Mutagenesis and generation of expression vectors Gene expression profiling Lentiviral production and infection of TF1 cells
|
|
|
- Susanna Gallagher
- 5 years ago
- Views:
Transcription
1 Mutagenesis and generation of expression vectors Human c-kit wild-type cdna encoding the short c-kit isoform was excised from vector pbshkitwt (kind gift from Dr. L Gros, France) by Sal I-Acc65 I digestion. The cdna fragment was subcloned into pentr1a vector (Invitrogen, France). c-kit mutations identified in patients were introduced into pentr1a-hkitwt vector with QuickChange site-directed mutagenesis Kit (Stratagene, Netherlands) according to the manufacturer s instructions. All the mutations were confirmed by bidirectional sequencing. The plxsn-gw vector was constructed with the plxsn retroviral vector (Clontech, Germany) by inserting a Gateway cassette (Invitrogen, France). The Gateway reaction was performed between pentr-kit and plxsn-gw vectors following the manufacturer s recommendations (Invitrogen, France) to obtain the c-kit expression vector. To generate cdnas encoding the hybrid protein EGFP-AKT or MyrEGFP-AKT separated by a 6 Glycines linker, fragments EGFP-6Gly, MyrEGFP-6Gly, and AKT were respectively cloned from three vectors (kindly provided by Dr. P Lecine, France) by using primers containing the Gateway recombination sequences AttB1 or AttB2 (Invitrogen, France). A second PCR was then performed to fuse EGFP-6Gly or MyrEGFP-6Gly to AKT. The resulting amplicons were cloned into the pdonr21 vector (Invitrogen, France) by Gateway technique. Finally, the chimeric cdnas were introduced into pmscvpuro expression vector (Clontech, Germany) which had been previously modified by inserting a Gateway cassette. Gene expression profiling RNA expression profiling of cell lines was done with Affymetrix M43 2. mouse oligonucleotide microarrays containing probe sets, representing transcripts and variants including well-characterized mouse genes. Preparation of crna, hybridizations, washes, detection, and quantification were done as recommended by the supplier. For each sample, synthesis of the first-strand cdna was done from 3 µg total RNA by T7-oligo(dT) priming, followed by second-strand cdna synthesis. After purification, in vitro transcription associated with amplification generated crna-containing biotinylated pseudouridine. Biotinylated crna was purified, quantified, and chemically fragmented (95 C for 35 min), then hybridized to microarrays in 2 µl hybridization buffer at 45 C for 16 h. Automated washes and staining with streptavidin-phycoerythrin were done as recommended. Signal amplification was done by biotinylated antistreptavidin antibody with goat-igg blocking antibody. Scanning was done with Affymetrix GeneArray scanner and quantification with Affymetrix GCOS software. Validation study using cdna-spotted arrays was done with Ipsogen Nylon microarrays containing 8, genes/expressed sequence tags (EST). Expression data were analyzed by the Robust Multichip Average method in R using Bioconductor and associated packages. Filtered and log 2 -transformed data were analyzed by hierarchical clustering using the Cluster program. Results were displayed using TreeView. Lentiviral production and infection of TF1 cells Human Kit cdnas were cloned in prrl lentiviral vector (*). For infection of TF-1 cells, 1 6 cells were incubated for 1 hour in 1 ml of media supplemented with GM-CSF and 8 µg/ml of polybrene (hexadimethrine bromide). Cells were then incubated for 3 hours with infectious particles, spun, and diluted in fresh media. GFP-positive infected cells were sorted 4 days later and immediately cultured in media without GM-CSF.
2 (*) Self-Inactivating Lentivirus Vector for Safe and Efficient In Vivo Gene Delivery. Romain Zufferey, Thomas Dull, Ronald J. Mandel, Anatoly Bukovsky, Dulce Quiroz, Luigi Naldini, and Didier Trono. J Virol December; 72(12): Figure S1. Growth curves of mutant c-kit expressed in Ba/F3 cells Ba/F3 cells expressing the different mutants were seeded in culture in the absence of SCF and counted every day by Trypan blue exclusion. Control native Ba/F3 cells were incubated in the presence of IL3 and counted with the same procedures. Figure S2. Effect of ECD and PTD mutant ectopic expression in human TF1 cells (A) TF1 cells expressing ECD and PTD mutants were seeded in culture in the absence of SCF and counted every day by Trypan blue exclusion. Control parental TF1 cells were incubated in the presence of GM-CSF (5 ng/ml) or SCF (25ng/mL) and were counted with the same procedures. (B) TF1 cells expressing WT or KIT mutants were introduced in a proliferation assay and incubated for 48 h with 5 ng/ml GM-CSF, 25ng/mL SCF or without growth factor and in the presence of 1 µm Imatinib or Dasatinib. Values are presented relative to cell proliferation in the absence of inhibitors.data represent mean values ± standard deviation (SD) of triplicates. Results shown are representative of two independent experiments. (C) Parental and KIT variants expressing TF1 cells were serum starved in RPMI.5% FBS for 3 h and stimulated with or without mscf for 5 min. Total lysates were analysed by SDS-PAGE followed by anti P- KIT (Y719) and anti phospho-akt (S473). The blot was stripped and reprobed with anti-akt antibodies to illustrate equal loading. Figure S3. ECD- or PTD-mutants display very different whole-genome transcriptional profiles Each EML cell line (WT-KIT, Del insy mutant and mutant) was cultured with or without 25 ng/ml SCF for 48 or 72h. Gene expression profile analysis was performed using whole-genome microarrays. Hierarchical clustering was applied to the four samples (2 for Del insy; 2 for ) and to the probe sets most variable across all samples. Samples were sorted into two large groups, which correlated with the type of mutation: Del insy in the right group and in the left group. The two groups appeared homogeneous (correlation coefficient.85 for mutants and.8 for Del insy mutants) and very distinct from each other (correlation coefficient.17). Figure S4. Differential activation of AKT pathway by ECD- and PTD-mutants in EML cells Parental and KIT variants expressing EML cells were serum starved in RPMI.5% FBS for 3 h and stimulated with or without mscf for 5 min. Total lysates were analysed by SDS-PAGE followed by anti phospho-akt (S473), anti-p STAT3, and anti-p ERK antibody blotting. The blot was stripped and reprobed with anti-akt, STAT3, and ERK2 antibody to illustrate equal loading. Figure S5. AKTVIII inhibits ECD-mutants dependent proliferation of EML cells and clonogenicity of Rat2 cells
3 (A) Differential sensitivity of ECD- and PTD-mutants towards AKTVIII in EML cells. EML cells expressing WT-KIT or mutants and Ba/F3 cells were seeded in 96-well plates and cultured for 48 h with or without mscf or mil-3 and in the presence of 5 µm AKTVIII. Cell growth was assessed using WST-1. Values are presented relative to cell proliferation in the absence of AKTVIII and represent mean values ± SD of triplicates. (B) Effect of AKTVIII on colony formation of Rat2 cells. Rat2 cells expressing WT-KIT, Del insy KIT mutant and RasV12 were plated at cells per dish in soft-agar medium with or without mscf and in the presence of 5 µm AKTVIII. Colonies were counted at day 1. Values are shown relative to colony number in the absence of AKTVIII. Figure S6. Analysis of surface expression of KIT by FACS Ba/F3 cells were transfected with WT or mutant forms of KIT. Surface expression of KIT was assessed by FACS using anti-kit monoclonal antibody (solid lines). Untransfected Ba/F3 cells (dotted lines) were used as a negative control.
