Data Mining in Bioinformatics Day 6: Classification in Bioinformatics
|
|
|
- Kathryn Strickland
- 6 years ago
- Views:
Transcription
1 Data Mining in Bioinformatics Day 6: Classification in Bioinformatics Karsten Borgwardt February 25 to March 10 Bioinformatics Group MPIs Tübingen Karsten Borgwardt: Data Mining in Bioinformatics, Page 1
2 Karsten M. Borgwardt Protein function prediction via graph kernels ISMB 2005 Joint work with Cheng Soon Ong and S.V.N. Vishwanathan, Stefan Schönauer, Hans-Peter Kriegel and Alex Smola Ludwig-Maximilians-Universität Munich, Germany and National ICT Australia, Canberra
3 Content Introduction The problem: protein function prediction The method: Support Vector Machines (SVM) Our approach to function prediction Protein graph model Protein graph kernel Experimental evaluation Technique to analyze our graph model Hyperkernels Discussion Karsten Borgwardt et al. - Protein function prediction via graph kernels 2
4 Current approaches to protein function prediction similar structures similar phylogenetic profiles similar motifs similar interaction partners similar function similar surface clefts similar sequences similar chemical properties Karsten Borgwardt et al. - Protein function prediction via graph kernels 3
5 Current approaches to protein function prediction similar structures similar phylogenetic profiles similar motifs similar interaction partners similar function similar sequences similar chemical properties similar surface clefts Karsten Borgwardt et al. - Protein function prediction via graph kernels 4
6 Support Vector Machines Are new data points (x) red or black? The blue decision boundary allows to predict class membership of new data points. Karsten Borgwardt et al. - Protein function prediction via graph kernels 5
7 Kernel trick input space feature space mapping Ф kernel function The kernel trick allows to introduce a separating hyperplane in feature space. Karsten Borgwardt et al. - Protein function prediction via graph kernels 6
8 Feature vectors for function prediction protein structure and/or protein sequence e.g. Cai et al. (2004), Dobson and Doig (2003) hydrophobicity polarity polarizability van der Waals volume fraction of amino acid types fraction of surface area disulphide bonds size of largest surface pocket 7
9 Our approach Sequence + Structure + Chemical properties Graph model SVMs + Graph models Protein function Karsten Borgwardt et al. - Protein function prediction via graph kernels 8
10 Protein graph model protein secondary structure sequence structure Karsten Borgwardt et al. - Protein function prediction via graph kernels 9
11 Protein graph model Node attributes hydrophobicity polarity polarizability van der Waals volume length helix, sheet, loop Edge attributes type (sequence, structure) length Karsten Borgwardt et al. - Protein function prediction via graph kernels 10
12 Protein graph kernel (Kashima et al. (2003) and Gärtner et al. (2003)) compares walks of identical length l l 1 k walk v 1,...,v l, w 1,..., w l = i =1 Walks are similar, if along both walks types of secondary structure elements (SSEs) are the same distances between SSEs are similar chemical properties of SSEs are similar k step v i,v i 1, w i, w i 1 11
13 Example: Protein kernel Protein A S S S S Protein B S Similar (H,10,S,1,S,3,H) (H,9,S,1,S,3,H) 12
14 Example: Protein kernel Protein A S S S S Protein B S Dissimilar (H,10,S,1,S) (S,3,H,5,S) 13
15 Evaluation: enzymes vs. non-enzymes 10-fold cross-validation on 1128 proteins from dataset by Dobson and Doig (2003); 59 % are enzymes. Kernel type accuracy SD Vector kernel Optimized vector kernel Graph kernel Graph kernel without structure Graph kernel with global info DALI classifier Karsten Borgwardt et al. - Protein function prediction via graph kernels 14
16 Attribute selection Which structural or chemical attribute is most important for correct classification? For this purpose, we employ hyperkernels (Ong et. al, 2003). Hyperkernels find an optimal linear combination of input kernel matrices : m i=1 β i K i minimizing training error and fulfilling regularization constraints Karsten Borgwardt et al. - Protein function prediction via graph kernels 15
17 Our approach: Attribute selection Calculate kernel matrix for 600 proteins on graph model with only ONE single attribute! Repeat this for all attributes Normalize these kernel matrices Determine hyperkernel combination Weights then reflect contribution of individual attributes to correct classification 16
18 Attribute selection Attribute EC 1 EC 2 EC 3 EC 4 EC 5 EC 6 Amino acid length bin van der Waals 3-bin Hydrophobicity 3-bin Polarity bin Polarizability d length 0.40 Total van der Waals Total Hydrophobicity Total Polarity Total Polarizability Karsten Borgwardt et al. - Protein function prediction via graph kernels 17
19 Discussion Novel combined approach to protein function prediction integrating sequence, structure and chemical information Reaches state-of-the-art classification accuracy on less information; higher accuracy levels on same amount of information Hyperkernels for finding most interesting protein characteristics Karsten Borgwardt et al. - Protein function prediction via graph kernels 18
20 Discussion More detailed graph models (amino acids, atoms) might be more interesting, yet raise computational difficulties (graphs too large!) Two directions of future research: Efficient, yet expressive graph kernels for structure Integrating more proteomic information, e.g. surface pockets, into our graph model Karsten Borgwardt et al. - Protein function prediction via graph kernels 19
21 The End Thank you! Questions? Karsten Borgwardt et al. - Protein function prediction via graph kernels 20
22 ARTS: Accurate Recognition of Transcription Starts in human Sören Sonnenburg, Alexander Zien,, Gunnar Rätsch Fraunhofer FIRST.IDA, Kekuléstr. 7, Berlin, Germany Friedrich Miescher Laboratory of the Max Planck Society, Max Planck Institute for Biological Cybernetics, Spemannstr , Tübingen, Germany
23 Promoter Detection Overview: Transcription Start Site (TSS) Features to describe the TSS Our approach Evaluation with current methods Example - Protocadherin-α Summary Sonnenburg, Zien, Rätsch 1
24 Promoter Detection Transcription Start Site - Properties POL II binds to a rather vague region of [ 20,+20] bp Upstream of TSS: promoter containing transcription factor binding sites Downstream of TSS: 5 UTR, and further downstream coding regions and introns (different statistics) 3D structure of the promoter must allow the transcription factors to bind Promoter Prediction is non-trivial Sonnenburg, Zien, Rätsch 2
25 Promoter Detection Features to describe the TSS TFBS in Promoter region condition: DNA should not be too twisted CpG islands (often over TSS/first exon; in most, but not all promoters) TSS with TATA box ( 30 bp upstream) Exon content in UTR 5 region Distance to first donor splice site Idea: Combine weak features to build strong promoter predictor Sonnenburg, Zien, Rätsch 3
26 Promoter Detection The ARTS Approach use SVM classifier ( Ns ) f(x) = sign y i α i k(x,x i ) + b i=1 key ingredient is kernel k(x,x ) similarity of two sequences use 5 sub-kernels suited to model the aforementioned features k(x, x ) = k TSS (x, x )+k CpG (x, x )+k coding (x, x )+k energy (x, x )+k twist (x, x ) Sonnenburg, Zien, Rätsch 4
