curriculum in Molecular Biology American Society of Cytopathology Core Curriculum in Molecular Biology Chapter 3 Molecular Techniques Separation and Detection, Part
American Society of Cytopathology Core Curriculum in Molecular Biology American Society of Cytopathology Core Curriculum in Molecular Biology Chapter 3 Molecular Techniques Separation and Detection, Part
Technical Review. Real time PCR
 Technical Review Real time PCR Normal PCR: Analyze with agarose gel Normal PCR vs Real time PCR Real-time PCR, also known as quantitative PCR (qpcr) or kinetic PCR Key feature: Used to amplify and simultaneously
Technical Review Real time PCR Normal PCR: Analyze with agarose gel Normal PCR vs Real time PCR Real-time PCR, also known as quantitative PCR (qpcr) or kinetic PCR Key feature: Used to amplify and simultaneously
Nature Structural & Molecular Biology: doi: /nsmb.2548
 Supplementary Figure 1. Structure of GltPhout. (a) Stereo view of a slice through a single GltPhout protomer shown in stick representation along with 2Fo-Fc and anomalous difference electron maps. The
Supplementary Figure 1. Structure of GltPhout. (a) Stereo view of a slice through a single GltPhout protomer shown in stick representation along with 2Fo-Fc and anomalous difference electron maps. The
Roche Molecular Biochemicals Technical Note No. LC 9/2000
 Roche Molecular Biochemicals Technical Note No. LC 9/2000 LightCycler Optimization Strategy Introduction Purpose of this Note Table of Contents The LightCycler system provides different detection formats
Roche Molecular Biochemicals Technical Note No. LC 9/2000 LightCycler Optimization Strategy Introduction Purpose of this Note Table of Contents The LightCycler system provides different detection formats
mmu-mir-200a-3p Real-time RT-PCR Detection and U6 Calibration Kit User Manual MyBioSource.com Catalog # MBS826230
 mmu-mir-200a-3p Real-time RT-PCR Detection and U6 Calibration Kit User Manual Catalog # MBS826230 For the detection and quantification of mirnas mmu-mir-200a-3p normalized by U6 snrna using Real-time RT-PCR
mmu-mir-200a-3p Real-time RT-PCR Detection and U6 Calibration Kit User Manual Catalog # MBS826230 For the detection and quantification of mirnas mmu-mir-200a-3p normalized by U6 snrna using Real-time RT-PCR
SELECTION OF MOLECULAR APTAMERS FOR
 SELECTION OF MOLECULAR APTAMERS FOR IDENTIFICATION OF LIVE CELLS OF RALSTONIA SOLANACEARUM: A NEW POTENTIAL METHOD IN PLANT PATHOLOGY Patrice Champoiseau, University of Florida 2009 APS ANNUAL MEETING
SELECTION OF MOLECULAR APTAMERS FOR IDENTIFICATION OF LIVE CELLS OF RALSTONIA SOLANACEARUM: A NEW POTENTIAL METHOD IN PLANT PATHOLOGY Patrice Champoiseau, University of Florida 2009 APS ANNUAL MEETING
SUPPLEMENTARY MATERIAL
 SUPPLEMENTARY MATERIAL Materials and Methods Circular dichroism (CD) spectroscopy. Far ultraviolet (UV) CD spectra of apo- and holo- CaM and the CaM mutants were recorded on a Jasco J-715 spectropolarimeter
SUPPLEMENTARY MATERIAL Materials and Methods Circular dichroism (CD) spectroscopy. Far ultraviolet (UV) CD spectra of apo- and holo- CaM and the CaM mutants were recorded on a Jasco J-715 spectropolarimeter
Supplementary Fig. S1. SAMHD1c has a more potent dntpase activity than. SAMHD1c. Purified recombinant SAMHD1c and SAMHD1c proteins (with
 Supplementary Fig. S1. SAMHD1c has a more potent dntpase activity than SAMHD1c. Purified recombinant SAMHD1c and SAMHD1c proteins (with concentration of 800nM) were incubated with 1mM dgtp for the indicated
Supplementary Fig. S1. SAMHD1c has a more potent dntpase activity than SAMHD1c. Purified recombinant SAMHD1c and SAMHD1c proteins (with concentration of 800nM) were incubated with 1mM dgtp for the indicated
Supplementary Figures
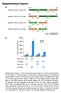 Supplementary Figures a HindIII XbaI pcdna3.1/v5-his A_Nfluc-VVD 1-398 37-186 Nfluc Linker 1 VVD HindIII XhoI pcdna3.1/v5-his A_VVD-Nfluc 37-186 1-398 VVD Linker 2 Nfluc HindIII XbaI pcdna3.1/v5-his A_Cfluc-VVD
Supplementary Figures a HindIII XbaI pcdna3.1/v5-his A_Nfluc-VVD 1-398 37-186 Nfluc Linker 1 VVD HindIII XhoI pcdna3.1/v5-his A_VVD-Nfluc 37-186 1-398 VVD Linker 2 Nfluc HindIII XbaI pcdna3.1/v5-his A_Cfluc-VVD
Polymerase Chain Reaction: Application and Practical Primer Probe Design qrt-pcr
 Polymerase Chain Reaction: Application and Practical Primer Probe Design qrt-pcr review Enzyme based DNA amplification Thermal Polymerarase derived from a thermophylic bacterium DNA dependant DNA polymerase