4 Figure S1 6 5 Number of cells x BaF3 IL3 D Y D816I D816V D816Y K59I Ins 53AY D D1 D2 D3 Growth curves in Ba/F3 cells
5 Figure S2 A B Cell count x1-6 2, 1,8 1,6 1,4 1,2 1,,8,6,4,2, D1 D2 D3 Days DMSO Imatinib Dasatinib TF-1 without GF TF-1 + GM-CSF TF-1 + SCF TF-1 Del InsY TF-1 TF-1 Cell proliferation in OD TF-1 +GM-CSF TF-1 +SCF D InsY TF-1 +GM-CSF TF-1 +SCF D InsY C TF1 cells WT WT D insy SCF P-KIT Y719 P-S 473 AKT AKT
6 Figure S3 r =.17 r =.85 r =.8-72h - 48h Del insy - 72h Del insy - 48h log 2-1 1
7 Figure S4 EML Cells EML Del insy D 816 I SCF P-S 473 AKT AKT P-STAT3 STAT3 P-ERK1/2 ERK1/2
8 Figure S5 A Proliferation (%DMSO control) AKTVIII (µm) EML+SCF Del insy D 816 I BaF3+IL-3 B % colonies of DMSO control AKTVIII (µm) WT+SCF Del insy RasV12
9 Figure S6 WT Del insy S 476 I Cell count K 59 I D 816 Y D 816 I KIT
Gene Expression Technology
 Gene Expression Technology Bing Zhang Department of Biomedical Informatics Vanderbilt University bing.zhang@vanderbilt.edu Gene expression Gene expression is the process by which information from a gene
Gene Expression Technology Bing Zhang Department of Biomedical Informatics Vanderbilt University bing.zhang@vanderbilt.edu Gene expression Gene expression is the process by which information from a gene
Cell proliferation was measured with Cell Counting Kit-8 (Dojindo Laboratories, Kumamoto, Japan).
 1 2 3 4 5 6 7 8 Supplemental Materials and Methods Cell proliferation assay Cell proliferation was measured with Cell Counting Kit-8 (Dojindo Laboratories, Kumamoto, Japan). GCs were plated at 96-well
1 2 3 4 5 6 7 8 Supplemental Materials and Methods Cell proliferation assay Cell proliferation was measured with Cell Counting Kit-8 (Dojindo Laboratories, Kumamoto, Japan). GCs were plated at 96-well
Partial list of differentially expressed genes from cdna microarray, comparing MUC18-
 Supplemental Figure legends Table-1 Partial list of differentially expressed genes from cdna microarray, comparing MUC18- silenced and NT-transduced A375SM cells. Supplemental Figure 1 Effect of MUC-18
Supplemental Figure legends Table-1 Partial list of differentially expressed genes from cdna microarray, comparing MUC18- silenced and NT-transduced A375SM cells. Supplemental Figure 1 Effect of MUC-18
RNA oligonucleotides and 2 -O-methylated oligonucleotides were synthesized by. 5 AGACACAAACACCAUUGUCACACUCCACAGC; Rand-2 OMe,
 Materials and methods Oligonucleotides and DNA constructs RNA oligonucleotides and 2 -O-methylated oligonucleotides were synthesized by Dharmacon Inc. (Lafayette, CO). The sequences were: 122-2 OMe, 5
Materials and methods Oligonucleotides and DNA constructs RNA oligonucleotides and 2 -O-methylated oligonucleotides were synthesized by Dharmacon Inc. (Lafayette, CO). The sequences were: 122-2 OMe, 5
Supplemental Figure 1
 Supplemental Fig. 1. Kinetics of,,, AKT and ERK activation in BMMCs following SCF stimulation. Starved BMMCs were stimulated with 250ng/mL of SCF for the indicated time. Soluble Cell Lysates (SCLs) were
Supplemental Fig. 1. Kinetics of,,, AKT and ERK activation in BMMCs following SCF stimulation. Starved BMMCs were stimulated with 250ng/mL of SCF for the indicated time. Soluble Cell Lysates (SCLs) were
Supplementary data. sienigma. F-Enigma F-EnigmaSM. a-p53
 Supplementary data Supplemental Figure 1 A sienigma #2 sienigma sicontrol a-enigma - + ++ - - - - - - + ++ - - - - - - ++ B sienigma F-Enigma F-EnigmaSM a-flag HLK3 cells - - - + ++ + ++ - + - + + - -
Supplementary data Supplemental Figure 1 A sienigma #2 sienigma sicontrol a-enigma - + ++ - - - - - - + ++ - - - - - - ++ B sienigma F-Enigma F-EnigmaSM a-flag HLK3 cells - - - + ++ + ++ - + - + + - -
supplementary information
 DOI: 10.1038/ncb2116 Figure S1 CDK phosphorylation of EZH2 in cells. (a) Comparison of candidate CDK phosphorylation sites on EZH2 with known CDK substrates by multiple sequence alignments. (b) CDK1 and
DOI: 10.1038/ncb2116 Figure S1 CDK phosphorylation of EZH2 in cells. (a) Comparison of candidate CDK phosphorylation sites on EZH2 with known CDK substrates by multiple sequence alignments. (b) CDK1 and
used at a final concentration of 5 ng/ml. Rabbit anti-bim and mouse anti-mkp2 antibodies were
 1 Supplemental Methods Reagents and chemicals: TGFβ was a generous gift from Genzyme Inc. (Cambridge, MA) and was used at a final concentration of 5 ng/ml. Rabbit anti-bim and mouse anti-mkp2 antibodies
1 Supplemental Methods Reagents and chemicals: TGFβ was a generous gift from Genzyme Inc. (Cambridge, MA) and was used at a final concentration of 5 ng/ml. Rabbit anti-bim and mouse anti-mkp2 antibodies
Microarray Experiment Design
 Microarray Experiment Design Samples used, extract preparation and labelling: AML blasts were isolated from bone marrow by centrifugation on a Ficoll- Hypaque gradient. Total RNA was extracted using TRIzol
Microarray Experiment Design Samples used, extract preparation and labelling: AML blasts were isolated from bone marrow by centrifugation on a Ficoll- Hypaque gradient. Total RNA was extracted using TRIzol
Recruitment of Grb2 to surface IgG and IgE provides antigen receptor-intrinsic costimulation to class-switched B cells
 SUPPLEMENTARY FIGURES Recruitment of Grb2 to surface IgG and IgE provides antigen receptor-intrinsic costimulation to class-switched B cells Niklas Engels, Lars Morten König, Christina Heemann, Johannes
SUPPLEMENTARY FIGURES Recruitment of Grb2 to surface IgG and IgE provides antigen receptor-intrinsic costimulation to class-switched B cells Niklas Engels, Lars Morten König, Christina Heemann, Johannes
Adenoviral Expression Systems. Lentivirus is not the only choice for gene delivery. Adeno-X
 Adenoviral Expression Systems Lentivirus is not the only choice for gene delivery 3 Adeno-X Why choose adenoviral gene delivery? Table I: Adenoviral vs. Lentiviral Gene Delivery Lentivirus Adenovirus Infects
Adenoviral Expression Systems Lentivirus is not the only choice for gene delivery 3 Adeno-X Why choose adenoviral gene delivery? Table I: Adenoviral vs. Lentiviral Gene Delivery Lentivirus Adenovirus Infects
Table S1. Nucleotide sequences of synthesized oligonucleotides for quantitative
 Table S1. Nucleotide sequences of synthesized oligonucleotides for quantitative RT-PCR For CD8-1 transcript, forward primer CD8-1F 5 -TAGTAACCAGAGGCCGCAAGA-3 reverse primer CD8-1R 5 -TCTACTAAGGTGTCCCATAGCATGAT-3
Table S1. Nucleotide sequences of synthesized oligonucleotides for quantitative RT-PCR For CD8-1 transcript, forward primer CD8-1F 5 -TAGTAACCAGAGGCCGCAAGA-3 reverse primer CD8-1R 5 -TCTACTAAGGTGTCCCATAGCATGAT-3
monoclonal antibody. (a) The specificity of the anti-rhbdd1 monoclonal antibody was examined in
 Supplementary information Supplementary figures Supplementary Figure 1 Determination of the s pecificity of in-house anti-rhbdd1 mouse monoclonal antibody. (a) The specificity of the anti-rhbdd1 monoclonal
Supplementary information Supplementary figures Supplementary Figure 1 Determination of the s pecificity of in-house anti-rhbdd1 mouse monoclonal antibody. (a) The specificity of the anti-rhbdd1 monoclonal
Quality Control Assays
 QUALITY CONTROL An integral part of the Penn Vector Core is its robust quality control program which is carried out by a separate quality control group. Quality control assays have been developed and optimized
QUALITY CONTROL An integral part of the Penn Vector Core is its robust quality control program which is carried out by a separate quality control group. Quality control assays have been developed and optimized
LINGO-1, A TRANSMEMBRANE SIGNALING PROTEIN, INHIBITS OLIGODENDROCYTE DIFFERENTIATION AND MYELINATION THROUGH INTERCELLULAR SELF- INTERACTIONS.
