Quantitative analysis of recombination in YFP and CFP gene of FRET biosensor induced by lentiviral or retroviral gene transfer.
|
|
|
- Olivia Phillips
- 6 years ago
- Views:
Transcription
1 Supplementary Information: Quantitative analysis of recombination in and gene of FRET biosensor induced by lentiviral or retroviral gene transfer. Akira T. Komatsubara, Michiyuki Matsuda,, and Kazuhiro Aoki 3,* Laboratory of Bioimaging and Cell Signaling, Graduate School of Biostudies, Kyoto University, Sakyo-ku, Kyoto , Japan Department of Pathology and Biology of Diseases, Graduate School of Medicine, Kyoto University, Sakyo-ku, Kyoto , Japan 3 Imaging Platform for Spatio-Temporal Information, Graduate School of Medicine, Kyoto University, Sakyo-ku, Kyoto , Japan
2 Supplementary Figure S FIGURE S Multiple alignment of nucleotide sequences in and genes. DNA sequences of e0ypet, h0ypet, nturquoise-gl, and TFP were aligned to highlight their sequence identity with the boxed area.
3 Supplementary Figure S FIGURE S Comparison of fluorescence intensities among chimeric GFP variants. (A) An alignment of amino acid sequences in YPet and nturquoise-gl. (B) Fluorescence intensities of h0ypet, chimeric GFP variants and were measured in HeLa cells transiently expressing the indicated biosensors (Left). Fluorescence intensities of channel were normalized by fluorescence intensities of channel. The averaged / ratios are represented with S.E. (N > 0 cells).
4 (a.u) (a.u) (a.u) (a.u) (a.u) (a.u) (a.u) (a.u) (a.u) (a.u) (a.u) (a.u) (a.u) (a.u) (a.u) (a.u) Supplementary Figure S3 Gene delivery: HIV-based lentivirus, Cell: HeLa A B C D h0 h7-e h0-e0 h-e (a.u.) 0 00 (a.u.) 0 00 (a.u.) 0 00 (a.u.) E F G H e0 e7-h e0-h0 e-h (a.u.) 0 00 (a.u.) 0 (a.u.) 0 00 (a.u.) Gene delivery: MuLV-based retrovirus, Cell: HeLa I J K L h0 h7-e h0-e0 h-e (a.u.) (a.u.) (a.u.) (a.u.) M N O P e0 e7-h e0-h0 e-h (a.u.) (a.u.) (a.u.) (a.u.) FIGURE S3 Recombination between and genes by lentiviral or retroviral gene transfer in HeLa cells. HeLa cells were infected with lentivirus (A-H) or retrovirus (I-P) encoding 8 different FRET biosensors as shown in the upper panel. At least 4 days after infection, the cells were imaged with an epi-fluorescence microscope. The average fluorescence intensities of and are represented as a log-log plot. Each dot corresponds to a HeLa cell. Three hundred cells were analyzed from two independent experiments. Red lines are the fitted line with the e0ypet data. Orange and cyan arrowheads indicate the T03Y and Y66W mutations, respectively.
5 (a.u.) (a.u.) (a.u.) (a.u.) (a.u.) (a.u.) (a.u.) (a.u.) (a.u.) (a.u.) (a.u.) (a.u.) (a.u.) (a.u.) (a.u.) (a.u.) Supplementary Figure S4 Gene delivery: Lipofection with lentivirus plasmid, Cell: HeLa A B C D h0 h7-e h0-e0 h-e (a.u.) 0 00 (a.u.) 0 00 (a.u.) 0 00 (a.u.) E F G H e0 e7-h e0-h0 e-h (a.u.) 0 00 (a.u.) 0 00 (a.u.) 0 00 (a.u.) Gene delivery: Lipofection with retrovirus plasmid, Cell: HeLa I J K L h0 h7-e h0-e0 h-e (a.u.) (a.u.) (a.u.) (a.u.) M N O P e0 e7-h e0-h0 e-h (a.u.) (a.u.) (a.u.) (a.u.) FIGURE S4 Transient expression of and in HeLa cells with lipofection. HeLa cells were transfected with lentiviral (A-H) or retroviral (I-P) vector plasmids encoding 8 different FRET biosensors as shown in the upper panel. At least days after transfection, the cells were imaged with an epi-fluorescence microscope. The average fluorescence intensities of and are represented as a log-log plot. Each dot corresponds to a HeLa cell. Three hundred cells were analyzed from two independent experiments. Red lines are the fitted line with the e0ypet data. Orange and cyan arrowheads indicate the T03Y and Y66W mutation, respectively.
6 Counts Supplementary Figure S A h0ypet ATGGTGAGCAAGGGCGAGGAGCTGTTCACCGGGGTGGTGCCCATCCTGGTCGAGCTGGACGGCGACGTAAACGGCCACAAGTTCAGCGTGTCCGGCGAGG 0 Sequence Data ATGGTGAGCAAGGGCGAGGAGCTGTTCACCGGGGTGGTGCCCATCCTGGTCGAGCTGGACGGCGACGTAAACGGCCACAAGTTCAGCGTGTCCGGCGAGG 0 nturquoise-gl ATGGTGAGCAAGGGCGAGGAGCTGTTCACCGGGGTGGTGCCCATCCTGGTCGAGCTGGACGGCGACGTAAACGGCCACAAGTTCAGCGTGTCCGGCGAGG 0 h0ypet GCGAGGGCGATGCCACCTACGGCAAGCTGACCCTGAAGCTTCTATGCACCACCGGCAAGCTGCCCGTGCCCTGGCCCACCCTCGTGACCACCCTGGGCTA 00 Sequence Data GCGAGGGCGATGCCACCTACGGCAAGCTGACCCTGAAGCTTCTATGCACCACCGGCAAGCTGCCCGTGCCCTGGCCCACCCTCGTGACCACCCTGGGCTA 00 nturquoise-gl GCGAGGGCGATGCCACCTACGGCAAGCTGACCCTGAAGTTCATCTGCACCACCGGCAAGCTGCCCGTGCCCTGGCCCACCCTCGTGACCACCCTGTCCTG 00 h0ypet 0 CGGCCTGCAGTGCTTCGCCCGCTACCCCGACCACATGAAGCAGCACGACTTCTTCAAGTCCGCCATGCCCGAAGGCTACGTCCAGGAGCGCACCATCTTC 300 Sequence Data 0 CGGCCTGCAGTGCTTCGCCCGCTACCCCGACCACATGAAGCAGCACGACTTCTTCAAGTCCGCCATGCCCGAAGGCTACGTCCAGGAGCGCACCATCTTC 300 nturquoise-gl 0 GGGCGTGCAGTGCTTCGCCCGCTACCCCGACCACATGAAGCAGCACGACTTCTTCAAGTCCGCCATGCCCGAAGGCTACGTCCAGGAGCGCACCATCTTC 300 h0ypet 30 TTCAAGGACGACGGCAACTACAAGACCCGCGCCGAGGTGAAGTTCGAGGGCGACACCCTGGTGAACCGCATCGAGCTGAAGGGCATCGACTTCAAGGAGG 400 Sequence Data 30 TTCAAGGACGACGGCAACTACAAGACCCGCGCCGAGGTGAAGTTCGAGGGCGACACCCTGGTGAACCGCATCGAGCTGAAGGGCATCGACTTCAAGGAGG 400 nturquoise-gl 30 TTCAAGGACGACGGCAACTACAAGACCCGCGCCGAGGTGAAGTTCGAGGGCGACACCCTGGTGAACCGCATCGAGCTGAAGGGCATCGACTTCAAGGAGG 400 h0ypet 40 ACGGCAACATCCTGGGGCACAAGCTGGAGTACAACTACAACAGCCACAACGTCTATATCACCGCCGACAAGCAGAAGAACGGCATCAAGGCCAACTTCAA 00 Sequence Data 40 ACGGCAACATCCTGGGGCACAAGCTGGAGTACAACTACAACAGCCACAACGTCTATATCACCGCCGACAAGCAGAAGAACGGCATCAAGGCCAACTTCAA 00 nturquoise-gl 40 ACGGCAACATCCTGGGGCACAAGCTGGAGTACAACTACATCAGCGGGAACGTCTATATCACCGCCGACAAGCAGAAGAACGGCATCAAGGCCAACTTCAA 00 h0ypet 0 GATCCGCCACAACATCGAGGACGGCGGCGTGCAGCTCGCCGACCACTACCAGCAGAACACCCCCATCGGCGACGGCCCCGTGCTGCTGCCCGACAACCAC 600 Sequence Data 0 GATCCGCCACAACATCGAGGACGGCGGCGTGCAGCTCGCCGACCACTACCAGCAGAACACCCCCATCGGCGACGGCCCCGTGCTGCTGCCCGACAACCAC 600 nturquoise-gl 0 GATCCGCCACAACATCGAGGACGGCGGCGTGCAGCTCGCCGACCACTACCAGCAGAACACCCCCATCGGCGACGGCCCCGTGCTGCTGCCCGACAACCAC 600 h0ypet 60 TACCTGAGCTACCAGTCCGCCCTGTTCAAAGACCCCAACGAGAAGCGCGATCACATGGTCCTGCTGGAGTTCCTGACCGCCGCCGGGATCACTGAGGGCA 700 Sequence Data 60 TACCTGAGCACCCAGTCCGCCTTAAGCAAAGACCCCAACGAGAAGCGCGATCACATGGTCCTGCTGGAGTTCTTGACCGCCGCCGGGATCACTCTCGGCA 700 nturquoise-gl 60 TACCTGAGCACCCAGTCCGCCTTAAGCAAAGACCCCAACGAGAAGCGCGATCACATGGTCCTGCTGGAGTTCTTGACCGCCGCCGGGATCACTCTCGGCA 700 h0ypet 70 TGAACGAGCTGTAC 74 Sequence Data 70 TGGACGAGCTG 7 nturquoise-gl 70 TGGACGAGCTG 7 B YPet nturquoise-gl aa aa Length (bp) Counts F47L V68L F46L T6G Y66W (99-0 bp) N46I H48G T03Y (6-6 bp) H3F S08F D34N 38 aa 37 aa C 0 y = 0.039x R² = Length (bp) FIGURE S Verification of the recombination. (A) An example of results of DNA sequencing in recombined YPet and nturquoise-gl genes. A49 cells infected with h0ypet-carrying lentivirus were sorted by FACS depending on fluorescence, followed by genomic DNA extraction, PCR amplification of the recombined genes, and sequencing. Nucleotide sequences of wild-type h0ypet, a representative recombined DNA sequence, and wild-type nturquoise-gl are aligned with black lines, which enclose identical nucleotides. The red line encloses the region where the recombination has
