New Statistical Algorithms for Monitoring Gene Expression on GeneChip Probe Arrays
|
|
|
- Dustin Flynn
- 6 years ago
- Views:
Transcription
1 GENE EXPRESSION MONITORING TECHNICAL NOTE New Statistical Algorithms for Monitoring Gene Expression on GeneChip Probe Arrays Introduction Affymetrix has designed new algorithms for monitoring GeneChip expression data. These statistical algorithms, created in response to customer input, were designed to accommodate the typical distribution of data found in microarray experiments. The new statistical algorithms employ standard statistical techniques and are optimized to accommodate advancements in array and probe selection technology. They provide accurate, high-quality analysis for GeneChip array data. This new, statistically based approach provides: Calculation of statistical significance for detection and change calls (p-values) and confidence limits for log ratio values (fold change). Easily tunable parameters that enable the user to vary the stringency of the analyses. Elimination of negative expression values observed with the empirical algorithms. Easily referenced standard statistical techniques. This technical note reviews the design and testing for Affymetrix new statistical algorithms and explores performance characteristics of the statistical algorithms versus the previous empirical algorithms. Experimental Design: How the New Algorithms Were Selected and Optimized To select components for the new algorithms and test for optimization, a training data set was required. To conduct this comprehensive testing, each transcript group was spiked into a labeled mixture of RNA from a tissue source in an GeneChip Experiment Groups of Transcripts pm Concentration experimental design known as a Latin Square. A Latin Square is used to accurately monitor the detectability of transcripts over a range of concentrations. It also allows the statistical analysis of patterns and variability in repeated measurements in a systematic fashion, thus revealing patterns in the data and allowing rigorous comparisons. The Latin Square experimental design used in the development of the statistical algorithms enabled thorough testing of a large set of transcripts over a broad range of concentrations. (Figure ) A B C D E F G H I J K L M N Figure. Latin Square used in algorithm development. Each row (numbered through 4) represents a GeneChip experiment. Each column (labeled A through N) represents a distinct set of transcripts. Each set contains a pool of transcripts distinct from the transcripts in any other set. In other words, no transcript is present in more than one set. Additionally, no set of transcripts is present at the same concentration in more than one experiment. For example, in Experiment 2 (shaded in blue), every transcript in Set J (boxed in red) is spiked along with the complex background into the target hybridization mixture at 28 pm. Then in Experiment 3, the same set of transcripts, (i.e., Set J boxed in green) is spiked in at 256 pm, twice the previous concentration, and is present at a different concentration in each subsequent experiment.
2 The Latin Square experimental design was used extensively in the algorithm development process to test a wide range of data sets, including transcript groups of E.coli, S. cerevisiae and H. sapiens. In the human Latin Square, each transcript group was designed to contain one distinct human transcript from to 24 pm in concentration. These were spiked into a labeled mixture of human RNA where these 4 transcripts showed no expression. In total, 2 transcripts were used to compute the results; two of the transcripts were removed from the final calculations due to low quality. In the yeast Latin Square, each transcript group contained eight different transcripts, labeled and spiked into a mixture of labeled human RNA. After hybridization to Human Genome U95Av2 or Yeast Genome S98 arrays respectively, Affymetrix Microarray Suite (MAS) 4. containing the empirical algorithms and MAS 5. containing the new statistical algorithms were used to analyze the data. In addition to Latin Square experiments, more conventional data sets were generated and analyzed where RNA from different sources was labeled and hybridized to GeneChip probe arrays, followed by analysis with MAS 4. and MAS 5.. A. MAS 5. Signal B. MAS 5. Signal Human Adrenal Gland RNA on HG-U95Av MAS 4. Average Difference Human Adrenal Gland RNA on HG-U95Av y =.3x R 2 = y =.3x R 2 = MAS 4. Average Difference 2 Figure 2. Expression values in experiments with human adrenal gland RNA. A. Human adrenal gland RNA was labeled and hybridized to a Human Genome U95Av2 GeneChip probe array. The scanned image was analyzed with MAS 4., as well as with MAS 5.. The Average Difference values for all 2,625 probe sets derived from MAS 4. were plotted on the x-axis against the Signal derived from MAS 5. on the y-axis. B. The lower left quadrant of A is enlarged to indicate the absence of negative signal values in MAS Results Comparison of Expression Values Generated by MAS 4. and MAS 5. Analyses were performed to study the concordance between expression values generated by empirical algorithms (MAS 4.) and statistical algorithms (MAS 5.). Experiments performed with human adrenal gland RNA on human genome U95A arrays are shown in Figure 2. As shown in Figure 2A, when results generated with MAS 4. and MAS 5. are compared, the results are highly similar, with a regression ratio of.94. Similar results were obtained across multiple tissues and with replicates (data not shown). This similarity was also observed on arrays representing different organisms. The regression ratios for these comparisons were in the range for M. musculus, A. thaliana and S. cerevisiae, and for D. melanogaster and E. coli. The occurrence of negative values in MAS 4., seen in the two left quadrants in Figure 2B, has been eliminated from MAS 5.. The corresponding output in MAS 5., termed Signal has no negative and zero values as is evident in the lower two quadrants of Figure 2B.
3 Fraction Called Present Added Statistical Quality Measures and Tunable Parameters An added benefit of the statistical algorithms in MAS 5. is the inclusion of probability values (statistical significance) associated with detection and comparison calls. This additional metric allows users to assess the significance of results, and if desired, adjust the balance between sensitivity and specificity. For example, higher confidence in expression calls (greater specificity) can be achieved by accepting fewer Present calls (lower sensitivity). Conversely, greater sensitivity may be achieved at the expense of lower specificity. The inverse relationship between sensitivity and specificity in MAS 5. was examined by monitoring detection calls in the set of experiments described below. As shown in Figure 3, a greater number of accurate Present calls is Human Latin Square on HG-U95A MAS 4. MAS 5. Figure 3. Expression calls in human Latin Square experiment. Analyses performed on the human Latin Square experiments in Figure 3 are shown here. The x-axis represents the different concentrations of spiked transcripts,and the y-axis represents the fraction of Present (P) calls.the MAS 5. analysis was performed at the default setting where transcripts with p values <.4 are assigned Present calls. made by MAS 5. at concentrations below 4 pm using the default p-value setting. However, the small number of false positives is also slightly greater for MAS 5. in this experiment, as log2 (signal) seen at the pm concentration. The sensitivity and specificity may be varied in a predictable fashion by varying the default setting for p-value cutoff. Figure 4 illustrates the linearity of dose response from the 4 spiked transcripts and the upper and lower confidence limits. It also demonstrates that signal calculation in the statistical algorithms (MAS 5.) generates signal values that accurately reflect the true concentration. As with Figure 2, no signal value in Figure 4 is ever negative or zero. This allows signal values to be easily transformed to a logarithmic scale. The effects of altering tunable confidence parameters on Present calls were studied in a yeast Latin Square experiment. The tunable parameter controlling the number of Present calls is termed a (alpha). Altering a results in a shift in the p-value cutoff used to make a Present call. Approximate 95% Confidence Interval for MAS 5. Signal on Human Latin Square (computed using median absolute deviation) Upper Limit Median Signal Lower Limit log2 (concentration) in picomolar Figure 4. Signal value compared to true concentration value. Three-fold replicates of the Latin Square design were performed for 4 human transcripts using 42 human U95A arrays. One outlier array was discarded and the remaining arrays were used to examine the relationship between signal and concentration. The observed variation between replicates of the same transcript at each concentration was used to estimate a 95% confidence interval. The median signal of all transcript-concentration pairs is shown in the figure.
