Computer aided modeling of a fructose repressor
|
|
|
- Chad Horn
- 6 years ago
- Views:
Transcription
1 Computer aided modeling of a fructose repressor Miia Helanto and Kristiina Kiviharju Abstract The bioconversion of fructose to mannitol is a commercially interesting bioprocess. Fructose can, however be utilized also otherwise, and this utilization is partly controlled by a fructose repressor (frur). In this study, we calculated a 3D-structure for the frur from the DNA and amino acid sequence and compared it to the structures of other repressors. The programs used in this study were ClustalW, Vectorette II, ORF finder, ExPASy Translate tool, BLASTP, InterProScan, PSIPRED, SWISS-MODEL and DeepView. The results showed rather good homology to other repressors, e.g. purine repressor. Introduction Heterofermentative lactic acid bacteria (LAB) utilize glucose in submerged cultures and produce lactate and ethanol into the culture medium. The ethanol production oxidizes NADH to the form that is needed for the cells to utilize glucose. This oxidization can also be done using other reactions preferably with commercial interest, for instance the reduction of fructose to mannitol. Fructose, however, can also be utilized by the LAB in the glucose pathway by certain enzymes produced when fructose is present in the culture medium, namely fructokinase and glucose-6-phosphate isomerase. The expression of these proteins is regulated by a fructose repressor (FruR). This repressor binds to the 1
2 DNA operator area, when no inducing agent is present in the medium. When an inducer, e.g. fructose is present, the repressor detaches from the DNA and RNA polymerases can begin the synthesis of mrna coding fructokinase and glucose-6-phosphate isomerase. The utilization of fructose in the LAB metabolic activities is not preferable in a process aiming at efficient bioconversion of fructose to mannitol and thus the DNA bound state of the repressor is preferred. As fructose tends to detach the repressor, an anti-inducer with a greater affinity to the enzyme than fructose has, would be valuable. The aim of this study is to compare the structure of the fructose repressor to other repressors and to use databases and computer programs to find out the possible 3D-structure of this enzyme based on its DNA sequence. Methods Cloning and sequencing of fructose operon from Lactobacillus fermentum NRRL-B-1932 Closely related fructokinase sequences from the bacteria Lactococcus lactis sp. lactis, Lc. lactis sp. cremoris, Streptococcus mutans, Pediococcus pentosaceus and Bacillus subtilis were identified using the BLAST search program on the NCBI server. Conserved regions of the fructokinases were then aligned using ClustalW with default settings. Degenerated primers were designed against the most conserved segments of alignment as described by Bartl (1997). A 0.7 kb fragment of L. fermentum fructokinase gene was amplified with PCR using degenerated primer pair, subcloned into cloning vector pgem-t Easy. Plasmids were sequenced by the University of Helsinki Institute of Biotechnology. The Vectorette II system (Sigma Genosys, Ltd., UK) was used to create a genomic library of L. fermentum. Chromosomal DNA of L. fermentum was digested either with BamHI, ClaI, EcoRI or HindIII (New England Biolabs, Beverly, MA), and ligated with 2
3 the respective Vectorette units. Vectorette amplicons were amplified with PCR using sequence specific and Vectorette unit specific primers, subcloned into cloning vector pgm-t Easy, and sequenced. ORF analysis and translation to amino acid sequences Sequence fragments were joined together and analyzed by GCG Wisconsin Package. Open reading frame (ORF) analyses were done with ORF-finder which identifies all possible ORFs in a DNA sequence by locating the standard and alternative stop and start codons. All possible ORFs were translated to amino acid sequences by Expasy Translate tool. Homology and function search The translated amino acid sequence was used in a BLASTP search against Swiss-Protand PDB-databases. The BLAST algorithm is a heuristic search method that seeks words of length W (default = 3 in BLASTP) that score at least T when aligned with the query and scored with a substitution matrix. Words in the database that score T or greater are extended in both directions in an attempt to find a locally optimal ungapped alignment or HSP (high scoring pair) with a score of at least S (alignment score) or an E value (number of hits with a score equal to or better than S that would be expected by chance (the background noise) when searching a database of a particular size) lower than the specified threshold. HSPs that meet these criteria will be reported by BLAST, provided they do not exceed the cutoff value specified for number of descriptions and/or alignments to report. 3
4 Protein function and structure can be determined by comparing so called conserved areas of the amino acid sequence. These are stored in databases e.g. PROSITE, which contains information on over 1000 protein families and domains. There are also programs that do database searches on many databases to find the same information on a broader basis, e.g. InterProScan, which is a European search engine using PROSITE, SwissProt and TrEMBL. In this study we used InterProScan. The program used the amino acid sequence of the FruR to search for similar structures and functions in database proteins. Secondary structure prediction Secondary structure prediction was done by PSIPRED protein structure prediction server. It is a simple and reliable secondary structure prediction method, incorporating two feedforward neural networks which perform an analysis on output obtained from PSI-BLAST (Position Specific Iterated - BLAST). 3D-structure prediction 3D-structure prediction was done by SWISS-MODEL server using the first approach mode. SWISS-MODEL is a server for automated comparative modeling of 3D protein structures. It pioneered the field of automated modeling starting in 1993 and is the most widely-used free web-based automated modeling facility today. Template selection, alignment and model building are done completely automated by the server. The first approach mode provides a simple interface and requires only an amino acid sequence as input data. The server will automatically select suitable templates. Optionally, the user can specify up to five template structures, either from the ExPDB library or uploaded coordinate files. The automated modeling procedure will start if at 4
5 least one modeling template is available that has a sequence identity of more than 25% with the submitted target sequence. However, users need to be aware that the model reliability decreases as the sequence identity decreases and that target-template pairs sharing less than 50% sequence identity may often require manual adjustment of the alignment. In the alignment mode the modeling procedure is initiated by submitting a sequence alignment. The user specifies which sequence in the given alignment is the target sequence and which one corresponds to a structurally known protein chain from the ExPDB template library. The server will build the model based on the given alignment. The project mode allows the user to submit a manually optimized modeling request to the SWISS-MODEL server. The starting point for this mode is a DeepView project file. It contains the superposed template structures, and the alignment between the target and the templates. This mode gives the user control over a wide range of parameters, e.g. template selection or gap placement in the alignment. Furthermore, the project mode can also be used to iteratively improve the output of the first approach mode Modeling procedure (Adapted from Schwede et al. (2003)) All homology-modeling methods consist of the following four steps: (i) template selection; (ii) target template alignment; (iii) model building; and (iv) evaluation. These steps can be iteratively repeated, until a satisfying model structure is achieved. Several different techniques for model building have been developed. The SWISS-MODEL server approach can be described as rigid fragment assembly, which will be outlined briefly. The SWISS-MODEL server template library ExPDB is extracted from the PDB. In order to allow a stable and automated workflow of the server, the PDB coordinate files are split into individual protein chains and unreliable entries, e.g. theoretical models and low quality structures providing only C coordinates, are removed. Additional information 5