7 occurred. *, synonymous mutation. (B) Schematic representation of the difference between YPet and nturquoise-gl amino acids. (Upper) The dotted lines represent different amino acid residues between YPet and nturquoise-gl. (Lower) We analyzed chimeric GFP and variants, in which Y66 amino acid was conserved. Red regions indicate potential recombination site. Length indicates the number of identical DNA sequence for each potential recombination sites. Counts indicate the number of recombined GFP or variants that have been generated as a result of recombination in between the identical DNA sequence. (C) The number of recombined GFP or variants are plotted as a function of the number of each identical DNA sequence, in which the recombination has occurred.
8 Supplementary Figure S6 r : recombination rate (/bp) (, ) = (, ) (, ) = (0, ) (, ) = (0.08, 0) (, ) = (0., 0) (, ) = (0.9, 0) (, ) = (, 0) Y66W L68V N46T (, ) = (0.6, 0) H48G T03Y 00 0 (0, 0.08 ~) (, 0) 0 00 (, ) = (, ) 00 0 ave SD Noise 0 00 Slope ave ave SD ave SD FIGURE S6 Scheme of the mathematical model of recombination between and genes. (Upper panel) Schematic representation of the recombination of and genes. Red lines indicate the critical amino acid residues for the substitution of and from GFP, Y66W and T03Y, respectively. Black arrowheads indicate the position of recombination. Of note, fluorescence intensity of chimeric GFP genes is changed in accord with the recombination sites (Supplementary Fig. SB). (Lower panels) The distribution of and intensities are illustrated in a log-log plot. The blue, orange, and green areas represent cells expressing,, or both, respectively (left). To recapitulate the experimental data, the indicated 9 parameters were extracted from the experimental data set (right).
9 Supplementary Figure S7 Computer simulation, Gene delivery: HIV-based retrovirus, Cell: A49 A h0 B h7-e C h0-e0 D h-e7,000,000,000,000,000,000,000, ,000 0,000 0,000 0,000 E e0 F e7-h G e0-h0 H e-h7,000,000,000,000,000,000,000, ,000 0,000 0,000 0,000 Computer simulation, Gene delivery: HIV-based lentivirus, Cell: HeLa I h0 J h7-e K h0-e0 L h-e7,000,000,000,000,000,000,000, ,000 0,000 0,000 0,000 M e0 N e7-h O e0-h0 P e-h7,000,000,000,000,000,000,000, ,000 0,000 0,000 0,000 Computer simulation, Gene delivery: MuLV-based retrovirus, Cell: HeLa Q h0 R h7-e S h0-e0 T h-e7,000,000,000,000,000,000,000, ,000 0,000 0,000 0,000 U e0 V e7-h W e0-h0 X e-h7,000,000,000,000,000,000,000, ,000 0,000 0,000 0,000
10 FIGURE S7 Computer simulation of the recombination between and genes. The recombination of and genes in A49 cells infected with the indicated retrovirus (A-H) or HeLa cells infected with the lentivirus (I-P) or retrovirus (Q-X) was simulated by computer with the recombination rates, which showed maximal likelihood estimation (Table ). To reproduce the experimental data, 9 parameters were extracted from the experimental data set in Figure 3, and Supplementary Figure S3 (see Supplementary Figure S6 and the Methods for details). Red lines are the fitted line with the e0ypet data. Orange and cyan arrowheads indicate the T03Y and Y66W mutations, respectively.
11 Log likelihood Log likelihood Log likelihood Log likelihood Supplementary Figure S8 A Gene delivery: HIV-based lentivirus Cell: A49 B Gene delivery: MuLV-based retrovirus 4 Cell: A Recombination rate, r (/bp) Recombination rate, r (/bp) C -0.9 Gene delivery: HIV-based lentivirus 4 Cell: HeLa D Gene delivery: MuLV-based retrovirus - 4 Cell: HeLa Recombination rate, r (/bp) Recombination rate, r (/bp) FIGURE S8 Maximal log-likelihood estimation of recombination rate. Log-likelihood values were calculated as described in Methods, and plotted as a function of recombination rate (r) in A49 cells infected with lentivirus (A) or retrovirus (B), or in HeLa cells infected with lentivirus (C) or retrovirus (D). The red circles indicate maximal log-likelihood values.
Supplementary Figure 1. Isolation of GFPHigh cells.
 Supplementary Figure 1. Isolation of GFP High cells. (A) Schematic diagram of cell isolation based on Wnt signaling activity. Colorectal cancer (CRC) cell lines were stably transduced with lentivirus encoding
Supplementary Figure 1. Isolation of GFP High cells. (A) Schematic diagram of cell isolation based on Wnt signaling activity. Colorectal cancer (CRC) cell lines were stably transduced with lentivirus encoding
Genome Sequence Assembly
 Genome Sequence Assembly Learning Goals: Introduce the field of bioinformatics Familiarize the student with performing sequence alignments Understand the assembly process in genome sequencing Introduction:
Genome Sequence Assembly Learning Goals: Introduce the field of bioinformatics Familiarize the student with performing sequence alignments Understand the assembly process in genome sequencing Introduction:
Multiplex Fluorescence Assays for Adherence Cells without Trypsinization
 Multiplex Fluorescence Assays for Adherence Cells without Trypsinization The combination of a bright field and three fluorescent channels allows the Celigo to perform many multiplexed assays. A gating
Multiplex Fluorescence Assays for Adherence Cells without Trypsinization The combination of a bright field and three fluorescent channels allows the Celigo to perform many multiplexed assays. A gating
SUPPLEMENTAL MATERIALS
 SUPPLEMENL MERILS Eh-seq: RISPR epitope tagging hip-seq of DN-binding proteins Daniel Savic, E. hristopher Partridge, Kimberly M. Newberry, Sophia. Smith, Sarah K. Meadows, rian S. Roberts, Mark Mackiewicz,
SUPPLEMENL MERILS Eh-seq: RISPR epitope tagging hip-seq of DN-binding proteins Daniel Savic, E. hristopher Partridge, Kimberly M. Newberry, Sophia. Smith, Sarah K. Meadows, rian S. Roberts, Mark Mackiewicz,
Supplementary Figure 1
 Supplementary Figure 1 Supplementary Figure 1: Vector maps of TRMPV and TRMPVIR variants. Many derivatives of TRMPV have been generated and tested. Unless otherwise noted, experiments in this paper use
Supplementary Figure 1 Supplementary Figure 1: Vector maps of TRMPV and TRMPVIR variants. Many derivatives of TRMPV have been generated and tested. Unless otherwise noted, experiments in this paper use
NOTES - CH 15 (and 14.3): DNA Technology ( Biotech )
 NOTES - CH 15 (and 14.3): DNA Technology ( Biotech ) Vocabulary Genetic Engineering Gene Recombinant DNA Transgenic Restriction Enzymes Vectors Plasmids Cloning Key Concepts What is genetic engineering?
