SIBC504: TRANSCRIPTION & RNA PROCESSING Assistant Professor Dr. Chatchawan Srisawat
|
|
|
- Blaise Burns
- 6 years ago
- Views:
Transcription
1 SIBC504: TRANSCRIPTION & RNA PROCESSING Assistant Professor Dr. Chatchawan Srisawat
2 TRANSCRIPTION: AN OVERVIEW Transcription: the synthesis of a single-stranded RNA from a doublestranded DNA template. The first stage in the overall process of gene expression. The most commonly-used step in regulating gene expression in organisms. Transcriptional control Switch gene on or off Phenotype
3 TRANSCRIPTION: AN OVERVIEW Important features of transcription RNA polymerase is the key enzyme: it requires DNA template, ribonucleotide precursors (ATP, GTP, CTP, and UTP) It catalyzes the synthesis of RNA from ribonucleotide precursors using one of the two DNA strands as a template. Coding or sense strand Template or antisense strand The sequence of the RNA transcript is the same as that of the coding strand (with U in place of T).
4 TRANSCRIPTION: AN OVERVIEW Important features of transcription Coding or sense strand Template or antisense strand Direction of synthesis occurs from the 5 - to the 3 -end of the RNA.
5 TRANSCRIPTION: AN OVERVIEW Important features of transcription Transcription process can be divided into three steps: 1. Initiation upstream downstream RNA polymerase transcription start site promoter A region of DNA involved in binding of RNA polymerase to initiate transcription; usually located upstream to the start site. Binding of RNA polymease to specific DNA sequences called promoters. Local unwinding of DNA and synthesizing the first few nucleotides of RNA (no primers required).
6 TRANSCRIPTION: AN OVERVIEW Important features of transcription Transcription process can be divided into three steps: 2. Elongation Successive addition of ribonucleotides into the RNA transcript from 5 to 3 direction using DNA as a template. DNA unwinding ahead of RNA polymerase and reannealing behind the enzyme.
7 TRANSCRIPTION: AN OVERVIEW Important features of transcription Transcription process can be divided into three steps: 3. Termination Synthesis of the RNA transcript is stopped at the terminator sequence. RNA polymerase dissociates from the DNA and releases the RNA transcript.
8 TRANSCRIPTION INITIATION Transcription initiation upstream downstream RNA polymerase promoter transcription start site The first step of transcription process Regulation of gene expression at transcriptional level are the most commonly used mode to determine which genes to be active or inactive. Most of regulatory mechanisms of gene expression occur at the transcriptional initiation step.
9 TRANSCRIPTION INITIATION IN PROKARYOTE In prokaryotes, RNA polymerase is a multisubunit enzyme. RNA polymerase holoenzyme Core enzyme σ (sigma) factor alpha (α) -2subunits/enzyme - required for core protein assembly; may be involved in promoter binding beta (β) - involved in catalysis; chain initiation and elongation. beta (β ) - involved in template DNA binding. omega (ω) - promotes assembly. Core enzyme can catalyze RNA synthesis and bind to DNA, but has no specificity; unable to start transcription at initiation sites alone.
10 TRANSCRIPTION INITIATION IN PROKARYOTE In prokaryotes, RNA polymerase is a multisubunit enzyme. RNA polymerase holoenzyme Core enzyme σ (sigma) factor Responsible for binding of DNA at promoters. Bring the core RNA polymerase enzyme to the initiation sites (promoters) to initiate transcription. Required for initiation and released from the core enzyme at the start of transcription elongation.
11 TRANSCRIPTION INITIATION IN PROKARYOTE σ factors and promoters Many bacteria, including E. coli, produce a set of different σ factors. σ 70, σ 32, σ 28, σ 38, σ 54 Number denotes the molecular weight of σ factor. σ 70 is the most common σ factor found in E. coli.
12 TRANSCRIPTION INITIATION IN PROKARYOTE σ factors and promoters upstream downstream promoter Promoter A region of DNA involved in binding of RNA polymerase to initiate transcription; usually located upstream to the start site. Different promoters contain conserved sequences, which can be specifically recognized and bound by different σ factors of RNA polymerase, thus facilitating assembly of the enzyme at the transcription initiation site.
13 TRANSCRIPTION INITIATION IN PROKARYOTE σ 70 factor and its promoter sequence The σ 70 promoter consists of a sequence of between bp (region from around -55 to +20), which is bound by RNA polymerase. consensus sequence of σ 70 promoter -35 sequence TTGACA Recognition region interacting with σ factor -10 sequence or Pribnow box TATAAT bp 5-8 bp CGT Important for DNA unwinding G A +1 Note - Different σ factors recognize specific promoters with different conserved sequences.
14 TRANSCRIPTION INITIATION IN PROKARYOTE σ 70 factor and its promoter sequence
15 TRANSCRIPTION INITIATION IN PROKARYOTE Steps in transcription initiation complex formation Promoter binding RNA polymerse core enzyme The RNA polymerase core enzyme has a nonspecific affinity for DNA, making it difficult and very slow to search and bind to the promoter by itself. With σ factor, the affinity for nonspecific sites on DNA is reduced by fold, while the affinity for promoters is enhanced by 100-fold. σ factor increases the specificity of the enzyme for promoter-binding sites.
16 TRANSCRIPTION INITIATION IN PROKARYOTE Steps in transcription initiation complex formation Promoter binding At the promoter, the RNA polymerase recognizes the promoter sequences at -35 and -10 regions (via the interaction with σ factor) Formation of the initial complex of the RNA polymerase and the basepaired DNA at the promoter: a closed complex.
17 TRANSCRIPTION INITIATION IN PROKARYOTE Steps in transcription initiation complex formation DNA unwinding Around 12 bp of the DNA (from -9 to +3) is unwound by the polymerase, forming an open comlex. -10 sequence region (AT-rich) is important for DNA unwinding
18 TRANSCRIPTION INITIATION IN PROKARYOTE Steps in transcription initiation complex formation RNA chain initiation -35 sequence TTGACA -10 sequence or Pribnow box TATAAT bp 5-8 bp CGT G A +1 RNA polymerase starts synthesizing a short RNA chain without the movement of the enzyme from the promoter. The synthesis does not require a primer. A short RNA chain of 9 nt in length is generated; the first base is usually started with G or A. The process is ineffective. It may abort and then restart until initiation succeeds.
19 TRANSCRIPTION INITIATION IN PROKARYOTE Steps in transcription initiation complex formation RNA chain initiation When initiation succeeds, the enzyme releases the σ factor (which is not required for elongation) and forms a ternary complex of polymerase-dna-rna transcript. Transcription elongation begins as the enzyme moves along the DNA to synthesize RNA transcript.
20 TRANSCRIPTION INITIATION IN PROKARYOTE Alternative sigma factors Sigma factors confer promoter specificity to the core RNA polymerase. The presence of alternate sigma factors provides the cell with a mechanism for turning on and off entire families of genes under different circumstances. Note- Deviation of a promoter sequence from the consensus may affect the promoter strength and efficiency of transcription
21 TRANSCRIPTION INITIATION IN EUKARYOTE In eukaryotes, there are many kinds of RNAs with different functions. There are also three different kinds of RNA polymerases (Pol I, II, and III) to transcribe these RNAs in eukaryotic cells.
22 TRANSCRIPTION INITIATION IN EUKARYOTE Eukaryotes have 3 types of RNA polymerases responsible for transcription of nuclear genes RNA polymerase I transcribes rrna RNA polymerase II - transcribes mrna (all protein-coding RNA) RNA polymerase III - transcribes trna, 5S RNA, snrna, 7SL RNA Distinguished by sensitivity to α-amanitin In addition, there are organelle-specific RNA polymerases (i.e., mitochondria and chloroplast) to transcribe genes in the organelles.
23 TRANSCRIPTION INITIATION IN EUKARYOTE Comparison of structures of eukaryotic and prokaryotic RNA polymerases Eukaryotic RNA polymerase (Pol I, II, and III) are multimeric enzymes and some of the subunits are conserved, showing similarity to the prokaryotic counterparts. Note- the carboxy terminus (CTD) of β like subunit of RNA pol II contains repeats of seven amino acids YSPTSPS - (up to 50 repeats), which can be phosphorylated during transcription.