 Supplemental Data: LINGO-1, A TRANSMEMBRANE SIGNALING PROTEIN, INHIBITS OLIGODENDROCYTE DIFFERENTIATION AND MYELINATION THROUGH INTERCELLULAR SELF- INTERACTIONS. Scott Jepson, Bryan Vought, Christian H.
Supplemental Data: LINGO-1, A TRANSMEMBRANE SIGNALING PROTEIN, INHIBITS OLIGODENDROCYTE DIFFERENTIATION AND MYELINATION THROUGH INTERCELLULAR SELF- INTERACTIONS. Scott Jepson, Bryan Vought, Christian H.
MicroRNAs Modulate Hematopoietic Lineage Differentiation
 Chen et al., page 1 MicroRNAs Modulate Hematopoietic Lineage Differentiation Chang-Zheng Chen, Ling Li, Harvey F. Lodish, David. Bartel Supplemental Online Material Methods Cell isolation Murine bone marrow
Chen et al., page 1 MicroRNAs Modulate Hematopoietic Lineage Differentiation Chang-Zheng Chen, Ling Li, Harvey F. Lodish, David. Bartel Supplemental Online Material Methods Cell isolation Murine bone marrow
Selected Techniques Part I
 1 Selected Techniques Part I Gel Electrophoresis Can be both qualitative and quantitative Qualitative About what size is the fragment? How many fragments are present? Is there in insert or not? Quantitative
1 Selected Techniques Part I Gel Electrophoresis Can be both qualitative and quantitative Qualitative About what size is the fragment? How many fragments are present? Is there in insert or not? Quantitative
Supplementary information to accompany: A novel role for the DNA repair gene Rad51 in Netrin-1 signalling
 Supplementary information to accompany: A novel role for the DNA repair gene Rad51 in Netrin-1 signalling Glendining KA 1, Markie D 2, Gardner RJM 4, Franz EA 3, Robertson SP 4, Jasoni CL 1 Supplementary
Supplementary information to accompany: A novel role for the DNA repair gene Rad51 in Netrin-1 signalling Glendining KA 1, Markie D 2, Gardner RJM 4, Franz EA 3, Robertson SP 4, Jasoni CL 1 Supplementary
At E17.5, the embryos were rinsed in phosphate-buffered saline (PBS) and immersed in
 Supplementary Materials and Methods Barrier function assays At E17.5, the embryos were rinsed in phosphate-buffered saline (PBS) and immersed in acidic X-gal mix (100 mm phosphate buffer at ph4.3, 3 mm
Supplementary Materials and Methods Barrier function assays At E17.5, the embryos were rinsed in phosphate-buffered saline (PBS) and immersed in acidic X-gal mix (100 mm phosphate buffer at ph4.3, 3 mm
Schematic representation of the endogenous PALB2 locus and gene-disruption constructs
 Supplementary Figures Supplementary Figure 1. Generation of PALB2 -/- and BRCA2 -/- /PALB2 -/- DT40 cells. (A) Schematic representation of the endogenous PALB2 locus and gene-disruption constructs carrying
Supplementary Figures Supplementary Figure 1. Generation of PALB2 -/- and BRCA2 -/- /PALB2 -/- DT40 cells. (A) Schematic representation of the endogenous PALB2 locus and gene-disruption constructs carrying
Supplementary Fig. 1. Schematic structure of TRAIP and RAP80. The prey line below TRAIP indicates bait and the two lines above RAP80 highlight the
 Supplementary Fig. 1. Schematic structure of TRAIP and RAP80. The prey line below TRAIP indicates bait and the two lines above RAP80 highlight the prey clones identified in the yeast two hybrid screen.
Supplementary Fig. 1. Schematic structure of TRAIP and RAP80. The prey line below TRAIP indicates bait and the two lines above RAP80 highlight the prey clones identified in the yeast two hybrid screen.
Design. Construction. Characterization
 Design Construction Characterization DNA mrna (messenger) A C C transcription translation C A C protein His A T G C T A C G Plasmids replicon copy number incompatibility selection marker origin of replication
Design Construction Characterization DNA mrna (messenger) A C C transcription translation C A C protein His A T G C T A C G Plasmids replicon copy number incompatibility selection marker origin of replication
Supplementary Methods
 Supplementary Methods Reverse transcribed Quantitative PCR. Total RNA was isolated from bone marrow derived macrophages using RNeasy Mini Kit (Qiagen), DNase-treated (Promega RQ1), and reverse transcribed
Supplementary Methods Reverse transcribed Quantitative PCR. Total RNA was isolated from bone marrow derived macrophages using RNeasy Mini Kit (Qiagen), DNase-treated (Promega RQ1), and reverse transcribed
CAP BIOINFORMATICS Su-Shing Chen CISE. 10/5/2005 Su-Shing Chen, CISE 1
 CAP 5510-9 BIOINFORMATICS Su-Shing Chen CISE 10/5/2005 Su-Shing Chen, CISE 1 Basic BioTech Processes Hybridization PCR Southern blotting (spot or stain) 10/5/2005 Su-Shing Chen, CISE 2 10/5/2005 Su-Shing
CAP 5510-9 BIOINFORMATICS Su-Shing Chen CISE 10/5/2005 Su-Shing Chen, CISE 1 Basic BioTech Processes Hybridization PCR Southern blotting (spot or stain) 10/5/2005 Su-Shing Chen, CISE 2 10/5/2005 Su-Shing
Supplementary Table 1. The Q-PCR primer sequence is summarized in the following table.
 Supplementary Table 1. The Q-PCR primer sequence is summarized in the following table. Name Sequence (5-3 ) Application Flag-u ggactacaaggacgacgatgac Shared upstream primer for all the amplifications of
Supplementary Table 1. The Q-PCR primer sequence is summarized in the following table. Name Sequence (5-3 ) Application Flag-u ggactacaaggacgacgatgac Shared upstream primer for all the amplifications of
DNA Arrays Affymetrix GeneChip System
 DNA Arrays Affymetrix GeneChip System chip scanner Affymetrix Inc. hybridization Affymetrix Inc. data analysis Affymetrix Inc. mrna 5' 3' TGTGATGGTGGGAATTGGGTCAGAAGGACTGTGGGCGCTGCC... GGAATTGGGTCAGAAGGACTGTGGC
DNA Arrays Affymetrix GeneChip System chip scanner Affymetrix Inc. hybridization Affymetrix Inc. data analysis Affymetrix Inc. mrna 5' 3' TGTGATGGTGGGAATTGGGTCAGAAGGACTGTGGGCGCTGCC... GGAATTGGGTCAGAAGGACTGTGGC
Protocols. We used a buffer mix with polymerase (either Mango Mix (forhandler) or 2x High Fidelity) and the appropriate primers and template DNA.
