MOLECULAR DETERMINANTS OF SELECTIVE CLEARANCE OF PROTEIN INCLUSIONS BY AUTOPHAGY
|
|
|
- Horatio Day
- 5 years ago
- Views:
Transcription
1 Nature Communications Supporting Information MOLECULAR DETERMINANTS OF SELECTIVE CLEARANCE OF PROTEIN INCLUSIONS BY AUTOPHAGY Esther Wong, Eloy Bejarano, Moumita Rakshit, Karen Lee, Hugo H. Hanson, Nava Zaarur, Greg R. Phillips, Michael Y. Sherman, and Ana Maria Cuervo Supplementary Figures 1-12 Supplementary Methods
2 Supplementary Figure S1. Sph1 contains molecular determinants for aggresome and aggregate formation. (a) Schematic diagram depicting different molecular domains in the full-length and various truncated Sph1 proteins used in this study. (b) Protein expression levels of various GFP-Sph1 proteins in SY5Y cells in the absence or presence of proteasome inhibitor, lactacystin. (-) indicates untransfected control. (c) Parameters utilized throughout this work for distinguishing aggresome (Agm) from aggregates (Agg). Representative examples using FL GFP-Sph1 protein are shown on the right. (d) Representative fluorescence images of Agm and Agg formed by various GFP-Sph1 proteins in SY5Y cells under proteasomal inhibition. (e) Relative propensity of the full-length and various truncated Sph1 proteins to form Agm and Agg. Graphs reflect the percentage of transfected cells containing either Agm or Agg and are mean + SE of 3 different experiments. 2
3 Supplementary Figure S2. Effect of modulating autophagy on the characteristics of aggregates formed by different Sph1 variants. Autophagy was blocked (by treatment with 3-methyladenine; 3-MA) or upregulated (by removal of serum; S-) in SY5Y cells expressing the indicated forms of Sph1 proteins. Graphs depict the number of aggregates per cell profile (a), percentage of cellular area occupied by the aggregates (b) and the average size of the aggregates (c). Values are mean + SE of the quantification of >10 cells in 2 different experiments. (*) Significant versus basal serum supplemented condition (S+) for P= 0.01 (t-test). 3
4 Supplementary Figure S3. Cells transfected with different GFP-Sph1 variants show similar levels of viability as untransfected cells under various treatment conditions used in this study. Viability of SY5Y cells untransfected or transfected with the indicated Sph1 constructs maintained in serumsupplemented (serum+) or serum-free (serum-) media in the absence or presence of lactacystin (Lacta) or combination of lactacystin and 3-methyladenine (Lacta/3-MA). Values are mean + SE of 3 different experiments. 4
5 Supplementary Figure S4. Only ANK1 containing GFP-Sph1 variants accumulate in the insoluble pellet fraction upon inhibition of autophagy. Left: Immunoblots for GFP and actin of cell extracts sequentially prepared with Triton X-100 (Soluble) and SDS (Pellet) from SY5Y cells transfected with ANK1-containing GFP-Sph1 ACA (a) and NA (b) or GFP-Sph1 CACT lacking the ANK1 domain (c) and maintained in serum-supplemented (serum+) or serum-free (serum-) media in the absence or presence of 3-methyladenine (3-MA). Right: Densitometric quantification of blots. Values are expressed relative to the levels of the proteins in cells maintained in the presence of serum. Values are mean + SE of 3 different experiments. 5
6 Supplementary Figure S5. Effect of latrunculin A on aggresome formation and basal autophagy. (a-b) SY5Y cells expressing the indicated Sph1 proteins were treated as in Fig. 1a, and then incubated in serum+ media in the absence or presence of 3-MA or 0.2µM latrunculin A (Lat A). (a) Prolonged 17h Lat A treatment perturbs formation of FL Sph1 Agm and induces accumulation of FL Sph1 Agg. (b) Short 6h Lat A treatment did not inhibit Agm formation but we observed significant accumulation of ANK1-containing Agg (right column) similar to when autophagy was blocked. Values are expressed as folds the amount of cells positive for Agm or Agg in untreated cells which were given an arbitrary value of 1. Values are mean + SE of 3 different experiments. (*) Significant versus untreated. P= 0.03 (t-test). 6
7 Figure S6. Effect of expression of GFP tagged p38 and ANK1-p38 on cellular autophagic activity. (a) GFP and p38 immunoblots of GFP-p38 (clones #3, 4 and 5) and GFP-ANK1-p38 (clones #9, 29, 31) fusion proteins expressed in SY5Y cells. Lane (-) represents untransfected control. (b) LC3 immunoblots of SY5Y cells transfected with an empty vector, p38 or ANK1-p38 in serum+ or serum- conditions treated or not with lysosomal inhibitors NH 4 CI and leupeptin (N/L). (c) Autophagic flux calculated as the increase in the amount of LC3-II upon blockage of lysosomal degradation in each condition. 7
8 Supplementary Figure S7. Effect of different Sph1 proteins on the association of LAMP1 with p38 aggresomes. SY5Y cells expressing HA-p38 together with the indicated Sph1 protein variants were maintained in the absence of serum and then immunostained for the lysosomal marker LAMP1. The Individual channels, merge of the three channels and the negative black and white images of the LAMP1 channel are shown. Yellow and black arrows indicate the association of Agm with LAMP1 in each image. 8
9 Supplementary Figure S8. FRAP analysis of the dynamics of mcherry-p62 in different types of aggresomes. Representative images before (prebleach) and after photobleaching to show the dynamics of mcherryp62 in the p62 sequestosome (control) and in Agm formed by the indicated aggregation-prone proteins. Time courses and mean values of diffusing fraction and t 1/2 are shown in Fig. 5b. 9
10 Figure S9. Autophagic activity remains intact in various p62 and NBR1 KO or KD cells. (a) Formation of p38 and ANK1-p38 Agm under proteasome inhibition is not significantly affected in p62 KD cells compared to control cells. (b-e) Comparison of autophagic activity in control and p62 KD SY5Y cells (b), WT and p62 KO MEFs (c), control and NBR1 KD SY5Y cells (d), and control and p62/nbr1 double KD cells (e) by measuring LC3 flux. Left: Representative immunoblots of LC3-II and actin in total cell lysates from the respective cells treated or not with lysosomal inhibitors NH 4 CI and leupeptin (N/L) maintained in the presence (serum+) or absence of serum (serum-) for 4h. Right: Net LC3-II autophagic flux in the control and various KO or KD cells calculated as the increase in the amount of LC3-II upon blockage of lysosomal degradation under serum+ and serum- conditions and expressed as times the values of the control cells under serum+ condition. Values are mean + SE of 3 different experiments and significance calculated using t-test. 10
11 Supplementary Figure S10. Autophagic clearance of ANK1 containing Sph1 Agm and Agg is not affected by decrease in p62 and NBR1 protein levels. (a-c) Autophagic clearance of ANK1-containing Sph1 Agm and Agg in control (Ctr), p62 knockdown (p62 KD), NBR1 KD and p62/nbr1 double KD SY5Y cells in response to serum removal. (d) Relative fold difference in the number of FL Agg under basal serum+ conditions treated or not with 3-MA. Values are mean + SE (n=3). (*) Significant versus serum+ condition. P= 0.04 (t-test) (e) Fluorescence images showing colocalization patterns of FL Agg (green) with endogenous p62 and NBR1 (red). 11
12 Supplementary Figure S11. Autophagic clearance of ANK1 containing Sph1 Agm is not affected in p62 knockout MEF. (a) Representative fluorescence images of Agm and Agg formed by various GFP-Sph1 proteins in p62 KO MEF under proteasomal inhibition. (b) Autophagic clearance of various GFP-Sph1 Agm and Agg in WT and p62 KO MEFs in response to serum removal. Values are mean + SE (n=3). (*) Significant versus serum+ condition. P= (t-test). 12
13 Supplementary Figure S12. FRAP analysis of the dynamics of LysoTracker in different types of Agm. FRAP analysis of the dynamics of LysoTracker in stained lysosomes alone or in lysosomes close to various types of Agm. Values are mean + SE of 10 cells analyzed over 3 different experiments. 13