27 Promoter Detection The 5 sub-kernels 1. TSS signal (including parts of core promoter with TATA box) use Weighted Degree Shift kernel 2. CpG Islands, distant enhancers and TFBS upstream of TSS use Spectrum kernel (large window upstream of TSS) 3. Model coding sequence TFBS downstream of TSS use another Spectrum kernel (small window downstream of TSS) 4. Stacking energy of DNA use btwist energy of dinucleotides with Linear kernel 5. Twistedness of DNA use btwist angle of dinucleotides with Linear kernel Sonnenburg, Zien, Rätsch 5
28 Promoter Detection Weighted Degree Shift Kernel x 1 k(x1,x2) = w6,3 + w6,-3 + w3,4 x 2 Count matching substrings of length 1...d Weight according to length of the match β 1...β d Position dependent but tolerates shifts of up to S k(x,x ) = d k=1 L k+1 β k l=1 S s=0 s+l L δ s (I(x[k : l + s]=x [k : l])+i(x[k : l]=x [k : l + s])) x[k : l] := subsequence of x of length k starting at position l Sonnenburg, Zien, Rätsch 6
29 Promoter Detection Training Data Generation True TSS: From dbtssv4 (based on hg16) extract putative TSS windows of size [ 1000, +1000] Decoy TSS: Annotate dbtssv4 with transcription-stop (via BLAT alignment of mrnas) From the interior of the gene (+100bp to gene end) sample negatives for training (10 per positive), again windows [ 1000,+1000] Processing: 8508 positive, negative examples Split into disjoint training and validation set (50% : 50%) Sonnenburg, Zien, Rätsch 7
30 Promoter Detection Training Model Selection 16 kernel parameters + SVM regularization to be tuned! Full grid search infeasible Local axis-parallel searches instead SVM training/evaluation on > 10, 000 examples computationally too demanding Speedup trick: f(x) = N s i=1 α i k(x i, x) + b = N s i=1 α i Φ(x i ) Φ(x) + b = w Φ(x) + b } {{ } w f(x) before: O(N s dls) now: = O(dL) speedup factor up to N s S Large Scale Training and Evaluation possible Sonnenburg, Zien, Rätsch 8
31 Promoter Detection Comparison Current state-of-the-art methods: FirstEF [Davuluri, Grosse, Zhang; 2001, Nat Genet] QDF: for promoter, donor, first exon, WM Range: [ 1500, +500] McPromoter [Ohler, Liao, Niemann, Rubin; 2002, Genome Biol] GHMM with IMC for 6 regions (e.g. upstream, TATA) NN Range: [ 250, +50] Eponine [Down, Hubbard; 2002 Genome Res] RVM: WM with positional distribution for 4 regions (e.g. TATA, CpG) Range: [ 200, +200] Do a genome wide evaluation! How to do a fair comparison? Sonnenburg, Zien, Rätsch 9
32 Promoter Detection Evaluation Idea: Only consider new TSS from dbtssv5-dbtssv4, with max 30% overlap 1. Compute genome wide outputs for each TSF 2. Decrease resolution: divide genome into non-overlapping fixed size chunks (e.g. 50 or 500) 3. Annotate dbtssv5 TSS with gene end 4. Label chunk positive if intersects with [T SS 20bp, T SS + 20bp] 5. Label chunk negative [T SS + 21bp, GeneEnd] Sonnenburg, Zien, Rätsch 10
33 Promoter Detection Results Receiver Operator Characteristic Curve and Precision Recall Curve 35% true positives at a false positive rate of 1/1000 (best other method find about a half (18%)) Sonnenburg, Zien, Rätsch 11
34 5.5 Promoter Detection What does ARTS do better? Entropy and Relative Entropy entropy auroc: 86.5% auprc: 49.8% entropy auroc: 86.5% auprc: 49.8% relative entropy auroc: 86.5% auprc: 49.8% Di-nucleotide Frequency strong discriminative signal around TSS Sonnenburg, Zien, Rätsch 12
35 Promoter Detection Which kernel captures most information? 96 using or removing single kernels area under ROC Curve (in %) TSS WD shift Promotor Spectrum 1st Exon Spectrum Angles Linear Most important Weighted Degree Shift kernel modelling the TSS signal Sonnenburg, Zien, Rätsch 13
36 Promoter Detection Alternative TSS - Protocadherin-α Sonnenburg, Zien, Rätsch 14
37 Promoter Detection Conclusion Developed a new TSF finder, ARTS In genome-wide evaluation achieves state-of-the-art results: ARTS about 35% true positives at a false positive rate of 1/1000 (best other method about a half, 18%) Reason: intensively modelling the TSS region, large scale svm training/evaluation with string kernels Future work: Drosophila, C.elegans, Zebrafish,... Poster: H56 Datasets, Genomebrowser custom track, a lot more details: Source code of SHOGUN toolbox used to train ARTS freely available: Sonnenburg, Zien, Rätsch 15
38 The end See you tomorrow! Next topic: Clustering in Bioinformatics Karsten Borgwardt: Data Mining in Bioinformatics, Page 2
ARTS: Accurate Recognition of Transcription Starts in human
 ARTS: Accurate Recognition of Transcription Starts in human Sören Sonnenburg, Alexander Zien,, Gunnar Rätsch Fraunhofer FIRST.IDA, Kekuléstr. 7, 12489 Berlin, Germany Friedrich Miescher Laboratory of the
ARTS: Accurate Recognition of Transcription Starts in human Sören Sonnenburg, Alexander Zien,, Gunnar Rätsch Fraunhofer FIRST.IDA, Kekuléstr. 7, 12489 Berlin, Germany Friedrich Miescher Laboratory of the
Discovering Common Sequence Variation in A. thaliana. Gunnar Rätsch
 Machine Learning Methods for Discovering Common Sequence Variation in A. thaliana Gunnar Rätsch Friedrich Miescher Laboratory, Max Planck Society, Tübingen Technical University Berlin March 31, 2008 Current
Machine Learning Methods for Discovering Common Sequence Variation in A. thaliana Gunnar Rätsch Friedrich Miescher Laboratory, Max Planck Society, Tübingen Technical University Berlin March 31, 2008 Current
Machine Learning Methods for RNA-seq-based Transcriptome Reconstruction
 Machine Learning Methods for RNA-seq-based Transcriptome Reconstruction Gunnar Rätsch Friedrich Miescher Laboratory Max Planck Society, Tübingen, Germany NGS Bioinformatics Meeting, Paris (March 24, 2010)
Machine Learning Methods for RNA-seq-based Transcriptome Reconstruction Gunnar Rätsch Friedrich Miescher Laboratory Max Planck Society, Tübingen, Germany NGS Bioinformatics Meeting, Paris (March 24, 2010)
Protein Synthesis Notes
 Protein Synthesis Notes Protein Synthesis: Overview Transcription: synthesis of mrna under the direction of DNA. Translation: actual synthesis of a polypeptide under the direction of mrna. Transcription
Protein Synthesis Notes Protein Synthesis: Overview Transcription: synthesis of mrna under the direction of DNA. Translation: actual synthesis of a polypeptide under the direction of mrna. Transcription
MATH 5610, Computational Biology
 MATH 5610, Computational Biology Lecture 2 Intro to Molecular Biology (cont) Stephen Billups University of Colorado at Denver MATH 5610, Computational Biology p.1/24 Announcements Error on syllabus Class
MATH 5610, Computational Biology Lecture 2 Intro to Molecular Biology (cont) Stephen Billups University of Colorado at Denver MATH 5610, Computational Biology p.1/24 Announcements Error on syllabus Class
CSE : Computational Issues in Molecular Biology. Lecture 19. Spring 2004
 CSE 397-497: Computational Issues in Molecular Biology Lecture 19 Spring 2004-1- Protein structure Primary structure of protein is determined by number and order of amino acids within polypeptide chain.
CSE 397-497: Computational Issues in Molecular Biology Lecture 19 Spring 2004-1- Protein structure Primary structure of protein is determined by number and order of amino acids within polypeptide chain.
Make the protein through the genetic dogma process.
