Structure determination and activity manipulation of the turfgrass ABA receptor FePYR1
|
|
|
- Chester Owen
- 6 years ago
- Views:
Transcription
1 Supplemental information for: Structure determination and activity manipulation of the turfgrass AA receptor FePYR1 Zhizhong Ren 1, 2,#, Zhen Wang 1, 2,#, X Edward Zhou 3, Huazhong Shi 4, Yechun Hong 1, 2, Minjie Cao 1, Zhulong Chan 5, Xue Liu 1, H Eric Xu 3, 6, * 1, 7, *, Jian-Kang Zhu 1 Shanghai Center for Plant Stress iology and Center for Excellence in Molecular Plant Sciences, 2 Chinese Academy of Sciences, Shanghai , China; University of Chinese Academy of Sciences (CAS), Shanghai , P.R. China; 3 Laboratory of Structural Sciences and Laboratory of Structural iology and iochemistry, Van Andel Research Institute, Grand Rapids, MI, USA; 4 Department of Chemistry and iochemistry, Texas Tech University, Lubbock, Texas 79409; 5 Key Laboratory of Horticultural Plant iology, Ministry of Education, College of Horticulture & Forest Sciences, Huazhong Agricultural University, Wuhan , China; 6 Key Laboratory of Receptor Research, VARI-SIMM Center, Center for Structure and Function of Drug Targets, Shanghai Institute of Materia Medica, Chinese Academy of Sciences, Shanghai, China; 7 Department of Horticulture and Landscape Architecture, Purdue University, West Lafayette, Indiana 47907
2 A subfamily-Ⅰ subfamily-Ⅱ subfamily-Ⅲ C Supplemental Figure 1. Determination of AA receptors in the two turfgrass species. (A) A phylogeny of multiple sequence alignment of the turfgrass and Arabidopsis PYR/PYLs using the neighbor joining method in MEGA5. These proteins were divided into three major subfamilies I, II and III to their phylogenic relationships. () Identification of Festuca elata AA receptors by Alpha-Screen assay. Interactions of putative receptor proteins with the Arabidopsis AtHA1 were evaluated by photon intensity (n=3, error bars are means ± SD). (C) Determination of Zoysia japonica AA receptor proteins using Alpha-Screen assay. Interactions between ZjPYR/PYLs candidates and AtHA1 were analyzed by photon counts (n=3, error bars represent ± SD).
3 A 40 KD 30 KD 25 KD 20 KD FePYL1 30 KD FePYR1 25 KD 20 KD C D 30 KD 25 KD 20 KD ZjPYL2 Supplemental Figure 2. Purification and crystallization of the three putative turfgrass AA receptor proteins. (A-C) The size-exclusion chromatography (SEC) elution profiles for FePYL1, FePYR1 and ZjPYL2 respectively. The proteins in the fractions of SEC eluents were visualized by Coomassie rilliant lue staining. (D) Crystals of FePYR1-AA complex after 4 days hanging drop optimization in the buffer containing 0.1 M is- Tris propane (ph 7.0) and 1.5 M Ammonium sulfate.
4 A Supplemental Figure 3. Comparison of FePYR1 and AtPYR1 on binding with and inhibition of AtHA1. (A) Competition binding of FePYR1 and AtPYR1 analyzed by using luminescent proximity assay. The interaction strength of 100 nm biotinylated AtHA1 and 100 nm 6 His-fusion AtPYR1 in the presence of 100 μm AA was used as the initial intensity (100%) in the Alpha-Screen assays described in the Materials and Methods. Addition of unlabeled FePYR1 (red) or AtPYR1 (blue) at the indicated concentrations competed with 6 His-tagged PYR1 for binding to biotinylated AtHA1 (n=3, error bars represent ± SD). () Comparison of inhibition of AtHA1 phosphatase activity by FePYR1 and AtPYR nm biotinylated AtHA1 and 500 nm H6-SUMO tagged FePYR1 (red) or AtPYR1 (blue) were incubated at indicated concentrations of AA. AtHA1 phosphatase activity was determined by colorimetric assay (iovision) (n=3, error bars mean ± SD).
5 A C H73 L185 L185 Supplemental Figure 4. Detailed conformational views of AA in the aligned structures of AA-FePYR1 and AA-AtPYL9. (A, ) The side view of AA bound to FePYR1 (A) and AtPYL9 (). The bound AA molecule (aurantia) in FePYR1 forms hydrogen bonds with Lys72 and has no other direct hydrogen bonds with amino acids at the bottom of FePYR1 pocket. An intra-molecule hydrogen bond is formed between the Asn186 and His73. AA molecule (green) makes intimate contacts with AtPYL9 by forming direct hydrogen bonds with Asn169 and Lys63. Direct hydrogen bonds between AA molecule and proteins were shown with broken lines, water molecules were shown as red balls. (C) The detailed structure of FePYR1 interface formed by α3 helix between two adjacent molecules. The backbone of two adjacent FePYR1 molecules were shown in cyan with semitransparent molecular surface and magenta respectively. The relevant residues were shown in yellow.
6 A Arabidopsis thaliana Festuca elata Supplemental Figure 5. Measurements of AA contents in leaves The AA levels of leaves were measured by using UPLC-MS in 4-week-old Arabidopsis thaliana () (A) and 1-week-old Festuca elata () under well-watered, and drought stress conditions (50%, 25%, and 10% water contents in soil). Error bar means ±SD of biological triplicates.
7 A C / 35S:FePYR1 Control Air dry 1 μm AA Control Air dry 1 μm AA :FePYR1 :FePYR1 :FePYR1 :FePYR1 :FePYR1 :FePYR1 / 35S:FePYR1 / 35S:FePYR1 / 35S:FePYR1 D E Supplemental Figure 6. FePYR1 expression in the transgenic plants. (A, ) Quantitative measurement of FePYR1 transcript levels in rosette leaves from 21-day-old Arabidopsis transgenic plants compared to and. Error bars represent ±SD of biological triplicates. (C - E) The protein levels of FePYR1-3 FLAG detected with anti- FLAG antibody in 21-day-old rosette leaves of 35S:FePYR1:3 FLAG transgenic and (C), RD29Apro:FePYR1:3 FLAG transgenic (D) and (E), with or without indicated treatments for 24 hours. Upper panel, FePYR1-3 FLAG protein. Lower panel, loading control (Coomassie rilliant lue staining). The full-length images for the blots and gels shown here were presented in Supplemental Figure 8, 9 and 10, respectively.
8 / 35S: FePYR1:3 FLAG A #1 #2 #1 #2 / RD29Apro :FePYR1:3 FLAG C #1 #2 D :FePYR1:3 FLAG #1 #2 1 μm AA 0.5 μm AA 0 μm AA / 35S: FePYR1:3 FLAG 2 cm 2 cm 2 cm 2 cm Supplemental Figure 7. FePYR1 partially rescues AA insensitive phenotype of mutant during seed germination. (A, ) Germination assay of wild type (),, and 35S:FePYR1:3 FLAG transgenic plants grown in 0.5 MS medium containing 0, 0.5, or 1 μm AA. The representative plants were photographed after growing for 7 days. ar, 2 cm. (C, D) Seed germination assay of wild type (),, and RD29Apro:FePYR1:3 FLAG transgenic lines grown in 0.5 MS medium supplemented with 0, 0.5, or 1 μm AA. The representative plants grown for 7 days before being photographed. ar, 2cm.
9 A Protein ladder / 35S:FePYR1 / 35S:FePYR1 / 35S:FePYR1 / 35S:FePYR1 50 kd 37 kd 25 kd / 35S:FePYR1 / 35S:FePYR1 / 35S:FePYR1 / 35S:FePYR1 250 kd 150 kd 100 kd 75 kd 50 kd 37 kd 25 kd 20 kd 15 kd 10 kd Supplemental Figure 8. The protein levels of FePYR1-3 FLAG detected in 21-day-old rosette leaves of 35S:FePYR1:3 FLAG transgenic and. (A, ) The full-length blot(a) and gel(coomassie rilliant lue staining)() images for Fig S6C. The images were generated by using ChemiDoc TM XRS+ system in Image Lab TM Software (Molecular Imager, IO-RAD) with 60 seconds exposure time and bands auto-exposure mode, respectively. Empty means no sample loaded.