NOTES - CH 15 (and 14.3): DNA Technology ( Biotech ) Vocabulary Genetic Engineering Gene Recombinant DNA Transgenic Restriction Enzymes Vectors Plasmids Cloning Key Concepts What is genetic engineering?
jetprime in vitro DNA & sirna transfection reagent PROTOCOL
 jetprime in vitro DNA & sirna transfection reagent PROTOCOL DESCRIPTION jetprime is a novel powerful transfection reagent based on a polymer formulation manufactured at Polyplus-transfection. jetprime
jetprime in vitro DNA & sirna transfection reagent PROTOCOL DESCRIPTION jetprime is a novel powerful transfection reagent based on a polymer formulation manufactured at Polyplus-transfection. jetprime
Supplementary Figure 1: sgrna library generation and the length of sgrnas for the functional screen. (a) A diagram of the retroviral vector for sgrna
 Supplementary Figure 1: sgrna library generation and the length of sgrnas for the functional screen. (a) A diagram of the retroviral vector for sgrna expression. It contains a U6-promoter-driven sgrna
Supplementary Figure 1: sgrna library generation and the length of sgrnas for the functional screen. (a) A diagram of the retroviral vector for sgrna expression. It contains a U6-promoter-driven sgrna
REGISTRATION DOCUMENT FOR RECOMBINANT DNA RESEARCH
 EHRS Date Received: Reg. Doc. No.: REGISTRATION DOCUMENT FOR RECOMBINANT DNA RESEARCH Principal Investigator: Penn ID#: Position Title: School: Department: Mailing Address: Mail Code: Telephone: FAX: E-mail:
EHRS Date Received: Reg. Doc. No.: REGISTRATION DOCUMENT FOR RECOMBINANT DNA RESEARCH Principal Investigator: Penn ID#: Position Title: School: Department: Mailing Address: Mail Code: Telephone: FAX: E-mail:
Supplementary Figure 1
 number of cells, normalized number of cells, normalized number of cells, normalized Supplementary Figure CD CD53 Cd3e fluorescence intensity fluorescence intensity fluorescence intensity Supplementary
number of cells, normalized number of cells, normalized number of cells, normalized Supplementary Figure CD CD53 Cd3e fluorescence intensity fluorescence intensity fluorescence intensity Supplementary
THE DELIVERY EXPERTS PROTOCOL
 THE DELIVERY EXPERTS jetprime in vitro DNA & sirna transfection reagent PROTOCOL DESCRIPTION jetprime is a novel powerful molecule based on a polymer formulation manufactured at Polyplustransfection. jetprime
THE DELIVERY EXPERTS jetprime in vitro DNA & sirna transfection reagent PROTOCOL DESCRIPTION jetprime is a novel powerful molecule based on a polymer formulation manufactured at Polyplustransfection. jetprime
(Hadlock, Journal of Virology, 2000), (Keck, Journal of Virology, 2008) CBH-5 Domain B 412, 416, 417, 418, 420, 421, 422, 423, 483, 484, 485, 488,
 Supplementary Table. mabs analyzed mab Antigenic Critical Binding Residues by Citations Domains Alanine Scanning 2 CBH-4B Domain A NA (Hadlock, Journal of Virology, 2), (Keck, Journal of Virology, 24)
Supplementary Table. mabs analyzed mab Antigenic Critical Binding Residues by Citations Domains Alanine Scanning 2 CBH-4B Domain A NA (Hadlock, Journal of Virology, 2), (Keck, Journal of Virology, 24)
NEW! CHOgro Expression System
 NEW! CHOgro Expression System At Mirus Bio, we know it s all about expression. Introducing the new CHOgro Expression System, a transient transfection platform that finally gets high protein titers with
NEW! CHOgro Expression System At Mirus Bio, we know it s all about expression. Introducing the new CHOgro Expression System, a transient transfection platform that finally gets high protein titers with
Introducing new DNA into the genome requires cloning the donor sequence, delivery of the cloned DNA into the cell, and integration into the genome.
 Key Terms Chapter 32: Genetic Engineering Cloning describes propagation of a DNA sequence by incorporating it into a hybrid construct that can be replicated in a host cell. A cloning vector is a plasmid
Key Terms Chapter 32: Genetic Engineering Cloning describes propagation of a DNA sequence by incorporating it into a hybrid construct that can be replicated in a host cell. A cloning vector is a plasmid
Synthetic Biology. Sustainable Energy. Therapeutics Industrial Enzymes. Agriculture. Accelerating Discoveries, Expanding Possibilities. Design.
 Synthetic Biology Accelerating Discoveries, Expanding Possibilities Sustainable Energy Therapeutics Industrial Enzymes Agriculture Design Build Generate Solutions to Advance Synthetic Biology Research
Synthetic Biology Accelerating Discoveries, Expanding Possibilities Sustainable Energy Therapeutics Industrial Enzymes Agriculture Design Build Generate Solutions to Advance Synthetic Biology Research
A CRISPR/Cas9 Vector System for Tissue-Specific Gene Disruption in Zebrafish
 Developmental Cell Supplemental Information A CRISPR/Cas9 Vector System for Tissue-Specific Gene Disruption in Zebrafish Julien Ablain, Ellen M. Durand, Song Yang, Yi Zhou, and Leonard I. Zon % larvae
Developmental Cell Supplemental Information A CRISPR/Cas9 Vector System for Tissue-Specific Gene Disruption in Zebrafish Julien Ablain, Ellen M. Durand, Song Yang, Yi Zhou, and Leonard I. Zon % larvae
Lecture 3 Mutagens and Mutagenesis. 1. Mutagens A. Physical and Chemical mutagens B. Transposons and retrotransposons C. T-DNA
 Lecture 3 Mutagens and Mutagenesis 1. Mutagens A. Physical and Chemical mutagens B. Transposons and retrotransposons C. T-DNA 2. Mutagenesis A. Screen B. Selection C. Lethal mutations Read: 508-514 Figs:
Lecture 3 Mutagens and Mutagenesis 1. Mutagens A. Physical and Chemical mutagens B. Transposons and retrotransposons C. T-DNA 2. Mutagenesis A. Screen B. Selection C. Lethal mutations Read: 508-514 Figs:
Color-Switch CRE recombinase stable cell line
 Color-Switch CRE recombinase stable cell line Catalog Number Product Name / Description Amount SC018-Bsd CRE reporter cell line (Bsd): HEK293-loxP-GFP- RFP (Bsd). RFP" cassette with blasticidin antibiotic
Color-Switch CRE recombinase stable cell line Catalog Number Product Name / Description Amount SC018-Bsd CRE reporter cell line (Bsd): HEK293-loxP-GFP- RFP (Bsd). RFP" cassette with blasticidin antibiotic
CHAPTER 9 DNA Technologies
 CHAPTER 9 DNA Technologies Recombinant DNA Artificially created DNA that combines sequences that do not occur together in the nature Basis of much of the modern molecular biology Molecular cloning of genes
CHAPTER 9 DNA Technologies Recombinant DNA Artificially created DNA that combines sequences that do not occur together in the nature Basis of much of the modern molecular biology Molecular cloning of genes
Molecular Cell Biology - Problem Drill 11: Recombinant DNA
 Molecular Cell Biology - Problem Drill 11: Recombinant DNA Question No. 1 of 10 1. Which of the following statements about the sources of DNA used for molecular cloning is correct? Question #1 (A) cdna
Molecular Cell Biology - Problem Drill 11: Recombinant DNA Question No. 1 of 10 1. Which of the following statements about the sources of DNA used for molecular cloning is correct? Question #1 (A) cdna
Supplementary Information Design of small molecule-responsive micrornas based on structural requirements for Drosha processing
 Supplementary Information Design of small molecule-responsive micrornas based on structural requirements for Drosha processing Chase L. Beisel, Yvonne Y. Chen, Stephanie J. Culler, Kevin G. Hoff, & Christina
Supplementary Information Design of small molecule-responsive micrornas based on structural requirements for Drosha processing Chase L. Beisel, Yvonne Y. Chen, Stephanie J. Culler, Kevin G. Hoff, & Christina
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Origin use and efficiency are similar among WT, rrm3, pif1-m2, and pif1-m2; rrm3 strains. A. Analysis of fork progression around confirmed and likely origins (from cerevisiae.oridb.org).