 Protocols PCRs Colony PCRs We used a buffer mix with polymerase (either Mango Mix (forhandler) or 2x High Fidelity) and the appropriate primers and template DNA. For a 10 µl reaction 2x Mango Mix/2x High
Protocols PCRs Colony PCRs We used a buffer mix with polymerase (either Mango Mix (forhandler) or 2x High Fidelity) and the appropriate primers and template DNA. For a 10 µl reaction 2x Mango Mix/2x High
Nature Medicine doi: /nm.3554
 SUPPLEMENTARY FIGURES LEGENDS Supplementary Figure 1: Generation, purification and characterization of recombinant mouse IL-35 (ril-35). High-Five insect cells expressing high levels of the bicistronic
SUPPLEMENTARY FIGURES LEGENDS Supplementary Figure 1: Generation, purification and characterization of recombinant mouse IL-35 (ril-35). High-Five insect cells expressing high levels of the bicistronic
NUCLEIC ACID PURIFICATION KITS FAST SIMPLE QUALITY CONVENIENT REPRODUCIBLE
 NUCLEIC ACID PURIFICATION KITS FAST SIMPLE QUALITY CONVENIENT REPRODUCIBLE COMPANY PROFILE Since its founding in 1998,, Inc. has been at the forefront of nucleic acid purification by offering products
NUCLEIC ACID PURIFICATION KITS FAST SIMPLE QUALITY CONVENIENT REPRODUCIBLE COMPANY PROFILE Since its founding in 1998,, Inc. has been at the forefront of nucleic acid purification by offering products
- 1 - Supplemental Data
 - 1-1 Supplemental Data 2 3 4 5 6 7 8 9 Supplemental Figure S1. Differential expression of AtPIP Genes in DC3000-inoculated plants. Gene expression in leaves was analyzed by real-time RT-PCR and expression
- 1-1 Supplemental Data 2 3 4 5 6 7 8 9 Supplemental Figure S1. Differential expression of AtPIP Genes in DC3000-inoculated plants. Gene expression in leaves was analyzed by real-time RT-PCR and expression
SUPPLEMENTARY MATERIAL AND METHODS
 SUPPLEMENTARY MATERIAL AND METHODS Amplification of HEV ORF1, ORF2 and ORF3 genome regions Total RNA was extracted from 200 µl EDTA plasma using Cobas AmpliPrep total nucleic acid isolation kit (Roche,
SUPPLEMENTARY MATERIAL AND METHODS Amplification of HEV ORF1, ORF2 and ORF3 genome regions Total RNA was extracted from 200 µl EDTA plasma using Cobas AmpliPrep total nucleic acid isolation kit (Roche,
SUPPLEMENTARY INFORMATION
 R2A MASNSEKNPLL-SDEKPKSTEENKSS-KPESASGSSTSSAMP---GLNFNAFDFSNMASIL 56 R2B MASSSEKTPLIPSDEKNDTKEESKSTTKPESGSGAPPSPS-PTDPGLDFNAFDFSGMAGIL 60 R2A NDPSIREMAEQIAKDPAFNQLAEQLQRSIPNAGQEGGFPNFDPQQYVNTMQQVMHNPEFK
R2A MASNSEKNPLL-SDEKPKSTEENKSS-KPESASGSSTSSAMP---GLNFNAFDFSNMASIL 56 R2B MASSSEKTPLIPSDEKNDTKEESKSTTKPESGSGAPPSPS-PTDPGLDFNAFDFSGMAGIL 60 R2A NDPSIREMAEQIAKDPAFNQLAEQLQRSIPNAGQEGGFPNFDPQQYVNTMQQVMHNPEFK
ASPP1 Fw GGTTGGGAATCCACGTGTTG ASPP1 Rv GCCATATCTTGGAGCTCTGAGAG
 Supplemental Materials and Methods Plasmids: the following plasmids were used in the supplementary data: pwzl-myc- Lats2 (Aylon et al, 2006), pretrosuper-vector and pretrosuper-shp53 (generous gift of
Supplemental Materials and Methods Plasmids: the following plasmids were used in the supplementary data: pwzl-myc- Lats2 (Aylon et al, 2006), pretrosuper-vector and pretrosuper-shp53 (generous gift of
supplementary information
 DOI: 10.1038/ncb1862 Figure S1 Identification of SCAI as a Dia1 associating factor. (a) Identification of SCAI (Riken cdna 930041I02) as a Dia1-FH3 interacting protein. Mouse brain lysate was incubated
DOI: 10.1038/ncb1862 Figure S1 Identification of SCAI as a Dia1 associating factor. (a) Identification of SCAI (Riken cdna 930041I02) as a Dia1-FH3 interacting protein. Mouse brain lysate was incubated
VOLUME 2. Molecular Clonin g A LABORATORY MANUA L THIRD EDITIO N. Joseph Sambrook. David W. Russell
 VOLUME 2 Molecular Clonin g A LABORATORY MANUA L THIRD EDITIO N Joseph Sambrook David W. Russell Chapter 8 In Vitro Amplification of DNA by the Polymerase 8. 1 Chain Reaction 1 The Basic Polymerase Chain
VOLUME 2 Molecular Clonin g A LABORATORY MANUA L THIRD EDITIO N Joseph Sambrook David W. Russell Chapter 8 In Vitro Amplification of DNA by the Polymerase 8. 1 Chain Reaction 1 The Basic Polymerase Chain
Supplemental Materials and Methods
 Supplemental Materials and Methods Antibodies: Anti-SRF (cat# Sc-335) and anti-igf1r (sc-712) (Santa Cruz Biotech), and anti- ADAM-10 (14-6211) were from e-bioscience, anti-ku70 (cat# MS-329-P) (Labvision),
Supplemental Materials and Methods Antibodies: Anti-SRF (cat# Sc-335) and anti-igf1r (sc-712) (Santa Cruz Biotech), and anti- ADAM-10 (14-6211) were from e-bioscience, anti-ku70 (cat# MS-329-P) (Labvision),
MeCP2. MeCP2/α-tubulin. GFP mir1-1 mir132
 Conservation Figure S1. Schematic showing 3 UTR (top; thick black line), mir132 MRE (arrow) and nucleotide sequence conservation (vertical black lines; http://genome.ucsc.edu). a GFP mir1-1 mir132 b GFP
Conservation Figure S1. Schematic showing 3 UTR (top; thick black line), mir132 MRE (arrow) and nucleotide sequence conservation (vertical black lines; http://genome.ucsc.edu). a GFP mir1-1 mir132 b GFP
Nature Biotechnology: doi: /nbt Supplementary Figure 1
 Supplementary Figure 1 Schematic and results of screening the combinatorial antibody library for Sox2 replacement activity. A single batch of MEFs were plated and transduced with doxycycline inducible
Supplementary Figure 1 Schematic and results of screening the combinatorial antibody library for Sox2 replacement activity. A single batch of MEFs were plated and transduced with doxycycline inducible
Fig. S1. Effect of p120-catenin overexpression on the interaction of SCUBE2 with E-cadherin. The expression plasmid encoding FLAG.
