PROTÉOMIQUE. Application de la protéomique à la cartographie de l interactome et la découverte de biomarqueurs. BENOIT COULOMBE, PhD
|
|
|
- James Porter
- 6 years ago
- Views:
Transcription
1 PROTÉOMIQUE Application de la protéomique à la cartographie de l interactome et la découverte de biomarqueurs BENOIT COULOMBE, PhD Laboratoire de protéomique & transcription génique BCM2003, mars 2012
2 Biomarkers and personalized medicine Early diagnosis and prognosis of disease Monitoring the response of patients to treatment - Known therapies - Development of new drugs Stratification of patient cohorts in order to provide personalized treatments for specific disease phenotypes (ex: IRESSA and lung cancer) Process of biomarker discovery can also identify new targets for drug discovery, and develop new knowledge on mechanisms of disease
3 The sequencing of the human genome has provided the list and linear sequence of all (most) human proteins Genome Proteins
4 Proteins rarely work alone, but rather assemble into protein complexes that integrate the function of multiple gene products Genome Proteins Complexes Machineries Function
5 Large-scale genomic studies identify an increasing number of genes associated with disease phenotypes Genome
6 Large-scale genomic studies identify an increasing number of genes associated with disease phenotypes Genome How the products of these genes participate in the establishment and the progression of disease is often unknown!
7 The interaction partners of disease-associated proteins can reveal their role in disease (guilt by association) Genome Proteins Complexes Machineries Role in disease
8 The interaction partners of disease-associated proteins are likely to act as modulators of the disease phenotype Genome Proteins Complexes Modulators Machineries Role in disease
9 The disease-associated proteins and their interactors are putative biomarkers and drug targets Genome Proteins Complexes Biomarkers Drug targets Machineries Role in disease
10 1. Laboratory Pipelines The P3MD platform Infrastructure, expertise, and data A. INTERACTOME B. BIOMARKERS Selection of Disease-Associated Proteins (Baits) AP-MS Selection of Peptides (transitions) SRM Cell Lines Candidate Biomarkers Patient Samples 2. Software Applications PROTEUS Candidate Biomarker Selection Engine P3MD Computational tools Computational classifier 3. Data List of Interactions (Network Maps) List of Validated Biomarkers
11 The human proteome
12 Identification of an association between a protein and a disease The case of hypothetical protein D D
13 The protein D interactome D
14 High-quality generic map of the protein D interactome Affinity purification coupled with MS D
15 Components of the protein D interactome are candidate drug targets and biomarkers D
16 Identification of interactors using LC-MS/MS + Determination of a probability (False Discovery Rate or FDR) for each interaction to be valid using computational algorithm trained using machine learning to differentiate valid vs. invalid interactions
17 Components of the protein D interactome are candidate drug targets and biomarkers D
18 Components of the protein D interactome are candidate drug targets and biomarkers D I
19 Components of the protein D interactome are candidate drug targets and biomarkers D I I
20 Preliminary validation of candidate biomarkers Modulations of components of protein D interactome in disease cells (SRM) D Upregulated Isoform Downregulated
21 Selected Reaction Monitoring (SRM, also termed MRM) Protein quantification is possible by spiking known amounts of chemically synthesized heavy-labelled peptides in the samples (standard)
22 Proof of concept
23 Cellular machineries involved in mrna synthesis GENERAL TRANSCRIPTION MACHINERY - RNA polymerase II (RNAP II) - General transcription factors - Elongation factors TFIID Mediator TFIIF RNAP II P-TEFb TFIIF RNAP II TFIIS 5 TATAAA +1 3 TFIIB TFIIE TFIIH Cleavage/polyA factors CHROMATIN-REMODELING MACHINERY - ATP dependent remodeling - Histone-modifying enzymes PRE-mRNA snrnps U1-U6 hnrnps PRE-mRNA PROCESSING MACHINERY Capping enzymes
24 Cellular machineries involved in mrna synthesis GENERAL TRANSCRIPTION MACHINERY - RNA polymerase II (RNAP II) - General transcription factors - Elongation factors TFIID Mediator TFIIF RNAP II P-TEFb TFIIF RNAP II TFIIS 5 TATAAA +1 3 TFIIB TFIIE TFIIH Cleavage/polyA factors CHROMATIN-REMODELING MACHINERY - ATP dependent remodeling - Histone-modifying enzymes PRE-mRNA snrnps U1-U6 hnrnps PRE-mRNA PROCESSING MACHINERY Capping enzymes
25 A platform for the systematic characterization of protein interaction networks in human cells Collection of 293 cell lines engineered to express physiological levels of affinity tagged polypeptides upon induction Soluble fraction (supernatant) Cell Lysis in Mild Conditions Insoluble fraction (pellet) Main Features of the Procedure: 1) Expression at physiological level 2) Cell lysis in mild conditions (soluble) 3) Purification in native conditions 4) Protein identification by sensitive MS 5) Efficient data analysis and validation Protein Identification Computational Analysis (Reliability, Connectivity)
26 A platform for the systematic characterization of protein interaction networks in human cells Human cdna cloned into an ecdysone-inducible expression vector encoding the TAP tag Transfection in EcR-293 cells and selection of expressing clones Induction with ponasterone A to obtain near physiological expression levels Cell lysis in mild conditions Centrifugation Soluble fraction (WCE) Insoluble fraction (pellet) Tandem Affinity Purification SDS gel analysis (high resolution) Protein identification (Orbitrap MS: high sensitivity & mass precision)
27 The ecdysone-inducible mammalian expression system pvgecr pind Modified from Vickers and Sharrocks, 2002 Advantages: very low basal expression ecdysone analogs are inert in mammalian cells high control on expression levels (physiological) Western blot
28 A platform for the systematic characterization of protein interaction networks in human cells Human cdna cloned into an ecdysone-inducible expression vector encoding the TAP tag Transfection in EcR-293 cells and selection of expressing clones Induction with ponasterone A to obtain near physiological expression levels Cell lysis in mild conditions Centrifugation Soluble fraction (WCE) Insoluble fraction (pellet) Tandem Affinity Purification SDS gel analysis (high resolution) Protein identification (Orbitrap MS: high sensitivity & mass precision)
29 Tandem affinity purification in native conditions Double-affinity tagged protein extract Protein of interest Calmodulin binding peptide (CBP) TEV protease cleavage site Protein A IgG-binding sites IgG sepharose Double-affinity Tag Unbound Bound TEV protease IgG Sepharose Beads Eluate Unbound Calmodulin Beads Bound TEV Beads EGTA Eluate Calmodulin Beads Purified Protein Complexes EGTA
30 A platform for the systematic characterization of protein interaction networks in human cells Human cdna cloned into an ecdysone-inducible expression vector encoding the TAP tag Transfection in EcR-293 cells and selection of expressing clones Induction with ponasterone A to obtain near physiological expression levels Cell lysis in mild conditions Centrifugation Soluble fraction (WCE) Insoluble fraction (pellet) Tandem Affinity Purification SDS gel analysis (high resolution) Protein identification (Orbitrap MS: high sensitivity & mass precision)
31 Identification of proteins by mass spectrometry SDS- PAGE Excised proteins Trypsin digestion Peptides mixture In-gel digestion N-I I Yarmush and Jayaraman, 2002
32 Mass spectrometry Peptide Sequence LC-MS/MS Computer Cluster
33 A platform for the systematic characterization of protein interaction networks in human cells Human cdna cloned into an ecdysone-inducible expression vector encoding the TAP tag Transfection in EcR-293 cells and selection of expressing clones Induction with ponasterone A to obtain near physiological expression levels Cell lysis in mild conditions Centrifugation Soluble fraction (WCE) Insoluble fraction (pellet) Tandem Affinity Purification SDS gel analysis (high resolution) Protein identification (Orbitrap MS: high sensitivity & mass precision)