Supplementary Figure 1 Origin use and efficiency are similar among WT, rrm3, pif1-m2, and pif1-m2; rrm3 strains. A. Analysis of fork progression around confirmed and likely origins (from cerevisiae.oridb.org).
Genetics Lecture 21 Recombinant DNA
 Genetics Lecture 21 Recombinant DNA Recombinant DNA In 1971, a paper published by Kathleen Danna and Daniel Nathans marked the beginning of the recombinant DNA era. The paper described the isolation of
Genetics Lecture 21 Recombinant DNA Recombinant DNA In 1971, a paper published by Kathleen Danna and Daniel Nathans marked the beginning of the recombinant DNA era. The paper described the isolation of
Nature Structural and Molecular Biology: doi: /nsmb.2937
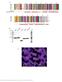 Supplementary Figure 1 Multiple sequence alignment of the CtIP N-terminal domain, purified CtIP protein constructs and details of the 2F o F c electron density map of CtIP-NTD. (a) Multiple sequence alignment,
Supplementary Figure 1 Multiple sequence alignment of the CtIP N-terminal domain, purified CtIP protein constructs and details of the 2F o F c electron density map of CtIP-NTD. (a) Multiple sequence alignment,
over time using live cell microscopy. The time post infection is indicated in the lower left corner.
 Title of file for HTML: Supplementary Information Description: Supplementary Figures and Supplementary Table Title of file for HTML: Supplementary Movie 1 Description: Fusion of NBs. BSR cells were infected
Title of file for HTML: Supplementary Information Description: Supplementary Figures and Supplementary Table Title of file for HTML: Supplementary Movie 1 Description: Fusion of NBs. BSR cells were infected
Supplementary information, Figure S1
 Supplementary information, Figure S1 (A) Schematic diagram of the sgrna and hspcas9 expression cassettes in a single binary vector designed for Agrobacterium-mediated stable transformation of Arabidopsis
Supplementary information, Figure S1 (A) Schematic diagram of the sgrna and hspcas9 expression cassettes in a single binary vector designed for Agrobacterium-mediated stable transformation of Arabidopsis
Retro-X qrt-pcr Titration Kit. User Manual. User Manual. Catalog No PT (091613)
 User Manual Retro-X qrt-pcr Titration Kit User Manual United States/Canada 800.662.2566 Asia Pacific +1.650.919.7300 Europe +33.(0)1.3904.6880 Japan +81.(0)77.543.6116 Clontech Laboratories, Inc. A Takara
User Manual Retro-X qrt-pcr Titration Kit User Manual United States/Canada 800.662.2566 Asia Pacific +1.650.919.7300 Europe +33.(0)1.3904.6880 Japan +81.(0)77.543.6116 Clontech Laboratories, Inc. A Takara
Chromosome-scale scaffolding of de novo genome assemblies based on chromatin interactions. Supplementary Material
 Chromosome-scale scaffolding of de novo genome assemblies based on chromatin interactions Joshua N. Burton 1, Andrew Adey 1, Rupali P. Patwardhan 1, Ruolan Qiu 1, Jacob O. Kitzman 1, Jay Shendure 1 1 Department
Chromosome-scale scaffolding of de novo genome assemblies based on chromatin interactions Joshua N. Burton 1, Andrew Adey 1, Rupali P. Patwardhan 1, Ruolan Qiu 1, Jacob O. Kitzman 1, Jay Shendure 1 1 Department
MIT Department of Biology 7.013: Introductory Biology - Spring 2005 Instructors: Professor Hazel Sive, Professor Tyler Jacks, Dr.
 MIT Department of Biology 7.01: Introductory Biology - Spring 2005 Instructors: Professor Hazel Sive, Professor Tyler Jacks, Dr. Claudette Gardel iv) Would Xba I be useful for cloning? Why or why not?
MIT Department of Biology 7.01: Introductory Biology - Spring 2005 Instructors: Professor Hazel Sive, Professor Tyler Jacks, Dr. Claudette Gardel iv) Would Xba I be useful for cloning? Why or why not?
TRANSFEX - SUPERIOR GENE EXPRESSION FOR HARD-TO-TRANSFECT CELL TYPES. Kevin Grady Product Line Business Manager ASCB Vendor Showcase Dec.
 TRANSFEX - SUPERIOR GENE EXPRESSION FOR HARD-TO-TRANSFECT CELL TYPES Kevin Grady Product Line Business Manager ASCB Vendor Showcase Dec. 15, 2013 Outline Overview of transfection TransfeX Primary/hTERT
TRANSFEX - SUPERIOR GENE EXPRESSION FOR HARD-TO-TRANSFECT CELL TYPES Kevin Grady Product Line Business Manager ASCB Vendor Showcase Dec. 15, 2013 Outline Overview of transfection TransfeX Primary/hTERT
NAME TA SEC Problem Set 4 FRIDAY October 15, Answers to this problem set must be inserted into the box outside
 MIT Biology Department 7.012: Introductory Biology - Fall 2004 Instructors: Professor Eric Lander, Professor Robert A. Weinberg, Dr. Claudette Gardel NAME TA SEC 7.012 Problem Set 4 FRIDAY October 15,
MIT Biology Department 7.012: Introductory Biology - Fall 2004 Instructors: Professor Eric Lander, Professor Robert A. Weinberg, Dr. Claudette Gardel NAME TA SEC 7.012 Problem Set 4 FRIDAY October 15,
Figure S1. Figure S2. Figure S3 HB Anti-FSP27 (COOH-terminal peptide) Ab. Anti-GST-FSP27(45-127) Ab.