 Fig. S1. Effect of p120-catenin overexpression on the interaction of SCUBE2 with E-cadherin. The expression plasmid encoding FLAG.SCUBE2, E-cadherin.Myc, or HA.p120-catenin was transfected in a combination
Fig. S1. Effect of p120-catenin overexpression on the interaction of SCUBE2 with E-cadherin. The expression plasmid encoding FLAG.SCUBE2, E-cadherin.Myc, or HA.p120-catenin was transfected in a combination
Hossain_Supplemental Figure 1
 Hossain_Supplemental Figure 1 GFP-PACT GFP-PACT Motif I GFP-PACT Motif II A. MG132 (1µM) GFP Tubulin GFP-PACT Pericentrin GFP-PACT GFP-PACT Pericentrin Fig. S1. Expression and localization of Orc1 PACT
Hossain_Supplemental Figure 1 GFP-PACT GFP-PACT Motif I GFP-PACT Motif II A. MG132 (1µM) GFP Tubulin GFP-PACT Pericentrin GFP-PACT GFP-PACT Pericentrin Fig. S1. Expression and localization of Orc1 PACT
Supplementary Fig. 1 Proteomic analysis of ATR-interacting proteins. ATR, ARID1A and
 Supplementary Figure Legend: Supplementary Fig. 1 Proteomic analysis of ATR-interacting proteins. ATR, ARID1A and ATRIP protein peptides identified from our mass spectrum analysis were shown. Supplementary
Supplementary Figure Legend: Supplementary Fig. 1 Proteomic analysis of ATR-interacting proteins. ATR, ARID1A and ATRIP protein peptides identified from our mass spectrum analysis were shown. Supplementary
Supporting Online Material for
 www.sciencemag.org/cgi/content/full/1137999/dc1 Supporting Online Material for Disrupting the Pairing Between let-7 and Enhances Oncogenic Transformation Christine Mayr, Michael T. Hemann, David P. Bartel*
www.sciencemag.org/cgi/content/full/1137999/dc1 Supporting Online Material for Disrupting the Pairing Between let-7 and Enhances Oncogenic Transformation Christine Mayr, Michael T. Hemann, David P. Bartel*
Supplementary Table 1. PCR amplification conditions for each primer pair. Primer sequence
 - 1 - Supplementary Tables Supplementary Table 1. PCR amplification conditions for each primer pair Primer sequence FN1 S - CAAAGCAAGCCCGGTTGT AS - CGCTCCCACTGTTGATTTATCTG ITGα2 S - TTAGGTTACTCTGTGGCTGCAATT
- 1 - Supplementary Tables Supplementary Table 1. PCR amplification conditions for each primer pair Primer sequence FN1 S - CAAAGCAAGCCCGGTTGT AS - CGCTCCCACTGTTGATTTATCTG ITGα2 S - TTAGGTTACTCTGTGGCTGCAATT
Establishing sirna assays in primary human peripheral blood lymphocytes
 page 1 of 7 Establishing sirna assays in primary human peripheral blood lymphocytes This protocol is designed to establish sirna assays in primary human peripheral blood lymphocytes using the Nucleofector
page 1 of 7 Establishing sirna assays in primary human peripheral blood lymphocytes This protocol is designed to establish sirna assays in primary human peripheral blood lymphocytes using the Nucleofector
CRISPR-based gene disruption in murine HSPCs. Ayumi Kitano and Daisuke Nakada
 CRISPR-based gene disruption in murine HSPCs Ayumi Kitano and Daisuke Nakada Nakada Lab Protocol INTRODUCTION We recently described fast, efficient, and cost-effective methods to directly modify the genomes
CRISPR-based gene disruption in murine HSPCs Ayumi Kitano and Daisuke Nakada Nakada Lab Protocol INTRODUCTION We recently described fast, efficient, and cost-effective methods to directly modify the genomes
Supplementary Information: Materials and Methods. Immunoblot and immunoprecipitation. Cells were washed in phosphate buffered
 Supplementary Information: Materials and Methods Immunoblot and immunoprecipitation. Cells were washed in phosphate buffered saline (PBS) and lysed in TNN lysis buffer (50mM Tris at ph 8.0, 120mM NaCl
Supplementary Information: Materials and Methods Immunoblot and immunoprecipitation. Cells were washed in phosphate buffered saline (PBS) and lysed in TNN lysis buffer (50mM Tris at ph 8.0, 120mM NaCl
Supplemental Fig. 1: PEA-15 knockdown efficiency assessed by immunohistochemistry and qpcr
 Supplemental figure legends Supplemental Fig. 1: PEA-15 knockdown efficiency assessed by immunohistochemistry and qpcr A, LβT2 cells were transfected with either scrambled or PEA-15 sirna. Cells were then
Supplemental figure legends Supplemental Fig. 1: PEA-15 knockdown efficiency assessed by immunohistochemistry and qpcr A, LβT2 cells were transfected with either scrambled or PEA-15 sirna. Cells were then
Supplementary Materials
 Supplementary Materials Supplementary Figure 1. PKM2 interacts with MLC2 in cytokinesis. a, U87, U87/EGFRvIII, and HeLa cells in cytokinesis were immunostained with DAPI and an anti-pkm2 antibody. Thirty
Supplementary Materials Supplementary Figure 1. PKM2 interacts with MLC2 in cytokinesis. a, U87, U87/EGFRvIII, and HeLa cells in cytokinesis were immunostained with DAPI and an anti-pkm2 antibody. Thirty
all samples of a band with a molecular weight close to that expected for the endogenous!-
 SUPPLEMENTAL FIGURE LEGENDS Supplemental Figure 1 : Specificity of anti-!-arrestin Abs and levels of!-arrestin in WHIM wt leukocytes. (A and B) HEK 293T cells were transiently transfected using the reagent
SUPPLEMENTAL FIGURE LEGENDS Supplemental Figure 1 : Specificity of anti-!-arrestin Abs and levels of!-arrestin in WHIM wt leukocytes. (A and B) HEK 293T cells were transiently transfected using the reagent
Protocol CRISPR Genome Editing In Cell Lines
 Protocol CRISPR Genome Editing In Cell Lines Protocol 2: HDR donor plasmid applications (gene knockout, gene mutagenesis, gene tagging, Safe Harbor ORF knock-in) Notes: 1. sgrna validation: GeneCopoeia
Protocol CRISPR Genome Editing In Cell Lines Protocol 2: HDR donor plasmid applications (gene knockout, gene mutagenesis, gene tagging, Safe Harbor ORF knock-in) Notes: 1. sgrna validation: GeneCopoeia
Supplemental Materials and Methods
 Supplemental Materials and Methods In situ hybridization In situ hybridization analysis of HFE2 and genin mrna in rat liver tissues was performed as previously described (1). Briefly, the digoxigenin-labeled
Supplemental Materials and Methods In situ hybridization In situ hybridization analysis of HFE2 and genin mrna in rat liver tissues was performed as previously described (1). Briefly, the digoxigenin-labeled
Standardized Assays and Reagents for GeneChip Expression Analysis
 Standardized Assays and Reagents for GeneChip Expression Analysis Tools to take you as far as your vision. Ensure Robust, Reproducible Results with Standardized GeneChip Assays and Reagents Standardized,
Standardized Assays and Reagents for GeneChip Expression Analysis Tools to take you as far as your vision. Ensure Robust, Reproducible Results with Standardized GeneChip Assays and Reagents Standardized,
Supplementary Information Alternative splicing of CD44 mrna by ESRP1 enhances lung colonization of metastatic cancer cell
 Supplementary Information Alternative splicing of CD44 mrna by ESRP1 enhances lung colonization of metastatic cancer cell Supplementary Figures S1-S3 Supplementary Methods Supplementary Figure S1. Identification
Supplementary Information Alternative splicing of CD44 mrna by ESRP1 enhances lung colonization of metastatic cancer cell Supplementary Figures S1-S3 Supplementary Methods Supplementary Figure S1. Identification
Short hairpin RNA (shrna) against MMP14. Lentiviral plasmids containing shrna
 Supplemental Materials and Methods Short hairpin RNA (shrna) against MMP14. Lentiviral plasmids containing shrna (Mission shrna, Sigma) against mouse MMP14 were transfected into HEK293 cells using FuGene6
Supplemental Materials and Methods Short hairpin RNA (shrna) against MMP14. Lentiviral plasmids containing shrna (Mission shrna, Sigma) against mouse MMP14 were transfected into HEK293 cells using FuGene6
Supplemental Methods Cell lines and culture
 Supplemental Methods Cell lines and culture AGS, CL5, BT549, and SKBR were propagated in RPMI 64 medium (Mediatech Inc., Manassas, VA) supplemented with % fetal bovine serum (FBS, Atlanta Biologicals,
Supplemental Methods Cell lines and culture AGS, CL5, BT549, and SKBR were propagated in RPMI 64 medium (Mediatech Inc., Manassas, VA) supplemented with % fetal bovine serum (FBS, Atlanta Biologicals,
Supplementary Figure 1: Overexpression of EBV-encoded proteins Western blot analysis of the expression levels of EBV-encoded latency III proteins in
 Supplementary Figure 1: Overexpression of EBV-encoded proteins Western blot analysis of the expression levels of EBV-encoded latency III proteins in BL2 cells. The Ponceau S staining of the membranes or
Supplementary Figure 1: Overexpression of EBV-encoded proteins Western blot analysis of the expression levels of EBV-encoded latency III proteins in BL2 cells. The Ponceau S staining of the membranes or
Supplementary Figure S1. The tetracycline-inducible CRISPR system. A) Hela cells stably
 Supplementary Information Supplementary Figure S1. The tetracycline-inducible CRISPR system. A) Hela cells stably expressing shrna sequences against TRF2 were examined by western blotting. shcon, shrna
Supplementary Information Supplementary Figure S1. The tetracycline-inducible CRISPR system. A) Hela cells stably expressing shrna sequences against TRF2 were examined by western blotting. shcon, shrna
Cat # Box1 Box2. DH5a Competent E. coli cells CCK-20 (20 rxns) 40 µl 40 µl 50 µl x 20 tubes. Choo-Choo Cloning TM Enzyme Mix
 Molecular Cloning Laboratories User Manual Version 3.3 Product name: Choo-Choo Cloning Kits Cat #: CCK-10, CCK-20, CCK-096, CCK-384 Description: Choo-Choo Cloning is a highly efficient directional PCR
Molecular Cloning Laboratories User Manual Version 3.3 Product name: Choo-Choo Cloning Kits Cat #: CCK-10, CCK-20, CCK-096, CCK-384 Description: Choo-Choo Cloning is a highly efficient directional PCR
Flow cytometry Stained cells were analyzed and sorted by SORP FACS Aria (BD Biosciences).
