Development of a cost-effective pyrosequencing approach for SNP genotyping in
|
|
|
- Cameron Sparks
- 6 years ago
- Views:
Transcription
1 Development of a cost-effective pyrosequencing approach for SNP genotyping in barley C. Silvar 1,2, D. Perovic 2,*, A. M. Casas 1, E. Igartua 1 and F. Ordon 2 1 Department of Genetics and Plant Production, Aula Dei Experimental Station, CSIC, E Zaragoza, Spain. 2 Julius Kühn-Institute (JKI), Federal Research Institute for Cultivated Plants, Institute for Resistance Research and Stress Tolerance, Erwin-Baur-Str. 27, Quedlinburg, Germany * Address for correspondence: Dragan Perovic, Julius Kuehn-Institute (JKI), Federal Research Institute for Cultivated Plants, Institute for Resistance Research and Stress Tolerance, Erwin-Baur-Str. 27, Quedlinburg, Germany dragan.perovic@jki.bund.de Telephone: +49(0) Fax: +49(0)
2 Abstract An improved, efficient, reliable and cost-effective pyrosequencing protocol for SNP genotyping in plants is described. Labelling of the PCR products, required for the single-stranded pyrosequencing assay is carried out in a one-step PCR reaction with a universal biotinalyted M13 primer of 19-bp. A ratio of 1:10 of tailed-primer to M13 primer in a 15 µl reaction volume turned out to be best suited for a successful identification of different alleles based on known SNPs. This technology allows cost effective SNP genotyping in plant genomes for loci of interest at an overall low cost in a short period of time and is therefore best suited for marker based selection procedures. Key words: SNP, pyrosequencing, M13 tail, barley, breeding, MAS, cost-effective 2
3 Introduction Molecular markers have been widely used in plant breeding for phylogenetic studies, comparative genomics, mapping of genes or quantitative trait locus (QTL) and marker assisted selection (MAS) (Varshney et al. 2005, Ganal et al. 2009). The most abundant molecular markers in plant and animal genomes are Single Nucleotide Polymorphism (SNP), which constitute single base-pair variations in the DNA sequence. They are becoming the marker of choice in plant breeding, as they are easily prone to automation and high-throughput. Consequently, genomic resources based on SNPs for crop plant species are growing fast (e.g. Close et al. 2009). Nevertheless, for breeding purposes it is often the case that one specific SNP has to be analysed on thousands of genotypes, because a certain allele is indicative for a desired trait (e.g. Tyrka et al. 2004). At this scale, the most popular method for this purpose is still SNP genotyping using Cleaved Amplified Polymorphic Sequences (CAPS) (Konieczny and Ausubel 1993), which is based on the digestion with a restriction endonuclease. This technology, however, has some drawbacks, as the possibility of false scorings due to partial restriction enzyme digestion and the limitation to SNPs for which a restriction enzyme is available. Therefore, the development of an accurate and low-cost methodology to assay SNP-based markers in plant breeding is still necessary. For this purpose we propose an improvement on the SNP detection procedure through pyrosequencing (Ronaghi et al. 1998). Pyrosequencing is a real-time sequencing technique based on the detection of released pyrophosphate during DNA synthesis. This method involves the conversion of the PCR product into single-strand DNA and isolation of one of the strands, usually through labelling with biotin. Pyrosequencing is accurate, flexible, and can be easily automated (Langaee and Ronaghi 2005). Several applications of this technique for SNP genotyping in plant genetic studies have been proposed (Mochida et al. 2003, Varshney et al. 2008, Hanemann et al. 2009, Wang et al. 2010). However, one major disadvantage of this technology, which notably increases the expenses, is the need for primers typically labelled with biotin. 3
4 The aim of this work is to develop a universal scheme for the conversion of SNP markers into easyto-handle and cost-effective pyrosequencing markers suitable for application in plant breeding. Material and Methods Plant Material. Seedlings from the barley landraces SBCC097 (Igartua et al. 1998) and MBR1012 and the cultivars Plaisant and Scarlett were harvested at the two leaf stage and frozen in liquid nitrogen. Total genomic DNA was extracted using either the NucleoSpin Plant XL kit (Macherey Nagel) or the CTAB method according to Stein et al. (2001). Primer design and PCR optimization. A set of SNP markers, previously analyzed in an Illumina Golden Gate assay (BOPA1, Close et al. 2009), were employed to set up the new protocol (Table 1). Each pyrosequencing marker comprises a set of three primers: forward, reverse and sequencing primers. All primers were designed using the Pyrosequencing Assay Design Software (Biotage). The primer design parameters included a primer length up to 21 bp and an optimal product size between 150 and 200 bp. An M13 tail (5 -CACGACGTTGTAAAACGAC-3 ) of 19 bp was added to the 5 end of the forward primer. The sequencing primer was always designed on the complementary strand to the M13-tagged primer. The biotin label was incorporated to a M13 universal primer with a complementary sequence to the M13 tail. First, we optimized the PCR conditions without the M13 primer. PCR reactions were performed in a total volume of 15 µl, containing 25 ng of genomic DNA, 1 PCR reaction buffer (Solis Biodyne, Tartu, Estonia), 2.5 mm Mg 2+, 0.2 mm of dntps (Fermentas, St. Leon-Rot, Germany), 0.2 mm of each primer (Biolegio, AB Nijmegen, Netherland) and 0.5 U Taq FIREPol polymerase (Solis Biodyne). Taq DNA polymerase (Qiagen), FIREPol polymerase (Solis Biodyne) and AmpliTaq DNA Polymerase (Applied Biosystems), respectively, with their corresponding reaction buffers were used for the parallel assessment of amplification quality and quantity. All fragments were amplified using a touchdown PCR profile in which the annealing temperature was decreased in 4
5 0.5 C increments from 62ºC to 56 C, followed by cycles at 56 C. When using a Hot-Start Taq FIREPol polymerase (Solis Biodyne), the PCR programme was modified by increasing the first step at 94 C up to 15 min and the number of cycles up to 45. Amplicons were analysed by electrophoresis on a 1.5% agarose gel in 1 TBE buffer and visualized using ethidium bromide (5µg/ml) staining. Once the presence of non-specific products was corroborated, the biotinylated M13 universal primer was incorporated to the reaction using different ratios (tailed forward primer: M13 primer): 1:2, 1:5 and 1:10. The amount of reverse primer was kept constant. A control reaction without the third primer was carried out at the same time. The presence of only one amplicon and the strength of amplification were checked by electrophoresis on a 1.5 % agorose gel. In instances where it was desirable to improve the PCR specificity and yield, a Hot-Start Polymerase (Solis Biodyne) was employed. Pyrosequencing assay. The pyrosequencing assay for SNP genotyping was carried out on a Pyromark ID system (Biotage). The experimental procedure for pyrosequencing assays, including annealing plate preparation, immobilization of PCR products to streptavidin beads and the preparation of single stranded pyrosequencing template DNA were done as described on manufacturer s instructions. Results and discussion Two markers (11_0761 and 11_0068) from the barley Illumina BOPA1 (Close et al. 2009), were employed for the optimization of the PCR and pyrosequencing approach. DNA from SBCC097 and Plaisant were successfully amplified with specific forward and reverse primers for each locus (data not shown). The first step of PCR represents the critical step for the further pyrosequencing assay since the reliability and reproducibility of the technique largely depends on the quality and quantity of the PCR product. The optimal concentration for M13 primer and the stoichiometry of 5
