#26 - Gene Prediction 10/22/07
|
|
|
- Nickolas Owens
- 5 years ago
- Views:
Transcription
1 BCB 444/544 Required Reading (before lecture) Lecture 26 Mon Oct 22 - Lecture 26 Gene Prediction Chp 8 - pp Gene Prediction Wed Oct 24 - Lecture 27 (will not be covered on Exam 2) Regulatory Element Prediction Chp 9 - pp Thurs Oct 25 - Review Session & Project Planning #26_Oct22 Fri Oct 26 - EXAM Assignments & Announcements BCB 544 "Team" Projects Sun Oct 21 - Study Guide for Exam 2 was posted Mon Oct 22 - HW#4 Due (no "correct" answer to post) Thu Oct 25 - Lab = Optional Review Session for Exam 544 Project Planning/Consult with DD & MT Fri Oct 26 - Exam 2 - Will cover: Lectures (thru Mon Sept 17) Labs 5-8 HW# 3 & 4 All assigned reading: Chps 6 (beginning with HMMs), 7-8, Eddy: What is an HMM Ginalski: Practical Lessons 544 Extra HW#2 is next step in Team Projects Write ~ 1 page outline Schedule meeting with Michael & Drena to discuss topic Read a few papers Write a more detailed plan You may work alone if you prefer Last week of classes will be devoted to Projects Written reports due: Mon Dec 3 (no class that day) Oral presentations (15-20') will be: Wed-Fri Dec 5,6,7 1 or 2 teams will present during each class period See Guidelines for Projects posted online 3 4 BCB 544 Only: New Homework Assignment Seminars this Week 544 Extra#2 (posted online Thurs?) No - sorry! sent by on Sat BCB List of URLs for Seminars related to Bioinformatics: Due: PART 1 - ASAP PART 2 - Fri Nov 2 by 5 PM Oct 25 Thur - BBMB Seminar 4:10 in 1414 MBB Dave Segal UC Davis Zinc Finger Protein Design Part 1 - Brief outline of Project, to Drena & Michael after response/approval, then: Part 2 - More detailed outline of project Read a few papers and summarize status of problem Schedule meeting with Drena & Michael to discuss ideas Oct 19 Fri - BCB Faculty Seminar 2:10 in 102 ScI Guang Song ComS, ISU Probing functional mechanisms by structure-based modeling and simulations 5 6 BCB 444/544 Fall 07 Dobbs 1
2 Chp 16 - RNA Structure Prediction Covalent & non-covalent bonds in RNA SECTION V STRUCTURAL BIOINFORMATICS Xiong: Chp 16 RNA Structure Prediction (Terribilini) RNA Function Types of RNA Structures RNA Secondary Structure Prediction Methods Ab Initio Approach Comparative Approach Performance Evaluation Primary: Covalent bonds Secondary/Tertiary Non-covalent bonds H-bonds (base-pairing) Base stacking 7 Fig 6.2 Baxevanis & Ouellette RNA Pseudoknots & Tetraloops Base Pairing in RNA Often have important regulatory or catalytic functions Pseudoknot Tetraloop G-C, A-U, G-U ("wobble") & many variants See: IMB Image Library of Biological Molecules Review/Annual-Reports/1995/images/rna.gif huang/qd/mckay_hr.gif 9 10 RNA Secondary Structure Prediction Methods Two (three, recently) main types of methods: 1. Ab initio - based on calculating most energetically favorable secondary structure(s) Energy minimization (thermodynamics) 2. Comparative approach - based on comparisons of multiple evolutionarily-related RNA sequences Sequence comparison (co-variation) 3. Combined computational & experimental Use experimental constraints when available RNA Secondary structure prediction - 3 3) Combined experimental & computational Experiments: Map single-stranded vs doublestranded regions in folded RNA How? Enzymes: S1 nuclease, T1 RNase Chemicals: kethoxal, DMS, OH Software: Mfold Sfold RNAStructure RNAFold RNAlifold Kethoxal modification (mild) (strong) DMS modification (mild) (strong) DMS G BCB 444/544 Fall 07 Dobbs 2
3 Ab Initio Prediction: Clarifications Energy minimization: What are the rules? Free energy is calculated based on parameters determined in the wet lab Correction: Use known energy associated with each type of nearest-neighbor pair (base-stacking) (not base-pair) Base-pair formation is not independent: multiple base-pairs adjacent to each other are more favorable than individual base-pairs - cooperative - because of base-stacking interactions Bulges and loops adjacent to base-pairs have a free energy penalty A U A U A U U A Basepair ΔG = -1.2 kcal/mole Basepair A=U A=U A=U U=A ΔG = -1.6 kcal/mole What gives here? 13 C Staben C Staben 2005 Energy minimization calculations: Base-stacking is critical AA UU -1.2 AU or UA UA AU -1.6 CG GC -3.0 GC CG -4.3 AG, AC, CA, GA UC, UG, GU, CU -2.1 GU UG -0.3 CC GG -4.8 XG, GX YU, UY 0 - Tinocco et al. 15 Ab Initio Energy Calculation Search for all possible base-pairing patterns Calculate total energy of each structure based on all stabilizing and destabilizing forces Total free energy for a specific RNA conformation = Sum of incremental energy terms for: helical stacking (sequence dependent) loop initiation unpaired stacking (favorable "increments" are < 0) Fig 6.3 Baxevanis & Ouellette Dynamic Programming 3 - Popular Programs that use Combined Computational Experimental Approaches Finding optimal secondary structure is difficult - lots of possibilities Compare RNA sequence with itself Apply scoring scheme based on energy parameters for base stacking, cooperativity, and penalties for destabilizing forces (loops, bulges) Find path that represents most energetically favorable secondary structure Mfold Sfold RNAStructure RNAFold RNAlifold BCB 444/544 Fall 07 Dobbs 3
4 Comparison of Predictions for Single RNA using Different Methods Comparison of Mfold Predictions: -/+ Constraints Mfold kcal/mol Sfold kcal/mol RNAstructure kcal/mol RNAfold kcal/mol Mfold kcal/mol Mfold plus constraints kcal/mol JH Lee 2007 JH Lee Performance Evaluation Chp 8 - Gene Prediction Ab initio methods? correlation coefficient = 20-60% Comparative approaches? correlation coefficient = 20-80% Programs that require user to supply MSA are more accurate Comparative programs are consistently more accurate than ab initio Base-pairs predicted by comparative sequence analysis for large & small subunit rrnas are 97% accurate when compared with high resolution crystal structures! - Gutell, Pace SECTION III GENE AND PROMOTER PREDICTION Xiong: Chp 8 Gene Prediction Categories of Gene Prediction Programs Gene Prediction in Prokaryotes Gene Prediction in Eukaryotes BEST APPROACH? Methods that combine computational prediction (ab initio & comparative) with experimental constraints (from chemical/enzymatic modification studies) What is a Gene? Gene Finding What is a gene? segment of DNA, some of which is "structural," i.e., transcribed to give a functional RNA product, & some of which is "regulatory" Problem: Given a new genomic DNA sequence, identify coding regions and their predicted RNA and protein sequences ATTACCATGGGGCAGGGTCAGATATAATGCCCTCATTTT Genes can encode: ATTACCATGGGGCAGGGTCAGATATAATGCCCTCATTTT mrna (for protein) other types of RNA (trna, rrna, mirna, etc.) Genes differ in eukaryotes vs prokaryotes (& archaea), both structure & regulation Steps: 1. Search against protein / EST database 2. Apply gene prediction programs (many programs available) 3. Analyze regulatory regions BCB 444/544 Fall 07 Dobbs 4