10 Control Air dry 1 μm AA A Protein ladder 50 kd 37 kd 25 kd Control Air dry 1 μm AA 250 kd 150 kd 100 kd 75 kd 50 kd 37 kd 25 kd 20 kd 15 kd Supplemental Figure 9. The protein levels of FePYR1-3 FLAG detected in 21-day-old rosette leaves of RD29Apro:FePYR1:3 FLAG transgenic plants (A, ) The full-length blot(a) and gel(coomassie rilliant lue staining)() images for Fig S6D. The plants were treated for 24 hours with or without indicated conditions. The images were generated by using ChemiDoc TM XRS+ system in Image Lab TM Software (Molecular Imager, IO-RAD) with 60 seconds exposure time and bands auto-exposure mode, respectively. Empty means no sample loaded.
11 Control Air dry 1 μm AA A Protein ladder :FePYR1 :FePYR1 :FePYR1 :FePYR1 :FePYR1 :FePYR1 50 kd 37 kd 25 kd Control Air dry 1 μm AA :FePYR1 :FePYR1 :FePYR1 :FePYR1 :FePYR1 :FePYR1 250 kd 150 kd 100 kd 75 kd 50 kd 37 kd 25 kd 20 kd 15 kd 10 kd Supplemental Figure 10. The protein levels of FePYR1-3 FLAG detected in 21-day-old rosette leaves of RD29Apro:FePYR1:3 FLAG transgenic plants. (A, ) The full-length blot(a) and gel(coomassie rilliant lue staining)() images for Fig S6E. The plants were treated for 24 hours with or without indicated conditions. The images were generated by using ChemiDoc TM XRS+ system in Image Lab TM Software (Molecular Imager, IO-RAD) with 60 seconds exposure time and bands auto-exposure mode, respectively. Empty means no sample loaded.
12 Table S1 Statistics of data sets and structure refinement Table S1. Statistics of data sets and structure refinement Space group P Resolution range, Å ( )* Cell parameters, Å, a=110.62, b=110.62, c=124.2; α=β=γ=90 Total/Unique reflections /20684 Completeness, % 96.3 (96.3) Mean I/σ 11.7 (1.3) Multiplicity 23.2 (21.9) Rmerge 0.29 (3.84) CC1/ (0.306) Refinement Resolution, Å No. reflections No. residues 578 No.solvent molecules 39 No. of non-h atoms 4418 Rcryst 22.8% Rfree 25.3% rmsd bonds, Å rmsd angles, 0.87 Average factor, Å *Values in the parentheses are for the highest resolution shell.
Supplemental Data. Wu et al. (2). Plant Cell..5/tpc RGLG Hormonal treatment H2O B RGLG µm ABA µm ACC µm GA Time (hours) µm µm MJ µm IA
 Supplemental Data. Wu et al. (2). Plant Cell..5/tpc..4. A B Supplemental Figure. Immunoblot analysis verifies the expression of the AD-PP2C and BD-RGLG proteins in the Y2H assay. Total proteins were extracted
Supplemental Data. Wu et al. (2). Plant Cell..5/tpc..4. A B Supplemental Figure. Immunoblot analysis verifies the expression of the AD-PP2C and BD-RGLG proteins in the Y2H assay. Total proteins were extracted
SUPPLEMENTARY INFORMATION. Small molecule activation of the TRAIL receptor DR5 in human cancer cells
 SUPPLEMENTARY INFORMATION Small molecule activation of the TRAIL receptor DR5 in human cancer cells Gelin Wang 1*, Xiaoming Wang 2, Hong Yu 1, Shuguang Wei 1, Noelle Williams 1, Daniel L. Holmes 1, Randal
SUPPLEMENTARY INFORMATION Small molecule activation of the TRAIL receptor DR5 in human cancer cells Gelin Wang 1*, Xiaoming Wang 2, Hong Yu 1, Shuguang Wei 1, Noelle Williams 1, Daniel L. Holmes 1, Randal
SUPPLEMENTAL FIGURE LEGENDS. Figure S1: Homology alignment of DDR2 amino acid sequence. Shown are
 SUPPLEMENTAL FIGURE LEGENDS Figure S1: Homology alignment of DDR2 amino acid sequence. Shown are the amino acid sequences of human DDR2, mouse DDR2 and the closest homologs in zebrafish and C. Elegans.
SUPPLEMENTAL FIGURE LEGENDS Figure S1: Homology alignment of DDR2 amino acid sequence. Shown are the amino acid sequences of human DDR2, mouse DDR2 and the closest homologs in zebrafish and C. Elegans.
Viral RNAi suppressor reversibly binds sirna to. outcompete Dicer and RISC via multiple-turnover
 Supplementary Data Viral RNAi suppressor reversibly binds sirna to outcompete Dicer and RISC via multiple-turnover Renata A. Rawlings 1,2, Vishalakshi Krishnan 2 and Nils G. Walter 2 * 1 Biophysics and
Supplementary Data Viral RNAi suppressor reversibly binds sirna to outcompete Dicer and RISC via multiple-turnover Renata A. Rawlings 1,2, Vishalakshi Krishnan 2 and Nils G. Walter 2 * 1 Biophysics and
Hossain_Supplemental Figure 1
 Hossain_Supplemental Figure 1 GFP-PACT GFP-PACT Motif I GFP-PACT Motif II A. MG132 (1µM) GFP Tubulin GFP-PACT Pericentrin GFP-PACT GFP-PACT Pericentrin Fig. S1. Expression and localization of Orc1 PACT
Hossain_Supplemental Figure 1 GFP-PACT GFP-PACT Motif I GFP-PACT Motif II A. MG132 (1µM) GFP Tubulin GFP-PACT Pericentrin GFP-PACT GFP-PACT Pericentrin Fig. S1. Expression and localization of Orc1 PACT
Supplementary Fig. S1. SAMHD1c has a more potent dntpase activity than. SAMHD1c. Purified recombinant SAMHD1c and SAMHD1c proteins (with
 Supplementary Fig. S1. SAMHD1c has a more potent dntpase activity than SAMHD1c. Purified recombinant SAMHD1c and SAMHD1c proteins (with concentration of 800nM) were incubated with 1mM dgtp for the indicated
Supplementary Fig. S1. SAMHD1c has a more potent dntpase activity than SAMHD1c. Purified recombinant SAMHD1c and SAMHD1c proteins (with concentration of 800nM) were incubated with 1mM dgtp for the indicated
Supplementary Material. TRIB3 inhibits proliferation and promotes osteogenesis in hbmscs by regulating the. ERK1/2 signaling pathway
 Supplementary Material TRIB3 inhibits proliferation and promotes osteogenesis in hbmscs by regulating the ERK1/2 signaling pathway Cui Zhang 1, Fan-Fan Hong 1, Cui-Cui Wang 1, Liang Li 1, Jian-Ling Chen
Supplementary Material TRIB3 inhibits proliferation and promotes osteogenesis in hbmscs by regulating the ERK1/2 signaling pathway Cui Zhang 1, Fan-Fan Hong 1, Cui-Cui Wang 1, Liang Li 1, Jian-Ling Chen
Supplemental Data. Steiner et al. Plant Cell. (2012) /tpc
 Supplemental Figure 1. SPY does not interact with free GST. Invitro pull-down assay using E. coli-expressed MBP-SPY and GST, GST-TCP14 and GST-TCP15. MBP-SPY was used as bait and incubated with equal amount
Supplemental Figure 1. SPY does not interact with free GST. Invitro pull-down assay using E. coli-expressed MBP-SPY and GST, GST-TCP14 and GST-TCP15. MBP-SPY was used as bait and incubated with equal amount
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION Conserved arginines on the rim of Hfq catalyze base pair formation and exchange Subrata Panja and Sarah A. Woodson T.C. Jenkins Department of Biophysics, Johns Hopkins University,
SUPPLEMENTARY INFORMATION Conserved arginines on the rim of Hfq catalyze base pair formation and exchange Subrata Panja and Sarah A. Woodson T.C. Jenkins Department of Biophysics, Johns Hopkins University,
Kinetics Review. Tonight at 7 PM Phys 204 We will do two problems on the board (additional ones than in the problem sets)
 Quiz 1 Kinetics Review Tonight at 7 PM Phys 204 We will do two problems on the board (additional ones than in the problem sets) I will post the problems with solutions on Toolkit for those that can t make
Quiz 1 Kinetics Review Tonight at 7 PM Phys 204 We will do two problems on the board (additional ones than in the problem sets) I will post the problems with solutions on Toolkit for those that can t make
Supplementary Table 1: List of CH3 domain interface residues in the first chain (A) and
 Supplementary Tables Supplementary Table 1: List of CH3 domain interface residues in the first chain (A) and their side chain contacting residues in the second chain (B) a Interface Res. in Contacting