14 Supplementary Methods Transfections. For microscopy experiments, 2.5 x 10 4 SY5Y cells and MEFs were seeded on coverslips for transient transfection with various expression vectors. 0.25µg DNA was used for all single transfection reactions except for GFP-CACT where 1µg DNA was used. For co-expression of two proteins, 0.25µg of each plasmid DNA was used. For immunoblotting and immunoprecipitation, 2 x 10 5 SY5Y cells were seeded in 6 well plates for transfection with 0.5µg of DNA in single transfection or 0.5µg of each plasmid DNA in co-transfection reactions. Inclusion formation and autophagic clearance. The susceptibility of various inclusions formed upon proteasomal inhibition toward autophagic removal was analyzed as described. 24h post-transfection, cells were first treated with 5μM lactacystin (12h), a proteasome inhibitor, to enhance aggresome-like inclusion formation. To analyze the effect of basal quality control autophagy on the inclusion number, the lactacystin treated cells were allowed to recover in serum containing media either in the presence or absence of autophagy inhibitor, 10mM 3-MA (Sigma) for 24h. Alternatively, basal quality control autophagy in the treated cells was inhibited by 6h treatment with latrunculin A (Invitrogen). To analyze the effect of stimulated autophagy on the inclusion number, a concurrent set of treated cells were washed extensively before incubating in serum free media to stimulate autophagy either in the presence or absence of 3-MA. Thereafter, the cells were fixed in 3% formaldehyde and the number of cells possessing aggresomes or aggregates were visualized using an Axiovert fluorescence microscope (Carl Zeiss Ltd., Thornwood, NY). Random fields of each slide were taken and a minimum of 200 SY5Y cells and 100 MEFs were counted for each experiment. Agm and Agg were scored according to the characteristics described in Fig. S1c. Briefly, an Agm-containing cell is that displaying a distinct large perinuclear structure of > 5μM, whereas Agg-containing cells were considering those containing 2 or more small aggregate structures of < 3μM disperse throughout the cells. For allocation purposes, those cells containing a distinct perinuclear large aggresomal structure, even in the presence of smaller aggregates, were scored as Agm-containing cells. In addition of counting the percent of cells containing Agg or Agm, where indicated, the percentage of cellular area occupied by these structures was calculated, by tracking individual cellular profiles and, after thresholding, comparing the total cellular area and the area of the fluorescent compartment. 14
15 Cell viability measurement. Viability of cells transfected with the different GFP-Sph1 variants as well as untransfected SY5Y control cells incubated under the various treatment conditions was measured using CellTiter-Blue Cell Viability Assay according to the manufacturer s instructions (Promega). Immunoblot and immunoprecipitation. Lysates were prepared by suspending the cells in ice cold RIPA lysis buffer. Protein was determined by the Lowry method, using BSA as a standard. 100ug of protein per sample was loaded for LC3-II flux analysis while 50-75µg of protein was used in other immunoblotting. In some cases, sequential fractionation of cell lysates into Triton-X-solubilized (Soluble) and SDS-solubilized (Pellet) fractions was performed as previously described 23. After SDS-PAGE and immunoblotting, the proteins recognized by the specific antibodies were visualized by chemiluminescence methods (Western lightning ECL plus, PerkinElmer). The intensity of the bands was quantified using the ImageJ software (NIH). For immunoprecipitation, cells co-transfected with either pcdna vector or various HA-ubiquitin constructs and the GFP-p38 variants or FL GFP-Sph1 variants were lysed in RIPA lysis buffer containing 6M Urea with brief sonication. Immunoprecipitation (IP) was performed with anti-gfp antibody at 4 C overnight. The IP mixtures were centrifuged at 800g for 5min to remove any precipitates prior to the addition of protein A/G beads (Amersham Biosciences) for further incubation of another 1h. Immunoprecipitates were washed three times with lysis buffer and subjected to SDS-PAGE and immunoblotting with the indicated antibodies. Live cell imaging and photobleaching. SY5Y cells expressing GFP tagged p38, ANK1-38, ANK1-p38 K385R mutant and mcherry-p62 separately or in combination were cultured on glass bottom 35mm culture dishes (MatTek). Cells were maintained in Minimum Essential Medium without phenol red supplemented with 10% FBS, sodium bicarbonate and HEPES at 37ºC for the duration of the experiment. Fluorescence recovery after photobleaching (FRAP) analysis was performed 24h after transfection using either the 488nm or 543nm laser wavelength for GFP and mcherry constructs respectively. Singles nonsaturated images were taken with a 63x 1.4 NA objective and a pinhole diameter equivalent to 4-5 Airy units at 512x512 pixels using a Leica TCS SP5 inverted confocal microscope (Leica Microsystems Inc.). Selective photobleaching of the aggresome area was carried out at % laser power until the fluorescence bleached away from the aggresome. The recovery of fluorescence in the region of interest was monitored by capturing images every 3 seconds at a low power of laser in a time course sequence. 15
16 Before bleaching, 3-4 images at low intensity illumination were taken to establish the fluorescence baseline. FRAP quantification analysis was performed as described previously 37. Average aggresomeassociated fluorescence intensity was determined using Leica Confocal software and the relative fluorescence intensity (RFI) was calculated as the average of at least three data points. RFI values were calculated using the equation (Ne t /N1 t )/(Ne 0 /N1 0 ) where Ne t represents the average intensity of bleachedaggresome at each time point and N1 t is the average intensity of non-bleached area in the neighboring cell at this given point which functions as control for global photobleaching. Ne 0 /N1 0 represents the ratio average intensity of the bleached and non-bleached area before photobleaching. FRAP data were fit using Sigma Plot Software to two-phase exponential method as described for other proteins in aggresomes 37. Same procedures were carried out for FRAP analysis of LysoTracker stained lysosomes. Correlative light and electron microscopy. SY5Y cells expressing GFP tagged NA Sph1, CACT Sph1, p38 and ANK1-p38 proteins grown on grid glass bottom 35mm culture dish were recovered in serum-free medium for 6h after proteasome inhibition. 5µM vinblastine was added for the last 2h of starvation to enhance the presence of autophagosomes. Thereafter the cells were fixed with 4% glutaraldehyde in 0.1M sodium cacodylate buffer with 1mM CaCl 2. Confocal stacks and DIC images of transfected cells were acquired on a Zeiss LSM 510 META microscope. Cells were embedded in Epon and processed as described The imaged cell was relocated in the block face and sectioned through. Sections were contrasted with lead citrate and uranyl acetate and serial sections of the cell of interest were documented at 10,000, 15,000 and 30,000x magnifications on a Hitachi 7000 electron microscope. Confocal, DIC and TEM images were realigned and oriented using nuclear and other morphological landmarks. References 37 Kim, S., Nollen, E. A. A., Kitagawa, K., Bindokas, V. P. & Morimoto, R. I. Polyglutamine protein aggregates are dynamic. Nat. Cell Biol. 4, (2002). 38 Hanson, H. H. et al. LC3-dependent intracellular membrane tubules induced by gammaprotocadherins A3 and B2: A role for intraluminal interactions. J. Biol. Chem. 285, (2010). 16
17 39 Hanson, H. H., Reilly, J. E., Lee, R., Janssen, W. G. & Phillips, G. R. Streamlined embedding of cell monolayers on gridded live imaging dishes for correlative light and electron microscopy. Microsc. Microanal. 20, 1-8 (2010). 17
T H E J O U R N A L O F C E L L B I O L O G Y
 T H E J O U R N A L O F C E L L B I O L O G Y Supplemental material Rainero et al., http://www.jcb.org/cgi/content/full/jcb.201109112/dc1 Figure S1. The expression of DGK- is reduced upon transfection
T H E J O U R N A L O F C E L L B I O L O G Y Supplemental material Rainero et al., http://www.jcb.org/cgi/content/full/jcb.201109112/dc1 Figure S1. The expression of DGK- is reduced upon transfection
SUPPLEMENTARY INFORMATION
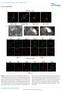 DOI: 10.1038/ncb2386 Figure 1 Src-containing puncta are not focal adhesions, podosomes or endosomes. (a) FAK-/- were stained with anti-py416 Src (green) and either (in red) the focal adhesion protein paxillin,
DOI: 10.1038/ncb2386 Figure 1 Src-containing puncta are not focal adhesions, podosomes or endosomes. (a) FAK-/- were stained with anti-py416 Src (green) and either (in red) the focal adhesion protein paxillin,
MEFs were treated with the indicated concentrations of LLOMe for three hours, washed
 Supplementary Materials and Methods Cell Fractionation MEFs were treated with the indicated concentrations of LLOMe for three hours, washed with ice-cold PBS, collected by centrifugation, and then homogenized
Supplementary Materials and Methods Cell Fractionation MEFs were treated with the indicated concentrations of LLOMe for three hours, washed with ice-cold PBS, collected by centrifugation, and then homogenized
SUPPLEMENTARY INFORMATION. Chemical modulation of Chaperone-mediated autophagy by novel
 SUPPLEMENTARY INFORMATION Chemical modulation of Chaperone-mediated autophagy by novel retinoic acid derivatives Jaime Anguiano 1, Thomas P Garner 2, Murugesan Mahalingam 1, Bhaskar C. Das 1, Evripidis