24 TRANSCRIPTION INITIATION IN EUKARYOTE Typical RNA polymerase II core promoter Core promoter: the minimal DNA region at which RNA polymerase II can bind and initiate basal transcription of the gene. 1. TATA box - located approximately bp upstream of the start site - the conserved sequence: 5 TATA A A T A T 3 2. Initiator element (Inr) - usually located at start site - the conserved sequence: 5 Py 2 CAPy 5 3
25 TRANSCRIPTION INITIATION IN EUKARYOTE Transcription initiation of RNA polymerase II Like the core enzyme of RNA polymerase in prokaryotes, the RNA polymerases in eukaryotes also cannot bind to the promoter by itself. General or basal transcription factors are required for RNA pol II to initiate transcription at the promoter.
26 TRANSCRIPTION INITIATION IN EUKARYOTE Transcription initiation of RNA polymerase II General or basal transcription factors Not subunits of the RNA polymerase II. Required for RNA polymerase to bind avidly and specifically to promoters. General transcription factors for RNA polymerase II are called TFIIx, where x = A, B, D,... Can have multiple subunits.
27 TRANSCRIPTION INITIATION IN EUKARYOTE Sequential model of initiation 1. TFIID assembles on the TATA box - possibly stimulated by TFIIA TFIID - a multisubunit complex One subunit called TBP (TATAbinding protein) and other components called TAF ll s (TBP-associated factors) Only TBP binds to DNA (at TATA box) in sequence specific manner
28 TRANSCRIPTION INITIATION IN EUKARYOTE Sequential model of initiation 1. TFIID assembles on the TATA box - possibly stimulated by TFIIA TBP binds to TATA box and bends the DNA ~80 o, forcing open the minor groove. Helps unwind the DNA and may allow interaction with distant elements. rasmol
29 TRANSCRIPTION INITIATION IN EUKARYOTE Sequential model of initiation 1. TFIID assembles on the TATA box - possibly stimulated by TFIIA TFIIA enhances and stabilizes the binding of TFIID to the TATA box. Probably stops inhibitory factors binding to TFIID, which could otherwise block further assembly of the transcription complex.
30 TRANSCRIPTION INITIATION IN EUKARYOTE Sequential model of initiation 2. Binding of TFIIB to the preinitiation complex. TFIIB binds to TFIID and acts as a bridge for RNA polymerase binding.
31 TRANSCRIPTION INITIATION IN EUKARYOTE Sequential model of initiation 3. RNA pol II is escorted to the preinitiation complex along with TFIIF. RNA polymerase II is in a complex with TFIIF. TFIIF delivers the RNA polymerase to the TFIID- TFIIA-TFIIB complex (via interactions with TFIIB). Could be involved in melting DNA at initiation.
32 TRANSCRIPTION INITIATION IN EUKARYOTE Sequential model of initiation 4. TFIIE, TFIIJ, and TFIIH are then recruited to the preinitiation complex. TFIIE acts as docking site for TFIIH and can stimulate transcription. TFIIJ functions are stil unknown.
33 TRANSCRIPTION INITIATION IN EUKARYOTE Sequential model of initiation 4. TFIIE, TFIIJ, and TFIIH are then recruited to the preinitiation complex. TFIIH a complex protein with 9 subunits Has DNA helicase/atpase activity for melting DNA during transcription initiation. Also has kinase activity: phosphorylates the carboxyl terminal domain (CTD) of the large β like subunit of RNA polymerase II
34 TRANSCRIPTION INITIATION IN EUKARYOTE Sequential model of initiation 5. Phosphorylation of C-terminal domain of RNA pol II by TFIIH. TFIIH phosphorylates the CTD of of the large β like subunit of RNA polymerase II. formation of a processive RNA polymerase complex. allows the RNA polymerase to leave the promoter region to begin transcription elongation. also serves as Landing pad for macromolecular assemblies that regulate mrna synthesis and processing, e.g., transcriptional elongation factors, RNA modifications enzymes.
35 TRANSCRIPTION INITIATION - SUMMARY Promoter binding of RNA polymerases prokaryote eukaryote Pol III RNA polymerase I promoters RNA polymerase II promoters RNA polymerase III promoters
36 TRANSCRIPTION ELONGATION RNA polymerase moves along DNA, unwinding it, exposing DNA bases to be used as a template -> transcription bubble Adds to the 3 end (grows from 5 to 3 direction of RNA transcript) As it passes, DNA helix rewinds and new RNA peels away. Add 60 nucleotides/sec in eukaryotes, 40 nucleotides/sec in prokaryotes.
37 TRANSCRIPTION ELONGATION In eukaryotes, TFIIF remains associated with RNA polymerase II through out transcription elongation. During elongation, the activity of RNA polymerase II are greatly enhanced by proteins called elongation factors: eg., ELL, p- TEFb, SII(TFIIS), Elongin (SIII). The elongation factors suppress pausing or arrest of transcription and also coordinate interactions between protein complexes involved in the posttranscriptional processing of mrnas. Once the RNA transcript is completed, transcription is terminated. Pol II is dephosphorylated and recycled, ready to initiate another transcription.
38 Termination in prokaryotes TRANSCRIPTION TERMINATION There are two ways to terminate transcription in E.coli ρ (rho)-dependent termination ρ (rho)-independent termination
39 Termination in prokaryotes TRANSCRIPTION TERMINATION ρ (rho)-dependent termination Some genes contain terminator sequences which require an additional protein factor, ρ, for efficient termination. ρ binds to specific sites in singlestranded RNA. Then it hydrolyzes ATP and moves along the RNA towards the transcription complex, enabling the polymerase to terminate transcription.
40 Termination in prokaryotes TRANSCRIPTION TERMINATION ρ (rho)-independent termination Elongation of RNA transcript continues until the RNA polymerase reaches the terminator sequence. The most common terminator is an RNA hairpin in which the RNA transcript is self-complementary.
41 Termination in prokaryotes TRANSCRIPTION TERMINATION ρ (rho)-independent termination RNA polymerase pauses immediately after it has synthesized the hairpin RNA. The subsequent stretch of U s basepairs only weakly with da s in the DNA template strand. Release of the RNA transcript. The two DNA strands reanneal. Dissociation of the core enzyme from the DNA.
42 Termination in eukaryotes TRANSCRIPTION TERMINATION Termination of RNA polymerase I and III transcription Both RNA polymerases, I and III, have discrete termination sites (like prokaryotic RNA polymerase). RNA polymerase I terminates at a discrete site, involving a recognition region of an 18-base terminator sequence by an ancillary factor. RNA polymerase III terminates at the sequence with a run of 4 U bases without the need of any accessory factors.
43 Termination in eukaryotes TRANSCRIPTION TERMINATION Termination of RNA polymerase II RNA polymerase II elongates past the cleavage site at AAUAAA (polyadenylation or cleavage signal) and ceases RNA synthesis within multiple termination sites located in a rather long terminator region. Sometimes >1000 bp downstream of the 3 end The nature of the individual termination sites and sequences is not known because the mature 3 end of the mrna is generated by cleavage at a specific sequence, not by the termination event.
44 RNA PROCESSING Messenger RNA processing in prokaryotes no significant modification Messenger RNA processing in eukaryotes 5 -capping Splicing 3 -cleavage and polyadenylation
45 RNA PROCESSING 5 -CAPPING Functions: Marks the 5 -end of the first exon and aids in the splicing process Essential for nucleo-cytoplasmic transport of mrnas through interaction with nuclear cap-binding proteins Increases the efficiency of translation by targeting formation of the preinitiation complex (cytoplasmic cap-binding proteins) protects the transcript from 5 3 exoribonucleolytic activities
46 5 -CAPPING RNA PROCESSING
47 RNA PROCESSING 3 -CLEAVAGE AND POLYADENYLATION An enzyme complex recognizes the polyadenylation or cleavage signal (AAUAAA) and a less well conserved G-U rich sequence located nucleotides downstream. An endonuclease cleaves the primary transcript nucleotides downstream of the AAUAAA signal. A series of A residues are added to the 3 -end of the cleaved transcript by polyadenylate polymerase.
48 RNA PROCESSING 3 -CLEAVAGE AND POLYADENYLATION Functions: Increase translational efficiency Protection of mrna from degradation
49 RNA PROCESSING SPLICING Two steps of transesterifications The 2 -OH from an adenine in intron attack 5 splicing site. 3 -OH at the 5 exon attach the 3 splicing site, joining the two exons together. Intron is excised as a lariat.