Polymerase Chain Reaction: Application and Practical Primer Probe Design qrt-pcr review Enzyme based DNA amplification Thermal Polymerarase derived from a thermophylic bacterium DNA dependant DNA polymerase
QuantiFluor ONE dsdna System
 TECHNICAL MANUAL QuantiFluor ONE dsdna System Instructions for Use of Products E4871 and E4870 Revised 6/17 TM405 QuantiFluor ONE dsdna System All technical literature is available at: www.promega.com/protocols/
TECHNICAL MANUAL QuantiFluor ONE dsdna System Instructions for Use of Products E4871 and E4870 Revised 6/17 TM405 QuantiFluor ONE dsdna System All technical literature is available at: www.promega.com/protocols/
Characterization of Aptamer Binding using SensíQ SPR Platforms
 Characterization of Aptamer Binding using SensíQ SPR Platforms APPLICATION NOTE INTRODUCTION Aptamers have the potential to provide a better solution in diagnostics and other research areas than traditional
Characterization of Aptamer Binding using SensíQ SPR Platforms APPLICATION NOTE INTRODUCTION Aptamers have the potential to provide a better solution in diagnostics and other research areas than traditional
Plant Molecular and Cellular Biology Lecture 9: Nuclear Genome Organization: Chromosome Structure, Chromatin, DNA Packaging, Mitosis Gary Peter
 Plant Molecular and Cellular Biology Lecture 9: Nuclear Genome Organization: Chromosome Structure, Chromatin, DNA Packaging, Mitosis Gary Peter 9/16/2008 1 Learning Objectives 1. List and explain how DNA
Plant Molecular and Cellular Biology Lecture 9: Nuclear Genome Organization: Chromosome Structure, Chromatin, DNA Packaging, Mitosis Gary Peter 9/16/2008 1 Learning Objectives 1. List and explain how DNA
Supplementary information for. An Ultrasensitive Biosensor for DNA Detection Based on. Hybridization Chain Reaction Coupled with the Efficient
 Supplementary information for An Ultrasensitive Biosensor for DNA Detection Based on Hybridization Chain Reaction Coupled with the Efficient Quenching of Ruthenium Complex to CdTe Quantum Dot Yufei Liu,
Supplementary information for An Ultrasensitive Biosensor for DNA Detection Based on Hybridization Chain Reaction Coupled with the Efficient Quenching of Ruthenium Complex to CdTe Quantum Dot Yufei Liu,
Introduction. Technical Note
 DNA and RNA quantification: fast and simple with PicoGreen dsdna and RiboGreen RNA quantification reagents Fluorescence intensity on Infinite F2 and Infinite M2 Introduction DNA quantification Detection
DNA and RNA quantification: fast and simple with PicoGreen dsdna and RiboGreen RNA quantification reagents Fluorescence intensity on Infinite F2 and Infinite M2 Introduction DNA quantification Detection
Quantitative Real Time PCR USING SYBR GREEN
 Quantitative Real Time PCR USING SYBR GREEN SYBR Green SYBR Green is a cyanine dye that binds to double stranded DNA. When it is bound to D.S. DNA it has a much greater fluorescence than when bound to
Quantitative Real Time PCR USING SYBR GREEN SYBR Green SYBR Green is a cyanine dye that binds to double stranded DNA. When it is bound to D.S. DNA it has a much greater fluorescence than when bound to
mmu-mir-34a Real-time RT-PCR Detection Kit User Manual
 mmu-mir-34a Real-time RT-PCR Detection Kit User Manual Catalog # CPK1272 For the detection and quantification of mirna mmu-mir-34a using Real-Time RT-PCR detection instruments. For research use only. Not
mmu-mir-34a Real-time RT-PCR Detection Kit User Manual Catalog # CPK1272 For the detection and quantification of mirna mmu-mir-34a using Real-Time RT-PCR detection instruments. For research use only. Not
Current ( pa) Current (pa) Voltage (mv) Voltage ( mv)
 Current ( pa) 3000 2000 1000 0-1000 -2000-3000 a -400-200 0 200 400 Voltage (mv) P1 P2 P3 P4 P5 P6 P7 P8 P9 P10 P11 P12 P13 P14 P15 P16 P17 P18 Average Current (pa) 4500 3000 1500 0-1500 -3000-4500 b -400-200
Current ( pa) 3000 2000 1000 0-1000 -2000-3000 a -400-200 0 200 400 Voltage (mv) P1 P2 P3 P4 P5 P6 P7 P8 P9 P10 P11 P12 P13 P14 P15 P16 P17 P18 Average Current (pa) 4500 3000 1500 0-1500 -3000-4500 b -400-200
DNA Hybridization and Detection
 Chapter 6 DNA Hybridization and Detection Fluorescence Polarization Detection of DNA Hybridization........................................................ 6-2 Introduction.............................................................................................................
Chapter 6 DNA Hybridization and Detection Fluorescence Polarization Detection of DNA Hybridization........................................................ 6-2 Introduction.............................................................................................................