34 Affinity purification of protein complexes
35 A platform for the systematic characterization of protein interaction networks in human cells Human cdna cloned into an ecdysone-inducible expression vector encoding the TAP tag Transfection in EcR-293 cells and selection of expressing clones Induction with ponasterone A to obtain near physiological expression levels Cell lysis in mild conditions Centrifugation Soluble fraction (WCE) Insoluble fraction (pellet) Tandem Affinity Purification Reciprocal tagging SDS gel analysis (high resolution) Protein identification (Orbitrap MS: high sensitivity & mass precision)
36 Reciprocal tagging allows to validate some interactions and to enrich the data set
37 A platform for the systematic characterization of protein interaction networks in human cells Human cdna cloned into an ecdysone-inducible expression vector encoding the TAP tag Transfection in EcR-293 cells and selection of expressing clones Induction with ponasterone A to obtain near physiological expression levels Cell lysis in mild conditions Centrifugation Soluble fraction (WCE) Insoluble fraction (pellet) Tandem Affinity Purification Reciprocal tagging SDS gel analysis (high resolution) Protein identification (Orbitrap MS: high sensitivity & mass precision) Computational analysis (reliability & connectivity)
38 A platform for the systematic characterization of protein interaction networks in human cells Human cdna cloned into an ecdysone-inducible expression vector encoding the TAP tag Transfection in EcR-293 cells and selection of expressing clones Induction with ponasterone A to obtain near physiological expression levels Cell lysis in mild conditions Centrifugation Soluble fraction (WCE) Insoluble fraction (pellet) Tandem Affinity Purification Reciprocal tagging SDS gel analysis (high resolution) Comprehensive Maps of Human Protein Interaction Networks Protein identification (Orbitrap MS: high sensitivity & mass precision) Computational analysis (reliability & connectivity)
39 The protein interaction database Proteus C. Poitras & D. Bergeron
40 Computational data analysis and validation A B C
41 Computational data analysis and validation keeping 83% of the interactions supported by the literature excluding 83% of the interactions judged likely false-positives
42 Summary of our survey of protein complexes (2007) 805 distinct interactions were selected as high-confidence protein interactions Jeronimo et al (2007) Mol Cell 27, 262
43 High-quality protein interaction network for the RNA polymerase II transcription machinery
44 Proof of concept
45 Myopathies The myopathies are a heterogeneous group of diseases of skeletal muscle They include: Sporadic Inclusion-Body Myositis (sibm) Dermatomyositis Polymyositis Myofibrillar myopathies Neurogenic muscular athrophy IBMPFD And many others
46 The RNA Polymerase II-Associated Proteins (RPAPs) are previously uncharacterized proteins
47 Expression of components of the RNAPII network is specifically modulated in tissues of patients suffering from various muscular diseases Muscle disease N biopsies RPAPx RPAPy RPAPz Normal controls 15 Normal Normal Normal Disease Phenotype A 12 Increased * Normal Normal Disease Phenotype B 8 Increased Increased * Increased Disease Phenotype C 10 Increased Normal Increased Increased expression of RPAPs is the only marker today that permits to discriminate between two specific muscular disorders
48 The RPAPs are dysregulated in muscular diseases
49 Diseases under study Myopathies Leucodystrophies Type 2 diabetes and insulin resistance Hypercholesterolemia Cancer
50 Thank you!
Genome Biology and Biotechnology
 Genome Biology and Biotechnology 10. The proteome Prof. M. Zabeau Department of Plant Systems Biology Flanders Interuniversity Institute for Biotechnology (VIB) University of Gent International course
Genome Biology and Biotechnology 10. The proteome Prof. M. Zabeau Department of Plant Systems Biology Flanders Interuniversity Institute for Biotechnology (VIB) University of Gent International course
However, only a fraction of these genes are transcribed in an individual cell at any given time.
 All cells in an organism contain the same set of genes. However, only a fraction of these genes are transcribed in an individual cell at any given time. It is the pattern of gene expression that determines
All cells in an organism contain the same set of genes. However, only a fraction of these genes are transcribed in an individual cell at any given time. It is the pattern of gene expression that determines
CONTENTS. LIST OF FIGURES... i. LIST OF TABLES... iii. SUMMARY OF THE WORK DONE... vi
 CONTENTS SECTION ~ LIST OF FIGURES... i LIST OF TABLES... iii LIST OF ABBREVIATIONS USED... ~... iv SUMMARY OF THE WORK DONE... vi 1. INTROD UCTION... :.................................... 1 1.1. AN OVERVIEW
CONTENTS SECTION ~ LIST OF FIGURES... i LIST OF TABLES... iii LIST OF ABBREVIATIONS USED... ~... iv SUMMARY OF THE WORK DONE... vi 1. INTROD UCTION... :.................................... 1 1.1. AN OVERVIEW
MBios 478: Mass Spectrometry Applications [Dr. Wyrick] Slide #1. Lecture 25: Mass Spectrometry Applications
![MBios 478: Mass Spectrometry Applications [Dr. Wyrick] Slide #1. Lecture 25: Mass Spectrometry Applications MBios 478: Mass Spectrometry Applications [Dr. Wyrick] Slide #1. Lecture 25: Mass Spectrometry Applications](/thumbs/74/71368870.jpg) MBios 478: Mass Spectrometry Applications [Dr. Wyrick] Slide #1 Lecture 25: Mass Spectrometry Applications Measuring Protein Abundance o ICAT o DIGE Identifying Post-Translational Modifications Protein-protein
MBios 478: Mass Spectrometry Applications [Dr. Wyrick] Slide #1 Lecture 25: Mass Spectrometry Applications Measuring Protein Abundance o ICAT o DIGE Identifying Post-Translational Modifications Protein-protein
Lecture 8: Affinity Chromatography-III
 Lecture 8: Affinity Chromatography-III Key words: Chromatography; Affinity chromatography; Protein Purification During this lecture, we shall be studying few more examples of affinity chromatography. The
Lecture 8: Affinity Chromatography-III Key words: Chromatography; Affinity chromatography; Protein Purification During this lecture, we shall be studying few more examples of affinity chromatography. The
NPTEL VIDEO COURSE PROTEOMICS PROF. SANJEEVA SRIVASTAVA
 LECTURE-26 Interactomics: Yeast two-hybrid immunoprecipitation protein microarrays TRANSCRIPT Welcome to the proteomics course. Today we will talk about interactomics, which is studying interactions of
LECTURE-26 Interactomics: Yeast two-hybrid immunoprecipitation protein microarrays TRANSCRIPT Welcome to the proteomics course. Today we will talk about interactomics, which is studying interactions of
Complex Transcription Machinery
 Complex Transcription Machinery Subunits of the Basal Txn Apparatus Stepwise Assembly of the Pre-initiation Complex Interplay of Activators, Co-regulators and RNA Polymerase at the Promoter Divide and
Complex Transcription Machinery Subunits of the Basal Txn Apparatus Stepwise Assembly of the Pre-initiation Complex Interplay of Activators, Co-regulators and RNA Polymerase at the Promoter Divide and
DISCOVERY AND VALIDATION OF TARGETS AND BIOMARKERS BY MASS SPECTROMETRY-BASED PROTEOMICS. September, 2011
 DISCOVERY AND VALIDATION OF TARGETS AND BIOMARKERS BY MASS SPECTROMETRY-BASED PROTEOMICS September, 2011 1 CAPRION PROTEOMICS Leading proteomics-based service provider - Biomarker and target discovery
DISCOVERY AND VALIDATION OF TARGETS AND BIOMARKERS BY MASS SPECTROMETRY-BASED PROTEOMICS September, 2011 1 CAPRION PROTEOMICS Leading proteomics-based service provider - Biomarker and target discovery
Eukaryotic Transcription
 Eukaryotic Transcription I. Differences between eukaryotic versus prokaryotic transcription. II. (core vs holoenzyme): RNA polymerase II - Promotor elements. - General Pol II transcription factors (GTF).