6 useful for template selection is gathered and added to the file header, e.g. probable quaternary structure, quality indicators like empirical force field energy or ANOLEA mean force potential scores. To select templates for a given protein, the sequences of the template structure library are searched. If these templates cover distinct regions of the target sequence, the modeling process will be split into separate independent batches. Up to five template structures per batch are superposed using an iterative least squares algorithm. A structural alignment is generated after removing incompatible templates, i.e. omitting structures with high C root mean square deviations to the first template. A local pair-wise alignment of the target sequence to the main template structures is calculated, followed by a heuristic step to improve the alignment for modeling purposes. The placement of insertions and deletions is optimized considering the template structure context. In particular, isolated residues in the alignment ( islands ) are moved to the flanks to facilitate the loop building process. To generate the core of the model, the backbone atom positions of the template structure are averaged. The templates are thereby weighted by their sequence similarity to the target sequence, while significantly deviating atom positions are excluded. The template coordinates cannot be used to model regions of insertions or deletions in the targettemplate alignment. To generate those parts, an ensemble of fragments compatible with the neighboring stems is constructed using constraint space programming (CSP). The best loop is selected using a scoring scheme, which accounts for force field energy, steric hindrance and favorable interactions like hydrogen bond formation. If no suitable loop can be identified, the flanking residues are included to the rebuilt fragment to allow for more flexibility. In cases where CSP does not give a satisfying solution and for loops above 10 residues, a loop library derived from experimental structures is searched to find compatible loop fragments. The reconstruction of the model side chains is based on the weighted positions of corresponding residues in the template structures. Starting with conserved residues, the model side chains are built by iso-sterically replacing template structure side chains. 6
7 Possible side chain conformations are selected from a backbone dependent rotamer library, which has been constructed carefully taking into account the quality of the source structures. A scoring function assessing favorable interactions (hydrogen bonds, disulfide bridges) and unfavorably close contacts is applied to select the most likely conformation. Deviations in the protein structure geometry, which have been introduced by the modeling algorithm when joining rigid fragments are regularized in the last modeling step by steepest descent energy minimization using the GROMOS96 force field. Empirical force fields are useful to detect parts of the model with conformationalerrors. In our own experience and the work of others, energy minimization or molecular dynamics methods are in general not able to improve the accuracy of the models, and are used in SWISS- MODEL only to regularize the structure. However, the successful application of restricted molecular dynamics for improving homology models has recently been reported for a few test cases. To derive more general rules of engagement of molecular dynamics, further systematic experiments have to be conducted. The four modeling steps template superposition, target-template alignment, model building and energy minimization have been implemented in the program ProModII in ANSI C. Comparative study Protein Data Base (PDB) The Protein Data Base (PDB) is a single worldwide repository for the processing and distribution of 3D-structure data of large molecules of proteins and nucleic acids. Repressor protein 3D-structures were downloaded from the PDB database to obtain model structures that can be compared with the frur model calculated by SWISS- MODEL. 7
8 DeepView The protein model was visualized and compared to other repressor structures with the DeepView program. The program DeepView (Swiss-PdbViewer) was designed to integrate functions for protein structure visualization, analysis and manipulation into a sequence-to-structure workbench with a user-friendlyinterface. It allows the user to manage complex modeling projects and is publicly available from the ExPASy server. With DeepView one can search for suitable modeling templates and download the corresponding PDB or ExPDB files directly from the DeepView server. Using the integrated sequence alignment tools and structural superposition algorithms, a target sequence can be mapped onto the modeling templates in one step. Then the initial sequence alignment can be optimized manually while the anticipated changes in the model backbone are reflected in real-time in the displayed structural superposition. The complete project file is then submitted to the SWISS-MODEL server for model building. The resulting protein model can be visualized and analyzed using the integrated tools in DeepView program. Results Analysis of the amino acid sequence The ORF finder found three open reading frames. The amino acid sequence analysis by BLAST showed that the operon consisted of two different genes coding fructokinase and phoshoglucose isomerase and their regulator protein frur. The analysis resulted in a too short frur sequence; the HTH motif was left away. This might have occurred because of the repressors transcription initiation codon TTG which was not recognized by the ORF finder program. In the literature we found that the HTH-motif was essential for a 8
9 functional repressor protein. It is the region that binds to DNA. The DNA sequence was then extended so that the HTH-motif coding region was also included in the frur sequence before further analysis. The ExPASy Translate tool found only one ORF that could code the amino acid sequence of the frur. This sequence was used for the rest of the study. Homology and function The InterProScan search result is shown in Figure 1. The program identified the beginning of the amino acid sequence as a Helix-Turn-Helix (HTH) motif, which classifies the protein as a LacI family protein. The LacI family consists of various regulatory proteins, such as ascg, ccpa, cytr, galr, laci, opnr, purf and scrr (Adhya and Weickert, 1992). The HTH motif is situated towards the N-terminus in the proteins of this family. Secondary structure prediction The predicted secondary structure obtained from PSIPRED is shown in Figure 2. The green tubes represent α-helixes, yellow arrows β-sheets and strands loop areas. The columns above the structure represent the probability of the structure. 3D-structure prediction The predicted 3D-structure of frur is shown in Figure 3. The α-helixes are shown in red and β-sheets in yellow. The blue-gray areas are loops or uncertainties of the model. The 9
10 frur has a headpiece containing three α-helixes (helix-turn-helix motif and hinge helix motif). The most probable active site is surrounded by the N-subdomain and C- subdomain, both containing α-helixes and β-sheets as well as uncertain structure. Figure 1. InterProScan results from the analysis of the frur amino acid sequence. Comparative study Figure 4 shows the surface models of frur and purr. The α-helixes are shown in red and β-sheets in yellow. Loops and uncertain areas of the model are shown in gray. Structural similarity of the two repressor proteins can clearly be observed. 10
11 Figure 2. The predicted secondary structure of the frur. The green tubes represent α-helixes and yellow arrows β-sheets. 11
12 Figure 3. SWISS-MODEL graphical representation of the frur. The red areas represent α-helixes and yellow areas β-sheets. The blue-gray areas are loops or uncertainties of the model. Figure 4. Surface models of the fructose repressor (left) and purine repressor (right). The purine repressor has a DNA strand attached to the headpiece. The red areas represent α-helixes and yellow areas β-sheets. Gray areas are loops or uncertainties of the model. 12
13 Discussion The predicted 3D-structure of the L. fermentum frur repressor protein still has many uncertain areas that will need more accurate modeling. Loop structures are normally very difficult to model, as they are quite flexible structures. Certain amino acids, like glycine and alanine, that are very small, can move easily and thus can not form rigid structures compared to the bigger amino acids, e.g. proline. The next step in modeling could be comparing the automatically predicted repressor model to known 3D-structures manually, one amino acid at a time, by using a superposition program. This can be done in a protein modeling program, e.g. Quanta. The model obtained in this work shows that this frur is a repressor with a typical DNA binding site. The model of the active site is still too uncertain for even guessing suitable inducers or anti-inducers that could be used for improvement of mannitol yield from fructose. References Adhya, S. and Weickert, M.J., A family of bacterial regulators homologous to gal and lac repressors, J. Biol. Chem. 267 (1992) BLAST, Bartl, S., Amplification using degenerate primers with multiple inosines to isolate genes with minimal sequence similarity. In: Methods in molecular biology, Vol 67: PCR cloning protocols: from molecular cloning to genetic engineering, Ed. B.A.White, Humana Press Inc., Totowa, NJ 1997, p
14 ExPASy Translate tool, InterProScan, ORF-finder, PROSITE, Protein Data Bank, PsiPred, Schwede, T., Kopp, J., Guex, N., and Peitsch, M.C., SWISS-MODEL: an automated protein homology-modeling server, Nucl. Acids Res. 31 (2003) SwissModel,
Multiple choice questions (numbers in brackets indicate the number of correct answers)
 1 Multiple choice questions (numbers in brackets indicate the number of correct answers) February 1, 2013 1. Ribose is found in Nucleic acids Proteins Lipids RNA DNA (2) 2. Most RNA in cells is transfer