5 Gene Prediction in Prokaryotes vs Eukaryotes Eukaryotes Large genomes bp Often less than 2% coding Complicated gene structure (splicing, long exons) Prediction success 50-95% ATG Splice sites TAA Prokaryotes Small genomes bp About 90% of genome is coding Simple gene structure Prediction success ~99% Start codon ATG Stop codon TAA DNA "Signals" Used by Gene Finding Algorithms 1. Exploit the regular gene structure ATG Exon1 Intron1 Exon2 ExonN STOP 2. Recognize coding bias CAG-CGA-GAC-TAT-TTA-GAT-AAC-ACA-CAT-GAA- 3. Recognize splice sites Intron cagt Exon ggtgag Intron 4. Model the duration of regions Introns tend to be much longer than exons, in mammals Exons are biased to have a given minimum length UTR Promotor Exons Introns UTR Promotor Open reading frame (ORF) 5. Use cross-species comparison Gene structure is conserved in mammals Exons are more similar (~85%) than introns Computational Gene Finding Approaches Examples of Gene Prediction Software Ab initio methods Search by signal: find DNA sequences involved in gene expression. Search by content: Test statistical properties distinguishing coding from non-coding DNA Similarity based methods Database search: exploit similarity to proteins, ESTs, and cdnas Comparative genomics: exploit aligned genomes Do other organisms have similar sequence? Hybrid methods - best 27 Ab initio Genscan, GeneMark.hmm, Genie, GeneID Similarity-based BLAST, Procrustes Hybrids GeneSeqer, GenomeScan, GenieEST, Twinscan, SGP, ROSETTA, CEM, TBLASTX, SLAM. BEST? Ab initio - Genescan (according to some assessments) Hybrid - GeneSeqer But depends on organism & specific task Lists of Gene Prediction Software Synthesis & Processing of Eukaryotic mrna DN Gene in DNA exon 1 intron exon 2 intron exon 3 1' transcript (RNA) Transcription Mature mrna 7Me G Splicing (remove introns) Capping & polyadenylation AAAAA Export to cytoplasm m What are cdnas & ESTs? cdna libraries are important for determining gene structure & studying regulation of gene expression Isolate RNA (always from a specific organism, region, and time point) insert Convert RNA to complementary DNA (with reverse transcriptase) Clone into cdna vector Sequence the cdna inserts vector Short cdnas are called ESTs or Expressed Sequence Tags ESTs are strong evidence for genes Full-length cdnas can be difficult to obtain BCB 444/544 Fall 07 Dobbs 5
6 UniGene: Unique genes via ESTs Gene Prediction Find UniGene at NCBI: UniGene clusters contain many ESTs UniGene data come from many cdna libraries. When you look up a gene in UniGene, you can obtain information re: level & tissue distribution of expression Overview of steps & strategies What sequence signals can be used? What other types of information can be used? Algorithms HMMs, Bayesian models, neural nets Gene prediction software 3 major types many, many programs! Overview of Gene Prediction Strategies What sequence signals can be used? Transcription: TF binding sites, promoter, initiation site, terminator, GC islands, etc. Processing signals: Splice donor/acceptors, polya signal Translation: Start (AUG = Met) & stop (UGA,UUA, UAG) ORFs, codon usage What other types of information can be used? Homology (sequence comparison, BLAST) cdnas & ESTs (experimental data, pairwise alignment) Gene prediction: Eukaryotes vs prokaryotes Gene prediction is easier in microbial genomes Why? Smaller genomes Simpler gene structures Many more sequenced genomes! (for comparative approaches) Many microbial genomes have been fully sequenced & whole-genome "gene structure" and "gene function" annotations are available e.g., GeneMark.hmm TIGR Comprehensive Microbial Resource (CMR) NCBI Microbial Genomes Predicting Genes - Basic steps: Predicting Genes - Details: Obtain genomic sequence BLAST it! Perform database similarity search (with EST & cdna databases, if available) Translate in all 6 reading frames (i.e., "6-frame translation") Compare with protein sequence databases Use Gene Prediction software to locate genes Analyze regulatory sequences Refine gene prediction 1. 1st, mask to "remove" repetitive elements (ALUs, etc.) 2. Perform database search on translated DNA (BlastX,TFasta) 3. Use several programs to predict genes (GENSCAN, GeneMark.hmm, GeneSeqer) 4. Search for functional motifs in translated ORFs (Blocks, Motifs, etc.) & in neighboring DNA sequences 5. Repeat BCB 444/544 Fall 07 Dobbs 6
7 GeneSeqer - Brendel et al.- ISU Spliced Alignment Algorithm Brendel et al (2004) Bioinformatics 20: 1157 Perform pairwise alignment with large gaps in one sequence (due to introns) Align genomic DNA with cdna, ESTs, protein sequences Score semi-conserved sequences at splice junctions Using Bayesian model or MM Score coding constraints in translated exons Using a Bayesian model or MM GT Donor Intron AG Acceptor Splice sites Genomic DNA Protein Brendel - Spliced Alignment II: Compare with protein probes Start codon Stop codon Splice Site Detection Information content vs position Do DNA sequences surrounding splice "consensus" sequences contribute to splicing signal? YES Information Content I i : I = + f log ( f ) i " 2 ib 2 B! U, C, A, G Extent of Splice Signal Window: I i! I " i: ith position in sequence Ī: avg information content over all positions >20 nt from splice site σ Ī : avg sample standard deviation of Ī I ib Human T2_GT Human T2_AG Which sequences are exons & which are introns? How can you tell? Brendel et al (2004) Bioinformatics 20: Markov Model for Spliced Alignment P ΔG P ΔG (1-P ΔG )(1-P D(n+1) ) e n e n+1 (1-P ΔG )P D(n+1) P A(n) P ΔG (1-P ΔG )P D(n+1) i n i n+1 1-P A(n) 41 BCB 444/544 Fall 07 Dobbs 7
Gene Prediction 10/21/05
 Gene Prediction 1/21/5 1/21/5 Gene Prediction Announcements Eam 2 - net Friday Posted online: Eam 2 Study Guide 544 Reading Assignment (2 papers) (formerly Gene Prediction - ) 1/21/5 D Dobbs ISU - BCB
Gene Prediction 1/21/5 1/21/5 Gene Prediction Announcements Eam 2 - net Friday Posted online: Eam 2 Study Guide 544 Reading Assignment (2 papers) (formerly Gene Prediction - ) 1/21/5 D Dobbs ISU - BCB
Gene Regulation 10/19/05
 10/19/05 Gene Regulation (formerly Gene Prediction - 2) Gene Prediction & Regulation Mon - Overview & Gene structure review: Eukaryotes vs prokaryotes Wed - Regulatory regions: Promoters & enhancers -
10/19/05 Gene Regulation (formerly Gene Prediction - 2) Gene Prediction & Regulation Mon - Overview & Gene structure review: Eukaryotes vs prokaryotes Wed - Regulatory regions: Promoters & enhancers -
#28 - Promoter Prediction 10/29/07
 BCB 444/544 Required Reading (before lecture) Lecture 28 Mon Oct 29 - Lecture 28 Promoter & Regulatory Element Prediction Chp 9 - pp 113-126 Gene Prediction - finish it Wed Oct 30 - Lecture 29 Phylogenetics
BCB 444/544 Required Reading (before lecture) Lecture 28 Mon Oct 29 - Lecture 28 Promoter & Regulatory Element Prediction Chp 9 - pp 113-126 Gene Prediction - finish it Wed Oct 30 - Lecture 29 Phylogenetics
GenBank Growth. In 2003 ~ 31 million sequences ~ 37 billion base pairs
 Gene Finding GenBank Growth GenBank Growth In 2003 ~ 31 million sequences ~ 37 billion base pairs GenBank: Exponential Growth Growth of GenBank in billions of base pairs from release 3 in April of 1994
Gene Finding GenBank Growth GenBank Growth In 2003 ~ 31 million sequences ~ 37 billion base pairs GenBank: Exponential Growth Growth of GenBank in billions of base pairs from release 3 in April of 1994
Gene Identification in silico
 Gene Identification in silico Nita Parekh, IIIT Hyderabad Presented at National Seminar on Bioinformatics and Functional Genomics, at Bioinformatics centre, Pondicherry University, Feb 15 17, 2006. Introduction
Gene Identification in silico Nita Parekh, IIIT Hyderabad Presented at National Seminar on Bioinformatics and Functional Genomics, at Bioinformatics centre, Pondicherry University, Feb 15 17, 2006. Introduction
Promoter Prediction (really) 10/26/05
 10/26/05 Promoter Prediction (really!) Announcements BCB Link for Seminar Schedules (updated) http://www.bcb.iastate.edu/seminars/inde.html Seminar (Fri Oct 28) 12:10 PM BCB Faculty Seminar in E164 Lagomarcino
10/26/05 Promoter Prediction (really!) Announcements BCB Link for Seminar Schedules (updated) http://www.bcb.iastate.edu/seminars/inde.html Seminar (Fri Oct 28) 12:10 PM BCB Faculty Seminar in E164 Lagomarcino
Genome annotation & EST
 Genome annotation & EST What is genome annotation? The process of taking the raw DNA sequence produced by the genome sequence projects and adding the layers of analysis and interpretation necessary
Genome annotation & EST What is genome annotation? The process of taking the raw DNA sequence produced by the genome sequence projects and adding the layers of analysis and interpretation necessary
Lecture 15. Promoters, TFs. #15_Sept26
 BCB 444/544 Lecture 15 More Review: RNA, Proteins, Promoters, TFs Next time: Profiles & Hidden Markov Models (HMMs) #15_Sept26 BCB 444/544 F07 ISU Dobbs #15 - RNA, Proteins, Promoters, TFs 9/26/07 1 Required
BCB 444/544 Lecture 15 More Review: RNA, Proteins, Promoters, TFs Next time: Profiles & Hidden Markov Models (HMMs) #15_Sept26 BCB 444/544 F07 ISU Dobbs #15 - RNA, Proteins, Promoters, TFs 9/26/07 1 Required
Gene Prediction. Mario Stanke. Institut für Mikrobiologie und Genetik Abteilung Bioinformatik. Gene Prediction p.