4 This parameter was varied in this analysis to determine the effect on the number of Present calls at different concentrations. Calls at higher concentrations are typically unaffected by altering a. At concentrations below 8 pm in this experiment, a decrease in stringency (i.e., increase in a from.4 to.) results in an increasing number of Present calls. As Figure 5 shows, a may be varied to obtain sensitivities using MAS 5. that are greater or lower than those obtained with MAS 4.. However, an increase in a produces a corresponding increase in the number of false positive calls, as seen at the pm concentration. It should be noted that false positives can easily be flagged by their high p-values. Comparison Calls Generated by MAS 4. and MAS 5. Show Concordance Analyses were performed to study the concordance between comparison calls Fraction Called Present generated by MAS 4. and MAS 5.. To assess No Change calls, comparison analysis was performed on replicate Human Genome U95A arrays from the human Latin Square experiment, where transcript group concentrations were identical between experiments. Therefore, a No Change call is expected for each transcript group between replicates. Results are shown in Figure 6. The results show that for both MAS 4. and MAS 5., over 95% (at some concentrations %) correct No Change calls are made. The concordance between Increase calls made by MAS 4. and MAS 5. are shown in Figure 7. The same transcript group was compared in pairs of experiments to evaluate the calls for a 2-fold and 4-fold change in concentration. For example, in the Latin Square represented in Figure, transcript group B was compared between Experiments and 2, then 2 and 3, then 3 and 4, and so on, to determine the fraction Yeast Latin Square on Yeast Genome S98 Arrays MAS 4. MAS 5. α =.4 MAS 5. α =.6 MAS 5. α = Fraction of Correct No Change Calls Human Latin Square: Replicate HG-U95 Arrays MAS 4. MAS Figure 6. Human Latin Square: No Change Calls. Replicate experiments from the Human Latin Square set were analyzed to assess the No Change calls. Of the 4 groups of experiments, groups had three replicates, one group had two replicates and two groups had 2 replicates each. The x-axis represents the concentration of the spiked transcripts. The y-axis represents the fraction of No Change calls. A. Human Latin Square, 2-fold Increase.2 MAS 4. MAS Fraction of Correct Increase Calls Fraction of Correct Increase Calls B. Human Latin Square, 4-fold Increase.2 MAS 4. MAS Figure 7. Increase calls in a human Latin Square. A. The fraction of Increase calls at the 2-fold change was plotted on the y-axis. The spiked transcript concentration is shown on the x-axis. B. The fraction of Increase calls at the 4-fold change was plotted on the y-axis. The spiked transcript concentration is shown on the x-axis. Figure 5. Expression calls in yeast Latin Square experiment: Altering p-value cutoff. Fourteen different sets of yeast transcripts were spiked into a mixture of labeled human RNA at concentrations ranging from to 24 pm and hybridized to 4 Yeast Genome S98 arrays. The x-axis represents the concentration of spiked transcripts. The y-axis represents the fraction of Present calls. The dark blue curve represents analyses performed with MAS 4., whereas the other three curves represent analyses performed at three different a settings of MAS 5.:.4,.5 and..
5 correctly assigned an Increase call at the expected 2-fold change. For the 4-fold change, transcript group B was compared between Experiments and 3, 2 and 4, 3 and 5 etc., such that all transcript groups were evaluated. In this data set, MAS 5. shows greater accuracy than does MAS 4. at both 2-fold and 4-fold Increase levels. For example, at the 2 pm spike in Figure 7B, MAS 5. gives a nearly % greater fraction of correct Increase calls than does MAS 4.. The drop in accuracy seen at very high concentration is due to the saturation of probe sets by the vast excess of spiked transcripts. To understand the difference in the performance of calls generated by MAS 4. and MAS 5. on a biological sample, we assessed the results using an independent method. We used the power of replicates and the wellknown student s t-test as a reference. We analyzed a number of replicates in two tissues and then asked, How well do the results from any single comparison match results from the group as a whole? The group was built from two sets of experiments mouse brain and mouse heart containing six replicates for each tissue. Each replicate was compared to every other replicate with both MAS 4. and MAS 5. (36 comparisons of 2488 probe sets). The t-test was then used to evaluate whether the mean of the replicates in one tissue was significantly different from the mean of the replicates in the other tissue. The results are shown in Figure 8. Overall, the Comparison calls generated by MAS 4. and MAS 5. are highly similar. The No Change calls of both MAS 4. and MAS 5. peak at approximately.9, as we would expect. The Decrease calls (D) peak at and the Increase calls (I) peak towards zero for both algorithms, while the lines follow each other very closely. The additional benefit of MAS 5. is that we now have p-values to assess statistical significance of the comparisons of every gene. The next technical note will contain more results on Comparison calls and their performance in MAS 5., together with discussions of tunable parameters and confidence limits. Conclusions In conclusion, our extensive validation studies, some results from which are shown above, allow us to state with confidence the following: The new statistical algorithms in MAS 5. perform as robustly and precisely as the previous algorithms, while providing the additional value of statistical significance (p values) and confidence limits, thereby offering GeneChip users the ability to evaluate the significance of their results. Negative expression values (signals) have been eliminated; only positive values are generated in MAS 5.. Comparative Calls with Mouse Brain RNA vs. Mouse Heart RNA on MG-U74Av2 Arrays Number of Calls (Increase, Decrease and No Change) E E E p-value of t-test MAS 4. D MAS 4. NC MAS 4. I MAS 5. D MAS 5. NC MAS 5. I Figure 8. Comparison Calls with Mouse Brain RNA vs. Mouse Heart RNA on U74Av2 arrays. Two sets of experiments were analyzed, each consisting of six replicate mouse U74Av2 arrays hybridized to samples derived from mouse brain RNA or from mouse heart RNA. These two sets of replicate experiments were analyzed with MAS 4. and MAS 5.. Thirty-six pair-wise comparisons were performed between the two sets of experiments. The y-axis represents the number of Increase, Decrease or No Change calls made by MAS 4. or MAS 5., for all 36 pair-wise comparisons, (i.e., for 36 x 2488 probe sets). A student's t-test was performed on the two sets of experiments and a p-value was generated for each comparison of individual probe sets. This p-value was plotted on the x-axis. The graph assesses the distribution of Comparison Calls made by MAS 4. and MAS 5. in comparison to a t-test.
6 AFFYMETRIX, INC. 338 Central Expressway Santa Clara, CA 955 USA Tel: (-888-DNA-CHIP) Fax: AFFYMETRIX UK Ltd., Voyager, Mercury Park, Wycombe Lane, Wooburn Green, High Wycombe HP HH United Kingdom Tel: +44 () Fax: +44 () For research use only. Not for use in diagnostic procedures. Part No. 797 Rev 3 2 Affymetrix, Inc. All rights reserved. Affymetrix,GeneChip,, HuSNP, Jaguar, EASI, MicroDB, GenFlex, 47, 48, 427, 428, Pin-and-Ring, Flying Objective,, CustomExpress, NetAffx and are trademarks owned or used by Affymetrix, Inc. Products may be covered by one or more of the following patents and/or sold under license from Oxford Gene Technology: U.S. Patent Nos. 5,445,934; 5,744,35; 6,26,776; 6,29,83; 5,7,637, and 5,945,334; and EP 69 32; and other U.S. or foreign patents.
Technical Note. Performance Review of the GeneChip AutoLoader for the Affymetrix GeneChip Scanner Introduction
 GeneChip AutoLoader AFFYMETRIX PRODUCT FAMILY > > Technical Note Performance Review of the GeneChip AutoLoader for the Affymetrix GeneChip ner 3000 Designed for use with the GeneChip ner 3000, the GeneChip
GeneChip AutoLoader AFFYMETRIX PRODUCT FAMILY > > Technical Note Performance Review of the GeneChip AutoLoader for the Affymetrix GeneChip ner 3000 Designed for use with the GeneChip ner 3000, the GeneChip
Data Sheet. GeneChip Human Genome U133 Arrays
 GeneChip Human Genome Arrays AFFYMETRIX PRODUCT FAMILY > ARRAYS > Data Sheet GeneChip Human Genome U133 Arrays The Most Comprehensive Coverage of the Human Genome in Two Flexible Formats: Single-array
GeneChip Human Genome Arrays AFFYMETRIX PRODUCT FAMILY > ARRAYS > Data Sheet GeneChip Human Genome U133 Arrays The Most Comprehensive Coverage of the Human Genome in Two Flexible Formats: Single-array
For research use only. Not for use in diagnostic procedures. AFFYMETRIX UK Ltd., AFFYMETRIX, INC.