 Mice C57BL/6-Ly5.1 or -Ly5.2 congenic mice were used for LSK transduction and competitive repopulation assays. Animal care was in accordance with the guidelines of Keio University for animal and recombinant
Mice C57BL/6-Ly5.1 or -Ly5.2 congenic mice were used for LSK transduction and competitive repopulation assays. Animal care was in accordance with the guidelines of Keio University for animal and recombinant
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature09732 Supplementary Figure 1: Depletion of Fbw7 results in elevated Mcl-1 abundance. a, Total thymocytes from 8-wk-old Lck-Cre/Fbw7 +/fl (Control) or Lck-Cre/Fbw7 fl/fl (Fbw7 KO) mice
doi:10.1038/nature09732 Supplementary Figure 1: Depletion of Fbw7 results in elevated Mcl-1 abundance. a, Total thymocytes from 8-wk-old Lck-Cre/Fbw7 +/fl (Control) or Lck-Cre/Fbw7 fl/fl (Fbw7 KO) mice
Supplementary Methods Plasmid constructs
 Supplementary Methods Plasmid constructs. Mouse cdna encoding SHP-1, amplified from mrna of RAW264.7 macrophages with primer 5'cgtgcctgcccagacaaactgt3' and 5'cggaattcagacgaatgcccagatcacttcc3', was cloned
Supplementary Methods Plasmid constructs. Mouse cdna encoding SHP-1, amplified from mrna of RAW264.7 macrophages with primer 5'cgtgcctgcccagacaaactgt3' and 5'cggaattcagacgaatgcccagatcacttcc3', was cloned
EPIGENTEK. EpiQuik Tissue Chromatin Immunoprecipitation Kit. Base Catalog # P-2003 PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE
 EpiQuik Tissue Chromatin Immunoprecipitation Kit Base Catalog # P-2003 PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE The EpiQuik Tissue Chromatin Immunoprecipitation Kit is suitable for combining the specificity
EpiQuik Tissue Chromatin Immunoprecipitation Kit Base Catalog # P-2003 PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE The EpiQuik Tissue Chromatin Immunoprecipitation Kit is suitable for combining the specificity
Supplementary Figure 1 - Characterization of rbag3 binding on macrophages cell surface.
 Supplementary Figure 1 - Characterization of rbag3 binding on macrophages cell surface. (a) Human PDAC cell lines were treated as indicated in Figure 1 panel F. Cells were analyzed for FITC-rBAG3 binding
Supplementary Figure 1 - Characterization of rbag3 binding on macrophages cell surface. (a) Human PDAC cell lines were treated as indicated in Figure 1 panel F. Cells were analyzed for FITC-rBAG3 binding
Supplemental Information
 Supplemental Information ATP-dependent unwinding of U4/U6 snrnas by the Brr2 helicase requires the C-terminus of Prp8 Corina Maeder 1,3, Alan K. Kutach 1,2,3, and Christine Guthrie 1 1 Department of Biochemistry
Supplemental Information ATP-dependent unwinding of U4/U6 snrnas by the Brr2 helicase requires the C-terminus of Prp8 Corina Maeder 1,3, Alan K. Kutach 1,2,3, and Christine Guthrie 1 1 Department of Biochemistry
Supplementary Table 1. Primers used to construct full-length or various truncated mutants of ISG12b2.
 Supplementary Table 1. Primers used to construct full-length or various truncated mutants of ISG12b2. Construct name ISG12b2 (No tag) HA-ISG12b2 (N-HA) ISG12b2-HA (C-HA; FL-HA) 94-283-HA (FL-GFP) 93-GFP
Supplementary Table 1. Primers used to construct full-length or various truncated mutants of ISG12b2. Construct name ISG12b2 (No tag) HA-ISG12b2 (N-HA) ISG12b2-HA (C-HA; FL-HA) 94-283-HA (FL-GFP) 93-GFP
(Supplementary Methods online)
 (Supplementary Methods online) Production and purification of either LC-antisense or control molecules Recombinant phagemids and the phagemid vector were transformed into XL-1 Blue competent bacterial
(Supplementary Methods online) Production and purification of either LC-antisense or control molecules Recombinant phagemids and the phagemid vector were transformed into XL-1 Blue competent bacterial
Transfection of CRISPR/Cas9 Nuclease NLS ribonucleoprotein (RNP) into adherent mammalian cells using Lipofectamine RNAiMAX
 Transfection of CRISPR/Cas9 Nuclease NLS ribonucleoprotein (RNP) into adherent mammalian cells using Lipofectamine RNAiMAX INTRODUCTION The CRISPR/Cas genome editing system consists of a single guide RNA
Transfection of CRISPR/Cas9 Nuclease NLS ribonucleoprotein (RNP) into adherent mammalian cells using Lipofectamine RNAiMAX INTRODUCTION The CRISPR/Cas genome editing system consists of a single guide RNA
Supporting Information
 Supporting Information Nan et al. 10.1073/pnas.1509123112 SI Materials and Methods Plasmids. PAmCherry1-KRas wild type and G12D mutants were generated by subcloning the respective KRas cdna fragments into
Supporting Information Nan et al. 10.1073/pnas.1509123112 SI Materials and Methods Plasmids. PAmCherry1-KRas wild type and G12D mutants were generated by subcloning the respective KRas cdna fragments into
Human IgG ELISpot BASIC
 Human IgG ELISpot BASIC Product Code: 3850-2H CONTENTS: Vial 1 (yellow top) Monoclonal antibodies MT91/145 (1.2 ml) Concentration: 0.5 mg/ml Vial 2 (green top) Biotinylated monoclonal antibodies MT78/145
Human IgG ELISpot BASIC Product Code: 3850-2H CONTENTS: Vial 1 (yellow top) Monoclonal antibodies MT91/145 (1.2 ml) Concentration: 0.5 mg/ml Vial 2 (green top) Biotinylated monoclonal antibodies MT78/145
SANTA CRUZ BIOTECHNOLOGY, INC.
 TECHNICAL SERVICE GUIDE: Western Blotting 2. What size bands were expected and what size bands were detected? 3. Was the blot blank or was a dark background or non-specific bands seen? 4. Did this same
TECHNICAL SERVICE GUIDE: Western Blotting 2. What size bands were expected and what size bands were detected? 3. Was the blot blank or was a dark background or non-specific bands seen? 4. Did this same
For quantitative detection of mouse IGF-1 in serum, body fluids, tissue lysates or cell culture supernatants.