 Make the protein through the genetic dogma process. Coding Strand 5 AGCAATCATGGATTGGGTACATTTGTAACTGT 3 Template Strand mrna Protein Complete the table. DNA strand DNA s strand G mrna A C U G T A T Amino
Make the protein through the genetic dogma process. Coding Strand 5 AGCAATCATGGATTGGGTACATTTGTAACTGT 3 Template Strand mrna Protein Complete the table. DNA strand DNA s strand G mrna A C U G T A T Amino
Year III Pharm.D Dr. V. Chitra
 Year III Pharm.D Dr. V. Chitra 1 Genome entire genetic material of an individual Transcriptome set of transcribed sequences Proteome set of proteins encoded by the genome 2 Only one strand of DNA serves
Year III Pharm.D Dr. V. Chitra 1 Genome entire genetic material of an individual Transcriptome set of transcribed sequences Proteome set of proteins encoded by the genome 2 Only one strand of DNA serves
Transcription in Eukaryotes
 Transcription in Eukaryotes Biology I Hayder A Giha Transcription Transcription is a DNA-directed synthesis of RNA, which is the first step in gene expression. Gene expression, is transformation of the
Transcription in Eukaryotes Biology I Hayder A Giha Transcription Transcription is a DNA-directed synthesis of RNA, which is the first step in gene expression. Gene expression, is transformation of the
2/23/16. Protein-Protein Interactions. Protein Interactions. Protein-Protein Interactions: The Interactome
 Protein-Protein Interactions Protein Interactions A Protein may interact with: Other proteins Nucleic Acids Small molecules Protein-Protein Interactions: The Interactome Experimental methods: Mass Spec,
Protein-Protein Interactions Protein Interactions A Protein may interact with: Other proteins Nucleic Acids Small molecules Protein-Protein Interactions: The Interactome Experimental methods: Mass Spec,
TIGR THE INSTITUTE FOR GENOMIC RESEARCH
 Introduction to Genome Annotation: Overview of What You Will Learn This Week C. Robin Buell May 21, 2007 Types of Annotation Structural Annotation: Defining genes, boundaries, sequence motifs e.g. ORF,
Introduction to Genome Annotation: Overview of What You Will Learn This Week C. Robin Buell May 21, 2007 Types of Annotation Structural Annotation: Defining genes, boundaries, sequence motifs e.g. ORF,
Eukaryotic Gene Structure
 Eukaryotic Gene Structure Terminology Genome entire genetic material of an individual Transcriptome set of transcribed sequences Proteome set of proteins encoded by the genome 2 Gene Basic physical and
Eukaryotic Gene Structure Terminology Genome entire genetic material of an individual Transcriptome set of transcribed sequences Proteome set of proteins encoded by the genome 2 Gene Basic physical and
BIOINFORMATICS Introduction
 BIOINFORMATICS Introduction Mark Gerstein, Yale University bioinfo.mbb.yale.edu/mbb452a 1 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu What is Bioinformatics? (Molecular) Bio -informatics One idea
BIOINFORMATICS Introduction Mark Gerstein, Yale University bioinfo.mbb.yale.edu/mbb452a 1 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu What is Bioinformatics? (Molecular) Bio -informatics One idea
ORTHOMINE - A dataset of Drosophila core promoters and its analysis. Sumit Middha Advisor: Dr. Peter Cherbas
 ORTHOMINE - A dataset of Drosophila core promoters and its analysis Sumit Middha Advisor: Dr. Peter Cherbas Introduction Challenges and Motivation D melanogaster Promoter Dataset Expanding promoter sequences
ORTHOMINE - A dataset of Drosophila core promoters and its analysis Sumit Middha Advisor: Dr. Peter Cherbas Introduction Challenges and Motivation D melanogaster Promoter Dataset Expanding promoter sequences
From Variants to Pathways: Agilent GeneSpring GX s Variant Analysis Workflow
 From Variants to Pathways: Agilent GeneSpring GX s Variant Analysis Workflow Technical Overview Import VCF Introduction Next-generation sequencing (NGS) studies have created unanticipated challenges with
From Variants to Pathways: Agilent GeneSpring GX s Variant Analysis Workflow Technical Overview Import VCF Introduction Next-generation sequencing (NGS) studies have created unanticipated challenges with
CMPS 6630: Introduction to Computational Biology and Bioinformatics. Secondary Structure Prediction
 CMPS 6630: Introduction to Computational Biology and Bioinformatics Secondary Structure Prediction Secondary Structure Annotation Given a macromolecular structure Identify the regions of secondary structure
CMPS 6630: Introduction to Computational Biology and Bioinformatics Secondary Structure Prediction Secondary Structure Annotation Given a macromolecular structure Identify the regions of secondary structure
Gene Structure & Gene Finding Part II
 Gene Structure & Gene Finding Part II David Wishart david.wishart@ualberta.ca 30,000 metabolite Gene Finding in Eukaryotes Eukaryotes Complex gene structure Large genomes (0.1 to 10 billion bp) Exons and
Gene Structure & Gene Finding Part II David Wishart david.wishart@ualberta.ca 30,000 metabolite Gene Finding in Eukaryotes Eukaryotes Complex gene structure Large genomes (0.1 to 10 billion bp) Exons and
Regulation of eukaryotic transcription:
 Promoter definition by mass genome annotation data: in silico primer extension EMBNET course Bioinformatics of transcriptional regulation Jan 28 2008 Christoph Schmid Regulation of eukaryotic transcription:
Promoter definition by mass genome annotation data: in silico primer extension EMBNET course Bioinformatics of transcriptional regulation Jan 28 2008 Christoph Schmid Regulation of eukaryotic transcription:
Machine learning applications in genomics: practical issues & challenges. Yuzhen Ye School of Informatics and Computing, Indiana University
 Machine learning applications in genomics: practical issues & challenges Yuzhen Ye School of Informatics and Computing, Indiana University Reference Machine learning applications in genetics and genomics
Machine learning applications in genomics: practical issues & challenges Yuzhen Ye School of Informatics and Computing, Indiana University Reference Machine learning applications in genetics and genomics
Multiple choice questions (numbers in brackets indicate the number of correct answers)
 1 Multiple choice questions (numbers in brackets indicate the number of correct answers) February 1, 2013 1. Ribose is found in Nucleic acids Proteins Lipids RNA DNA (2) 2. Most RNA in cells is transfer
1 Multiple choice questions (numbers in brackets indicate the number of correct answers) February 1, 2013 1. Ribose is found in Nucleic acids Proteins Lipids RNA DNA (2) 2. Most RNA in cells is transfer
The Double Helix. DNA and RNA, part 2. Part A. Hint 1. The difference between purines and pyrimidines. Hint 2. Distinguish purines from pyrimidines
 DNA and RNA, part 2 Due: 3:00pm on Wednesday, September 24, 2014 You will receive no credit for items you complete after the assignment is due. Grading Policy The Double Helix DNA, or deoxyribonucleic
DNA and RNA, part 2 Due: 3:00pm on Wednesday, September 24, 2014 You will receive no credit for items you complete after the assignment is due. Grading Policy The Double Helix DNA, or deoxyribonucleic
Outline. Introduction to ab initio and evidence-based gene finding. Prokaryotic gene predictions
 Outline Introduction to ab initio and evidence-based gene finding Overview of computational gene predictions Different types of eukaryotic gene predictors Common types of gene prediction errors Wilson
Outline Introduction to ab initio and evidence-based gene finding Overview of computational gene predictions Different types of eukaryotic gene predictors Common types of gene prediction errors Wilson
Genome Sequence Assembly
 Genome Sequence Assembly Learning Goals: Introduce the field of bioinformatics Familiarize the student with performing sequence alignments Understand the assembly process in genome sequencing Introduction:
Genome Sequence Assembly Learning Goals: Introduce the field of bioinformatics Familiarize the student with performing sequence alignments Understand the assembly process in genome sequencing Introduction:
Transcription & post transcriptional modification
 Transcription & post transcriptional modification Transcription The synthesis of RNA molecules using DNA strands as the templates so that the genetic information can be transferred from DNA to RNA Similarity
Transcription & post transcriptional modification Transcription The synthesis of RNA molecules using DNA strands as the templates so that the genetic information can be transferred from DNA to RNA Similarity
Gene Expression Technology
 Gene Expression Technology Bing Zhang Department of Biomedical Informatics Vanderbilt University bing.zhang@vanderbilt.edu Gene expression Gene expression is the process by which information from a gene
Gene Expression Technology Bing Zhang Department of Biomedical Informatics Vanderbilt University bing.zhang@vanderbilt.edu Gene expression Gene expression is the process by which information from a gene
GENETICS الفريق الطبي االكاديمي. DNA Genes & Chromosomes. DONE BY : Buthaina Al-masaeed & Yousef Qandeel. Page 0
 GENETICS ومن أحياها DNA Genes & Chromosomes الفريق الطبي االكاديمي DNA Genes & Chromosomes DONE BY : Buthaina Al-masaeed & Yousef Qandeel Page 0 T(0:44 min) In the pre lecture we take about the back bone
GENETICS ومن أحياها DNA Genes & Chromosomes الفريق الطبي االكاديمي DNA Genes & Chromosomes DONE BY : Buthaina Al-masaeed & Yousef Qandeel Page 0 T(0:44 min) In the pre lecture we take about the back bone
Discovery of Transcription Factor Binding Sites with Deep Convolutional Neural Networks
 Discovery of Transcription Factor Binding Sites with Deep Convolutional Neural Networks Reesab Pathak Dept. of Computer Science Stanford University rpathak@stanford.edu Abstract Transcription factors are
Discovery of Transcription Factor Binding Sites with Deep Convolutional Neural Networks Reesab Pathak Dept. of Computer Science Stanford University rpathak@stanford.edu Abstract Transcription factors are
STRUCTURAL BIOLOGY. α/β structures Closed barrels Open twisted sheets Horseshoe folds
 STRUCTURAL BIOLOGY α/β structures Closed barrels Open twisted sheets Horseshoe folds The α/β domains Most frequent domain structures are α/β domains: A central parallel or mixed β sheet Surrounded by α
STRUCTURAL BIOLOGY α/β structures Closed barrels Open twisted sheets Horseshoe folds The α/β domains Most frequent domain structures are α/β domains: A central parallel or mixed β sheet Surrounded by α
Predicting the Coupling Specif icity of G-protein Coupled Receptors to G-proteins by Support Vector Machines
 Article Predicting the Coupling Specif icity of G-protein Coupled Receptors to G-proteins by Support Vector Machines Cui-Ping Guan, Zhen-Ran Jiang, and Yan-Hong Zhou* Hubei Bioinformatics and Molecular
Article Predicting the Coupling Specif icity of G-protein Coupled Receptors to G-proteins by Support Vector Machines Cui-Ping Guan, Zhen-Ran Jiang, and Yan-Hong Zhou* Hubei Bioinformatics and Molecular
ProSOM: core promoter prediction based on unsupervised clustering of DNA physical profiles
 BIOINFORMATICS Vol. 24 ISMB 2008, pages i24 i31 doi:1093/bioinformatics/btn172 ProSOM: core promoter prediction based on unsupervised clustering of DNA physical profiles Thomas Abeel 1,2, Yvan Saeys 1,2,
BIOINFORMATICS Vol. 24 ISMB 2008, pages i24 i31 doi:1093/bioinformatics/btn172 ProSOM: core promoter prediction based on unsupervised clustering of DNA physical profiles Thomas Abeel 1,2, Yvan Saeys 1,2,
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Origin use and efficiency are similar among WT, rrm3, pif1-m2, and pif1-m2; rrm3 strains. A. Analysis of fork progression around confirmed and likely origins (from cerevisiae.oridb.org).
Supplementary Figure 1 Origin use and efficiency are similar among WT, rrm3, pif1-m2, and pif1-m2; rrm3 strains. A. Analysis of fork progression around confirmed and likely origins (from cerevisiae.oridb.org).
Fig Ch 17: From Gene to Protein
 Fig. 17-1 Ch 17: From Gene to Protein Basic Principles of Transcription and Translation RNA is the intermediate between genes and the proteins for which they code Transcription is the synthesis of RNA
Fig. 17-1 Ch 17: From Gene to Protein Basic Principles of Transcription and Translation RNA is the intermediate between genes and the proteins for which they code Transcription is the synthesis of RNA
Genie Gene Finding in Drosophila melanogaster
 Methods Gene Finding in Drosophila melanogaster Martin G. Reese, 1,2,4 David Kulp, 2 Hari Tammana, 2 and David Haussler 2,3 1 Berkeley Drosophila Genome Project, Department of Molecular and Cell Biology,
Methods Gene Finding in Drosophila melanogaster Martin G. Reese, 1,2,4 David Kulp, 2 Hari Tammana, 2 and David Haussler 2,3 1 Berkeley Drosophila Genome Project, Department of Molecular and Cell Biology,
Bi 8 Lecture 7. Ellen Rothenberg 26 January Reading: Ch. 3, pp ; panel 3-1
 Bi 8 Lecture 7 PROTEIN STRUCTURE, Functional analysis, and evolution Ellen Rothenberg 26 January 2016 Reading: Ch. 3, pp. 109-134; panel 3-1 (end with free amine) aromatic, hydrophobic small, hydrophilic
Bi 8 Lecture 7 PROTEIN STRUCTURE, Functional analysis, and evolution Ellen Rothenberg 26 January 2016 Reading: Ch. 3, pp. 109-134; panel 3-1 (end with free amine) aromatic, hydrophobic small, hydrophilic
Gene Signal Estimates from Exon Arrays
 Gene Signal Estimates from Exon Arrays I. Introduction: With exon arrays like the GeneChip Human Exon 1.0 ST Array, researchers can examine the transcriptional profile of an entire gene (Figure 1). Being
Gene Signal Estimates from Exon Arrays I. Introduction: With exon arrays like the GeneChip Human Exon 1.0 ST Array, researchers can examine the transcriptional profile of an entire gene (Figure 1). Being
UCSC Genome Browser. Introduction to ab initio and evidence-based gene finding
 UCSC Genome Browser Introduction to ab initio and evidence-based gene finding Wilson Leung 06/2006 Outline Introduction to annotation ab initio gene finding Basics of the UCSC Browser Evidence-based gene
UCSC Genome Browser Introduction to ab initio and evidence-based gene finding Wilson Leung 06/2006 Outline Introduction to annotation ab initio gene finding Basics of the UCSC Browser Evidence-based gene
Parameters tuning boosts hypersmurf predictions of rare deleterious non-coding genetic variants
 Parameters tuning boosts hypersmurf predictions of rare deleterious non-coding genetic variants The regulatory code that determines whether and how a given genetic variant affects the function of a regulatory
Parameters tuning boosts hypersmurf predictions of rare deleterious non-coding genetic variants The regulatory code that determines whether and how a given genetic variant affects the function of a regulatory
TRANSCRIPTION AND PROCESSING OF RNA
 TRANSCRIPTION AND PROCESSING OF RNA 1. The steps of gene expression. 2. General characterization of transcription: steps, components of transcription apparatus. 3. Transcription of eukaryotic structural
TRANSCRIPTION AND PROCESSING OF RNA 1. The steps of gene expression. 2. General characterization of transcription: steps, components of transcription apparatus. 3. Transcription of eukaryotic structural
Predicting DNA Methylation State of CpG Dinucleotide Using Genome Topological Features and Deep Networks
 The University of Southern Mississippi The Aquila Digital Community Master's Theses Spring 5-2016 Predicting DNA Methylation State of CpG Dinucleotide Using Genome Topological Features and Deep Networks
The University of Southern Mississippi The Aquila Digital Community Master's Theses Spring 5-2016 Predicting DNA Methylation State of CpG Dinucleotide Using Genome Topological Features and Deep Networks
Chapter 24: Promoters and Enhancers
 Chapter 24: Promoters and Enhancers A typical gene transcribed by RNA polymerase II has a promoter that usually extends upstream from the site where transcription is initiated the (#1) of transcription