Eukaryotic Transcription I. Differences between eukaryotic versus prokaryotic transcription. II. (core vs holoenzyme): RNA polymerase II - Promotor elements. - General Pol II transcription factors (GTF).
Experimental Techniques 2
 Experimental Techniques 2 High-throughput interaction detection Yeast two-hybrid - pairwise organisms as machines to learn about organisms yeast, worm, fly, human,... low intersection between repeated
Experimental Techniques 2 High-throughput interaction detection Yeast two-hybrid - pairwise organisms as machines to learn about organisms yeast, worm, fly, human,... low intersection between repeated
Eukaryotic & Prokaryotic Transcription. RNA polymerases
 Eukaryotic & Prokaryotic Transcription RNA polymerases RNA Polymerases A. E. coli RNA polymerase 1. core enzyme = ββ'(α)2 has catalytic activity but cannot recognize start site of transcription ~500,000
Eukaryotic & Prokaryotic Transcription RNA polymerases RNA Polymerases A. E. coli RNA polymerase 1. core enzyme = ββ'(α)2 has catalytic activity but cannot recognize start site of transcription ~500,000
TRANSCRIPTION AND PROCESSING OF RNA
 TRANSCRIPTION AND PROCESSING OF RNA 1. The steps of gene expression. 2. General characterization of transcription: steps, components of transcription apparatus. 3. Transcription of eukaryotic structural
TRANSCRIPTION AND PROCESSING OF RNA 1. The steps of gene expression. 2. General characterization of transcription: steps, components of transcription apparatus. 3. Transcription of eukaryotic structural
30 Gene expression: Transcription
 30 Gene expression: Transcription Gene structure. o Exons coding region of DNA. o Introns non-coding region of DNA. o Introns are interspersed between exons of a single gene. o Promoter region helps enzymes
30 Gene expression: Transcription Gene structure. o Exons coding region of DNA. o Introns non-coding region of DNA. o Introns are interspersed between exons of a single gene. o Promoter region helps enzymes
Expressed genes profiling (Microarrays) Overview Of Gene Expression Control Profiling Of Expressed Genes
 Expressed genes profiling (Microarrays) Overview Of Gene Expression Control Profiling Of Expressed Genes Genes can be regulated at many levels Usually, gene regulation, are referring to transcriptional
Expressed genes profiling (Microarrays) Overview Of Gene Expression Control Profiling Of Expressed Genes Genes can be regulated at many levels Usually, gene regulation, are referring to transcriptional
Practice Exam A. Briefly describe how IL-25 treatment might be able to help this responder subgroup of liver cancer patients.
 Practice Exam 2007 1. A special JAK-STAT signaling system (JAK5-STAT5) was recently identified in which a gene called TS5 becomes selectively transcribed and expressed in the liver upon induction by a
Practice Exam 2007 1. A special JAK-STAT signaling system (JAK5-STAT5) was recently identified in which a gene called TS5 becomes selectively transcribed and expressed in the liver upon induction by a
Quantitative Proteomics: From Technology to Cancer Biology
 Quantitative Proteomics: From Technology to Cancer Biology Beyond the Genetic Prescription Pad: Personalizing Cancer Medicine in 2014 February 10-11, 2014 Thomas Kislinger Molecular Biomarkers in Body
Quantitative Proteomics: From Technology to Cancer Biology Beyond the Genetic Prescription Pad: Personalizing Cancer Medicine in 2014 February 10-11, 2014 Thomas Kislinger Molecular Biomarkers in Body
Lecture 5: 8/31. CHAPTER 5 Techniques in Protein Biochemistry
 Lecture 5: 8/31 CHAPTER 5 Techniques in Protein Biochemistry Chapter 5 Outline The proteome is the entire set of proteins expressed and modified by a cell under a particular set of biochemical conditions.
Lecture 5: 8/31 CHAPTER 5 Techniques in Protein Biochemistry Chapter 5 Outline The proteome is the entire set of proteins expressed and modified by a cell under a particular set of biochemical conditions.
Post-translational modification
 Protein expression Western blotting, is a widely used and accepted technique to detect levels of protein expression in a cell or tissue extract. This technique measures protein levels in a biological sample
Protein expression Western blotting, is a widely used and accepted technique to detect levels of protein expression in a cell or tissue extract. This technique measures protein levels in a biological sample
Motivation From Protein to Gene
 MOLECULAR BIOLOGY 2003-4 Topic B Recombinant DNA -principles and tools Construct a library - what for, how Major techniques +principles Bioinformatics - in brief Chapter 7 (MCB) 1 Motivation From Protein
MOLECULAR BIOLOGY 2003-4 Topic B Recombinant DNA -principles and tools Construct a library - what for, how Major techniques +principles Bioinformatics - in brief Chapter 7 (MCB) 1 Motivation From Protein
Use of a Label-Free Quantitative Platform Based on MS/MS Average TIC to Calculate Dynamics of Protein Complexes in Insulin Signaling
 ABRF 2009 SELECTED PRESENTATIONS Use of a Label-Free Quantitative Platform Based on MS/MS Average TIC to Calculate Dynamics of Protein Complexes in Insulin Signaling Xuemei Yang, 1 Adam Friedman, 2 Shailender
ABRF 2009 SELECTED PRESENTATIONS Use of a Label-Free Quantitative Platform Based on MS/MS Average TIC to Calculate Dynamics of Protein Complexes in Insulin Signaling Xuemei Yang, 1 Adam Friedman, 2 Shailender
Proteomic analysis of biological fluids
 Proteomic analysis of biological fluids Anne Gonzalez, CNRS-IPBS, Toulouse Équipe «Analyse Protéomique et Spectrométrie de Masse des Biomolécules» Odile Schiltz-Bernard Monsarrat Workshop «Protéomique
Proteomic analysis of biological fluids Anne Gonzalez, CNRS-IPBS, Toulouse Équipe «Analyse Protéomique et Spectrométrie de Masse des Biomolécules» Odile Schiltz-Bernard Monsarrat Workshop «Protéomique
Strategies in proteomics
 Strategies in proteomics Systems biology - understand cellpathways, network, and complex interacting (includes Genomics, Proteomics, Metabolomics..) Biological processes - characterize protein complexes,
Strategies in proteomics Systems biology - understand cellpathways, network, and complex interacting (includes Genomics, Proteomics, Metabolomics..) Biological processes - characterize protein complexes,
Proteomics And Cancer Biomarker Discovery. Dr. Zahid Khan Institute of chemical Sciences (ICS) University of Peshawar. Overview. Cancer.