 Gene Prediction Mario Stanke mstanke@gwdg.de Institut für Mikrobiologie und Genetik Abteilung Bioinformatik Gene Prediction p.1/23 Why Predict Genes with a Computer? tons of data 39/250 eukaryotic/prokaryotic
Gene Prediction Mario Stanke mstanke@gwdg.de Institut für Mikrobiologie und Genetik Abteilung Bioinformatik Gene Prediction p.1/23 Why Predict Genes with a Computer? tons of data 39/250 eukaryotic/prokaryotic
132 Grundlagen der Bioinformatik, SoSe 14, D. Huson, June 22, This exposition is based on the following source, which is recommended reading:
 132 Grundlagen der Bioinformatik, SoSe 14, D. Huson, June 22, 214 1 Gene Prediction Using HMMs This exposition is based on the following source, which is recommended reading: 1. Chris Burge and Samuel
132 Grundlagen der Bioinformatik, SoSe 14, D. Huson, June 22, 214 1 Gene Prediction Using HMMs This exposition is based on the following source, which is recommended reading: 1. Chris Burge and Samuel
Grundlagen der Bioinformatik, SoSe 11, D. Huson, July 4, This exposition is based on the following source, which is recommended reading:
 Grundlagen der Bioinformatik, SoSe 11, D. Huson, July 4, 211 155 12 Gene Prediction Using HMMs This exposition is based on the following source, which is recommended reading: 1. Chris Burge and Samuel
Grundlagen der Bioinformatik, SoSe 11, D. Huson, July 4, 211 155 12 Gene Prediction Using HMMs This exposition is based on the following source, which is recommended reading: 1. Chris Burge and Samuel
Genscan. The Genscan HMM model Training Genscan Validating Genscan. (c) Devika Subramanian,
 Genscan The Genscan HMM model Training Genscan Validating Genscan (c) Devika Subramanian, 2009 96 Gene structure assumed by Genscan donor site acceptor site (c) Devika Subramanian, 2009 97 A simple model
Genscan The Genscan HMM model Training Genscan Validating Genscan (c) Devika Subramanian, 2009 96 Gene structure assumed by Genscan donor site acceptor site (c) Devika Subramanian, 2009 97 A simple model
Annotating the Genome (H)
 Annotating the Genome (H) Annotation principles (H1) What is annotation? In general: annotation = explanatory note* What could be useful as an annotation of a DNA sequence? an amino acid sequence? What
Annotating the Genome (H) Annotation principles (H1) What is annotation? In general: annotation = explanatory note* What could be useful as an annotation of a DNA sequence? an amino acid sequence? What
Computational gene finding
 Computational gene finding Devika Subramanian Comp 470 Outline (3 lectures) Lec 1 Lec 2 Lec 3 The biological context Markov models and Hidden Markov models Ab-initio methods for gene finding Comparative
Computational gene finding Devika Subramanian Comp 470 Outline (3 lectures) Lec 1 Lec 2 Lec 3 The biological context Markov models and Hidden Markov models Ab-initio methods for gene finding Comparative
Lecture 10. Ab initio gene finding
 Lecture 10 Ab initio gene finding Uses of probabilistic sequence Segmentation models/hmms Multiple alignment using profile HMMs Prediction of sequence function (gene family models) ** Gene finding ** Review
Lecture 10 Ab initio gene finding Uses of probabilistic sequence Segmentation models/hmms Multiple alignment using profile HMMs Prediction of sequence function (gene family models) ** Gene finding ** Review
How to design an HMM for a new problem. HMM model structure. Inherent limitation of HMMs. Duration modeling. Duration modeling
 How to design an HMM for a new problem Architecture/topology design: What are the states, observation symbols, and the topology of the state transition graph? Learning/Training: Fully annotated or partially
How to design an HMM for a new problem Architecture/topology design: What are the states, observation symbols, and the topology of the state transition graph? Learning/Training: Fully annotated or partially
DNA is normally found in pairs, held together by hydrogen bonds between the bases
 Bioinformatics Biology Review The genetic code is stored in DNA Deoxyribonucleic acid. DNA molecules are chains of four nucleotide bases Guanine, Thymine, Cytosine, Adenine DNA is normally found in pairs,
Bioinformatics Biology Review The genetic code is stored in DNA Deoxyribonucleic acid. DNA molecules are chains of four nucleotide bases Guanine, Thymine, Cytosine, Adenine DNA is normally found in pairs,
Profile HMMs. 2/10/05 CAP5510/CGS5166 (Lec 10) 1 START STATE 1 STATE 2 STATE 3 STATE 4 STATE 5 STATE 6 END
 Profile HMMs START STATE 1 STATE 2 STATE 3 STATE 4 STATE 5 STATE 6 END 2/10/05 CAP5510/CGS5166 (Lec 10) 1 Profile HMMs with InDels Insertions Deletions Insertions & Deletions DELETE 1 DELETE 2 DELETE 3
Profile HMMs START STATE 1 STATE 2 STATE 3 STATE 4 STATE 5 STATE 6 END 2/10/05 CAP5510/CGS5166 (Lec 10) 1 Profile HMMs with InDels Insertions Deletions Insertions & Deletions DELETE 1 DELETE 2 DELETE 3
Computational gene finding. Devika Subramanian Comp 470
 Computational gene finding Devika Subramanian Comp 470 Outline (3 lectures) The biological context Lec 1 Lec 2 Lec 3 Markov models and Hidden Markov models Ab-initio methods for gene finding Comparative
Computational gene finding Devika Subramanian Comp 470 Outline (3 lectures) The biological context Lec 1 Lec 2 Lec 3 Markov models and Hidden Markov models Ab-initio methods for gene finding Comparative
Reading Lecture 8: Lecture 9: Lecture 8. DNA Libraries. Definition Types Construction
 Lecture 8 Reading Lecture 8: 96-110 Lecture 9: 111-120 DNA Libraries Definition Types Construction 142 DNA Libraries A DNA library is a collection of clones of genomic fragments or cdnas from a certain
Lecture 8 Reading Lecture 8: 96-110 Lecture 9: 111-120 DNA Libraries Definition Types Construction 142 DNA Libraries A DNA library is a collection of clones of genomic fragments or cdnas from a certain
The Ensembl Database. Dott.ssa Inga Prokopenko. Corso di Genomica
 The Ensembl Database Dott.ssa Inga Prokopenko Corso di Genomica 1 www.ensembl.org Lecture 7.1 2 What is Ensembl? Public annotation of mammalian and other genomes Open source software Relational database
The Ensembl Database Dott.ssa Inga Prokopenko Corso di Genomica 1 www.ensembl.org Lecture 7.1 2 What is Ensembl? Public annotation of mammalian and other genomes Open source software Relational database
Eukaryotic Gene Prediction. Wei Zhu May 2007
 Eukaryotic Gene Prediction Wei Zhu May 2007 In nature, nothing is perfect... - Alice Walker Gene Structure What is Gene Prediction? Gene prediction is the problem of parsing a sequence into nonoverlapping
Eukaryotic Gene Prediction Wei Zhu May 2007 In nature, nothing is perfect... - Alice Walker Gene Structure What is Gene Prediction? Gene prediction is the problem of parsing a sequence into nonoverlapping
Outline. Gene Finding Questions. Recap: Prokaryotic gene finding Eukaryotic gene finding The human gene complement Regulation
 Tues, Nov 29: Gene Finding 1 Online FCE s: Thru Dec 12 Thurs, Dec 1: Gene Finding 2 Tues, Dec 6: PS5 due Project presentations 1 (see course web site for schedule) Thurs, Dec 8 Final papers due Project