50 RNA PROCESSING SPLICING The chemistry is simple, but the in vivo process in not. Need to accurately define where exons end and introns begin. Catalysis of splicing reactions. SPLICEOSOME A large RNA/protein complex consists of specialized RNA-protein complexes, small nuclear ribonucleoproteins (snrnps). U1, U2, U4, U5, and U6 snrnp Each snrnp contains one of a class of eukaryotic RNAs, 100 to 200 nucleotides long, known as small nuclear RNAs (snrnas).
51 RNA PROCESSING SPLICING Recognition of exon/intron boundaries Exon - Ex pressed Sequences Intron - In tervening Sequences Polypyrimidine tract (U or C) consensus sequence for 5 splice site (donor site) consensus sequence for 3 splice site (acceptor site) Most introns start from the sequence GU and end with the sequence AG (in the 5' to 3' direction); the splice donor (5 splice site) and splice acceptor (3 splice site), respectively. In over 60% of cases, the exon sequence is (A/C)AG at the donor site, and G at the acceptor site. Another important sequence is called the branch site located bases upstream of the acceptor site; "CU(A/G)A(C/U)", where A is conserved in all genes.
52 RNA PROCESSING SPLICING Recognition of exon/intron boundaries Exon - Ex pressed Sequences Intron - In tervening Sequences Polypyrimidine tract (U or C) consensus sequence for 5 splice site (donor site) consensus sequence for 3 splice site (acceptor site)
53 RNA PROCESSING SPLICING Splicing mechanism in mrna primary transcripts
54 RNA PROCESSING SPLICING Splicing mechanism in mrna primary transcripts
55 RNA PROCESSING
56 Cotranscriptional pre -mrna processing RNA PROCESSING
57 RNA PROCESSING
30 Gene expression: Transcription
 30 Gene expression: Transcription Gene structure. o Exons coding region of DNA. o Introns non-coding region of DNA. o Introns are interspersed between exons of a single gene. o Promoter region helps enzymes
30 Gene expression: Transcription Gene structure. o Exons coding region of DNA. o Introns non-coding region of DNA. o Introns are interspersed between exons of a single gene. o Promoter region helps enzymes
GENETICS - CLUTCH CH.10 TRANSCRIPTION.
 !! www.clutchprep.com CONCEPT: OVERVIEW OF TRANSCRIPTION Transcription is the process of using DNA as a template to RNA RNA polymerase is the enzyme that transcribes DNA - There are many different types
!! www.clutchprep.com CONCEPT: OVERVIEW OF TRANSCRIPTION Transcription is the process of using DNA as a template to RNA RNA polymerase is the enzyme that transcribes DNA - There are many different types
DNA Transcription. Dr Aliwaini
 DNA Transcription 1 DNA Transcription-Introduction The synthesis of an RNA molecule from DNA is called Transcription. All eukaryotic cells have five major classes of RNA: ribosomal RNA (rrna), messenger
DNA Transcription 1 DNA Transcription-Introduction The synthesis of an RNA molecule from DNA is called Transcription. All eukaryotic cells have five major classes of RNA: ribosomal RNA (rrna), messenger
Biochemistry Eukaryotic Transcription
 1 Description of Module Subject Name Paper Name Module Name/Title Dr. Vijaya Khader Dr. MC Varadaraj 2 1. Objectives 1. Understand and have an overview of eucaryotic transcriptional regulation. 2. Explain
1 Description of Module Subject Name Paper Name Module Name/Title Dr. Vijaya Khader Dr. MC Varadaraj 2 1. Objectives 1. Understand and have an overview of eucaryotic transcriptional regulation. 2. Explain
Transcription. By : Lucia Dhiantika Witasari M.Biotech., Apt
 Transcription By : Lucia Dhiantika Witasari M.Biotech., Apt REGULATION OF GENE EXPRESSION 11/26/2010 2 RNA Messenger RNAs (mrnas) encode the amino acid sequence of one or more polypeptides specified by
Transcription By : Lucia Dhiantika Witasari M.Biotech., Apt REGULATION OF GENE EXPRESSION 11/26/2010 2 RNA Messenger RNAs (mrnas) encode the amino acid sequence of one or more polypeptides specified by
TRANSCRIPTION AND PROCESSING OF RNA
 TRANSCRIPTION AND PROCESSING OF RNA 1. The steps of gene expression. 2. General characterization of transcription: steps, components of transcription apparatus. 3. Transcription of eukaryotic structural
TRANSCRIPTION AND PROCESSING OF RNA 1. The steps of gene expression. 2. General characterization of transcription: steps, components of transcription apparatus. 3. Transcription of eukaryotic structural
Transcription. Manzur Ali PP, DBT,M.E.S College,Marampally
 Transcription Manzur Ali PP, DBT,M.E.S College,Marampally manzursir@gmail.com RNA transcription is actively regulated Not all DNA is transcribed in a given cell (less than 50% even in prokaryotes) For
Transcription Manzur Ali PP, DBT,M.E.S College,Marampally manzursir@gmail.com RNA transcription is actively regulated Not all DNA is transcribed in a given cell (less than 50% even in prokaryotes) For
Eukaryotic & Prokaryotic Transcription. RNA polymerases
 Eukaryotic & Prokaryotic Transcription RNA polymerases RNA Polymerases A. E. coli RNA polymerase 1. core enzyme = ββ'(α)2 has catalytic activity but cannot recognize start site of transcription ~500,000
Eukaryotic & Prokaryotic Transcription RNA polymerases RNA Polymerases A. E. coli RNA polymerase 1. core enzyme = ββ'(α)2 has catalytic activity but cannot recognize start site of transcription ~500,000
Mechanisms of Transcription. School of Life Science Shandong University
 Mechanisms of Transcription School of Life Science Shandong University Ch 12: Mechanisms of Transcription 1. RNA polymerase and the transcription cycle 2. The transcription cycle in bacteria 3. Transcription
Mechanisms of Transcription School of Life Science Shandong University Ch 12: Mechanisms of Transcription 1. RNA polymerase and the transcription cycle 2. The transcription cycle in bacteria 3. Transcription
Transcription in Eukaryotes
 Transcription in Eukaryotes Biology I Hayder A Giha Transcription Transcription is a DNA-directed synthesis of RNA, which is the first step in gene expression. Gene expression, is transformation of the
Transcription in Eukaryotes Biology I Hayder A Giha Transcription Transcription is a DNA-directed synthesis of RNA, which is the first step in gene expression. Gene expression, is transformation of the
Chapter 11. Transcription. The biochemistry and molecular biology department of CMU
 Chapter 11 Transcription The biochemistry and molecular biology department of CMU Transcription The synthesis of RNA molecules using DNA strands as the templates so that the genetic information can be
Chapter 11 Transcription The biochemistry and molecular biology department of CMU Transcription The synthesis of RNA molecules using DNA strands as the templates so that the genetic information can be
Lecture Summary: Regulation of transcription. General mechanisms-what are the major regulatory points?