 / 36B4 mrna ratio Figure S1 * 2. 1.6 1.2.8 *.4 control TNFα BRL49653 Figure S2 Su bw AT p iw Anti- (COOH-terminal peptide) Ab Blot : Anti-GST-(45-127) Ab β-actin Figure S3 HB2 HW AT BA T Figure S4 A TAG
/ 36B4 mrna ratio Figure S1 * 2. 1.6 1.2.8 *.4 control TNFα BRL49653 Figure S2 Su bw AT p iw Anti- (COOH-terminal peptide) Ab Blot : Anti-GST-(45-127) Ab β-actin Figure S3 HB2 HW AT BA T Figure S4 A TAG
TransIT -Lenti Transfection Reagent
 Quick Reference Protocol, SDS and Certificate of Analysis available at mirusbio.com/6600 INTRODUCTION Lentivirus is an enveloped, single-stranded RNA virus from the Retroviridae family capable of infecting
Quick Reference Protocol, SDS and Certificate of Analysis available at mirusbio.com/6600 INTRODUCTION Lentivirus is an enveloped, single-stranded RNA virus from the Retroviridae family capable of infecting
Antibiotic Resistance: Ampicillin Bacterial Backbone: pbluescriptksii(+), Agilent Technologies
 G0656 pfiv3.2rsvmcs Coordinates Plasmid Features 1-682 CMV/5 LTR hybrid 958-1468 Partial Gag 1494-1653 Central Polypurine Tract 1695-2075 RSV Promoter 2104-2222 Multiple Cloning Sites 2244-2839 WPRE 2868-3009
G0656 pfiv3.2rsvmcs Coordinates Plasmid Features 1-682 CMV/5 LTR hybrid 958-1468 Partial Gag 1494-1653 Central Polypurine Tract 1695-2075 RSV Promoter 2104-2222 Multiple Cloning Sites 2244-2839 WPRE 2868-3009
Supplementary Material. Manuscript title: Cross-immunity and community structure of a multiple-strain pathogen in the
 Supplementary Material Manuscript title: Cross-immunity and community structure of a multiple-strain pathogen in the tick vector Description of the primers used in the second PCR: The forward and the reverse
Supplementary Material Manuscript title: Cross-immunity and community structure of a multiple-strain pathogen in the tick vector Description of the primers used in the second PCR: The forward and the reverse
Orflo Application Brief 3 /2017 GFP Transfection Efficiency Monitoring with Orflo s Moxi GO Next Generation Flow Cytometer. Introduction/Background
 Introduction/Background Cell transfection and transduction refer to an array of techniques used to introduce foreign genetic material, or cloning vectors, into cell genomes. The application of these methods
Introduction/Background Cell transfection and transduction refer to an array of techniques used to introduce foreign genetic material, or cloning vectors, into cell genomes. The application of these methods
Color-Switch CRE Reporter Stable Cell Line
 Color-Switch CRE Reporter Stable Cell Line Catalog Number SC018-Bsd SC018-Neo SC018-Puro Product Name / Description CRE reporter cell line (Bsd): HEK293-loxP-GFP- RFP (Bsd). Cell line expresses "LoxP-GFP-stop-
Color-Switch CRE Reporter Stable Cell Line Catalog Number SC018-Bsd SC018-Neo SC018-Puro Product Name / Description CRE reporter cell line (Bsd): HEK293-loxP-GFP- RFP (Bsd). Cell line expresses "LoxP-GFP-stop-
The neutral theory of molecular evolution
 The neutral theory of molecular evolution Objectives the neutral theory detecting natural selection exercises 1 - learn about the neutral theory 2 - be able to detect natural selection at the molecular
The neutral theory of molecular evolution Objectives the neutral theory detecting natural selection exercises 1 - learn about the neutral theory 2 - be able to detect natural selection at the molecular
Learning Objectives :
 Learning Objectives : Understand the basic differences between genomic and cdna libraries Understand how genomic libraries are constructed Understand the purpose for having overlapping DNA fragments in
Learning Objectives : Understand the basic differences between genomic and cdna libraries Understand how genomic libraries are constructed Understand the purpose for having overlapping DNA fragments in
ViraBind PLUS Retrovirus Concentration and Purification Mega Kit
 Product Manual ViraBind PLUS Retrovirus Concentration and Purification Mega Kit Catalog Number VPK-136 VPK-136-5 2 preps 10 preps FOR RESEARCH USE ONLY Not for use in diagnostic procedures Introduction
Product Manual ViraBind PLUS Retrovirus Concentration and Purification Mega Kit Catalog Number VPK-136 VPK-136-5 2 preps 10 preps FOR RESEARCH USE ONLY Not for use in diagnostic procedures Introduction
Supplementary Information
 Journal : Nature Biotechnology Supplementary Information Targeted genome engineering in human cells with RNA-guided endonucleases Seung Woo Cho, Sojung Kim, Jong Min Kim, and Jin-Soo Kim* National Creative
Journal : Nature Biotechnology Supplementary Information Targeted genome engineering in human cells with RNA-guided endonucleases Seung Woo Cho, Sojung Kim, Jong Min Kim, and Jin-Soo Kim* National Creative
Technical Review. Real time PCR
 Technical Review Real time PCR Normal PCR: Analyze with agarose gel Normal PCR vs Real time PCR Real-time PCR, also known as quantitative PCR (qpcr) or kinetic PCR Key feature: Used to amplify and simultaneously
Technical Review Real time PCR Normal PCR: Analyze with agarose gel Normal PCR vs Real time PCR Real-time PCR, also known as quantitative PCR (qpcr) or kinetic PCR Key feature: Used to amplify and simultaneously
Supplementary Figure 1 qrt-pcr expression analysis of NLP8 with and without KNO 3 during germination.
 Supplementary Figure 1 qrt-pcr expression analysis of NLP8 with and without KNO 3 during germination. Seeds of Col-0 were harvested from plants grown at 16 C, stored for 2 months, imbibed for indicated
Supplementary Figure 1 qrt-pcr expression analysis of NLP8 with and without KNO 3 during germination. Seeds of Col-0 were harvested from plants grown at 16 C, stored for 2 months, imbibed for indicated
Discovery and Humanization of Novel High Affinity Neutralizing Monoclonal Antibodies to Human IL-17A
 Discovery and Humanization of Novel High Affinity Neutralizing Monoclonal Antibodies to Human IL-17A Contacts: Marty Simonetti martysimonetti@gmail.com Kirby Alton kirby.alton@abeomecorp.com Rick Shimkets
Discovery and Humanization of Novel High Affinity Neutralizing Monoclonal Antibodies to Human IL-17A Contacts: Marty Simonetti martysimonetti@gmail.com Kirby Alton kirby.alton@abeomecorp.com Rick Shimkets
How Do You Clone a Gene?
 S-20 Edvo-Kit #S-20 How Do You Clone a Gene? Experiment Objective: The objective of this experiment is to gain an understanding of the structure of DNA, a genetically engineered clone, and how genes are
S-20 Edvo-Kit #S-20 How Do You Clone a Gene? Experiment Objective: The objective of this experiment is to gain an understanding of the structure of DNA, a genetically engineered clone, and how genes are
Supplementary Methods. Plasmid construction. NS5A-fluorophore fusion Jc1 genomes were generated
 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 Supplementary Methods Plasmid construction. NS5A-fluorophore fusion Jc1 genomes were generated by introducing a linker containing an XbaI and
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 Supplementary Methods Plasmid construction. NS5A-fluorophore fusion Jc1 genomes were generated by introducing a linker containing an XbaI and
Supplementary Data: Fig. 1S Detailed description of In vivo experimental design
 1 2 Supplementary Data: Fig. 1S Detailed description of In vivo experimental design 3 4 5 6 7 8 9 Relative Expression Studies 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 The
1 2 Supplementary Data: Fig. 1S Detailed description of In vivo experimental design 3 4 5 6 7 8 9 Relative Expression Studies 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 The
Figure 1: E. Coli lysate transfer using liquid handling automation
 Figure 1: E. Coli lysate transfer using liquid handling automation Figure 1 - E. coli lysate transfer using liquid handling automation. Following the manufacturer s procedures, a 96-well plate miniprep
Figure 1: E. Coli lysate transfer using liquid handling automation Figure 1 - E. coli lysate transfer using liquid handling automation. Following the manufacturer s procedures, a 96-well plate miniprep
Improving CRISPR-Cas9 Gene Knockout with a Validated Guide RNA Algorithm
 Improving CRISPR-Cas9 Gene Knockout with a Validated Guide RNA Algorithm Anja Smith Director R&D Dharmacon, part of GE Healthcare Imagination at work crrna:tracrrna program Cas9 nuclease Active crrna is
Improving CRISPR-Cas9 Gene Knockout with a Validated Guide RNA Algorithm Anja Smith Director R&D Dharmacon, part of GE Healthcare Imagination at work crrna:tracrrna program Cas9 nuclease Active crrna is
Bio-Reagent Services. Custom Gene Services. Gateway to Smooth Molecular Biology! Your Innovation Partner in Drug Discovery!
 Bio-Reagent Services Custom Gene Services Gateway to Smooth Molecular Biology! Gene Synthesis Mutagenesis Mutant Libraries Plasmid Preparation sirna and mirna Services Large-scale DNA Sequencing GenPool
Bio-Reagent Services Custom Gene Services Gateway to Smooth Molecular Biology! Gene Synthesis Mutagenesis Mutant Libraries Plasmid Preparation sirna and mirna Services Large-scale DNA Sequencing GenPool
CHAPTER 20 DNA TECHNOLOGY AND GENOMICS. Section A: DNA Cloning
 Section A: DNA Cloning 1. DNA technology makes it possible to clone genes for basic research and commercial applications: an overview 2. Restriction enzymes are used to make recombinant DNA 3. Genes can
Section A: DNA Cloning 1. DNA technology makes it possible to clone genes for basic research and commercial applications: an overview 2. Restriction enzymes are used to make recombinant DNA 3. Genes can
AP Biology Gene Expression/Biotechnology REVIEW
 AP Biology Gene Expression/Biotechnology REVIEW Multiple Choice Identify the choice that best completes the statement or answers the question. 1. Gene expression can be a. regulated before transcription.