Tues, Nov 29: Gene Finding 1 Online FCE s: Thru Dec 12 Thurs, Dec 1: Gene Finding 2 Tues, Dec 6: PS5 due Project presentations 1 (see course web site for schedule) Thurs, Dec 8 Final papers due Project
Computational Gene Finding
 Computational Gene Finding Dong Xu Digital Biology Laboratory Computer Science Department Christopher S. Life Sciences Center University of Missouri, Columbia E-mail: xudong@missouri.edu http://digbio.missouri.edu
Computational Gene Finding Dong Xu Digital Biology Laboratory Computer Science Department Christopher S. Life Sciences Center University of Missouri, Columbia E-mail: xudong@missouri.edu http://digbio.missouri.edu
I. Gene Expression Figure 1: Central Dogma of Molecular Biology
 I. Gene Expression Figure 1: Central Dogma of Molecular Biology Central Dogma: Gene Expression: RNA Structure RNA nucleotides contain the pentose sugar Ribose instead of deoxyribose. Contain the bases
I. Gene Expression Figure 1: Central Dogma of Molecular Biology Central Dogma: Gene Expression: RNA Structure RNA nucleotides contain the pentose sugar Ribose instead of deoxyribose. Contain the bases
Genome annotation. Erwin Datema (2011) Sandra Smit (2012, 2013)
 Genome annotation Erwin Datema (2011) Sandra Smit (2012, 2013) Genome annotation AGACAAAGATCCGCTAAATTAAATCTGGACTTCACATATTGAAGTGATATCACACGTTTCTCTAAT AATCTCCTCACAATATTATGTTTGGGATGAACTTGTCGTGATTTGCCATTGTAGCAATCACTTGAA
Genome annotation Erwin Datema (2011) Sandra Smit (2012, 2013) Genome annotation AGACAAAGATCCGCTAAATTAAATCTGGACTTCACATATTGAAGTGATATCACACGTTTCTCTAAT AATCTCCTCACAATATTATGTTTGGGATGAACTTGTCGTGATTTGCCATTGTAGCAATCACTTGAA
Genomics and Gene Recognition Genes and Blue Genes
 Genomics and Gene Recognition Genes and Blue Genes November 1, 2004 Prokaryotic Gene Structure prokaryotes are simplest free-living organisms studying prokaryotes can give us a sense what is the minimum
Genomics and Gene Recognition Genes and Blue Genes November 1, 2004 Prokaryotic Gene Structure prokaryotes are simplest free-living organisms studying prokaryotes can give us a sense what is the minimum
From RNA To Protein
 From RNA To Protein 22-11-2016 Introduction mrna Processing heterogeneous nuclear RNA (hnrna) RNA that comprises transcripts of nuclear genes made by RNA polymerase II; it has a wide size distribution
From RNA To Protein 22-11-2016 Introduction mrna Processing heterogeneous nuclear RNA (hnrna) RNA that comprises transcripts of nuclear genes made by RNA polymerase II; it has a wide size distribution
Outline. Introduction to ab initio and evidence-based gene finding. Prokaryotic gene predictions
 Outline Introduction to ab initio and evidence-based gene finding Overview of computational gene predictions Different types of eukaryotic gene predictors Common types of gene prediction errors Wilson
Outline Introduction to ab initio and evidence-based gene finding Overview of computational gene predictions Different types of eukaryotic gene predictors Common types of gene prediction errors Wilson
RNA folding & ncrna discovery
 I519 Introduction to Bioinformatics RNA folding & ncrna discovery Yuzhen Ye (yye@indiana.edu) School of Informatics & Computing, IUB Contents Non-coding RNAs and their functions RNA structures RNA folding
I519 Introduction to Bioinformatics RNA folding & ncrna discovery Yuzhen Ye (yye@indiana.edu) School of Informatics & Computing, IUB Contents Non-coding RNAs and their functions RNA structures RNA folding
Transcription in Eukaryotes
 Transcription in Eukaryotes Biology I Hayder A Giha Transcription Transcription is a DNA-directed synthesis of RNA, which is the first step in gene expression. Gene expression, is transformation of the
Transcription in Eukaryotes Biology I Hayder A Giha Transcription Transcription is a DNA-directed synthesis of RNA, which is the first step in gene expression. Gene expression, is transformation of the
RNA : functional role
 RNA : functional role Hamad Yaseen, PhD MLS Department, FAHS Hamad.ali@hsc.edu.kw RNA mrna rrna trna 1 From DNA to Protein -Outline- From DNA to RNA From RNA to Protein From DNA to RNA Transcription: Copying
RNA : functional role Hamad Yaseen, PhD MLS Department, FAHS Hamad.ali@hsc.edu.kw RNA mrna rrna trna 1 From DNA to Protein -Outline- From DNA to RNA From RNA to Protein From DNA to RNA Transcription: Copying
SCBC203 Gene Expression. Assoc. Prof. Rutaiwan Tohtong Department of Biochemistry Faculty of Science PR318
 SCBC203 Gene Expression Assoc. Prof. Rutaiwan Tohtong Department of Biochemistry Faculty of Science PR318 Rutaiwan.toh@mahidol.ac.th 1 Gene Expression Gene expression is a process where by the genetic
SCBC203 Gene Expression Assoc. Prof. Rutaiwan Tohtong Department of Biochemistry Faculty of Science PR318 Rutaiwan.toh@mahidol.ac.th 1 Gene Expression Gene expression is a process where by the genetic
Genes and How They Work. Chapter 15
 Genes and How They Work Chapter 15 The Nature of Genes They proposed the one gene one enzyme hypothesis. Today we know this as the one gene one polypeptide hypothesis. 2 The Nature of Genes The central
Genes and How They Work Chapter 15 The Nature of Genes They proposed the one gene one enzyme hypothesis. Today we know this as the one gene one polypeptide hypothesis. 2 The Nature of Genes The central
30 Gene expression: Transcription
 30 Gene expression: Transcription Gene structure. o Exons coding region of DNA. o Introns non-coding region of DNA. o Introns are interspersed between exons of a single gene. o Promoter region helps enzymes
30 Gene expression: Transcription Gene structure. o Exons coding region of DNA. o Introns non-coding region of DNA. o Introns are interspersed between exons of a single gene. o Promoter region helps enzymes
RNA secondary structure prediction and analysis
 RNA secondary structure prediction and analysis 1 Resources Lecture Notes from previous years: Takis Benos Covariance algorithm: Eddy and Durbin, Nucleic Acids Research, v22: 11, 2079 Useful lecture slides
RNA secondary structure prediction and analysis 1 Resources Lecture Notes from previous years: Takis Benos Covariance algorithm: Eddy and Durbin, Nucleic Acids Research, v22: 11, 2079 Useful lecture slides
DNA makes RNA makes Proteins. The Central Dogma
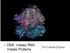 DNA makes RNA makes Proteins The Central Dogma TRANSCRIPTION DNA RNA transcript RNA polymerase RNA PROCESSING Exon RNA transcript (pre-mrna) Intron Aminoacyl-tRNA synthetase NUCLEUS CYTOPLASM FORMATION
DNA makes RNA makes Proteins The Central Dogma TRANSCRIPTION DNA RNA transcript RNA polymerase RNA PROCESSING Exon RNA transcript (pre-mrna) Intron Aminoacyl-tRNA synthetase NUCLEUS CYTOPLASM FORMATION
Transcription is the first stage of gene expression
 Transcription is the first stage of gene expression RNA synthesis is catalyzed by RNA polymerase, which pries the DNA strands apart and hooks together the RNA nucleotides The RNA is complementary to the
Transcription is the first stage of gene expression RNA synthesis is catalyzed by RNA polymerase, which pries the DNA strands apart and hooks together the RNA nucleotides The RNA is complementary to the
BIO 311C Spring Lecture 36 Wednesday 28 Apr.