Supplementary Tables Supplementary Table 1: List of CH3 domain interface residues in the first chain (A) and their side chain contacting residues in the second chain (B) a Interface Res. in Contacting
A subclass of HSP70s regulate development and abiotic stress responses in Arabidopsis thaliana
 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 Journal of Plant Research A subclass of HSP70s regulate development and abiotic stress responses in Arabidopsis thaliana Linna Leng 1 Qianqian Liang
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 Journal of Plant Research A subclass of HSP70s regulate development and abiotic stress responses in Arabidopsis thaliana Linna Leng 1 Qianqian Liang
A) B) Ladder. Supplementary Figure 1. Recombinant calpain 14 purification analysis. A) The general domain structure of classical
 A) B) Ladder C) r4 r4 Nt- -Ct 78 kda Supplementary Figure 1. Recombinant calpain 14 purification analysis. A) The general domain structure of classical calpains is shown. The protease core consists of
A) B) Ladder C) r4 r4 Nt- -Ct 78 kda Supplementary Figure 1. Recombinant calpain 14 purification analysis. A) The general domain structure of classical calpains is shown. The protease core consists of
Coleman et al., Supplementary Figure 1
 Coleman et al., Supplementary Figure 1 BrdU Merge G1 Early S Mid S Supplementary Figure 1. Sequential destruction of CRL4 Cdt2 targets during the G1/S transition. HCT116 cells were synchronized by sequential
Coleman et al., Supplementary Figure 1 BrdU Merge G1 Early S Mid S Supplementary Figure 1. Sequential destruction of CRL4 Cdt2 targets during the G1/S transition. HCT116 cells were synchronized by sequential
X-ray structures of fructosyl peptide oxidases revealing residues responsible for gating oxygen access in the oxidative half reaction
 X-ray structures of fructosyl peptide oxidases revealing residues responsible for gating oxygen access in the oxidative half reaction Tomohisa Shimasaki 1, Hiromi Yoshida 2, Shigehiro Kamitori 2 & Koji
X-ray structures of fructosyl peptide oxidases revealing residues responsible for gating oxygen access in the oxidative half reaction Tomohisa Shimasaki 1, Hiromi Yoshida 2, Shigehiro Kamitori 2 & Koji
Supplemental Data. Sun et al. Plant Cell. (2012) /tpc FLS2 -cmyc -GFP FLS2 -HA. FLS2-FLAG FLS2 -HA BAK1-HA flg22 (10mM)
 Supplemental Data. Sun et al. Plant ell. ()..5/tpc..9599 A FLS-FLA FLS -HA BAK-HA flg (mm) - B FLS -cmyc -FP FLS -HA IP antibody cmyc FLA FP cmyc IP ; WB: α-ha IP; WB: α-ha IP: α-fla; WB: α-ha IP ; WB:
Supplemental Data. Sun et al. Plant ell. ()..5/tpc..9599 A FLS-FLA FLS -HA BAK-HA flg (mm) - B FLS -cmyc -FP FLS -HA IP antibody cmyc FLA FP cmyc IP ; WB: α-ha IP; WB: α-ha IP: α-fla; WB: α-ha IP ; WB:
Supplemental Data. Lee et al. Plant Cell. (2010) /tpc Supplemental Figure 1. Protein and Gene Structures of DWA1 and DWA2.
 Supplemental Figure 1. Protein and Gene Structures of DWA1 and DWA2. (A) Protein structures of DWA1 and DWA2. WD40 region was determined based on the NCBI conserved domain databases (B, C) Schematic representation
Supplemental Figure 1. Protein and Gene Structures of DWA1 and DWA2. (A) Protein structures of DWA1 and DWA2. WD40 region was determined based on the NCBI conserved domain databases (B, C) Schematic representation
Supplementary Figure S1. Immunodetection of full-length XA21 and the XA21 C-terminal cleavage product.
 Supplementary Information Supplementary Figure S1. Immunodetection of full-length XA21 and the XA21 C-terminal cleavage product. Total protein extracted from Kitaake wild type and rice plants carrying
Supplementary Information Supplementary Figure S1. Immunodetection of full-length XA21 and the XA21 C-terminal cleavage product. Total protein extracted from Kitaake wild type and rice plants carrying
Randomly arrayed G-rich DNA sequence for label-free and realtime. assay of enzyme activity
 Electronic Supplementary Material (ESI) for ChemComm. This journal is The Royal Society of Chemistry 2014 Randomly arrayed G-rich DNA sequence for label-free and realtime assay of enzyme activity Zhuoliang
Electronic Supplementary Material (ESI) for ChemComm. This journal is The Royal Society of Chemistry 2014 Randomly arrayed G-rich DNA sequence for label-free and realtime assay of enzyme activity Zhuoliang
BIOC 463A Expt. 4: Column Chromatographic Methods Column Chromatography
 Column Chromatography Chromatography is the process use to separate molecules based on SOME physical property of the molecule: Mass (i.e. size) Charge Affinity for ligands or substrates Hydrophobic interactions
Column Chromatography Chromatography is the process use to separate molecules based on SOME physical property of the molecule: Mass (i.e. size) Charge Affinity for ligands or substrates Hydrophobic interactions
Purification of Lactate Dehydrogenase
 Dominican University of California Dominican Scholar Scholarly & Creative Works Conference 2018 Scholarly and Creative Works Conference 2016 Apr 15th, 1:30 PM - 2:00 PM Purification of Lactate Dehydrogenase
Dominican University of California Dominican Scholar Scholarly & Creative Works Conference 2018 Scholarly and Creative Works Conference 2016 Apr 15th, 1:30 PM - 2:00 PM Purification of Lactate Dehydrogenase
6-Foot Mini Toober Activity
 Big Idea The interaction between the substrate and enzyme is highly specific. Even a slight change in shape of either the substrate or the enzyme may alter the efficient and selective ability of the enzyme
Big Idea The interaction between the substrate and enzyme is highly specific. Even a slight change in shape of either the substrate or the enzyme may alter the efficient and selective ability of the enzyme
Electronic Supplementary Information
 Electronic Supplementary Material (ESI) for Molecular BioSystems. This journal is The Royal Society of Chemistry 2017 Electronic Supplementary Information Dissecting binding of a β-barrel outer membrane
Electronic Supplementary Material (ESI) for Molecular BioSystems. This journal is The Royal Society of Chemistry 2017 Electronic Supplementary Information Dissecting binding of a β-barrel outer membrane
GTTCGGGTTCC TTTTGAGCAG
 Supplementary Figures Splice variants of the SIP1 transcripts play a role in nodule organogenesis in Lotus japonicus. Wang C, Zhu H, Jin L, Chen T, Wang L, Kang H, Hong Z, Zhang Z. 5 UTR CDS 3 UTR TCTCAACCATCCTTTGTCTGCTTCCGCCGCATGGGTGAGGTCATTTTGTCTAGATGACGTGCAATTTACAATGA
Supplementary Figures Splice variants of the SIP1 transcripts play a role in nodule organogenesis in Lotus japonicus. Wang C, Zhu H, Jin L, Chen T, Wang L, Kang H, Hong Z, Zhang Z. 5 UTR CDS 3 UTR TCTCAACCATCCTTTGTCTGCTTCCGCCGCATGGGTGAGGTCATTTTGTCTAGATGACGTGCAATTTACAATGA
Electrophoresis and transfer
 Electrophoresis and transfer Electrophoresis Cation = positively charged ion, it moves toward the cathode (-) Anion = negatively charged ion, it moves toward the anode (+) Amphoteric substance = can have
Electrophoresis and transfer Electrophoresis Cation = positively charged ion, it moves toward the cathode (-) Anion = negatively charged ion, it moves toward the anode (+) Amphoteric substance = can have
Product. Ni-NTA His Bind Resin. Ni-NTA His Bind Superflow. His Bind Resin. His Bind Magnetic Agarose Beads. His Bind Column. His Bind Quick Resin
 Novagen offers a large variety of affinity supports and kits for the purification of recombinant proteins containing popular peptide fusion tags, including His Tag, GST Tag, S Tag and T7 Tag sequences.