AP Biology Gene Expression/Biotechnology REVIEW Multiple Choice Identify the choice that best completes the statement or answers the question. 1. Gene expression can be a. regulated before transcription.
Green Fluorescent Protein. Avinash Bayya Varun Maturi Nikhileswar Reddy Mukkamala Aravindh Subhramani
 Green Fluorescent Protein Avinash Bayya Varun Maturi Nikhileswar Reddy Mukkamala Aravindh Subhramani Introduction The Green fluorescent protein (GFP) was first isolated from the Jellyfish Aequorea victoria,
Green Fluorescent Protein Avinash Bayya Varun Maturi Nikhileswar Reddy Mukkamala Aravindh Subhramani Introduction The Green fluorescent protein (GFP) was first isolated from the Jellyfish Aequorea victoria,
Sept 2. Structure and Organization of Genomes. Today: Genetic and Physical Mapping. Sept 9. Forward and Reverse Genetics. Genetic and Physical Mapping
 Sept 2. Structure and Organization of Genomes Today: Genetic and Physical Mapping Assignments: Gibson & Muse, pp.4-10 Brown, pp. 126-160 Olson et al., Science 245: 1434 New homework:due, before class,
Sept 2. Structure and Organization of Genomes Today: Genetic and Physical Mapping Assignments: Gibson & Muse, pp.4-10 Brown, pp. 126-160 Olson et al., Science 245: 1434 New homework:due, before class,
Xfect Protein Transfection Reagent
 Xfect Protein Transfection Reagent Mammalian Expression Systems Rapid, high-efficiency, low-toxicity protein transfection Transfect a large amount of active protein Virtually no cytotoxicity, unlike lipofection
Xfect Protein Transfection Reagent Mammalian Expression Systems Rapid, high-efficiency, low-toxicity protein transfection Transfect a large amount of active protein Virtually no cytotoxicity, unlike lipofection
Recombination between Two Identical Sequences within the Same Retroviral RNA Molecule
 JOURNAL OF VIROLOGY, July 1999, p. 5912 5917 Vol. 73, No. 7 0022-538X/99/$04.00 0 Copyright 1999, American Society for Microbiology. All Rights Reserved. Recombination between Two Identical Sequences within
JOURNAL OF VIROLOGY, July 1999, p. 5912 5917 Vol. 73, No. 7 0022-538X/99/$04.00 0 Copyright 1999, American Society for Microbiology. All Rights Reserved. Recombination between Two Identical Sequences within
Evolutionary optimization of fluorescent proteins for intracellular FRET
 Evolutionary optimization of fluorescent proteins for intracellular FRET Annalee W Nguyen 1 & Patrick S Daugherty 1 Fluorescent proteins that exhibit Förster resonance energy transfer (FRET) have made
Evolutionary optimization of fluorescent proteins for intracellular FRET Annalee W Nguyen 1 & Patrick S Daugherty 1 Fluorescent proteins that exhibit Förster resonance energy transfer (FRET) have made
Optimizing Synthetic DNA for Metabolic Engineering Applications. Howard Salis Penn State University
 Optimizing Synthetic DNA for Metabolic Engineering Applications Howard Salis Penn State University Synthetic Biology Specify a function Build a genetic system (a DNA molecule) Genetic Pseudocode call producequorumsignal(luxi
Optimizing Synthetic DNA for Metabolic Engineering Applications Howard Salis Penn State University Synthetic Biology Specify a function Build a genetic system (a DNA molecule) Genetic Pseudocode call producequorumsignal(luxi
BIBC 103 Learning Goals with Supporting Learning Outcomes
 BIBC 103 Learning Goals with Supporting Learning Outcomes 1) Basic Lab Skills A. Conceptual understanding and moderate level of hands-on proficiency in making laboratory solutions, including understanding
BIBC 103 Learning Goals with Supporting Learning Outcomes 1) Basic Lab Skills A. Conceptual understanding and moderate level of hands-on proficiency in making laboratory solutions, including understanding
MATH 5610, Computational Biology
 MATH 5610, Computational Biology Lecture 2 Intro to Molecular Biology (cont) Stephen Billups University of Colorado at Denver MATH 5610, Computational Biology p.1/24 Announcements Error on syllabus Class
MATH 5610, Computational Biology Lecture 2 Intro to Molecular Biology (cont) Stephen Billups University of Colorado at Denver MATH 5610, Computational Biology p.1/24 Announcements Error on syllabus Class
User s Guide MegaTran 1.0 Transfection Reagent
 User s Guide MegaTran 1.0 Transfection Reagent Package Contents and Storage Conditions... 2 Related products... 2 Introduction... 2 Production and Quality Assurance:... 3 Experimental Procedures... 3 Transfection
User s Guide MegaTran 1.0 Transfection Reagent Package Contents and Storage Conditions... 2 Related products... 2 Introduction... 2 Production and Quality Assurance:... 3 Experimental Procedures... 3 Transfection
American Society of Cytopathology Core Curriculum in Molecular Biology
 American Society of Cytopathology Core Curriculum in Molecular Biology American Society of Cytopathology Core Curriculum in Molecular Biology Chapter 3 Molecular Techniques Separation and Detection, Part
American Society of Cytopathology Core Curriculum in Molecular Biology American Society of Cytopathology Core Curriculum in Molecular Biology Chapter 3 Molecular Techniques Separation and Detection, Part
Bio Rad PCR Song Lyrics
 Bio Rad PCR Song Lyrics There was a time when to amplify DNA, You had to grow tons and tons of tiny cells. (Oooh) Then along came a guy named Dr. Kary Mullis, Said you can amplify in vitro just as well.
Bio Rad PCR Song Lyrics There was a time when to amplify DNA, You had to grow tons and tons of tiny cells. (Oooh) Then along came a guy named Dr. Kary Mullis, Said you can amplify in vitro just as well.
BENG 183 Trey Ideker. Genome Assembly and Physical Mapping
 BENG 183 Trey Ideker Genome Assembly and Physical Mapping Reasons for sequencing Complete genome sequencing!!! Resequencing (Confirmatory) E.g., short regions containing single nucleotide polymorphisms
BENG 183 Trey Ideker Genome Assembly and Physical Mapping Reasons for sequencing Complete genome sequencing!!! Resequencing (Confirmatory) E.g., short regions containing single nucleotide polymorphisms
Nature Biotechnology: doi: /nbt Supplementary Figure 1. Number and length distributions of the inferred fosmids.
 Supplementary Figure 1 Number and length distributions of the inferred fosmids. Fosmid were inferred by mapping each pool s sequence reads to hg19. We retained only those reads that mapped to within a
Supplementary Figure 1 Number and length distributions of the inferred fosmids. Fosmid were inferred by mapping each pool s sequence reads to hg19. We retained only those reads that mapped to within a
Sort-seq under the hood: implications of design choices on largescale characterization of sequence-function relations
 Sort-seq under the hood: implications of design choices on largescale characterization of sequence-function relations The Harvard community has made this article openly available. Please share how this
Sort-seq under the hood: implications of design choices on largescale characterization of sequence-function relations The Harvard community has made this article openly available. Please share how this
Masayoshi Honda, Jeehae Park, Robert A. Pugh, Taekjip Ha, and Maria Spies
 Molecular Cell, Volume 35 Supplemental Data Single-Molecule Analysis Reveals Differential Effect of ssdna-binding Proteins on DNA Translocation by XPD Helicase Masayoshi Honda, Jeehae Park, Robert A. Pugh,
Molecular Cell, Volume 35 Supplemental Data Single-Molecule Analysis Reveals Differential Effect of ssdna-binding Proteins on DNA Translocation by XPD Helicase Masayoshi Honda, Jeehae Park, Robert A. Pugh,
Phenotypic response conferred by the Lr22a leaf rust resistance gene against ten Swiss P. triticina isolates.