6 three primers was assayed to avoid the amplification of unspecific products and optimize the yield of labelled PCR product. A touchdown programme, as recommended in Schuelke (2000), turned out to be very suitable for obtaining good-quality and strong fragments. Amplification with three primers for these two markers gave a single and specific product of expected size. No differences were observed between different ratios in an agarose gel by visual inspection (Fig. 1a). The standard reaction volume used for PCR was 15 µl. If a weak PCR product was obtained, the volume was increased up to 25 µl. We did not notice differences in amplification with cheap (FIREPol) and more expensive polymerases (AmpliTaq and Qiagen Taq) (data not shown); therefore all further steps were performed by using the cheaper FIREPol polymerase. To assess which concentration of universal primer produces the highest amount of biotinylated amplicons, the above PCR products were used as templates for the pyrosequencing assay. All different ratios tested produced a pyrogram, in which each allele is clearly identified. The different primer stoichiometries did not produce a visible effect on the resulting pyrograms (Fig. 1b). The new approach described in this paper is similar to the M13-tailed primer method reported earlier by Oetting et al. (1995). That technique has been mostly used in plant and animal genetics for microsatellite genotyping using fluorescent dye labels and capillary electrophoresis or laser detection systems (Anca et al. 2007, Santra et al. 2008). However, to our knowledge, no report exists on the application of an M13 tail for the genotyping of SNP markers. Only Royo et al. (2007) described a method for mutation detection in humans based on an M13 tail and a pyrosequencing scheme. In plants, only a short report by Pacey-Miller and Henry (2003) showed preliminary data on using a universal biotinylated primer in a pyrosequencing approach. The main drawback of the latter method is that two separate PCRs were performed to incorporate the biotinylated tag sequence to the template. In the present work, we overcome such inconvenience by labelling the PCR product with biotin in a single-tube step. Compared to the procedure reported by Royo et al. (2007), one major advantage of our method lies in the length of the M13 tail. Nowadays, many companies offer 6
7 the cheapest oligo synthesis tier up to 40 bp. The 24 bp long universal primer proposed by Royo et al (2007) leaves only 16 bp for forward primer design in the cheap tier. In most cases, longer primers will be needed, thus entailing significant cost increases of oligo synthesis. Our 19-bp sequence of M13 universal primer allows the design of forward primers up to 21 bp without increasing the cost of the primer synthesis. Additionally, smaller PCR volumes can be used in our approach compared to Royo et al. (2007) and thereby the cost of chemicals per reaction will be notably reduced. To evaluate the reproducibility and robustness of the method, we selected the ratio 1:10 (0.2 µm M13 primer: 0.02 µm forward primer) for further work on another four BOPA1 SNP markers and two different lines ( MBR1012 and Scarlett ). No other PCR components or conditions for these markers were optimized. PCR products of expected size and expected pyrograms were obtained for all 4 different markers and a polymorphisms was detected in three of them (Fig. S1). Therefore, we can conclude that all the time-consuming preliminary steps for primer optimization do not need to be undertaken for each new primer set or DNA material. This feature constitutes a considerable improvement compared to previously available procedures and a notable profit for rapid and successful implementation of this technology into the laboratory, since individual optimization of each SNP assay is not necessary. Additionally, the method proposed successfully identified SNPs that differed in the composition of adjacent sequence (which frequently entails the development of specific procedures to assay each SNP). Therefore, this method facilitates to assay a panel of SNPs with a single procedure, avoiding the need to deploy a suite of different genotyping methods for each specific SNP. The technology presented here provides an opportunity for laboratories possessing a pyrosequencer to optimize its use by reducing notably the expenses. This method will be best suited for all cases in which a limited number of SNPs has to be tested on a large amount of genotypes, e.g. in high resolution mapping procedures and in marker based selection schemes. Additionally, the flexibility 7
8 provided by our method to explore a large number of genotypes, together with the sequencing by synthesis technology inherent in the pyrosequencer, opens possibilities for SNP discovery in a much cheaper way. It would be possible to identify SNPs which are known (as it has been demonstrated in MBR1012 and Scarlet) but also to discover new SNPs in the neighbourhood of known SNPs while analyzing new genotypes in germplasm collections (see SNP 11_0537 in Fig. S1). Thus, new alleles for known genes may be discovered. In conclusion, the major advantages of the approach presented here are a reduction of the overall cost of pyrosequencing (cheap Taq polymerase, only one general biotinylated primer, small PCR volumes and only one single-tube reaction for labelling of PCR products) and of the requirements for individual assay optimization, while maintaining the specificity, flexibility and accuracy of a typical pyrosequencing approach. Acknowledgments This study was partially carried out within project AGL , funded by the Spanish Ministry for Science and Innovation. C.S. holds an I3P contract from CSIC. C.S. was supported by mobility fellowships from the Deutsche Forschungsgemeinschaft (DFG), CSIC, Fundacion Caja Inmaculada and COST Action FA0604 (Tritigen). References Anca, M. V., S. B. Andersen, D. Banga, M. Ardelean, and A. M. Torp, 2007: Genetic mapping of wheat 2bl chromosome using SSR markers. Buletin USAMV-CN 64, Close, T. J., R. B. Prasanna, L. S. Lonardi, Y. Wu, N. Rostoks, L. Ramsay, A. Druka, N. Stein, J. T. Svensson, S. Wanamaker, S. Bozdag, M. L. Roose, M. J. Moscou, S. Chao, R. K. Varshney, P. Szűcs, K. Sato, P. M. Hayes, D. E. Matthews, A. Kleinhofs, G. J. Muehlbauer, J. DeYoung, D. F. Marshall, K. Madishetty, R. D. Fenton, P. Condamine, A. Graner, and R. Waugh, 2009: Development and implementation of high-throughput SNP genotyping in barley. BMC Genomics 10,
9 Ganal, M. W., T. Altmann, and M. S. Röder, 2009: SNP identification in crop plants. Curr. Opin. Plant Biol. 12, Hanemann, A., G. F. Schweizer, R. Cossu, T. Wicker, and M. S. Röder, 2009: Fine mapping, physical mapping and development of diagnostic markers for the Rrs2 scald resistance gene in barley. Theor. Appl. Genet. 119, Igartua E, M. P. Gracia, J. M. Lasa, B. Medina, J. L. Molina-Cano, J. L. Montoya, and I. Romagosa, 1998: The Spanish barley core collection. Genet. Resour. Crop Ev. 45, Konieczny, A., and F. M. Ausubel, 1993: A procedure for mapping Arabidopsis mutations using co-dominant ecotype-specific PCR-based markers. Plant J. 4, Langaee, T., and R. M. Ronaghi, 2005: Genetic variation analyses by Pyrosequencing. Mutation Res. 573, Mochida, K., Y. Yamazaki, and Y. Ogihara, 2003: Discrimination of homoeologous gene expression in hexaploid wheat by SNP analysis of contigs grouped from a large number of expressed sequence tags. Mol. Gen. Genomics 270, Oetting, W.S., H. K. Lee, D. J. Flanders, G. L. Wiesner, T. A. Sellers, and R. A. King, 1995: Linkage analysis with multiplexed short tandem repeat polymorphisms using infrared fluorescen and M13 tailed primers. Genomics 30, Pacey-Miller, T., and R. Henry, 2002: Single-nucleotide polymorphism detection in plants using a single-stranded pyrosequencing protocol with a universal biotinylated primer. Anal. Biochem. 317, Ronaghi, R. M., M. Uhlen, and P. Nyren, 1998: A sequencing method based on real-time pyrophosphate. Science 281, Royo, J. L., M. Hidalgo, and A. Ruiz, 2007: Pyrosequencing protocol using a universal biotinylated primer for mutation detection and SNP genotyping. Nat. Protoc. 2, Santra, D. K., X. M. Chen, M. Santra, K. G. Campbell, and K. K. Kidwell, 2008: IdentiWcation and mapping QTL for high-temperature adult-plant resistance to stripe rust in winter wheat (Triticum aestivum L.) cultivar Stephens. Theor. Appl. Genet. 117,