 AFFYMETRIX, INC. 3380 Central Expressway Santa Clara, CA 95051 USA Tel: 1-888-362-2447 (1-888-DNA-CHIP) Fax: 1-408-731-5441 sales@affymetrix.com support@affymetrix.com AFFYMETRIX UK Ltd., Voyager, Mercury
AFFYMETRIX, INC. 3380 Central Expressway Santa Clara, CA 95051 USA Tel: 1-888-362-2447 (1-888-DNA-CHIP) Fax: 1-408-731-5441 sales@affymetrix.com support@affymetrix.com AFFYMETRIX UK Ltd., Voyager, Mercury
GeneChip Eukaryotic Small Sample Target Labeling Assay Version II *
 GENE EXPRESSION MONITORING TECHNICAL NOTE GeneChip Eukaryotic Small Sample Target Labeling Assay Version II * Introduction There is an overwhelming and continuing demand for a well characterized protocol
GENE EXPRESSION MONITORING TECHNICAL NOTE GeneChip Eukaryotic Small Sample Target Labeling Assay Version II * Introduction There is an overwhelming and continuing demand for a well characterized protocol
STATC 141 Spring 2005, April 5 th Lecture notes on Affymetrix arrays. Materials are from
 STATC 141 Spring 2005, April 5 th Lecture notes on Affymetrix arrays Materials are from http://www.ohsu.edu/gmsr/amc/amc_technology.html The GeneChip high-density oligonucleotide arrays are fabricated
STATC 141 Spring 2005, April 5 th Lecture notes on Affymetrix arrays Materials are from http://www.ohsu.edu/gmsr/amc/amc_technology.html The GeneChip high-density oligonucleotide arrays are fabricated
Quality Control Assessment in Genotyping Console
 Quality Control Assessment in Genotyping Console Introduction Prior to the release of Genotyping Console (GTC) 2.1, quality control (QC) assessment of the SNP Array 6.0 assay was performed using the Dynamic
Quality Control Assessment in Genotyping Console Introduction Prior to the release of Genotyping Console (GTC) 2.1, quality control (QC) assessment of the SNP Array 6.0 assay was performed using the Dynamic
FACTORS CONTRIBUTING TO VARIABILITY IN DNA MICROARRAY RESULTS: THE ABRF MICROARRAY RESEARCH GROUP 2002 STUDY
 FACTORS CONTRIBUTING TO VARIABILITY IN DNA MICROARRAY RESULTS: THE ABRF MICROARRAY RESEARCH GROUP 2002 STUDY K. L. Knudtson 1, C. Griffin 2, A. I. Brooks 3, D. A. Iacobas 4, K. Johnson 5, G. Khitrov 6,
FACTORS CONTRIBUTING TO VARIABILITY IN DNA MICROARRAY RESULTS: THE ABRF MICROARRAY RESEARCH GROUP 2002 STUDY K. L. Knudtson 1, C. Griffin 2, A. I. Brooks 3, D. A. Iacobas 4, K. Johnson 5, G. Khitrov 6,
Affymetrix GeneChip Arrays. Lecture 3 (continued) Computational and Statistical Aspects of Microarray Analysis June 21, 2005 Bressanone, Italy
 Affymetrix GeneChip Arrays Lecture 3 (continued) Computational and Statistical Aspects of Microarray Analysis June 21, 2005 Bressanone, Italy Affymetrix GeneChip Design 5 3 Reference sequence TGTGATGGTGGGGAATGGGTCAGAAGGCCTCCGATGCGCCGATTGAGAAT
Affymetrix GeneChip Arrays Lecture 3 (continued) Computational and Statistical Aspects of Microarray Analysis June 21, 2005 Bressanone, Italy Affymetrix GeneChip Design 5 3 Reference sequence TGTGATGGTGGGGAATGGGTCAGAAGGCCTCCGATGCGCCGATTGAGAAT
Introduction to gene expression microarray data analysis
 Introduction to gene expression microarray data analysis Outline Brief introduction: Technology and data. Statistical challenges in data analysis. Preprocessing data normalization and transformation. Useful
Introduction to gene expression microarray data analysis Outline Brief introduction: Technology and data. Statistical challenges in data analysis. Preprocessing data normalization and transformation. Useful
EECS730: Introduction to Bioinformatics
 EECS730: Introduction to Bioinformatics Lecture 14: Microarray Some slides were adapted from Dr. Luke Huan (University of Kansas), Dr. Shaojie Zhang (University of Central Florida), and Dr. Dong Xu and
EECS730: Introduction to Bioinformatics Lecture 14: Microarray Some slides were adapted from Dr. Luke Huan (University of Kansas), Dr. Shaojie Zhang (University of Central Florida), and Dr. Dong Xu and
Preprocessing Affymetrix GeneChip Data. Affymetrix GeneChip Design. Terminology TGTGATGGTGGGGAATGGGTCAGAAGGCCTCCGATGCGCCGATTGAGAAT
 Preprocessing Affymetrix GeneChip Data Credit for some of today s materials: Ben Bolstad, Leslie Cope, Laurent Gautier, Terry Speed and Zhijin Wu Affymetrix GeneChip Design 5 3 Reference sequence TGTGATGGTGGGGAATGGGTCAGAAGGCCTCCGATGCGCCGATTGAGAAT
Preprocessing Affymetrix GeneChip Data Credit for some of today s materials: Ben Bolstad, Leslie Cope, Laurent Gautier, Terry Speed and Zhijin Wu Affymetrix GeneChip Design 5 3 Reference sequence TGTGATGGTGGGGAATGGGTCAGAAGGCCTCCGATGCGCCGATTGAGAAT
A Distribution Free Summarization Method for Affymetrix GeneChip Arrays
 A Distribution Free Summarization Method for Affymetrix GeneChip Arrays Zhongxue Chen 1,2, Monnie McGee 1,*, Qingzhong Liu 3, and Richard Scheuermann 2 1 Department of Statistical Science, Southern Methodist
A Distribution Free Summarization Method for Affymetrix GeneChip Arrays Zhongxue Chen 1,2, Monnie McGee 1,*, Qingzhong Liu 3, and Richard Scheuermann 2 1 Department of Statistical Science, Southern Methodist
Background Correction and Normalization. Lecture 3 Computational and Statistical Aspects of Microarray Analysis June 21, 2005 Bressanone, Italy
 Background Correction and Normalization Lecture 3 Computational and Statistical Aspects of Microarray Analysis June 21, 2005 Bressanone, Italy Feature Level Data Outline Affymetrix GeneChip arrays Two
Background Correction and Normalization Lecture 3 Computational and Statistical Aspects of Microarray Analysis June 21, 2005 Bressanone, Italy Feature Level Data Outline Affymetrix GeneChip arrays Two
Analysis of Microarray Data
 Analysis of Microarray Data Lecture 1: Experimental Design and Data Normalization George Bell, Ph.D. Senior Bioinformatics Scientist Bioinformatics and Research Computing Whitehead Institute Outline Introduction
Analysis of Microarray Data Lecture 1: Experimental Design and Data Normalization George Bell, Ph.D. Senior Bioinformatics Scientist Bioinformatics and Research Computing Whitehead Institute Outline Introduction
Gene Signal Estimates from Exon Arrays
 Gene Signal Estimates from Exon Arrays I. Introduction: With exon arrays like the GeneChip Human Exon 1.0 ST Array, researchers can examine the transcriptional profile of an entire gene (Figure 1). Being
Gene Signal Estimates from Exon Arrays I. Introduction: With exon arrays like the GeneChip Human Exon 1.0 ST Array, researchers can examine the transcriptional profile of an entire gene (Figure 1). Being
Outline. Analysis of Microarray Data. Most important design question. General experimental issues
 Outline Analysis of Microarray Data Lecture 1: Experimental Design and Data Normalization Introduction to microarrays Experimental design Data normalization Other data transformation Exercises George Bell,
Outline Analysis of Microarray Data Lecture 1: Experimental Design and Data Normalization Introduction to microarrays Experimental design Data normalization Other data transformation Exercises George Bell,
Analysis of a Tiling Regulation Study in Partek Genomics Suite 6.6
 Analysis of a Tiling Regulation Study in Partek Genomics Suite 6.6 The example data set used in this tutorial consists of 6 technical replicates from the same human cell line, 3 are SP1 treated, and 3
Analysis of a Tiling Regulation Study in Partek Genomics Suite 6.6 The example data set used in this tutorial consists of 6 technical replicates from the same human cell line, 3 are SP1 treated, and 3
What does PLIER really do?