 Proteomics And Cancer Biomarker Discovery Dr. Zahid Khan Institute of chemical Sciences (ICS) University of Peshawar Overview Proteomics Cancer Aims Tools Data Base search Challenges Summary 1 Overview
Proteomics And Cancer Biomarker Discovery Dr. Zahid Khan Institute of chemical Sciences (ICS) University of Peshawar Overview Proteomics Cancer Aims Tools Data Base search Challenges Summary 1 Overview
Proteomics. Manickam Sugumaran. Department of Biology University of Massachusetts Boston, MA 02125
 Proteomics Manickam Sugumaran Department of Biology University of Massachusetts Boston, MA 02125 Genomic studies produced more than 75,000 potential gene sequence targets. (The number may be even higher
Proteomics Manickam Sugumaran Department of Biology University of Massachusetts Boston, MA 02125 Genomic studies produced more than 75,000 potential gene sequence targets. (The number may be even higher
Chapter 14 Regulation of Transcription
 Chapter 14 Regulation of Transcription Cis-acting sequences Distance-independent cis-acting elements Dissecting regulatory elements Transcription factors Overview transcriptional regulation Transcription
Chapter 14 Regulation of Transcription Cis-acting sequences Distance-independent cis-acting elements Dissecting regulatory elements Transcription factors Overview transcriptional regulation Transcription
Design. Construction. Characterization
 Design Construction Characterization DNA mrna (messenger) A C C transcription translation C A C protein His A T G C T A C G Plasmids replicon copy number incompatibility selection marker origin of replication
Design Construction Characterization DNA mrna (messenger) A C C transcription translation C A C protein His A T G C T A C G Plasmids replicon copy number incompatibility selection marker origin of replication
Corporate Medical Policy
 Corporate Medical Policy Proteogenomic Testing for Patients with Cancer (GPS Cancer Test) File Name: Origination: Last CAP Review: Next CAP Review: Last Review: proteogenomic_testing_for_patients_with_cancer_gps_cancer_test
Corporate Medical Policy Proteogenomic Testing for Patients with Cancer (GPS Cancer Test) File Name: Origination: Last CAP Review: Next CAP Review: Last Review: proteogenomic_testing_for_patients_with_cancer_gps_cancer_test
Parallel analysis of translated ORF (PLATO)
 Parallel analysis of translated ORF (PLATO) Technical Journal Club 16.04.2013 Mary-Aude Rochat, PhD student Speck group Proteomics Rationale: Increase in the DNA sequence information Understand biological
Parallel analysis of translated ORF (PLATO) Technical Journal Club 16.04.2013 Mary-Aude Rochat, PhD student Speck group Proteomics Rationale: Increase in the DNA sequence information Understand biological
ENCODE DCC Antibody Validation Document
 ENCODE DCC Antibody Validation Document Date of Submission 09/12/12 Name: Trupti Kawli Email: trupti@stanford.edu Lab Snyder Antibody Name: SREBP1 (sc-8984) Target: SREBP1 Company/ Source: Santa Cruz Biotechnology
ENCODE DCC Antibody Validation Document Date of Submission 09/12/12 Name: Trupti Kawli Email: trupti@stanford.edu Lab Snyder Antibody Name: SREBP1 (sc-8984) Target: SREBP1 Company/ Source: Santa Cruz Biotechnology
Supplementary information
 Supplementary information Table of Content: Supplementary Results... 2 Supplementary Figure S1: Experimental validation of AP-MS results by coimmunprecipitation Western blot analysis.... 3 Supplementary
Supplementary information Table of Content: Supplementary Results... 2 Supplementary Figure S1: Experimental validation of AP-MS results by coimmunprecipitation Western blot analysis.... 3 Supplementary
Transcription Eukaryotic Cells
 Transcription Eukaryotic Cells Packet #20 1 Introduction Transcription is the process in which genetic information, stored in a strand of DNA (gene), is copied into a strand of RNA. Protein-encoding genes
Transcription Eukaryotic Cells Packet #20 1 Introduction Transcription is the process in which genetic information, stored in a strand of DNA (gene), is copied into a strand of RNA. Protein-encoding genes
Differential Gene Expression
 Biology 4361 Developmental Biology Differential Gene Expression September 28, 2006 Chromatin Structure ~140 bp ~60 bp Transcriptional Regulation: 1. Packing prevents access CH 3 2. Acetylation ( C O )
Biology 4361 Developmental Biology Differential Gene Expression September 28, 2006 Chromatin Structure ~140 bp ~60 bp Transcriptional Regulation: 1. Packing prevents access CH 3 2. Acetylation ( C O )
Strategies for Quantitative Proteomics. Atelier "Protéomique Quantitative" La Grande Motte, France - June 26, 2007
 Strategies for Quantitative Proteomics Atelier "Protéomique Quantitative", France - June 26, 2007 Bruno Domon, Ph.D. Institut of Molecular Systems Biology ETH Zurich Zürich, Switzerland OUTLINE Introduction
Strategies for Quantitative Proteomics Atelier "Protéomique Quantitative", France - June 26, 2007 Bruno Domon, Ph.D. Institut of Molecular Systems Biology ETH Zurich Zürich, Switzerland OUTLINE Introduction
462 BCH. Biotechnology & Genetic engineering. (Practical)
 462 BCH Biotechnology & Genetic engineering (Practical) Nora Aljebrin Office: Building 5, 3 rd floor, 5T304 E.mail: naljebrin@ksu.edu.sa All lectures and lab sheets are available on my website: Fac.ksu.edu.sa\naljebrin
462 BCH Biotechnology & Genetic engineering (Practical) Nora Aljebrin Office: Building 5, 3 rd floor, 5T304 E.mail: naljebrin@ksu.edu.sa All lectures and lab sheets are available on my website: Fac.ksu.edu.sa\naljebrin
Gene expression analysis. Gene expression analysis. Total RNA. Rare and abundant transcripts. Expression levels. Transcriptional output of the genome
 Gene expression analysis Gene expression analysis Biology of the transcriptome Observing the transcriptome Computational biology of gene expression sven.nelander@wlab.gu.se Recent examples Transcriptonal
Gene expression analysis Gene expression analysis Biology of the transcriptome Observing the transcriptome Computational biology of gene expression sven.nelander@wlab.gu.se Recent examples Transcriptonal
Glutathione Resin. (Cat. # , , , ) think proteins! think G-Biosciences
 191PR-05 G-Biosciences 1-800-628-7730 1-314-991-6034 technical@gbiosciences.com A Geno Technology, Inc. (USA) brand name Glutathione Resin (Cat. # 786-280, 786-310, 786-311, 786-312) think proteins! think
191PR-05 G-Biosciences 1-800-628-7730 1-314-991-6034 technical@gbiosciences.com A Geno Technology, Inc. (USA) brand name Glutathione Resin (Cat. # 786-280, 786-310, 786-311, 786-312) think proteins! think
Chromatographic Separation of the three forms of RNA Polymerase II.
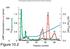 Chromatographic Separation of the three forms of RNA Polymerase II. α-amanitin α-amanitin bound to Pol II Function of the three enzymes. Yeast Pol II. RNA Polymerase Subunit Structures 10-7 Subunit structure.
Chromatographic Separation of the three forms of RNA Polymerase II. α-amanitin α-amanitin bound to Pol II Function of the three enzymes. Yeast Pol II. RNA Polymerase Subunit Structures 10-7 Subunit structure.