10 Schuelke, M., 2000: An economic method for the fluorescent labelling of PCR fragments. Nature Biotech. 18, Stein, N., G. Herren, and B. Keller, 2001: A new DNA extraction method for high throughput marker in a large genome species such as Triticum aestivum. Plant Breed. 120, Tyrka, M., L. Blaszcyk, J. Chelkowski, V. Lind, L. Kramer, M. Weilepp, H. Wisniewska, and F. Ordon, 2004: Development of the single nucleotide polymorphism marker of the wheat Lr1 leaf rust resistance gene. Cell Mol. Biol. Lett. 9, Varshney, R. K., A. Graner, and M.E. Sorrells, 2005: Genic microsatellite markers in plants: features and applications. Trends Biotech. 23, Varshney, R. K., T. Thiel, T. Sretenovic-Rajicic, M. Baum, J. Valkoun, P. Guo, S, Grando, S. Ceccarelli, and A. Graner, 2008: Identification and validation of a core set of informative genic SSR and SNP markers for assaying functional diversity in barley. Mol. Breed. 22, Wang, G., I. Schmalenbach, M. von Korff, J. Leon, B. Kilian, J. Rode, and K. Pillen, 2010: Association of barley photoperiod and vernalization genes with QTLs for flowering time and agronomic traits in a BC2DH population and a set of wild barley introgression lines. Theor. Appl. Genet. 120,
11 Table 1. Marker name and primer sequences for BOPA1 SNPs employed for optimization of the pyrosequencing assay. Marker 1 Unigene 2 SNP Forward primer 3 PCR product (5 3 ) Reverse primer (5 3 ) Sequencing primer (5 3 ) Sequence to analyze (bp) 11_ C/G GCTGTCAGAGTCATCAACAC AATAATGGTGCACGACAAGT CAAGCTCCATCAATCG GC/GCATGGTACCA _ C/G CATTTTCTGGGCATTACG AAGGACTACGACGAGAATG CTGGGAGGATCTGGT CTC/GAAGAAGAGTG _ A/G AATTCTTGGCGATTACTC GCTACCATGTGATAGAGCT TCCAGTGCCCATCAT C/TTATTCCAGTA _ A/C TCCCTTGTTCATAGAGTCG GCGGTTTTGATAATTTAGAAG CTCAGAGAGATTCCCG WG/TCGGAAACAAT _ A/G CCAGCAGACAAATGACAC AAGGGTGGATACACCATCT CCATCTCCGACAACTC GAGC/TGGAAACAAG _ A/G GCCATCCATCTTGTTGTC TTACAGCACGAACACCAT ACCCTAGCTTGGTAGTTC C/TGCCATCAGAA Illumina OPA markers are designated according to BOPA1 name (Close et al. 2009) 2 Number of Unigene in HarvEST Assembly #35 ( 3 The biotynilated M13 (CACGACGTTGTAAAACGAC) was added at the 5 end of each forward primer 11
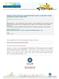
12 M 11_0761 P 11_0068 S P P Figure 1. (A) S PCR S amplification of two BOPA1 markers using three primers (M13, tailed-forward and reverse primers). S, SBCC097; P, Plaisant; M, 50 bp marker. Control indicates a PCR reaction A without M13 primer. (B) Pyrograms obtained with two different SNP BOPA1 markers; 11_0068 (right) and 11_0761 (left) on lines SBCC097 and Plaisant. The pyrograms observed by pyrosequencing analysis represent each possible genotype. I, II and III indicate the ratio of tailed forward primer to M13 primer; 1:2, 1:5 and 1:10, respectively. Letters on the X-axis indicate B the dispensation order established for the machine according to the sequence to analyze. E and S stand for enzyme and substrate reagents, respectively. 12
Marker types. Potato Association of America Frederiction August 9, Allen Van Deynze
 Marker types Potato Association of America Frederiction August 9, 2009 Allen Van Deynze Use of DNA Markers in Breeding Germplasm Analysis Fingerprinting of germplasm Arrangement of diversity (clustering,
Marker types Potato Association of America Frederiction August 9, 2009 Allen Van Deynze Use of DNA Markers in Breeding Germplasm Analysis Fingerprinting of germplasm Arrangement of diversity (clustering,
Lecture 8: Sequencing and SNP. Sept 15, 2006
 Lecture 8: Sequencing and SNP Sept 15, 2006 Announcements Random questioning during literature discussion sessions starts next week for real! Schedule changes Moved QTL lecture up Removed landscape genetics
Lecture 8: Sequencing and SNP Sept 15, 2006 Announcements Random questioning during literature discussion sessions starts next week for real! Schedule changes Moved QTL lecture up Removed landscape genetics
SolCAP. Executive Commitee : David Douches Walter De Jong Robin Buell David Francis Alexandra Stone Lukas Mueller AllenVan Deynze
 SolCAP Solanaceae Coordinated Agricultural Project Supported by the National Research Initiative Plant Genome Program of USDA CSREES for the Improvement of Potato and Tomato Executive Commitee : David
SolCAP Solanaceae Coordinated Agricultural Project Supported by the National Research Initiative Plant Genome Program of USDA CSREES for the Improvement of Potato and Tomato Executive Commitee : David
Association genetics in barley
 Association genetics in barley Ildikó Karsai, Hungarian Academy of Sciences, Martonvásár, Hungary Frank Ordon, Dragan Perovic, Julius Kühn Institute, Quedlinburg, Germany Günther Schweizer, Bianca Büttner,
Association genetics in barley Ildikó Karsai, Hungarian Academy of Sciences, Martonvásár, Hungary Frank Ordon, Dragan Perovic, Julius Kühn Institute, Quedlinburg, Germany Günther Schweizer, Bianca Büttner,
2. Pyrosequencing Assay Design
 2. Pyrosequencing Assay Design 2.1 Guidelines for PCR set-up and primer design 2.1.1 PCR primer design Design of PCR primers follows standard rules, i.e. calculated Tm of 62-65 C, primer length of about
2. Pyrosequencing Assay Design 2.1 Guidelines for PCR set-up and primer design 2.1.1 PCR primer design Design of PCR primers follows standard rules, i.e. calculated Tm of 62-65 C, primer length of about
Authors: Vivek Sharma and Ram Kunwar
 Molecular markers types and applications A genetic marker is a gene or known DNA sequence on a chromosome that can be used to identify individuals or species. Why we need Molecular Markers There will be
Molecular markers types and applications A genetic marker is a gene or known DNA sequence on a chromosome that can be used to identify individuals or species. Why we need Molecular Markers There will be
INTERNATIONAL UNION FOR THE PROTECTION OF NEW VARIETIES OF PLANTS GENEVA
 E BMT Guidelines (proj.4) ORIGINAL: English DATE: December 21, 2005 INTERNATIONAL UNION FOR THE PROTECTION OF NEW VARIETIES OF PLANTS GENEVA GUIDELINES FOR DNA-PROFILING: MOLECULAR MARKER SELECTION AND
E BMT Guidelines (proj.4) ORIGINAL: English DATE: December 21, 2005 INTERNATIONAL UNION FOR THE PROTECTION OF NEW VARIETIES OF PLANTS GENEVA GUIDELINES FOR DNA-PROFILING: MOLECULAR MARKER SELECTION AND
PlantDirect TM Multiplex PCR System
 PlantDirect TM Multiplex PCR System Technical Manual No. 0178 Version 10112010 I Description.. 1 II Applications 2 III Key Features.. 3 IV Shipping and Storage. 3 V Simplified Procedures. 3 VI Detailed
PlantDirect TM Multiplex PCR System Technical Manual No. 0178 Version 10112010 I Description.. 1 II Applications 2 III Key Features.. 3 IV Shipping and Storage. 3 V Simplified Procedures. 3 VI Detailed
INTERNATIONAL UNION FOR THE PROTECTION OF NEW VARIETIES OF PLANTS
 ORIGINAL: English DATE: October 21, 2010 INTERNATIONAL UNION FOR THE PROTECTION OF NEW VARIETIES OF PLANTS GENEVA E GUIDELINES FOR DNA-PROFILING: MOLECULAR MARKER SELECTION AND DATABASE CONSTRUCTION (
ORIGINAL: English DATE: October 21, 2010 INTERNATIONAL UNION FOR THE PROTECTION OF NEW VARIETIES OF PLANTS GENEVA E GUIDELINES FOR DNA-PROFILING: MOLECULAR MARKER SELECTION AND DATABASE CONSTRUCTION (
Positive control kit for enzymatic mismatch cleavage and agarose gel visualization (version 2.4)
 28 January, 2010 Positive control kit for enzymatic mismatch cleavage and agarose gel visualization (version 2.4) Plant Breeding Unit Brad Till and Owen Huynh January, 2010 Kit Contents: Genomic DNA from
28 January, 2010 Positive control kit for enzymatic mismatch cleavage and agarose gel visualization (version 2.4) Plant Breeding Unit Brad Till and Owen Huynh January, 2010 Kit Contents: Genomic DNA from
Genomic Sequencing. Genomic Sequencing. Maj Gen (R) Suhaib Ahmed, HI (M)
 Maj Gen (R) Suhaib Ahmed, HI (M) The process of determining the sequence of an unknown DNA is called sequencing. There are many approaches for DNA sequencing. In the last couple of decades automated Sanger
Maj Gen (R) Suhaib Ahmed, HI (M) The process of determining the sequence of an unknown DNA is called sequencing. There are many approaches for DNA sequencing. In the last couple of decades automated Sanger
Introduction to some aspects of molecular genetics
 Introduction to some aspects of molecular genetics Julius van der Werf (partly based on notes from Margaret Katz) University of New England, Armidale, Australia Genetic and Physical maps of the genome...
Introduction to some aspects of molecular genetics Julius van der Werf (partly based on notes from Margaret Katz) University of New England, Armidale, Australia Genetic and Physical maps of the genome...