 BIO 311C Spring 2010 1 Lecture 36 Wednesday 28 Apr. Synthesis of a Polypeptide Chain 5 direction of ribosome movement along the mrna 3 ribosome mrna NH 2 polypeptide chain direction of mrna movement through
BIO 311C Spring 2010 1 Lecture 36 Wednesday 28 Apr. Synthesis of a Polypeptide Chain 5 direction of ribosome movement along the mrna 3 ribosome mrna NH 2 polypeptide chain direction of mrna movement through
From DNA to Protein: Genotype to Phenotype
 12 From DNA to Protein: Genotype to Phenotype 12.1 What Is the Evidence that Genes Code for Proteins? The gene-enzyme relationship is one-gene, one-polypeptide relationship. Example: In hemoglobin, each
12 From DNA to Protein: Genotype to Phenotype 12.1 What Is the Evidence that Genes Code for Proteins? The gene-enzyme relationship is one-gene, one-polypeptide relationship. Example: In hemoglobin, each
Computational gene finding
 Computational gene finding Devika Subramanian Comp 470 Outline (3 lectures) Lec 1 Lec 2 Lec 3 The biological context Markov models and Hidden Markov models Ab-initio methods for gene finding Comparative
Computational gene finding Devika Subramanian Comp 470 Outline (3 lectures) Lec 1 Lec 2 Lec 3 The biological context Markov models and Hidden Markov models Ab-initio methods for gene finding Comparative
The Nature of Genes. The Nature of Genes. Genes and How They Work. Chapter 15/16
 Genes and How They Work Chapter 15/16 The Nature of Genes Beadle and Tatum proposed the one gene one enzyme hypothesis. Today we know this as the one gene one polypeptide hypothesis. 2 The Nature of Genes
Genes and How They Work Chapter 15/16 The Nature of Genes Beadle and Tatum proposed the one gene one enzyme hypothesis. Today we know this as the one gene one polypeptide hypothesis. 2 The Nature of Genes
Lecture Summary: Regulation of transcription. General mechanisms-what are the major regulatory points?
 BCH 401G Lecture 37 Andres Lecture Summary: Regulation of transcription. General mechanisms-what are the major regulatory points? RNA processing: Capping, polyadenylation, splicing. Why process mammalian
BCH 401G Lecture 37 Andres Lecture Summary: Regulation of transcription. General mechanisms-what are the major regulatory points? RNA processing: Capping, polyadenylation, splicing. Why process mammalian
Lecture for Wednesday. Dr. Prince BIOL 1408
 Lecture for Wednesday Dr. Prince BIOL 1408 THE FLOW OF GENETIC INFORMATION FROM DNA TO RNA TO PROTEIN Copyright 2009 Pearson Education, Inc. Genes are expressed as proteins A gene is a segment of DNA that
Lecture for Wednesday Dr. Prince BIOL 1408 THE FLOW OF GENETIC INFORMATION FROM DNA TO RNA TO PROTEIN Copyright 2009 Pearson Education, Inc. Genes are expressed as proteins A gene is a segment of DNA that
There are four major types of introns. Group I introns, found in some rrna genes, are self-splicing: they can catalyze their own removal.
 1 2 Continuous genes - Intron: Many eukaryotic genes contain coding regions called exons and noncoding regions called intervening sequences or introns. The average human gene contains from eight to nine
1 2 Continuous genes - Intron: Many eukaryotic genes contain coding regions called exons and noncoding regions called intervening sequences or introns. The average human gene contains from eight to nine
Unit 1: DNA and the Genome. Sub-Topic (1.3) Gene Expression
 Unit 1: DNA and the Genome Sub-Topic (1.3) Gene Expression Unit 1: DNA and the Genome Sub-Topic (1.3) Gene Expression On completion of this subtopic I will be able to State the meanings of the terms genotype,
Unit 1: DNA and the Genome Sub-Topic (1.3) Gene Expression Unit 1: DNA and the Genome Sub-Topic (1.3) Gene Expression On completion of this subtopic I will be able to State the meanings of the terms genotype,
Improved Splice Site Detection in Genie
 Improved Splice Site Detection in Genie Martin Reese Informatics Group Human Genome Center Lawrence Berkeley National Laboratory MGReese@lbl.gov http://www-hgc.lbl.gov/inf Santa Fe, 1/23/97 Database Homologies
Improved Splice Site Detection in Genie Martin Reese Informatics Group Human Genome Center Lawrence Berkeley National Laboratory MGReese@lbl.gov http://www-hgc.lbl.gov/inf Santa Fe, 1/23/97 Database Homologies
The Central Dogma. DNA makes RNA makes Proteins
 The Central Dogma DNA makes RNA makes Proteins TRANSCRIPTION DNA RNA transcript RNA polymerase RNA PROCESSING Exon RNA transcript (pre-) Intron Aminoacyl-tRNA synthetase NUCLEUS CYTOPLASM FORMATION OF
The Central Dogma DNA makes RNA makes Proteins TRANSCRIPTION DNA RNA transcript RNA polymerase RNA PROCESSING Exon RNA transcript (pre-) Intron Aminoacyl-tRNA synthetase NUCLEUS CYTOPLASM FORMATION OF
BIOL 300 Foundations of Biology Summer 2017 Telleen Lecture Outline
 BIOL 300 Foundations of Biology Summer 2017 Telleen Lecture Outline RNA, the Genetic Code, Proteins I. How RNA differs from DNA A. The sugar ribose replaces deoxyribose. The presence of the oxygen on the
BIOL 300 Foundations of Biology Summer 2017 Telleen Lecture Outline RNA, the Genetic Code, Proteins I. How RNA differs from DNA A. The sugar ribose replaces deoxyribose. The presence of the oxygen on the
Textbook Reading Guidelines
 Understanding Bioinformatics by Marketa Zvelebil and Jeremy Baum Last updated: January 16, 2013 Textbook Reading Guidelines Preface: Read the whole preface, and especially: For the students with Life Science
Understanding Bioinformatics by Marketa Zvelebil and Jeremy Baum Last updated: January 16, 2013 Textbook Reading Guidelines Preface: Read the whole preface, and especially: For the students with Life Science
Molecular Cell Biology - Problem Drill 08: Transcription, Translation and the Genetic Code
 Molecular Cell Biology - Problem Drill 08: Transcription, Translation and the Genetic Code Question No. 1 of 10 1. Which of the following statements about how genes function is correct? Question #1 (A)