 Supplementary Figure 1 Phenotypic response conferred by the leaf rust resistance gene against ten Swiss P. triticina isolates. The third leaf of Thatcher (left) and RL6044 (right) is shown ten days after
Supplementary Figure 1 Phenotypic response conferred by the leaf rust resistance gene against ten Swiss P. triticina isolates. The third leaf of Thatcher (left) and RL6044 (right) is shown ten days after
3 Designing Primers for Site-Directed Mutagenesis
 3 Designing Primers for Site-Directed Mutagenesis 3.1 Learning Objectives During the next two labs you will learn the basics of site-directed mutagenesis: you will design primers for the mutants you designed
3 Designing Primers for Site-Directed Mutagenesis 3.1 Learning Objectives During the next two labs you will learn the basics of site-directed mutagenesis: you will design primers for the mutants you designed
Some types of Mutagenesis
 Mutagenesis What Is a Mutation? Genetic information is encoded by the sequence of the nucleotide bases in DNA of the gene. The four nucleotides are: adenine (A), thymine (T), guanine (G), and cytosine
Mutagenesis What Is a Mutation? Genetic information is encoded by the sequence of the nucleotide bases in DNA of the gene. The four nucleotides are: adenine (A), thymine (T), guanine (G), and cytosine
psmpuw-ires-blasticidin Lentiviral Expression Vector
 Product Data Sheet psmpuw-ires-blasticidin Lentiviral Expression Vector CATALOG NUMBER: VPK-219 STORAGE: -20ºC QUANTITY AND CONCENTRATION: 10 µg at 0.25 µg/µl in TE Background Lentivirus vector based on
Product Data Sheet psmpuw-ires-blasticidin Lentiviral Expression Vector CATALOG NUMBER: VPK-219 STORAGE: -20ºC QUANTITY AND CONCENTRATION: 10 µg at 0.25 µg/µl in TE Background Lentivirus vector based on
Cre Stoplight with Living Colors is a faster, brighter
 Cre Stoplight with Living Colors is a faster, brighter reporter for Cre recombinase. Drago A Guggiana-Nilo 1, Anne Marie Quinn 2,Thomas E. Hughes 1 1 Department of Cell Biology and Neuroscience, Montana
Cre Stoplight with Living Colors is a faster, brighter reporter for Cre recombinase. Drago A Guggiana-Nilo 1, Anne Marie Quinn 2,Thomas E. Hughes 1 1 Department of Cell Biology and Neuroscience, Montana
Introduction to Next Generation Sequencing (NGS) Andrew Parrish Exeter, 2 nd November 2017
 Introduction to Next Generation Sequencing (NGS) Andrew Parrish Exeter, 2 nd November 2017 Topics to cover today What is Next Generation Sequencing (NGS)? Why do we need NGS? Common approaches to NGS NGS
Introduction to Next Generation Sequencing (NGS) Andrew Parrish Exeter, 2 nd November 2017 Topics to cover today What is Next Generation Sequencing (NGS)? Why do we need NGS? Common approaches to NGS NGS
High-Resolution Oligonucleotide- Based acgh Analysis of Single Cells in Under 24 Hours
 High-Resolution Oligonucleotide- Based acgh Analysis of Single Cells in Under 24 Hours Application Note Authors Paula Costa and Anniek De Witte Agilent Technologies, Inc. Santa Clara, CA USA Abstract As
High-Resolution Oligonucleotide- Based acgh Analysis of Single Cells in Under 24 Hours Application Note Authors Paula Costa and Anniek De Witte Agilent Technologies, Inc. Santa Clara, CA USA Abstract As
Manipulating DNA. Nucleic acids are chemically different from other macromolecules such as proteins and carbohydrates.
 Lesson Overview 14.3 Studying the Human Genome Nucleic acids are chemically different from other macromolecules such as proteins and carbohydrates. Nucleic acids are chemically different from other macromolecules
Lesson Overview 14.3 Studying the Human Genome Nucleic acids are chemically different from other macromolecules such as proteins and carbohydrates. Nucleic acids are chemically different from other macromolecules
Supplemental Information. Boundary Formation through a Direct. Threshold-Based Readout. of Mobile Small RNA Gradients
 Developmental Cell, Volume 43 Supplemental Information Boundary Formation through a Direct Threshold-Based Readout of Mobile Small RNA Gradients Damianos S. Skopelitis, Anna H. Benkovics, Aman Y. Husbands,
Developmental Cell, Volume 43 Supplemental Information Boundary Formation through a Direct Threshold-Based Readout of Mobile Small RNA Gradients Damianos S. Skopelitis, Anna H. Benkovics, Aman Y. Husbands,
Supplementary Figures Montero et al._supplementary Figure 1
 Montero et al_suppl. Info 1 Supplementary Figures Montero et al._supplementary Figure 1 Montero et al_suppl. Info 2 Supplementary Figure 1. Transcripts arising from the structurally conserved subtelomeres
Montero et al_suppl. Info 1 Supplementary Figures Montero et al._supplementary Figure 1 Montero et al_suppl. Info 2 Supplementary Figure 1. Transcripts arising from the structurally conserved subtelomeres
F 11/23 Happy Thanksgiving! 8 M 11/26 Gene identification in the genomic era Bamshad et al. Nature Reviews Genetics 12: , 2011
 3 rd Edition 4 th Edition Lecture Day Date Topic Reading Problems Reading Problems 1 M 11/5 Complementation testing reveals that genes are distinct entities Ch. 7 224-232 2 W 11/7 One gene makes one protein
3 rd Edition 4 th Edition Lecture Day Date Topic Reading Problems Reading Problems 1 M 11/5 Complementation testing reveals that genes are distinct entities Ch. 7 224-232 2 W 11/7 One gene makes one protein
Supplemental Information. A Versatile Tool for Live-Cell Imaging. and Super-Resolution Nanoscopy Studies. of HIV-1 Env Distribution and Mobility
 Cell Chemical Biology, Volume 24 Supplemental Information A Versatile Tool for Live-Cell Imaging and Super-Resolution Nanoscopy Studies of HIV-1 Env Distribution and Mobility Volkan Sakin, Janina Hanne,
Cell Chemical Biology, Volume 24 Supplemental Information A Versatile Tool for Live-Cell Imaging and Super-Resolution Nanoscopy Studies of HIV-1 Env Distribution and Mobility Volkan Sakin, Janina Hanne,
Nature Immunology: doi: /ni Supplementary Figure 1
 Supplementary Figure 1 BALB/c LYVE1-deficient mice exhibited reduced lymphatic trafficking of all DC subsets after oxazolone-induced sensitization. (a) Schematic overview of the mouse skin oxazolone contact
Supplementary Figure 1 BALB/c LYVE1-deficient mice exhibited reduced lymphatic trafficking of all DC subsets after oxazolone-induced sensitization. (a) Schematic overview of the mouse skin oxazolone contact
SUPPLEMENTARY INFORMATION
 In the format provided by the authors and unedited. SUPPLEMENTARY INFORMATION ARTICLE NUMBER: 16206 DOI: 10.1038/NMICROBIOL.2016.206 Single cell RNA seq ties macrophage polarization to growth rate of intracellular
In the format provided by the authors and unedited. SUPPLEMENTARY INFORMATION ARTICLE NUMBER: 16206 DOI: 10.1038/NMICROBIOL.2016.206 Single cell RNA seq ties macrophage polarization to growth rate of intracellular
Application Note. NGS Analysis of B-Cell Receptors & Antibodies by AptaAnalyzer -BCR
 Reduce to the Best Application Note NGS Analysis of B-Cell Receptors & Antibodies by AptaAnalyzer -BCR The software AptaAnalyzer harnesses next generation sequencing (NGS) data to monitor the immune response
Reduce to the Best Application Note NGS Analysis of B-Cell Receptors & Antibodies by AptaAnalyzer -BCR The software AptaAnalyzer harnesses next generation sequencing (NGS) data to monitor the immune response
Neutral theory: The neutral theory does not say that all evolution is neutral and everything is only due to to genetic drift.
 Neutral theory: The vast majority of observed sequence differences between members of a population are neutral (or close to neutral). These differences can be fixed in the population through random genetic
Neutral theory: The vast majority of observed sequence differences between members of a population are neutral (or close to neutral). These differences can be fixed in the population through random genetic
Applications of the Ion AmpliSeq Immune Repertoire Assay Plus TCRβ
 Applications of the Ion AmpliSeq Immune Repertoire Assay Plus TCRβ Timothy Looney, PhD Staff Scientist, Clinical Next-Generation Sequencing Division Thermo Fisher Scientific The world leader in serving
Applications of the Ion AmpliSeq Immune Repertoire Assay Plus TCRβ Timothy Looney, PhD Staff Scientist, Clinical Next-Generation Sequencing Division Thermo Fisher Scientific The world leader in serving
Supporting Information
 Supporting Information Yuan et al. 10.1073/pnas.0906869106 Fig. S1. Heat map showing that Populus ICS is coregulated with orthologs of Arabidopsis genes involved in PhQ biosynthesis and PSI function, but
Supporting Information Yuan et al. 10.1073/pnas.0906869106 Fig. S1. Heat map showing that Populus ICS is coregulated with orthologs of Arabidopsis genes involved in PhQ biosynthesis and PSI function, but
Supplementary Table 1. The Q-PCR primer sequence is summarized in the following table.