1 Multiple choice questions (numbers in brackets indicate the number of correct answers) February 1, 2013 1. Ribose is found in Nucleic acids Proteins Lipids RNA DNA (2) 2. Most RNA in cells is transfer
3 Designing Primers for Site-Directed Mutagenesis
 3 Designing Primers for Site-Directed Mutagenesis 3.1 Learning Objectives During the next two labs you will learn the basics of site-directed mutagenesis: you will design primers for the mutants you designed
3 Designing Primers for Site-Directed Mutagenesis 3.1 Learning Objectives During the next two labs you will learn the basics of site-directed mutagenesis: you will design primers for the mutants you designed
DNA Structure and Properties Basic Properties Predicting Melting Temperature. Dinesh Yadav
 DNA Structure and Properties Basic Properties Predicting Melting Temperature Dinesh Yadav Nucleic Acid Structure Question: Is this RNA or DNA? Molecules of Life, pp. 15 2 Nucleic Acid Bases Molecules of
DNA Structure and Properties Basic Properties Predicting Melting Temperature Dinesh Yadav Nucleic Acid Structure Question: Is this RNA or DNA? Molecules of Life, pp. 15 2 Nucleic Acid Bases Molecules of
Lecture Four. Molecular Approaches I: Nucleic Acids
 Lecture Four. Molecular Approaches I: Nucleic Acids I. Recombinant DNA and Gene Cloning Recombinant DNA is DNA that has been created artificially. DNA from two or more sources is incorporated into a single
Lecture Four. Molecular Approaches I: Nucleic Acids I. Recombinant DNA and Gene Cloning Recombinant DNA is DNA that has been created artificially. DNA from two or more sources is incorporated into a single
Antisense RNA Insert Design for Plasmid Construction to Knockdown Target Gene Expression
 Vol. 1:7-15 Antisense RNA Insert Design for Plasmid Construction to Knockdown Target Gene Expression Ji, Tom, Lu, Aneka, Wu, Kaylee Department of Microbiology and Immunology, University of British Columbia
Vol. 1:7-15 Antisense RNA Insert Design for Plasmid Construction to Knockdown Target Gene Expression Ji, Tom, Lu, Aneka, Wu, Kaylee Department of Microbiology and Immunology, University of British Columbia
Synthetic Biology for
 Synthetic Biology for Plasmids and DNA Digestion Plasmids Plasmids are small DNA molecules that are separate from chromosomal DNA They are most commonly found as double stranded, circular DNA Typical plasmids
Synthetic Biology for Plasmids and DNA Digestion Plasmids Plasmids are small DNA molecules that are separate from chromosomal DNA They are most commonly found as double stranded, circular DNA Typical plasmids
CHAPTER 20 DNA TECHNOLOGY AND GENOMICS. Section A: DNA Cloning
 Section A: DNA Cloning 1. DNA technology makes it possible to clone genes for basic research and commercial applications: an overview 2. Restriction enzymes are used to make recombinant DNA 3. Genes can
Section A: DNA Cloning 1. DNA technology makes it possible to clone genes for basic research and commercial applications: an overview 2. Restriction enzymes are used to make recombinant DNA 3. Genes can
DNA Technology. Asilomar Singer, Zinder, Brenner, Berg
 DNA Technology Asilomar 1973. Singer, Zinder, Brenner, Berg DNA Technology The following are some of the most important molecular methods we will be using in this course. They will be used, among other
DNA Technology Asilomar 1973. Singer, Zinder, Brenner, Berg DNA Technology The following are some of the most important molecular methods we will be using in this course. They will be used, among other
Prokaryotic Transcription
 Prokaryotic Transcription Transcription Basics DNA is the genetic material Nucleic acid Capable of self-replication and synthesis of RNA RNA is the middle man Nucleic acid Structure and base sequence are
Prokaryotic Transcription Transcription Basics DNA is the genetic material Nucleic acid Capable of self-replication and synthesis of RNA RNA is the middle man Nucleic acid Structure and base sequence are
Sequence Based Function Annotation. Qi Sun Bioinformatics Facility Biotechnology Resource Center Cornell University
 Sequence Based Function Annotation Qi Sun Bioinformatics Facility Biotechnology Resource Center Cornell University Usage scenarios for sequence based function annotation Function prediction of newly cloned
Sequence Based Function Annotation Qi Sun Bioinformatics Facility Biotechnology Resource Center Cornell University Usage scenarios for sequence based function annotation Function prediction of newly cloned
Sequence Databases and database scanning
 Sequence Databases and database scanning Marjolein Thunnissen Lund, 2012 Types of databases: Primary sequence databases (proteins and nucleic acids). Composite protein sequence databases. Secondary databases.
Sequence Databases and database scanning Marjolein Thunnissen Lund, 2012 Types of databases: Primary sequence databases (proteins and nucleic acids). Composite protein sequence databases. Secondary databases.
Chapter 8 DNA Recognition in Prokaryotes by Helix-Turn-Helix Motifs
 Chapter 8 DNA Recognition in Prokaryotes by Helix-Turn-Helix Motifs 1. Helix-turn-helix proteins 2. Zinc finger proteins 3. Leucine zipper proteins 4. Beta-scaffold factors 5. Others λ-repressor AND CRO
Chapter 8 DNA Recognition in Prokaryotes by Helix-Turn-Helix Motifs 1. Helix-turn-helix proteins 2. Zinc finger proteins 3. Leucine zipper proteins 4. Beta-scaffold factors 5. Others λ-repressor AND CRO
DNA Structure and Analysis. Chapter 4: Background
 DNA Structure and Analysis Chapter 4: Background Molecular Biology Three main disciplines of biotechnology Biochemistry Genetics Molecular Biology # Biotechnology: A Laboratory Skills Course explorer.bio-rad.com
DNA Structure and Analysis Chapter 4: Background Molecular Biology Three main disciplines of biotechnology Biochemistry Genetics Molecular Biology # Biotechnology: A Laboratory Skills Course explorer.bio-rad.com
BIOLOGY LTF DIAGNOSTIC TEST DNA to PROTEIN & BIOTECHNOLOGY
 Biology Multiple Choice 016074 BIOLOGY LTF DIAGNOSTIC TEST DNA to PROTEIN & BIOTECHNOLOGY Test Code: 016074 Directions: Each of the questions or incomplete statements below is followed by five suggested
Biology Multiple Choice 016074 BIOLOGY LTF DIAGNOSTIC TEST DNA to PROTEIN & BIOTECHNOLOGY Test Code: 016074 Directions: Each of the questions or incomplete statements below is followed by five suggested
Database Searching and BLAST Dannie Durand
 Computational Genomics and Molecular Biology, Fall 2013 1 Database Searching and BLAST Dannie Durand Tuesday, October 8th Review: Karlin-Altschul Statistics Recall that a Maximal Segment Pair (MSP) is
Computational Genomics and Molecular Biology, Fall 2013 1 Database Searching and BLAST Dannie Durand Tuesday, October 8th Review: Karlin-Altschul Statistics Recall that a Maximal Segment Pair (MSP) is
How Do You Clone a Gene?
 S-20 Edvo-Kit #S-20 How Do You Clone a Gene? Experiment Objective: The objective of this experiment is to gain an understanding of the structure of DNA, a genetically engineered clone, and how genes are
S-20 Edvo-Kit #S-20 How Do You Clone a Gene? Experiment Objective: The objective of this experiment is to gain an understanding of the structure of DNA, a genetically engineered clone, and how genes are
translation The building blocks of proteins are? amino acids nitrogen containing bases like A, G, T, C, and U Complementary base pairing links
 The actual process of assembling the proteins on the ribosome is called? translation The building blocks of proteins are? Complementary base pairing links Define and name the Purines amino acids nitrogen
The actual process of assembling the proteins on the ribosome is called? translation The building blocks of proteins are? Complementary base pairing links Define and name the Purines amino acids nitrogen
Why learn sequence database searching? Searching Molecular Databases with BLAST
 Why learn sequence database searching? Searching Molecular Databases with BLAST What have I cloned? Is this really!my gene"? Basic Local Alignment Search Tool How BLAST works Interpreting search results
Why learn sequence database searching? Searching Molecular Databases with BLAST What have I cloned? Is this really!my gene"? Basic Local Alignment Search Tool How BLAST works Interpreting search results
Genome Sequence Assembly
 Genome Sequence Assembly Learning Goals: Introduce the field of bioinformatics Familiarize the student with performing sequence alignments Understand the assembly process in genome sequencing Introduction:
Genome Sequence Assembly Learning Goals: Introduce the field of bioinformatics Familiarize the student with performing sequence alignments Understand the assembly process in genome sequencing Introduction:
Ch. 10 Notes DNA: Transcription and Translation
 Ch. 10 Notes DNA: Transcription and Translation GOALS Compare the structure of RNA with that of DNA Summarize the process of transcription Relate the role of codons to the sequence of amino acids that
Ch. 10 Notes DNA: Transcription and Translation GOALS Compare the structure of RNA with that of DNA Summarize the process of transcription Relate the role of codons to the sequence of amino acids that
Bio11 Announcements. Ch 21: DNA Biology and Technology. DNA Functions. DNA and RNA Structure. How do DNA and RNA differ? What are genes?