Molecular Cell Biology - Problem Drill 08: Transcription, Translation and the Genetic Code Question No. 1 of 10 1. Which of the following statements about how genes function is correct? Question #1 (A)
Bi 8 Lecture 5. Ellen Rothenberg 19 January 2016
 Bi 8 Lecture 5 MORE ON HOW WE KNOW WHAT WE KNOW and intro to the protein code Ellen Rothenberg 19 January 2016 SIZE AND PURIFICATION BY SYNTHESIS: BASIS OF EARLY SEQUENCING complex mixture of aborted DNA
Bi 8 Lecture 5 MORE ON HOW WE KNOW WHAT WE KNOW and intro to the protein code Ellen Rothenberg 19 January 2016 SIZE AND PURIFICATION BY SYNTHESIS: BASIS OF EARLY SEQUENCING complex mixture of aborted DNA
From DNA to Protein: Genotype to Phenotype
 12 From DNA to Protein: Genotype to Phenotype 12.1 What Is the Evidence that Genes Code for Proteins? The gene-enzyme relationship is one-gene, one-polypeptide relationship. Example: In hemoglobin, each
12 From DNA to Protein: Genotype to Phenotype 12.1 What Is the Evidence that Genes Code for Proteins? The gene-enzyme relationship is one-gene, one-polypeptide relationship. Example: In hemoglobin, each
Applications of hidden Markov models to sequence analysis. Lior Pachter
 Applications of hidden Markov models to sequence analysis Lior Pachter Outline Why do we analyze sequences? What are we looking for? Annotation of DNA sequences I (and HMMs) Alignment Annotation of DNA
Applications of hidden Markov models to sequence analysis Lior Pachter Outline Why do we analyze sequences? What are we looking for? Annotation of DNA sequences I (and HMMs) Alignment Annotation of DNA
Gene Structure & Gene Finding Part II
 Gene Structure & Gene Finding Part II David Wishart david.wishart@ualberta.ca 30,000 metabolite Gene Finding in Eukaryotes Eukaryotes Complex gene structure Large genomes (0.1 to 10 billion bp) Exons and
Gene Structure & Gene Finding Part II David Wishart david.wishart@ualberta.ca 30,000 metabolite Gene Finding in Eukaryotes Eukaryotes Complex gene structure Large genomes (0.1 to 10 billion bp) Exons and
Translation BIT 220 Chapter 13
 Translation BIT 220 Chapter 13 Making protein from mrna Most genes encode for proteins -some make RNA as end product Proteins -Monomer Amino Acid 20 amino acids -peptides -polypeptides -Structure of Amino
Translation BIT 220 Chapter 13 Making protein from mrna Most genes encode for proteins -some make RNA as end product Proteins -Monomer Amino Acid 20 amino acids -peptides -polypeptides -Structure of Amino
Regulation of bacterial gene expression
 Regulation of bacterial gene expression Gene Expression Gene Expression: RNA and protein synthesis DNA ----------> RNA ----------> Protein transcription translation! DNA replication only occurs in cells
Regulation of bacterial gene expression Gene Expression Gene Expression: RNA and protein synthesis DNA ----------> RNA ----------> Protein transcription translation! DNA replication only occurs in cells
Analysis of Biological Sequences SPH
 Analysis of Biological Sequences SPH 140.638 swheelan@jhmi.edu nuts and bolts meet Tuesdays & Thursdays, 3:30-4:50 no exam; grade derived from 3-4 homework assignments plus a final project (open book,
Analysis of Biological Sequences SPH 140.638 swheelan@jhmi.edu nuts and bolts meet Tuesdays & Thursdays, 3:30-4:50 no exam; grade derived from 3-4 homework assignments plus a final project (open book,
Gene Expression: Transcription, Translation, RNAs and the Genetic Code
 Lecture 28-29 Gene Expression: Transcription, Translation, RNAs and the Genetic Code Central dogma of molecular biology During transcription, the information in a DNA sequence (a gene) is copied into a
Lecture 28-29 Gene Expression: Transcription, Translation, RNAs and the Genetic Code Central dogma of molecular biology During transcription, the information in a DNA sequence (a gene) is copied into a
Biology A: Chapter 9 Annotating Notes Protein Synthesis
 Name: Pd: Biology A: Chapter 9 Annotating Notes Protein Synthesis -As you read your textbook, please fill out these notes. -Read each paragraph state the big/main idea on the left side. -On the right side
Name: Pd: Biology A: Chapter 9 Annotating Notes Protein Synthesis -As you read your textbook, please fill out these notes. -Read each paragraph state the big/main idea on the left side. -On the right side
BIOCHEMISTRY REVIEW. Overview of Biomolecules. Chapter 12 Transcription
 BIOCHEMISTRY REVIEW Overview of Biomolecules Chapter 12 Transcription 2 3 4 5 Are You Getting It?? Which are general characteristics of transcription? (multiple answers) a) An entire DNA molecule is transcribed
BIOCHEMISTRY REVIEW Overview of Biomolecules Chapter 12 Transcription 2 3 4 5 Are You Getting It?? Which are general characteristics of transcription? (multiple answers) a) An entire DNA molecule is transcribed
Videos. Lesson Overview. Fermentation
 Lesson Overview Fermentation Videos Bozeman Transcription and Translation: https://youtu.be/h3b9arupxzg Drawing transcription and translation: https://youtu.be/6yqplgnjr4q Objectives 29a) I can contrast
Lesson Overview Fermentation Videos Bozeman Transcription and Translation: https://youtu.be/h3b9arupxzg Drawing transcription and translation: https://youtu.be/6yqplgnjr4q Objectives 29a) I can contrast
UCSC Genome Browser. Introduction to ab initio and evidence-based gene finding
 UCSC Genome Browser Introduction to ab initio and evidence-based gene finding Wilson Leung 06/2006 Outline Introduction to annotation ab initio gene finding Basics of the UCSC Browser Evidence-based gene
UCSC Genome Browser Introduction to ab initio and evidence-based gene finding Wilson Leung 06/2006 Outline Introduction to annotation ab initio gene finding Basics of the UCSC Browser Evidence-based gene
Themes: RNA and RNA Processing. Messenger RNA (mrna) What is a gene? RNA is very versatile! RNA-RNA interactions are very important!