Alizarin Red S for Online Pyrophosphate Detection Identified by a Rapid Screening Method
 Supporting Information Alizarin Red S for Online Pyrophosphate Detection Identified by a Rapid Screening Method Jens Fischbach, Qiuting Loh, Frank F. Bier, Theam Soon Lim, Marcus Frohme, Jörn Glökler Table
Supporting Information Alizarin Red S for Online Pyrophosphate Detection Identified by a Rapid Screening Method Jens Fischbach, Qiuting Loh, Frank F. Bier, Theam Soon Lim, Marcus Frohme, Jörn Glökler Table
microrna-122 stimulates translation of Hepatitis C Virus RNA
 Supplementary information for: microrna-122 stimulates translation of Hepatitis C Virus RNA Jura Inga Henke 1,5, Dagmar Goergen 1,5, Junfeng Zheng 3, Yutong Song 4, Christian G. Schüttler 2, Carmen Fehr
Supplementary information for: microrna-122 stimulates translation of Hepatitis C Virus RNA Jura Inga Henke 1,5, Dagmar Goergen 1,5, Junfeng Zheng 3, Yutong Song 4, Christian G. Schüttler 2, Carmen Fehr
Product Specifications & Manual
 Product Specifications & Manual Custom Oligo Synthesis, antisense oligos, RNA oligos, chimeric oligos, Fluorescent dye labeled oligos, Molecular Beacons, sirna, phosphonates Affinity Ligands, 2-5 linked
Product Specifications & Manual Custom Oligo Synthesis, antisense oligos, RNA oligos, chimeric oligos, Fluorescent dye labeled oligos, Molecular Beacons, sirna, phosphonates Affinity Ligands, 2-5 linked
Detecting Rare Genomic Copies or Events
 Detecting Rare Genomic Copies or Events Integrated DNA Technologies Nick Downey, PhD Common Diagnostics Challenges Detecting targets early with accuracy Detecting different strains (e.g., antibiotic resistance)
Detecting Rare Genomic Copies or Events Integrated DNA Technologies Nick Downey, PhD Common Diagnostics Challenges Detecting targets early with accuracy Detecting different strains (e.g., antibiotic resistance)
Supplementary Figure 1. Pressure dependence of the self-cleavage reaction of the modified hairpin ribozyme. Time evolution of the modified hairpin
 Supplementary Figure 1. Pressure dependence of the self-cleavage reaction of the modified hairpin ribozyme. Time evolution of the modified hairpin ribozyme s fraction cleaved for the reaction in the presence
Supplementary Figure 1. Pressure dependence of the self-cleavage reaction of the modified hairpin ribozyme. Time evolution of the modified hairpin ribozyme s fraction cleaved for the reaction in the presence
Module1TheBasicsofRealTimePCR Monday, March 19, 2007
 Objectives Slide notes: Page 1 of 41 Module 1: The Basics Of Real Time PCR Slide notes: Module 1: The Basics of real time PCR Page 2 of 41 Polymerase Chain Reaction Slide notes: Here is a review of PCR,
Objectives Slide notes: Page 1 of 41 Module 1: The Basics Of Real Time PCR Slide notes: Module 1: The Basics of real time PCR Page 2 of 41 Polymerase Chain Reaction Slide notes: Here is a review of PCR,
Improving the Limit of Detection of Lateral Flow Assays using 3DNA Technology
 Improving the Limit of Detection of Lateral Flow Assays using 3DNA Technology Introduction Lateral Flow (LF) and similar assays represent a unique growing class of Point of Care (POC) tests designed to
Improving the Limit of Detection of Lateral Flow Assays using 3DNA Technology Introduction Lateral Flow (LF) and similar assays represent a unique growing class of Point of Care (POC) tests designed to
INTRODUCTION TO REVERSE TRANSCRIPTION PCR (RT-PCR) ABCF 2016 BecA-ILRI Hub, Nairobi 21 st September 2016 Roger Pelle Principal Scientist
 INTRODUCTION TO REVERSE TRANSCRIPTION PCR (RT-PCR) ABCF 2016 BecA-ILRI Hub, Nairobi 21 st September 2016 Roger Pelle Principal Scientist Objective of PCR To provide a solution to one of the most pressing
INTRODUCTION TO REVERSE TRANSCRIPTION PCR (RT-PCR) ABCF 2016 BecA-ILRI Hub, Nairobi 21 st September 2016 Roger Pelle Principal Scientist Objective of PCR To provide a solution to one of the most pressing
SOP: SYBR Green-based real-time RT-PCR
 SOP: SYBR Green-based real-time RT-PCR By Richard Yu Research fellow Centre for Marine Environmental Research and Innovative Technology (MERIT) Department of Biology and Chemistry City University of Hong
SOP: SYBR Green-based real-time RT-PCR By Richard Yu Research fellow Centre for Marine Environmental Research and Innovative Technology (MERIT) Department of Biology and Chemistry City University of Hong
Reverse transcription-pcr (rt-pcr) Dr. Hani Alhadrami
 Reverse transcription-pcr (rt-pcr) Dr. Hani Alhadrami hanialhadrami@kau.edu.sa www.hanialhadrami.kau.edu.sa Overview Several techniques are available to detect and analyse RNA. Examples of these techniques
Reverse transcription-pcr (rt-pcr) Dr. Hani Alhadrami hanialhadrami@kau.edu.sa www.hanialhadrami.kau.edu.sa Overview Several techniques are available to detect and analyse RNA. Examples of these techniques
Quantitation of mrna Using Real-Time Reverse Transcription PCR (RT-PCR)
 Quantitation of mrna Using Real-Time Reverse Transcription PCR (RT-PCR) Quantitative Real-Time RT-PCR Versus RT-PCR In Real-Time RT- PCR, DNA amplification monitored at each cycle but RT-PCR measures the
Quantitation of mrna Using Real-Time Reverse Transcription PCR (RT-PCR) Quantitative Real-Time RT-PCR Versus RT-PCR In Real-Time RT- PCR, DNA amplification monitored at each cycle but RT-PCR measures the
Protein Stability Analysis Using the Optim Patrick Celie NKI Protein Facility, Amsterdam
 Protein Stability Analysis Using the Optim 1000 Patrick Celie NKI Protein Facility, Amsterdam NKI Protein Facility Fundamental and translation cancer research ~ 650 scientists + supporting personnel Connected
Protein Stability Analysis Using the Optim 1000 Patrick Celie NKI Protein Facility, Amsterdam NKI Protein Facility Fundamental and translation cancer research ~ 650 scientists + supporting personnel Connected
Confocal Microscopy Analyzes Cells
 Choosing Filters for Fluorescence A Laurin Publication Photonic Solutions for Biotechnology and Medicine November 2002 Confocal Microscopy Analyzes Cells Reprinted from the November 2002 issue of Biophotonics
Choosing Filters for Fluorescence A Laurin Publication Photonic Solutions for Biotechnology and Medicine November 2002 Confocal Microscopy Analyzes Cells Reprinted from the November 2002 issue of Biophotonics
Systematic Evolution of Ligands by Exponential Enrichment SELEX