 What does PLIER really do? Terry M. Therneau Karla V. Ballman Technical Report #75 November 2005 Copyright 2005 Mayo Foundation 1 Abstract Motivation: Our goal was to understand why the PLIER algorithm
What does PLIER really do? Terry M. Therneau Karla V. Ballman Technical Report #75 November 2005 Copyright 2005 Mayo Foundation 1 Abstract Motivation: Our goal was to understand why the PLIER algorithm
Overview. General. RNA & Microarrays. Information Systems in the Life Sciences
 Overview Information Systems in the Life Sciences PERP GeneChip data warehouse; Implementation of a dynamic time series query tool with graphical interface General PERP GeneChip data warehouse Affymetrix
Overview Information Systems in the Life Sciences PERP GeneChip data warehouse; Implementation of a dynamic time series query tool with graphical interface General PERP GeneChip data warehouse Affymetrix
Probe-Level Data Normalisation: RMA and GC-RMA Sam Robson Images courtesy of Neil Ward, European Application Engineer, Agilent Technologies.
 Probe-Level Data Normalisation: RMA and GC-RMA Sam Robson Images courtesy of Neil Ward, European Application Engineer, Agilent Technologies. References Summaries of Affymetrix Genechip Probe Level Data,
Probe-Level Data Normalisation: RMA and GC-RMA Sam Robson Images courtesy of Neil Ward, European Application Engineer, Agilent Technologies. References Summaries of Affymetrix Genechip Probe Level Data,
New Stringent Two-Color Gene Expression Workflow Enables More Accurate and Reproducible Microarray Data
 Application Note GENOMICS INFORMATICS PROTEOMICS METABOLOMICS A T C T GATCCTTC T G AAC GGAAC T AATTTC AA G AATCTGATCCTTG AACTACCTTCCAAGGTG New Stringent Two-Color Gene Expression Workflow Enables More
Application Note GENOMICS INFORMATICS PROTEOMICS METABOLOMICS A T C T GATCCTTC T G AAC GGAAC T AATTTC AA G AATCTGATCCTTG AACTACCTTCCAAGGTG New Stringent Two-Color Gene Expression Workflow Enables More
Technical Note. SNP Selection Process
 GeneChip SNP Selection AFFYMETRIX PRODUCT FAMILY > DNA ARRAYS AND REAGENTS > Technical Note SNP Selection Criteria for the GeneChip Human Mapping 10K Array Xba 131 The GeneChip Human Mapping 10K Array
GeneChip SNP Selection AFFYMETRIX PRODUCT FAMILY > DNA ARRAYS AND REAGENTS > Technical Note SNP Selection Criteria for the GeneChip Human Mapping 10K Array Xba 131 The GeneChip Human Mapping 10K Array
Exploration, Normalization, Summaries, and Software for Affymetrix Probe Level Data
 Exploration, Normalization, Summaries, and Software for Affymetrix Probe Level Data Rafael A. Irizarry Department of Biostatistics, JHU March 12, 2003 Outline Review of technology Why study probe level
Exploration, Normalization, Summaries, and Software for Affymetrix Probe Level Data Rafael A. Irizarry Department of Biostatistics, JHU March 12, 2003 Outline Review of technology Why study probe level
Expression Array System
 Integrated Science for Gene Expression Applied Biosystems Expression Array System Expression Array System SEE MORE GENES The most complete, most sensitive system for whole genome expression analysis. The
Integrated Science for Gene Expression Applied Biosystems Expression Array System Expression Array System SEE MORE GENES The most complete, most sensitive system for whole genome expression analysis. The
Mixed effects model for assessing RNA degradation in Affymetrix GeneChip experiments
 Mixed effects model for assessing RNA degradation in Affymetrix GeneChip experiments Kellie J. Archer, Ph.D. Suresh E. Joel Viswanathan Ramakrishnan,, Ph.D. Department of Biostatistics Virginia Commonwealth
Mixed effects model for assessing RNA degradation in Affymetrix GeneChip experiments Kellie J. Archer, Ph.D. Suresh E. Joel Viswanathan Ramakrishnan,, Ph.D. Department of Biostatistics Virginia Commonwealth
Predicting Microarray Signals by Physical Modeling. Josh Deutsch. University of California. Santa Cruz
 Predicting Microarray Signals by Physical Modeling Josh Deutsch University of California Santa Cruz Predicting Microarray Signals by Physical Modeling p.1/39 Collaborators Shoudan Liang NASA Ames Onuttom
Predicting Microarray Signals by Physical Modeling Josh Deutsch University of California Santa Cruz Predicting Microarray Signals by Physical Modeling p.1/39 Collaborators Shoudan Liang NASA Ames Onuttom
DNA Microarray Data Oligonucleotide Arrays
 DNA Microarray Data Oligonucleotide Arrays Sandrine Dudoit, Robert Gentleman, Rafael Irizarry, and Yee Hwa Yang Bioconductor Short Course 2003 Copyright 2002, all rights reserved Biological question Experimental
DNA Microarray Data Oligonucleotide Arrays Sandrine Dudoit, Robert Gentleman, Rafael Irizarry, and Yee Hwa Yang Bioconductor Short Course 2003 Copyright 2002, all rights reserved Biological question Experimental
Humboldt Universität zu Berlin. Grundlagen der Bioinformatik SS Microarrays. Lecture
 Humboldt Universität zu Berlin Microarrays Grundlagen der Bioinformatik SS 2017 Lecture 6 09.06.2017 Agenda 1.mRNA: Genomic background 2.Overview: Microarray 3.Data-analysis: Quality control & normalization
Humboldt Universität zu Berlin Microarrays Grundlagen der Bioinformatik SS 2017 Lecture 6 09.06.2017 Agenda 1.mRNA: Genomic background 2.Overview: Microarray 3.Data-analysis: Quality control & normalization
User Bulletin. ABI PRISM 7000 Sequence Detection System DRAFT. SUBJECT: Software Upgrade from Version 1.0 to 1.1. Version 1.
 User Bulletin ABI PRISM 7000 Sequence Detection System 9/15/03 SUBJECT: Software Upgrade from Version 1.0 to 1.1 The ABI PRISM 7000 Sequence Detection System version 1.1 software is a free upgrade. Also
User Bulletin ABI PRISM 7000 Sequence Detection System 9/15/03 SUBJECT: Software Upgrade from Version 1.0 to 1.1 The ABI PRISM 7000 Sequence Detection System version 1.1 software is a free upgrade. Also
% Viability. isw2 ino isw2 ino isw2 ino isw2 ino mM HU 4-NQO CPT
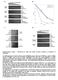 a Drug concentration b 1.3% MMS nhp1 nhp1 8 nhp1 mag1.5% MMS.3% MMS nhp1 nhp1 ino8 9 ino8 9 % Viability 4.5% MMS ino8 9 ino8 9 2.5.1.15 % MMS c d nhp1 nhp1 nhp1 nhp1 nhp1 nhp1 Control (YPD) γ IR (1 gy)
a Drug concentration b 1.3% MMS nhp1 nhp1 8 nhp1 mag1.5% MMS.3% MMS nhp1 nhp1 ino8 9 ino8 9 % Viability 4.5% MMS ino8 9 ino8 9 2.5.1.15 % MMS c d nhp1 nhp1 nhp1 nhp1 nhp1 nhp1 Control (YPD) γ IR (1 gy)
Detection and Restoration of Hybridization Problems in Affymetrix GeneChip Data by Parametric Scanning
 100 Genome Informatics 17(2): 100-109 (2006) Detection and Restoration of Hybridization Problems in Affymetrix GeneChip Data by Parametric Scanning Tomokazu Konishi konishi@akita-pu.ac.jp Faculty of Bioresource
100 Genome Informatics 17(2): 100-109 (2006) Detection and Restoration of Hybridization Problems in Affymetrix GeneChip Data by Parametric Scanning Tomokazu Konishi konishi@akita-pu.ac.jp Faculty of Bioresource
Microarray Technique. Some background. M. Nath
 Microarray Technique Some background M. Nath Outline Introduction Spotting Array Technique GeneChip Technique Data analysis Applications Conclusion Now Blind Guess? Functional Pathway Microarray Technique
Microarray Technique Some background M. Nath Outline Introduction Spotting Array Technique GeneChip Technique Data analysis Applications Conclusion Now Blind Guess? Functional Pathway Microarray Technique
A learned comparative expression measure for Affymetrix GeneChip DNA microarrays
 Proceedings of the Computational Systems Bioinformatics Conference, August 8-11, 2005, Stanford, CA. pp. 144-154. A learned comparative expression measure for Affymetrix GeneChip DNA microarrays Will Sheffler
Proceedings of the Computational Systems Bioinformatics Conference, August 8-11, 2005, Stanford, CA. pp. 144-154. A learned comparative expression measure for Affymetrix GeneChip DNA microarrays Will Sheffler
Tech Note. Using the ncounter Analysis System with FFPE Samples for Gene Expression Analysis. ncounter Gene Expression. Molecules That Count
 ncounter Gene Expression Tech Note Using the ncounter Analysis System with FFPE Samples for Gene Expression Analysis Introduction For the past several decades, pathologists have kept samples obtained from
ncounter Gene Expression Tech Note Using the ncounter Analysis System with FFPE Samples for Gene Expression Analysis Introduction For the past several decades, pathologists have kept samples obtained from
Use of DNA microarrays, wherein the expression levels of
 Modeling of DNA microarray data by using physical properties of hybridization G. A. Held*, G. Grinstein, and Y. Tu IBM Thomas J. Watson Research Center, Yorktown Heights, NY 10598 Communicated by Charles
Modeling of DNA microarray data by using physical properties of hybridization G. A. Held*, G. Grinstein, and Y. Tu IBM Thomas J. Watson Research Center, Yorktown Heights, NY 10598 Communicated by Charles
Description of Logit-t: Detecting Differentially Expressed Genes Using Probe-Level Data
 Description of Logit-t: Detecting Differentially Expressed Genes Using Probe-Level Data Tobias Guennel October 22, 2008 Contents 1 Introduction 2 2 What s new in this version 3 3 Preparing data for use
Description of Logit-t: Detecting Differentially Expressed Genes Using Probe-Level Data Tobias Guennel October 22, 2008 Contents 1 Introduction 2 2 What s new in this version 3 3 Preparing data for use
SIMS2003. Instructors:Rus Yukhananov, Alex Loguinov BWH, Harvard Medical School. Introduction to Microarray Technology.