SUPPLEMENTARY INFORMATION Chemical modulation of Chaperone-mediated autophagy by novel retinoic acid derivatives Jaime Anguiano 1, Thomas P Garner 2, Murugesan Mahalingam 1, Bhaskar C. Das 1, Evripidis
Figure 1: TDP-43 is subject to lysine acetylation within the RNA-binding domain a) QBI-293 cells were transfected with TDP-43 in the presence or
 Figure 1: TDP-43 is subject to lysine acetylation within the RNA-binding domain a) QBI-293 cells were transfected with TDP-43 in the presence or absence of the acetyltransferase CBP and acetylated TDP-43
Figure 1: TDP-43 is subject to lysine acetylation within the RNA-binding domain a) QBI-293 cells were transfected with TDP-43 in the presence or absence of the acetyltransferase CBP and acetylated TDP-43
JCB. Supplemental material THE JOURNAL OF CELL BIOLOGY. Hong et al.,
 Supplemental material JCB Hong et al., http://www.jcb.org/cgi/content/full/jcb.201412127/dc1 THE JOURNAL OF CELL BIOLOGY Figure S1. Analysis of purified proteins by SDS-PAGE and pull-down assays. (A) Coomassie-stained
Supplemental material JCB Hong et al., http://www.jcb.org/cgi/content/full/jcb.201412127/dc1 THE JOURNAL OF CELL BIOLOGY Figure S1. Analysis of purified proteins by SDS-PAGE and pull-down assays. (A) Coomassie-stained
Supporting Information
 Supporting Information Stavru et al. 0.073/pnas.357840 SI Materials and Methods Immunofluorescence. For immunofluorescence, cells were fixed for 0 min in 4% (wt/vol) paraformaldehyde (Electron Microscopy
Supporting Information Stavru et al. 0.073/pnas.357840 SI Materials and Methods Immunofluorescence. For immunofluorescence, cells were fixed for 0 min in 4% (wt/vol) paraformaldehyde (Electron Microscopy
Sarker et al. Supplementary Material. Subcellular Fractionation
 Supplementary Material Subcellular Fractionation Transfected 293T cells were harvested with phosphate buffered saline (PBS) and centrifuged at 2000 rpm (500g) for 3 min. The pellet was washed, re-centrifuged
Supplementary Material Subcellular Fractionation Transfected 293T cells were harvested with phosphate buffered saline (PBS) and centrifuged at 2000 rpm (500g) for 3 min. The pellet was washed, re-centrifuged
Supplemental Information. Pacer Mediates the Function of Class III PI3K. and HOPS Complexes in Autophagosome. Maturation by Engaging Stx17
 Molecular Cell, Volume 65 Supplemental Information Pacer Mediates the Function of Class III PI3K and HOPS Complexes in Autophagosome Maturation by Engaging Stx17 Xiawei Cheng, Xiuling Ma, Xianming Ding,
Molecular Cell, Volume 65 Supplemental Information Pacer Mediates the Function of Class III PI3K and HOPS Complexes in Autophagosome Maturation by Engaging Stx17 Xiawei Cheng, Xiuling Ma, Xianming Ding,
Supplementary data. sienigma. F-Enigma F-EnigmaSM. a-p53
 Supplementary data Supplemental Figure 1 A sienigma #2 sienigma sicontrol a-enigma - + ++ - - - - - - + ++ - - - - - - ++ B sienigma F-Enigma F-EnigmaSM a-flag HLK3 cells - - - + ++ + ++ - + - + + - -
Supplementary data Supplemental Figure 1 A sienigma #2 sienigma sicontrol a-enigma - + ++ - - - - - - + ++ - - - - - - ++ B sienigma F-Enigma F-EnigmaSM a-flag HLK3 cells - - - + ++ + ++ - + - + + - -
Supplementary methods Shoc2 In Vitro Ubiquitination Assay
 Supplementary methods Shoc2 In Vitro Ubiquitination Assay 35 S-labelled Shoc2 was prepared using a TNT quick Coupled transcription/ translation System (Promega) as recommended by manufacturer. For the
Supplementary methods Shoc2 In Vitro Ubiquitination Assay 35 S-labelled Shoc2 was prepared using a TNT quick Coupled transcription/ translation System (Promega) as recommended by manufacturer. For the
Supplementary Figure 1. Intracellular distribution of the EPE peptide. HeLa cells were serum-starved (16 h, 0.1%), and treated with EPE peptide,
 Supplementary Figure 1. Intracellular distribution of the EPE peptide. HeLa cells were serum-starved (16 h, 0.1%), and treated with EPE peptide, conjugated with either TAT or Myristic acid and biotin for
Supplementary Figure 1. Intracellular distribution of the EPE peptide. HeLa cells were serum-starved (16 h, 0.1%), and treated with EPE peptide, conjugated with either TAT or Myristic acid and biotin for
Supplementary Figure 1. Confirmation of sirna in PC3 and H1299 cells PC3 (a) and H1299 (b) cells were transfected with sirna oligonucleotides
 Supplementary Figure 1. Confirmation of sirna in PC3 and H1299 cells PC3 (a) and H1299 (b) cells were transfected with sirna oligonucleotides targeting RCP (SMARTPool (RCP) or two individual oligos (RCP#1
Supplementary Figure 1. Confirmation of sirna in PC3 and H1299 cells PC3 (a) and H1299 (b) cells were transfected with sirna oligonucleotides targeting RCP (SMARTPool (RCP) or two individual oligos (RCP#1
Supplementary Figure 1 Phosphorylated tau accumulates in Nrf2 (-/-) mice. Hippocampal tissues obtained from Nrf2 (-/-) (10 months old, 4 male; 2
 Supplementary Figure 1 Phosphorylated tau accumulates in Nrf2 (-/-) mice. Hippocampal tissues obtained from Nrf2 (-/-) (10 months old, 4 male; 2 female) or wild-type (5 months old, 1 male; 11 months old,
Supplementary Figure 1 Phosphorylated tau accumulates in Nrf2 (-/-) mice. Hippocampal tissues obtained from Nrf2 (-/-) (10 months old, 4 male; 2 female) or wild-type (5 months old, 1 male; 11 months old,
Supplementary Figure legends. Supplementary Methods
 Supplementary Methods Transmission electron microscopy. For transmission electron microscopy (TEM), the cell populations were rinsed with 0.1 Sorensen s buffer (ph 7.5), fixed in 2.5% glutaraldehyde for
Supplementary Methods Transmission electron microscopy. For transmission electron microscopy (TEM), the cell populations were rinsed with 0.1 Sorensen s buffer (ph 7.5), fixed in 2.5% glutaraldehyde for
Grb2-Mediated Alteration in the Trafficking of AβPP: Insights from Grb2-AICD Interaction
 Journal of Alzheimer s Disease 20 (2010) 1 9 1 IOS Press Supplementary Material Grb2-Mediated Alteration in the Trafficking of AβPP: Insights from Grb2-AICD Interaction Mithu Raychaudhuri and Debashis
Journal of Alzheimer s Disease 20 (2010) 1 9 1 IOS Press Supplementary Material Grb2-Mediated Alteration in the Trafficking of AβPP: Insights from Grb2-AICD Interaction Mithu Raychaudhuri and Debashis
T H E J O U R N A L O F C E L L B I O L O G Y
 Supplemental material Thompson et al., http://www.jcb.org/cgi/content/full/jcb.200909067/dc1 T H E J O U R N A L O F C E L L B I O L O G Y Figure S1. Modification-specific antibodies do not detect unmodified
Supplemental material Thompson et al., http://www.jcb.org/cgi/content/full/jcb.200909067/dc1 T H E J O U R N A L O F C E L L B I O L O G Y Figure S1. Modification-specific antibodies do not detect unmodified
Supplementary Material
 Supplementary Material Supplementary Methods Cell synchronization. For synchronized cell growth, thymidine was added to 30% confluent U2OS cells to a final concentration of 2.5mM. Cells were incubated
Supplementary Material Supplementary Methods Cell synchronization. For synchronized cell growth, thymidine was added to 30% confluent U2OS cells to a final concentration of 2.5mM. Cells were incubated
. Viability of colonies was then assessed using the WST-1 reagent as described above, and normalized relative to untreated controls.
 Cell viability analysis in the absence of disaggregation To assess cell viability in the absence of disaggregation, quintuplicate samples of cells at 5 x 1 5 /ml were treated with mab (1 µg/ml) for 24
Cell viability analysis in the absence of disaggregation To assess cell viability in the absence of disaggregation, quintuplicate samples of cells at 5 x 1 5 /ml were treated with mab (1 µg/ml) for 24
Stargazin regulates AMPA receptor trafficking through adaptor protein. complexes during long term depression
 Supplementary Information Stargazin regulates AMPA receptor trafficking through adaptor protein complexes during long term depression Shinji Matsuda, Wataru Kakegawa, Timotheus Budisantoso, Toshihiro Nomura,
Supplementary Information Stargazin regulates AMPA receptor trafficking through adaptor protein complexes during long term depression Shinji Matsuda, Wataru Kakegawa, Timotheus Budisantoso, Toshihiro Nomura,
Figure legends for supplement
 Figure legends for supplement Supplemental Figure 1 Characterization of purified and recombinant proteins Relevant fractions related the final stage of the purification protocol(bingham et al., 1998; Toba
Figure legends for supplement Supplemental Figure 1 Characterization of purified and recombinant proteins Relevant fractions related the final stage of the purification protocol(bingham et al., 1998; Toba
Supplementary Figure 1. α-synuclein is truncated in PD and LBD brains. Nature Structural & Molecular Biology: doi: /nsmb.