 Themes: RNA is very versatile! RNA and RNA Processing Chapter 14 RNA-RNA interactions are very important! Prokaryotes and Eukaryotes have many important differences. Messenger RNA (mrna) Carries genetic
Themes: RNA is very versatile! RNA and RNA Processing Chapter 14 RNA-RNA interactions are very important! Prokaryotes and Eukaryotes have many important differences. Messenger RNA (mrna) Carries genetic
RNA Genomics II. BME 110: CompBio Tools Todd Lowe & Andrew Uzilov May 17, 2011
 RNA Genomics II BME 110: CompBio Tools Todd Lowe & Andrew Uzilov May 17, 2011 1 TIME Why RNA? An evolutionary perspective The RNA World hypotheses: life arose as self-replicating non-coding RNA (ncrna)
RNA Genomics II BME 110: CompBio Tools Todd Lowe & Andrew Uzilov May 17, 2011 1 TIME Why RNA? An evolutionary perspective The RNA World hypotheses: life arose as self-replicating non-coding RNA (ncrna)
DNA Replication and Repair
 DNA Replication and Repair http://hyperphysics.phy-astr.gsu.edu/hbase/organic/imgorg/cendog.gif Overview of DNA Replication SWYK CNs 1, 2, 30 Explain how specific base pairing enables existing DNA strands
DNA Replication and Repair http://hyperphysics.phy-astr.gsu.edu/hbase/organic/imgorg/cendog.gif Overview of DNA Replication SWYK CNs 1, 2, 30 Explain how specific base pairing enables existing DNA strands
MODULE 5: TRANSLATION
 MODULE 5: TRANSLATION Lesson Plan: CARINA ENDRES HOWELL, LEOCADIA PALIULIS Title Translation Objectives Determine the codons for specific amino acids and identify reading frames by looking at the Base
MODULE 5: TRANSLATION Lesson Plan: CARINA ENDRES HOWELL, LEOCADIA PALIULIS Title Translation Objectives Determine the codons for specific amino acids and identify reading frames by looking at the Base
Videos. Bozeman Transcription and Translation: Drawing transcription and translation:
 Videos Bozeman Transcription and Translation: https://youtu.be/h3b9arupxzg Drawing transcription and translation: https://youtu.be/6yqplgnjr4q Objectives 29a) I can contrast RNA and DNA. 29b) I can explain
Videos Bozeman Transcription and Translation: https://youtu.be/h3b9arupxzg Drawing transcription and translation: https://youtu.be/6yqplgnjr4q Objectives 29a) I can contrast RNA and DNA. 29b) I can explain
Introduction to RNA-Seq. David Wood Winter School in Mathematics and Computational Biology July 1, 2013
 Introduction to RNA-Seq David Wood Winter School in Mathematics and Computational Biology July 1, 2013 Abundance RNA is... Diverse Dynamic Central DNA rrna Epigenetics trna RNA mrna Time Protein Abundance
Introduction to RNA-Seq David Wood Winter School in Mathematics and Computational Biology July 1, 2013 Abundance RNA is... Diverse Dynamic Central DNA rrna Epigenetics trna RNA mrna Time Protein Abundance
Homework 4. Due in class, Wednesday, November 10, 2004
 1 GCB 535 / CIS 535 Fall 2004 Homework 4 Due in class, Wednesday, November 10, 2004 Comparative genomics 1. (6 pts) In Loots s paper (http://www.seas.upenn.edu/~cis535/lab/sciences-loots.pdf), the authors
1 GCB 535 / CIS 535 Fall 2004 Homework 4 Due in class, Wednesday, November 10, 2004 Comparative genomics 1. (6 pts) In Loots s paper (http://www.seas.upenn.edu/~cis535/lab/sciences-loots.pdf), the authors
Human Gene,cs 06: Gene Expression. Diversity of cell types. How do cells become different? 9/19/11. neuron
 Human Gene,cs 06: Gene Expression 20110920 Diversity of cell types neuron How do cells become different? A. Each type of cell has different DNA in its nucleus B. Each cell has different genes C. Each type
Human Gene,cs 06: Gene Expression 20110920 Diversity of cell types neuron How do cells become different? A. Each type of cell has different DNA in its nucleus B. Each cell has different genes C. Each type
The Flow of Genetic Information
 Chapter 17 The Flow of Genetic Information The DNA inherited by an organism leads to specific traits by dictating the synthesis of proteins and of RNA molecules involved in protein synthesis. Proteins
Chapter 17 The Flow of Genetic Information The DNA inherited by an organism leads to specific traits by dictating the synthesis of proteins and of RNA molecules involved in protein synthesis. Proteins
Computational analysis of non-coding RNA. Andrew Uzilov BME110 Tue, Nov 16, 2010
 Computational analysis of non-coding RNA Andrew Uzilov auzilov@ucsc.edu BME110 Tue, Nov 16, 2010 1 Corrected/updated talk slides are here: http://tinyurl.com/uzilovrna redirects to: http://users.soe.ucsc.edu/~auzilov/bme110/fall2010/
Computational analysis of non-coding RNA Andrew Uzilov auzilov@ucsc.edu BME110 Tue, Nov 16, 2010 1 Corrected/updated talk slides are here: http://tinyurl.com/uzilovrna redirects to: http://users.soe.ucsc.edu/~auzilov/bme110/fall2010/
Transcription steps. Transcription steps. Eukaryote RNA processing
 Transcription steps Initiation at 5 end of gene binding of RNA polymerase to promoter unwinding of DNA Elongation addition of nucleotides to 3 end rules of base pairing requires Mg 2+ energy from NTP substrates
Transcription steps Initiation at 5 end of gene binding of RNA polymerase to promoter unwinding of DNA Elongation addition of nucleotides to 3 end rules of base pairing requires Mg 2+ energy from NTP substrates
Fermentation. Lesson Overview. Lesson Overview 13.1 RNA
 13.1 RNA THINK ABOUT IT DNA is the genetic material of cells. The sequence of nucleotide bases in the strands of DNA carries some sort of code. In order for that code to work, the cell must be able to
13.1 RNA THINK ABOUT IT DNA is the genetic material of cells. The sequence of nucleotide bases in the strands of DNA carries some sort of code. In order for that code to work, the cell must be able to
Bioinformatics: Sequence Analysis. COMP 571 Luay Nakhleh, Rice University
 Bioinformatics: Sequence Analysis COMP 571 Luay Nakhleh, Rice University Course Information Instructor: Luay Nakhleh (nakhleh@rice.edu); office hours by appointment (office: DH 3119) TA: Leo Elworth (DH
Bioinformatics: Sequence Analysis COMP 571 Luay Nakhleh, Rice University Course Information Instructor: Luay Nakhleh (nakhleh@rice.edu); office hours by appointment (office: DH 3119) TA: Leo Elworth (DH
Chapter 13. From DNA to Protein
 Chapter 13 From DNA to Protein Proteins All proteins consist of polypeptide chains A linear sequence of amino acids Each chain corresponds to the nucleotide base sequenceof a gene The Path From Genes to
Chapter 13 From DNA to Protein Proteins All proteins consist of polypeptide chains A linear sequence of amino acids Each chain corresponds to the nucleotide base sequenceof a gene The Path From Genes to
DNA Function: Information Transmission
 DNA Function: Information Transmission DNA is called the code of life. What does it code for? *the information ( code ) to make proteins! Why are proteins so important? Nearly every function of a living
DNA Function: Information Transmission DNA is called the code of life. What does it code for? *the information ( code ) to make proteins! Why are proteins so important? Nearly every function of a living
Genomics and Gene Recognition Genes and Blue Genes
 Genomics and Gene Recognition Genes and Blue Genes November 3, 2004 Eukaryotic Gene Structure eukaryotic genomes are considerably more complex than those of prokaryotes eukaryotic cells have organelles
Genomics and Gene Recognition Genes and Blue Genes November 3, 2004 Eukaryotic Gene Structure eukaryotic genomes are considerably more complex than those of prokaryotes eukaryotic cells have organelles
May 16. Gene Finding
 Gene Finding j T[j,k] k i Q is a set of states T is a matrix of transition probabilities T[j,k]: probability of moving from state j to state k Σ is a set of symbols e j (S) is the probability of emitting
Gene Finding j T[j,k] k i Q is a set of states T is a matrix of transition probabilities T[j,k]: probability of moving from state j to state k Σ is a set of symbols e j (S) is the probability of emitting
High-throughput Transcriptome analysis
 High-throughput Transcriptome analysis CAGE and beyond Dr. Rimantas Kodzius, Singapore, A*STAR, IMCB rkodzius@imcb.a-star.edu.sg for KAUST 2008 Agenda 1. Current research - PhD work on discovery of new
High-throughput Transcriptome analysis CAGE and beyond Dr. Rimantas Kodzius, Singapore, A*STAR, IMCB rkodzius@imcb.a-star.edu.sg for KAUST 2008 Agenda 1. Current research - PhD work on discovery of new
Key Area 1.3: Gene Expression
 Key Area 1.3: Gene Expression RNA There is a second type of nucleic acid in the cell, called RNA. RNA plays a vital role in the production of protein from the code in the DNA. What is gene expression?
Key Area 1.3: Gene Expression RNA There is a second type of nucleic acid in the cell, called RNA. RNA plays a vital role in the production of protein from the code in the DNA. What is gene expression?