Existing potato markers and marker conversions. Walter De Jong PAA Workshop August 2009
 Existing potato markers and marker conversions Walter De Jong PAA Workshop August 2009 1 What makes for a good marker? diagnostic for trait of interest robust works even with DNA of poor quality or low
Existing potato markers and marker conversions Walter De Jong PAA Workshop August 2009 1 What makes for a good marker? diagnostic for trait of interest robust works even with DNA of poor quality or low
STUDY OF VNTR HUMAN POLYMORPHISMS BY PCR
 STUDY OF VNTR HUMAN POLYMORPHISMS BY PCR Ref. PCR1 1. OBJECTIVE OF THE EXPERIMENT The objective of this experiment is to introduce students to the principles and practice of Polymerase Chain Reaction (PCR)
STUDY OF VNTR HUMAN POLYMORPHISMS BY PCR Ref. PCR1 1. OBJECTIVE OF THE EXPERIMENT The objective of this experiment is to introduce students to the principles and practice of Polymerase Chain Reaction (PCR)
DETERMINATION OF THE Rh FACTOR BY PCR
 DETERMINATION OF THE Rh FACTOR BY PCR Ref.: PCR2 1. EXPERIMENT OBJECTIVE The aim of this experiment is to introduce students to the principles and practice of the Polymerase Chain Reaction (PCR) by studying
DETERMINATION OF THE Rh FACTOR BY PCR Ref.: PCR2 1. EXPERIMENT OBJECTIVE The aim of this experiment is to introduce students to the principles and practice of the Polymerase Chain Reaction (PCR) by studying
Comparative study of EST-SSR, SSR, RAPD, and ISSR and their transferability analysis in pea, chickpea and mungbean
 EUROPEAN ACADEMIC RESEARCH Vol. IV, Issue 2/ May 2016 ISSN 2286-4822 www.euacademic.org Impact Factor: 3.4546 (UIF) DRJI Value: 5.9 (B+) Comparative study of EST-SSR, SSR, RAPD, and ISSR and their transferability
EUROPEAN ACADEMIC RESEARCH Vol. IV, Issue 2/ May 2016 ISSN 2286-4822 www.euacademic.org Impact Factor: 3.4546 (UIF) DRJI Value: 5.9 (B+) Comparative study of EST-SSR, SSR, RAPD, and ISSR and their transferability
Microsatellite markers
 Microsatellite markers Review of repetitive sequences 25% 45% 8% 21% 13% 3% Mobile genetic elements: = dispersed repeat included: transposition: moving in the form of DNA by element coding for transposases.
Microsatellite markers Review of repetitive sequences 25% 45% 8% 21% 13% 3% Mobile genetic elements: = dispersed repeat included: transposition: moving in the form of DNA by element coding for transposases.
Plant microsatellite genotyping with 4-color fluorescent detection using multiple-tailed primers
 A. Missiaggia and D. Grattapaglia 72 Plant microsatellite genotyping with 4-color fluorescent detection using multiple-tailed primers Alexandre Missiaggia 1,2 and Dario Grattapaglia 1,3 1 Laboratório de
A. Missiaggia and D. Grattapaglia 72 Plant microsatellite genotyping with 4-color fluorescent detection using multiple-tailed primers Alexandre Missiaggia 1,2 and Dario Grattapaglia 1,3 1 Laboratório de
Using molecular marker technology in studies on plant genetic diversity Final considerations
 Using molecular marker technology in studies on plant genetic diversity Final considerations Copyright: IPGRI and Cornell University, 2003 Final considerations 1 Contents! When choosing a technique...!
Using molecular marker technology in studies on plant genetic diversity Final considerations Copyright: IPGRI and Cornell University, 2003 Final considerations 1 Contents! When choosing a technique...!
GDMS Templates Documentation GDMS Templates Release 1.0
 GDMS Templates Documentation GDMS Templates Release 1.0 1 Table of Contents 1. SSR Genotyping Template 03 2. DArT Genotyping Template... 05 3. SNP Genotyping Template.. 08 4. QTL Template.. 09 5. Map Template..
GDMS Templates Documentation GDMS Templates Release 1.0 1 Table of Contents 1. SSR Genotyping Template 03 2. DArT Genotyping Template... 05 3. SNP Genotyping Template.. 08 4. QTL Template.. 09 5. Map Template..
Pyrosequencing for quantitative analysis of methylation at multiple CpG sites
 Application Note for Nature Methods Pyrosequencing for quantitative analysis of methylation at multiple CpG sites Robert England and Monica Pettersson Pyrosequencing from Biotage 1 improves virtually every
Application Note for Nature Methods Pyrosequencing for quantitative analysis of methylation at multiple CpG sites Robert England and Monica Pettersson Pyrosequencing from Biotage 1 improves virtually every
Single- and double-ssr primer combined analyses in rice
 Single- and double-ssr primer combined analyses in rice J. Ma, S.C. Guan, Z. Zhang and P.W. Wang Biotechnology Center of Jilin Agricultural University, Changchun, China Corresponding author: P.W. Wang
Single- and double-ssr primer combined analyses in rice J. Ma, S.C. Guan, Z. Zhang and P.W. Wang Biotechnology Center of Jilin Agricultural University, Changchun, China Corresponding author: P.W. Wang
BIOLOGY - CLUTCH CH.20 - BIOTECHNOLOGY.
 !! www.clutchprep.com CONCEPT: DNA CLONING DNA cloning is a technique that inserts a foreign gene into a living host to replicate the gene and produce gene products. Transformation the process by which
!! www.clutchprep.com CONCEPT: DNA CLONING DNA cloning is a technique that inserts a foreign gene into a living host to replicate the gene and produce gene products. Transformation the process by which
Comparative assessment of wheat landraces from AWCC, ICARDA and VIR germplasm collections based on the analysis of SSR markers
 epublications@scu Southern Cross Plant Science 2008 Comparative assessment of wheat landraces from AWCC, ICARDA and VIR germplasm collections based on the analysis of SSR markers P Strelchenko K Street
epublications@scu Southern Cross Plant Science 2008 Comparative assessment of wheat landraces from AWCC, ICARDA and VIR germplasm collections based on the analysis of SSR markers P Strelchenko K Street
Procomcure Biotech. PCR Reagents
 Procomcure Biotech PCR Reagents valid for 2018 VitaTaq DNA Polymerase is a standard Taq DNA polymerase suitable for all common PCR applications like colonypcr, cloning applications, high-throughput PCR
Procomcure Biotech PCR Reagents valid for 2018 VitaTaq DNA Polymerase is a standard Taq DNA polymerase suitable for all common PCR applications like colonypcr, cloning applications, high-throughput PCR
P HENIX. PHENIX PCR Enzyme Guide Tools For Life Science Discovery RESEARCH PRODUCTS
 PHENIX PCR Enzyme Guide PHENIX offers a broad line of premium quality PCR Enzymes. This PCR Enzyme Guide will help simplify your polymerase selection process. Each DNA Polymerase has different characteristics
PHENIX PCR Enzyme Guide PHENIX offers a broad line of premium quality PCR Enzymes. This PCR Enzyme Guide will help simplify your polymerase selection process. Each DNA Polymerase has different characteristics
Quantitative analysis of methylation at multiple CpG sites by Pyrosequencing TM
 Quantitative analysis of methylation at multiple CpG sites by Pyrosequencing TM Robert England and Monica Pettersson Pyrosequencing from Biotage 1 improves virtually every aspect of CpG methylation analysis:
Quantitative analysis of methylation at multiple CpG sites by Pyrosequencing TM Robert England and Monica Pettersson Pyrosequencing from Biotage 1 improves virtually every aspect of CpG methylation analysis:
WORKING GROUP ON BIOCHEMICAL AND MOLECULAR TECHNIQUES AND DNA PROFILING IN PARTICULAR. Eleventh Session Madrid, September 16 to 18, 2008
 ORIGINAL: English DATE: September 16, 2008 INTERNATIONAL UNION FOR THE PROTECTION OF NEW VARIETIES OF PLANTS GENEVA E WORKING GROUP ON BIOCHEMICAL AND MOLECULAR TECHNIQUES AND DNA PROFILING IN PARTICULAR
ORIGINAL: English DATE: September 16, 2008 INTERNATIONAL UNION FOR THE PROTECTION OF NEW VARIETIES OF PLANTS GENEVA E WORKING GROUP ON BIOCHEMICAL AND MOLECULAR TECHNIQUES AND DNA PROFILING IN PARTICULAR
Analysis of the expression of homoeologous genes in polyploid cereals using fluorescence cdna-sscp
 Analysis of the expression of homoeologous genes in polyploid cereals using fluorescence cdna-sscp De Bustos A., Perez R., Jouve N. in Molina-Cano J.L. (ed.), Christou P. (ed.), Graner A. (ed.), Hammer
Analysis of the expression of homoeologous genes in polyploid cereals using fluorescence cdna-sscp De Bustos A., Perez R., Jouve N. in Molina-Cano J.L. (ed.), Christou P. (ed.), Graner A. (ed.), Hammer
HCS806 Summer 2010 Methods in Plant Biology: Breeding with Molecular Markers.