Introduction to Proteomics
 Introduction to Proteomics Tasso Miliotis, PhD AstraZeneca R&D Gothenburg Translational Sciences tasso.miliotis@astrazeneca.com 1 Site Management Mölndal May 2008 Outline Drug Discovery AZ R&D Sample Preparation
Introduction to Proteomics Tasso Miliotis, PhD AstraZeneca R&D Gothenburg Translational Sciences tasso.miliotis@astrazeneca.com 1 Site Management Mölndal May 2008 Outline Drug Discovery AZ R&D Sample Preparation
DNA Transcription. Dr Aliwaini
 DNA Transcription 1 DNA Transcription-Introduction The synthesis of an RNA molecule from DNA is called Transcription. All eukaryotic cells have five major classes of RNA: ribosomal RNA (rrna), messenger
DNA Transcription 1 DNA Transcription-Introduction The synthesis of an RNA molecule from DNA is called Transcription. All eukaryotic cells have five major classes of RNA: ribosomal RNA (rrna), messenger
Executing T-REX in mammalian cells
 Supplementary Figure 1 Executing T-REX in mammalian cells HEK-293 cells cultured (a) in 2x 55 cm 2 adherent cell culture plates, and (b and c) in a 48-well multi-well adherent cell culture plate. No cover
Supplementary Figure 1 Executing T-REX in mammalian cells HEK-293 cells cultured (a) in 2x 55 cm 2 adherent cell culture plates, and (b and c) in a 48-well multi-well adherent cell culture plate. No cover
Glutathione Resin. (Cat. # , , , ) think proteins! think G-Biosciences
 191PR 05 G-Biosciences 1-800-628-7730 1-314-991-6034 technical@gbiosciences.com A Geno Technology, Inc. (USA) brand name Glutathione Resin (Cat. # 786 280, 786 310, 786 311, 786 312) think proteins! think
191PR 05 G-Biosciences 1-800-628-7730 1-314-991-6034 technical@gbiosciences.com A Geno Technology, Inc. (USA) brand name Glutathione Resin (Cat. # 786 280, 786 310, 786 311, 786 312) think proteins! think
Genomics and Proteomics *
 OpenStax-CNX module: m44558 1 Genomics and Proteomics * OpenStax This work is produced by OpenStax-CNX and licensed under the Creative Commons Attribution License 3.0 By the end of this section, you will
OpenStax-CNX module: m44558 1 Genomics and Proteomics * OpenStax This work is produced by OpenStax-CNX and licensed under the Creative Commons Attribution License 3.0 By the end of this section, you will
ENCODE DCC Antibody Validation Document
 ENCODE DCC Antibody Validation Document Date of Submission 06/15/2012 Name: Trupti Kawli Email: trupti@stanford.edu Lab Snyder Antibody Name: CDP (sc6327) Target: CDP Company/ Source: Santa Cruz Catalog
ENCODE DCC Antibody Validation Document Date of Submission 06/15/2012 Name: Trupti Kawli Email: trupti@stanford.edu Lab Snyder Antibody Name: CDP (sc6327) Target: CDP Company/ Source: Santa Cruz Catalog
Synthetic Biology. Sustainable Energy. Therapeutics Industrial Enzymes. Agriculture. Accelerating Discoveries, Expanding Possibilities. Design.
 Synthetic Biology Accelerating Discoveries, Expanding Possibilities Sustainable Energy Therapeutics Industrial Enzymes Agriculture Design Build Generate Solutions to Advance Synthetic Biology Research
Synthetic Biology Accelerating Discoveries, Expanding Possibilities Sustainable Energy Therapeutics Industrial Enzymes Agriculture Design Build Generate Solutions to Advance Synthetic Biology Research
Mechanisms of Transcription. School of Life Science Shandong University
 Mechanisms of Transcription School of Life Science Shandong University Ch 12: Mechanisms of Transcription 1. RNA polymerase and the transcription cycle 2. The transcription cycle in bacteria 3. Transcription
Mechanisms of Transcription School of Life Science Shandong University Ch 12: Mechanisms of Transcription 1. RNA polymerase and the transcription cycle 2. The transcription cycle in bacteria 3. Transcription
Computational Biology I LSM5191 (2003/4)
 Computational Biology I LSM5191 (2003/4) Aylwin Ng, D.Phil Lecture Notes: Transcriptome: Molecular Biology of Gene Expression I Flow of information: DNA to polypeptide DNA Start Exon1 Intron Exon2 Termination
Computational Biology I LSM5191 (2003/4) Aylwin Ng, D.Phil Lecture Notes: Transcriptome: Molecular Biology of Gene Expression I Flow of information: DNA to polypeptide DNA Start Exon1 Intron Exon2 Termination
Corporate Medical Policy
 Corporate Medical Policy Proteogenomic Testing for Patients with Cancer (GPS Cancer Test) File Name: Origination: Last CAP Review: Next CAP Review: Last Review: proteogenomic_testing_for_patients_with_cancer_gps_cancer_test
Corporate Medical Policy Proteogenomic Testing for Patients with Cancer (GPS Cancer Test) File Name: Origination: Last CAP Review: Next CAP Review: Last Review: proteogenomic_testing_for_patients_with_cancer_gps_cancer_test
Corporate Medical Policy
 Corporate Medical Policy Proteogenomic Testing for Patients with Cancer (GPS Cancer Test) File Name: Origination: Last CAP Review: Next CAP Review: Last Review: proteogenomic_testing_for_patients_with_cancer_gps_cancer_test
Corporate Medical Policy Proteogenomic Testing for Patients with Cancer (GPS Cancer Test) File Name: Origination: Last CAP Review: Next CAP Review: Last Review: proteogenomic_testing_for_patients_with_cancer_gps_cancer_test
GST Elution Buffer. (Cat. # ) think proteins! think G-Biosciences
 191PR-05 G-Biosciences 1-800-628-7730 1-314-991-6034 technical@gbiosciences.com A Geno Technology, Inc. (USA) brand name GST Elution Buffer (Cat. #786-541) think proteins! think G-Biosciences www.gbiosciences.com
191PR-05 G-Biosciences 1-800-628-7730 1-314-991-6034 technical@gbiosciences.com A Geno Technology, Inc. (USA) brand name GST Elution Buffer (Cat. #786-541) think proteins! think G-Biosciences www.gbiosciences.com
Chapter 5: Proteins: Primary Structure
 Instant download and all chapters Test Bank Fundamentals of Biochemistry Life at the Molecular Level 4th Edition Donald Voet https://testbanklab.com/download/test-bank-fundamentals-biochemistry-life-molecular-level-
Instant download and all chapters Test Bank Fundamentals of Biochemistry Life at the Molecular Level 4th Edition Donald Voet https://testbanklab.com/download/test-bank-fundamentals-biochemistry-life-molecular-level-
Enhancers mutations that make the original mutant phenotype more extreme. Suppressors mutations that make the original mutant phenotype less extreme
 Interactomics and Proteomics 1. Interactomics The field of interactomics is concerned with interactions between genes or proteins. They can be genetic interactions, in which two genes are involved in the
Interactomics and Proteomics 1. Interactomics The field of interactomics is concerned with interactions between genes or proteins. They can be genetic interactions, in which two genes are involved in the
CAP BIOINFORMATICS Su-Shing Chen CISE. 10/5/2005 Su-Shing Chen, CISE 1
 CAP 5510-9 BIOINFORMATICS Su-Shing Chen CISE 10/5/2005 Su-Shing Chen, CISE 1 Basic BioTech Processes Hybridization PCR Southern blotting (spot or stain) 10/5/2005 Su-Shing Chen, CISE 2 10/5/2005 Su-Shing
CAP 5510-9 BIOINFORMATICS Su-Shing Chen CISE 10/5/2005 Su-Shing Chen, CISE 1 Basic BioTech Processes Hybridization PCR Southern blotting (spot or stain) 10/5/2005 Su-Shing Chen, CISE 2 10/5/2005 Su-Shing
Differential Gene Expression
 Biology 4361 Developmental Biology Differential Gene Expression June 19, 2008 Differential Gene Expression Overview Chromatin structure Gene anatomy RNA processing and protein production Initiating transcription:
Biology 4361 Developmental Biology Differential Gene Expression June 19, 2008 Differential Gene Expression Overview Chromatin structure Gene anatomy RNA processing and protein production Initiating transcription:
Transcriptional Regulation in Eukaryotes
 Transcriptional Regulation in Eukaryotes Concepts, Strategies, and Techniques Michael Carey Stephen T. Smale COLD SPRING HARBOR LABORATORY PRESS NEW YORK 2000 Cold Spring Harbor Laboratory Press, 0-87969-537-4
Transcriptional Regulation in Eukaryotes Concepts, Strategies, and Techniques Michael Carey Stephen T. Smale COLD SPRING HARBOR LABORATORY PRESS NEW YORK 2000 Cold Spring Harbor Laboratory Press, 0-87969-537-4
Yesterday s Picture UNIT 3B
 Warm-Up Plasmids are circular pieces of DNA which bacterial cells are able to take up from the environment, then replicate and transcribe. Eukaryotic cells, by contrast, contain large, linear (non-circular)
Warm-Up Plasmids are circular pieces of DNA which bacterial cells are able to take up from the environment, then replicate and transcribe. Eukaryotic cells, by contrast, contain large, linear (non-circular)
Introduction to Microarray Analysis
 Introduction to Microarray Analysis Methods Course: Gene Expression Data Analysis -Day One Rainer Spang Microarrays Highly parallel measurement devices for gene expression levels 1. How does the microarray
Introduction to Microarray Analysis Methods Course: Gene Expression Data Analysis -Day One Rainer Spang Microarrays Highly parallel measurement devices for gene expression levels 1. How does the microarray
Recitation CHAPTER 9 DNA Technologies
 Recitation CHAPTER 9 DNA Technologies DNA Cloning: General Scheme A cloning vector and eukaryotic chromosomes are separately cleaved with the same restriction endonuclease. (A single chromosome is shown
Recitation CHAPTER 9 DNA Technologies DNA Cloning: General Scheme A cloning vector and eukaryotic chromosomes are separately cleaved with the same restriction endonuclease. (A single chromosome is shown
Peptide libraries: applications, design options and considerations. Laura Geuss, PhD May 5, 2015, 2:00-3:00 pm EST
 Peptide libraries: applications, design options and considerations Laura Geuss, PhD May 5, 2015, 2:00-3:00 pm EST Overview 1 2 3 4 5 Introduction Peptide library basics Peptide library design considerations
Peptide libraries: applications, design options and considerations Laura Geuss, PhD May 5, 2015, 2:00-3:00 pm EST Overview 1 2 3 4 5 Introduction Peptide library basics Peptide library design considerations
Tissue Acetyl-Histone H4 ChIP Kit
 Tissue Acetyl-Histone H4 ChIP Kit Catalog Number KA0672 48 assays Version: 03 Intended for research use only www.abnova.com Table of Contents Introduction... 3 Intended Use... 3 Background... 3 Principle
Tissue Acetyl-Histone H4 ChIP Kit Catalog Number KA0672 48 assays Version: 03 Intended for research use only www.abnova.com Table of Contents Introduction... 3 Intended Use... 3 Background... 3 Principle
GENETICS - CLUTCH CH.15 GENOMES AND GENOMICS.
 !! www.clutchprep.com CONCEPT: OVERVIEW OF GENOMICS Genomics is the study of genomes in their entirety Bioinformatics is the analysis of the information content of genomes - Genes, regulatory sequences,
!! www.clutchprep.com CONCEPT: OVERVIEW OF GENOMICS Genomics is the study of genomes in their entirety Bioinformatics is the analysis of the information content of genomes - Genes, regulatory sequences,
SGN-6106 Computational Systems Biology I
 SGN-6106 Computational Systems Biology I A View of Modern Measurement Systems in Cell Biology Kaisa-Leena Taattola 21.2.2007 1 The cell a complex system (Source: Lehninger Principles of Biochemistry 4th
SGN-6106 Computational Systems Biology I A View of Modern Measurement Systems in Cell Biology Kaisa-Leena Taattola 21.2.2007 1 The cell a complex system (Source: Lehninger Principles of Biochemistry 4th
Proteomics and some of its Mass Spectrometric Applications
 Proteomics and some of its Mass Spectrometric Applications What? Large scale screening of proteins, their expression, modifications and interactions by using high-throughput approaches 2 1 Why? The number
Proteomics and some of its Mass Spectrometric Applications What? Large scale screening of proteins, their expression, modifications and interactions by using high-throughput approaches 2 1 Why? The number
Technical tips Session 4
 Technical tips Session 4 Biotinylation assay: Streptavidin is a small bacterial protein that binds with high affinity to the vitamin biotin. This streptavidin-biotin combination can be used to link molecules
Technical tips Session 4 Biotinylation assay: Streptavidin is a small bacterial protein that binds with high affinity to the vitamin biotin. This streptavidin-biotin combination can be used to link molecules
Chapter 3. DNA, RNA, and Protein Synthesis
 Chapter 3. DNA, RNA, and Protein Synthesis 4. Transcription Gene Expression Regulatory region (promoter) 5 flanking region Upstream region Coding region 3 flanking region Downstream region Transcription
Chapter 3. DNA, RNA, and Protein Synthesis 4. Transcription Gene Expression Regulatory region (promoter) 5 flanking region Upstream region Coding region 3 flanking region Downstream region Transcription
Non-Organic-Based Isolation of Mammalian microrna using Norgen s microrna Purification Kit
 Application Note 13 RNA Sample Preparation Non-Organic-Based Isolation of Mammalian microrna using Norgen s microrna Purification Kit B. Lam, PhD 1, P. Roberts, MSc 1 Y. Haj-Ahmad, M.Sc., Ph.D 1,2 1 Norgen
Application Note 13 RNA Sample Preparation Non-Organic-Based Isolation of Mammalian microrna using Norgen s microrna Purification Kit B. Lam, PhD 1, P. Roberts, MSc 1 Y. Haj-Ahmad, M.Sc., Ph.D 1,2 1 Norgen
Identification of Microprotein-Protein Interactions via APEX Tagging
 Supporting Information Identification of Microprotein-Protein Interactions via APEX Tagging Qian Chu, Annie Rathore,, Jolene K. Diedrich,, Cynthia J. Donaldson, John R. Yates III, and Alan Saghatelian
Supporting Information Identification of Microprotein-Protein Interactions via APEX Tagging Qian Chu, Annie Rathore,, Jolene K. Diedrich,, Cynthia J. Donaldson, John R. Yates III, and Alan Saghatelian
Genetic Engineering & Recombinant DNA
 Genetic Engineering & Recombinant DNA Chapter 10 Copyright The McGraw-Hill Companies, Inc) Permission required for reproduction or display. Applications of Genetic Engineering Basic science vs. Applied
Genetic Engineering & Recombinant DNA Chapter 10 Copyright The McGraw-Hill Companies, Inc) Permission required for reproduction or display. Applications of Genetic Engineering Basic science vs. Applied
For Research Use Only. Not for use in diagnostic procedures.
 Printed December 13, 2011 Version 1.0 For Research Use Only. Not for use in diagnostic procedures. DDDDK-tagged Protein PURIFICATION GEL with Elution Peptide (MoAb. clone FLA-1) CODE No. 3326 / 3327 PURIFICATION
Printed December 13, 2011 Version 1.0 For Research Use Only. Not for use in diagnostic procedures. DDDDK-tagged Protein PURIFICATION GEL with Elution Peptide (MoAb. clone FLA-1) CODE No. 3326 / 3327 PURIFICATION
Gene Expression Technology
 Gene Expression Technology Bing Zhang Department of Biomedical Informatics Vanderbilt University bing.zhang@vanderbilt.edu Gene expression Gene expression is the process by which information from a gene
Gene Expression Technology Bing Zhang Department of Biomedical Informatics Vanderbilt University bing.zhang@vanderbilt.edu Gene expression Gene expression is the process by which information from a gene
BIOLOGY - CLUTCH CH.20 - BIOTECHNOLOGY.