 Mouse IGF-1 ELISA Kit (migf-1-elisa) Cat. No. EK0378 96 Tests in 8 x 12 divisible strips Background Insulin-like growth factor 1 (IGF-1), also known as somatomedin C, is a polypeptide protein hormone similar
Mouse IGF-1 ELISA Kit (migf-1-elisa) Cat. No. EK0378 96 Tests in 8 x 12 divisible strips Background Insulin-like growth factor 1 (IGF-1), also known as somatomedin C, is a polypeptide protein hormone similar
Supplementary Methods
 Supplementary Methods Microarray Data Analysis Gene expression data were obtained by hybridising a total of 24 samples from 6 experimental groups (n=4 per group) to Illumina HumanHT-12 Expression BeadChips.
Supplementary Methods Microarray Data Analysis Gene expression data were obtained by hybridising a total of 24 samples from 6 experimental groups (n=4 per group) to Illumina HumanHT-12 Expression BeadChips.
Supporting Information
 Supporting Information Tal et al. 10.1073/pnas.0807694106 SI Materials and Methods VSV Infection and Quantification. Infection was carried out by seeding 5 10 5 MEF cells per well in a 6-well plate and
Supporting Information Tal et al. 10.1073/pnas.0807694106 SI Materials and Methods VSV Infection and Quantification. Infection was carried out by seeding 5 10 5 MEF cells per well in a 6-well plate and
Data Sheet GITR / NF-ĸB-Luciferase Reporter (Luc) - Jurkat Cell Line Catalog #60546
 Data Sheet GITR / NF-ĸB-Luciferase Reporter (Luc) - Jurkat Cell Line Catalog #60546 Description This cell line expresses a surface human GITR (glucocorticoid-induced TNFR family related gene; TNFRSF18;
Data Sheet GITR / NF-ĸB-Luciferase Reporter (Luc) - Jurkat Cell Line Catalog #60546 Description This cell line expresses a surface human GITR (glucocorticoid-induced TNFR family related gene; TNFRSF18;
Online Supporting Material for. The Bisecting GlcNAc on N-Glycans Inhibits Growth. Factor Signaling and Retards Mammary Tumor
 Online Supporting Material for The Bisecting GlcNAc on N-Glycans Inhibits Growth Factor Signaling and Retards Mammary Tumor Progression Yinghui Song 1, Jason A. Aglipay 1, Joshua D. Bernstein 2, Sumanta
Online Supporting Material for The Bisecting GlcNAc on N-Glycans Inhibits Growth Factor Signaling and Retards Mammary Tumor Progression Yinghui Song 1, Jason A. Aglipay 1, Joshua D. Bernstein 2, Sumanta
INVESTIGATION OF THE BINDING SPECIFICITY OF IGF-IR USING MONOCLONAL ANTIBODIES
 INVESTIGATION OF THE BINDING SPECIFICITY OF IGF-IR USING MONOCLONAL ANTIBODIES By Mehrnaz Keyhanfar, Pharm.D. A thesis submitted to the University of Adelaide, South Australia in fulfilment of the requirements
INVESTIGATION OF THE BINDING SPECIFICITY OF IGF-IR USING MONOCLONAL ANTIBODIES By Mehrnaz Keyhanfar, Pharm.D. A thesis submitted to the University of Adelaide, South Australia in fulfilment of the requirements
Supplementary information
 Supplementary information Identification of E-cadherin signature sites functioning as cleavage sites for Helicobacter pylori HtrA Thomas P. Schmidt 1*, Anna M. Perna 2*, Tim Fugmann 3, Manja Böhm 4, Jan
Supplementary information Identification of E-cadherin signature sites functioning as cleavage sites for Helicobacter pylori HtrA Thomas P. Schmidt 1*, Anna M. Perna 2*, Tim Fugmann 3, Manja Böhm 4, Jan
Mouse IgM ELISpot BASIC
 Mouse IgM ELISpot BASIC Product Code: 3885-2H CONTENTS: Vial 1 (green top) Monoclonal antibody MT6A3 (1.2 ml) Concentration: 0.5 mg/ml Vial 2 (yellow top) Biotinylated monoclonal antibody MT9A2 (100 μl)
Mouse IgM ELISpot BASIC Product Code: 3885-2H CONTENTS: Vial 1 (green top) Monoclonal antibody MT6A3 (1.2 ml) Concentration: 0.5 mg/ml Vial 2 (yellow top) Biotinylated monoclonal antibody MT9A2 (100 μl)
Rat IGF-1 ELISA Kit (rigf-1-elisa)
 Rat IGF-1 ELISA Kit (rigf-1-elisa) Cat. No. EK0377 96 Tests in 8 x 12 divisible strips Background Insulin-like growth factor 1 (IGF-1), also known as somatomedin C, is a polypeptide protein hormone similar
Rat IGF-1 ELISA Kit (rigf-1-elisa) Cat. No. EK0377 96 Tests in 8 x 12 divisible strips Background Insulin-like growth factor 1 (IGF-1), also known as somatomedin C, is a polypeptide protein hormone similar
Applicazioni biotecnologiche
 Applicazioni biotecnologiche Analisi forense Sintesi di proteine ricombinanti Restriction Fragment Length Polymorphism (RFLP) Polymorphism (more fully genetic polymorphism) refers to the simultaneous occurrence
Applicazioni biotecnologiche Analisi forense Sintesi di proteine ricombinanti Restriction Fragment Length Polymorphism (RFLP) Polymorphism (more fully genetic polymorphism) refers to the simultaneous occurrence
supplementary information
 DOI: 10.1038/ncb1816 A Gal4 Gal4 B Kinase domain C (1502) N (5111191) Interaction with SIAH1 C SIAH1 SIAH1 C (182) SIAH1 N (82282) RING domain Interaction with Input GST GSTSIAH1 1a a N C Input GST GSTSIAH1
DOI: 10.1038/ncb1816 A Gal4 Gal4 B Kinase domain C (1502) N (5111191) Interaction with SIAH1 C SIAH1 SIAH1 C (182) SIAH1 N (82282) RING domain Interaction with Input GST GSTSIAH1 1a a N C Input GST GSTSIAH1
Supplementary Information
 Journal : Nature Biotechnology Supplementary Information Targeted genome engineering in human cells with RNA-guided endonucleases Seung Woo Cho, Sojung Kim, Jong Min Kim, and Jin-Soo Kim* National Creative
Journal : Nature Biotechnology Supplementary Information Targeted genome engineering in human cells with RNA-guided endonucleases Seung Woo Cho, Sojung Kim, Jong Min Kim, and Jin-Soo Kim* National Creative
Supplementary Information to: Genome-wide Real-time in vivo Transcriptional Dynamics During Plasmodium falciparum. Blood-stage Development
 Supplementary Information to: Genome-wide Real-time in vivo Transcriptional Dynamics During Plasmodium falciparum Blood-stage Development Painter et al. 1 of 8 Supplementary Figure 1: Supplementary Figure
Supplementary Information to: Genome-wide Real-time in vivo Transcriptional Dynamics During Plasmodium falciparum Blood-stage Development Painter et al. 1 of 8 Supplementary Figure 1: Supplementary Figure
Chapter 20 Recombinant DNA Technology. Copyright 2009 Pearson Education, Inc.