 HCS806 Summer 2010 Methods in Plant Biology: Breeding with Molecular Markers. DNA, the stuff of life. We can think of DNA as a biochemical entity or as a data string. With the advent of high throughput
HCS806 Summer 2010 Methods in Plant Biology: Breeding with Molecular Markers. DNA, the stuff of life. We can think of DNA as a biochemical entity or as a data string. With the advent of high throughput
7. Troubleshooting Guidelines
 7. Troubleshooting Guidelines The Pyrosequencing TM technique provides the user with sequence information for several nucleotides 3 of the sequencing primer apart from the polymorphic site. This gives
7. Troubleshooting Guidelines The Pyrosequencing TM technique provides the user with sequence information for several nucleotides 3 of the sequencing primer apart from the polymorphic site. This gives
Applied Biosystems Real-Time PCR Rapid Assay Development Guidelines
 Applied Biosystems Real-Time PCR Rapid Assay Development Guidelines Description This tutorial will discuss recommended guidelines for designing and running real-time PCR quantification and SNP Genotyping
Applied Biosystems Real-Time PCR Rapid Assay Development Guidelines Description This tutorial will discuss recommended guidelines for designing and running real-time PCR quantification and SNP Genotyping
Lecture Four. Molecular Approaches I: Nucleic Acids
 Lecture Four. Molecular Approaches I: Nucleic Acids I. Recombinant DNA and Gene Cloning Recombinant DNA is DNA that has been created artificially. DNA from two or more sources is incorporated into a single
Lecture Four. Molecular Approaches I: Nucleic Acids I. Recombinant DNA and Gene Cloning Recombinant DNA is DNA that has been created artificially. DNA from two or more sources is incorporated into a single
7.1 Techniques for Producing and Analyzing DNA. SBI4U Ms. Ho-Lau
 7.1 Techniques for Producing and Analyzing DNA SBI4U Ms. Ho-Lau What is Biotechnology? From Merriam-Webster: the manipulation of living organisms or their components to produce useful usually commercial
7.1 Techniques for Producing and Analyzing DNA SBI4U Ms. Ho-Lau What is Biotechnology? From Merriam-Webster: the manipulation of living organisms or their components to produce useful usually commercial
*Author for correspondence (
 Received: 22 nd Oct-2012 Revised: 17 th Nov-2012 Accepted: 10 th Oct -2013 Research article EVALUATION OF RAPD AND SSR MOLECULAR MARKERS IN BREAD WHEAT GENOTYPES Khavarinejad M.S. 1 *, Karimov.M 2 and
Received: 22 nd Oct-2012 Revised: 17 th Nov-2012 Accepted: 10 th Oct -2013 Research article EVALUATION OF RAPD AND SSR MOLECULAR MARKERS IN BREAD WHEAT GENOTYPES Khavarinejad M.S. 1 *, Karimov.M 2 and
MICROSATELLITE MARKER AND ITS UTILITY
 Your full article ( between 500 to 5000 words) - - Do check for grammatical errors or spelling mistakes MICROSATELLITE MARKER AND ITS UTILITY 1 Prasenjit, D., 2 Anirudha, S. K. and 3 Mallar, N.K. 1,2 M.Sc.(Agri.),
Your full article ( between 500 to 5000 words) - - Do check for grammatical errors or spelling mistakes MICROSATELLITE MARKER AND ITS UTILITY 1 Prasenjit, D., 2 Anirudha, S. K. and 3 Mallar, N.K. 1,2 M.Sc.(Agri.),
GENOTYPING BY PCR PROTOCOL FORM MUTANT MOUSE REGIONAL RESOURCE CENTER North America, International
 Please provide the following information required for genetic analysis of your mutant mice. Please fill in form electronically by tabbing through the text fields. The first 2 pages are protected with gray
Please provide the following information required for genetic analysis of your mutant mice. Please fill in form electronically by tabbing through the text fields. The first 2 pages are protected with gray
AmpFlSTR Identifiler PCR Amplification Kit
 Product Bulletin Human Identification AmpFlSTR Identifiler PCR Amplification Kit Fifteen STR (short tandem repeat) loci and Amelogenin co-amplified in a single tube Incorporation of the same proven, reliable
Product Bulletin Human Identification AmpFlSTR Identifiler PCR Amplification Kit Fifteen STR (short tandem repeat) loci and Amelogenin co-amplified in a single tube Incorporation of the same proven, reliable
SUPPLEMENTARY MATERIAL AND METHODS
 SUPPLEMENTARY MATERIAL AND METHODS Amplification of HEV ORF1, ORF2 and ORF3 genome regions Total RNA was extracted from 200 µl EDTA plasma using Cobas AmpliPrep total nucleic acid isolation kit (Roche,
SUPPLEMENTARY MATERIAL AND METHODS Amplification of HEV ORF1, ORF2 and ORF3 genome regions Total RNA was extracted from 200 µl EDTA plasma using Cobas AmpliPrep total nucleic acid isolation kit (Roche,
Bio Rad PCR Song Lyrics
 Bio Rad PCR Song Lyrics There was a time when to amplify DNA, You had to grow tons and tons of tiny cells. (Oooh) Then along came a guy named Dr. Kary Mullis, Said you can amplify in vitro just as well.
Bio Rad PCR Song Lyrics There was a time when to amplify DNA, You had to grow tons and tons of tiny cells. (Oooh) Then along came a guy named Dr. Kary Mullis, Said you can amplify in vitro just as well.
A comparative performance evaluation of illustra Ready-To-Go GenomiPhi V3 and illustra GenomiPhi V2 DNA amplification kits
 GE Healthcare Life Sciences Application note 29-0245-46 AA Nucleic acid sample preparation A comparative performance evaluation of illustra Ready-To-Go GenomiPhi V3 and illustra GenomiPhi V2 DNA amplification
GE Healthcare Life Sciences Application note 29-0245-46 AA Nucleic acid sample preparation A comparative performance evaluation of illustra Ready-To-Go GenomiPhi V3 and illustra GenomiPhi V2 DNA amplification
GreenMasterMix (2X) b i o s c i e n c e. G E N A X X O N b i o s c i e n c e. High ROX (500nM)
 G:\products\productflyer\pcr\polymerasen\hotstart\manu_m3052_green_en.docx GreenMasterMix (2) High RO (500nM) qpcr master mix with fluorescence dye and passive reference dye Contact & Technical support
G:\products\productflyer\pcr\polymerasen\hotstart\manu_m3052_green_en.docx GreenMasterMix (2) High RO (500nM) qpcr master mix with fluorescence dye and passive reference dye Contact & Technical support
White Paper: High Throughput SNP Genotyping Using Array Tape in Place of Microplates
 White Paper High Throughput SNP Genotyping White Paper: High Throughput SNP Genotyping Using Array Tape in Place of Microplates Miniaturization and Automation Using a Novel New Reaction Substrate Originally
White Paper High Throughput SNP Genotyping White Paper: High Throughput SNP Genotyping Using Array Tape in Place of Microplates Miniaturization and Automation Using a Novel New Reaction Substrate Originally
Report of Analyzing Short Tandem Repeats for Parentage Testing
 1 Alex Michael Tseng Department of Forensic Medicine, College of Medicine, National Taiwan University Report of Analyzing Short Tandem Repeats for Parentage Testing Introduction In the three billion letter
1 Alex Michael Tseng Department of Forensic Medicine, College of Medicine, National Taiwan University Report of Analyzing Short Tandem Repeats for Parentage Testing Introduction In the three billion letter
Functional Genomics Research Stream. Research Meetings: November 2 & 3, 2009 Next Generation Sequencing
 Functional Genomics Research Stream Research Meetings: November 2 & 3, 2009 Next Generation Sequencing Current Issues Research Meetings: Meet with me this Thursday or Friday. (bring laboratory notebook
Functional Genomics Research Stream Research Meetings: November 2 & 3, 2009 Next Generation Sequencing Current Issues Research Meetings: Meet with me this Thursday or Friday. (bring laboratory notebook
Multiplex SNP Genotyping of Field Corn Crude Samples with Probe-Based Assays using the IntelliQube from Douglas Scientific INTRODUCTION
 PARTNERING WITH YOU TO MAKE THE WORLD A BETTER PLACE with Probe-Based Assays using the IntelliQube from ABSTRACT Single Nucleotide Polymorphism (SNP) genotyping must be accurate, reliable, cost-effective,
PARTNERING WITH YOU TO MAKE THE WORLD A BETTER PLACE with Probe-Based Assays using the IntelliQube from ABSTRACT Single Nucleotide Polymorphism (SNP) genotyping must be accurate, reliable, cost-effective,
PowerPlex. Y System Validation
 PowerPlex Y System Validation By Patricia M. Fulmer, Dawn Rabbach, Kimberly Huston, Curtis Knox and Cynthia Sprecher, Promega Corporation Abstract We have improved the manufacturing process for the PowerPlex
PowerPlex Y System Validation By Patricia M. Fulmer, Dawn Rabbach, Kimberly Huston, Curtis Knox and Cynthia Sprecher, Promega Corporation Abstract We have improved the manufacturing process for the PowerPlex
Methods in virus diagnosis PCR techniques
 Methods in virus diagnosis PCR techniques 450 MBIO PRACTICAL LESSON 5 Molecular Methods Methods based on the detection of viral genome are also commonly known as molecular methods. It is often said that