 Bio11 Announcements TODAY Genetics (review) and quiz (CP #4) Structure and function of DNA Extra credit due today Next week in lab: Case study presentations Following week: Lab Quiz 2 Ch 21: DNA Biology
Bio11 Announcements TODAY Genetics (review) and quiz (CP #4) Structure and function of DNA Extra credit due today Next week in lab: Case study presentations Following week: Lab Quiz 2 Ch 21: DNA Biology
Product Applications for the Sequence Analysis Collection
 Product Applications for the Sequence Analysis Collection Pipeline Pilot Contents Introduction... 1 Pipeline Pilot and Bioinformatics... 2 Sequence Searching with Profile HMM...2 Integrating Data in a
Product Applications for the Sequence Analysis Collection Pipeline Pilot Contents Introduction... 1 Pipeline Pilot and Bioinformatics... 2 Sequence Searching with Profile HMM...2 Integrating Data in a
Nucleic acids deoxyribonucleic acid (DNA) ribonucleic acid (RNA) nucleotide
 Nucleic Acids Nucleic acids are molecules that store information for cellular growth and reproduction There are two types of nucleic acids: - deoxyribonucleic acid (DNA) and ribonucleic acid (RNA) These
Nucleic Acids Nucleic acids are molecules that store information for cellular growth and reproduction There are two types of nucleic acids: - deoxyribonucleic acid (DNA) and ribonucleic acid (RNA) These
Lecture 2: Central Dogma of Molecular Biology & Intro to Programming
 Lecture 2: Central Dogma of Molecular Biology & Intro to Programming Central Dogma of Molecular Biology Proteins: workhorse molecules of biological systems Proteins are synthesized from the genetic blueprints
Lecture 2: Central Dogma of Molecular Biology & Intro to Programming Central Dogma of Molecular Biology Proteins: workhorse molecules of biological systems Proteins are synthesized from the genetic blueprints
CSE : Computational Issues in Molecular Biology. Lecture 19. Spring 2004
 CSE 397-497: Computational Issues in Molecular Biology Lecture 19 Spring 2004-1- Protein structure Primary structure of protein is determined by number and order of amino acids within polypeptide chain.
CSE 397-497: Computational Issues in Molecular Biology Lecture 19 Spring 2004-1- Protein structure Primary structure of protein is determined by number and order of amino acids within polypeptide chain.
DNA is the genetic material. DNA structure. Chapter 7: DNA Replication, Transcription & Translation; Mutations & Ames test
 DNA is the genetic material Chapter 7: DNA Replication, Transcription & Translation; Mutations & Ames test Dr. Amy Rogers Bio 139 General Microbiology Hereditary information is carried by DNA Griffith/Avery
DNA is the genetic material Chapter 7: DNA Replication, Transcription & Translation; Mutations & Ames test Dr. Amy Rogers Bio 139 General Microbiology Hereditary information is carried by DNA Griffith/Avery
Computational Biology I LSM5191
 Computational Biology I LSM5191 Lecture 5 Notes: Genetic manipulation & Molecular Biology techniques Broad Overview of: Enzymatic tools in Molecular Biology Gel electrophoresis Restriction mapping DNA
Computational Biology I LSM5191 Lecture 5 Notes: Genetic manipulation & Molecular Biology techniques Broad Overview of: Enzymatic tools in Molecular Biology Gel electrophoresis Restriction mapping DNA
CHAPTER 9 DNA Technologies
 CHAPTER 9 DNA Technologies Recombinant DNA Artificially created DNA that combines sequences that do not occur together in the nature Basis of much of the modern molecular biology Molecular cloning of genes
CHAPTER 9 DNA Technologies Recombinant DNA Artificially created DNA that combines sequences that do not occur together in the nature Basis of much of the modern molecular biology Molecular cloning of genes
MIT Department of Biology 7.013: Introductory Biology - Spring 2005 Instructors: Professor Hazel Sive, Professor Tyler Jacks, Dr.
 MIT Department of Biology 7.01: Introductory Biology - Spring 2005 Instructors: Professor Hazel Sive, Professor Tyler Jacks, Dr. Claudette Gardel iv) Would Xba I be useful for cloning? Why or why not?
MIT Department of Biology 7.01: Introductory Biology - Spring 2005 Instructors: Professor Hazel Sive, Professor Tyler Jacks, Dr. Claudette Gardel iv) Would Xba I be useful for cloning? Why or why not?
Higher Human Biology Unit 1: Human Cells Pupils Learning Outcomes
 Higher Human Biology Unit 1: Human Cells Pupils Learning Outcomes 1.1 Division and Differentiation in Human Cells I can state that cellular differentiation is the process by which a cell develops more
Higher Human Biology Unit 1: Human Cells Pupils Learning Outcomes 1.1 Division and Differentiation in Human Cells I can state that cellular differentiation is the process by which a cell develops more
Self-test Quiz for Chapter 12 (From DNA to Protein: Genotype to Phenotype)
 Self-test Quiz for Chapter 12 (From DNA to Protein: Genotype to Phenotype) Question#1: One-Gene, One-Polypeptide The figure below shows the results of feeding trials with one auxotroph strain of Neurospora
Self-test Quiz for Chapter 12 (From DNA to Protein: Genotype to Phenotype) Question#1: One-Gene, One-Polypeptide The figure below shows the results of feeding trials with one auxotroph strain of Neurospora
Lecture 3 (FW) January 28, 2009 Cloning of DNA; PCR amplification Reading assignment: Cloning, ; ; 330 PCR, ; 329.
 Lecture 3 (FW) January 28, 2009 Cloning of DNA; PCR amplification Reading assignment: Cloning, 240-245; 286-87; 330 PCR, 270-274; 329. Take Home Lesson(s) from Lecture 2: 1. DNA is a double helix of complementary
Lecture 3 (FW) January 28, 2009 Cloning of DNA; PCR amplification Reading assignment: Cloning, 240-245; 286-87; 330 PCR, 270-274; 329. Take Home Lesson(s) from Lecture 2: 1. DNA is a double helix of complementary
Answers to Module 1. An obligate aerobe is an organism that has an absolute requirement of oxygen for growth.