Novagen offers a large variety of affinity supports and kits for the purification of recombinant proteins containing popular peptide fusion tags, including His Tag, GST Tag, S Tag and T7 Tag sequences.
Chemical hijacking of auxin signaling with an engineered auxin-tir1
 1 SUPPLEMENTARY INFORMATION Chemical hijacking of auxin signaling with an engineered auxin-tir1 pair Naoyuki Uchida 1,2*, Koji Takahashi 2*, Rie Iwasaki 1, Ryotaro Yamada 2, Masahiko Yoshimura 2, Takaho
1 SUPPLEMENTARY INFORMATION Chemical hijacking of auxin signaling with an engineered auxin-tir1 pair Naoyuki Uchida 1,2*, Koji Takahashi 2*, Rie Iwasaki 1, Ryotaro Yamada 2, Masahiko Yoshimura 2, Takaho
Stabilization of a virus-like particle and its application as a nanoreactor at physiological conditions
 Supporting Information Stabilization of a virus-like particle and its application as a nanoreactor at physiological conditions Lise Schoonen, b Sjors Maassen, b Roeland J. M. Nolte b and Jan C. M. van
Supporting Information Stabilization of a virus-like particle and its application as a nanoreactor at physiological conditions Lise Schoonen, b Sjors Maassen, b Roeland J. M. Nolte b and Jan C. M. van
Nucleic acids. How DNA works. DNA RNA Protein. DNA (deoxyribonucleic acid) RNA (ribonucleic acid) Central Dogma of Molecular Biology
 Nucleic acid chemistry and basic molecular theory Nucleic acids DNA (deoxyribonucleic acid) RNA (ribonucleic acid) Central Dogma of Molecular Biology Cell cycle DNA RNA Protein Transcription Translation
Nucleic acid chemistry and basic molecular theory Nucleic acids DNA (deoxyribonucleic acid) RNA (ribonucleic acid) Central Dogma of Molecular Biology Cell cycle DNA RNA Protein Transcription Translation
FRAUNHOFER IME SCREENINGPORT
 FRAUNHOFER IME SCREENINGPORT Detection technologies used in drug discovery Introduction Detection technologies in drug discovery is unlimited only a biased snapshot can be presented Differences can presented
FRAUNHOFER IME SCREENINGPORT Detection technologies used in drug discovery Introduction Detection technologies in drug discovery is unlimited only a biased snapshot can be presented Differences can presented
INSECT CELL/BACULOVIRUS PRODUCTION
 INSECT CELL/BACULOVIRUS PRODUCTION PEF # GENE NAME TRANSFER VECTOR BEVS MOLECULAR WEIGHT 2015-XXXX XXXX pbac1 flashbacultra TM 36.0 kda EXPRESSION METHOD OVERVIEW: Insect cells Spodoptera frugiperda (Sf9)
INSECT CELL/BACULOVIRUS PRODUCTION PEF # GENE NAME TRANSFER VECTOR BEVS MOLECULAR WEIGHT 2015-XXXX XXXX pbac1 flashbacultra TM 36.0 kda EXPRESSION METHOD OVERVIEW: Insect cells Spodoptera frugiperda (Sf9)
Supplementary information for. An Ultrasensitive Biosensor for DNA Detection Based on. Hybridization Chain Reaction Coupled with the Efficient
 Supplementary information for An Ultrasensitive Biosensor for DNA Detection Based on Hybridization Chain Reaction Coupled with the Efficient Quenching of Ruthenium Complex to CdTe Quantum Dot Yufei Liu,
Supplementary information for An Ultrasensitive Biosensor for DNA Detection Based on Hybridization Chain Reaction Coupled with the Efficient Quenching of Ruthenium Complex to CdTe Quantum Dot Yufei Liu,
Control of secondary cell wall patterning involves xylan deacetylation by a GDSL esterase
 In the format provided by the authors and unedited. SUPPLEMENTARY INFORMATION VOLUME: 3 ARTICLE NUMBER: 17017 Control of secondary cell wall patterning involves xylan deacetylation by a GDSL esterase Baocai
In the format provided by the authors and unedited. SUPPLEMENTARY INFORMATION VOLUME: 3 ARTICLE NUMBER: 17017 Control of secondary cell wall patterning involves xylan deacetylation by a GDSL esterase Baocai
Efficient Multi-Well Protein Purification Strategies
 Application Note PN 33576 Efficient Multi-Well Protein Purification Strategies Introduction Many tools and techniques are available today for protein purification. Development of a purification process
Application Note PN 33576 Efficient Multi-Well Protein Purification Strategies Introduction Many tools and techniques are available today for protein purification. Development of a purification process
SUPPLEMENTAL MATERIALS
 SUPPLEMENL MERILS Eh-seq: RISPR epitope tagging hip-seq of DN-binding proteins Daniel Savic, E. hristopher Partridge, Kimberly M. Newberry, Sophia. Smith, Sarah K. Meadows, rian S. Roberts, Mark Mackiewicz,
SUPPLEMENL MERILS Eh-seq: RISPR epitope tagging hip-seq of DN-binding proteins Daniel Savic, E. hristopher Partridge, Kimberly M. Newberry, Sophia. Smith, Sarah K. Meadows, rian S. Roberts, Mark Mackiewicz,
Strep-tag Technology for Molecular Weight (MW) Determinations on Blots Using Precision Plus Protein Standards
 blotting tech note 2847 Strep-tag Technology for Molecular Weight (MW) Determinations on Blots Using Precision Plus Protein Standards Introduction Bio-Rad has consistently provided innovative standards
blotting tech note 2847 Strep-tag Technology for Molecular Weight (MW) Determinations on Blots Using Precision Plus Protein Standards Introduction Bio-Rad has consistently provided innovative standards
Supplemental Data. Osakabe et al. (2013). Plant Cell /tpc
 Supplemental Figure 1. Phylogenetic analysis of KUP in various species. The amino acid sequences of KUPs from green algae and land plants were identified with a BLAST search and aligned using ClustalW
Supplemental Figure 1. Phylogenetic analysis of KUP in various species. The amino acid sequences of KUPs from green algae and land plants were identified with a BLAST search and aligned using ClustalW
Nature Structural and Molecular Biology: doi: /nsmb.2937
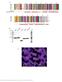 Supplementary Figure 1 Multiple sequence alignment of the CtIP N-terminal domain, purified CtIP protein constructs and details of the 2F o F c electron density map of CtIP-NTD. (a) Multiple sequence alignment,
Supplementary Figure 1 Multiple sequence alignment of the CtIP N-terminal domain, purified CtIP protein constructs and details of the 2F o F c electron density map of CtIP-NTD. (a) Multiple sequence alignment,
Appendix IV Version
 APPENDIX IV. Gel Electrophoresis. Migration of biological molecules in the presence of an electric field through a gel matrix is the heart of many biochemistry experiments. The variety of electrophoresis
APPENDIX IV. Gel Electrophoresis. Migration of biological molecules in the presence of an electric field through a gel matrix is the heart of many biochemistry experiments. The variety of electrophoresis
Case 7 A Storage Protein From Seeds of Brassica nigra is a Serine Protease Inhibitor Last modified 29 September 2005
 Case 7 A Storage Protein From Seeds of Brassica nigra is a Serine Protease Inhibitor Last modified 9 September 005 Focus concept Purification of a novel seed storage protein allows sequence analysis and
Case 7 A Storage Protein From Seeds of Brassica nigra is a Serine Protease Inhibitor Last modified 9 September 005 Focus concept Purification of a novel seed storage protein allows sequence analysis and
ARBRE-P4EU Consensus Protein Quality Guidelines for Biophysical and Biochemical Studies Minimal information to provide
 ARBRE-P4EU Consensus Protein Quality Guidelines for Biophysical and Biochemical Studies Minimal information to provide Protein name and full primary structure, by providing a NCBI (or UniProt) accession
ARBRE-P4EU Consensus Protein Quality Guidelines for Biophysical and Biochemical Studies Minimal information to provide Protein name and full primary structure, by providing a NCBI (or UniProt) accession
Supplementary information, Figure S1
 Supplementary information, Figure S1 (A) Schematic diagram of the sgrna and hspcas9 expression cassettes in a single binary vector designed for Agrobacterium-mediated stable transformation of Arabidopsis
Supplementary information, Figure S1 (A) Schematic diagram of the sgrna and hspcas9 expression cassettes in a single binary vector designed for Agrobacterium-mediated stable transformation of Arabidopsis
Head of College Scholars List Scheme. Summer Studentship. Report Form
 Head of College Scholars List Scheme Summer Studentship Report Form This report should be completed by the student with his/her project supervisor. It should summarise the work undertaken during the project
Head of College Scholars List Scheme Summer Studentship Report Form This report should be completed by the student with his/her project supervisor. It should summarise the work undertaken during the project
Purification: Step 1. Lecture 11 Protein and Peptide Chemistry. Cells: Break them open! Crude Extract
 Purification: Step 1 Lecture 11 Protein and Peptide Chemistry Cells: Break them open! Crude Extract Total contents of cell Margaret A. Daugherty Fall 2003 Big Problem: Crude extract is not the natural
Purification: Step 1 Lecture 11 Protein and Peptide Chemistry Cells: Break them open! Crude Extract Total contents of cell Margaret A. Daugherty Fall 2003 Big Problem: Crude extract is not the natural
Purification: Step 1. Protein and Peptide Chemistry. Lecture 11. Big Problem: Crude extract is not the natural environment. Cells: Break them open!