 SIMS2003 Instructors:Rus Yukhananov, Alex Loguinov BWH, Harvard Medical School Introduction to Microarray Technology. Lecture 1 I. EXPERIMENTAL DETAILS II. ARRAY CONSTRUCTION III. IMAGE ANALYSIS Lecture
SIMS2003 Instructors:Rus Yukhananov, Alex Loguinov BWH, Harvard Medical School Introduction to Microarray Technology. Lecture 1 I. EXPERIMENTAL DETAILS II. ARRAY CONSTRUCTION III. IMAGE ANALYSIS Lecture
Recent technology allow production of microarrays composed of 70-mers (essentially a hybrid of the two techniques)
 Microarrays and Transcript Profiling Gene expression patterns are traditionally studied using Northern blots (DNA-RNA hybridization assays). This approach involves separation of total or polya + RNA on
Microarrays and Transcript Profiling Gene expression patterns are traditionally studied using Northern blots (DNA-RNA hybridization assays). This approach involves separation of total or polya + RNA on
Roche Molecular Biochemicals Technical Note No. LC 10/2000
 Roche Molecular Biochemicals Technical Note No. LC 10/2000 LightCycler Overview of LightCycler Quantification Methods 1. General Introduction Introduction Content Definitions This Technical Note will introduce
Roche Molecular Biochemicals Technical Note No. LC 10/2000 LightCycler Overview of LightCycler Quantification Methods 1. General Introduction Introduction Content Definitions This Technical Note will introduce
6. GENE EXPRESSION ANALYSIS MICROARRAYS
 6. GENE EXPRESSION ANALYSIS MICROARRAYS BIOINFORMATICS COURSE MTAT.03.239 16.10.2013 GENE EXPRESSION ANALYSIS MICROARRAYS Slides adapted from Konstantin Tretyakov s 2011/2012 and Priit Adlers 2010/2011
6. GENE EXPRESSION ANALYSIS MICROARRAYS BIOINFORMATICS COURSE MTAT.03.239 16.10.2013 GENE EXPRESSION ANALYSIS MICROARRAYS Slides adapted from Konstantin Tretyakov s 2011/2012 and Priit Adlers 2010/2011
Real-Time PCR Workshop Gene Expression. Applications Absolute and Relative Quantitation
 Real-Time PCR Workshop Gene Expression Applications Absolute and Relative Quantitation Absolute Quantitation Easy to understand the data, difficult to develop/qualify the standards Relative Quantitation
Real-Time PCR Workshop Gene Expression Applications Absolute and Relative Quantitation Absolute Quantitation Easy to understand the data, difficult to develop/qualify the standards Relative Quantitation
FEATURE-LEVEL EXPLORATION OF THE CHOE ET AL. AFFYMETRIX GENECHIP CONTROL DATASET
 Johns Hopkins University, Dept. of Biostatistics Working Papers 3-17-2006 FEATURE-LEVEL EXPLORATION OF THE CHOE ET AL. AFFYETRIX GENECHIP CONTROL DATASET Rafael A. Irizarry Johns Hopkins Bloomberg School
Johns Hopkins University, Dept. of Biostatistics Working Papers 3-17-2006 FEATURE-LEVEL EXPLORATION OF THE CHOE ET AL. AFFYETRIX GENECHIP CONTROL DATASET Rafael A. Irizarry Johns Hopkins Bloomberg School
Parameter Estimation for the Exponential-Normal Convolution Model
 Parameter Estimation for the Exponential-Normal Convolution Model Monnie McGee & Zhongxue Chen cgee@smu.edu, zhongxue@smu.edu. Department of Statistical Science Southern Methodist University ENAR Spring
Parameter Estimation for the Exponential-Normal Convolution Model Monnie McGee & Zhongxue Chen cgee@smu.edu, zhongxue@smu.edu. Department of Statistical Science Southern Methodist University ENAR Spring
Quality Measures for CytoChip Microarrays
 Quality Measures for CytoChip Microarrays How to evaluate CytoChip Oligo data quality in BlueFuse Multi software. Data quality is one of the most important aspects of any microarray experiment. This technical
Quality Measures for CytoChip Microarrays How to evaluate CytoChip Oligo data quality in BlueFuse Multi software. Data quality is one of the most important aspects of any microarray experiment. This technical
Outline. Array platform considerations: Comparison between the technologies available in microarrays
 Microarray overview Outline Array platform considerations: Comparison between the technologies available in microarrays Differences in array fabrication Differences in array organization Applications of
Microarray overview Outline Array platform considerations: Comparison between the technologies available in microarrays Differences in array fabrication Differences in array organization Applications of
Validation Study of FUJIFILM QuickGene System for Affymetrix GeneChip
 Validation Study of FUJIFILM QuickGene System for Affymetrix GeneChip Reproducibility of Extraction of Genomic DNA from Whole Blood samples in EDTA using FUJIFILM membrane technology on the QuickGene-810
Validation Study of FUJIFILM QuickGene System for Affymetrix GeneChip Reproducibility of Extraction of Genomic DNA from Whole Blood samples in EDTA using FUJIFILM membrane technology on the QuickGene-810
CS 5984: Application of Basic Clustering Algorithms to Find Expression Modules in Cancer
 CS 5984: Application of Basic Clustering Algorithms to Find Expression Modules in Cancer T. M. Murali January 31, 2006 Innovative Application of Hierarchical Clustering A module map showing conditional
CS 5984: Application of Basic Clustering Algorithms to Find Expression Modules in Cancer T. M. Murali January 31, 2006 Innovative Application of Hierarchical Clustering A module map showing conditional
RNA spike-in controls & analysis methods for trustworthy genome-scale measurements
 RNA spike-in controls & analysis methods for trustworthy genome-scale measurements Sarah A. Munro, Ph.D. Genome-Scale Measurements Group ABRF Meeting March 29, 2015 Overview External RNA Controls Consortium
RNA spike-in controls & analysis methods for trustworthy genome-scale measurements Sarah A. Munro, Ph.D. Genome-Scale Measurements Group ABRF Meeting March 29, 2015 Overview External RNA Controls Consortium
Lecture #1. Introduction to microarray technology
 Lecture #1 Introduction to microarray technology Outline General purpose Microarray assay concept Basic microarray experimental process cdna/two channel arrays Oligonucleotide arrays Exon arrays Comparing
Lecture #1 Introduction to microarray technology Outline General purpose Microarray assay concept Basic microarray experimental process cdna/two channel arrays Oligonucleotide arrays Exon arrays Comparing
Analysis of Microarray Data
 Analysis of Microarray Data Lecture 3: Visualization and Functional Analysis George Bell, Ph.D. Senior Bioinformatics Scientist Bioinformatics and Research Computing Whitehead Institute Outline Review
Analysis of Microarray Data Lecture 3: Visualization and Functional Analysis George Bell, Ph.D. Senior Bioinformatics Scientist Bioinformatics and Research Computing Whitehead Institute Outline Review
Estimating Cell Cycle Phase Distribution of Yeast from Time Series Gene Expression Data
 2011 International Conference on Information and Electronics Engineering IPCSIT vol.6 (2011) (2011) IACSIT Press, Singapore Estimating Cell Cycle Phase Distribution of Yeast from Time Series Gene Expression
2011 International Conference on Information and Electronics Engineering IPCSIT vol.6 (2011) (2011) IACSIT Press, Singapore Estimating Cell Cycle Phase Distribution of Yeast from Time Series Gene Expression
reverse transcription! RT 1! RT 2! RT 3!