 BCH 401G Lecture 37 Andres Lecture Summary: Regulation of transcription. General mechanisms-what are the major regulatory points? RNA processing: Capping, polyadenylation, splicing. Why process mammalian
BCH 401G Lecture 37 Andres Lecture Summary: Regulation of transcription. General mechanisms-what are the major regulatory points? RNA processing: Capping, polyadenylation, splicing. Why process mammalian
Transcription & post transcriptional modification
 Transcription & post transcriptional modification Transcription The synthesis of RNA molecules using DNA strands as the templates so that the genetic information can be transferred from DNA to RNA Similarity
Transcription & post transcriptional modification Transcription The synthesis of RNA molecules using DNA strands as the templates so that the genetic information can be transferred from DNA to RNA Similarity
Chapter 3. DNA, RNA, and Protein Synthesis
 Chapter 3. DNA, RNA, and Protein Synthesis 4. Transcription Gene Expression Regulatory region (promoter) 5 flanking region Upstream region Coding region 3 flanking region Downstream region Transcription
Chapter 3. DNA, RNA, and Protein Synthesis 4. Transcription Gene Expression Regulatory region (promoter) 5 flanking region Upstream region Coding region 3 flanking region Downstream region Transcription
Transcription Eukaryotic Cells
 Transcription Eukaryotic Cells Packet #20 1 Introduction Transcription is the process in which genetic information, stored in a strand of DNA (gene), is copied into a strand of RNA. Protein-encoding genes
Transcription Eukaryotic Cells Packet #20 1 Introduction Transcription is the process in which genetic information, stored in a strand of DNA (gene), is copied into a strand of RNA. Protein-encoding genes
Transcription steps. Transcription steps. Eukaryote RNA processing
 Transcription steps Initiation at 5 end of gene binding of RNA polymerase to promoter unwinding of DNA Elongation addition of nucleotides to 3 end rules of base pairing requires Mg 2+ energy from NTP substrates
Transcription steps Initiation at 5 end of gene binding of RNA polymerase to promoter unwinding of DNA Elongation addition of nucleotides to 3 end rules of base pairing requires Mg 2+ energy from NTP substrates
RNA Expression of the information in a gene generally involves production of an RNA molecule transcribed from a DNA template. RNA differs from DNA
 RNA Expression of the information in a gene generally involves production of an RNA molecule transcribed from a DNA template. RNA differs from DNA that it has a hydroxyl group at the 2 position of the
RNA Expression of the information in a gene generally involves production of an RNA molecule transcribed from a DNA template. RNA differs from DNA that it has a hydroxyl group at the 2 position of the
Lecture 11. Initiation of RNA Pol II transcription. Transcription Initiation Complex
 Lecture 11 *Eukaryotic Transcription Gene Organization RNA Processing 5 cap 3 polyadenylation splicing Translation Initiation of RNA Pol II transcription Consensus sequence of promoter TATA Transcription
Lecture 11 *Eukaryotic Transcription Gene Organization RNA Processing 5 cap 3 polyadenylation splicing Translation Initiation of RNA Pol II transcription Consensus sequence of promoter TATA Transcription
M1 - Biochemistry. Nucleic Acid Structure II/Transcription I
 M1 - Biochemistry Nucleic Acid Structure II/Transcription I PH Ratz, PhD (Resources: Lehninger et al., 5th ed., Chapters 8, 24 & 26) 1 Nucleic Acid Structure II/Transcription I Learning Objectives: 1.
M1 - Biochemistry Nucleic Acid Structure II/Transcription I PH Ratz, PhD (Resources: Lehninger et al., 5th ed., Chapters 8, 24 & 26) 1 Nucleic Acid Structure II/Transcription I Learning Objectives: 1.
Expression of the genome. Books: 1. Molecular biology of the gene: Watson et al 2. Genetics: Peter J. Russell
 Expression of the genome Books: 1. Molecular biology of the gene: Watson et al 2. Genetics: Peter J. Russell 1 Transcription 1. Francis Crick (1956) named the flow of information from DNA RNA protein the
Expression of the genome Books: 1. Molecular biology of the gene: Watson et al 2. Genetics: Peter J. Russell 1 Transcription 1. Francis Crick (1956) named the flow of information from DNA RNA protein the
Transcription is the first stage of gene expression
 Transcription is the first stage of gene expression RNA synthesis is catalyzed by RNA polymerase, which pries the DNA strands apart and hooks together the RNA nucleotides The RNA is complementary to the
Transcription is the first stage of gene expression RNA synthesis is catalyzed by RNA polymerase, which pries the DNA strands apart and hooks together the RNA nucleotides The RNA is complementary to the
We can now identify three major pathways of information flow in the cell (in replication, information passes from one DNA molecule to other DNA
 1 We can now identify three major pathways of information flow in the cell (in replication, information passes from one DNA molecule to other DNA molecules; in transcription, information passes from DNA
1 We can now identify three major pathways of information flow in the cell (in replication, information passes from one DNA molecule to other DNA molecules; in transcription, information passes from DNA
Computational Biology I LSM5191 (2003/4)
 Computational Biology I LSM5191 (2003/4) Aylwin Ng, D.Phil Lecture Notes: Transcriptome: Molecular Biology of Gene Expression I Flow of information: DNA to polypeptide DNA Start Exon1 Intron Exon2 Termination
Computational Biology I LSM5191 (2003/4) Aylwin Ng, D.Phil Lecture Notes: Transcriptome: Molecular Biology of Gene Expression I Flow of information: DNA to polypeptide DNA Start Exon1 Intron Exon2 Termination
Transcription. The sugar molecule found in RNA is ribose, rather than the deoxyribose found in DNA.
 Transcription RNA (ribonucleic acid) is a key intermediary between a DNA sequence and a polypeptide. RNA is an informational polynucleotide similar to DNA, but it differs from DNA in three ways: RNA generally
Transcription RNA (ribonucleic acid) is a key intermediary between a DNA sequence and a polypeptide. RNA is an informational polynucleotide similar to DNA, but it differs from DNA in three ways: RNA generally
The discovery of the role of RNA RNA structure, synthesis and function
 Central Dogma The discovery of the role of RNA RNA structure, synthesis and function! Fundamental observations in genetics!! Genes are located in nuclei (in eukaryotes)!! Polypeptides are synthesised in
Central Dogma The discovery of the role of RNA RNA structure, synthesis and function! Fundamental observations in genetics!! Genes are located in nuclei (in eukaryotes)!! Polypeptides are synthesised in
Chromatographic Separation of the three forms of RNA Polymerase II.
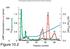 Chromatographic Separation of the three forms of RNA Polymerase II. α-amanitin α-amanitin bound to Pol II Function of the three enzymes. Yeast Pol II. RNA Polymerase Subunit Structures 10-7 Subunit structure.
Chromatographic Separation of the three forms of RNA Polymerase II. α-amanitin α-amanitin bound to Pol II Function of the three enzymes. Yeast Pol II. RNA Polymerase Subunit Structures 10-7 Subunit structure.
RNA synthesis/transcription I Biochemistry 302. February 6, 2004 Bob Kelm
 RNA synthesis/transcription I Biochemistry 302 February 6, 2004 Bob Kelm Overview of RNA classes Messenger RNA (mrna) Encodes protein Relatively short half-life ( 3 min in E. coli, 30 min in eukaryotic
RNA synthesis/transcription I Biochemistry 302 February 6, 2004 Bob Kelm Overview of RNA classes Messenger RNA (mrna) Encodes protein Relatively short half-life ( 3 min in E. coli, 30 min in eukaryotic
RNA metabolism. DNA dependent synthesis of RNA RNA processing RNA dependent synthesis of RNA and DNA.
 RNA metabolism DNA dependent synthesis of RNA RNA processing RNA dependent synthesis of RNA and DNA http://www.youtube.com/watch?v=ovc8nxobxmq DNA dependent synthesis of RNA : production of an RNA molecule
RNA metabolism DNA dependent synthesis of RNA RNA processing RNA dependent synthesis of RNA and DNA http://www.youtube.com/watch?v=ovc8nxobxmq DNA dependent synthesis of RNA : production of an RNA molecule
Gene Expression: Transcription, Translation, RNAs and the Genetic Code
 Lecture 28-29 Gene Expression: Transcription, Translation, RNAs and the Genetic Code Central dogma of molecular biology During transcription, the information in a DNA sequence (a gene) is copied into a
Lecture 28-29 Gene Expression: Transcription, Translation, RNAs and the Genetic Code Central dogma of molecular biology During transcription, the information in a DNA sequence (a gene) is copied into a
Transcription & RNA Processing
 Chapter 10. Transcription & RNA Processing 1. Transfer of Genetic Information: the Central Dogma 2. The Process of Gene Expression 3. Transcription & RNA Processing in Eukaryotes 4. Interrupted Genes in
Chapter 10. Transcription & RNA Processing 1. Transfer of Genetic Information: the Central Dogma 2. The Process of Gene Expression 3. Transcription & RNA Processing in Eukaryotes 4. Interrupted Genes in
Transcription in Prokaryotes. Jörg Bungert, PhD Phone:
 Transcription in Prokaryotes Jörg Bungert, PhD Phone: 352-273-8098 Email: jbungert@ufl.edu Objectives Understand the basic mechanism of transcription. Know the function of promoter elements and associating
Transcription in Prokaryotes Jörg Bungert, PhD Phone: 352-273-8098 Email: jbungert@ufl.edu Objectives Understand the basic mechanism of transcription. Know the function of promoter elements and associating
Make the protein through the genetic dogma process.