Methods in virus diagnosis PCR techniques 450 MBIO PRACTICAL LESSON 5 Molecular Methods Methods based on the detection of viral genome are also commonly known as molecular methods. It is often said that
Platinum II Taq Hot-Start DNA Polymerase for high-throughput PCR
 WHITE PAPER Platinum II Taq Hot-Start DNA Polymerase Platinum II Taq Hot-Start DNA Polymerase for high-throughput PCR Abstract The advances in thermal cycler technology permit a substantial increase in
WHITE PAPER Platinum II Taq Hot-Start DNA Polymerase Platinum II Taq Hot-Start DNA Polymerase for high-throughput PCR Abstract The advances in thermal cycler technology permit a substantial increase in
Mycobacterium paratuberculosis
 BACTOTYPE PCR Amplification Kit Mycobacterium paratuberculosis Labor Diagnostik Leipzig Manual Technology The product group BACTOTYPE PCR Amplification Kit comprises optimised systems for the identification
BACTOTYPE PCR Amplification Kit Mycobacterium paratuberculosis Labor Diagnostik Leipzig Manual Technology The product group BACTOTYPE PCR Amplification Kit comprises optimised systems for the identification
Development of transcript-associated microsatellite markers in Ancherythoculter nigrocauda and cross-amplification in Culter alburnus
 Development of transcript-associated microsatellite markers in Ancherythoculter nigrocauda and cross-amplification in Culter alburnus Y. Sun 1,2, Q. Li 1, G. Wang 1, D. Zhu 1, J. Chen 1 and P. Li 1 1 Wuhan
Development of transcript-associated microsatellite markers in Ancherythoculter nigrocauda and cross-amplification in Culter alburnus Y. Sun 1,2, Q. Li 1, G. Wang 1, D. Zhu 1, J. Chen 1 and P. Li 1 1 Wuhan
HiPer Random Amplification of Polymorphic DNA (RAPD) Teaching Kit
 HiPer Random Amplification of Polymorphic DNA (RAPD) Teaching Kit Product Code: HTBM031 Number of experiments that can be performed: 5 Duration of Experiment: Protocol: 3.5 hours Agarose Gel Electrophoresis:
HiPer Random Amplification of Polymorphic DNA (RAPD) Teaching Kit Product Code: HTBM031 Number of experiments that can be performed: 5 Duration of Experiment: Protocol: 3.5 hours Agarose Gel Electrophoresis:
BIOLOGY Dr.Locke Lecture# 27 An Introduction to Polymerase Chain Reaction (PCR)
 BIOLOGY 207 - Dr.Locke Lecture# 27 An Introduction to Polymerase Chain Reaction (PCR) Required readings and problems: Reading: Open Genetics, Chapter 8.1 Problems: Chapter 8 Optional Griffiths (2008) 9
BIOLOGY 207 - Dr.Locke Lecture# 27 An Introduction to Polymerase Chain Reaction (PCR) Required readings and problems: Reading: Open Genetics, Chapter 8.1 Problems: Chapter 8 Optional Griffiths (2008) 9
Pyrosequencing. Alix Groom
 Pyrosequencing Alix Groom Pyrosequencing high-throughput CpG methylation analysis platform real-time, sequence-based detection and quantification % methylation at multiple adjacent CpG sites 80-100 bases
Pyrosequencing Alix Groom Pyrosequencing high-throughput CpG methylation analysis platform real-time, sequence-based detection and quantification % methylation at multiple adjacent CpG sites 80-100 bases
POLYMERASE CHAIN REACTION PCR. (Biotechnology and Genetic Engineering) Edemhanria, Lawrence
 POLYMERASE CHAIN REACTION PCR. (Biotechnology and Genetic Engineering) Edemhanria, Lawrence Biochemistry Unit Chemical Sciences Department Samuel Adegboyega University Ogwa, Edo State, Nigeria. Outline
POLYMERASE CHAIN REACTION PCR. (Biotechnology and Genetic Engineering) Edemhanria, Lawrence Biochemistry Unit Chemical Sciences Department Samuel Adegboyega University Ogwa, Edo State, Nigeria. Outline
WORKING GROUP ON BIOCHEMICAL AND MOLECULAR TECHNIQUES AND DNA PROFILING IN PARTICULAR. Twelfth Session Ottawa, Canada, May 11 to 13, 2010
 E BMT/12/9 ORIGINAL: English DATE: April 9, 2010 INTERNATIONAL UNION FOR THE PROTECTION OF NEW VARIETIES OF PLANTS GENEVA WORKING GROUP ON BIOCHEMICAL AND MOLECULAR TECHNIQUES AND DNA PROFILING IN PARTICULAR
E BMT/12/9 ORIGINAL: English DATE: April 9, 2010 INTERNATIONAL UNION FOR THE PROTECTION OF NEW VARIETIES OF PLANTS GENEVA WORKING GROUP ON BIOCHEMICAL AND MOLECULAR TECHNIQUES AND DNA PROFILING IN PARTICULAR
Product Name : Simple mirna Detection Kit
 Product Name : Simple mirna Detection Kit Code No. : DS700 This product is for research use only Kit Contents This kit provides sufficient reagents to perform 20 reactions for detecting microrna. Components
Product Name : Simple mirna Detection Kit Code No. : DS700 This product is for research use only Kit Contents This kit provides sufficient reagents to perform 20 reactions for detecting microrna. Components
Hy-Fy High Fidelity Mix (x2)
 Hy-Fy High Fidelity Mix (x2) #EZ-2021 1ml, 100rxn of 20μl Contents: 2X High Fidelity Mix 1ml Nuclease-free water 1ml Store at -20 C Shelf life: 2 years Description Hy-Fy High Fidelity Mix (x2) is a premixed,
Hy-Fy High Fidelity Mix (x2) #EZ-2021 1ml, 100rxn of 20μl Contents: 2X High Fidelity Mix 1ml Nuclease-free water 1ml Store at -20 C Shelf life: 2 years Description Hy-Fy High Fidelity Mix (x2) is a premixed,
Characterisation of microsatellite DNA markers for Mirbelia bursarioides A.M.Monro & Crisp ms.
 1 2 3 Microsatellite Letter for Conservation Genetic Resources Word Count 870 Characterisation of microsatellite DNA markers for Mirbelia bursarioides A.M.Monro & Crisp ms. 4 5 Melissa A Millar 1, 2, Margaret
1 2 3 Microsatellite Letter for Conservation Genetic Resources Word Count 870 Characterisation of microsatellite DNA markers for Mirbelia bursarioides A.M.Monro & Crisp ms. 4 5 Melissa A Millar 1, 2, Margaret
DNA Sequencing by Ion Torrent. Marc Lavergne CHEM 4590
 DNA Sequencing by Ion Torrent Marc Lavergne CHEM 4590 OVERVIEW History DNA Synthesis and First-Gen Sequencing Technology Sequencing Signal Detection Advantages/Disadvantages Applications Current Research
DNA Sequencing by Ion Torrent Marc Lavergne CHEM 4590 OVERVIEW History DNA Synthesis and First-Gen Sequencing Technology Sequencing Signal Detection Advantages/Disadvantages Applications Current Research
HiPer RT-PCR Teaching Kit
 HiPer RT-PCR Teaching Kit Product Code: HTBM024 Number of experiments that can be performed: 5 Duration of Experiment: Protocol: 4 hours Agarose Gel Electrophoresis: 45 minutes Storage Instructions: The
HiPer RT-PCR Teaching Kit Product Code: HTBM024 Number of experiments that can be performed: 5 Duration of Experiment: Protocol: 4 hours Agarose Gel Electrophoresis: 45 minutes Storage Instructions: The
SuperiorScript III cdna Synthesis Kit Instruction Manual
 SuperiorScript III cdna Synthesis Kit Instruction Manual Cat.# EZ405S, EZ405M SuperiorScript III cdna Synthesis Kit Table of Contents I. Description... 3 II. Kit... 4 III. Procedure... 5 IV. Control Experiment
SuperiorScript III cdna Synthesis Kit Instruction Manual Cat.# EZ405S, EZ405M SuperiorScript III cdna Synthesis Kit Table of Contents I. Description... 3 II. Kit... 4 III. Procedure... 5 IV. Control Experiment
Puro. Knockout Detection (KOD) Kit
 Puro Knockout Detection (KOD) Kit Cat. No. CC-03 18 Oct. 2016 Contents I. Kit Contents and Storage II. Product Overview III. Methods Experimental Outline Genomic DNA Preparation Obtain Hybrid DNA Digest
Puro Knockout Detection (KOD) Kit Cat. No. CC-03 18 Oct. 2016 Contents I. Kit Contents and Storage II. Product Overview III. Methods Experimental Outline Genomic DNA Preparation Obtain Hybrid DNA Digest
Polymerase Chain Reaction (PCR) and Its Applications
 Polymerase Chain Reaction (PCR) and Its Applications What is PCR? PCR is an exponentially progressing synthesis of the defined target DNA sequences in vitro. It was invented in 1983 by Dr. Kary Mullis,
Polymerase Chain Reaction (PCR) and Its Applications What is PCR? PCR is an exponentially progressing synthesis of the defined target DNA sequences in vitro. It was invented in 1983 by Dr. Kary Mullis,
Pasteurella multocida
 BACTOTYPE PCR Amplification Kit Pasteurella multocida Labor Diagnostik Leipzig Manual Technology The product group BACTOTYPE PCR Amplification Kit comprises optimised systems for the identification of
BACTOTYPE PCR Amplification Kit Pasteurella multocida Labor Diagnostik Leipzig Manual Technology The product group BACTOTYPE PCR Amplification Kit comprises optimised systems for the identification of
Chapter 7. DNA Microarrays
 Bioinformatics III Structural Bioinformatics and Genome Analysis Chapter 7. DNA Microarrays 7.9 Next Generation Sequencing 454 Sequencing Solexa Illumina Solid TM System Sequencing Process of determining
Bioinformatics III Structural Bioinformatics and Genome Analysis Chapter 7. DNA Microarrays 7.9 Next Generation Sequencing 454 Sequencing Solexa Illumina Solid TM System Sequencing Process of determining
Towards whole genome association genetic scans in barley