 Supplementary Figure 1 α-synuclein is truncated in PD and LBD brains. (a) Specificity of anti-n103 antibody. Anti-N103 antibody was coated on an ELISA plate and different concentrations of full-length
Supplementary Figure 1 α-synuclein is truncated in PD and LBD brains. (a) Specificity of anti-n103 antibody. Anti-N103 antibody was coated on an ELISA plate and different concentrations of full-length
B. ADM: C. D. Apoptosis: 1.68% 2.99% 1.31% Figure.S1,Li et al. number. invaded cells. HuH7 BxPC-3 DLD-1.
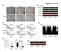 A. - Figure.S1,Li et al. B. : - + - + - + E-cadherin CK19 α-sma vimentin β -actin C. D. Apoptosis: 1.68% 2.99% 1.31% - : - + - + - + Apoptosis: 48.33% 45.32% 44.59% E. invaded cells number 400 300 200
A. - Figure.S1,Li et al. B. : - + - + - + E-cadherin CK19 α-sma vimentin β -actin C. D. Apoptosis: 1.68% 2.99% 1.31% - : - + - + - + Apoptosis: 48.33% 45.32% 44.59% E. invaded cells number 400 300 200
Supplemental Materials and Methods
 Supplemental Materials and Methods Co-immunoprecipitation (Co-IP) assay Cells were lysed with NETN buffer (20 mm Tris-HCl, ph 8.0, 0 mm NaCl, 1 mm EDT, 0.5% Nonidet P-40) containing 50 mm β-glycerophosphate,
Supplemental Materials and Methods Co-immunoprecipitation (Co-IP) assay Cells were lysed with NETN buffer (20 mm Tris-HCl, ph 8.0, 0 mm NaCl, 1 mm EDT, 0.5% Nonidet P-40) containing 50 mm β-glycerophosphate,
Supplementary Materials and Methods
 Supplementary Materials and Methods sirna sequences used in this study The sequences of Stealth Select RNAi for ALK and FLOT-1 were as follows: ALK sense no.1 (ALK): 5 -AAUACUGACAGCCACAGGCAAUGUC-3 ; ALK
Supplementary Materials and Methods sirna sequences used in this study The sequences of Stealth Select RNAi for ALK and FLOT-1 were as follows: ALK sense no.1 (ALK): 5 -AAUACUGACAGCCACAGGCAAUGUC-3 ; ALK
< Supporting Information >
 SUPPORTING INFORMATION 1 < Supporting Information > Discovery of autophagy modulators through the construction of high-content screening platform via monitoring of lipid droplets Sanghee Lee, Eunha Kim,
SUPPORTING INFORMATION 1 < Supporting Information > Discovery of autophagy modulators through the construction of high-content screening platform via monitoring of lipid droplets Sanghee Lee, Eunha Kim,
Supplementary Figure 1. APP cleavage assay. HEK293 cells were transfected with various
 Supplementary Figure 1. APP cleavage assay. HEK293 cells were transfected with various GST-tagged N-terminal truncated APP fragments including GST-APP full-length (FL), APP (123-695), APP (189-695), or
Supplementary Figure 1. APP cleavage assay. HEK293 cells were transfected with various GST-tagged N-terminal truncated APP fragments including GST-APP full-length (FL), APP (123-695), APP (189-695), or
Journal of Cell Science Supplementary Material
 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 SUPPLEMENTARY FIGURE LEGENDS Figure S1: Eps8 is localized at focal adhesions and binds directly to FAK (A) Focal
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 SUPPLEMENTARY FIGURE LEGENDS Figure S1: Eps8 is localized at focal adhesions and binds directly to FAK (A) Focal
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/9/429/ra54/dc1 Supplementary Materials for Dephosphorylation of the adaptor LAT and phospholipase C by SHP-1 inhibits natural killer cell cytotoxicity Omri Matalon,
www.sciencesignaling.org/cgi/content/full/9/429/ra54/dc1 Supplementary Materials for Dephosphorylation of the adaptor LAT and phospholipase C by SHP-1 inhibits natural killer cell cytotoxicity Omri Matalon,
GFP CCD2 GFP IP:GFP
 D1 D2 1 75 95 148 178 492 GFP CCD1 CCD2 CCD2 GFP D1 D2 GFP D1 D2 Beclin 1 IB:GFP IP:GFP Supplementary Figure 1: Mapping domains required for binding to HEK293T cells are transfected with EGFP-tagged mutant
D1 D2 1 75 95 148 178 492 GFP CCD1 CCD2 CCD2 GFP D1 D2 GFP D1 D2 Beclin 1 IB:GFP IP:GFP Supplementary Figure 1: Mapping domains required for binding to HEK293T cells are transfected with EGFP-tagged mutant
Methanol fixation allows better visualization of Kal7. To compare methods for
 Supplementary Data Methanol fixation allows better visualization of Kal7. To compare methods for visualizing Kal7 in dendrites, mature cultures of dissociated hippocampal neurons (DIV21) were fixed with
Supplementary Data Methanol fixation allows better visualization of Kal7. To compare methods for visualizing Kal7 in dendrites, mature cultures of dissociated hippocampal neurons (DIV21) were fixed with
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION Contents: Supplementary Figure 1. Additional structural and binding data for designed tuim peptides. Supplementary Figure 2. Subcellular localization patterns of designed tuim
SUPPLEMENTARY INFORMATION Contents: Supplementary Figure 1. Additional structural and binding data for designed tuim peptides. Supplementary Figure 2. Subcellular localization patterns of designed tuim
Flag-Rac Vector V12 V12 N17 C40. Vector C40 pakt (T308) Akt1. Myc-DN-PAK1 (N-SP)
 a b FlagRac FlagRac V2 V2 N7 C4 V2 V2 N7 C4 p (T38) p (S99, S24) p Flag (Rac) NIH 3T3 COS c +Serum p (T38) MycDN (NSP) Mycp27 3 6 2 3 6 2 3 6 2 min p Myc ( or p27) Figure S (a) Effects of Rac mutants on
a b FlagRac FlagRac V2 V2 N7 C4 V2 V2 N7 C4 p (T38) p (S99, S24) p Flag (Rac) NIH 3T3 COS c +Serum p (T38) MycDN (NSP) Mycp27 3 6 2 3 6 2 3 6 2 min p Myc ( or p27) Figure S (a) Effects of Rac mutants on
Supplementary Figure S1. Immunodetection of full-length XA21 and the XA21 C-terminal cleavage product.
 Supplementary Information Supplementary Figure S1. Immunodetection of full-length XA21 and the XA21 C-terminal cleavage product. Total protein extracted from Kitaake wild type and rice plants carrying
Supplementary Information Supplementary Figure S1. Immunodetection of full-length XA21 and the XA21 C-terminal cleavage product. Total protein extracted from Kitaake wild type and rice plants carrying
Transcriptional regulation of BRCA1 expression by a metabolic switch: Di, Fernandez, De Siervi, Longo, and Gardner. H3K4Me3
 ChIP H3K4Me3 enrichment.25.2.15.1.5 H3K4Me3 H3K4Me3 ctrl H3K4Me3 + E2 NS + E2 1. kb kb +82 kb Figure S1. Estrogen promotes entry of MCF-7 into the cell cycle but does not significantly change activation-associated
ChIP H3K4Me3 enrichment.25.2.15.1.5 H3K4Me3 H3K4Me3 ctrl H3K4Me3 + E2 NS + E2 1. kb kb +82 kb Figure S1. Estrogen promotes entry of MCF-7 into the cell cycle but does not significantly change activation-associated
T H E J O U R N A L O F C E L L B I O L O G Y
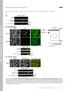 T H E J O U R N A L O F C E L L B I O L O G Y Supplemental material Nakajima and Tanoue, http://www.jcb.org/cgi/content/full/jcb.201104118/dc1 Figure S1. DLD-1 cells exhibit the characteristic morphology
T H E J O U R N A L O F C E L L B I O L O G Y Supplemental material Nakajima and Tanoue, http://www.jcb.org/cgi/content/full/jcb.201104118/dc1 Figure S1. DLD-1 cells exhibit the characteristic morphology
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb2271 Supplementary Figure a! WM266.4 mock WM266.4 #7 sirna WM266.4 #10 sirna SKMEL28 mock SKMEL28 #7 sirna SKMEL28 #10 sirna WM1361 mock WM1361 #7 sirna WM1361 #10 sirna 9 WM266. WM136
DOI: 10.1038/ncb2271 Supplementary Figure a! WM266.4 mock WM266.4 #7 sirna WM266.4 #10 sirna SKMEL28 mock SKMEL28 #7 sirna SKMEL28 #10 sirna WM1361 mock WM1361 #7 sirna WM1361 #10 sirna 9 WM266. WM136
ASPP1 Fw GGTTGGGAATCCACGTGTTG ASPP1 Rv GCCATATCTTGGAGCTCTGAGAG
 Supplemental Materials and Methods Plasmids: the following plasmids were used in the supplementary data: pwzl-myc- Lats2 (Aylon et al, 2006), pretrosuper-vector and pretrosuper-shp53 (generous gift of
Supplemental Materials and Methods Plasmids: the following plasmids were used in the supplementary data: pwzl-myc- Lats2 (Aylon et al, 2006), pretrosuper-vector and pretrosuper-shp53 (generous gift of
Supplementary Figure 1. The Hsp70 acetylation level is related to the co-chaperone binding of Hsp70 under various stress conditions.