 Chapter 20 Recombinant DNA Technology Copyright 2009 Pearson Education, Inc. 20.1 Recombinant DNA Technology Began with Two Key Tools: Restriction Enzymes and DNA Cloning Vectors Recombinant DNA refers
Chapter 20 Recombinant DNA Technology Copyright 2009 Pearson Education, Inc. 20.1 Recombinant DNA Technology Began with Two Key Tools: Restriction Enzymes and DNA Cloning Vectors Recombinant DNA refers
Creating pentr vectors by BP reaction
 Creating pentr vectors by BP reaction Tanya Lepikhova and Rafael Martinez 15072011 Overview: 1. Design primers to add the attb sites to gene of interest 2. Perform PCR with a high fidelity DNA polymerase
Creating pentr vectors by BP reaction Tanya Lepikhova and Rafael Martinez 15072011 Overview: 1. Design primers to add the attb sites to gene of interest 2. Perform PCR with a high fidelity DNA polymerase
Bi 8 Lecture 4. Ellen Rothenberg 14 January Reading: from Alberts Ch. 8
 Bi 8 Lecture 4 DNA approaches: How we know what we know Ellen Rothenberg 14 January 2016 Reading: from Alberts Ch. 8 Central concept: DNA or RNA polymer length as an identifying feature RNA has intrinsically
Bi 8 Lecture 4 DNA approaches: How we know what we know Ellen Rothenberg 14 January 2016 Reading: from Alberts Ch. 8 Central concept: DNA or RNA polymer length as an identifying feature RNA has intrinsically
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb3240 Supplementary Figure 1 GBM cell lines display similar levels of p100 to p52 processing but respond differentially to TWEAK-induced TERT expression according to TERT promoter mutation
DOI: 10.1038/ncb3240 Supplementary Figure 1 GBM cell lines display similar levels of p100 to p52 processing but respond differentially to TWEAK-induced TERT expression according to TERT promoter mutation
The retroviral vectors encoding WT human SIRT1 or a mutant of SIRT in which a
 Supporting online material Constructs The retroviral vectors encoding WT human SIRT1 or a mutant of SIRT in which a critical histidine in the deacetylase domain of SIRT1 has been replaced by a tyrosine
Supporting online material Constructs The retroviral vectors encoding WT human SIRT1 or a mutant of SIRT in which a critical histidine in the deacetylase domain of SIRT1 has been replaced by a tyrosine
Enhancers mutations that make the original mutant phenotype more extreme. Suppressors mutations that make the original mutant phenotype less extreme
 Interactomics and Proteomics 1. Interactomics The field of interactomics is concerned with interactions between genes or proteins. They can be genetic interactions, in which two genes are involved in the
Interactomics and Proteomics 1. Interactomics The field of interactomics is concerned with interactions between genes or proteins. They can be genetic interactions, in which two genes are involved in the
Product Manual. GFP ELISA Kit. Catalog Number. FOR RESEARCH USE ONLY Not for use in diagnostic procedures
 Product Manual GFP ELISA Kit Catalog Number AKR-121 96 assays FOR RESEARCH USE ONLY Not for use in diagnostic procedures Introduction Green fluorescent protein (GFP) is a spontaneously fluorescent protein
Product Manual GFP ELISA Kit Catalog Number AKR-121 96 assays FOR RESEARCH USE ONLY Not for use in diagnostic procedures Introduction Green fluorescent protein (GFP) is a spontaneously fluorescent protein
ViraBind Lentivirus Concentration and Purification Kit
 Product Manual ViraBind Lentivirus Concentration and Purification Kit Catalog Number VPK-091 VPK-091-5 5 preps 25 preps FOR RESEARCH USE ONLY Not for use in diagnostic procedures Introduction Lentivirus
Product Manual ViraBind Lentivirus Concentration and Purification Kit Catalog Number VPK-091 VPK-091-5 5 preps 25 preps FOR RESEARCH USE ONLY Not for use in diagnostic procedures Introduction Lentivirus
TransIT -Lenti Transfection Reagent
 Quick Reference Protocol, SDS and Certificate of Analysis available at mirusbio.com/6600 INTRODUCTION Lentivirus is an enveloped, single-stranded RNA virus from the Retroviridae family capable of infecting
Quick Reference Protocol, SDS and Certificate of Analysis available at mirusbio.com/6600 INTRODUCTION Lentivirus is an enveloped, single-stranded RNA virus from the Retroviridae family capable of infecting
ViraBind Lentivirus Concentration and Purification Kit
 Product Manual ViraBind Lentivirus Concentration and Purification Kit Catalog Number VPK-090 2 preps FOR RESEARCH USE ONLY Not for use in diagnostic procedures Introduction Lentivirus vectors based on
Product Manual ViraBind Lentivirus Concentration and Purification Kit Catalog Number VPK-090 2 preps FOR RESEARCH USE ONLY Not for use in diagnostic procedures Introduction Lentivirus vectors based on
pdsipher and pdsipher -GFP shrna Vector User s Guide
 pdsipher and pdsipher -GFP shrna Vector User s Guide NOTE: PLEASE READ THE ENTIRE PROTOCOL CAREFULLY BEFORE USE Page 1. Introduction... 1 2. Vector Overview... 1 3. Vector Maps 2 4. Materials Provided...
pdsipher and pdsipher -GFP shrna Vector User s Guide NOTE: PLEASE READ THE ENTIRE PROTOCOL CAREFULLY BEFORE USE Page 1. Introduction... 1 2. Vector Overview... 1 3. Vector Maps 2 4. Materials Provided...
Recent technology allow production of microarrays composed of 70-mers (essentially a hybrid of the two techniques)
 Microarrays and Transcript Profiling Gene expression patterns are traditionally studied using Northern blots (DNA-RNA hybridization assays). This approach involves separation of total or polya + RNA on
Microarrays and Transcript Profiling Gene expression patterns are traditionally studied using Northern blots (DNA-RNA hybridization assays). This approach involves separation of total or polya + RNA on
Supplementary Figure 1 PARP1 is involved in regulating the stability of mrnas from pro-inflammatory cytokine/chemokine mediators.
 Supplementary Figure 1 PARP1 is involved in regulating the stability of mrnas from pro-inflammatory cytokine/chemokine mediators. (a) A graphic depiction of the approach to determining the stability of
Supplementary Figure 1 PARP1 is involved in regulating the stability of mrnas from pro-inflammatory cytokine/chemokine mediators. (a) A graphic depiction of the approach to determining the stability of
Supplemental Figure 1
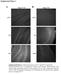 Supplemental Figure 1 A C19 B Mα p27 TL p27-/- p27-/- Supplemental Figure 1: Altered expression of p27 and p27 in the brain. Immunostaining for p27 was performed on brain frozen sections. Brain sections
Supplemental Figure 1 A C19 B Mα p27 TL p27-/- p27-/- Supplemental Figure 1: Altered expression of p27 and p27 in the brain. Immunostaining for p27 was performed on brain frozen sections. Brain sections
SUPPLEMENTARY INFORMATION
 Local auxin metabolism regulates environment-induced hypocotyl elongation Zuyu Zheng 1,2, Yongxia Guo 3, Ondřej Novák 4,5, William Chen 2, Karin Ljung 4, Joseph P. Noel 1,3, *, and Joanne Chory 1,2, *
Local auxin metabolism regulates environment-induced hypocotyl elongation Zuyu Zheng 1,2, Yongxia Guo 3, Ondřej Novák 4,5, William Chen 2, Karin Ljung 4, Joseph P. Noel 1,3, *, and Joanne Chory 1,2, *
Microarrays: since we use probes we obviously must know the sequences we are looking at!
 These background are needed: 1. - Basic Molecular Biology & Genetics DNA replication Transcription Post-transcriptional RNA processing Translation Post-translational protein modification Gene expression
These background are needed: 1. - Basic Molecular Biology & Genetics DNA replication Transcription Post-transcriptional RNA processing Translation Post-translational protein modification Gene expression
Supplementary Figure 1. (A) Cell proliferative ability and (B) the invasiveness of
 LEGEND FOR SUPPLEMENTARY FIGURES Supplementary Figure 1. (A) Cell proliferative ability and (B) the invasiveness of LLC-1, LLC-3, and LLC-5 cell line series were quantified by BrdU assay after 72 h of
LEGEND FOR SUPPLEMENTARY FIGURES Supplementary Figure 1. (A) Cell proliferative ability and (B) the invasiveness of LLC-1, LLC-3, and LLC-5 cell line series were quantified by BrdU assay after 72 h of