 Make the protein through the genetic dogma process. Coding Strand 5 AGCAATCATGGATTGGGTACATTTGTAACTGT 3 Template Strand mrna Protein Complete the table. DNA strand DNA s strand G mrna A C U G T A T Amino
Make the protein through the genetic dogma process. Coding Strand 5 AGCAATCATGGATTGGGTACATTTGTAACTGT 3 Template Strand mrna Protein Complete the table. DNA strand DNA s strand G mrna A C U G T A T Amino
Eukaryotic Transcription
 Eukaryotic Transcription I. Differences between eukaryotic versus prokaryotic transcription. II. (core vs holoenzyme): RNA polymerase II - Promotor elements. - General Pol II transcription factors (GTF).
Eukaryotic Transcription I. Differences between eukaryotic versus prokaryotic transcription. II. (core vs holoenzyme): RNA polymerase II - Promotor elements. - General Pol II transcription factors (GTF).
Basi s c i Fea e tu t re r s s of f R NA N Sy S nth t esi s s i s
 Transcription Dr.H.B.Mahesha, Yuvaraja s College, University of Mysore, Mysuru. It is the process of transcribing or making a copy of Genetic information stored in a DNA strand into a Complementary strand
Transcription Dr.H.B.Mahesha, Yuvaraja s College, University of Mysore, Mysuru. It is the process of transcribing or making a copy of Genetic information stored in a DNA strand into a Complementary strand
Initiation and termination of transcription, Post transcription modification of the RNA. Mitesh Shrestha
 Initiation and termination of transcription, Post transcription modification of the RNA Mitesh Shrestha Transcription: overview In prokaryotes transcription and translation are coupled. Proteins are synthesized
Initiation and termination of transcription, Post transcription modification of the RNA Mitesh Shrestha Transcription: overview In prokaryotes transcription and translation are coupled. Proteins are synthesized
The Flow of Genetic Information
 Chapter 17 The Flow of Genetic Information The DNA inherited by an organism leads to specific traits by dictating the synthesis of proteins and of RNA molecules involved in protein synthesis. Proteins
Chapter 17 The Flow of Genetic Information The DNA inherited by an organism leads to specific traits by dictating the synthesis of proteins and of RNA molecules involved in protein synthesis. Proteins
BIOCHEMISTRY REVIEW. Overview of Biomolecules. Chapter 12 Transcription
 BIOCHEMISTRY REVIEW Overview of Biomolecules Chapter 12 Transcription 2 3 4 5 Are You Getting It?? Which are general characteristics of transcription? (multiple answers) a) An entire DNA molecule is transcribed
BIOCHEMISTRY REVIEW Overview of Biomolecules Chapter 12 Transcription 2 3 4 5 Are You Getting It?? Which are general characteristics of transcription? (multiple answers) a) An entire DNA molecule is transcribed
DNA Prokaryote Transcription Steps (updated February 2013)
 URS AACTGT ATATTA - 35-10 transcription Pribnow Box discriminator +1 AGGAGGT TTA TCCTCCA ATT Gene C TGA TAG ACT ATC rho or GC hairpin loop transcription termination DNA Prokaryote Transcription Steps (updated
URS AACTGT ATATTA - 35-10 transcription Pribnow Box discriminator +1 AGGAGGT TTA TCCTCCA ATT Gene C TGA TAG ACT ATC rho or GC hairpin loop transcription termination DNA Prokaryote Transcription Steps (updated
BIO 311C Spring Lecture 36 Wednesday 28 Apr.
 BIO 311C Spring 2010 1 Lecture 36 Wednesday 28 Apr. Synthesis of a Polypeptide Chain 5 direction of ribosome movement along the mrna 3 ribosome mrna NH 2 polypeptide chain direction of mrna movement through
BIO 311C Spring 2010 1 Lecture 36 Wednesday 28 Apr. Synthesis of a Polypeptide Chain 5 direction of ribosome movement along the mrna 3 ribosome mrna NH 2 polypeptide chain direction of mrna movement through
Biochemistry 111. Carl Parker x A Braun
 Biochemistry 111 Carl Parker x6368 101A Braun csp@caltech.edu Central Dogma of Molecular Biology DNA-Dependent RNA Polymerase Requires a DNA Template Synthesizes RNA in a 5 to 3 direction Requires ribonucleoside
Biochemistry 111 Carl Parker x6368 101A Braun csp@caltech.edu Central Dogma of Molecular Biology DNA-Dependent RNA Polymerase Requires a DNA Template Synthesizes RNA in a 5 to 3 direction Requires ribonucleoside
RNA : functional role
 RNA : functional role Hamad Yaseen, PhD MLS Department, FAHS Hamad.ali@hsc.edu.kw RNA mrna rrna trna 1 From DNA to Protein -Outline- From DNA to RNA From RNA to Protein From DNA to RNA Transcription: Copying
RNA : functional role Hamad Yaseen, PhD MLS Department, FAHS Hamad.ali@hsc.edu.kw RNA mrna rrna trna 1 From DNA to Protein -Outline- From DNA to RNA From RNA to Protein From DNA to RNA Transcription: Copying
RNA Metabolism Chap 26, part I
 RNA Metabolism Chap 26, part I mrna (selective and regulated) trna rrna other (specialized) RNAs (eukaryotes!!!) processing transcriptome (Surprisingly, much of your genome is transcribed!) RNA is the
RNA Metabolism Chap 26, part I mrna (selective and regulated) trna rrna other (specialized) RNAs (eukaryotes!!!) processing transcriptome (Surprisingly, much of your genome is transcribed!) RNA is the
Chapters 31-32: Ribonucleic Acid (RNA)
 Chapters 31-32: Ribonucleic Acid (RNA) Short segments from the transcription, processing and translation sections of each chapter Slide 1 RNA In comparison with DNA RNA utilizes uracil in place of thymine
Chapters 31-32: Ribonucleic Acid (RNA) Short segments from the transcription, processing and translation sections of each chapter Slide 1 RNA In comparison with DNA RNA utilizes uracil in place of thymine
Feedback D. Incorrect! No, although this is a correct characteristic of RNA, this is not the best response to the questions.
 Biochemistry - Problem Drill 23: RNA No. 1 of 10 1. Which of the following statements best describes the structural highlights of RNA? (A) RNA can be single or double stranded. (B) G-C pairs have 3 hydrogen
Biochemistry - Problem Drill 23: RNA No. 1 of 10 1. Which of the following statements best describes the structural highlights of RNA? (A) RNA can be single or double stranded. (B) G-C pairs have 3 hydrogen
Chapter 17. From Gene to Protein
 Chapter 17 From Gene to Protein One Gene One Enzyme Hypothesis Archibald Garrod 1 st to suggest that genes dictate phenotypes through enzymes that catalyze specific chemical reactions ; alkaptonuria Beadle
Chapter 17 From Gene to Protein One Gene One Enzyme Hypothesis Archibald Garrod 1 st to suggest that genes dictate phenotypes through enzymes that catalyze specific chemical reactions ; alkaptonuria Beadle
FROM GENE TO PROTEIN. One Gene One Enzyme Hypothesis 3/12/2013. Basic Principles of Transcription & Translation
 One Gene One Enzyme Hypothesis FROM GENE TO PROTEIN C H A P T E R 1 7 Archibald Garrod 1 st to suggest that genes dictate phenotypes through enzymes that catalyze specific chemical reactions ; alkaptonuria
One Gene One Enzyme Hypothesis FROM GENE TO PROTEIN C H A P T E R 1 7 Archibald Garrod 1 st to suggest that genes dictate phenotypes through enzymes that catalyze specific chemical reactions ; alkaptonuria
Transcription and Post Transcript Modification
 Transcription and Post Transcript Modification You Should Be Able To 1. Describe transcription. 2. Compare and contrast eukaryotic + prokaryotic transcription. 3. Explain mrna processing in eukaryotes.
Transcription and Post Transcript Modification You Should Be Able To 1. Describe transcription. 2. Compare and contrast eukaryotic + prokaryotic transcription. 3. Explain mrna processing in eukaryotes.