 Answers to Module 1 Short Answers 1) What is an obligate aerobe? An obligate aerobe is an organism that has an absolute requirement of oxygen for growth. What about facultative anaerobe? 2) Distinguish
Answers to Module 1 Short Answers 1) What is an obligate aerobe? An obligate aerobe is an organism that has an absolute requirement of oxygen for growth. What about facultative anaerobe? 2) Distinguish
Protein Bioinformatics Part I: Access to information
 Protein Bioinformatics Part I: Access to information 260.655 April 6, 2006 Jonathan Pevsner, Ph.D. pevsner@kennedykrieger.org Outline [1] Proteins at NCBI RefSeq accession numbers Cn3D to visualize structures
Protein Bioinformatics Part I: Access to information 260.655 April 6, 2006 Jonathan Pevsner, Ph.D. pevsner@kennedykrieger.org Outline [1] Proteins at NCBI RefSeq accession numbers Cn3D to visualize structures
Adv Biology: DNA and RNA Study Guide
 Adv Biology: DNA and RNA Study Guide Chapter 12 Vocabulary -Notes What experiments led up to the discovery of DNA being the hereditary material? o The discovery that DNA is the genetic code involved many
Adv Biology: DNA and RNA Study Guide Chapter 12 Vocabulary -Notes What experiments led up to the discovery of DNA being the hereditary material? o The discovery that DNA is the genetic code involved many
Microbiology: The Blueprint of Life, from DNA to protein
 Microbiology: The Blueprint of Life, from DNA to protein I. Overview A. DNA ultimately determines every aspect of a cell from shape to function 1. DNA = 2. Nucleotides of DNA have three units a. A nitrogen-containing
Microbiology: The Blueprint of Life, from DNA to protein I. Overview A. DNA ultimately determines every aspect of a cell from shape to function 1. DNA = 2. Nucleotides of DNA have three units a. A nitrogen-containing
Name Class Date. Practice Test
 Name Class Date 12 DNA Practice Test Multiple Choice Write the letter that best answers the question or completes the statement on the line provided. 1. What do bacteriophages infect? a. mice. c. viruses.
Name Class Date 12 DNA Practice Test Multiple Choice Write the letter that best answers the question or completes the statement on the line provided. 1. What do bacteriophages infect? a. mice. c. viruses.
Protein Synthesis
 HEBISD Student Expectations: Identify that RNA Is a nucleic acid with a single strand of nucleotides Contains the 5-carbon sugar ribose Contains the nitrogen bases A, G, C and U instead of T. The U is
HEBISD Student Expectations: Identify that RNA Is a nucleic acid with a single strand of nucleotides Contains the 5-carbon sugar ribose Contains the nitrogen bases A, G, C and U instead of T. The U is
Bio 101 Sample questions: Chapter 10
 Bio 101 Sample questions: Chapter 10 1. Which of the following is NOT needed for DNA replication? A. nucleotides B. ribosomes C. Enzymes (like polymerases) D. DNA E. all of the above are needed 2 The information
Bio 101 Sample questions: Chapter 10 1. Which of the following is NOT needed for DNA replication? A. nucleotides B. ribosomes C. Enzymes (like polymerases) D. DNA E. all of the above are needed 2 The information
REGULATION OF GENE EXPRESSION
 REGULATION OF GENE EXPRESSION Each cell of a living organism contains thousands of genes. But all genes do not function at a time. Genes function according to requirements of the cell. Genes control the
REGULATION OF GENE EXPRESSION Each cell of a living organism contains thousands of genes. But all genes do not function at a time. Genes function according to requirements of the cell. Genes control the
Genetics Lecture 21 Recombinant DNA
 Genetics Lecture 21 Recombinant DNA Recombinant DNA In 1971, a paper published by Kathleen Danna and Daniel Nathans marked the beginning of the recombinant DNA era. The paper described the isolation of
Genetics Lecture 21 Recombinant DNA Recombinant DNA In 1971, a paper published by Kathleen Danna and Daniel Nathans marked the beginning of the recombinant DNA era. The paper described the isolation of
AP Biology Gene Expression/Biotechnology REVIEW
 AP Biology Gene Expression/Biotechnology REVIEW Multiple Choice Identify the choice that best completes the statement or answers the question. 1. Gene expression can be a. regulated before transcription.
AP Biology Gene Expression/Biotechnology REVIEW Multiple Choice Identify the choice that best completes the statement or answers the question. 1. Gene expression can be a. regulated before transcription.
Molecular Cell Biology - Problem Drill 11: Recombinant DNA
 Molecular Cell Biology - Problem Drill 11: Recombinant DNA Question No. 1 of 10 1. Which of the following statements about the sources of DNA used for molecular cloning is correct? Question #1 (A) cdna
Molecular Cell Biology - Problem Drill 11: Recombinant DNA Question No. 1 of 10 1. Which of the following statements about the sources of DNA used for molecular cloning is correct? Question #1 (A) cdna
Nucleic acids and protein synthesis
 THE FUNCTIONS OF DNA Nucleic acids and protein synthesis The full name of DNA is deoxyribonucleic acid. Every nucleotide has the same sugar molecule and phosphate group, but each nucleotide contains one
THE FUNCTIONS OF DNA Nucleic acids and protein synthesis The full name of DNA is deoxyribonucleic acid. Every nucleotide has the same sugar molecule and phosphate group, but each nucleotide contains one
DNA. translation. base pairing rules for DNA Replication. thymine. cytosine. amino acids. The building blocks of proteins are?
 2 strands, has the 5-carbon sugar deoxyribose, and has the nitrogen base Thymine. The actual process of assembling the proteins on the ribosome is called? DNA translation Adenine pairs with Thymine, Thymine
2 strands, has the 5-carbon sugar deoxyribose, and has the nitrogen base Thymine. The actual process of assembling the proteins on the ribosome is called? DNA translation Adenine pairs with Thymine, Thymine
BLAST. compared with database sequences Sequences with many matches to high- scoring words are used for final alignments
 BLAST 100 times faster than dynamic programming. Good for database searches. Derive a list of words of length w from query (e.g., 3 for protein, 11 for DNA) High-scoring words are compared with database
BLAST 100 times faster than dynamic programming. Good for database searches. Derive a list of words of length w from query (e.g., 3 for protein, 11 for DNA) High-scoring words are compared with database
ONLINE BIOINFORMATICS RESOURCES
 Dedan Githae Email: d.githae@cgiar.org BecA-ILRI Hub; Nairobi, Kenya 16 May, 2014 ONLINE BIOINFORMATICS RESOURCES Introduction to Molecular Biology and Bioinformatics (IMBB) 2014 The larger picture.. Lower
Dedan Githae Email: d.githae@cgiar.org BecA-ILRI Hub; Nairobi, Kenya 16 May, 2014 ONLINE BIOINFORMATICS RESOURCES Introduction to Molecular Biology and Bioinformatics (IMBB) 2014 The larger picture.. Lower
Protein Synthesis Notes
 Protein Synthesis Notes Protein Synthesis: Overview Transcription: synthesis of mrna under the direction of DNA. Translation: actual synthesis of a polypeptide under the direction of mrna. Transcription
Protein Synthesis Notes Protein Synthesis: Overview Transcription: synthesis of mrna under the direction of DNA. Translation: actual synthesis of a polypeptide under the direction of mrna. Transcription
Synthetic Biology. Sustainable Energy. Therapeutics Industrial Enzymes. Agriculture. Accelerating Discoveries, Expanding Possibilities. Design.
 Synthetic Biology Accelerating Discoveries, Expanding Possibilities Sustainable Energy Therapeutics Industrial Enzymes Agriculture Design Build Generate Solutions to Advance Synthetic Biology Research
Synthetic Biology Accelerating Discoveries, Expanding Possibilities Sustainable Energy Therapeutics Industrial Enzymes Agriculture Design Build Generate Solutions to Advance Synthetic Biology Research
MATH 5610, Computational Biology
 MATH 5610, Computational Biology Lecture 2 Intro to Molecular Biology (cont) Stephen Billups University of Colorado at Denver MATH 5610, Computational Biology p.1/24 Announcements Error on syllabus Class
MATH 5610, Computational Biology Lecture 2 Intro to Molecular Biology (cont) Stephen Billups University of Colorado at Denver MATH 5610, Computational Biology p.1/24 Announcements Error on syllabus Class
GENE REGULATION IN PROKARYOTES
 GENE REGULATION IN PROKARYOTES Prepared by Brenda Leady, University of Toledo Copyright (c) The McGraw-Hill Companies, Inc. Permission required for reproduction or display. 1 Gene regulation refers to
GENE REGULATION IN PROKARYOTES Prepared by Brenda Leady, University of Toledo Copyright (c) The McGraw-Hill Companies, Inc. Permission required for reproduction or display. 1 Gene regulation refers to
Manipulating DNA. Nucleic acids are chemically different from other macromolecules such as proteins and carbohydrates.