 Supplementary Figure 1. The Hsp70 acetylation level is related to the co-chaperone binding of Hsp70 under various stress conditions. 1 (a) Etoposide treatment gradually changes acetylation level and co-chaperone
Supplementary Figure 1. The Hsp70 acetylation level is related to the co-chaperone binding of Hsp70 under various stress conditions. 1 (a) Etoposide treatment gradually changes acetylation level and co-chaperone
Regulation of autophagic activity by ζ proteins associated with class III. phosphatidylinositol-3 kinase. Mercedes Pozuelo Rubio
 ONLINE SUPPORTING INFORMATION Regulation of autophagic activity by 14-3-3ζ proteins associated with class III phosphatidylinositol-3 kinase Mercedes Pozuelo Rubio entro Andaluz de Biología Molecular y
ONLINE SUPPORTING INFORMATION Regulation of autophagic activity by 14-3-3ζ proteins associated with class III phosphatidylinositol-3 kinase Mercedes Pozuelo Rubio entro Andaluz de Biología Molecular y
Apoptosis And Anti-tumor Effect Induced By Mtor Inhibitor And Autophagy Inhibitor In Human Osteosarcoma Cells
 Apoptosis And Anti-tumor Effect Induced By Mtor Inhibitor And Autophagy Inhibitor In Human Osteosarcoma Cells Ryosuke Horie. Kagawa University of medecine, Kita-gun, Japan. Disclosures: R. Horie: None.
Apoptosis And Anti-tumor Effect Induced By Mtor Inhibitor And Autophagy Inhibitor In Human Osteosarcoma Cells Ryosuke Horie. Kagawa University of medecine, Kita-gun, Japan. Disclosures: R. Horie: None.
Supplementary Table 1. The Q-PCR primer sequence is summarized in the following table.
 Supplementary Table 1. The Q-PCR primer sequence is summarized in the following table. Name Sequence (5-3 ) Application Flag-u ggactacaaggacgacgatgac Shared upstream primer for all the amplifications of
Supplementary Table 1. The Q-PCR primer sequence is summarized in the following table. Name Sequence (5-3 ) Application Flag-u ggactacaaggacgacgatgac Shared upstream primer for all the amplifications of
Four different active promoter genes were chosen, ATXN7L2, PSRC1, CELSR2 and
 SUPPLEMENTARY MATERIALS AND METHODS Chromatin Immunoprecipitation for qpcr analysis Four different active promoter genes were chosen, ATXN7L2, PSRC1, CELSR2 and IL24, all located on chromosome 1. Primer
SUPPLEMENTARY MATERIALS AND METHODS Chromatin Immunoprecipitation for qpcr analysis Four different active promoter genes were chosen, ATXN7L2, PSRC1, CELSR2 and IL24, all located on chromosome 1. Primer
Supplemental Materials and Methods
 Supplemental Materials and Methods In situ hybridization In situ hybridization analysis of HFE2 and genin mrna in rat liver tissues was performed as previously described (1). Briefly, the digoxigenin-labeled
Supplemental Materials and Methods In situ hybridization In situ hybridization analysis of HFE2 and genin mrna in rat liver tissues was performed as previously described (1). Briefly, the digoxigenin-labeled
Segments of the obstructed intestinal loops were fixed in 4% paraformaldehyde
 Supplementary text Supplementary materials and methods Histopathological examination Segments of the obstructed intestinal loops were fixed in 4% paraformaldehyde (PFA) and embedded in paraffin wax with
Supplementary text Supplementary materials and methods Histopathological examination Segments of the obstructed intestinal loops were fixed in 4% paraformaldehyde (PFA) and embedded in paraffin wax with
J. Cell Sci. 128: doi: /jcs : Supplementary Material. Supplemental Figures. Journal of Cell Science Supplementary Material
 Supplemental Figures Figure S1. Trio controls endothelial barrier function. (A) TagRFP-shTrio constructs were expressed in ECs. Western blot shows efficient Trio knockdown in TagRFP-expressing ECs. (B)
Supplemental Figures Figure S1. Trio controls endothelial barrier function. (A) TagRFP-shTrio constructs were expressed in ECs. Western blot shows efficient Trio knockdown in TagRFP-expressing ECs. (B)
CD93 and dystroglycan cooperation in human endothelial cell adhesion and migration
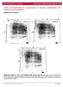 /, Supplementary Advance Publications Materials 2016 CD93 and dystroglycan cooperation in human endothelial cell adhesion and migration Supplementary Materials Supplementary Figure S1: In ECs CD93 silencing
/, Supplementary Advance Publications Materials 2016 CD93 and dystroglycan cooperation in human endothelial cell adhesion and migration Supplementary Materials Supplementary Figure S1: In ECs CD93 silencing
SUPPLEMENTAL MATERIAL. Supplemental Methods:
 SUPPLEMENTAL MATERIAL Supplemental Methods: Immunoprecipitation- As we described but with some modifications [22]. As part of another ongoing project, lysate from human umbilical vein endothelial cells
SUPPLEMENTAL MATERIAL Supplemental Methods: Immunoprecipitation- As we described but with some modifications [22]. As part of another ongoing project, lysate from human umbilical vein endothelial cells
Supplementary Figure 1 Structural modeling and purification of V. cholerae ABH. (a) The migration of the purified rabh and catalytically inactive
 Supplementary Figure 1 Structural modeling and purification of V. cholerae ABH. (a) The migration of the purified rabh and catalytically inactive variants rabhs, rabhd, and rabhh are shown on 12% SDS-PAGE
Supplementary Figure 1 Structural modeling and purification of V. cholerae ABH. (a) The migration of the purified rabh and catalytically inactive variants rabhs, rabhd, and rabhh are shown on 12% SDS-PAGE
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature10016 Supplementary discussion on binding site density for protein complexes on the surface: The density of biotin sites on the chip is ~10 3 biotin-peg per µm 2. The biotin sites are
doi:10.1038/nature10016 Supplementary discussion on binding site density for protein complexes on the surface: The density of biotin sites on the chip is ~10 3 biotin-peg per µm 2. The biotin sites are
Confocal immunofluorescence microscopy
 Confocal immunofluorescence microscopy HL-6 and cells were cultured and cytospun onto glass slides. The cells were double immunofluorescence stained for Mt NPM1 and fibrillarin (nucleolar marker). Briefly,
Confocal immunofluorescence microscopy HL-6 and cells were cultured and cytospun onto glass slides. The cells were double immunofluorescence stained for Mt NPM1 and fibrillarin (nucleolar marker). Briefly,
SUPPLEMENTARY INFORMATION
 The Supplementary Information (SI) Methods Cell culture and transfections H1299, U2OS, 293, HeLa cells were maintained in DMEM medium supplemented with 10% fetal bovine serum. H1299 and 293 cells were
The Supplementary Information (SI) Methods Cell culture and transfections H1299, U2OS, 293, HeLa cells were maintained in DMEM medium supplemented with 10% fetal bovine serum. H1299 and 293 cells were
Supplementary Information (Ha, et. al) Supplementary Figures Supplementary Fig. S1
 Supplementary Information (Ha, et. al) Supplementary Figures Supplementary Fig. S1 a His-ORMDL3 ~ 17 His-ORMDL3 GST-ORMDL3 - + - + IPTG GST-ORMDL3 ~ b Integrated Density (ORMDL3/ -actin) 0.4 0.3 0.2 0.1
Supplementary Information (Ha, et. al) Supplementary Figures Supplementary Fig. S1 a His-ORMDL3 ~ 17 His-ORMDL3 GST-ORMDL3 - + - + IPTG GST-ORMDL3 ~ b Integrated Density (ORMDL3/ -actin) 0.4 0.3 0.2 0.1
supplementary information
 DOI: 10.1038/ncb2172 Figure S1 p53 regulates cellular NADPH and lipid levels via inhibition of G6PD. (a) U2OS cells stably expressing p53 shrna or a control shrna were transfected with control sirna or
DOI: 10.1038/ncb2172 Figure S1 p53 regulates cellular NADPH and lipid levels via inhibition of G6PD. (a) U2OS cells stably expressing p53 shrna or a control shrna were transfected with control sirna or
T H E J O U R N A L O F C E L L B I O L O G Y
 T H E J O U R N A L O F C E L L B I O L O G Y Supplemental material Han et al., http://www.jcb.org/cgi/content/full/jcb.201311007/dc1 Figure S1. SIVA1 interacts with PCNA. (A) HEK293T cells were transiently
T H E J O U R N A L O F C E L L B I O L O G Y Supplemental material Han et al., http://www.jcb.org/cgi/content/full/jcb.201311007/dc1 Figure S1. SIVA1 interacts with PCNA. (A) HEK293T cells were transiently
λ N -GFP: an RNA reporter system for live-cell imaging
 λ N -GFP: an RNA reporter system for live-cell imaging Nathalie Daigle & Jan Ellenberg Supplementary Figures and Text: Supplementary Figure 1 localization in the cytoplasm. 4 λ N22-3 megfp-m9 serves as
λ N -GFP: an RNA reporter system for live-cell imaging Nathalie Daigle & Jan Ellenberg Supplementary Figures and Text: Supplementary Figure 1 localization in the cytoplasm. 4 λ N22-3 megfp-m9 serves as
Supplementary Table 1. Sequences for BTG2 and BRCA1 sirnas.