Chapter 8 Lecture Outline. Transcription, Translation, and Bioinformatics
 Chapter 8 Lecture Outline Transcription, Translation, and Bioinformatics Replication, Transcription, Translation n Repetitive processes Build polymers of nucleotides or amino acids n All have 3 major steps
Chapter 8 Lecture Outline Transcription, Translation, and Bioinformatics Replication, Transcription, Translation n Repetitive processes Build polymers of nucleotides or amino acids n All have 3 major steps
Proofreading, post-replication modification of DNA. Mitesh Shrestha
 Proofreading, post-replication modification of DNA Mitesh Shrestha Proofreading During DNA replication (copying), most DNA polymerases can check their work with each base that they add. This process is
Proofreading, post-replication modification of DNA Mitesh Shrestha Proofreading During DNA replication (copying), most DNA polymerases can check their work with each base that they add. This process is
Molecular Genetics Principles of Gene Expression: Transcription
 Paper No. : 16 Module : 12 Principles of gene expression: Transcription Development Team Principal Investigator: Prof. Neeta Sehgal Head, Department of Zoology, University of Delhi Paper Coordinator: Prof.
Paper No. : 16 Module : 12 Principles of gene expression: Transcription Development Team Principal Investigator: Prof. Neeta Sehgal Head, Department of Zoology, University of Delhi Paper Coordinator: Prof.
CLASS 3.5: 03/29/07 EUKARYOTIC TRANSCRIPTION I: PROMOTERS AND ENHANCERS
 CLASS 3.5: 03/29/07 EUKARYOTIC TRANSCRIPTION I: PROMOTERS AND ENHANCERS A. Promoters and Polymerases (RNA pols): 1. General characteristics - Initiation of transcription requires a. Transcription factors
CLASS 3.5: 03/29/07 EUKARYOTIC TRANSCRIPTION I: PROMOTERS AND ENHANCERS A. Promoters and Polymerases (RNA pols): 1. General characteristics - Initiation of transcription requires a. Transcription factors
TRANSCRIPTION COMPARISON OF DNA & RNA TRANSCRIPTION. Umm AL Qura University. Sugar Ribose Deoxyribose. Bases AUCG ATCG. Strand length Short Long
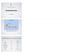 Umm AL Qura University TRANSCRIPTION Dr Neda Bogari TRANSCRIPTION COMPARISON OF DNA & RNA RNA DNA Sugar Ribose Deoxyribose Bases AUCG ATCG Strand length Short Long No. strands One Two Helix Single Double
Umm AL Qura University TRANSCRIPTION Dr Neda Bogari TRANSCRIPTION COMPARISON OF DNA & RNA RNA DNA Sugar Ribose Deoxyribose Bases AUCG ATCG Strand length Short Long No. strands One Two Helix Single Double
DNA Replication and Repair
 DNA Replication and Repair http://hyperphysics.phy-astr.gsu.edu/hbase/organic/imgorg/cendog.gif Overview of DNA Replication SWYK CNs 1, 2, 30 Explain how specific base pairing enables existing DNA strands
DNA Replication and Repair http://hyperphysics.phy-astr.gsu.edu/hbase/organic/imgorg/cendog.gif Overview of DNA Replication SWYK CNs 1, 2, 30 Explain how specific base pairing enables existing DNA strands
DNA Transcription. Visualizing Transcription. The Transcription Process
 DNA Transcription By: Suzanne Clancy, Ph.D. 2008 Nature Education Citation: Clancy, S. (2008) DNA transcription. Nature Education 1(1) If DNA is a book, then how is it read? Learn more about the DNA transcription
DNA Transcription By: Suzanne Clancy, Ph.D. 2008 Nature Education Citation: Clancy, S. (2008) DNA transcription. Nature Education 1(1) If DNA is a book, then how is it read? Learn more about the DNA transcription
BIOLOGY - CLUTCH CH.17 - GENE EXPRESSION.
 !! www.clutchprep.com CONCEPT: GENES Beadle and Tatum develop the one gene one enzyme hypothesis through their work with Neurospora (bread mold). This idea was later revised as the one gene one polypeptide
!! www.clutchprep.com CONCEPT: GENES Beadle and Tatum develop the one gene one enzyme hypothesis through their work with Neurospora (bread mold). This idea was later revised as the one gene one polypeptide
Molecular Cell Biology - Problem Drill 08: Transcription, Translation and the Genetic Code
 Molecular Cell Biology - Problem Drill 08: Transcription, Translation and the Genetic Code Question No. 1 of 10 1. Which of the following statements about how genes function is correct? Question #1 (A)
Molecular Cell Biology - Problem Drill 08: Transcription, Translation and the Genetic Code Question No. 1 of 10 1. Which of the following statements about how genes function is correct? Question #1 (A)
DIFFERENT ASPECTS OF GENE REGULATION
 TARTU UNIVERSITY FACULTY OF BILOGY AND GEOGRAPHY DIFFERENT ASPECTS OF GENE REGULATION TARTU 2005 Sten Ilmjärv 2 TABLE OF CONTENT TABLE OF CONTENT...2 GLOSSARY...3 INTRODUCTION...4 1. THE GENE...5 2. GENE
TARTU UNIVERSITY FACULTY OF BILOGY AND GEOGRAPHY DIFFERENT ASPECTS OF GENE REGULATION TARTU 2005 Sten Ilmjärv 2 TABLE OF CONTENT TABLE OF CONTENT...2 GLOSSARY...3 INTRODUCTION...4 1. THE GENE...5 2. GENE
Review of Protein (one or more polypeptide) A polypeptide is a long chain of..
 Gene expression Review of Protein (one or more polypeptide) A polypeptide is a long chain of.. In a protein, the sequence of amino acid determines its which determines the protein s A protein with an enzymatic
Gene expression Review of Protein (one or more polypeptide) A polypeptide is a long chain of.. In a protein, the sequence of amino acid determines its which determines the protein s A protein with an enzymatic
Classes of eukaryotic cellular RNAs
 Classes of eukaryotic cellular RNAs ribosomal RNA (rrna) 18S (small subunit) 28S (large subunit) 5.8S (large subunit) 5S (large subunit) transfer RNA (trna) messenger RNA (mrna) heterogeneous nuclear RNA
Classes of eukaryotic cellular RNAs ribosomal RNA (rrna) 18S (small subunit) 28S (large subunit) 5.8S (large subunit) 5S (large subunit) transfer RNA (trna) messenger RNA (mrna) heterogeneous nuclear RNA
Gene function at the level of traits Gene function at the molecular level
 Gene expression Gene function at the level of traits Gene function at the molecular level Two levels tied together since the molecular level affects the structure and function of cells which determines
Gene expression Gene function at the level of traits Gene function at the molecular level Two levels tied together since the molecular level affects the structure and function of cells which determines
Fig Ch 17: From Gene to Protein
 Fig. 17-1 Ch 17: From Gene to Protein Basic Principles of Transcription and Translation RNA is the intermediate between genes and the proteins for which they code Transcription is the synthesis of RNA
Fig. 17-1 Ch 17: From Gene to Protein Basic Principles of Transcription and Translation RNA is the intermediate between genes and the proteins for which they code Transcription is the synthesis of RNA
Biological information flow
 BCMB 3100 Chapters 36-38 Transcription & RNA Processing Definition of gene RNA Polymerase Gene coding vs template strand Promoter Transcription in E. coli Transcription factors mrna processing Biological
BCMB 3100 Chapters 36-38 Transcription & RNA Processing Definition of gene RNA Polymerase Gene coding vs template strand Promoter Transcription in E. coli Transcription factors mrna processing Biological
BCH 4054 Fall 2000 Chapter 31 Lecture Notes
 BCH 4054 Fall 2000 Chapter 31 Lecture Notes 1 Chapter 31 Transcription and Regulation of Gene Expression 2 Messenger RNA Central Dogma (Francis Crick, 1958) DNA RNA Protein (Fig 31.1) Jacob-Monod Hypothesis:
BCH 4054 Fall 2000 Chapter 31 Lecture Notes 1 Chapter 31 Transcription and Regulation of Gene Expression 2 Messenger RNA Central Dogma (Francis Crick, 1958) DNA RNA Protein (Fig 31.1) Jacob-Monod Hypothesis:
Chapter 11 Part A: Metabolism: The synthesis of nucleic acids and proteins
 Chapter 11 Part A: Metabolism: The synthesis of nucleic acids and proteins I. Synthesis of DNA = REPLICATION A. Components of DNA (Fig. 11-1) 1. Composed of 4 different nucleotides that are joined by the
Chapter 11 Part A: Metabolism: The synthesis of nucleic acids and proteins I. Synthesis of DNA = REPLICATION A. Components of DNA (Fig. 11-1) 1. Composed of 4 different nucleotides that are joined by the
Chapter 3 Gene Function. Transcription Prokaroyotes Eukaryotes Transcript processing Proteins Translation Genetic nomenclature
 Chapter 3 Gene Function Transcription Prokaroyotes Eukaryotes Transcript processing Proteins Translation Genetic nomenclature Transcription RNA composition ATP, GTP, UTP, CTP are substrates for RNA polymerase.