 Lesson Overview 14.3 Studying the Human Genome Nucleic acids are chemically different from other macromolecules such as proteins and carbohydrates. Nucleic acids are chemically different from other macromolecules
Lesson Overview 14.3 Studying the Human Genome Nucleic acids are chemically different from other macromolecules such as proteins and carbohydrates. Nucleic acids are chemically different from other macromolecules
From Gene to Protein Transcription and Translation i
 How do genes influence our characteristics? From Gene to Protein Transcription and Translation i A gene is a segment of DNA that provides the instructions for making a protein. Proteins have many different
How do genes influence our characteristics? From Gene to Protein Transcription and Translation i A gene is a segment of DNA that provides the instructions for making a protein. Proteins have many different
DNA & Protein Synthesis #21
 Name: Period: Date: Living Environment Lab DNA & Protein Synthesis #21 Introduction Of all the molecules that is in the body, DNA is perhaps the most important. DNA or dioxiribosenucleic acid is important
Name: Period: Date: Living Environment Lab DNA & Protein Synthesis #21 Introduction Of all the molecules that is in the body, DNA is perhaps the most important. DNA or dioxiribosenucleic acid is important
From Gene to Protein Transcription and Translation
 Name: Hour: From Gene to Protein Transcription and Translation Introduction: In this activity you will learn how the genes in our DNA influence our characteristics. For example, how can a gene cause albinism
Name: Hour: From Gene to Protein Transcription and Translation Introduction: In this activity you will learn how the genes in our DNA influence our characteristics. For example, how can a gene cause albinism
2. From the first paragraph in this section, find three ways in which RNA differs from DNA.
 Name Chapter 17: From Gene to Protein Begin reading at page 328 Basic Principles of Transcription and Translation. Work on this chapter a single concept at a time, and expect to spend at least 6 hours
Name Chapter 17: From Gene to Protein Begin reading at page 328 Basic Principles of Transcription and Translation. Work on this chapter a single concept at a time, and expect to spend at least 6 hours
Chapter 13 - Concept Mapping
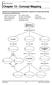 Chapter 13 - Concept Mapping Using the terms and phrases provided below, complete the concept map showing the discovery of DNA structure. amount of base pairs five-carbon sugar purine DNA polymerases Franklin
Chapter 13 - Concept Mapping Using the terms and phrases provided below, complete the concept map showing the discovery of DNA structure. amount of base pairs five-carbon sugar purine DNA polymerases Franklin
Study Guide for Chapter 12 Exam DNA, RNA, & Protein Synthesis
 Name: Date: Period: Study Guide for Chapter 12 Exam DNA, RNA, & Protein Synthesis ***Completing this study guide in its entirety will result in extra credit on the exam. You must show me the DAY OF the
Name: Date: Period: Study Guide for Chapter 12 Exam DNA, RNA, & Protein Synthesis ***Completing this study guide in its entirety will result in extra credit on the exam. You must show me the DAY OF the
Genomics and Gene Recognition Genes and Blue Genes
 Genomics and Gene Recognition Genes and Blue Genes November 1, 2004 Prokaryotic Gene Structure prokaryotes are simplest free-living organisms studying prokaryotes can give us a sense what is the minimum
Genomics and Gene Recognition Genes and Blue Genes November 1, 2004 Prokaryotic Gene Structure prokaryotes are simplest free-living organisms studying prokaryotes can give us a sense what is the minimum
Sequence Analysis Lab Protocol
 Sequence Analysis Lab Protocol You will need this handout of instructions The sequence of your plasmid from the ABI The Accession number for Lambda DNA J02459 The Accession number for puc 18 is L09136
Sequence Analysis Lab Protocol You will need this handout of instructions The sequence of your plasmid from the ABI The Accession number for Lambda DNA J02459 The Accession number for puc 18 is L09136
Fig Ch 17: From Gene to Protein
 Fig. 17-1 Ch 17: From Gene to Protein Basic Principles of Transcription and Translation RNA is the intermediate between genes and the proteins for which they code Transcription is the synthesis of RNA
Fig. 17-1 Ch 17: From Gene to Protein Basic Principles of Transcription and Translation RNA is the intermediate between genes and the proteins for which they code Transcription is the synthesis of RNA
Student Exploration: RNA and Protein Synthesis Due Wednesday 11/27/13
 http://www.explorelearning.com Name: Period : Student Exploration: RNA and Protein Synthesis Due Wednesday 11/27/13 Vocabulary: Define these terms in complete sentences on a separate piece of paper: amino
http://www.explorelearning.com Name: Period : Student Exploration: RNA and Protein Synthesis Due Wednesday 11/27/13 Vocabulary: Define these terms in complete sentences on a separate piece of paper: amino
PUC Vikasana Program- 2012
 Chromosome Nucleus DNA PUC Vikasana Program- 2012 Introduction Molecular biology is the study of biology at a molecular level. Macromolecules and the macromolecular mechanisms. Interactions between the
Chromosome Nucleus DNA PUC Vikasana Program- 2012 Introduction Molecular biology is the study of biology at a molecular level. Macromolecules and the macromolecular mechanisms. Interactions between the
8/21/2014. From Gene to Protein
 From Gene to Protein Chapter 17 Objectives Describe the contributions made by Garrod, Beadle, and Tatum to our understanding of the relationship between genes and enzymes Briefly explain how information
From Gene to Protein Chapter 17 Objectives Describe the contributions made by Garrod, Beadle, and Tatum to our understanding of the relationship between genes and enzymes Briefly explain how information
2054, Chap. 14, page 1
 2054, Chap. 14, page 1 I. Recombinant DNA technology (Chapter 14) A. recombinant DNA technology = collection of methods used to perform genetic engineering 1. genetic engineering = deliberate modification
2054, Chap. 14, page 1 I. Recombinant DNA technology (Chapter 14) A. recombinant DNA technology = collection of methods used to perform genetic engineering 1. genetic engineering = deliberate modification
Structural Bioinformatics (C3210) DNA and RNA Structure
 Structural Bioinformatics (C3210) DNA and RNA Structure Importance of DNA/RNA 3D Structure Nucleic acids are essential materials found in all living organisms. Their main function is to maintain and transmit
Structural Bioinformatics (C3210) DNA and RNA Structure Importance of DNA/RNA 3D Structure Nucleic acids are essential materials found in all living organisms. Their main function is to maintain and transmit
Non-Organic-Based Isolation of Mammalian microrna using Norgen s microrna Purification Kit
 Application Note 13 RNA Sample Preparation Non-Organic-Based Isolation of Mammalian microrna using Norgen s microrna Purification Kit B. Lam, PhD 1, P. Roberts, MSc 1 Y. Haj-Ahmad, M.Sc., Ph.D 1,2 1 Norgen
Application Note 13 RNA Sample Preparation Non-Organic-Based Isolation of Mammalian microrna using Norgen s microrna Purification Kit B. Lam, PhD 1, P. Roberts, MSc 1 Y. Haj-Ahmad, M.Sc., Ph.D 1,2 1 Norgen
DNA/RNA STUDY GUIDE. Match the following scientists with their accomplishments in discovering DNA using the statement in the box below.