 Lecture 11 Protein and Peptide Chemistry Margaret A. Daugherty Fall 2003 Purification: Step 1 Cells: Break them open! Crude Extract Total contents of cell Big Problem: Crude extract is not the natural
Lecture 11 Protein and Peptide Chemistry Margaret A. Daugherty Fall 2003 Purification: Step 1 Cells: Break them open! Crude Extract Total contents of cell Big Problem: Crude extract is not the natural
T H E J O U R N A L O F C E L L B I O L O G Y
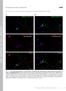 T H E J O U R N A L O F C E L L B I O L O G Y Supplemental material Kanaani et al., http://www.jcb.org/cgi/content/full/jcb.200912101/dc1 Figure S1. The K2 rabbit polyclonal antibody is specific for GAD67,
T H E J O U R N A L O F C E L L B I O L O G Y Supplemental material Kanaani et al., http://www.jcb.org/cgi/content/full/jcb.200912101/dc1 Figure S1. The K2 rabbit polyclonal antibody is specific for GAD67,
Supplementary Table 1. The Q-PCR primer sequence is summarized in the following table.
 Supplementary Table 1. The Q-PCR primer sequence is summarized in the following table. Name Sequence (5-3 ) Application Flag-u ggactacaaggacgacgatgac Shared upstream primer for all the amplifications of
Supplementary Table 1. The Q-PCR primer sequence is summarized in the following table. Name Sequence (5-3 ) Application Flag-u ggactacaaggacgacgatgac Shared upstream primer for all the amplifications of
ENCODE DCC Antibody Validation Document
 ENCODE DCC Antibody Validation Document Date of Submission 09/12/12 Name: Trupti Kawli Email: trupti@stanford.edu Lab Snyder Antibody Name: SREBP1 (sc-8984) Target: SREBP1 Company/ Source: Santa Cruz Biotechnology
ENCODE DCC Antibody Validation Document Date of Submission 09/12/12 Name: Trupti Kawli Email: trupti@stanford.edu Lab Snyder Antibody Name: SREBP1 (sc-8984) Target: SREBP1 Company/ Source: Santa Cruz Biotechnology
Supplementary Figure 1 Two distinct conformational states of the HNH domain in crystal structures. a
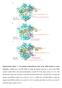 Supplementary Figure 1 Two distinct conformational states of the HNH domain in crystal structures. a HNH-state 1 in PDB 4OO8, in which the distance from the C atom of the HNH catalytic residue 840 to the
Supplementary Figure 1 Two distinct conformational states of the HNH domain in crystal structures. a HNH-state 1 in PDB 4OO8, in which the distance from the C atom of the HNH catalytic residue 840 to the
Purification of Recombinant Proteins on Nuvia TM cprime TM Hydrophobic Cation Exchange Media: A Simple Approach to Method Development
 Purification of Recombinant Proteins on Nuvia TM cprime TM Hydrophobic Cation Exchange Media: A Simple Approach to Method Development Xuemei M. He, Sherif Hanala, and Mark Snyder, Bio-Rad Laboratories,
Purification of Recombinant Proteins on Nuvia TM cprime TM Hydrophobic Cation Exchange Media: A Simple Approach to Method Development Xuemei M. He, Sherif Hanala, and Mark Snyder, Bio-Rad Laboratories,
ASPP1 Fw GGTTGGGAATCCACGTGTTG ASPP1 Rv GCCATATCTTGGAGCTCTGAGAG
 Supplemental Materials and Methods Plasmids: the following plasmids were used in the supplementary data: pwzl-myc- Lats2 (Aylon et al, 2006), pretrosuper-vector and pretrosuper-shp53 (generous gift of
Supplemental Materials and Methods Plasmids: the following plasmids were used in the supplementary data: pwzl-myc- Lats2 (Aylon et al, 2006), pretrosuper-vector and pretrosuper-shp53 (generous gift of
Small-Molecule Drug Target Identification/Deconvolution Technologies
 Small-Molecule Drug Target Identification/Deconvolution Technologies Case-Studies Shantani Target ID Technology Tool Box Target Deconvolution is not Trivial = A single Tool / Technology May Not necessarily
Small-Molecule Drug Target Identification/Deconvolution Technologies Case-Studies Shantani Target ID Technology Tool Box Target Deconvolution is not Trivial = A single Tool / Technology May Not necessarily
Figure 1: TDP-43 is subject to lysine acetylation within the RNA-binding domain a) QBI-293 cells were transfected with TDP-43 in the presence or
 Figure 1: TDP-43 is subject to lysine acetylation within the RNA-binding domain a) QBI-293 cells were transfected with TDP-43 in the presence or absence of the acetyltransferase CBP and acetylated TDP-43
Figure 1: TDP-43 is subject to lysine acetylation within the RNA-binding domain a) QBI-293 cells were transfected with TDP-43 in the presence or absence of the acetyltransferase CBP and acetylated TDP-43
Drug Discovery Research Clinical Screening. Comparison of ELISA and AlphaScreen Assay Technologies for Measurement of Protein Expression Levels
 Drug Discovery Research Clinical Screening FLAG Expression Measurement Application Note Comparison of ELISA and AlphaScreen Assay Technologies for Measurement of Protein Expression Levels ALPHASCREEN APPLICATION
Drug Discovery Research Clinical Screening FLAG Expression Measurement Application Note Comparison of ELISA and AlphaScreen Assay Technologies for Measurement of Protein Expression Levels ALPHASCREEN APPLICATION
NPTEL VIDEO COURSE PROTEOMICS PROF. SANJEEVA SRIVASTAVA
 LECTURE-06 PROTEIN PURIFICATION AND PEPTIDE ISOLATION USING CHROMATOGRAPHY TRANSCRIPT Welcome to the proteomics course. Today, we will talk about protein purification and peptide isolation using chromatography
LECTURE-06 PROTEIN PURIFICATION AND PEPTIDE ISOLATION USING CHROMATOGRAPHY TRANSCRIPT Welcome to the proteomics course. Today, we will talk about protein purification and peptide isolation using chromatography
Structural Bioinformatics (C3210) DNA and RNA Structure
 Structural Bioinformatics (C3210) DNA and RNA Structure Importance of DNA/RNA 3D Structure Nucleic acids are essential materials found in all living organisms. Their main function is to maintain and transmit
Structural Bioinformatics (C3210) DNA and RNA Structure Importance of DNA/RNA 3D Structure Nucleic acids are essential materials found in all living organisms. Their main function is to maintain and transmit
BIRKBECK COLLEGE (University of London)
 BIRKBECK COLLEGE (University of London) SCHOOL OF BIOLOGICAL SCIENCES M.Sc. EXAMINATION FOR INTERNAL STUDENTS ON: Postgraduate Certificate in Principles of Protein Structure MSc Structural Molecular Biology
BIRKBECK COLLEGE (University of London) SCHOOL OF BIOLOGICAL SCIENCES M.Sc. EXAMINATION FOR INTERNAL STUDENTS ON: Postgraduate Certificate in Principles of Protein Structure MSc Structural Molecular Biology
Electro refers to electron flow or current. Thus Electrophoresis is movement under electric current.