Chapter 3 Gene Function Transcription Prokaroyotes Eukaryotes Transcript processing Proteins Translation Genetic nomenclature Transcription RNA composition ATP, GTP, UTP, CTP are substrates for RNA polymerase.
Transcription. DNA to RNA
 Transcription from DNA to RNA The Central Dogma of Molecular Biology replication DNA RNA Protein transcription translation Why call it transcription and translation? transcription is such a direct copy
Transcription from DNA to RNA The Central Dogma of Molecular Biology replication DNA RNA Protein transcription translation Why call it transcription and translation? transcription is such a direct copy
Transcription is the first step of gene expression, in which a particular segment of DNA is copied into RNA by the enzyme, RNA polymerase.
 Transcription in Bacteria Transcription in Bacteria Transcription in Bacteria Transcription is the first step of gene expression, in which a particular segment of DNA is copied into RNA by the enzyme,
Transcription in Bacteria Transcription in Bacteria Transcription in Bacteria Transcription is the first step of gene expression, in which a particular segment of DNA is copied into RNA by the enzyme,
There are four major types of introns. Group I introns, found in some rrna genes, are self-splicing: they can catalyze their own removal.
 1 2 Continuous genes - Intron: Many eukaryotic genes contain coding regions called exons and noncoding regions called intervening sequences or introns. The average human gene contains from eight to nine
1 2 Continuous genes - Intron: Many eukaryotic genes contain coding regions called exons and noncoding regions called intervening sequences or introns. The average human gene contains from eight to nine
The 5' cap (red) is added before synthesis of the primary transcript is complete. A non coding sequence following the last exon is shown in orange.
 RNA PROCESSING The 5' cap (red) is added before synthesis of the primary transcript is complete. A non coding sequence following the last exon is shown in orange. Splicing can occur either before or after
RNA PROCESSING The 5' cap (red) is added before synthesis of the primary transcript is complete. A non coding sequence following the last exon is shown in orange. Splicing can occur either before or after
DNA Topoisomerases relieve the supercoiling stress ahead of the fork
 DNA Topoisomerases relieve the supercoiling stress ahead of the fork Tw 1) T w : # of turns around the central axis 2) W r : # of times the double helix crosses itself 3) Linking Number: L k = T w + W
DNA Topoisomerases relieve the supercoiling stress ahead of the fork Tw 1) T w : # of turns around the central axis 2) W r : # of times the double helix crosses itself 3) Linking Number: L k = T w + W
BEADLE & TATUM EXPERIMENT
 FROM DNA TO PROTEINS: gene expression Chapter 14 LECTURE OBJECTIVES What Is the Evidence that Genes Code for Proteins? How Does Information Flow from Genes to Proteins? How Is the Information Content in
FROM DNA TO PROTEINS: gene expression Chapter 14 LECTURE OBJECTIVES What Is the Evidence that Genes Code for Proteins? How Does Information Flow from Genes to Proteins? How Is the Information Content in
Bis2A 12.0 Transcription *
 OpenStax-CNX module: m56068 1 Bis2A 12.0 Transcription * Mitch Singer Based on Transcription by OpenStax This work is produced by OpenStax-CNX and licensed under the Creative Commons Attribution License
OpenStax-CNX module: m56068 1 Bis2A 12.0 Transcription * Mitch Singer Based on Transcription by OpenStax This work is produced by OpenStax-CNX and licensed under the Creative Commons Attribution License
I. Gene Expression Figure 1: Central Dogma of Molecular Biology
 I. Gene Expression Figure 1: Central Dogma of Molecular Biology Central Dogma: Gene Expression: RNA Structure RNA nucleotides contain the pentose sugar Ribose instead of deoxyribose. Contain the bases
I. Gene Expression Figure 1: Central Dogma of Molecular Biology Central Dogma: Gene Expression: RNA Structure RNA nucleotides contain the pentose sugar Ribose instead of deoxyribose. Contain the bases
CH 17 :From Gene to Protein
 CH 17 :From Gene to Protein Defining a gene gene gene Defining a gene is problematic because one gene can code for several protein products, some genes code only for RNA, two genes can overlap, and there
CH 17 :From Gene to Protein Defining a gene gene gene Defining a gene is problematic because one gene can code for several protein products, some genes code only for RNA, two genes can overlap, and there
From Gene to Protein transcription, messenger RNA (mrna) translation, RNA processing triplet code, template strand, codons,
 From Gene to Protein I. Transcription and translation are the two main processes linking gene to protein. A. RNA is chemically similar to DNA, except that it contains ribose as its sugar and substitutes
From Gene to Protein I. Transcription and translation are the two main processes linking gene to protein. A. RNA is chemically similar to DNA, except that it contains ribose as its sugar and substitutes
Information Readout: Transcription and Post-transcriptional Processing Translation
 Information Readout: Transcription and Post-transcriptional Processing Translation Copyright 2013 Pearson Canada Inc. 27-1 DNA as the Template for RNA Synthesis Enzymology of RNA Synthesis: RNA Polymerase
Information Readout: Transcription and Post-transcriptional Processing Translation Copyright 2013 Pearson Canada Inc. 27-1 DNA as the Template for RNA Synthesis Enzymology of RNA Synthesis: RNA Polymerase
Themes: RNA and RNA Processing. Messenger RNA (mrna) What is a gene? RNA is very versatile! RNA-RNA interactions are very important!
 Themes: RNA is very versatile! RNA and RNA Processing Chapter 14 RNA-RNA interactions are very important! Prokaryotes and Eukaryotes have many important differences. Messenger RNA (mrna) Carries genetic
Themes: RNA is very versatile! RNA and RNA Processing Chapter 14 RNA-RNA interactions are very important! Prokaryotes and Eukaryotes have many important differences. Messenger RNA (mrna) Carries genetic
Differential Gene Expression
 Developmental Biology Biology 4361 Differential Gene Expression October 13, 2005 core transcription initiation site 5 promoter 3 TATAT +1 upstream downstream Basal transcription factors (eukaryotes) TFIID
Developmental Biology Biology 4361 Differential Gene Expression October 13, 2005 core transcription initiation site 5 promoter 3 TATAT +1 upstream downstream Basal transcription factors (eukaryotes) TFIID
Biological information flow
 BCMB 3100 Chapters 36-38 Transcription & RNA Processing Biological information flow Definition of gene RNA Polymerase Gene coding vs template strand Promoter Transcription in E. coli Transcription factors
BCMB 3100 Chapters 36-38 Transcription & RNA Processing Biological information flow Definition of gene RNA Polymerase Gene coding vs template strand Promoter Transcription in E. coli Transcription factors
Videos. Bozeman Transcription and Translation: Drawing transcription and translation:
 Videos Bozeman Transcription and Translation: https://youtu.be/h3b9arupxzg Drawing transcription and translation: https://youtu.be/6yqplgnjr4q Objectives 29a) I can contrast RNA and DNA. 29b) I can explain
Videos Bozeman Transcription and Translation: https://youtu.be/h3b9arupxzg Drawing transcription and translation: https://youtu.be/6yqplgnjr4q Objectives 29a) I can contrast RNA and DNA. 29b) I can explain
RNA: Structure & Synthesis. Amr S. Moustafa, M.D.; Ph.D.
 RNA: Structure & Synthesis By Amr S. Moustafa, M.D.; Ph.D. Objectives The differences between DNA and RNA The structure and functions of RNAs RNA synthesis (Transcription) Post-transcriptional events (modifications)
RNA: Structure & Synthesis By Amr S. Moustafa, M.D.; Ph.D. Objectives The differences between DNA and RNA The structure and functions of RNAs RNA synthesis (Transcription) Post-transcriptional events (modifications)
SSA Signal Search Analysis II
 SSA Signal Search Analysis II SSA other applications - translation In contrast to translation initiation in bacteria, translation initiation in eukaryotes is not guided by a Shine-Dalgarno like motif.