 Supplementary Figure! Entire workflow repeated 3 times! mirna! stock! -fold dilution series! to yield - copies! per µl in PCR reaction! reverse transcription! RT! pre-pcr! mastermix!!!! 3!!! 3! ddpcr!
Supplementary Figure! Entire workflow repeated 3 times! mirna! stock! -fold dilution series! to yield - copies! per µl in PCR reaction! reverse transcription! RT! pre-pcr! mastermix!!!! 3!!! 3! ddpcr!
Data Analysis on the ABI PRISM 7700 Sequence Detection System: Setting Baselines and Thresholds. Overview. Data Analysis Tutorial
 Data Analysis on the ABI PRISM 7700 Sequence Detection System: Setting Baselines and Thresholds Overview In order for accuracy and precision to be optimal, the assay must be properly evaluated and a few
Data Analysis on the ABI PRISM 7700 Sequence Detection System: Setting Baselines and Thresholds Overview In order for accuracy and precision to be optimal, the assay must be properly evaluated and a few
Comparison of Affymetrix GeneChip Expression Measures
 Johns Hopkins University, Dept. of Biostatistics Working Papers 9-1-2005 Comparison of Affymetrix GeneChip Expression Measures Rafael A. Irizarry Johns Hopkins Bloomberg School of Public Health, Department
Johns Hopkins University, Dept. of Biostatistics Working Papers 9-1-2005 Comparison of Affymetrix GeneChip Expression Measures Rafael A. Irizarry Johns Hopkins Bloomberg School of Public Health, Department
Agilent RNA Spike-In Kit
 Agilent RNA Spike-In Kit Agilent RNA Spike-In Kit Product Number 5188-5279 Introduction The Agilent RNA Spike-In Kit was developed to provide positive controls for monitoring the microarray workflow from
Agilent RNA Spike-In Kit Agilent RNA Spike-In Kit Product Number 5188-5279 Introduction The Agilent RNA Spike-In Kit was developed to provide positive controls for monitoring the microarray workflow from
Introduction to Bioinformatics and Gene Expression Technologies
 Introduction to Bioinformatics and Gene Expression Technologies Utah State University Fall 2017 Statistical Bioinformatics (Biomedical Big Data) Notes 1 1 Vocabulary Gene: hereditary DNA sequence at a
Introduction to Bioinformatics and Gene Expression Technologies Utah State University Fall 2017 Statistical Bioinformatics (Biomedical Big Data) Notes 1 1 Vocabulary Gene: hereditary DNA sequence at a
Technical Review. Real time PCR
 Technical Review Real time PCR Normal PCR: Analyze with agarose gel Normal PCR vs Real time PCR Real-time PCR, also known as quantitative PCR (qpcr) or kinetic PCR Key feature: Used to amplify and simultaneously
Technical Review Real time PCR Normal PCR: Analyze with agarose gel Normal PCR vs Real time PCR Real-time PCR, also known as quantitative PCR (qpcr) or kinetic PCR Key feature: Used to amplify and simultaneously
Calculation of Spot Reliability Evaluation Scores (SRED) for DNA Microarray Data
 Protocol Calculation of Spot Reliability Evaluation Scores (SRED) for DNA Microarray Data Kazuro Shimokawa, Rimantas Kodzius, Yonehiro Matsumura, and Yoshihide Hayashizaki This protocol was adapted from
Protocol Calculation of Spot Reliability Evaluation Scores (SRED) for DNA Microarray Data Kazuro Shimokawa, Rimantas Kodzius, Yonehiro Matsumura, and Yoshihide Hayashizaki This protocol was adapted from
MulCom: a Multiple Comparison statistical test for microarray data in Bioconductor.
 MulCom: a Multiple Comparison statistical test for microarray data in Bioconductor. Claudio Isella, Tommaso Renzulli, Davide Corà and Enzo Medico May 3, 2016 Abstract Many microarray experiments compare
MulCom: a Multiple Comparison statistical test for microarray data in Bioconductor. Claudio Isella, Tommaso Renzulli, Davide Corà and Enzo Medico May 3, 2016 Abstract Many microarray experiments compare
3. human genomics clone genes associated with genetic disorders. 4. many projects generate ordered clones that cover genome
 Lectures 30 and 31 Genome analysis I. Genome analysis A. two general areas 1. structural 2. functional B. genome projects a status report 1. 1 st sequenced: several viral genomes 2. mitochondria and chloroplasts
Lectures 30 and 31 Genome analysis I. Genome analysis A. two general areas 1. structural 2. functional B. genome projects a status report 1. 1 st sequenced: several viral genomes 2. mitochondria and chloroplasts
Introduction to Bioinformatics and Gene Expression Technology
 Vocabulary Introduction to Bioinformatics and Gene Expression Technology Utah State University Spring 2014 STAT 5570: Statistical Bioinformatics Notes 1.1 Gene: Genetics: Genome: Genomics: hereditary DNA
Vocabulary Introduction to Bioinformatics and Gene Expression Technology Utah State University Spring 2014 STAT 5570: Statistical Bioinformatics Notes 1.1 Gene: Genetics: Genome: Genomics: hereditary DNA
Bioinformatics and Genomics: A New SP Frontier?
 Bioinformatics and Genomics: A New SP Frontier? A. O. Hero University of Michigan - Ann Arbor http://www.eecs.umich.edu/ hero Collaborators: G. Fleury, ESE - Paris S. Yoshida, A. Swaroop UM - Ann Arbor
Bioinformatics and Genomics: A New SP Frontier? A. O. Hero University of Michigan - Ann Arbor http://www.eecs.umich.edu/ hero Collaborators: G. Fleury, ESE - Paris S. Yoshida, A. Swaroop UM - Ann Arbor
Intro to Microarray Analysis. Courtesy of Professor Dan Nettleton Iowa State University (with some edits)
 Intro to Microarray Analysis Courtesy of Professor Dan Nettleton Iowa State University (with some edits) Some Basic Biology Genes are DNA sequences that code for proteins. (e.g. gene lengths perhaps 1000
Intro to Microarray Analysis Courtesy of Professor Dan Nettleton Iowa State University (with some edits) Some Basic Biology Genes are DNA sequences that code for proteins. (e.g. gene lengths perhaps 1000
Bayesian Variable Selection and Data Integration for Biological Regulatory Networks
 Bayesian Variable Selection and Data Integration for Biological Regulatory Networks Shane T. Jensen Department of Statistics The Wharton School, University of Pennsylvania stjensen@wharton.upenn.edu Gary
Bayesian Variable Selection and Data Integration for Biological Regulatory Networks Shane T. Jensen Department of Statistics The Wharton School, University of Pennsylvania stjensen@wharton.upenn.edu Gary
DNA Microarray Data Analysis and Mining: Affymetrix Software Package and In-House Complementary Packages
 University of New Orleans ScholarWorks@UNO University of New Orleans Theses and Dissertations Dissertations and Theses 12-19-2003 DNA Microarray Data Analysis and Mining: Affymetrix Software Package and
University of New Orleans ScholarWorks@UNO University of New Orleans Theses and Dissertations Dissertations and Theses 12-19-2003 DNA Microarray Data Analysis and Mining: Affymetrix Software Package and
Introduction to Bioinformatics. Fabian Hoti 6.10.
 Introduction to Bioinformatics Fabian Hoti 6.10. Analysis of Microarray Data Introduction Different types of microarrays Experiment Design Data Normalization Feature selection/extraction Clustering Introduction
Introduction to Bioinformatics Fabian Hoti 6.10. Analysis of Microarray Data Introduction Different types of microarrays Experiment Design Data Normalization Feature selection/extraction Clustering Introduction
less sensitive than RNA-seq but more robust analysis pipelines expensive but quantitiatve standard but typically not high throughput
 Chapter 11: Gene Expression The availability of an annotated genome sequence enables massively parallel analysis of gene expression. The expression of all genes in an organism can be measured in one experiment.