 Towards whole genome association genetic scans in barley Varshney R., Kearsey M., Luo Z., Potokina E., Close T.J., Waugh R., Rostoks N., Ramsay L., Marshall D., Thomas B., Druka A., Wannamaker S., Svensson
Towards whole genome association genetic scans in barley Varshney R., Kearsey M., Luo Z., Potokina E., Close T.J., Waugh R., Rostoks N., Ramsay L., Marshall D., Thomas B., Druka A., Wannamaker S., Svensson
Phenotype analysis: biological-biochemical analysis. Genotype analysis: molecular and physical analysis
 1 Genetic Analysis Phenotype analysis: biological-biochemical analysis Behaviour under specific environmental conditions Behaviour of specific genetic configurations Behaviour of progeny in crosses - Genotype
1 Genetic Analysis Phenotype analysis: biological-biochemical analysis Behaviour under specific environmental conditions Behaviour of specific genetic configurations Behaviour of progeny in crosses - Genotype
Overview. Background ~30 min. Lab activity ~50 min. DNA profiling Polymerase Chain Reaction (PCR) Gel Electrophoresis PCR
 Overview Day 1: Tuesday Introduction to DNA profiling How do we use DNA to solve crimes? Background Polymerase Chain Reaction (PCR) Gel Electrophoresis Set up PCR Day 2: Wednesday Make and Run Agarose
Overview Day 1: Tuesday Introduction to DNA profiling How do we use DNA to solve crimes? Background Polymerase Chain Reaction (PCR) Gel Electrophoresis Set up PCR Day 2: Wednesday Make and Run Agarose
Real-Time PCR Principles and Applications
 Real-Time PCR Principles and Applications Dr Esam Ibraheem Azhar (BSc, MSc, Ph.D Molecular Medical Virology) Asst. Prof. Medical Laboratory Technology Department Objectives Real-Time PCR Principles and
Real-Time PCR Principles and Applications Dr Esam Ibraheem Azhar (BSc, MSc, Ph.D Molecular Medical Virology) Asst. Prof. Medical Laboratory Technology Department Objectives Real-Time PCR Principles and
Laboratory Exercise 4. Multiplex PCR of Short Tandem Repeats and Vertical Polyacrylamide Gel Electrophoresis.
 Laboratory Exercise 4 4 Multiplex PCR of Short Tandem Repeats and Vertical Polyacrylamide Gel Electrophoresis B A C K G R O U N D The human genome contains over 3000 million base pairs, which are distributed
Laboratory Exercise 4 4 Multiplex PCR of Short Tandem Repeats and Vertical Polyacrylamide Gel Electrophoresis B A C K G R O U N D The human genome contains over 3000 million base pairs, which are distributed
Reverse Transcription & RT-PCR
 Creating Gene Expression Solutions Reverse Transcription & RT-PCR Reverse transcription, a process that involves a reverse transcriptase (RTase) which uses RNA as the template to make complementary DNA
Creating Gene Expression Solutions Reverse Transcription & RT-PCR Reverse transcription, a process that involves a reverse transcriptase (RTase) which uses RNA as the template to make complementary DNA
Phenotype analysis: biological-biochemical analysis. Genotype analysis: molecular and physical analysis
 1 Genetic Analysis Phenotype analysis: biological-biochemical analysis Behaviour under specific environmental conditions Behaviour of specific genetic configurations Behaviour of progeny in crosses - Genotype
1 Genetic Analysis Phenotype analysis: biological-biochemical analysis Behaviour under specific environmental conditions Behaviour of specific genetic configurations Behaviour of progeny in crosses - Genotype
Discovery and development of exome-based, co-dominant single nucleotide polymorphism markers in hexaploid wheat (Triticum aestivum L.
 Application note GAPP-0001 Discovery and development of exome-based, co-dominant single nucleotide polymorphism markers in hexaploid wheat (Triticum aestivum L.) Keith J. Edwards, University of Bristol,
Application note GAPP-0001 Discovery and development of exome-based, co-dominant single nucleotide polymorphism markers in hexaploid wheat (Triticum aestivum L.) Keith J. Edwards, University of Bristol,
Ah, Lou! There really are differences between us!
 Name Per Ah, Lou! There really are differences between us! Introduction The human genome (the total sum of our genetic makeup) is made up of approximately 6 billion base pairs distributed on 46 chromosomes.
Name Per Ah, Lou! There really are differences between us! Introduction The human genome (the total sum of our genetic makeup) is made up of approximately 6 billion base pairs distributed on 46 chromosomes.
Exploring Genetic Variation in a Caffeine Metabolism gene LAB TWO: POLYMERASE CHAIN REACTION
 Exploring Genetic Variation in a Caffeine Metabolism gene LAB TWO: POLYMERASE CHAIN REACTION Purpose: In this laboratory, we will set up a polymerase chain reaction to amplify the region of the caffeine
Exploring Genetic Variation in a Caffeine Metabolism gene LAB TWO: POLYMERASE CHAIN REACTION Purpose: In this laboratory, we will set up a polymerase chain reaction to amplify the region of the caffeine
I.1 The Principle: Identification and Application of Molecular Markers
 I.1 The Principle: Identification and Application of Molecular Markers P. Langridge and K. Chalmers 1 1 Introduction Plant breeding is based around the identification and utilisation of genetic variation.
I.1 The Principle: Identification and Application of Molecular Markers P. Langridge and K. Chalmers 1 1 Introduction Plant breeding is based around the identification and utilisation of genetic variation.
PCR. CSIBD Molecular Genetics Course July 12, 2011 Michael Choi, M.D.
 PCR CSIBD Molecular Genetics Course July 12, 2011 Michael Choi, M.D. General Outline of the Lecture I. Background II. Basic Principles III. Detection and Analysis of PCR Products IV. Common Applications
PCR CSIBD Molecular Genetics Course July 12, 2011 Michael Choi, M.D. General Outline of the Lecture I. Background II. Basic Principles III. Detection and Analysis of PCR Products IV. Common Applications
Overview. Introduction
 Genetics 101: Introduction Overview Important terminology DNA extraction, gel electrophoresis, PCR Allozymes (Protein electrophoresis) RFLP AFLP Sequencing Microsatellites SNPs Costs, Sample Collection
Genetics 101: Introduction Overview Important terminology DNA extraction, gel electrophoresis, PCR Allozymes (Protein electrophoresis) RFLP AFLP Sequencing Microsatellites SNPs Costs, Sample Collection
Breeding rye (Secale cereale L.) for early fall-winter forage production
 Breeding rye (Secale cereale L.) for early fall-winter forage production M. C. Saha, J. L. Baker, and J. H. Bouton Forage Improvement Division, The Samuel Roberts Noble Foundation, Inc., 2510 Sam Noble
Breeding rye (Secale cereale L.) for early fall-winter forage production M. C. Saha, J. L. Baker, and J. H. Bouton Forage Improvement Division, The Samuel Roberts Noble Foundation, Inc., 2510 Sam Noble
Automated DNA isolation system GENE PREP STAR. March 16th, 2010 KURABO INDUSTRIES LTD. 1
 Automated DNA isolation system GENE PREP STAR PI-80X March 16th, 2010 KURABO INDUSTRIES LTD. 1 GENE PREP STAR PI-80X is a fully automated and flexible DNA isolation system. Key features Ease-of-use Built-in
Automated DNA isolation system GENE PREP STAR PI-80X March 16th, 2010 KURABO INDUSTRIES LTD. 1 GENE PREP STAR PI-80X is a fully automated and flexible DNA isolation system. Key features Ease-of-use Built-in
Comparison of ExoSAP-IT and ExoSAP-IT Express reagents to alternative PCR cleanup methods
 WHITE PAPER ExoSAP-IT PCR cleanup reagents Comparison of ExoSAP-IT and ExoSAP-IT Express reagents to alternative PCR cleanup methods Abstract Here we present superior workflow advantages of enzymatic PCR
WHITE PAPER ExoSAP-IT PCR cleanup reagents Comparison of ExoSAP-IT and ExoSAP-IT Express reagents to alternative PCR cleanup methods Abstract Here we present superior workflow advantages of enzymatic PCR
500U. Unit Definition. Storage Buffer. 10X PCR Buffer with Mg 2+
 Hy-Taq 500U + dntps #EZ1012 500U Concentration: 5U/μl Contents: Hy-Taq DNA Polymerase 100μl 10xPCR Buffer(Mg 2+ Plus) 1.25ml dntps(2.5mm each) 1ml 6xLoading Buffer 1ml Store at -20 C For research only
Hy-Taq 500U + dntps #EZ1012 500U Concentration: 5U/μl Contents: Hy-Taq DNA Polymerase 100μl 10xPCR Buffer(Mg 2+ Plus) 1.25ml dntps(2.5mm each) 1ml 6xLoading Buffer 1ml Store at -20 C For research only
 http://fire.biol.wwu.edu/trent/trent/direct_detection_of_genotype.html 1 Like most other model organism Arabidopsis thaliana has a sequenced genome? What do we mean by sequenced genome? What sort of info
http://fire.biol.wwu.edu/trent/trent/direct_detection_of_genotype.html 1 Like most other model organism Arabidopsis thaliana has a sequenced genome? What do we mean by sequenced genome? What sort of info
PrimeSTAR Max DNA Polymerase
 Cat. # R045A For Research Use PrimeSTAR Max DNA Polymerase Product Manual Table of Contents I. Description...3 II. III. IV. Components...3 Storage...3 General Composition of PCR Reaction Mixture...3 V.