Chapter 24: Promoters and Enhancers A typical gene transcribed by RNA polymerase II has a promoter that usually extends upstream from the site where transcription is initiated the (#1) of transcription
Biotechnology Explorer
 Biotechnology Explorer C. elegans Behavior Kit Bioinformatics Supplement explorer.bio-rad.com Catalog #166-5120EDU This kit contains temperature-sensitive reagents. Open immediately and see individual
Biotechnology Explorer C. elegans Behavior Kit Bioinformatics Supplement explorer.bio-rad.com Catalog #166-5120EDU This kit contains temperature-sensitive reagents. Open immediately and see individual
Convolutional Kitchen Sinks for Transcription Factor Binding Site Prediction
 Convolutional Kitchen Sinks for Transcription Factor Binding Site Prediction Alyssa Morrow*, Vaishaal Shankar* Anthony Joseph, Benjamin Recht, Nir Yosef Transcription Factor A protein that binds to DNA
Convolutional Kitchen Sinks for Transcription Factor Binding Site Prediction Alyssa Morrow*, Vaishaal Shankar* Anthony Joseph, Benjamin Recht, Nir Yosef Transcription Factor A protein that binds to DNA
The common structure of a DNA nucleotide. Hewitt
 GENETICS Unless otherwise noted* the artwork and photographs in this slide show are original and by Burt Carter. Permission is granted to use them for non-commercial, non-profit educational purposes provided
GENETICS Unless otherwise noted* the artwork and photographs in this slide show are original and by Burt Carter. Permission is granted to use them for non-commercial, non-profit educational purposes provided
Exploring Similarities of Conserved Domains/Motifs
 Exploring Similarities of Conserved Domains/Motifs Sotiria Palioura Abstract Traditionally, proteins are represented as amino acid sequences. There are, though, other (potentially more exciting) representations;
Exploring Similarities of Conserved Domains/Motifs Sotiria Palioura Abstract Traditionally, proteins are represented as amino acid sequences. There are, though, other (potentially more exciting) representations;
Chimp Sequence Annotation: Region 2_3
 Chimp Sequence Annotation: Region 2_3 Jeff Howenstein March 30, 2007 BIO434W Genomics 1 Introduction We received region 2_3 of the ChimpChunk sequence, and the first step we performed was to run RepeatMasker
Chimp Sequence Annotation: Region 2_3 Jeff Howenstein March 30, 2007 BIO434W Genomics 1 Introduction We received region 2_3 of the ChimpChunk sequence, and the first step we performed was to run RepeatMasker
Original article: IDENTIFYING DNA SPLICE SITES USING PATTERNS STATISTICAL PROPERTIES AND FUZZY NEURAL NETWORKS
 Original article: IDENTIFYING DNA SPLICE SITES USING PATTERNS STATISTICAL PROPERTIES AND FUZZY NEURAL NETWORKS Essam Al-Daoud Computer Science Department, Faculty of Science and Information Technology,
Original article: IDENTIFYING DNA SPLICE SITES USING PATTERNS STATISTICAL PROPERTIES AND FUZZY NEURAL NETWORKS Essam Al-Daoud Computer Science Department, Faculty of Science and Information Technology,
RNA : functional role
 RNA : functional role Hamad Yaseen, PhD MLS Department, FAHS Hamad.ali@hsc.edu.kw RNA mrna rrna trna 1 From DNA to Protein -Outline- From DNA to RNA From RNA to Protein From DNA to RNA Transcription: Copying
RNA : functional role Hamad Yaseen, PhD MLS Department, FAHS Hamad.ali@hsc.edu.kw RNA mrna rrna trna 1 From DNA to Protein -Outline- From DNA to RNA From RNA to Protein From DNA to RNA Transcription: Copying
How to view Results with Scaffold. Proteomics Shared Resource
 How to view Results with Scaffold Proteomics Shared Resource Starting out Download Scaffold from http://www.proteomes oftware.com/proteom e_software_prod_sca ffold_download.html Follow installation instructions
How to view Results with Scaffold Proteomics Shared Resource Starting out Download Scaffold from http://www.proteomes oftware.com/proteom e_software_prod_sca ffold_download.html Follow installation instructions
Bio11 Announcements. Ch 21: DNA Biology and Technology. DNA Functions. DNA and RNA Structure. How do DNA and RNA differ? What are genes?
 Bio11 Announcements TODAY Genetics (review) and quiz (CP #4) Structure and function of DNA Extra credit due today Next week in lab: Case study presentations Following week: Lab Quiz 2 Ch 21: DNA Biology
Bio11 Announcements TODAY Genetics (review) and quiz (CP #4) Structure and function of DNA Extra credit due today Next week in lab: Case study presentations Following week: Lab Quiz 2 Ch 21: DNA Biology
Annotating 7G24-63 Justin Richner May 4, Figure 1: Map of my sequence
 Annotating 7G24-63 Justin Richner May 4, 2005 Zfh2 exons Thd1 exons Pur-alpha exons 0 40 kb 8 = 1 kb = LINE, Penelope = DNA/Transib, Transib1 = DINE = Novel Repeat = LTR/PAO, Diver2 I = LTR/Gypsy, Invader
Annotating 7G24-63 Justin Richner May 4, 2005 Zfh2 exons Thd1 exons Pur-alpha exons 0 40 kb 8 = 1 kb = LINE, Penelope = DNA/Transib, Transib1 = DINE = Novel Repeat = LTR/PAO, Diver2 I = LTR/Gypsy, Invader
Structural Bioinformatics (C3210) DNA and RNA Structure
 Structural Bioinformatics (C3210) DNA and RNA Structure Importance of DNA/RNA 3D Structure Nucleic acids are essential materials found in all living organisms. Their main function is to maintain and transmit
Structural Bioinformatics (C3210) DNA and RNA Structure Importance of DNA/RNA 3D Structure Nucleic acids are essential materials found in all living organisms. Their main function is to maintain and transmit
Transcription Gene regulation
 Transcription Gene regulation The machine that transcribes a gene is composed of perhaps 50 proteins, including RNA polymerase, the enzyme that converts DNA code into RNA code. A crew of transcription
Transcription Gene regulation The machine that transcribes a gene is composed of perhaps 50 proteins, including RNA polymerase, the enzyme that converts DNA code into RNA code. A crew of transcription
SSA Signal Search Analysis II
 SSA Signal Search Analysis II SSA other applications - translation In contrast to translation initiation in bacteria, translation initiation in eukaryotes is not guided by a Shine-Dalgarno like motif.
SSA Signal Search Analysis II SSA other applications - translation In contrast to translation initiation in bacteria, translation initiation in eukaryotes is not guided by a Shine-Dalgarno like motif.
A Propagation-based Algorithm for Inferring Gene-Disease Associations
 A Propagation-based Algorithm for Inferring Gene-Disease Associations Oron Vanunu Roded Sharan Abstract: A fundamental challenge in human health is the identification of diseasecausing genes. Recently,
A Propagation-based Algorithm for Inferring Gene-Disease Associations Oron Vanunu Roded Sharan Abstract: A fundamental challenge in human health is the identification of diseasecausing genes. Recently,
Mapping strategies for sequence reads
 Mapping strategies for sequence reads Ernest Turro University of Cambridge 21 Oct 2013 Quantification A basic aim in genomics is working out the contents of a biological sample. 1. What distinct elements
Mapping strategies for sequence reads Ernest Turro University of Cambridge 21 Oct 2013 Quantification A basic aim in genomics is working out the contents of a biological sample. 1. What distinct elements
CLASS 3.5: 03/29/07 EUKARYOTIC TRANSCRIPTION I: PROMOTERS AND ENHANCERS
 CLASS 3.5: 03/29/07 EUKARYOTIC TRANSCRIPTION I: PROMOTERS AND ENHANCERS A. Promoters and Polymerases (RNA pols): 1. General characteristics - Initiation of transcription requires a. Transcription factors
CLASS 3.5: 03/29/07 EUKARYOTIC TRANSCRIPTION I: PROMOTERS AND ENHANCERS A. Promoters and Polymerases (RNA pols): 1. General characteristics - Initiation of transcription requires a. Transcription factors
SIBC504: TRANSCRIPTION & RNA PROCESSING Assistant Professor Dr. Chatchawan Srisawat
 SIBC504: TRANSCRIPTION & RNA PROCESSING Assistant Professor Dr. Chatchawan Srisawat TRANSCRIPTION: AN OVERVIEW Transcription: the synthesis of a single-stranded RNA from a doublestranded DNA template.