 Systematic Evolution of Ligands by Exponential Enrichment SELEX SELEX Systematic evolution of ligands by exponential amplification * Isolation of new RNs with novel properties * New therapeutics, New diagnostics
Systematic Evolution of Ligands by Exponential Enrichment SELEX SELEX Systematic evolution of ligands by exponential amplification * Isolation of new RNs with novel properties * New therapeutics, New diagnostics
Molecular Cell Biology - Problem Drill 11: Recombinant DNA
 Molecular Cell Biology - Problem Drill 11: Recombinant DNA Question No. 1 of 10 1. Which of the following statements about the sources of DNA used for molecular cloning is correct? Question #1 (A) cdna
Molecular Cell Biology - Problem Drill 11: Recombinant DNA Question No. 1 of 10 1. Which of the following statements about the sources of DNA used for molecular cloning is correct? Question #1 (A) cdna
X-ray structures of fructosyl peptide oxidases revealing residues responsible for gating oxygen access in the oxidative half reaction
 X-ray structures of fructosyl peptide oxidases revealing residues responsible for gating oxygen access in the oxidative half reaction Tomohisa Shimasaki 1, Hiromi Yoshida 2, Shigehiro Kamitori 2 & Koji
X-ray structures of fructosyl peptide oxidases revealing residues responsible for gating oxygen access in the oxidative half reaction Tomohisa Shimasaki 1, Hiromi Yoshida 2, Shigehiro Kamitori 2 & Koji
Roche Molecular Biochemicals Technical Note No. LC 10/2000
 Roche Molecular Biochemicals Technical Note No. LC 10/2000 LightCycler Overview of LightCycler Quantification Methods 1. General Introduction Introduction Content Definitions This Technical Note will introduce
Roche Molecular Biochemicals Technical Note No. LC 10/2000 LightCycler Overview of LightCycler Quantification Methods 1. General Introduction Introduction Content Definitions This Technical Note will introduce
Mechanism of Transcription Termination by RNA Polymerase III Utilizes a Non-template Strand Sequence-Specific Signal Element
 Molecular Cell Supplemental Information Mechanism of ranscription ermination by RNA Polymerase III Utilizes a Non-template Strand Sequence-Specific Signal Element Aneeshkumar G. Arimbasseri and Richard
Molecular Cell Supplemental Information Mechanism of ranscription ermination by RNA Polymerase III Utilizes a Non-template Strand Sequence-Specific Signal Element Aneeshkumar G. Arimbasseri and Richard
Diagnosis Sanger. Interpreting and Troubleshooting Chromatograms. Volume 1: Help! No Data! GENEWIZ Technical Support
 Diagnosis Sanger Interpreting and Troubleshooting Chromatograms GENEWIZ Technical Support DNAseq@genewiz.com Troubleshooting This troubleshooting guide is based on common issues seen from samples within
Diagnosis Sanger Interpreting and Troubleshooting Chromatograms GENEWIZ Technical Support DNAseq@genewiz.com Troubleshooting This troubleshooting guide is based on common issues seen from samples within
Nature Methods: doi: /nmeth Supplementary Figure 1. Retention of RNA with LabelX.
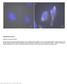 Supplementary Figure 1 Retention of RNA with LabelX. (a) Epi-fluorescence image of single molecule FISH (smfish) against GAPDH on HeLa cells expanded without LabelX treatment. (b) Epi-fluorescence image
Supplementary Figure 1 Retention of RNA with LabelX. (a) Epi-fluorescence image of single molecule FISH (smfish) against GAPDH on HeLa cells expanded without LabelX treatment. (b) Epi-fluorescence image
Empty virus-like particles (evlps) for plant synthetic biology
 Empty virus-like particles (evlps) for plant synthetic biology George Lomonossoff Dept. of Biological Chemistry John Innes Centre, Norwich, U.K. Introduction to Opportunities in Plant Synthetic Biology
Empty virus-like particles (evlps) for plant synthetic biology George Lomonossoff Dept. of Biological Chemistry John Innes Centre, Norwich, U.K. Introduction to Opportunities in Plant Synthetic Biology
Supplementary Figure 1: Two modes of low concentration of BsSMC on a DNA (a) Protein staining (left) and fluorescent imaging of Cy3 (right) confirm
 Supplementary Figure 1: Two modes of low concentration of BsSMC on a DNA (a) Protein staining (left) and fluorescent imaging of Cy3 (right) confirm that BsSMC was labeled with Cy3 NHS-Ester. In each panel,
Supplementary Figure 1: Two modes of low concentration of BsSMC on a DNA (a) Protein staining (left) and fluorescent imaging of Cy3 (right) confirm that BsSMC was labeled with Cy3 NHS-Ester. In each panel,
The practical task of monoclonal IgG purification with CHT TM ceramic hydroxyapatite
 The practical task of monoclonal IgG purification with CHT TM ceramic hydroxyapatite Pete Gagnon, Jie He, Paul Ng, Julia Zhen, Cheryl Aberin, Heather Mekosh 11th Annual Waterside Conference, Chicago, May
The practical task of monoclonal IgG purification with CHT TM ceramic hydroxyapatite Pete Gagnon, Jie He, Paul Ng, Julia Zhen, Cheryl Aberin, Heather Mekosh 11th Annual Waterside Conference, Chicago, May
MagSi Beads. Magnetic Silica Beads for Life Science and Biotechnology study
 MagSi Beads Magnetic Silica Beads for Life Science and Biotechnology study MagnaMedics Diagnostics B.V. / Rev. 9.2 / 2012 Wide range of products for numerous applications MagnaMedics separation solutions
MagSi Beads Magnetic Silica Beads for Life Science and Biotechnology study MagnaMedics Diagnostics B.V. / Rev. 9.2 / 2012 Wide range of products for numerous applications MagnaMedics separation solutions
Introduction To Real-Time Quantitative PCR (qpcr)
 Introduction To Real-Time Quantitative PCR (qpcr) Samuel Rulli, Ph.D. Samuel.Rulli@QIAGEN.com Technical Support: BRCsupport@qiagen.com The products described in this webinar are intended for molecular
Introduction To Real-Time Quantitative PCR (qpcr) Samuel Rulli, Ph.D. Samuel.Rulli@QIAGEN.com Technical Support: BRCsupport@qiagen.com The products described in this webinar are intended for molecular
Molecular characterization, detection & quantitation of biological products Purin Charoensuksai, PhD
 Molecular characterization, detection & quantitation of biological products Purin Charoensuksai, PhD Department of Biopharmacy, Faculty of Pharmacy, Silpakorn University Example of critical checkpoints
Molecular characterization, detection & quantitation of biological products Purin Charoensuksai, PhD Department of Biopharmacy, Faculty of Pharmacy, Silpakorn University Example of critical checkpoints
Supplementary Figures. Figure S1. Design and operation of the microfluidic cartridge.