Eukaryotic Gene Structure
 Eukaryotic Gene Structure Terminology Genome entire genetic material of an individual Transcriptome set of transcribed sequences Proteome set of proteins encoded by the genome 2 Gene Basic physical and
Eukaryotic Gene Structure Terminology Genome entire genetic material of an individual Transcriptome set of transcribed sequences Proteome set of proteins encoded by the genome 2 Gene Basic physical and
MATH 5610, Computational Biology
 MATH 5610, Computational Biology Lecture 2 Intro to Molecular Biology (cont) Stephen Billups University of Colorado at Denver MATH 5610, Computational Biology p.1/24 Announcements Error on syllabus Class
MATH 5610, Computational Biology Lecture 2 Intro to Molecular Biology (cont) Stephen Billups University of Colorado at Denver MATH 5610, Computational Biology p.1/24 Announcements Error on syllabus Class
CLEP Biology - Problem Drill 11: Transcription, Translation and The Genetic Code
 CLEP Biology - Problem Drill 11: Transcription, Translation and The Genetic Code No. 1 of 10 1. Three types of RNA comprise the structural and functional core for protein synthesis, serving as a template
CLEP Biology - Problem Drill 11: Transcription, Translation and The Genetic Code No. 1 of 10 1. Three types of RNA comprise the structural and functional core for protein synthesis, serving as a template
Prediction of noncoding RNAs with RNAz
 Prediction of noncoding RNAs with RNAz John Dzmil, III Steve Griesmer Philip Murillo April 4, 2007 What is non-coding RNA (ncrna)? RNA molecules that are not translated into proteins Size range from 20
Prediction of noncoding RNAs with RNAz John Dzmil, III Steve Griesmer Philip Murillo April 4, 2007 What is non-coding RNA (ncrna)? RNA molecules that are not translated into proteins Size range from 20
Chapter 3. DNA, RNA, and Protein Synthesis
 Chapter 3. DNA, RNA, and Protein Synthesis 4. Transcription Gene Expression Regulatory region (promoter) 5 flanking region Upstream region Coding region 3 flanking region Downstream region Transcription
Chapter 3. DNA, RNA, and Protein Synthesis 4. Transcription Gene Expression Regulatory region (promoter) 5 flanking region Upstream region Coding region 3 flanking region Downstream region Transcription
Gene function at the level of traits Gene function at the molecular level
 Gene expression Gene function at the level of traits Gene function at the molecular level Two levels tied together since the molecular level affects the structure and function of cells which determines
Gene expression Gene function at the level of traits Gene function at the molecular level Two levels tied together since the molecular level affects the structure and function of cells which determines
From Gene to Protein. How Genes Work
 From Gene to Protein How Genes Work 2007-2008 The Central Dogma Flow of genetic information in a cell How do we move information from DNA to proteins? DNA RNA protein replication phenotype You! Step 1:
From Gene to Protein How Genes Work 2007-2008 The Central Dogma Flow of genetic information in a cell How do we move information from DNA to proteins? DNA RNA protein replication phenotype You! Step 1:
PROTEIN SYNTHESIS Flow of Genetic Information The flow of genetic information can be symbolized as: DNA RNA Protein
 PROTEIN SYNTHESIS Flow of Genetic Information The flow of genetic information can be symbolized as: DNA RNA Protein This is also known as: The central dogma of molecular biology Protein Proteins are made
PROTEIN SYNTHESIS Flow of Genetic Information The flow of genetic information can be symbolized as: DNA RNA Protein This is also known as: The central dogma of molecular biology Protein Proteins are made
Lecture 7 Motif Databases and Gene Finding
 Introduction to Bioinformatics for Medical Research Gideon Greenspan gdg@cs.technion.ac.il Lecture 7 Motif Databases and Gene Finding Motif Databases & Gene Finding Motifs Recap Motif Databases TRANSFAC
Introduction to Bioinformatics for Medical Research Gideon Greenspan gdg@cs.technion.ac.il Lecture 7 Motif Databases and Gene Finding Motif Databases & Gene Finding Motifs Recap Motif Databases TRANSFAC
Gene & genome organisation. Computational gene identification
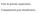 Gene & genome organisation Computational gene identification Eubacterial gene Eukaryotic gene Regulatory elements Promoter Translation start Introns polya signal Transcription stop DNA Transcription
Gene & genome organisation Computational gene identification Eubacterial gene Eukaryotic gene Regulatory elements Promoter Translation start Introns polya signal Transcription stop DNA Transcription
Sequence Analysis. II: Sequence Patterns and Matrices. George Bell, Ph.D. WIBR Bioinformatics and Research Computing
 Sequence Analysis II: Sequence Patterns and Matrices George Bell, Ph.D. WIBR Bioinformatics and Research Computing Sequence Patterns and Matrices Multiple sequence alignments Sequence patterns Sequence
Sequence Analysis II: Sequence Patterns and Matrices George Bell, Ph.D. WIBR Bioinformatics and Research Computing Sequence Patterns and Matrices Multiple sequence alignments Sequence patterns Sequence
Protein Synthesis Notes
 Protein Synthesis Notes Protein Synthesis: Overview Transcription: synthesis of mrna under the direction of DNA. Translation: actual synthesis of a polypeptide under the direction of mrna. Transcription
Protein Synthesis Notes Protein Synthesis: Overview Transcription: synthesis of mrna under the direction of DNA. Translation: actual synthesis of a polypeptide under the direction of mrna. Transcription
CH 17 :From Gene to Protein
 CH 17 :From Gene to Protein Defining a gene gene gene Defining a gene is problematic because one gene can code for several protein products, some genes code only for RNA, two genes can overlap, and there
CH 17 :From Gene to Protein Defining a gene gene gene Defining a gene is problematic because one gene can code for several protein products, some genes code only for RNA, two genes can overlap, and there
Multiple choice questions (numbers in brackets indicate the number of correct answers)
 1 February 15, 2013 Multiple choice questions (numbers in brackets indicate the number of correct answers) 1. Which of the following statements are not true Transcriptomes consist of mrnas Proteomes consist
1 February 15, 2013 Multiple choice questions (numbers in brackets indicate the number of correct answers) 1. Which of the following statements are not true Transcriptomes consist of mrnas Proteomes consist
Gene Expression Transcription/Translation Protein Synthesis
 Gene Expression Transcription/Translation Protein Synthesis 1. Describe how genetic information is transcribed into sequences of bases in RNA molecules and is finally translated into sequences of amino
Gene Expression Transcription/Translation Protein Synthesis 1. Describe how genetic information is transcribed into sequences of bases in RNA molecules and is finally translated into sequences of amino
MULTIPLE CHOICE. Choose the one alternative that best completes the statement or answers the question.
 Ch 17 Practice Questions MULTIPLE CHOICE. Choose the one alternative that best completes the statement or answers the question. 1) Garrod hypothesized that "inborn errors of metabolism" such as alkaptonuria
Ch 17 Practice Questions MULTIPLE CHOICE. Choose the one alternative that best completes the statement or answers the question. 1) Garrod hypothesized that "inborn errors of metabolism" such as alkaptonuria
Bio 101 Sample questions: Chapter 10
 Bio 101 Sample questions: Chapter 10 1. Which of the following is NOT needed for DNA replication? A. nucleotides B. ribosomes C. Enzymes (like polymerases) D. DNA E. all of the above are needed 2 The information
Bio 101 Sample questions: Chapter 10 1. Which of the following is NOT needed for DNA replication? A. nucleotides B. ribosomes C. Enzymes (like polymerases) D. DNA E. all of the above are needed 2 The information
Hello! Outline. Cell Biology: RNA and Protein synthesis. In all living cells, DNA molecules are the storehouses of information. 6.
 Cell Biology: RNA and Protein synthesis In all living cells, DNA molecules are the storehouses of information Hello! Outline u 1. Key concepts u 2. Central Dogma u 3. RNA Types u 4. RNA (Ribonucleic Acid)
Cell Biology: RNA and Protein synthesis In all living cells, DNA molecules are the storehouses of information Hello! Outline u 1. Key concepts u 2. Central Dogma u 3. RNA Types u 4. RNA (Ribonucleic Acid)