 ELECTROPHORESIS Electrophoresis Electro refers to electron flow or current. Phoresis refers to movement. Thus Electrophoresis is movement under electric current. This technique therefore can separate molecules
ELECTROPHORESIS Electrophoresis Electro refers to electron flow or current. Phoresis refers to movement. Thus Electrophoresis is movement under electric current. This technique therefore can separate molecules
Supplementary Figure 1: Two modes of low concentration of BsSMC on a DNA (a) Protein staining (left) and fluorescent imaging of Cy3 (right) confirm
 Supplementary Figure 1: Two modes of low concentration of BsSMC on a DNA (a) Protein staining (left) and fluorescent imaging of Cy3 (right) confirm that BsSMC was labeled with Cy3 NHS-Ester. In each panel,
Supplementary Figure 1: Two modes of low concentration of BsSMC on a DNA (a) Protein staining (left) and fluorescent imaging of Cy3 (right) confirm that BsSMC was labeled with Cy3 NHS-Ester. In each panel,
Notes to accompany the slidecast on theory of SDS PAGE and Western blotting
 S317 Biological science: from genes to species Notes to accompany the slidecast on theory of SDS PAGE and Western blotting SDS PAGE SDS PAGE is a standard technique for determining the molecular size of
S317 Biological science: from genes to species Notes to accompany the slidecast on theory of SDS PAGE and Western blotting SDS PAGE SDS PAGE is a standard technique for determining the molecular size of
The microtubule-associated tau protein has intrinsic acetyltransferase activity. Todd J. Cohen, Dave Friedmann, Andrew W. Hwang, Ronen Marmorstein and
 SUPPLEMENTARY INFORMATION: The microtubule-associated tau protein has intrinsic acetyltransferase activity Todd J. Cohen, Dave Friedmann, Andrew W. Hwang, Ronen Marmorstein and Virginia M.Y. Lee Cohen
SUPPLEMENTARY INFORMATION: The microtubule-associated tau protein has intrinsic acetyltransferase activity Todd J. Cohen, Dave Friedmann, Andrew W. Hwang, Ronen Marmorstein and Virginia M.Y. Lee Cohen
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION A human XRCC4-XLF complex bridges DNA ends. Sara N. Andres 1, Alexandra Vergnes 2, Dejan Ristic 3, Claire Wyman 3, Mauro Modesti 2,4, and Murray Junop 2,4 1 Department of Biochemistry
SUPPLEMENTARY INFORMATION A human XRCC4-XLF complex bridges DNA ends. Sara N. Andres 1, Alexandra Vergnes 2, Dejan Ristic 3, Claire Wyman 3, Mauro Modesti 2,4, and Murray Junop 2,4 1 Department of Biochemistry
Lecture 25: Introduction to Chromatography and Gel Filtration
 Biological Chemistry Laboratory Biology 3515/Chemistry 3515 Spring 2018 Lecture 25: Introduction to Chromatography and Gel Filtration 10 April 2018 c David P. Goldenberg University of Utah goldenberg@biology.utah.edu
Biological Chemistry Laboratory Biology 3515/Chemistry 3515 Spring 2018 Lecture 25: Introduction to Chromatography and Gel Filtration 10 April 2018 c David P. Goldenberg University of Utah goldenberg@biology.utah.edu
Masayoshi Honda, Jeehae Park, Robert A. Pugh, Taekjip Ha, and Maria Spies
 Molecular Cell, Volume 35 Supplemental Data Single-Molecule Analysis Reveals Differential Effect of ssdna-binding Proteins on DNA Translocation by XPD Helicase Masayoshi Honda, Jeehae Park, Robert A. Pugh,
Molecular Cell, Volume 35 Supplemental Data Single-Molecule Analysis Reveals Differential Effect of ssdna-binding Proteins on DNA Translocation by XPD Helicase Masayoshi Honda, Jeehae Park, Robert A. Pugh,
Jung-Nam Cho, Jee-Youn Ryu, Young-Min Jeong, Jihye Park, Ji-Joon Song, Richard M. Amasino, Bosl Noh, and Yoo-Sun Noh
 Developmental Cell, Volume 22 Supplemental Information Control of Seed Germination by Light-Induced Histone Arginine Demethylation Activity Jung-Nam Cho, Jee-Youn Ryu, Young-Min Jeong, Jihye Park, Ji-Joon
Developmental Cell, Volume 22 Supplemental Information Control of Seed Germination by Light-Induced Histone Arginine Demethylation Activity Jung-Nam Cho, Jee-Youn Ryu, Young-Min Jeong, Jihye Park, Ji-Joon
Supplementary materials. Computational study of β-n-acetylhexosaminidase from. Talaromyces flavus, a glycosidase with high substrate flexibility
 Supplementary materials Computational study of β-n-acetylhexosaminidase from Talaromyces flavus, a glycosidase with high substrate flexibility Natallia Kulik 1,*, Kristýna Slámová 2, Rüdiger Ettrich 1,3,
Supplementary materials Computational study of β-n-acetylhexosaminidase from Talaromyces flavus, a glycosidase with high substrate flexibility Natallia Kulik 1,*, Kristýna Slámová 2, Rüdiger Ettrich 1,3,
CBI Toolbox Tour 2015
 CBI Toolbox Tour 2015 Thermophoresis (NanoTemper) NT.115 & NT.LabelFree Images: NanoTemper Circular Dichroism Jasco J-1500 Spectrometer Six Position Turreted Peltier Temperature Control System Automated
CBI Toolbox Tour 2015 Thermophoresis (NanoTemper) NT.115 & NT.LabelFree Images: NanoTemper Circular Dichroism Jasco J-1500 Spectrometer Six Position Turreted Peltier Temperature Control System Automated
Size Exclusion BioHPLC Columns
 Size Exclusion BioHPLC Columns Size Exclusion Product Families Particle Porosity Functionalities Particle Pore Size Application Sizes Agilent Bio SEC- Silica Fully porous N/A um 00A, 0A, 00A High efficiency
Size Exclusion BioHPLC Columns Size Exclusion Product Families Particle Porosity Functionalities Particle Pore Size Application Sizes Agilent Bio SEC- Silica Fully porous N/A um 00A, 0A, 00A High efficiency
42 fl organelles = 34.5 fl (1) 3.5X X 0.93 = 78,000 (2)
 SUPPLEMENTAL DATA Supplementary Experimental Procedures Fluorescence Microscopy - A Zeiss Axiovert 200M microscope equipped with a Zeiss 100x Plan- Apochromat (1.40 NA) DIC objective and Hamamatsu Orca
SUPPLEMENTAL DATA Supplementary Experimental Procedures Fluorescence Microscopy - A Zeiss Axiovert 200M microscope equipped with a Zeiss 100x Plan- Apochromat (1.40 NA) DIC objective and Hamamatsu Orca
MSD Immuno-Dot-Blot Assays. A division of Meso Scale Diagnostics, LLC.