Chapter 11: Gene Expression The availability of an annotated genome sequence enables massively parallel analysis of gene expression. The expression of all genes in an organism can be measured in one experiment.
3.1.4 DNA Microarray Technology
 3.1.4 DNA Microarray Technology Scientists have discovered that one of the differences between healthy and cancer is which genes are turned on in each. Scientists can compare the gene expression patterns
3.1.4 DNA Microarray Technology Scientists have discovered that one of the differences between healthy and cancer is which genes are turned on in each. Scientists can compare the gene expression patterns
Using Low Input of Poly (A) + RNA and Total RNA for Oligonucleotide Microarrays Application
 Using Low Input of Poly (A) + RNA and Total RNA for Oligonucleotide Microarrays Application Gene Expression Author Michelle M. Chen Agilent Technologies, Inc. 3500 Deer Creek Road, MS 25U-7 Palo Alto,
Using Low Input of Poly (A) + RNA and Total RNA for Oligonucleotide Microarrays Application Gene Expression Author Michelle M. Chen Agilent Technologies, Inc. 3500 Deer Creek Road, MS 25U-7 Palo Alto,
Reliability Engineering. RIT Software Engineering
 Reliability Engineering Reliability Engineering Practices Define reliability objectives Use operational profiles to guide test execution Preferable to create automated test infrastructure Track failures
Reliability Engineering Reliability Engineering Practices Define reliability objectives Use operational profiles to guide test execution Preferable to create automated test infrastructure Track failures
Designing Complex Omics Experiments
 Designing Complex Omics Experiments Xiangqin Cui Section on Statistical Genetics Department of Biostatistics University of Alabama at Birmingham 6/15/2015 Some slides are from previous lectures given by
Designing Complex Omics Experiments Xiangqin Cui Section on Statistical Genetics Department of Biostatistics University of Alabama at Birmingham 6/15/2015 Some slides are from previous lectures given by
Chapter 5 Evaluating Classification & Predictive Performance
 Chapter 5 Evaluating Classification & Predictive Performance Data Mining for Business Intelligence Shmueli, Patel & Bruce Galit Shmueli and Peter Bruce 2010 Why Evaluate? Multiple methods are available
Chapter 5 Evaluating Classification & Predictive Performance Data Mining for Business Intelligence Shmueli, Patel & Bruce Galit Shmueli and Peter Bruce 2010 Why Evaluate? Multiple methods are available
Chromosome Analysis Suite 3.0 (ChAS 3.0)
 Chromosome Analysis Suite 3.0 (ChAS 3.0) FAQs related to CytoScan Cytogenetics Suite 1. What is required for processing and viewing CytoScan array CEL files? CytoScan array CEL files are processed and
Chromosome Analysis Suite 3.0 (ChAS 3.0) FAQs related to CytoScan Cytogenetics Suite 1. What is required for processing and viewing CytoScan array CEL files? CytoScan array CEL files are processed and
A comprehensive assessment of RNA-seq accuracy, reproducibility and information content by the Sequencing Quality Control Consortium
 A comprehensive assessment of RNA-seq accuracy, reproducibility and information content by the Sequencing Quality Control Consortium SEQC/MAQC-III Consortium* 214 Nature America, Inc. All rights reserved.
A comprehensive assessment of RNA-seq accuracy, reproducibility and information content by the Sequencing Quality Control Consortium SEQC/MAQC-III Consortium* 214 Nature America, Inc. All rights reserved.
Systematic comparison of CRISPR/Cas9 and RNAi screens for essential genes
 CORRECTION NOTICE Nat. Biotechnol. doi:10.1038/nbt. 3567 Systematic comparison of CRISPR/Cas9 and RNAi screens for essential genes David W Morgens, Richard M Deans, Amy Li & Michael C Bassik In the version
CORRECTION NOTICE Nat. Biotechnol. doi:10.1038/nbt. 3567 Systematic comparison of CRISPR/Cas9 and RNAi screens for essential genes David W Morgens, Richard M Deans, Amy Li & Michael C Bassik In the version
Supporting Online Material for
 www.sciencemag.org/cgi/content/full/313/5795/1929/dc1 Supporting Online Material for The Connectivity Map: Using Gene-Expression Signatures to Connect Small Molecules, Genes, and Disease Justin Lamb, *
www.sciencemag.org/cgi/content/full/313/5795/1929/dc1 Supporting Online Material for The Connectivity Map: Using Gene-Expression Signatures to Connect Small Molecules, Genes, and Disease Justin Lamb, *
Gene Expression Technology
 Gene Expression Technology Bing Zhang Department of Biomedical Informatics Vanderbilt University bing.zhang@vanderbilt.edu Gene expression Gene expression is the process by which information from a gene
Gene Expression Technology Bing Zhang Department of Biomedical Informatics Vanderbilt University bing.zhang@vanderbilt.edu Gene expression Gene expression is the process by which information from a gene
O C. 5 th C. 3 rd C. the national health museum
 Elements of Molecular Biology Cells Cells is a basic unit of all living organisms. It stores all information to replicate itself Nucleus, chromosomes, genes, All living things are made of cells Prokaryote,
Elements of Molecular Biology Cells Cells is a basic unit of all living organisms. It stores all information to replicate itself Nucleus, chromosomes, genes, All living things are made of cells Prokaryote,
Chromosome-scale scaffolding of de novo genome assemblies based on chromatin interactions. Supplementary Material
 Chromosome-scale scaffolding of de novo genome assemblies based on chromatin interactions Joshua N. Burton 1, Andrew Adey 1, Rupali P. Patwardhan 1, Ruolan Qiu 1, Jacob O. Kitzman 1, Jay Shendure 1 1 Department
Chromosome-scale scaffolding of de novo genome assemblies based on chromatin interactions Joshua N. Burton 1, Andrew Adey 1, Rupali P. Patwardhan 1, Ruolan Qiu 1, Jacob O. Kitzman 1, Jay Shendure 1 1 Department
Introduction to BioMEMS & Medical Microdevices DNA Microarrays and Lab-on-a-Chip Methods
 Introduction to BioMEMS & Medical Microdevices DNA Microarrays and Lab-on-a-Chip Methods Companion lecture to the textbook: Fundamentals of BioMEMS and Medical Microdevices, by Prof., http://saliterman.umn.edu/
Introduction to BioMEMS & Medical Microdevices DNA Microarrays and Lab-on-a-Chip Methods Companion lecture to the textbook: Fundamentals of BioMEMS and Medical Microdevices, by Prof., http://saliterman.umn.edu/
Quantitative Real time PCR. Only for teaching purposes - not for reproduction or sale
 Quantitative Real time PCR PCR reaction conventional versus real time PCR real time PCR principles threshold cycle C T efficiency relative quantification reference genes primers detection chemistry GLP
Quantitative Real time PCR PCR reaction conventional versus real time PCR real time PCR principles threshold cycle C T efficiency relative quantification reference genes primers detection chemistry GLP
Exploration, normalization, and summaries of high density oligonucleotide array probe level data
 Biostatistics (2003), 4, 2,pp. 249 264 Printed in Great Britain Exploration, normalization, and summaries of high density oligonucleotide array probe level data RAFAEL A. IRIZARRY Department of Biostatistics,
Biostatistics (2003), 4, 2,pp. 249 264 Printed in Great Britain Exploration, normalization, and summaries of high density oligonucleotide array probe level data RAFAEL A. IRIZARRY Department of Biostatistics,
Spreadsheets in Education (ejsie)
 Spreadsheets in Education (ejsie) Volume 2, Issue 2 2005 Article 5 Forecasting with Excel: Suggestions for Managers Scott Nadler John F. Kros East Carolina University, nadlers@mail.ecu.edu East Carolina
Spreadsheets in Education (ejsie) Volume 2, Issue 2 2005 Article 5 Forecasting with Excel: Suggestions for Managers Scott Nadler John F. Kros East Carolina University, nadlers@mail.ecu.edu East Carolina
affy: Built-in Processing Methods
 affy: Built-in Processing Methods Ben Bolstad October 30, 2017 Contents 1 Introduction 2 2 Background methods 2 2.1 none...................................... 2 2.2 rma/rma2...................................
affy: Built-in Processing Methods Ben Bolstad October 30, 2017 Contents 1 Introduction 2 2 Background methods 2 2.1 none...................................... 2 2.2 rma/rma2...................................