SIBC504: TRANSCRIPTION & RNA PROCESSING Assistant Professor Dr. Chatchawan Srisawat TRANSCRIPTION: AN OVERVIEW Transcription: the synthesis of a single-stranded RNA from a doublestranded DNA template.
Transcription factor binding site prediction in vivo using DNA sequence and shape features
 Transcription factor binding site prediction in vivo using DNA sequence and shape features Anthony Mathelier, Lin Yang, Tsu-Pei Chiu, Remo Rohs, and Wyeth Wasserman anthony.mathelier@gmail.com @AMathelier
Transcription factor binding site prediction in vivo using DNA sequence and shape features Anthony Mathelier, Lin Yang, Tsu-Pei Chiu, Remo Rohs, and Wyeth Wasserman anthony.mathelier@gmail.com @AMathelier
Chapter 8 DNA Recognition in Prokaryotes by Helix-Turn-Helix Motifs
 Chapter 8 DNA Recognition in Prokaryotes by Helix-Turn-Helix Motifs 1. Helix-turn-helix proteins 2. Zinc finger proteins 3. Leucine zipper proteins 4. Beta-scaffold factors 5. Others λ-repressor AND CRO
Chapter 8 DNA Recognition in Prokaryotes by Helix-Turn-Helix Motifs 1. Helix-turn-helix proteins 2. Zinc finger proteins 3. Leucine zipper proteins 4. Beta-scaffold factors 5. Others λ-repressor AND CRO
Regulation of gene expression. (Lehninger pg )
 Regulation of gene expression (Lehninger pg. 1072-1085) Today s lecture Gene expression Constitutive, inducible, repressible genes Specificity factors, activators, repressors Negative and positive gene
Regulation of gene expression (Lehninger pg. 1072-1085) Today s lecture Gene expression Constitutive, inducible, repressible genes Specificity factors, activators, repressors Negative and positive gene
Review of Protein (one or more polypeptide) A polypeptide is a long chain of..
 Gene expression Review of Protein (one or more polypeptide) A polypeptide is a long chain of.. In a protein, the sequence of amino acid determines its which determines the protein s A protein with an enzymatic
Gene expression Review of Protein (one or more polypeptide) A polypeptide is a long chain of.. In a protein, the sequence of amino acid determines its which determines the protein s A protein with an enzymatic
Lecture 10: Motif Finding Regulatory element detection using correlation with expression
 CS5238 Combinatorial methods in bioinformatics 2006/2007 Semester 1 Lecture 10: Motif Finding Lecturer: Wing-Kin Sung Scribe: Zhang Jingbo, Shrikant Kashyap 10.1 Regulatory element detection using correlation
CS5238 Combinatorial methods in bioinformatics 2006/2007 Semester 1 Lecture 10: Motif Finding Lecturer: Wing-Kin Sung Scribe: Zhang Jingbo, Shrikant Kashyap 10.1 Regulatory element detection using correlation
Outline. Annotation of Drosophila Primer. Gene structure nomenclature. Muller element nomenclature. GEP Drosophila annotation projects 01/04/2018
 Outline Overview of the GEP annotation projects Annotation of Drosophila Primer January 2018 GEP annotation workflow Practice applying the GEP annotation strategy Wilson Leung and Chris Shaffer AAACAACAATCATAAATAGAGGAAGTTTTCGGAATATACGATAAGTGAAATATCGTTCT
Outline Overview of the GEP annotation projects Annotation of Drosophila Primer January 2018 GEP annotation workflow Practice applying the GEP annotation strategy Wilson Leung and Chris Shaffer AAACAACAATCATAAATAGAGGAAGTTTTCGGAATATACGATAAGTGAAATATCGTTCT
Guided tour to Ensembl
 Guided tour to Ensembl Introduction Introduction to the Ensembl project Walk-through of the browser Variations and Functional Genomics Comparative Genomics BioMart Ensembl Genome browser http://www.ensembl.org
Guided tour to Ensembl Introduction Introduction to the Ensembl project Walk-through of the browser Variations and Functional Genomics Comparative Genomics BioMart Ensembl Genome browser http://www.ensembl.org
Supplementary Fig. 1 related to Fig. 1 Clinical relevance of lncrna candidate
 Supplementary Figure Legends Supplementary Fig. 1 related to Fig. 1 Clinical relevance of lncrna candidate BC041951 in gastric cancer. (A) The flow chart for selected candidate lncrnas in 660 up-regulated
Supplementary Figure Legends Supplementary Fig. 1 related to Fig. 1 Clinical relevance of lncrna candidate BC041951 in gastric cancer. (A) The flow chart for selected candidate lncrnas in 660 up-regulated
Transcription Eukaryotic Cells
 Transcription Eukaryotic Cells Packet #20 1 Introduction Transcription is the process in which genetic information, stored in a strand of DNA (gene), is copied into a strand of RNA. Protein-encoding genes
Transcription Eukaryotic Cells Packet #20 1 Introduction Transcription is the process in which genetic information, stored in a strand of DNA (gene), is copied into a strand of RNA. Protein-encoding genes
DNA Transcription. Dr Aliwaini
 DNA Transcription 1 DNA Transcription-Introduction The synthesis of an RNA molecule from DNA is called Transcription. All eukaryotic cells have five major classes of RNA: ribosomal RNA (rrna), messenger
DNA Transcription 1 DNA Transcription-Introduction The synthesis of an RNA molecule from DNA is called Transcription. All eukaryotic cells have five major classes of RNA: ribosomal RNA (rrna), messenger
Ab Initio SERVER PROTOTYPE FOR PREDICTION OF PHOSPHORYLATION SITES IN PROTEINS*
 COMPUTATIONAL METHODS IN SCIENCE AND TECHNOLOGY 9(1-2) 93-100 (2003/2004) Ab Initio SERVER PROTOTYPE FOR PREDICTION OF PHOSPHORYLATION SITES IN PROTEINS* DARIUSZ PLEWCZYNSKI AND LESZEK RYCHLEWSKI BiolnfoBank
COMPUTATIONAL METHODS IN SCIENCE AND TECHNOLOGY 9(1-2) 93-100 (2003/2004) Ab Initio SERVER PROTOTYPE FOR PREDICTION OF PHOSPHORYLATION SITES IN PROTEINS* DARIUSZ PLEWCZYNSKI AND LESZEK RYCHLEWSKI BiolnfoBank
Integrating Genomic Data to Predict Transcription Factor Binding
 Genome Informatics 16(1): 83 94 (2005) 83 Integrating Genomic Data to Predict Transcription Factor Binding Dustin T. Holloway 1 Mark Kon 2 Charles DeLisi 3 dth128@bu.edu mkon@bu.edu delisi@bu.edu 1 Molecular
Genome Informatics 16(1): 83 94 (2005) 83 Integrating Genomic Data to Predict Transcription Factor Binding Dustin T. Holloway 1 Mark Kon 2 Charles DeLisi 3 dth128@bu.edu mkon@bu.edu delisi@bu.edu 1 Molecular
Supporting Information
 Supporting Information Ho et al. 1.173/pnas.81288816 SI Methods Sequences of shrna hairpins: Brg shrna #1: ccggcggctcaagaaggaagttgaactcgagttcaacttccttcttgacgnttttg (TRCN71383; Open Biosystems). Brg shrna
Supporting Information Ho et al. 1.173/pnas.81288816 SI Methods Sequences of shrna hairpins: Brg shrna #1: ccggcggctcaagaaggaagttgaactcgagttcaacttccttcttgacgnttttg (TRCN71383; Open Biosystems). Brg shrna
Chapter 8 Lecture Outline. Transcription, Translation, and Bioinformatics
 Chapter 8 Lecture Outline Transcription, Translation, and Bioinformatics Replication, Transcription, Translation n Repetitive processes Build polymers of nucleotides or amino acids n All have 3 major steps
Chapter 8 Lecture Outline Transcription, Translation, and Bioinformatics Replication, Transcription, Translation n Repetitive processes Build polymers of nucleotides or amino acids n All have 3 major steps
Chromatographic Separation of the three forms of RNA Polymerase II.