 MSD Immuno-Dot-Blot Assays Example: High Throughput Western Blots Replacements Traditional Western Blots High content Molecular weight and immunoreactivity Labor and protein intensive Inherently low throughput
MSD Immuno-Dot-Blot Assays Example: High Throughput Western Blots Replacements Traditional Western Blots High content Molecular weight and immunoreactivity Labor and protein intensive Inherently low throughput
pgbkt7 Anti- Myc AH109 strain (KDa) 50
 pgbkt7 (KDa) 50 37 Anti- Myc AH109 strain Supplementary Figure 1. Protein expression of CRN and TDR in yeast. To analyse the protein expression of CRNKD and TDRKD, total proteins extracted from yeast culture
pgbkt7 (KDa) 50 37 Anti- Myc AH109 strain Supplementary Figure 1. Protein expression of CRN and TDR in yeast. To analyse the protein expression of CRNKD and TDRKD, total proteins extracted from yeast culture
Supplemental Information. Role of phosphatase of regenerating liver 1 (PRL1) in spermatogenesis
 Supplemental Information Role of phosphatase of regenerating liver 1 (PRL1) in spermatogenesis Yunpeng Bai ;, Lujuan Zhang #, Hongming Zhou #, Yuanshu Dong #, Qi Zeng, Weinian Shou, and Zhong-Yin Zhang
Supplemental Information Role of phosphatase of regenerating liver 1 (PRL1) in spermatogenesis Yunpeng Bai ;, Lujuan Zhang #, Hongming Zhou #, Yuanshu Dong #, Qi Zeng, Weinian Shou, and Zhong-Yin Zhang
Supplementary Figure 1. BES1 specifically inhibits ABA responses in early seedling
 Supplementary Figure 1. BES1 specifically inhibits ABA responses in early seedling development. a. Exogenous BR application overcomes the hypersensitivity of bzr1-1d seedlings to ABA. Seed germination
Supplementary Figure 1. BES1 specifically inhibits ABA responses in early seedling development. a. Exogenous BR application overcomes the hypersensitivity of bzr1-1d seedlings to ABA. Seed germination
MOLEBIO LAB #3: Electrophoretic Separation of Proteins
 MOLEBIO LAB #3: Electrophoretic Separation of Proteins Introduction: Proteins occupy a central position in the structure and function of all living organisms. Some proteins serve as structural components
MOLEBIO LAB #3: Electrophoretic Separation of Proteins Introduction: Proteins occupy a central position in the structure and function of all living organisms. Some proteins serve as structural components
electrophoresis tech Methods Materials
 electrophoresis tech note 3176 Monitoring the Expression, Purification, and Processing of -Tagged Proteins Using the Experion Automated Electrophoresis System Xuemei He and William Strong, Bio-Rad Laboratories,
electrophoresis tech note 3176 Monitoring the Expression, Purification, and Processing of -Tagged Proteins Using the Experion Automated Electrophoresis System Xuemei He and William Strong, Bio-Rad Laboratories,
MBMB451A Section1 Fall 2008 KEY These questions may have more than one correct answer
 MBMB451A Section1 Fall 2008 KEY These questions may have more than one correct answer 1. In a double stranded molecule of DNA, the ratio of purines : pyrimidines is (a) variable (b) determined by the base
MBMB451A Section1 Fall 2008 KEY These questions may have more than one correct answer 1. In a double stranded molecule of DNA, the ratio of purines : pyrimidines is (a) variable (b) determined by the base
The practical task of monoclonal IgG purification with CHT TM ceramic hydroxyapatite
 The practical task of monoclonal IgG purification with CHT TM ceramic hydroxyapatite Pete Gagnon, Jie He, Paul Ng, Julia Zhen, Cheryl Aberin, Heather Mekosh 11th Annual Waterside Conference, Chicago, May
The practical task of monoclonal IgG purification with CHT TM ceramic hydroxyapatite Pete Gagnon, Jie He, Paul Ng, Julia Zhen, Cheryl Aberin, Heather Mekosh 11th Annual Waterside Conference, Chicago, May
SDS-PAGE and Western Blot. Molecular Basis of Evolution
 1 SDS-PAGE and Western Blot Molecular Basis of Evolution Homology high level of DNA and protein sequence similarity due to common ancestry. Evidence Genomes of related organisms are very similar. Even
1 SDS-PAGE and Western Blot Molecular Basis of Evolution Homology high level of DNA and protein sequence similarity due to common ancestry. Evidence Genomes of related organisms are very similar. Even
Supplementary Figure 1 qrt-pcr expression analysis of NLP8 with and without KNO 3 during germination.
 Supplementary Figure 1 qrt-pcr expression analysis of NLP8 with and without KNO 3 during germination. Seeds of Col-0 were harvested from plants grown at 16 C, stored for 2 months, imbibed for indicated
Supplementary Figure 1 qrt-pcr expression analysis of NLP8 with and without KNO 3 during germination. Seeds of Col-0 were harvested from plants grown at 16 C, stored for 2 months, imbibed for indicated
Purification of alpha-1 antitrypsin using an antibody based affinity chromatography medium
 Purification of alpha-1 antitrypsin using an antibody based affinity chromatography medium Ulrika Meyer a, Hanna Wlad a, Sven Blokland b, Frank J.M. Detmers b and Henrik Ihre a a GE Healthcare Bio-Sciences
Purification of alpha-1 antitrypsin using an antibody based affinity chromatography medium Ulrika Meyer a, Hanna Wlad a, Sven Blokland b, Frank J.M. Detmers b and Henrik Ihre a a GE Healthcare Bio-Sciences
Nanobiotechnology. Place: IOP 1 st Meeting Room Time: 9:30-12:00. Reference: Review Papers. Grade: 50% midterm, 50% final.