 Supplementary Figures Figure S1. Design and operation of the microfluidic cartridge. (a) The device has three inlets for DNA and capture beads, MNP probes and washing buffer. Each inlet is gated by the
Supplementary Figures Figure S1. Design and operation of the microfluidic cartridge. (a) The device has three inlets for DNA and capture beads, MNP probes and washing buffer. Each inlet is gated by the
Light Sensitization of DNA Nanostructures via Incorporation of Photo-Cleavable Spacers
 Electronic Supplementary Material (ESI) for Chemical Communications. This journal is The Royal Society of Chemistry 2016 Light Sensitization of DNA Nanostructures via Incorporation of Photo-Cleavable Spacers
Electronic Supplementary Material (ESI) for Chemical Communications. This journal is The Royal Society of Chemistry 2016 Light Sensitization of DNA Nanostructures via Incorporation of Photo-Cleavable Spacers
Principals of Real-Time PCR. Amira A. T. AL-Hosary Lecturer of Infectious Diseases, Faculty of Veterinary Medicine, Assiut University, Egypt
 Principals of Real-Time PCR Amira A. T. AL-Hosary Lecturer of Infectious Diseases, Faculty of Veterinary Medicine, Assiut University, Egypt What Is Real-Time PCR? Nucleic acid (DNA) amplification and detection
Principals of Real-Time PCR Amira A. T. AL-Hosary Lecturer of Infectious Diseases, Faculty of Veterinary Medicine, Assiut University, Egypt What Is Real-Time PCR? Nucleic acid (DNA) amplification and detection
Supplemental Information. Natural RNA Polymerase Aptamers. Regulate Transcription in E. coli
 Molecular Cell, Volume 67 Supplemental Information Natural RNA Polymerase Aptamers Regulate Transcription in E. coli Nadezda Sedlyarova, Philipp Rescheneder, Andrés Magán, Niko Popitsch, Natascha Rziha,
Molecular Cell, Volume 67 Supplemental Information Natural RNA Polymerase Aptamers Regulate Transcription in E. coli Nadezda Sedlyarova, Philipp Rescheneder, Andrés Magán, Niko Popitsch, Natascha Rziha,
Introduction to Real-Time PCR: Basic Principles and Chemistries
 Introduction to Real-Time PCR: Basic Principles and Chemistries Leta Steffen, PhD Applications Scientist Promega Corporation Outline I. Real-Time PCR overview Basics of Real-Time PCR Understanding the
Introduction to Real-Time PCR: Basic Principles and Chemistries Leta Steffen, PhD Applications Scientist Promega Corporation Outline I. Real-Time PCR overview Basics of Real-Time PCR Understanding the
Gene Expression Technology
 Gene Expression Technology Bing Zhang Department of Biomedical Informatics Vanderbilt University bing.zhang@vanderbilt.edu Gene expression Gene expression is the process by which information from a gene
Gene Expression Technology Bing Zhang Department of Biomedical Informatics Vanderbilt University bing.zhang@vanderbilt.edu Gene expression Gene expression is the process by which information from a gene
Electronic Supplementary Information Sensitive detection of polynucleotide kinase using rolling circle amplification-induced chemiluminescence
 Electronic Supplementary Material (ESI) for Chemical Communications. This journal is The Royal Society of Chemistry 2014 Electronic Supplementary Information Sensitive detection of polynucleotide kinase
Electronic Supplementary Material (ESI) for Chemical Communications. This journal is The Royal Society of Chemistry 2014 Electronic Supplementary Information Sensitive detection of polynucleotide kinase
Hepatitis B Virus genesig Easy Kit for use on the genesig q16
 TM Primerdesign Ltd genesig Easy Kit for use on the genesig q16 50 reaction For general laboratory and research use only 1 genesig Easy: at a glance guide For each DNA test Component Volume Lab-in-a-box
TM Primerdesign Ltd genesig Easy Kit for use on the genesig q16 50 reaction For general laboratory and research use only 1 genesig Easy: at a glance guide For each DNA test Component Volume Lab-in-a-box
Amino-allyl Dye Coupling Protocol
 Amino-allyl Dye Coupling Protocol Joseph DeRisi, June 2001 Typically, fluorescently labeled cdna is generated by incorporation of dyeconjugated nucleotide analogs during the reverse transcription process.
Amino-allyl Dye Coupling Protocol Joseph DeRisi, June 2001 Typically, fluorescently labeled cdna is generated by incorporation of dyeconjugated nucleotide analogs during the reverse transcription process.
Supplementary Table 1. The Q-PCR primer sequence is summarized in the following table.
 Supplementary Table 1. The Q-PCR primer sequence is summarized in the following table. Name Sequence (5-3 ) Application Flag-u ggactacaaggacgacgatgac Shared upstream primer for all the amplifications of
Supplementary Table 1. The Q-PCR primer sequence is summarized in the following table. Name Sequence (5-3 ) Application Flag-u ggactacaaggacgacgatgac Shared upstream primer for all the amplifications of
Viral RNAi suppressor reversibly binds sirna to. outcompete Dicer and RISC via multiple-turnover
 Supplementary Data Viral RNAi suppressor reversibly binds sirna to outcompete Dicer and RISC via multiple-turnover Renata A. Rawlings 1,2, Vishalakshi Krishnan 2 and Nils G. Walter 2 * 1 Biophysics and
Supplementary Data Viral RNAi suppressor reversibly binds sirna to outcompete Dicer and RISC via multiple-turnover Renata A. Rawlings 1,2, Vishalakshi Krishnan 2 and Nils G. Walter 2 * 1 Biophysics and
Figure S1: NUN preparation yields nascent, unadenylated RNA with a different profile from Total RNA.