 !! www.clutchprep.com CONCEPT: DNA CLONING DNA cloning is a technique that inserts a foreign gene into a living host to replicate the gene and produce gene products. Transformation the process by which
!! www.clutchprep.com CONCEPT: DNA CLONING DNA cloning is a technique that inserts a foreign gene into a living host to replicate the gene and produce gene products. Transformation the process by which
Chapter 11: Applications of Biotechnology
 Chapter 11: Applications of Biotechnology Lecture Outline Enger, E. D., Ross, F. C., & Bailey, D. B. (2012). Concepts in biology (14th ed.). New York: McGraw- Hill. 11-1 Why Biotechnology Works 11-2 Biotechnology
Chapter 11: Applications of Biotechnology Lecture Outline Enger, E. D., Ross, F. C., & Bailey, D. B. (2012). Concepts in biology (14th ed.). New York: McGraw- Hill. 11-1 Why Biotechnology Works 11-2 Biotechnology
Microarrays & Gene Expression Analysis
 Microarrays & Gene Expression Analysis Contents DNA microarray technique Why measure gene expression Clustering algorithms Relation to Cancer SAGE SBH Sequencing By Hybridization DNA Microarrays 1. Developed
Microarrays & Gene Expression Analysis Contents DNA microarray technique Why measure gene expression Clustering algorithms Relation to Cancer SAGE SBH Sequencing By Hybridization DNA Microarrays 1. Developed
Combining Techniques to Answer Molecular Questions
 Combining Techniques to Answer Molecular Questions UNIT FM02 How to cite this article: Curr. Protoc. Essential Lab. Tech. 9:FM02.1-FM02.5. doi: 10.1002/9780470089941.etfm02s9 INTRODUCTION This manual is
Combining Techniques to Answer Molecular Questions UNIT FM02 How to cite this article: Curr. Protoc. Essential Lab. Tech. 9:FM02.1-FM02.5. doi: 10.1002/9780470089941.etfm02s9 INTRODUCTION This manual is
12/11/12. Dr. Sanjeeva Srivastava. IIT Bombay 2. Interactomics Yeast Two-Hybrid Immunoprecipitation Protein microarrays. Proteomics Course NPTEL
 Dr. Sanjeeva Srivastava IIT Bombay Interactomics Yeast Two-Hybrid Immunoprecipitation Protein microarrays IIT Bombay 2 1 IIT Bombay Tremendous effect of interactions IIT Bombay 4 2 Complex formation Protein
Dr. Sanjeeva Srivastava IIT Bombay Interactomics Yeast Two-Hybrid Immunoprecipitation Protein microarrays IIT Bombay 2 1 IIT Bombay Tremendous effect of interactions IIT Bombay 4 2 Complex formation Protein
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature11070 Supplementary Figure 1 Purification of FLAG-tagged proteins. a, Purification of FLAG-RNF12 by FLAG-affinity from nuclear extracts of wild-type (WT) and two FLAG- RNF12 transgenic
doi:10.1038/nature11070 Supplementary Figure 1 Purification of FLAG-tagged proteins. a, Purification of FLAG-RNF12 by FLAG-affinity from nuclear extracts of wild-type (WT) and two FLAG- RNF12 transgenic
Appendix A. Exogenous matrix binding assays with. various putative MARs
 Appendix A Exogenous matrix binding assays with various putative MARs Introduction Matrix attachment regions of DNA are defined operationally by their affinity for the nuclear matrix. Identification of
Appendix A Exogenous matrix binding assays with various putative MARs Introduction Matrix attachment regions of DNA are defined operationally by their affinity for the nuclear matrix. Identification of
From RNA To Protein
 From RNA To Protein 22-11-2016 Introduction mrna Processing heterogeneous nuclear RNA (hnrna) RNA that comprises transcripts of nuclear genes made by RNA polymerase II; it has a wide size distribution
From RNA To Protein 22-11-2016 Introduction mrna Processing heterogeneous nuclear RNA (hnrna) RNA that comprises transcripts of nuclear genes made by RNA polymerase II; it has a wide size distribution
SIBC504: TRANSCRIPTION & RNA PROCESSING Assistant Professor Dr. Chatchawan Srisawat
 SIBC504: TRANSCRIPTION & RNA PROCESSING Assistant Professor Dr. Chatchawan Srisawat TRANSCRIPTION: AN OVERVIEW Transcription: the synthesis of a single-stranded RNA from a doublestranded DNA template.
SIBC504: TRANSCRIPTION & RNA PROCESSING Assistant Professor Dr. Chatchawan Srisawat TRANSCRIPTION: AN OVERVIEW Transcription: the synthesis of a single-stranded RNA from a doublestranded DNA template.
Pushing the Leading Edge in Protein Quantitation: Integrated, Precise, and Reproducible Protein Quantitation Workflow Solutions
 2017 Metabolomics Seminars Pushing the Leading Edge in Protein Quantitation: Integrated, Precise, and Reproducible Protein Quantitation Workflow Solutions The world leader in serving science 2 3 Cancer
2017 Metabolomics Seminars Pushing the Leading Edge in Protein Quantitation: Integrated, Precise, and Reproducible Protein Quantitation Workflow Solutions The world leader in serving science 2 3 Cancer
So.. Let us say you have an impure solution containing a protein of interest. Q: How do you (a) analyze what you have and (b) purify what you want?
 So.. Let us say you have an impure solution containing a protein of interest. Q: How do you (a) analyze what you have and (b) purify what you want? Polyacrylamide Gel Electrophoresis (PAGE) Note: proteins
So.. Let us say you have an impure solution containing a protein of interest. Q: How do you (a) analyze what you have and (b) purify what you want? Polyacrylamide Gel Electrophoresis (PAGE) Note: proteins
Transcriptional Regulation (Gene Regulation)
 Experimental Techniques in Biomedical Sciences 의생명과학실험기법 Transcriptional Regulation (Gene Regulation) 4/17/13 Jeong Hoon Kim (jeongkim@skku.edu) Department of Health Sciences and Technology, SKKU Graduate
Experimental Techniques in Biomedical Sciences 의생명과학실험기법 Transcriptional Regulation (Gene Regulation) 4/17/13 Jeong Hoon Kim (jeongkim@skku.edu) Department of Health Sciences and Technology, SKKU Graduate
Lecture 23: Clinical and Biomedical Applications of Proteomics; Proteomics Industry
 Lecture 23: Clinical and Biomedical Applications of Proteomics; Proteomics Industry Clinical proteomics is the application of proteomic approach to the field of medicine. Proteome of an organism changes
Lecture 23: Clinical and Biomedical Applications of Proteomics; Proteomics Industry Clinical proteomics is the application of proteomic approach to the field of medicine. Proteome of an organism changes
Gene expression DNA RNA. Protein DNA. Replication. Initiation Elongation Processing Export. DNA RNA Protein. Transcription. Degradation.