 Nanobiotechnology Place: IOP 1 st Meeting Room Time: 9:30-12:00 Reference: Review Papers Grade: 50% midterm, 50% final Midterm: 5/15 History Atom Earth, Air, Water Fire SEM: 20-40 nm Silver 66.2% Gold
Nanobiotechnology Place: IOP 1 st Meeting Room Time: 9:30-12:00 Reference: Review Papers Grade: 50% midterm, 50% final Midterm: 5/15 History Atom Earth, Air, Water Fire SEM: 20-40 nm Silver 66.2% Gold
Technical tips Session 5
 Technical tips Session 5 Chromatine Immunoprecipitation (ChIP): This is a powerful in vivo method to quantitate interaction of proteins associated with specific regions of the genome. It involves the immunoprecipitation
Technical tips Session 5 Chromatine Immunoprecipitation (ChIP): This is a powerful in vivo method to quantitate interaction of proteins associated with specific regions of the genome. It involves the immunoprecipitation
Lecture 23: SDS Gel Electrophoresis and Stacking Gels
 Biological Chemistry Laboratory Biology 3515/Chemistry 3515 Spring 2018 Lecture 23: SDS Gel Electrophoresis and Stacking Gels 3 April 2018 c David P. Goldenberg University of Utah goldenberg@biology.utah.edu
Biological Chemistry Laboratory Biology 3515/Chemistry 3515 Spring 2018 Lecture 23: SDS Gel Electrophoresis and Stacking Gels 3 April 2018 c David P. Goldenberg University of Utah goldenberg@biology.utah.edu
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION Dynamic Phosphorylation of HP1 Regulates Mitotic Progression in Human Cells Supplementary Figures Supplementary Figure 1. NDR1 interacts with HP1. (a) Immunoprecipitation using
SUPPLEMENTARY INFORMATION Dynamic Phosphorylation of HP1 Regulates Mitotic Progression in Human Cells Supplementary Figures Supplementary Figure 1. NDR1 interacts with HP1. (a) Immunoprecipitation using
Strep-tag detection in Western blots
 Strep-tag detection in Western blots General protocol for the detection of Strep-tag fusion proteins Last date of revision April 2012 Version PR07-0010 www.strep-tag.com For research use only Important
Strep-tag detection in Western blots General protocol for the detection of Strep-tag fusion proteins Last date of revision April 2012 Version PR07-0010 www.strep-tag.com For research use only Important
PHT1;2-CFP YFP-PHF + PHT1;2-CFP YFP-PHF
 YFP-PHF1 CFP-PHT1;2 PHT1;2-CFP YFP-PHF + PHT1;2-CFP YFP-PHF + CFP-PHT1;2 Negative control!-gfp Supplemental Figure 1: PHT1;2 accumulation is PHF1 dependent. Immunoblot analysis on total protein extract
YFP-PHF1 CFP-PHT1;2 PHT1;2-CFP YFP-PHF + PHT1;2-CFP YFP-PHF + CFP-PHT1;2 Negative control!-gfp Supplemental Figure 1: PHT1;2 accumulation is PHF1 dependent. Immunoblot analysis on total protein extract
Polyclonal ARHGAP25 antibody was prepared from rabbit serum after intracutaneous
 Preparation and purification of polyclonal antibodies Polyclonal ARHGAP25 antibody was prepared from rabbit serum after intracutaneous injections of glutathione S-transferase-ARHGAP25-(509-619) (GST-coiled
Preparation and purification of polyclonal antibodies Polyclonal ARHGAP25 antibody was prepared from rabbit serum after intracutaneous injections of glutathione S-transferase-ARHGAP25-(509-619) (GST-coiled
Specificity: Induced Fit
 Specificity: Induced Fit Conformational changes may occur upon ligand binding (Daniel Koshland in 1958) This adaptation is called the induced fit Induced fit allows for tighter binding of the ligand Induced
Specificity: Induced Fit Conformational changes may occur upon ligand binding (Daniel Koshland in 1958) This adaptation is called the induced fit Induced fit allows for tighter binding of the ligand Induced
Rapid Learning Center Presents. Teach Yourself AP Biology in 24 Hours
 Rapid Learning Center Chemistry :: Biology :: Physics :: Math Rapid Learning Center Presents Teach Yourself AP Biology in 24 Hours 1/35 *AP is a registered trademark of the College Board, which does not
Rapid Learning Center Chemistry :: Biology :: Physics :: Math Rapid Learning Center Presents Teach Yourself AP Biology in 24 Hours 1/35 *AP is a registered trademark of the College Board, which does not
STRUCTURAL BIOLOGY. α/β structures Closed barrels Open twisted sheets Horseshoe folds
 STRUCTURAL BIOLOGY α/β structures Closed barrels Open twisted sheets Horseshoe folds The α/β domains Most frequent domain structures are α/β domains: A central parallel or mixed β sheet Surrounded by α
STRUCTURAL BIOLOGY α/β structures Closed barrels Open twisted sheets Horseshoe folds The α/β domains Most frequent domain structures are α/β domains: A central parallel or mixed β sheet Surrounded by α
Molecular Biology (1)
 Molecular Biology (1) DNA structure and basic applications Mamoun Ahram, PhD Second semester, 2017-2018 Resources This lecture Cooper, pp. 49-52, 118-119, 130 What is molecular biology? Central dogma
Molecular Biology (1) DNA structure and basic applications Mamoun Ahram, PhD Second semester, 2017-2018 Resources This lecture Cooper, pp. 49-52, 118-119, 130 What is molecular biology? Central dogma
Figure S1. Figure S2. Figure S3 HB Anti-FSP27 (COOH-terminal peptide) Ab. Anti-GST-FSP27(45-127) Ab.
 / 36B4 mrna ratio Figure S1 * 2. 1.6 1.2.8 *.4 control TNFα BRL49653 Figure S2 Su bw AT p iw Anti- (COOH-terminal peptide) Ab Blot : Anti-GST-(45-127) Ab β-actin Figure S3 HB2 HW AT BA T Figure S4 A TAG
/ 36B4 mrna ratio Figure S1 * 2. 1.6 1.2.8 *.4 control TNFα BRL49653 Figure S2 Su bw AT p iw Anti- (COOH-terminal peptide) Ab Blot : Anti-GST-(45-127) Ab β-actin Figure S3 HB2 HW AT BA T Figure S4 A TAG
Zool 3200: Cell Biology Exam 3 3/6/15
 Name: Trask Zool 3200: Cell Biology Exam 3 3/6/15 Answer each of the following questions in the space provided; circle the correct answer or answers for each multiple choice question and circle either
Name: Trask Zool 3200: Cell Biology Exam 3 3/6/15 Answer each of the following questions in the space provided; circle the correct answer or answers for each multiple choice question and circle either
Newsletter Issue 7 One-STrEP Analysis of Protein:Protein-Interactions
 www.iba-biotagnology.com Newsletter Issue 7 One-STrEP Analysis of Protein:Protein-Interactions Strep-tag and One-STrEP-tag PPI Analysis with the co-precipitation/ mass spectrometry approach 3 Background
www.iba-biotagnology.com Newsletter Issue 7 One-STrEP Analysis of Protein:Protein-Interactions Strep-tag and One-STrEP-tag PPI Analysis with the co-precipitation/ mass spectrometry approach 3 Background
* ** ** * IB: p-p90rsk. p90rsk (Ser380) (arbitrary units) (Ser380) p90rsk. IB: p90rsk. Tubulin. IB: Tubulin. Ang II (200 nm) Ang II (200 nm)
 I: p-p9rsk I: p9rsk I: C I: p-p9rsk I: p9rsk 5 (ka) 5 5 (min) Ang II ( nm) p-p9rsk (Ser8) p9rsk p-p9rsk (Ser8) p9rsk (h) Mannitol 5 mm -Glucose 5 mm p9rsk (Ser8) (arbitrary units) p-p9rsk (Ser8) (arbitrary
I: p-p9rsk I: p9rsk I: C I: p-p9rsk I: p9rsk 5 (ka) 5 5 (min) Ang II ( nm) p-p9rsk (Ser8) p9rsk p-p9rsk (Ser8) p9rsk (h) Mannitol 5 mm -Glucose 5 mm p9rsk (Ser8) (arbitrary units) p-p9rsk (Ser8) (arbitrary
BIBC 103 Learning Goals with Supporting Learning Outcomes
 BIBC 103 Learning Goals with Supporting Learning Outcomes 1) Basic Lab Skills A. Conceptual understanding and moderate level of hands-on proficiency in making laboratory solutions, including understanding
BIBC 103 Learning Goals with Supporting Learning Outcomes 1) Basic Lab Skills A. Conceptual understanding and moderate level of hands-on proficiency in making laboratory solutions, including understanding
A General Protocol for GST Pull-down Lili Jing *
 A General Protocol for GST Pull-down Lili Jing * Department of Cell and Molecular Biology, University of Pennsylvania, Philadelphia, USA *For correspondence: lilijingcn@gmail.com [Abstract] GST pull-down
A General Protocol for GST Pull-down Lili Jing * Department of Cell and Molecular Biology, University of Pennsylvania, Philadelphia, USA *For correspondence: lilijingcn@gmail.com [Abstract] GST pull-down
Detection and identification of body fluid stains using antibodynanoparticle
 Electronic Supplementary Information Detection and identification of body fluid stains using antibodynanoparticle conjugates Nunzianda Frascione,* a Richard Thorogate a, Barbara Daniel a and Sue Jickells
Electronic Supplementary Information Detection and identification of body fluid stains using antibodynanoparticle conjugates Nunzianda Frascione,* a Richard Thorogate a, Barbara Daniel a and Sue Jickells
Molecular characterization, detection & quantitation of biological products Purin Charoensuksai, PhD
 Molecular characterization, detection & quantitation of biological products Purin Charoensuksai, PhD Department of Biopharmacy, Faculty of Pharmacy, Silpakorn University Example of critical checkpoints
Molecular characterization, detection & quantitation of biological products Purin Charoensuksai, PhD Department of Biopharmacy, Faculty of Pharmacy, Silpakorn University Example of critical checkpoints