 Summary of Supplemental Information Figure S1: NUN preparation yields nascent, unadenylated RNA with a different profile from Total RNA. Figure S2: rrna removal procedure is effective for clearing out
Summary of Supplemental Information Figure S1: NUN preparation yields nascent, unadenylated RNA with a different profile from Total RNA. Figure S2: rrna removal procedure is effective for clearing out
Chapter 17. PCR the polymerase chain reaction and its many uses. Prepared by Woojoo Choi
 Chapter 17. PCR the polymerase chain reaction and its many uses Prepared by Woojoo Choi Polymerase chain reaction 1) Polymerase chain reaction (PCR): artificial amplification of a DNA sequence by repeated
Chapter 17. PCR the polymerase chain reaction and its many uses Prepared by Woojoo Choi Polymerase chain reaction 1) Polymerase chain reaction (PCR): artificial amplification of a DNA sequence by repeated
Methods of Biomaterials Testing Lesson 3-5. Biochemical Methods - Molecular Biology -
 Methods of Biomaterials Testing Lesson 3-5 Biochemical Methods - Molecular Biology - Chromosomes in the Cell Nucleus DNA in the Chromosome Deoxyribonucleic Acid (DNA) DNA has double-helix structure The
Methods of Biomaterials Testing Lesson 3-5 Biochemical Methods - Molecular Biology - Chromosomes in the Cell Nucleus DNA in the Chromosome Deoxyribonucleic Acid (DNA) DNA has double-helix structure The
Recent technology allow production of microarrays composed of 70-mers (essentially a hybrid of the two techniques)
 Microarrays and Transcript Profiling Gene expression patterns are traditionally studied using Northern blots (DNA-RNA hybridization assays). This approach involves separation of total or polya + RNA on
Microarrays and Transcript Profiling Gene expression patterns are traditionally studied using Northern blots (DNA-RNA hybridization assays). This approach involves separation of total or polya + RNA on
Fluorescence Microscopy. Terms and concepts to know: 10/11/2011. Visible spectrum (of light) and energy
 Fluorescence Microscopy Louisiana Tech University Ruston, Louisiana Microscopy Workshop Dr. Mark DeCoster Associate Professor Biomedical Engineering 1 Terms and concepts to know: Signal to Noise Excitation
Fluorescence Microscopy Louisiana Tech University Ruston, Louisiana Microscopy Workshop Dr. Mark DeCoster Associate Professor Biomedical Engineering 1 Terms and concepts to know: Signal to Noise Excitation
CHAPTER 9 DNA Technologies
 CHAPTER 9 DNA Technologies Recombinant DNA Artificially created DNA that combines sequences that do not occur together in the nature Basis of much of the modern molecular biology Molecular cloning of genes
CHAPTER 9 DNA Technologies Recombinant DNA Artificially created DNA that combines sequences that do not occur together in the nature Basis of much of the modern molecular biology Molecular cloning of genes
QIAcube RNA isolation from stool samples using the RNeasy PowerMicrobiome Kit
 Application Note QIAcube RNA isolation from stool samples using the RNeasy PowerMicrobiome Kit Patrick Smith, Eva Haenssler, Eddie Adams, and Dominic O Neil QIAGEN GmbH, QIAGEN Strasse 1, 4724 Hilden,
Application Note QIAcube RNA isolation from stool samples using the RNeasy PowerMicrobiome Kit Patrick Smith, Eva Haenssler, Eddie Adams, and Dominic O Neil QIAGEN GmbH, QIAGEN Strasse 1, 4724 Hilden,
American Society of Cytopathology Core Curriculum in Molecular Biology
 Brought American Society of Cytopathology Core Curriculum in Molecular Biology Brought American Society of Cytopathology Core Curriculum in Molecular Biology Chapter 3 Molecular Techniques Alternatives
Brought American Society of Cytopathology Core Curriculum in Molecular Biology Brought American Society of Cytopathology Core Curriculum in Molecular Biology Chapter 3 Molecular Techniques Alternatives
Agilent s Mx3000P and Mx3005P
 Agilent s Mx3000P and Mx3005P Realtime PCR just got better Dr. Ivan Bendezu Genomics Agent Andalucia Real-time PCR Chemistries SYBR Green SYBR Green: Dye attaches to the minor groove of double-stranded
Agilent s Mx3000P and Mx3005P Realtime PCR just got better Dr. Ivan Bendezu Genomics Agent Andalucia Real-time PCR Chemistries SYBR Green SYBR Green: Dye attaches to the minor groove of double-stranded
Non-Organic-Based Isolation of Mammalian microrna using Norgen s microrna Purification Kit
 Application Note 13 RNA Sample Preparation Non-Organic-Based Isolation of Mammalian microrna using Norgen s microrna Purification Kit B. Lam, PhD 1, P. Roberts, MSc 1 Y. Haj-Ahmad, M.Sc., Ph.D 1,2 1 Norgen
Application Note 13 RNA Sample Preparation Non-Organic-Based Isolation of Mammalian microrna using Norgen s microrna Purification Kit B. Lam, PhD 1, P. Roberts, MSc 1 Y. Haj-Ahmad, M.Sc., Ph.D 1,2 1 Norgen
Quantitative analysis of recombination in YFP and CFP gene of FRET biosensor induced by lentiviral or retroviral gene transfer.
 Supplementary Information: Quantitative analysis of recombination in and gene of FRET biosensor induced by lentiviral or retroviral gene transfer. Akira T. Komatsubara, Michiyuki Matsuda,, and Kazuhiro
Supplementary Information: Quantitative analysis of recombination in and gene of FRET biosensor induced by lentiviral or retroviral gene transfer. Akira T. Komatsubara, Michiyuki Matsuda,, and Kazuhiro
Microarrays: since we use probes we obviously must know the sequences we are looking at!