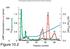 Chromatographic Separation of the three forms of RNA Polymerase II. α-amanitin α-amanitin bound to Pol II Function of the three enzymes. Yeast Pol II. RNA Polymerase Subunit Structures 10-7 Subunit structure.
Chromatographic Separation of the three forms of RNA Polymerase II. α-amanitin α-amanitin bound to Pol II Function of the three enzymes. Yeast Pol II. RNA Polymerase Subunit Structures 10-7 Subunit structure.
Protein Synthesis. OpenStax College
 OpenStax-CNX module: m46032 1 Protein Synthesis OpenStax College This work is produced by OpenStax-CNX and licensed under the Creative Commons Attribution License 3.0 By the end of this section, you will
OpenStax-CNX module: m46032 1 Protein Synthesis OpenStax College This work is produced by OpenStax-CNX and licensed under the Creative Commons Attribution License 3.0 By the end of this section, you will
Themes: RNA and RNA Processing. Messenger RNA (mrna) What is a gene? RNA is very versatile! RNA-RNA interactions are very important!
 Themes: RNA is very versatile! RNA and RNA Processing Chapter 14 RNA-RNA interactions are very important! Prokaryotes and Eukaryotes have many important differences. Messenger RNA (mrna) Carries genetic
Themes: RNA is very versatile! RNA and RNA Processing Chapter 14 RNA-RNA interactions are very important! Prokaryotes and Eukaryotes have many important differences. Messenger RNA (mrna) Carries genetic
Figure S1: NUN preparation yields nascent, unadenylated RNA with a different profile from Total RNA.
 Summary of Supplemental Information Figure S1: NUN preparation yields nascent, unadenylated RNA with a different profile from Total RNA. Figure S2: rrna removal procedure is effective for clearing out
Summary of Supplemental Information Figure S1: NUN preparation yields nascent, unadenylated RNA with a different profile from Total RNA. Figure S2: rrna removal procedure is effective for clearing out
Differential Gene Expression
 Biology 4361 Developmental Biology Differential Gene Expression September 28, 2006 Chromatin Structure ~140 bp ~60 bp Transcriptional Regulation: 1. Packing prevents access CH 3 2. Acetylation ( C O )
Biology 4361 Developmental Biology Differential Gene Expression September 28, 2006 Chromatin Structure ~140 bp ~60 bp Transcriptional Regulation: 1. Packing prevents access CH 3 2. Acetylation ( C O )
Whole Transcriptome Analysis of Illumina RNA- Seq Data. Ryan Peters Field Application Specialist
 Whole Transcriptome Analysis of Illumina RNA- Seq Data Ryan Peters Field Application Specialist Partek GS in your NGS Pipeline Your Start-to-Finish Solution for Analysis of Next Generation Sequencing Data
Whole Transcriptome Analysis of Illumina RNA- Seq Data Ryan Peters Field Application Specialist Partek GS in your NGS Pipeline Your Start-to-Finish Solution for Analysis of Next Generation Sequencing Data
Ana Teresa Freitas 2016/2017
 Finding Regulatory Motifs in DNA Sequences Ana Teresa Freitas 2016/2017 Combinatorial Gene Regulation A recent microarray experiment showed that when gene X is knocked out, 20 other genes are not expressed
Finding Regulatory Motifs in DNA Sequences Ana Teresa Freitas 2016/2017 Combinatorial Gene Regulation A recent microarray experiment showed that when gene X is knocked out, 20 other genes are not expressed
#28 - Promoter Prediction 10/29/07
 BCB 444/544 Required Reading (before lecture) Lecture 28 Mon Oct 29 - Lecture 28 Promoter & Regulatory Element Prediction Chp 9 - pp 113-126 Gene Prediction - finish it Wed Oct 30 - Lecture 29 Phylogenetics
BCB 444/544 Required Reading (before lecture) Lecture 28 Mon Oct 29 - Lecture 28 Promoter & Regulatory Element Prediction Chp 9 - pp 113-126 Gene Prediction - finish it Wed Oct 30 - Lecture 29 Phylogenetics
Structure/function relationship in DNA-binding proteins
 PHRM 836 September 22, 2015 Structure/function relationship in DNA-binding proteins Devlin Chapter 8.8-9 u General description of transcription factors (TFs) u Sequence-specific interactions between DNA
PHRM 836 September 22, 2015 Structure/function relationship in DNA-binding proteins Devlin Chapter 8.8-9 u General description of transcription factors (TFs) u Sequence-specific interactions between DNA
Lecture 11: Gene Prediction
 Lecture 11: Gene Prediction Study Chapter 6.11-6.14 1 Gene: A sequence of nucleotides coding for protein Gene Prediction Problem: Determine the beginning and end positions of genes in a genome Where are
Lecture 11: Gene Prediction Study Chapter 6.11-6.14 1 Gene: A sequence of nucleotides coding for protein Gene Prediction Problem: Determine the beginning and end positions of genes in a genome Where are
Chapter 3 Nucleic Acids, Proteins, and Enzymes
 3 Nucleic Acids, Proteins, and Enzymes Chapter 3 Nucleic Acids, Proteins, and Enzymes Key Concepts 3.1 Nucleic Acids Are Informational Macromolecules 3.2 Proteins Are Polymers with Important Structural
3 Nucleic Acids, Proteins, and Enzymes Chapter 3 Nucleic Acids, Proteins, and Enzymes Key Concepts 3.1 Nucleic Acids Are Informational Macromolecules 3.2 Proteins Are Polymers with Important Structural
Introduction to the UCSC genome browser
 Introduction to the UCSC genome browser Dominik Beck NHMRC Peter Doherty and CINSW ECR Fellow, Senior Lecturer Lowy Cancer Research Centre, UNSW and Centre for Health Technology, UTS SYDNEY NSW AUSTRALIA
Introduction to the UCSC genome browser Dominik Beck NHMRC Peter Doherty and CINSW ECR Fellow, Senior Lecturer Lowy Cancer Research Centre, UNSW and Centre for Health Technology, UTS SYDNEY NSW AUSTRALIA
Prokaryotic Transcription
 Prokaryotic Transcription Transcription Basics DNA is the genetic material Nucleic acid Capable of self-replication and synthesis of RNA RNA is the middle man Nucleic acid Structure and base sequence are
Prokaryotic Transcription Transcription Basics DNA is the genetic material Nucleic acid Capable of self-replication and synthesis of RNA RNA is the middle man Nucleic acid Structure and base sequence are
Neural Networks and Applications in Bioinformatics. Yuzhen Ye School of Informatics and Computing, Indiana University
 Neural Networks and Applications in Bioinformatics Yuzhen Ye School of Informatics and Computing, Indiana University Contents Biological problem: promoter modeling Basics of neural networks Perceptrons
Neural Networks and Applications in Bioinformatics Yuzhen Ye School of Informatics and Computing, Indiana University Contents Biological problem: promoter modeling Basics of neural networks Perceptrons
Chimp Chunk 3-14 Annotation by Matthew Kwong, Ruth Howe, and Hao Yang
 Chimp Chunk 3-14 Annotation by Matthew Kwong, Ruth Howe, and Hao Yang Ruth Howe Bio 434W April 1, 2010 INTRODUCTION De novo annotation is the process by which a finished genomic sequence is searched for
Chimp Chunk 3-14 Annotation by Matthew Kwong, Ruth Howe, and Hao Yang Ruth Howe Bio 434W April 1, 2010 INTRODUCTION De novo annotation is the process by which a finished genomic sequence is searched for