SSA Signal Search Analysis II SSA other applications - translation In contrast to translation initiation in bacteria, translation initiation in eukaryotes is not guided by a Shine-Dalgarno like motif.
Biological information flow
 BCMB 3100 Chapters 36-38 Transcription & RNA Processing Definition of gene RNA Polymerase Gene coding vs template strand Promoter Transcription in E. coli Transcription factors mrna processing Biological
BCMB 3100 Chapters 36-38 Transcription & RNA Processing Definition of gene RNA Polymerase Gene coding vs template strand Promoter Transcription in E. coli Transcription factors mrna processing Biological
Protein Synthesis Notes
 Protein Synthesis Notes Protein Synthesis: Overview Transcription: synthesis of mrna under the direction of DNA. Translation: actual synthesis of a polypeptide under the direction of mrna. Transcription
Protein Synthesis Notes Protein Synthesis: Overview Transcription: synthesis of mrna under the direction of DNA. Translation: actual synthesis of a polypeptide under the direction of mrna. Transcription
However, only a fraction of these genes are transcribed in an individual cell at any given time.
 All cells in an organism contain the same set of genes. However, only a fraction of these genes are transcribed in an individual cell at any given time. It is the pattern of gene expression that determines
All cells in an organism contain the same set of genes. However, only a fraction of these genes are transcribed in an individual cell at any given time. It is the pattern of gene expression that determines
Chapter 17. From Gene to Protein
 Chapter 17 From Gene to Protein Overview: The Flow of Genetic Information The information content of DNA is in the form of specific sequences of nucleotides The DNA inherited by an organism leads to specific
Chapter 17 From Gene to Protein Overview: The Flow of Genetic Information The information content of DNA is in the form of specific sequences of nucleotides The DNA inherited by an organism leads to specific
DNA Function: Information Transmission
 DNA Function: Information Transmission DNA is called the code of life. What does it code for? *the information ( code ) to make proteins! Why are proteins so important? Nearly every function of a living
DNA Function: Information Transmission DNA is called the code of life. What does it code for? *the information ( code ) to make proteins! Why are proteins so important? Nearly every function of a living
Figure A summary of spontaneous alterations likely to require DNA repair.
 DNA Damage Figure 5-46. A summary of spontaneous alterations likely to require DNA repair. The sites on each nucleotide that are known to be modified by spontaneous oxidative damage (red arrows), hydrolytic
DNA Damage Figure 5-46. A summary of spontaneous alterations likely to require DNA repair. The sites on each nucleotide that are known to be modified by spontaneous oxidative damage (red arrows), hydrolytic
Resources. This lecture Campbell and Farrell's Biochemistry, Chapter 11
 Transcription Resources This lecture Campbell and Farrell's Biochemistry, Chapter 11 2 Definition of a gene The entire nucleic acid sequence that is necessary for the synthesis of a functional polypeptide
Transcription Resources This lecture Campbell and Farrell's Biochemistry, Chapter 11 2 Definition of a gene The entire nucleic acid sequence that is necessary for the synthesis of a functional polypeptide
Differential Gene Expression
 Biology 4361 Developmental Biology Differential Gene Expression June 19, 2008 Differential Gene Expression Overview Chromatin structure Gene anatomy RNA processing and protein production Initiating transcription:
Biology 4361 Developmental Biology Differential Gene Expression June 19, 2008 Differential Gene Expression Overview Chromatin structure Gene anatomy RNA processing and protein production Initiating transcription:
Chapter 13. From DNA to Protein
 Chapter 13 From DNA to Protein Proteins All proteins consist of polypeptide chains A linear sequence of amino acids Each chain corresponds to the nucleotide base sequenceof a gene The Path From Genes to
Chapter 13 From DNA to Protein Proteins All proteins consist of polypeptide chains A linear sequence of amino acids Each chain corresponds to the nucleotide base sequenceof a gene The Path From Genes to
Analyzed Fungi Neurospora crassa mutants. Mutants were UNABLE to grow without Arginine (an amino acid) Other biochemical experiments indicated:
 From Gene to Protein Beadle and Tatum Analyzed Fungi Neurospora crassa mutants Mutants were UNABLE to grow without Arginine (an amino acid) Other biochemical experiments indicated: Precursor Ornithine
From Gene to Protein Beadle and Tatum Analyzed Fungi Neurospora crassa mutants Mutants were UNABLE to grow without Arginine (an amino acid) Other biochemical experiments indicated: Precursor Ornithine
IN E. COLI WHAT IS THE FUNCTION OF DNA POLYMERASE III
 10 January, 2018 IN E. COLI WHAT IS THE FUNCTION OF DNA POLYMERASE III Document Filetype: PDF 312 KB 0 IN E. COLI WHAT IS THE FUNCTION OF DNA POLYMERASE III The actual replication enzyme in E. Both will
10 January, 2018 IN E. COLI WHAT IS THE FUNCTION OF DNA POLYMERASE III Document Filetype: PDF 312 KB 0 IN E. COLI WHAT IS THE FUNCTION OF DNA POLYMERASE III The actual replication enzyme in E. Both will
Biological information flow
 BCMB 3100 Chapters 36-38 Transcription & RNA Processing Biological information flow Definition of gene RNA Polymerase Gene coding vs template strand Promoter Transcription in E. coli Transcription factors
BCMB 3100 Chapters 36-38 Transcription & RNA Processing Biological information flow Definition of gene RNA Polymerase Gene coding vs template strand Promoter Transcription in E. coli Transcription factors
Chapter 26 RNA Metabolism
 Chapter 26 RNA Metabolism 1. How is RNA synthesized using DNA templates (transcription 转录 )? 2. How is newly synthesized primary RNA transcripts further processed to make functional RNA molecules (RNA
Chapter 26 RNA Metabolism 1. How is RNA synthesized using DNA templates (transcription 转录 )? 2. How is newly synthesized primary RNA transcripts further processed to make functional RNA molecules (RNA
DNA REPLICATION. DNA structure. Semiconservative replication. DNA structure. Origin of replication. Replication bubbles and forks.
 DNA REPLICATION 5 4 Phosphate 3 DNA structure Nitrogenous base 1 Deoxyribose 2 Nucleotide DNA strand = DNA polynucleotide 2004 Biology Olympiad Preparation Program 2 2004 Biology Olympiad Preparation Program
DNA REPLICATION 5 4 Phosphate 3 DNA structure Nitrogenous base 1 Deoxyribose 2 Nucleotide DNA strand = DNA polynucleotide 2004 Biology Olympiad Preparation Program 2 2004 Biology Olympiad Preparation Program
Chapter 3 Expression of Genes
 Part I Relationship between Cells and Genetic Information A protein gene is a piece of DNA that determines the amino acid sequence of a protein, and the synthesis of a protein based on genetic information
Part I Relationship between Cells and Genetic Information A protein gene is a piece of DNA that determines the amino acid sequence of a protein, and the synthesis of a protein based on genetic information
Chapter 17. From Gene to Protein. Slide 1. Slide 2. Slide 3. Gene Expression. Which of the following is the best example of gene expression? Why?
 Slide 1 Chapter 17 From Gene to Protein PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley with contributions from
Slide 1 Chapter 17 From Gene to Protein PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley with contributions from
The Nature of Genes. The Nature of Genes. The Nature of Genes. The Nature of Genes. The Nature of Genes. The Genetic Code. Genes and How They Work
 Genes and How They Work Chapter 15 Early ideas to explain how genes work came from studying human diseases. Archibald Garrod studied alkaptonuria, 1902 Garrod recognized that the disease is inherited via
Genes and How They Work Chapter 15 Early ideas to explain how genes work came from studying human diseases. Archibald Garrod studied alkaptonuria, 1902 Garrod recognized that the disease is inherited via
Genes and How They Work. Chapter 15
 Genes and How They Work Chapter 15 1 The Nature of Genes Early ideas to explain how genes work came from studying human diseases Archibald Garrod 1902 Recognized that alkaptonuria is inherited via a recessive
Genes and How They Work Chapter 15 1 The Nature of Genes Early ideas to explain how genes work came from studying human diseases Archibald Garrod 1902 Recognized that alkaptonuria is inherited via a recessive