 These background are needed: 1. - Basic Molecular Biology & Genetics DNA replication Transcription Post-transcriptional RNA processing Translation Post-translational protein modification Gene expression
These background are needed: 1. - Basic Molecular Biology & Genetics DNA replication Transcription Post-transcriptional RNA processing Translation Post-translational protein modification Gene expression
RNA Clean-Up and Concentration Kit Product # 23600, 43200
 3430 Schmon Parkway Thorold, ON, Canada L2V 4Y6 Phone: 866-667-4362 (905) 227-8848 Fax: (905) 227-1061 Email: techsupport@norgenbiotek.com RNA Clean-Up and Concentration Kit Product # 23600, 43200 Product
3430 Schmon Parkway Thorold, ON, Canada L2V 4Y6 Phone: 866-667-4362 (905) 227-8848 Fax: (905) 227-1061 Email: techsupport@norgenbiotek.com RNA Clean-Up and Concentration Kit Product # 23600, 43200 Product
MMG 301, Lec. 25 Mutations and Bacteriophage
 MMG 301, Lec. 25 Mutations and Bacteriophage Questions for today: 1. What are mutations and how do they form? 2. How are mutant bacteria used in research? 3. What are the general properties of bacteriophage
MMG 301, Lec. 25 Mutations and Bacteriophage Questions for today: 1. What are mutations and how do they form? 2. How are mutant bacteria used in research? 3. What are the general properties of bacteriophage
Boost Your Real-Time Results
 Boost Your Real-Time Results Refresh your knowledge of the fundamentals of real-time PCR Sample & Assay Technologies 1 Introduction Real-time PCR and RT-PCR are highly sensitive techniques that enable
Boost Your Real-Time Results Refresh your knowledge of the fundamentals of real-time PCR Sample & Assay Technologies 1 Introduction Real-time PCR and RT-PCR are highly sensitive techniques that enable
Supplementary Figure 1 Two distinct conformational states of the HNH domain in crystal structures. a
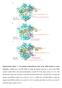 Supplementary Figure 1 Two distinct conformational states of the HNH domain in crystal structures. a HNH-state 1 in PDB 4OO8, in which the distance from the C atom of the HNH catalytic residue 840 to the
Supplementary Figure 1 Two distinct conformational states of the HNH domain in crystal structures. a HNH-state 1 in PDB 4OO8, in which the distance from the C atom of the HNH catalytic residue 840 to the
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Origin use and efficiency are similar among WT, rrm3, pif1-m2, and pif1-m2; rrm3 strains. A. Analysis of fork progression around confirmed and likely origins (from cerevisiae.oridb.org).
Supplementary Figure 1 Origin use and efficiency are similar among WT, rrm3, pif1-m2, and pif1-m2; rrm3 strains. A. Analysis of fork progression around confirmed and likely origins (from cerevisiae.oridb.org).
over time using live cell microscopy. The time post infection is indicated in the lower left corner.
 Title of file for HTML: Supplementary Information Description: Supplementary Figures and Supplementary Table Title of file for HTML: Supplementary Movie 1 Description: Fusion of NBs. BSR cells were infected
Title of file for HTML: Supplementary Information Description: Supplementary Figures and Supplementary Table Title of file for HTML: Supplementary Movie 1 Description: Fusion of NBs. BSR cells were infected
NAME TA SEC Problem Set 4 FRIDAY October 15, Answers to this problem set must be inserted into the box outside
 MIT Biology Department 7.012: Introductory Biology - Fall 2004 Instructors: Professor Eric Lander, Professor Robert A. Weinberg, Dr. Claudette Gardel NAME TA SEC 7.012 Problem Set 4 FRIDAY October 15,
MIT Biology Department 7.012: Introductory Biology - Fall 2004 Instructors: Professor Eric Lander, Professor Robert A. Weinberg, Dr. Claudette Gardel NAME TA SEC 7.012 Problem Set 4 FRIDAY October 15,
Quiz Submissions Quiz 4
 Quiz Submissions Quiz 4 Attempt 1 Written: Nov 1, 2015 17:35 Nov 1, 2015 22:19 Submission View Released: Nov 4, 2015 20:24 Question 1 0 / 1 point Three RNA polymerases synthesize most of the RNA present
Quiz Submissions Quiz 4 Attempt 1 Written: Nov 1, 2015 17:35 Nov 1, 2015 22:19 Submission View Released: Nov 4, 2015 20:24 Question 1 0 / 1 point Three RNA polymerases synthesize most of the RNA present
One Step SYBR PrimeScript RT-PCR Kit II (Perfect Real Time)
 Cat. # RR086A For Research Use One Step SYBR PrimeScript RT-PCR Kit II Product Manual Table of Contents I. Description...3 II. III. IV. Principle...3 Components...5 Storage...6 V. Features...6 VI. VII.
Cat. # RR086A For Research Use One Step SYBR PrimeScript RT-PCR Kit II Product Manual Table of Contents I. Description...3 II. III. IV. Principle...3 Components...5 Storage...6 V. Features...6 VI. VII.