 Name: Period: Date: DNA/RNA STUDY GUIDE Part A: DNA History Match the following scientists with their accomplishments in discovering DNA using the statement in the box below. Used a technique called x-ray
Name: Period: Date: DNA/RNA STUDY GUIDE Part A: DNA History Match the following scientists with their accomplishments in discovering DNA using the statement in the box below. Used a technique called x-ray
Recombinant DNA Technology
 History of recombinant DNA technology Recombinant DNA Technology (DNA cloning) Majid Mojarrad Recombinant DNA technology is one of the recent advances in biotechnology, which was developed by two scientists
History of recombinant DNA technology Recombinant DNA Technology (DNA cloning) Majid Mojarrad Recombinant DNA technology is one of the recent advances in biotechnology, which was developed by two scientists
SAMPLE LITERATURE Please refer to included weblink for correct version.
 Edvo-Kit #340 DNA Informatics Experiment Objective: In this experiment, students will explore the popular bioninformatics tool BLAST. First they will read sequences from autoradiographs of automated gel
Edvo-Kit #340 DNA Informatics Experiment Objective: In this experiment, students will explore the popular bioninformatics tool BLAST. First they will read sequences from autoradiographs of automated gel
Genetic Engineering for Biofuels Production
 Genetic Engineering for Biofuels Production WSE 573 Spring 2013 Greeley Beck INTRODUCTION Alternative transportation fuels are needed in the United States because of oil supply insecurity, oil price increases,
Genetic Engineering for Biofuels Production WSE 573 Spring 2013 Greeley Beck INTRODUCTION Alternative transportation fuels are needed in the United States because of oil supply insecurity, oil price increases,
RNA : functional role
 RNA : functional role Hamad Yaseen, PhD MLS Department, FAHS Hamad.ali@hsc.edu.kw RNA mrna rrna trna 1 From DNA to Protein -Outline- From DNA to RNA From RNA to Protein From DNA to RNA Transcription: Copying
RNA : functional role Hamad Yaseen, PhD MLS Department, FAHS Hamad.ali@hsc.edu.kw RNA mrna rrna trna 1 From DNA to Protein -Outline- From DNA to RNA From RNA to Protein From DNA to RNA Transcription: Copying
Comparative Bioinformatics. BSCI348S Fall 2003 Midterm 1
 BSCI348S Fall 2003 Midterm 1 Multiple Choice: select the single best answer to the question or completion of the phrase. (5 points each) 1. The field of bioinformatics a. uses biomimetic algorithms to
BSCI348S Fall 2003 Midterm 1 Multiple Choice: select the single best answer to the question or completion of the phrase. (5 points each) 1. The field of bioinformatics a. uses biomimetic algorithms to
Gene Expression Transcription/Translation Protein Synthesis
 Gene Expression Transcription/Translation Protein Synthesis 1. Describe how genetic information is transcribed into sequences of bases in RNA molecules and is finally translated into sequences of amino
Gene Expression Transcription/Translation Protein Synthesis 1. Describe how genetic information is transcribed into sequences of bases in RNA molecules and is finally translated into sequences of amino
Total Test Questions: 66 Levels: Grades Units of Credit: 1.0 STANDARD 2. Demonstrate appropriate use of personal protective devices.
 DESCRIPTION Biotechnology is designed to create an awareness of career possibilities in the field of biotechnology. Students are introduced to diagnostic and therapeutic laboratory procedures that support
DESCRIPTION Biotechnology is designed to create an awareness of career possibilities in the field of biotechnology. Students are introduced to diagnostic and therapeutic laboratory procedures that support
The common structure of a DNA nucleotide. Hewitt
 GENETICS Unless otherwise noted* the artwork and photographs in this slide show are original and by Burt Carter. Permission is granted to use them for non-commercial, non-profit educational purposes provided
GENETICS Unless otherwise noted* the artwork and photographs in this slide show are original and by Burt Carter. Permission is granted to use them for non-commercial, non-profit educational purposes provided
The Polymerase Chain Reaction. Chapter 6: Background
 The Polymerase Chain Reaction Chapter 6: Background Invention of PCR Kary Mullis Mile marker 46.58 in April of 1983 Pulled off the road and outlined a way to conduct DNA replication in a tube Worked for
The Polymerase Chain Reaction Chapter 6: Background Invention of PCR Kary Mullis Mile marker 46.58 in April of 1983 Pulled off the road and outlined a way to conduct DNA replication in a tube Worked for
DNA and RNA. Chapter 12
 DNA and RNA Chapter 12 History of DNA Late 1800 s scientists discovered that DNA is in the nucleus of the cell 1902 Walter Sutton proposed that hereditary material resided in the chromosomes in the nucleus
DNA and RNA Chapter 12 History of DNA Late 1800 s scientists discovered that DNA is in the nucleus of the cell 1902 Walter Sutton proposed that hereditary material resided in the chromosomes in the nucleus
Methods of Biomaterials Testing Lesson 3-5. Biochemical Methods - Molecular Biology -
 Methods of Biomaterials Testing Lesson 3-5 Biochemical Methods - Molecular Biology - Chromosomes in the Cell Nucleus DNA in the Chromosome Deoxyribonucleic Acid (DNA) DNA has double-helix structure The
Methods of Biomaterials Testing Lesson 3-5 Biochemical Methods - Molecular Biology - Chromosomes in the Cell Nucleus DNA in the Chromosome Deoxyribonucleic Acid (DNA) DNA has double-helix structure The
Nucleic Acids: DNA and RNA
 Nucleic Acids: DNA and RNA Living organisms are complex systems. Hundreds of thousands of proteins exist inside each one of us to help carry out our daily functions. These proteins are produced locally,
Nucleic Acids: DNA and RNA Living organisms are complex systems. Hundreds of thousands of proteins exist inside each one of us to help carry out our daily functions. These proteins are produced locally,
Genetic Engineering & Recombinant DNA
 Genetic Engineering & Recombinant DNA Chapter 10 Copyright The McGraw-Hill Companies, Inc) Permission required for reproduction or display. Applications of Genetic Engineering Basic science vs. Applied
Genetic Engineering & Recombinant DNA Chapter 10 Copyright The McGraw-Hill Companies, Inc) Permission required for reproduction or display. Applications of Genetic Engineering Basic science vs. Applied
AP BIOLOGY RNA, DNA, & Proteins Chapters 16 & 17 Review
 AP BIOLOGY RNA, DNA, & Proteins Chapters 16 & 17 Review Enzyme that adds nucleotide subunits to an RNA primer during replication DNA polymerase III Another name for protein synthesis translation Sugar
AP BIOLOGY RNA, DNA, & Proteins Chapters 16 & 17 Review Enzyme that adds nucleotide subunits to an RNA primer during replication DNA polymerase III Another name for protein synthesis translation Sugar
Transcription in Prokaryotes. Jörg Bungert, PhD Phone:
 Transcription in Prokaryotes Jörg Bungert, PhD Phone: 352-273-8098 Email: jbungert@ufl.edu Objectives Understand the basic mechanism of transcription. Know the function of promoter elements and associating
Transcription in Prokaryotes Jörg Bungert, PhD Phone: 352-273-8098 Email: jbungert@ufl.edu Objectives Understand the basic mechanism of transcription. Know the function of promoter elements and associating
ELE4120 Bioinformatics. Tutorial 5
 ELE4120 Bioinformatics Tutorial 5 1 1. Database Content GenBank RefSeq TPA UniProt 2. Database Searches 2 Databases A common situation for alignment is to search through a database to retrieve the similar
ELE4120 Bioinformatics Tutorial 5 1 1. Database Content GenBank RefSeq TPA UniProt 2. Database Searches 2 Databases A common situation for alignment is to search through a database to retrieve the similar
Chapter 18: Regulation of Gene Expression. 1. Gene Regulation in Bacteria 2. Gene Regulation in Eukaryotes 3. Gene Regulation & Cancer
 Chapter 18: Regulation of Gene Expression 1. Gene Regulation in Bacteria 2. Gene Regulation in Eukaryotes 3. Gene Regulation & Cancer Gene Regulation Gene regulation refers to all aspects of controlling
Chapter 18: Regulation of Gene Expression 1. Gene Regulation in Bacteria 2. Gene Regulation in Eukaryotes 3. Gene Regulation & Cancer Gene Regulation Gene regulation refers to all aspects of controlling
Structural bioinformatics
 Structural bioinformatics Why structures? The representation of the molecules in 3D is more informative New properties of the molecules are revealed, which can not be detected by sequences Eran Eyal Plant
Structural bioinformatics Why structures? The representation of the molecules in 3D is more informative New properties of the molecules are revealed, which can not be detected by sequences Eran Eyal Plant
Zool 3200: Cell Biology Exam 3 3/6/15
 Name: Trask Zool 3200: Cell Biology Exam 3 3/6/15 Answer each of the following questions in the space provided; circle the correct answer or answers for each multiple choice question and circle either
Name: Trask Zool 3200: Cell Biology Exam 3 3/6/15 Answer each of the following questions in the space provided; circle the correct answer or answers for each multiple choice question and circle either
DNA replication. Begins at specific sites on a double helix. Proceeds in both directions. Is initiated at many points in eukaryotic chromosomes.