 Supplementary Table 1. The Q-PCR primer sequence is summarized in the following table. Name Sequence (5-3 ) Application Flag-u ggactacaaggacgacgatgac Shared upstream primer for all the amplifications of
Supplementary Table 1. The Q-PCR primer sequence is summarized in the following table. Name Sequence (5-3 ) Application Flag-u ggactacaaggacgacgatgac Shared upstream primer for all the amplifications of
Application of next generation sequencing of a begomovirusresistant. KASPar assay for SNP detection of the Ty1-Ty3 region
 Application of next generation sequencing of a begomovirusresistant inbred to design a KASPar assay for SNP detection of the Ty1-Ty3 region Menda, N. 1, S. Strickler 1, D.M. Dunham 1, D.P. Maxwell 2, L.
Application of next generation sequencing of a begomovirusresistant inbred to design a KASPar assay for SNP detection of the Ty1-Ty3 region Menda, N. 1, S. Strickler 1, D.M. Dunham 1, D.P. Maxwell 2, L.
psmpuw-puro Lentiviral Expression Vector
 Product Data Sheet psmpuw-puro Lentiviral Expression Vector CATALOG NUMBER: VPK-212 STORAGE: -20ºC QUANTITY AND CONCENTRATION: 10 µg at 0.25 µg/µl in TE Background Lentivirus vector based on the human
Product Data Sheet psmpuw-puro Lentiviral Expression Vector CATALOG NUMBER: VPK-212 STORAGE: -20ºC QUANTITY AND CONCENTRATION: 10 µg at 0.25 µg/µl in TE Background Lentivirus vector based on the human
Amplicon Sequencing Template Preparation
 Amplicon Sequencing Template Preparation The DNA sample preparation procedure for Amplicon Sequencing consists of a simple PCR amplification reaction, but uses special Fusion Primers (Figure 1-1). The
Amplicon Sequencing Template Preparation The DNA sample preparation procedure for Amplicon Sequencing consists of a simple PCR amplification reaction, but uses special Fusion Primers (Figure 1-1). The
ViraBind PLUS Retrovirus Concentration and Purification Kit
 Product Manual ViraBind PLUS Retrovirus Concentration and Purification Kit Catalog Number VPK-135 2 preps FOR RESEARCH USE ONLY Not for use in diagnostic procedures Introduction Retroviral gene transfer
Product Manual ViraBind PLUS Retrovirus Concentration and Purification Kit Catalog Number VPK-135 2 preps FOR RESEARCH USE ONLY Not for use in diagnostic procedures Introduction Retroviral gene transfer
Biosafety Level Host Range Propagation Comments
 Guidelines BSL for Commonly used Viral Vectors Version 1.0 Office of Animal Care and Institutional Biosafety (OACIB) 1737 West Polk Street (MC 672) 206 Administrative Office Building Chicago, IL 60612
Guidelines BSL for Commonly used Viral Vectors Version 1.0 Office of Animal Care and Institutional Biosafety (OACIB) 1737 West Polk Street (MC 672) 206 Administrative Office Building Chicago, IL 60612
Lecture 25 (11/15/17)
 Lecture 25 (11/15/17) Reading: Ch9; 328-332 Ch25; 990-995, 1005-1012 Problems: Ch9 (study-guide: applying); 1,2 Ch9 (study-guide: facts); 7,8 Ch25 (text); 1-3,5-7,9,10,13-15 Ch25 (study-guide: applying);
Lecture 25 (11/15/17) Reading: Ch9; 328-332 Ch25; 990-995, 1005-1012 Problems: Ch9 (study-guide: applying); 1,2 Ch9 (study-guide: facts); 7,8 Ch25 (text); 1-3,5-7,9,10,13-15 Ch25 (study-guide: applying);
X-ray structures of fructosyl peptide oxidases revealing residues responsible for gating oxygen access in the oxidative half reaction
 X-ray structures of fructosyl peptide oxidases revealing residues responsible for gating oxygen access in the oxidative half reaction Tomohisa Shimasaki 1, Hiromi Yoshida 2, Shigehiro Kamitori 2 & Koji
X-ray structures of fructosyl peptide oxidases revealing residues responsible for gating oxygen access in the oxidative half reaction Tomohisa Shimasaki 1, Hiromi Yoshida 2, Shigehiro Kamitori 2 & Koji
Exam MOL3007 Functional Genomics
 Faculty of Medicine Department of Cancer Research and Molecular Medicine Exam MOL3007 Functional Genomics Tuesday May 29 th 9.00-13.00 ECTS credits: 7.5 Number of pages (included front-page): 5 Supporting
Faculty of Medicine Department of Cancer Research and Molecular Medicine Exam MOL3007 Functional Genomics Tuesday May 29 th 9.00-13.00 ECTS credits: 7.5 Number of pages (included front-page): 5 Supporting
Selection of Target Sites for Mobile DNA Integration in the Human Genome
 Selection of Target Sites for Mobile DNA Integration in the Human Genome Charles Berry 1, Sridhar Hannenhalli 2, Jeremy Leipzig 3, Frederic D. Bushman 3* 1 Department of Family and Preventive Medicine,
Selection of Target Sites for Mobile DNA Integration in the Human Genome Charles Berry 1, Sridhar Hannenhalli 2, Jeremy Leipzig 3, Frederic D. Bushman 3* 1 Department of Family and Preventive Medicine,
Thermo Scientific Dharmacon SMARTvector 2.0 Lentiviral shrna Particles
 Thermo Scientific Dharmacon SMARTvector 2.0 Lentiviral shrna Particles Long-term gene silencing shrna-specific design algorithm High titer, purified particles Thermo Scientific Dharmacon SMARTvector shrna
Thermo Scientific Dharmacon SMARTvector 2.0 Lentiviral shrna Particles Long-term gene silencing shrna-specific design algorithm High titer, purified particles Thermo Scientific Dharmacon SMARTvector shrna
Using mutants to clone genes
 Using mutants to clone genes Objectives 1. What is positional cloning? 2. What is insertional tagging? 3. How can one confirm that the gene cloned is the same one that is mutated to give the phenotype
Using mutants to clone genes Objectives 1. What is positional cloning? 2. What is insertional tagging? 3. How can one confirm that the gene cloned is the same one that is mutated to give the phenotype
BIOCHEMISTRY 551: BIOCHEMICAL METHODS SYLLABUS
 BIOCHEMISTRY 551: BIOCHEMICAL METHODS SYLLABUS Course Description: Biochemistry 551 is an integrated lecture, lab and seminar course that covers biochemistry-centered theory and techniques. The course
BIOCHEMISTRY 551: BIOCHEMICAL METHODS SYLLABUS Course Description: Biochemistry 551 is an integrated lecture, lab and seminar course that covers biochemistry-centered theory and techniques. The course
Supplemental Information. Natural RNA Polymerase Aptamers. Regulate Transcription in E. coli
 Molecular Cell, Volume 67 Supplemental Information Natural RNA Polymerase Aptamers Regulate Transcription in E. coli Nadezda Sedlyarova, Philipp Rescheneder, Andrés Magán, Niko Popitsch, Natascha Rziha,
Molecular Cell, Volume 67 Supplemental Information Natural RNA Polymerase Aptamers Regulate Transcription in E. coli Nadezda Sedlyarova, Philipp Rescheneder, Andrés Magán, Niko Popitsch, Natascha Rziha,
Genome Engineering. Brian Petuch
 Genome Engineering Brian Petuch Guiding Principles An understanding of the details of gene engineering is essential for understanding protocols when making recommendations on: Risk assessment Containment
Genome Engineering Brian Petuch Guiding Principles An understanding of the details of gene engineering is essential for understanding protocols when making recommendations on: Risk assessment Containment