ChIP-Seq Data Analysis. J Fass UCD Genome Center Bioinformatics Core Wednesday 15 June 2015
 ChIP-Seq Data Analysis J Fass UCD Genome Center Bioinformatics Core Wednesday 15 June 2015 What s the Question? Where do Transcription Factors (TFs) bind genomic DNA 1? (Where do other things bind DNA
ChIP-Seq Data Analysis J Fass UCD Genome Center Bioinformatics Core Wednesday 15 June 2015 What s the Question? Where do Transcription Factors (TFs) bind genomic DNA 1? (Where do other things bind DNA
Please purchase PDFcamp Printer on to remove this watermark. DNA microarray
 DNA microarray Example of an approximately 40,000 probe spotted oligo microarray with enlarged inset to show detail. A DNA microarray is a multiplex technology used in molecular biology. It consists of
DNA microarray Example of an approximately 40,000 probe spotted oligo microarray with enlarged inset to show detail. A DNA microarray is a multiplex technology used in molecular biology. It consists of
Gene Expression Data Analysis (I)
 Gene Expression Data Analysis (I) Bing Zhang Department of Biomedical Informatics Vanderbilt University bing.zhang@vanderbilt.edu Bioinformatics tasks Biological question Experiment design Microarray experiment
Gene Expression Data Analysis (I) Bing Zhang Department of Biomedical Informatics Vanderbilt University bing.zhang@vanderbilt.edu Bioinformatics tasks Biological question Experiment design Microarray experiment
Gene Expression Profiling and Validation Using Agilent SurePrint G3 Gene Expression Arrays
 Gene Expression Profiling and Validation Using Agilent SurePrint G3 Gene Expression Arrays Application Note Authors Bahram Arezi, Nilanjan Guha and Anne Bergstrom Lucas Agilent Technologies Inc. Santa
Gene Expression Profiling and Validation Using Agilent SurePrint G3 Gene Expression Arrays Application Note Authors Bahram Arezi, Nilanjan Guha and Anne Bergstrom Lucas Agilent Technologies Inc. Santa
MICROARRAYS+SEQUENCING
 MICROARRAYS+SEQUENCING The most efficient way to advance genomics research Down to a Science. www.affymetrix.com/downtoascience Affymetrix GeneChip Expression Technology Complementing your Next-Generation
MICROARRAYS+SEQUENCING The most efficient way to advance genomics research Down to a Science. www.affymetrix.com/downtoascience Affymetrix GeneChip Expression Technology Complementing your Next-Generation
DNA Microarray Technology
 CHAPTER 1 DNA Microarray Technology All living organisms are composed of cells. As a functional unit, each cell can make copies of itself, and this process depends on a proper replication of the genetic
CHAPTER 1 DNA Microarray Technology All living organisms are composed of cells. As a functional unit, each cell can make copies of itself, and this process depends on a proper replication of the genetic
Inherent variation in the reactions, type of enzymes used. Depends on the type of labeling and procedures, as well as the age of the labels.
 332 Experimental design, analysis of variance and slide quality assessment in gene expression arrays Sorin Draghici*, Alexander Kuklin, Bruce Hoff & Soheil Shams Address BioDiscovery Inc 11150 West Olympic
332 Experimental design, analysis of variance and slide quality assessment in gene expression arrays Sorin Draghici*, Alexander Kuklin, Bruce Hoff & Soheil Shams Address BioDiscovery Inc 11150 West Olympic
SPH 247 Statistical Analysis of Laboratory Data
 SPH 247 Statistical Analysis of Laboratory Data April 14, 2015 SPH 247 Statistical Analysis of Laboratory Data 1 Basic Design of Expression Arrays For each gene that is a target for the array, we have
SPH 247 Statistical Analysis of Laboratory Data April 14, 2015 SPH 247 Statistical Analysis of Laboratory Data 1 Basic Design of Expression Arrays For each gene that is a target for the array, we have
RNA-Sequencing analysis
 RNA-Sequencing analysis Markus Kreuz 25. 04. 2012 Institut für Medizinische Informatik, Statistik und Epidemiologie Content: Biological background Overview transcriptomics RNA-Seq RNA-Seq technology Challenges
RNA-Sequencing analysis Markus Kreuz 25. 04. 2012 Institut für Medizinische Informatik, Statistik und Epidemiologie Content: Biological background Overview transcriptomics RNA-Seq RNA-Seq technology Challenges
Same TaqMan Assay quality, new format
 Product Bulletin TaqMan Array Plates TaqMan Array Plates TaqMan Gene Expression Assays delivered in ready-to-use 96-well Custom Array Plates or predefined Gene Signature Plates Flexible choose from customizable
Product Bulletin TaqMan Array Plates TaqMan Array Plates TaqMan Gene Expression Assays delivered in ready-to-use 96-well Custom Array Plates or predefined Gene Signature Plates Flexible choose from customizable
Bioinformatics III Structural Bioinformatics and Genome Analysis. PART II: Genome Analysis. Chapter 7. DNA Microarrays
 Bioinformatics III Structural Bioinformatics and Genome Analysis PART II: Genome Analysis Chapter 7. DNA Microarrays 7.1 Motivation 7.2 DNA Microarray History and current states 7.3 DNA Microarray Techniques
Bioinformatics III Structural Bioinformatics and Genome Analysis PART II: Genome Analysis Chapter 7. DNA Microarrays 7.1 Motivation 7.2 DNA Microarray History and current states 7.3 DNA Microarray Techniques
Methods of Biomaterials Testing Lesson 3-5. Biochemical Methods - Molecular Biology -
 Methods of Biomaterials Testing Lesson 3-5 Biochemical Methods - Molecular Biology - Chromosomes in the Cell Nucleus DNA in the Chromosome Deoxyribonucleic Acid (DNA) DNA has double-helix structure The
Methods of Biomaterials Testing Lesson 3-5 Biochemical Methods - Molecular Biology - Chromosomes in the Cell Nucleus DNA in the Chromosome Deoxyribonucleic Acid (DNA) DNA has double-helix structure The
Oligonucleotide microarray data are not normally distributed
 Oligonucleotide microarray data are not normally distributed Johanna Hardin Jason Wilson John Kloke Abstract Novel techniques for analyzing microarray data are constantly being developed. Though many of
Oligonucleotide microarray data are not normally distributed Johanna Hardin Jason Wilson John Kloke Abstract Novel techniques for analyzing microarray data are constantly being developed. Though many of
PeakSeq enables systematic scoring of ChIP-seq experiments relative to controls
 PeakSeq enables systematic scoring of ChIP-seq experiments relative to controls Joel Rozowsky, Ghia Euskirchen 2, Raymond K Auerbach 3, Zhengdong D Zhang, Theodore Gibson, Robert Bjornson 4, Nicholas Carriero
PeakSeq enables systematic scoring of ChIP-seq experiments relative to controls Joel Rozowsky, Ghia Euskirchen 2, Raymond K Auerbach 3, Zhengdong D Zhang, Theodore Gibson, Robert Bjornson 4, Nicholas Carriero
RNA-Seq analysis using R: Differential expression and transcriptome assembly
 RNA-Seq analysis using R: Differential expression and transcriptome assembly Beibei Chen Ph.D BICF 12/7/2016 Agenda Brief about RNA-seq and experiment design Gene oriented analysis Gene quantification
RNA-Seq analysis using R: Differential expression and transcriptome assembly Beibei Chen Ph.D BICF 12/7/2016 Agenda Brief about RNA-seq and experiment design Gene oriented analysis Gene quantification
Phosphate buffered saline (PBS) for washing the cells TE buffer (nuclease-free) ph 7.5 for resuspending the SingleShot RNA control template
 Catalog # Description 172-5085 SingleShot SYBR Green Kit, 100 x 50 µl reactions For research purposes only. Introduction The SingleShot SYBR Green Kit prepares genomic DNA (gdna) free RNA directly from
Catalog # Description 172-5085 SingleShot SYBR Green Kit, 100 x 50 µl reactions For research purposes only. Introduction The SingleShot SYBR Green Kit prepares genomic DNA (gdna) free RNA directly from