 Supplementary Table 1. Sequences for BTG2 and BRCA1 sirnas. Target Gene Non-target / Control BTG2 BRCA1 NFE2L2 Target Sequence ON-TARGET plus Non-targeting sirna # 1 (Cat# D-001810-01-05) sirna1: GAACCGACAUGCUCCCGGA
Supplementary Table 1. Sequences for BTG2 and BRCA1 sirnas. Target Gene Non-target / Control BTG2 BRCA1 NFE2L2 Target Sequence ON-TARGET plus Non-targeting sirna # 1 (Cat# D-001810-01-05) sirna1: GAACCGACAUGCUCCCGGA
Supporting Information
 Supporting Information Üstün and Börnke, 2015 Figure S1: XopJ does not display acetyltransferase activity. Autoacetylation activity in vitro. Acetylation reactions using MBP, MBP-XopJ, GST and GST-HopZ1a
Supporting Information Üstün and Börnke, 2015 Figure S1: XopJ does not display acetyltransferase activity. Autoacetylation activity in vitro. Acetylation reactions using MBP, MBP-XopJ, GST and GST-HopZ1a
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/8/404/ra120/dc1 Supplementary Materials for The subcellular localization and activity of cortactin is regulated by acetylation and interaction with Keap1 Akihiro
www.sciencesignaling.org/cgi/content/full/8/404/ra120/dc1 Supplementary Materials for The subcellular localization and activity of cortactin is regulated by acetylation and interaction with Keap1 Akihiro
Supplemental Fig. 1: PEA-15 knockdown efficiency assessed by immunohistochemistry and qpcr
 Supplemental figure legends Supplemental Fig. 1: PEA-15 knockdown efficiency assessed by immunohistochemistry and qpcr A, LβT2 cells were transfected with either scrambled or PEA-15 sirna. Cells were then
Supplemental figure legends Supplemental Fig. 1: PEA-15 knockdown efficiency assessed by immunohistochemistry and qpcr A, LβT2 cells were transfected with either scrambled or PEA-15 sirna. Cells were then
Figure S1. Verification of ihog Mutation by Protein Immunoblotting Figure S2. Verification of ihog and boi
 Figure S1. Verification of ihog Mutation by Protein Immunoblotting Extracts from S2R+ cells, embryos, and adults were analyzed by immunoprecipitation and immunoblotting with anti-ihog antibody. The Ihog
Figure S1. Verification of ihog Mutation by Protein Immunoblotting Extracts from S2R+ cells, embryos, and adults were analyzed by immunoprecipitation and immunoblotting with anti-ihog antibody. The Ihog
pt7ht vector and over-expressed in E. coli as inclusion bodies. Cells were lysed in 6 M
 Supplementary Methods MIG6 production, purification, inhibition, and kinase assays MIG6 segment 1 (30mer, residues 334 364) peptide was synthesized using standard solid-phase peptide synthesis as described
Supplementary Methods MIG6 production, purification, inhibition, and kinase assays MIG6 segment 1 (30mer, residues 334 364) peptide was synthesized using standard solid-phase peptide synthesis as described
transcription and the promoter occupancy of Smad proteins. (A) HepG2 cells were co-transfected with the wwp-luc reporter, and FLAG-tagged FHL1,
 Supplementary Data Supplementary Figure Legends Supplementary Figure 1 FHL-mediated TGFβ-responsive reporter transcription and the promoter occupancy of Smad proteins. (A) HepG2 cells were co-transfected
Supplementary Data Supplementary Figure Legends Supplementary Figure 1 FHL-mediated TGFβ-responsive reporter transcription and the promoter occupancy of Smad proteins. (A) HepG2 cells were co-transfected
Nature Structural & Molecular Biology: doi: /nsmb.1583
 Acetylation by GCN5 regulates CDC6 phosphorylation in the S-phase of the cell cycle Roberta Paolinelli 1,2, Ramiro Mendoza-Maldonado 2, Anna Cereseto 1 and Mauro Giacca 2 1 Molecular Biology Laboratory,
Acetylation by GCN5 regulates CDC6 phosphorylation in the S-phase of the cell cycle Roberta Paolinelli 1,2, Ramiro Mendoza-Maldonado 2, Anna Cereseto 1 and Mauro Giacca 2 1 Molecular Biology Laboratory,
 DOI: 10.1038/ncb3259 A Ismail et al. Supplementary Figure 1 B 60000 45000 SSC 30000 15000 Live cells 0 0 15000 30000 45000 60000 FSC- PARR 60000 45000 PARR Width 30000 FSC- 15000 Single cells 0 0 15000
DOI: 10.1038/ncb3259 A Ismail et al. Supplementary Figure 1 B 60000 45000 SSC 30000 15000 Live cells 0 0 15000 30000 45000 60000 FSC- PARR 60000 45000 PARR Width 30000 FSC- 15000 Single cells 0 0 15000
Supplementary information
 Supplementary information The E3 ligase RNF8 regulates KU80 removal and NHEJ repair Lin Feng 1, Junjie Chen 1 1 Department of Experimental Radiation Oncology, The University of Texas M. D. Anderson Cancer
Supplementary information The E3 ligase RNF8 regulates KU80 removal and NHEJ repair Lin Feng 1, Junjie Chen 1 1 Department of Experimental Radiation Oncology, The University of Texas M. D. Anderson Cancer
Supplementary Information
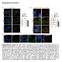 Supplementary Information Supplementary Figure S1 (a) P-cRAF colocalizes with LC3 puncta. Immunofluorescence (IF) depicting colocalization of P-cRAF (green) and LC3 puncta (red) in NIH/3T3 cells treated
Supplementary Information Supplementary Figure S1 (a) P-cRAF colocalizes with LC3 puncta. Immunofluorescence (IF) depicting colocalization of P-cRAF (green) and LC3 puncta (red) in NIH/3T3 cells treated
X2-C/X1-Y X2-C/VCAM-Y. FRET efficiency. Ratio YFP/CFP
 FRET efficiency.7.6..4.3.2 X2-C/X1-Y X2-C/VCAM-Y.1 1 2 3 Ratio YFP/CFP Supplemental Data 1. Analysis of / heterodimers in live cells using FRET. FRET saturation curves were obtained using cells transiently
FRET efficiency.7.6..4.3.2 X2-C/X1-Y X2-C/VCAM-Y.1 1 2 3 Ratio YFP/CFP Supplemental Data 1. Analysis of / heterodimers in live cells using FRET. FRET saturation curves were obtained using cells transiently
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION Legends for Supplementary Tables. Supplementary Table 1. An excel file containing primary screen data. Worksheet 1, Normalized quantification data from a duplicated screen: valid
SUPPLEMENTARY INFORMATION Legends for Supplementary Tables. Supplementary Table 1. An excel file containing primary screen data. Worksheet 1, Normalized quantification data from a duplicated screen: valid
Supporting information
 Supporting information Construction of strains and plasmids To create ptc67, a PCR product obtained with primers cc2570-162f (gcatgggcaagcttgaggacggcgtcatgt) and cc2570+512f (gaggccgtggtaccatagaggcgggcg),
Supporting information Construction of strains and plasmids To create ptc67, a PCR product obtained with primers cc2570-162f (gcatgggcaagcttgaggacggcgtcatgt) and cc2570+512f (gaggccgtggtaccatagaggcgggcg),
SUPPLEMENTARY INFORMATION
 a 14 12 Densitometry (AU) 1 8 6 4 2 t b 16 NMHC-IIA GAPDH NMHC-IIB Densitometry (AU) 14 12 1 8 6 4 2 1 nm 1 nm 1 nm 1 nm sirna 1 nm 1 nm Figure S1 S4 Quantification of protein levels. (a) The microtubule
a 14 12 Densitometry (AU) 1 8 6 4 2 t b 16 NMHC-IIA GAPDH NMHC-IIB Densitometry (AU) 14 12 1 8 6 4 2 1 nm 1 nm 1 nm 1 nm sirna 1 nm 1 nm Figure S1 S4 Quantification of protein levels. (a) The microtubule
SUPPLEMENTARY INFORMATION. Small molecule activation of the TRAIL receptor DR5 in human cancer cells
 SUPPLEMENTARY INFORMATION Small molecule activation of the TRAIL receptor DR5 in human cancer cells Gelin Wang 1*, Xiaoming Wang 2, Hong Yu 1, Shuguang Wei 1, Noelle Williams 1, Daniel L. Holmes 1, Randal
SUPPLEMENTARY INFORMATION Small molecule activation of the TRAIL receptor DR5 in human cancer cells Gelin Wang 1*, Xiaoming Wang 2, Hong Yu 1, Shuguang Wei 1, Noelle Williams 1, Daniel L. Holmes 1, Randal
Fig. S1. Effect of p120-catenin overexpression on the interaction of SCUBE2 with E-cadherin. The expression plasmid encoding FLAG.