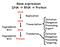 Gene expression DNA RNA Protein DNA DNA Degradation RNA Degradation Protein Replication Transcription Translation Initiation Elongation Processing Export Initiation Elongation Processing Targeting Chapter
Gene expression DNA RNA Protein DNA DNA Degradation RNA Degradation Protein Replication Transcription Translation Initiation Elongation Processing Export Initiation Elongation Processing Targeting Chapter
Biology: the basis for smart proteomic approaches to protein analysis
 QuickTime and a TIFF (Uncompressed) decompressor are needed to see this picture. Biology: the basis for smart proteomic approaches to protein analysis Helen Kim, PhD Department of Pharmacology & Toxicology
QuickTime and a TIFF (Uncompressed) decompressor are needed to see this picture. Biology: the basis for smart proteomic approaches to protein analysis Helen Kim, PhD Department of Pharmacology & Toxicology
EPIGENTEK. EpiQuik Chromatin Immunoprecipitation Kit. Base Catalog # P-2002 PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE
 EpiQuik Chromatin Immunoprecipitation Kit Base Catalog # P-2002 PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE The EpiQuik Chromatin Immunoprecipitation Kit is suitable for combining the specificity of
EpiQuik Chromatin Immunoprecipitation Kit Base Catalog # P-2002 PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE The EpiQuik Chromatin Immunoprecipitation Kit is suitable for combining the specificity of
STSs and ESTs. Sequence-Tagged Site: short, unique sequence Expressed Sequence Tag: short, unique sequence from a coding region
 STSs and ESTs Sequence-Tagged Site: short, unique sequence Expressed Sequence Tag: short, unique sequence from a coding region 1991: 609 ESTs [Adams et al.] June 2000: 4.6 million in dbest Genome sequencing
STSs and ESTs Sequence-Tagged Site: short, unique sequence Expressed Sequence Tag: short, unique sequence from a coding region 1991: 609 ESTs [Adams et al.] June 2000: 4.6 million in dbest Genome sequencing
EPIGENTEK. EpiQuik Tissue Chromatin Immunoprecipitation Kit. Base Catalog # P-2003 PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE
 EpiQuik Tissue Chromatin Immunoprecipitation Kit Base Catalog # P-2003 PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE The EpiQuik Tissue Chromatin Immunoprecipitation Kit is suitable for combining the specificity
EpiQuik Tissue Chromatin Immunoprecipitation Kit Base Catalog # P-2003 PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE The EpiQuik Tissue Chromatin Immunoprecipitation Kit is suitable for combining the specificity
Capabilities & Services
 Capabilities & Services Accelerating Research & Development Table of Contents Introduction to DHMRI 3 Services and Capabilites: Genomics 4 Proteomics & Protein Characterization 5 Metabolomics 6 In Vitro
Capabilities & Services Accelerating Research & Development Table of Contents Introduction to DHMRI 3 Services and Capabilites: Genomics 4 Proteomics & Protein Characterization 5 Metabolomics 6 In Vitro
Control of Eukaryotic Gene Expression (Learning Objectives)
 Control of Eukaryotic Gene Expression (Learning Objectives) 1. Compare and contrast chromatin and chromosome: composition, proteins involved and level of packing. Explain the structure and function of
Control of Eukaryotic Gene Expression (Learning Objectives) 1. Compare and contrast chromatin and chromosome: composition, proteins involved and level of packing. Explain the structure and function of
Differential Gene Expression
 Developmental Biology Biology 4361 Differential Gene Expression October 13, 2005 core transcription initiation site 5 promoter 3 TATAT +1 upstream downstream Basal transcription factors (eukaryotes) TFIID
Developmental Biology Biology 4361 Differential Gene Expression October 13, 2005 core transcription initiation site 5 promoter 3 TATAT +1 upstream downstream Basal transcription factors (eukaryotes) TFIID
Figure S1. TT and MZ-CRC-1 cell viability and proliferation after Palbociclib treatment. Cell viability assays (A) and cell counting assays (B) were
 Figure S1. TT and MZ-CRC-1 cell viability and proliferation after Palbociclib treatment. Cell viability assays (A) and cell counting assays (B) were performed with TT and MZ-CRC-1 cells treated with Palbociclib
Figure S1. TT and MZ-CRC-1 cell viability and proliferation after Palbociclib treatment. Cell viability assays (A) and cell counting assays (B) were performed with TT and MZ-CRC-1 cells treated with Palbociclib
Understanding life WITH NEXT GENERATION PROTEOMICS SOLUTIONS
 Innovative services and products for highly multiplexed protein discovery and quantification to help you understand the biological processes that shape life. Understanding life WITH NEXT GENERATION PROTEOMICS
Innovative services and products for highly multiplexed protein discovery and quantification to help you understand the biological processes that shape life. Understanding life WITH NEXT GENERATION PROTEOMICS
Computing with large data sets
 Computing with large data sets Richard Bonneau, spring 2009 Lecture 14 (week 8): genomics 1 Central dogma Gene expression DNA RNA Protein v22.0480: computing with data, Richard Bonneau Lecture 14 places
Computing with large data sets Richard Bonneau, spring 2009 Lecture 14 (week 8): genomics 1 Central dogma Gene expression DNA RNA Protein v22.0480: computing with data, Richard Bonneau Lecture 14 places
Bioreactor Scale-Up of BacMam-Transduced CHO-S Cells for the Production of Soluble Recombinant Proteins
 Bioreactor Scale-Up of BacMam-Transduced CHO-S Cells for the Production of Soluble Recombinant Proteins Chris Kemp and April Birch Kempbio, Inc. Frederick, MD USA Presentation Summary BacMam Overview BacMam
Bioreactor Scale-Up of BacMam-Transduced CHO-S Cells for the Production of Soluble Recombinant Proteins Chris Kemp and April Birch Kempbio, Inc. Frederick, MD USA Presentation Summary BacMam Overview BacMam
Studying the Human Genome. Lesson Overview. Lesson Overview Studying the Human Genome
 Lesson Overview 14.3 Studying the Human Genome THINK ABOUT IT Just a few decades ago, computers were gigantic machines found only in laboratories and universities. Today, many of us carry small, powerful
Lesson Overview 14.3 Studying the Human Genome THINK ABOUT IT Just a few decades ago, computers were gigantic machines found only in laboratories and universities. Today, many of us carry small, powerful
3. Translation. 2. Transcription. 1. Replication. and functioning through their expression in. Genes are units perpetuating themselves
 Central Dogma Genes are units perpetuating themselves and functioning through their expression in the form of proteins 1 DNA RNA Protein 2 3 1. Replication 2. Transcription 3. Translation Spring 2002 21
Central Dogma Genes are units perpetuating themselves and functioning through their expression in the form of proteins 1 DNA RNA Protein 2 3 1. Replication 2. Transcription 3. Translation Spring 2002 21
Lecture 21 Regulation of transcription
 Lecture 21 Regulation of transcription Key learning goals: Understand what a promoter sequence is Understand the role of sigma factors in prokaryotic RNAP initiation Understand what an open complex is,
Lecture 21 Regulation of transcription Key learning goals: Understand what a promoter sequence is Understand the role of sigma factors in prokaryotic RNAP initiation Understand what an open complex is,
Biochemistry Eukaryotic Transcription
 1 Description of Module Subject Name Paper Name Module Name/Title Dr. Vijaya Khader Dr. MC Varadaraj 2 1. Objectives 1. Understand and have an overview of eucaryotic transcriptional regulation. 2. Explain
1 Description of Module Subject Name Paper Name Module Name/Title Dr. Vijaya Khader Dr. MC Varadaraj 2 1. Objectives 1. Understand and have an overview of eucaryotic transcriptional regulation. 2. Explain
CELL BIOLOGY - CLUTCH CH TECHNIQUES IN CELL BIOLOGY.
 !! www.clutchprep.com CONCEPT: LIGHT MICROSCOPE A light microscope uses light waves and magnification to view specimens Can be used to visualize transparent, and some cellular components - Can visualize
!! www.clutchprep.com CONCEPT: LIGHT MICROSCOPE A light microscope uses light waves and magnification to view specimens Can be used to visualize transparent, and some cellular components - Can visualize
Bis2A 12.2 Eukaryotic Transcription
 OpenStax-CNX module: m56061 1 Bis2A 12.2 Eukaryotic Transcription Mitch Singer Based on Eukaryotic Transcription by OpenStax College This work is produced by OpenStax-CNX and licensed under the Creative
OpenStax-CNX module: m56061 1 Bis2A 12.2 Eukaryotic Transcription Mitch Singer Based on Eukaryotic Transcription by OpenStax College This work is produced by OpenStax-CNX and licensed under the Creative