Detection and quantification of hepatitis E virus by real-time reverse transcriptase PCR
 Page 1 of 6 EU FP VII PROJECT VITAL STANDARD OPERATING Detection and quantification of hepatitis E virus by real-time reverse transcriptase PCR CREATED: REVISED: APPROVED: David Rodríguez Lázaro: 18-2-2010
Page 1 of 6 EU FP VII PROJECT VITAL STANDARD OPERATING Detection and quantification of hepatitis E virus by real-time reverse transcriptase PCR CREATED: REVISED: APPROVED: David Rodríguez Lázaro: 18-2-2010
Contents... vii. List of Figures... xii. List of Tables... xiv. Abbreviatons... xv. Summary... xvii. 1. Introduction In vitro evolution...
 vii Contents Contents... vii List of Figures... xii List of Tables... xiv Abbreviatons... xv Summary... xvii 1. Introduction...1 1.1 In vitro evolution... 1 1.2 Phage Display Technology... 3 1.3 Cell surface
vii Contents Contents... vii List of Figures... xii List of Tables... xiv Abbreviatons... xv Summary... xvii 1. Introduction...1 1.1 In vitro evolution... 1 1.2 Phage Display Technology... 3 1.3 Cell surface
Flow Cytometry - The Essentials
 Flow Cytometry - The Essentials Pocket Guide to Flow Cytometry: 1. Know your Cytometer 2. Understanding Fluorescence and Fluorophores 3. Gating Process 4. Controls 5. Optimization 6. Panel Building 7.
Flow Cytometry - The Essentials Pocket Guide to Flow Cytometry: 1. Know your Cytometer 2. Understanding Fluorescence and Fluorophores 3. Gating Process 4. Controls 5. Optimization 6. Panel Building 7.
Different types of PCR and principles of Real Time PCR. Prof. Dr. Hamdy M. El-Aref Assiut University, Faculty of Agriculture Genetics Department
 Different types of PC and principles of eal Time PC. Prof. Dr. Hamdy M. El-Aref Assiut University, Faculty of Agriculture Genetics Department I N T O D U C T I O N PC Cycle (round) I N T O D U C T I O
Different types of PC and principles of eal Time PC. Prof. Dr. Hamdy M. El-Aref Assiut University, Faculty of Agriculture Genetics Department I N T O D U C T I O N PC Cycle (round) I N T O D U C T I O
Sensitivity vs Specificity
 Viral Detection Animal Inoculation Culturing the Virus Definitive Length of time Serology Detecting antibodies to the infectious agent Detecting Viral Proteins Western Blot ELISA Detecting the Viral Genome
Viral Detection Animal Inoculation Culturing the Virus Definitive Length of time Serology Detecting antibodies to the infectious agent Detecting Viral Proteins Western Blot ELISA Detecting the Viral Genome
Hepatitis B Virus. Core Protein Region. 150 tests. Quantification of Hepatitis B Virus genomes. Advanced kit handbook HB
 Techne qpcr test Hepatitis B Virus Core Protein Region 150 tests For general laboratory and research use only 1 Introduction to Hepatitis B Virus Originally known as serum hepatitis, hepatitis B has only
Techne qpcr test Hepatitis B Virus Core Protein Region 150 tests For general laboratory and research use only 1 Introduction to Hepatitis B Virus Originally known as serum hepatitis, hepatitis B has only
SideStep Lysis and Stabilization Buffer
 SideStep Lysis and Stabilization Buffer INSTRUCTION MANUAL Catalog #400900 Revision B.0 For Research Use Only. Not for use in diagnostic procedures. 400900-12 LIMITED PRODUCT WARRANTY This warranty limits
SideStep Lysis and Stabilization Buffer INSTRUCTION MANUAL Catalog #400900 Revision B.0 For Research Use Only. Not for use in diagnostic procedures. 400900-12 LIMITED PRODUCT WARRANTY This warranty limits
TECH NOTE SMARTer T-cell receptor profiling in single cells
 TECH NOTE SMARTer T-cell receptor profiling in single cells Flexible workflow: Illumina-ready libraries from FACS or manually sorted single cells Ease of use: Optimized indexing allows for pooling 96 cells
TECH NOTE SMARTer T-cell receptor profiling in single cells Flexible workflow: Illumina-ready libraries from FACS or manually sorted single cells Ease of use: Optimized indexing allows for pooling 96 cells
Module I: Introduction
 Module I: Introduction 20.109 Lecture 1 3 February, 2011 Introduction to: Module Overview Fundamental concepts and techniques in molecular biology Appreciating nucleic acids (RNA in particular) as more
Module I: Introduction 20.109 Lecture 1 3 February, 2011 Introduction to: Module Overview Fundamental concepts and techniques in molecular biology Appreciating nucleic acids (RNA in particular) as more
Performance characteristics of the High Sensitivity DNA kit for the Agilent 2100 Bioanalyzer
 Performance characteristics of the High Sensitivity DNA kit for the Agilent 2100 Bioanalyzer Technical Note 10 Measured conc. [ng/µl] 1 Y intercept = 0.09 r 2 = 0.993 0.1 0.1 1 10 Reference concentration
Performance characteristics of the High Sensitivity DNA kit for the Agilent 2100 Bioanalyzer Technical Note 10 Measured conc. [ng/µl] 1 Y intercept = 0.09 r 2 = 0.993 0.1 0.1 1 10 Reference concentration
Application Note Detecting low copy numbers. Introduction. Methods A08-005B
 Application Note Detecting low copy numbers A08-005B Introduction Sensitivity of a qpcr assay is highly dependent on primer efficiency. Not all assays will be capable of detecting a single copy of template
Application Note Detecting low copy numbers A08-005B Introduction Sensitivity of a qpcr assay is highly dependent on primer efficiency. Not all assays will be capable of detecting a single copy of template
Kit Specifications 45 g. 45 g of RNA 8 L
 RNA Clean-Up and Concentration Micro-Elute Kit Product # 61000 Product Insert Norgen s RNA Clean-Up and Concentration Micro-Elution Kit provides a rapid method for the purification, cleanup and concentration
RNA Clean-Up and Concentration Micro-Elute Kit Product # 61000 Product Insert Norgen s RNA Clean-Up and Concentration Micro-Elution Kit provides a rapid method for the purification, cleanup and concentration
Recombinant DNA Technology
 Recombinant DNA Technology Common General Cloning Strategy Target DNA from donor organism extracted, cut with restriction endonuclease and ligated into a cloning vector cut with compatible restriction
Recombinant DNA Technology Common General Cloning Strategy Target DNA from donor organism extracted, cut with restriction endonuclease and ligated into a cloning vector cut with compatible restriction