 DNA replication Begins at specific sites on a double helix. Proceeds in both directions. Is initiated at many points in eukaryotic chromosomes. Figure 10.8 http://www.hhmi.org/biointeractive/media/ DNAi_replication_schematic-lg.mov
DNA replication Begins at specific sites on a double helix. Proceeds in both directions. Is initiated at many points in eukaryotic chromosomes. Figure 10.8 http://www.hhmi.org/biointeractive/media/ DNAi_replication_schematic-lg.mov
Nucleic Acids, Proteins, and Enzymes
 3 Nucleic Acids, Proteins, and Enzymes Chapter 3 Nucleic Acids, Proteins, and Enzymes Key Concepts 3.1 Nucleic Acids Are Informational Macromolecules 3.2 Proteins Are Polymers with Important Structural
3 Nucleic Acids, Proteins, and Enzymes Chapter 3 Nucleic Acids, Proteins, and Enzymes Key Concepts 3.1 Nucleic Acids Are Informational Macromolecules 3.2 Proteins Are Polymers with Important Structural
Roche Molecular Biochemicals Technical Note No. LC 10/2000
 Roche Molecular Biochemicals Technical Note No. LC 10/2000 LightCycler Overview of LightCycler Quantification Methods 1. General Introduction Introduction Content Definitions This Technical Note will introduce
Roche Molecular Biochemicals Technical Note No. LC 10/2000 LightCycler Overview of LightCycler Quantification Methods 1. General Introduction Introduction Content Definitions This Technical Note will introduce
The GeneEditor TM in vitro Mutagenesis System: Site- Directed Mutagenesis Using Altered Beta-Lactamase Specificity
 Promega Notes Magazine Number 62, 1997, p. 02 The GeneEditor TM in vitro Mutagenesis System: Site- Directed Mutagenesis Using Altered Beta-Lactamase Specificity By Christine Andrews and Scott Lesley Promega
Promega Notes Magazine Number 62, 1997, p. 02 The GeneEditor TM in vitro Mutagenesis System: Site- Directed Mutagenesis Using Altered Beta-Lactamase Specificity By Christine Andrews and Scott Lesley Promega
Introduction to BioMEMS & Medical Microdevices DNA Microarrays and Lab-on-a-Chip Methods
 Introduction to BioMEMS & Medical Microdevices DNA Microarrays and Lab-on-a-Chip Methods Companion lecture to the textbook: Fundamentals of BioMEMS and Medical Microdevices, by Prof., http://saliterman.umn.edu/
Introduction to BioMEMS & Medical Microdevices DNA Microarrays and Lab-on-a-Chip Methods Companion lecture to the textbook: Fundamentals of BioMEMS and Medical Microdevices, by Prof., http://saliterman.umn.edu/
CHAPTER 21 LECTURE SLIDES
 CHAPTER 21 LECTURE SLIDES Prepared by Brenda Leady University of Toledo To run the animations you must be in Slideshow View. Use the buttons on the animation to play, pause, and turn audio/text on or off.
CHAPTER 21 LECTURE SLIDES Prepared by Brenda Leady University of Toledo To run the animations you must be in Slideshow View. Use the buttons on the animation to play, pause, and turn audio/text on or off.
Bundle 5 Test Review
 Bundle 5 Test Review DNA vs. RNA DNA Replication Gene Mutations- Protein Synthesis 1. Label the different components and complete the complimentary base pairing. What is this molecule called? _Nucleic
Bundle 5 Test Review DNA vs. RNA DNA Replication Gene Mutations- Protein Synthesis 1. Label the different components and complete the complimentary base pairing. What is this molecule called? _Nucleic
Protein 3D Structure Prediction
 Protein 3D Structure Prediction Michael Tress CNIO ?? MREYKLVVLGSGGVGKSALTVQFVQGIFVDE YDPTIEDSYRKQVEVDCQQCMLEILDTAGTE QFTAMRDLYMKNGQGFALVYSITAQSTFNDL QDLREQILRVKDTEDVPMILVGNKCDLEDER VVGKEQGQNLARQWCNCAFLESSAKSKINVN
Protein 3D Structure Prediction Michael Tress CNIO ?? MREYKLVVLGSGGVGKSALTVQFVQGIFVDE YDPTIEDSYRKQVEVDCQQCMLEILDTAGTE QFTAMRDLYMKNGQGFALVYSITAQSTFNDL QDLREQILRVKDTEDVPMILVGNKCDLEDER VVGKEQGQNLARQWCNCAFLESSAKSKINVN
STUDY GUIDE SECTION 10-1 Discovery of DNA
 STUDY GUIDE SECTION 10-1 Discovery of DNA Name Period Date Multiple Choice-Write the correct letter in the blank. 1. The virulent strain of the bacterium S. pneumoniae causes disease because it a. has
STUDY GUIDE SECTION 10-1 Discovery of DNA Name Period Date Multiple Choice-Write the correct letter in the blank. 1. The virulent strain of the bacterium S. pneumoniae causes disease because it a. has
Regulation of gene expression. (Lehninger pg )
 Regulation of gene expression (Lehninger pg. 1072-1085) Today s lecture Gene expression Constitutive, inducible, repressible genes Specificity factors, activators, repressors Negative and positive gene
Regulation of gene expression (Lehninger pg. 1072-1085) Today s lecture Gene expression Constitutive, inducible, repressible genes Specificity factors, activators, repressors Negative and positive gene
Pre-Lab: Molecular Biology
 Pre-Lab: Molecular Biology Name 1. What are the three chemical parts of a nucleotide. Draw a simple sketch to show how the three parts are arranged. 2. What are the rules of base pairing? 3. In double
Pre-Lab: Molecular Biology Name 1. What are the three chemical parts of a nucleotide. Draw a simple sketch to show how the three parts are arranged. 2. What are the rules of base pairing? 3. In double
DNA REPLICATION REVIEW
 Biology Ms. Ye DNA REPLICATION REVIEW 1. Number the steps of DNA replication the correct order (1, 2, 3): Name Date Block Daughter strands are formed using complementary base pairing DNA unwinds The DNA
Biology Ms. Ye DNA REPLICATION REVIEW 1. Number the steps of DNA replication the correct order (1, 2, 3): Name Date Block Daughter strands are formed using complementary base pairing DNA unwinds The DNA