 Fig. S1. Effect of p120-catenin overexpression on the interaction of SCUBE2 with E-cadherin. The expression plasmid encoding FLAG.SCUBE2, E-cadherin.Myc, or HA.p120-catenin was transfected in a combination
Fig. S1. Effect of p120-catenin overexpression on the interaction of SCUBE2 with E-cadherin. The expression plasmid encoding FLAG.SCUBE2, E-cadherin.Myc, or HA.p120-catenin was transfected in a combination
hnrnp C promotes APP translation by competing with FMRP for APP mrna recruitment to P bodies
 hnrnp C promotes APP translation by competing with for APP mrna recruitment to P bodies Eun Kyung Lee 1, Hyeon Ho Kim 1, Yuki Kuwano 1, Kotb Abdelmohsen 1, Subramanya Srikantan 1, Sarah S. Subaran 2, Marc
hnrnp C promotes APP translation by competing with for APP mrna recruitment to P bodies Eun Kyung Lee 1, Hyeon Ho Kim 1, Yuki Kuwano 1, Kotb Abdelmohsen 1, Subramanya Srikantan 1, Sarah S. Subaran 2, Marc
supplementary information
 DOI: 1.138/ncb1839 a b Control 1 2 3 Control 1 2 3 Fbw7 Smad3 1 2 3 4 1 2 3 4 c d IGF-1 IGF-1Rβ IGF-1Rβ-P Control / 1 2 3 4 Real-time RT-PCR Relative quantity (IGF-1/ mrna) 2 1 IGF-1 1 2 3 4 Control /
DOI: 1.138/ncb1839 a b Control 1 2 3 Control 1 2 3 Fbw7 Smad3 1 2 3 4 1 2 3 4 c d IGF-1 IGF-1Rβ IGF-1Rβ-P Control / 1 2 3 4 Real-time RT-PCR Relative quantity (IGF-1/ mrna) 2 1 IGF-1 1 2 3 4 Control /
HPV E6 oncoprotein targets histone methyltransferases for modulating specific. Chih-Hung Hsu, Kai-Lin Peng, Hua-Ci Jhang, Chia-Hui Lin, Shwu-Yuan Wu,
 1 HPV E oncoprotein targets histone methyltransferases for modulating specific gene transcription 3 5 Chih-Hung Hsu, Kai-Lin Peng, Hua-Ci Jhang, Chia-Hui Lin, Shwu-Yuan Wu, Cheng-Ming Chiang, Sheng-Chung
1 HPV E oncoprotein targets histone methyltransferases for modulating specific gene transcription 3 5 Chih-Hung Hsu, Kai-Lin Peng, Hua-Ci Jhang, Chia-Hui Lin, Shwu-Yuan Wu, Cheng-Ming Chiang, Sheng-Chung
Supplementary methods
 Supplementary methods Cell culture, infection, transfection, and RNA interference HEK293 cells and its derivatives were grown in DMEM supplemented with 10% FBS. Various constructs were introduced into
Supplementary methods Cell culture, infection, transfection, and RNA interference HEK293 cells and its derivatives were grown in DMEM supplemented with 10% FBS. Various constructs were introduced into
SUPPLEMENTARY INFORMATION
 doi: 10.1038/nature06721 SUPPLEMENTARY INFORMATION. Supplemental Figure Legends Supplemental Figure 1 The distribution of hatx-1[82q] in Cos7 cells. Cos7 cells are co-transfected with hatx-1[82q]-gfp (green)
doi: 10.1038/nature06721 SUPPLEMENTARY INFORMATION. Supplemental Figure Legends Supplemental Figure 1 The distribution of hatx-1[82q] in Cos7 cells. Cos7 cells are co-transfected with hatx-1[82q]-gfp (green)
Supplemental Table 1: Sequences of real time PCR primers. Primers were intronspanning
 Symbol Accession Number Sense-primer (5-3 ) Antisense-primer (5-3 ) T a C ACTB NM_001101.3 CCAGAGGCGTACAGGGATAG CCAACCGCGAGAAGATGA 57 HSD3B2 NM_000198.3 CTTGGACAAGGCCTTCAGAC TCAAGTACAGTCAGCTTGGTCCT 60
Symbol Accession Number Sense-primer (5-3 ) Antisense-primer (5-3 ) T a C ACTB NM_001101.3 CCAGAGGCGTACAGGGATAG CCAACCGCGAGAAGATGA 57 HSD3B2 NM_000198.3 CTTGGACAAGGCCTTCAGAC TCAAGTACAGTCAGCTTGGTCCT 60
42 fl organelles = 34.5 fl (1) 3.5X X 0.93 = 78,000 (2)
 SUPPLEMENTAL DATA Supplementary Experimental Procedures Fluorescence Microscopy - A Zeiss Axiovert 200M microscope equipped with a Zeiss 100x Plan- Apochromat (1.40 NA) DIC objective and Hamamatsu Orca
SUPPLEMENTAL DATA Supplementary Experimental Procedures Fluorescence Microscopy - A Zeiss Axiovert 200M microscope equipped with a Zeiss 100x Plan- Apochromat (1.40 NA) DIC objective and Hamamatsu Orca
T H E J O U R N A L O F C E L L B I O L O G Y
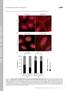 T H E J O U R N A L O F C E L L B I O L O G Y Supplemental material Feng et al., http://www.jcb.org/cgi/content/full/jcb.201408079/dc1 Figure S1. A modest elevation of disulfide-bonded K14 in primary mouse
T H E J O U R N A L O F C E L L B I O L O G Y Supplemental material Feng et al., http://www.jcb.org/cgi/content/full/jcb.201408079/dc1 Figure S1. A modest elevation of disulfide-bonded K14 in primary mouse
Supplementary Figure 1 - Characterization of rbag3 binding on macrophages cell surface.