ALGORITHMS IN BIO INFORMATICS. Chapman & Hall/CRC Mathematical and Computational Biology Series A PRACTICAL INTRODUCTION. CRC Press WING-KIN SUNG
 Chapman & Hall/CRC Mathematical and Computational Biology Series ALGORITHMS IN BIO INFORMATICS A PRACTICAL INTRODUCTION WING-KIN SUNG CRC Press Taylor & Francis Group Boca Raton London New York CRC Press
Chapman & Hall/CRC Mathematical and Computational Biology Series ALGORITHMS IN BIO INFORMATICS A PRACTICAL INTRODUCTION WING-KIN SUNG CRC Press Taylor & Francis Group Boca Raton London New York CRC Press
Introduction to genome biology
 Introduction to genome biology Lisa Stubbs We ve found most genes; but what about the rest of the genome? Genome size* 12 Mb 95 Mb 170 Mb 1500 Mb 2700 Mb 3200 Mb #coding genes ~7000 ~20000 ~14000 ~26000
Introduction to genome biology Lisa Stubbs We ve found most genes; but what about the rest of the genome? Genome size* 12 Mb 95 Mb 170 Mb 1500 Mb 2700 Mb 3200 Mb #coding genes ~7000 ~20000 ~14000 ~26000
Sam68 STARR Sam68 QUA1- KH. p(r ) [Å] [Å] TSTAR STAR. Sam68 QUA1-KH and. constructs are
![Sam68 STARR Sam68 QUA1- KH. p(r ) [Å] [Å] TSTAR STAR. Sam68 QUA1-KH and. constructs are Sam68 STARR Sam68 QUA1- KH. p(r ) [Å] [Å] TSTAR STAR. Sam68 QUA1-KH and. constructs are](/thumbs/73/69166612.jpg) a b Sam68 STARR Sam68 QUA1- KH c d e ) p(r p(r ) r [Å] r [Å] Supplementary Figure 1: The QUA2 domain is not involved in i the overall conformation of the STAR domain (a) Overlay of T-STAR QUA1-KH in complex
a b Sam68 STARR Sam68 QUA1- KH c d e ) p(r p(r ) r [Å] r [Å] Supplementary Figure 1: The QUA2 domain is not involved in i the overall conformation of the STAR domain (a) Overlay of T-STAR QUA1-KH in complex
Figure S4 A-H : Initiation site properties and evolutionary changes
 A 0.3 Figure S4 A-H : Initiation site properties and evolutionary changes G-correction not used 0.25 Fraction of total counts 0.2 0.5 0. tag 2 tags 3 tags 4 tags 5 tags 6 tags 7tags 8tags 9 tags >9 tags
A 0.3 Figure S4 A-H : Initiation site properties and evolutionary changes G-correction not used 0.25 Fraction of total counts 0.2 0.5 0. tag 2 tags 3 tags 4 tags 5 tags 6 tags 7tags 8tags 9 tags >9 tags
REGULATION OF PROTEIN SYNTHESIS. II. Eukaryotes
 REGULATION OF PROTEIN SYNTHESIS II. Eukaryotes Complexities of eukaryotic gene expression! Several steps needed for synthesis of mrna! Separation in space of transcription and translation! Compartmentation
REGULATION OF PROTEIN SYNTHESIS II. Eukaryotes Complexities of eukaryotic gene expression! Several steps needed for synthesis of mrna! Separation in space of transcription and translation! Compartmentation
Predictive and Causal Modeling in the Health Sciences. Sisi Ma MS, MS, PhD. New York University, Center for Health Informatics and Bioinformatics
 Predictive and Causal Modeling in the Health Sciences Sisi Ma MS, MS, PhD. New York University, Center for Health Informatics and Bioinformatics 1 Exponentially Rapid Data Accumulation Protein Sequencing
Predictive and Causal Modeling in the Health Sciences Sisi Ma MS, MS, PhD. New York University, Center for Health Informatics and Bioinformatics 1 Exponentially Rapid Data Accumulation Protein Sequencing
Molecular Biology Primer. CptS 580, Computational Genomics, Spring 09
 Molecular Biology Primer pts 580, omputational enomics, Spring 09 Starting 19 th century What do we know of cellular biology? ell as a fundamental building block 1850s+: ``DNA was discovered by Friedrich
Molecular Biology Primer pts 580, omputational enomics, Spring 09 Starting 19 th century What do we know of cellular biology? ell as a fundamental building block 1850s+: ``DNA was discovered by Friedrich
Genome Annotation. What Does Annotation Describe??? Genome duplications Genes Mobile genetic elements Small repeats Genetic diversity
 Genome Annotation Genome Sequencing Costliest aspect of sequencing the genome o But Devoid of content Genome must be annotated o Annotation definition Analyzing the raw sequence of a genome and describing
Genome Annotation Genome Sequencing Costliest aspect of sequencing the genome o But Devoid of content Genome must be annotated o Annotation definition Analyzing the raw sequence of a genome and describing
Plant genome annotation using bioinformatics
 Plant genome annotation using bioinformatics ghorbani mandolakani Hossein, khodarahmi manouchehr darvish farrokh, taeb mohammad ghorbani24sma@yahoo.com islamic azad university of science and research branch
Plant genome annotation using bioinformatics ghorbani mandolakani Hossein, khodarahmi manouchehr darvish farrokh, taeb mohammad ghorbani24sma@yahoo.com islamic azad university of science and research branch
Data Mining in Bioinformatics. Prof. André de Carvalho ICMC-Universidade de São Paulo
 Data Mining in Bioinformatics Prof. André de Carvalho ICMC-Universidade de São Paulo Main topics Motivation Data Mining Prediction Bioinformatics Molecular Biology Using DM in Molecular Biology Case studies
Data Mining in Bioinformatics Prof. André de Carvalho ICMC-Universidade de São Paulo Main topics Motivation Data Mining Prediction Bioinformatics Molecular Biology Using DM in Molecular Biology Case studies
Click here to read the case study about protein synthesis.
 Click here to read the case study about protein synthesis. Big Question: How do cells use the genetic information stored in DNA to make millions of different proteins the body needs? Key Concept: Genetics
Click here to read the case study about protein synthesis. Big Question: How do cells use the genetic information stored in DNA to make millions of different proteins the body needs? Key Concept: Genetics
Conserved elements with potential to form polymorphic G-quadruplex structures in the first intron of human genes
 Published online 10 January 2008 Nucleic Acids Research, 2008, Vol. 36, No. 4 1321 1333 doi:10.1093/nar/gkm1138 Conserved elements with potential to form polymorphic G-quadruplex structures in the first
Published online 10 January 2008 Nucleic Acids Research, 2008, Vol. 36, No. 4 1321 1333 doi:10.1093/nar/gkm1138 Conserved elements with potential to form polymorphic G-quadruplex structures in the first
Gene Expression and Heritable Phenotype. CBS520 Eric Nabity
 Gene Expression and Heritable Phenotype CBS520 Eric Nabity DNA is Just the Beginning DNA was determined to be the genetic material, and the structure was identified as a (double stranded) double helix.
Gene Expression and Heritable Phenotype CBS520 Eric Nabity DNA is Just the Beginning DNA was determined to be the genetic material, and the structure was identified as a (double stranded) double helix.
MODULE 1: INTRODUCTION TO THE GENOME BROWSER: WHAT IS A GENE?
 MODULE 1: INTRODUCTION TO THE GENOME BROWSER: WHAT IS A GENE? Lesson Plan: Title Introduction to the Genome Browser: what is a gene? JOYCE STAMM Objectives Demonstrate basic skills in using the UCSC Genome
MODULE 1: INTRODUCTION TO THE GENOME BROWSER: WHAT IS A GENE? Lesson Plan: Title Introduction to the Genome Browser: what is a gene? JOYCE STAMM Objectives Demonstrate basic skills in using the UCSC Genome