 Supplementary Figure 1 - Characterization of rbag3 binding on macrophages cell surface. (a) Human PDAC cell lines were treated as indicated in Figure 1 panel F. Cells were analyzed for FITC-rBAG3 binding
Supplementary Figure 1 - Characterization of rbag3 binding on macrophages cell surface. (a) Human PDAC cell lines were treated as indicated in Figure 1 panel F. Cells were analyzed for FITC-rBAG3 binding
Attenuation of synaptic toxicity and MARK4/PAR1-mediated Tau phosphorylation by
 Supplementary Methods and Figures Attenuation of synaptic toxicity and MARK4/PAR1-mediated Tau phosphorylation by methylene blue for Alzheimer s disease treatment Wenchao Sun 1, Seongsoo Lee 1,2, Xiaoran
Supplementary Methods and Figures Attenuation of synaptic toxicity and MARK4/PAR1-mediated Tau phosphorylation by methylene blue for Alzheimer s disease treatment Wenchao Sun 1, Seongsoo Lee 1,2, Xiaoran
T H E J O U R N A L O F C E L L B I O L O G Y
 Supplemental material Moutin et al., http://www.jcb.org/cgi/content/full/jcb.201110101/dc1 T H E J O U R N A L O F C E L L B I O L O G Y Figure S1. Tagged Homer1a and Homer are functional and display different
Supplemental material Moutin et al., http://www.jcb.org/cgi/content/full/jcb.201110101/dc1 T H E J O U R N A L O F C E L L B I O L O G Y Figure S1. Tagged Homer1a and Homer are functional and display different
Supplementary information to accompany: A novel role for the DNA repair gene Rad51 in Netrin-1 signalling
 Supplementary information to accompany: A novel role for the DNA repair gene Rad51 in Netrin-1 signalling Glendining KA 1, Markie D 2, Gardner RJM 4, Franz EA 3, Robertson SP 4, Jasoni CL 1 Supplementary
Supplementary information to accompany: A novel role for the DNA repair gene Rad51 in Netrin-1 signalling Glendining KA 1, Markie D 2, Gardner RJM 4, Franz EA 3, Robertson SP 4, Jasoni CL 1 Supplementary
Antibodies against PCNA were previously described [1]. To deplete PCNA from Xenopus egg
![Antibodies against PCNA were previously described [1]. To deplete PCNA from Xenopus egg Antibodies against PCNA were previously described [1]. To deplete PCNA from Xenopus egg](/thumbs/84/90687567.jpg) Supplementary information Supplementary methods PCNA antibody and immunodepletion Antibodies against PCNA were previously described [1]. To deplete PCNA from Xenopus egg extracts, one volume of protein
Supplementary information Supplementary methods PCNA antibody and immunodepletion Antibodies against PCNA were previously described [1]. To deplete PCNA from Xenopus egg extracts, one volume of protein
supplementary information
 DOI: 10.1038/ncb1977 Figure S1 a. Immunofluorescence analysis of IFT20 localization in PBL costained with anti-β-tubulin antibodies. b. Immunofluorescence analysis of IFT20 localization in Jurkat cells,
DOI: 10.1038/ncb1977 Figure S1 a. Immunofluorescence analysis of IFT20 localization in PBL costained with anti-β-tubulin antibodies. b. Immunofluorescence analysis of IFT20 localization in Jurkat cells,
Cytotoxicity of Botulinum Neurotoxins Reveals a Direct Role of
 Supplementary Information Cytotoxicity of Botulinum Neurotoxins Reveals a Direct Role of Syntaxin 1 and SNAP-25 in Neuron Survival Lisheng Peng, Huisheng Liu, Hongyu Ruan, William H. Tepp, William H. Stoothoff,
Supplementary Information Cytotoxicity of Botulinum Neurotoxins Reveals a Direct Role of Syntaxin 1 and SNAP-25 in Neuron Survival Lisheng Peng, Huisheng Liu, Hongyu Ruan, William H. Tepp, William H. Stoothoff,
Supporting Online Material Material and Methods
 Supporting Online Material Material and Methods Live-cell DIC recordings PtK 1 cells (ATCC, Manassas, VA) were filmed on a Nikon (Nikon Instruments, Melville, NY) inverted microscope equipped with 60x
Supporting Online Material Material and Methods Live-cell DIC recordings PtK 1 cells (ATCC, Manassas, VA) were filmed on a Nikon (Nikon Instruments, Melville, NY) inverted microscope equipped with 60x
Sequence of C-terminal tail regions of myosin heavy chains in class XI of Nicotiana benthamiana (Nb
 Fig. S1 Sequence of C-terminal tail regions of myosin heavy chains in class XI of Nicotiana benthamiana (Nb myosin XI-2, -F and K) and BY-2 cell (Nt 170-kD myosin and Nt 175-kD myosin). Amino acids identical
Fig. S1 Sequence of C-terminal tail regions of myosin heavy chains in class XI of Nicotiana benthamiana (Nb myosin XI-2, -F and K) and BY-2 cell (Nt 170-kD myosin and Nt 175-kD myosin). Amino acids identical
15 June 2011 Supplementary material Bagriantsev et al.
 Supplementary Figure S1 Characterization of K 2P 2.1 (TREK-1) GOF mutants A, Distribution of the positions of mutated nucleotides, represented by a red x, from a pool of 18 unselected K 2P 2.1 (KCNK2)
Supplementary Figure S1 Characterization of K 2P 2.1 (TREK-1) GOF mutants A, Distribution of the positions of mutated nucleotides, represented by a red x, from a pool of 18 unselected K 2P 2.1 (KCNK2)
Otefin, a Nuclear Membrane Protein, Determines
 Developmental Cell 14 Supplemental Data Otefin, a Nuclear Membrane Protein, Determines the Fate of Germline Stem Cells in Drosophila via Interaction with Smad Complexes Xiaoyong Jiang, Laixin Xia, Dongsheng
Developmental Cell 14 Supplemental Data Otefin, a Nuclear Membrane Protein, Determines the Fate of Germline Stem Cells in Drosophila via Interaction with Smad Complexes Xiaoyong Jiang, Laixin Xia, Dongsheng
Supporting Information
 Supporting Information Tal et al. 10.1073/pnas.0807694106 SI Materials and Methods VSV Infection and Quantification. Infection was carried out by seeding 5 10 5 MEF cells per well in a 6-well plate and
Supporting Information Tal et al. 10.1073/pnas.0807694106 SI Materials and Methods VSV Infection and Quantification. Infection was carried out by seeding 5 10 5 MEF cells per well in a 6-well plate and
Supplementary Fig.1 Luton
 Supplementary Fig.1 Luton a 175 Brain Thymus Spleen Small Intestine Kidney Testis HeLa b 250 Lung Kidney MDCK c EFA6B si Control si Mismatch #637 #1564 #1770 83 62 47.5 175 IB: anti-efa6b #B1 130 66 Lysates
Supplementary Fig.1 Luton a 175 Brain Thymus Spleen Small Intestine Kidney Testis HeLa b 250 Lung Kidney MDCK c EFA6B si Control si Mismatch #637 #1564 #1770 83 62 47.5 175 IB: anti-efa6b #B1 130 66 Lysates
(a) Immunoblotting to show the migration position of Flag-tagged MAVS
 Supplementary Figure 1 Characterization of six MAVS isoforms. (a) Immunoblotting to show the migration position of Flag-tagged MAVS isoforms. HEK293T Mavs -/- cells were transfected with constructs expressing
Supplementary Figure 1 Characterization of six MAVS isoforms. (a) Immunoblotting to show the migration position of Flag-tagged MAVS isoforms. HEK293T Mavs -/- cells were transfected with constructs expressing
Supplemental Figure 1. Alignment of the NbGAPC amino acid sequences with their Arabidopsis homologues.
 Supplemental Figure 1. Alignment of the NbGAPC amino acid sequences with their Arabidopsis homologues. Homologs from N. benthamiana (NbGAPC1, NbGAPC2, NbGAPC3), Arabidopsis (AtGAPC1, AT3G04120; AtGAPC2,
Supplemental Figure 1. Alignment of the NbGAPC amino acid sequences with their Arabidopsis homologues. Homologs from N. benthamiana (NbGAPC1, NbGAPC2, NbGAPC3), Arabidopsis (AtGAPC1, AT3G04120; AtGAPC2,
Supplementary Information: Materials and Methods. Immunoblot and immunoprecipitation. Cells were washed in phosphate buffered
 Supplementary Information: Materials and Methods Immunoblot and immunoprecipitation. Cells were washed in phosphate buffered saline (PBS) and lysed in TNN lysis buffer (50mM Tris at ph 8.0, 120mM NaCl
Supplementary Information: Materials and Methods Immunoblot and immunoprecipitation. Cells were washed in phosphate buffered saline (PBS) and lysed in TNN lysis buffer (50mM Tris at ph 8.0, 120mM NaCl
Supplementary Figure 1. Immunoprecipitation of synthetic SUMOm-remnant peptides using UMO monoclonal antibody. (a) LC-MS analyses of tryptic
 Supplementary Figure 1. Immunoprecipitation of synthetic SUMOm-remnant peptides using UMO 1-7-7 monoclonal antibody. (a) LC-MS analyses of tryptic digest from HEK293 cells spiked with 6 SUMOmremnant peptides
Supplementary Figure 1. Immunoprecipitation of synthetic SUMOm-remnant peptides using UMO 1-7-7 monoclonal antibody. (a) LC-MS analyses of tryptic digest from HEK293 cells spiked with 6 SUMOmremnant peptides
Title: The cleaved FAS ligand activates the Na + /H + exchanger NHE1 through. Akt/ROCK1 to stimulate cell motility.
 Title: The cleaved FAS ligand activates the Na + /H + exchanger NHE through Akt/ROCK to stimulate cell motility. Authors : Monet Michael, Poët Mallorie, Tauzin Sébastien 2,#, Fouqué Amélie 3, Cophignon
Title: The cleaved FAS ligand activates the Na + /H + exchanger NHE through Akt/ROCK to stimulate cell motility. Authors : Monet Michael, Poët Mallorie, Tauzin Sébastien 2,#, Fouqué Amélie 3, Cophignon
Supplementary Information
 Supplementary Information COP1 E3 ligase protects HYL1 to retain microrna biogenesis Seok Keun Cho 1, Samir Ben Chaabane 1, Pratik Shah 1, Christian Peter Poulsen 1, and Seong Wook Yang 1 * 1 Laboratory
Supplementary Information COP1 E3 ligase protects HYL1 to retain microrna biogenesis Seok Keun Cho 1, Samir Ben Chaabane 1, Pratik Shah 1, Christian Peter Poulsen 1, and Seong Wook Yang 1 * 1 Laboratory