Cat. # R045A For Research Use PrimeSTAR Max DNA Polymerase Product Manual Table of Contents I. Description...3 II. III. IV. Components...3 Storage...3 General Composition of PCR Reaction Mixture...3 V.
Blood direct 2x PCR Mastermix. Data sheet. Order No. BS reactions x 20 µl. (For research and in vitro applications only) Batch No.
 Data sheet Order No. BS91.222.0250 250 reactions x 20 µl Order No. BS91.222.1250 1250 reactions x 20 µl (For research and in vitro applications only) Batch No.: Best before: Appearance: Colour: 1 Description
Data sheet Order No. BS91.222.0250 250 reactions x 20 µl Order No. BS91.222.1250 1250 reactions x 20 µl (For research and in vitro applications only) Batch No.: Best before: Appearance: Colour: 1 Description
The Polymerase Chain Reaction. Chapter 6: Background
 The Polymerase Chain Reaction Chapter 6: Background Invention of PCR Kary Mullis Mile marker 46.58 in April of 1983 Pulled off the road and outlined a way to conduct DNA replication in a tube Worked for
The Polymerase Chain Reaction Chapter 6: Background Invention of PCR Kary Mullis Mile marker 46.58 in April of 1983 Pulled off the road and outlined a way to conduct DNA replication in a tube Worked for
NZYGene Synthesis kit
 Kit components Component Concentration Amount NZYGene Synthesis kit Catalogue number: MB33901, 10 reactions GS DNA Polymerase 1U/ μl 30 μl Reaction Buffer for GS DNA Polymerase 10 150 μl dntp mix 2 mm
Kit components Component Concentration Amount NZYGene Synthesis kit Catalogue number: MB33901, 10 reactions GS DNA Polymerase 1U/ μl 30 μl Reaction Buffer for GS DNA Polymerase 10 150 μl dntp mix 2 mm
Characterization of novel polymorphic genomic microsatellite markers of Boehmeria tricuspis (Hance) Makino
 Characterization of novel polymorphic genomic microsatellite markers of Boehmeria tricuspis (Hance) Makino Q. Tang, J.H. Chen, G.G. Zang and M.B. Luan Key Laboratory of Stem-fiber Biomass and Engineering
Characterization of novel polymorphic genomic microsatellite markers of Boehmeria tricuspis (Hance) Makino Q. Tang, J.H. Chen, G.G. Zang and M.B. Luan Key Laboratory of Stem-fiber Biomass and Engineering
What is DNA? Deoxyribonucleic Acid The inherited genetic material that makes us what we are
 DNA Basic Genetics What is DNA? DNA is Deoxyribonucleic Acid The inherited genetic material that makes us what we are DNA in the Cell Human Genome ~3 billion base pairs of DNA 30,000-35,000 genes Population-each
DNA Basic Genetics What is DNA? DNA is Deoxyribonucleic Acid The inherited genetic material that makes us what we are DNA in the Cell Human Genome ~3 billion base pairs of DNA 30,000-35,000 genes Population-each
Basic Steps of the DNA process
 As time pasted technology has improve the methods of analyzing DNA. One of the first methods for the analysis of DNA is known as Restriction Fragment Length Polymorphism (RFLP). This technique analyzed
As time pasted technology has improve the methods of analyzing DNA. One of the first methods for the analysis of DNA is known as Restriction Fragment Length Polymorphism (RFLP). This technique analyzed
EZ-GENXpress TM Normalized Gene Expression Detection Kit (Cat. #: EP-10001, EP & EP-10003)
 EZ-GENXpress TM Normalized Gene Expression Detection Kit (Cat. #: EP-10001, EP-10002 & EP-10003) INSTRUCTION MANUAL *These products are designed and sold for use in the Polymerase Chain Reaction (PCR)
EZ-GENXpress TM Normalized Gene Expression Detection Kit (Cat. #: EP-10001, EP-10002 & EP-10003) INSTRUCTION MANUAL *These products are designed and sold for use in the Polymerase Chain Reaction (PCR)
Functional Genomics Research Stream. Research Meeting: June 19, 2012 SYBR Green qpcr, Research Update
 Functional Genomics Research Stream Research Meeting: June 19, 2012 SYBR Green qpcr, Research Update Updates Alternate Lab Meeting Fridays 11:30-1:00 WEL 4.224 Welcome to attend either one Lab Log thanks
Functional Genomics Research Stream Research Meeting: June 19, 2012 SYBR Green qpcr, Research Update Updates Alternate Lab Meeting Fridays 11:30-1:00 WEL 4.224 Welcome to attend either one Lab Log thanks
Polymerase Chain Reaction PCR
 Polymerase Chain Reaction PCR What is PCR? An in vitro process that detects, identifies, and copies (amplifies) a specific piece of DNA in a biological sample. Discovered by Dr. Kary Mullis in 1983. A
Polymerase Chain Reaction PCR What is PCR? An in vitro process that detects, identifies, and copies (amplifies) a specific piece of DNA in a biological sample. Discovered by Dr. Kary Mullis in 1983. A
GenScript TissueDirect TM Multiplex PCR System
 GenScript TissueDirect TM Multiplex PCR System Technical Manual No. 0173 Version 20040908 I Description.. 1 II Applications 2 III Key Features.. 2 IV Shipping and Storage. 2 V Simplified Procedures. 2
GenScript TissueDirect TM Multiplex PCR System Technical Manual No. 0173 Version 20040908 I Description.. 1 II Applications 2 III Key Features.. 2 IV Shipping and Storage. 2 V Simplified Procedures. 2
Cat. # R100A. For Research Use. EpiScope MSP Kit. Product Manual. v201712da_2
 Cat. # R100A For Research Use EpiScope MSP Kit Product Manual Table of Contents I. Description... 3 II. MSP Principle... 3 III. Components... 4 IV. Storage... 4 V. Materials Required but not Provided...
Cat. # R100A For Research Use EpiScope MSP Kit Product Manual Table of Contents I. Description... 3 II. MSP Principle... 3 III. Components... 4 IV. Storage... 4 V. Materials Required but not Provided...
Development of Kompetitive Allele Specific PCR Markers for Submergence Tolerant Gene Sub1 in Rice
 Plant Breed. Biotech. 2019 (March) 7(1):62~66 https://doi.org/10.9787/pbb.2019.7.1.62 RAPID COMMUNICATION Online ISSN: 2287-9366 Print ISSN: 2287-9358 Development of Kompetitive Allele Specific PCR Markers
Plant Breed. Biotech. 2019 (March) 7(1):62~66 https://doi.org/10.9787/pbb.2019.7.1.62 RAPID COMMUNICATION Online ISSN: 2287-9366 Print ISSN: 2287-9358 Development of Kompetitive Allele Specific PCR Markers
Amplicon Sequencing Template Preparation
 Amplicon Sequencing Template Preparation The DNA sample preparation procedure for Amplicon Sequencing consists of a simple PCR amplification reaction, but uses special Fusion Primers (Figure 1-1). The
Amplicon Sequencing Template Preparation The DNA sample preparation procedure for Amplicon Sequencing consists of a simple PCR amplification reaction, but uses special Fusion Primers (Figure 1-1). The
FMF NIRCA PROTOCOL STEP 1.
 FMF NIRCA PROTOCOL STEP 1. After you have isolated patient s DNA and DNA from a healthy donor (wild type), you perform a nested PCR. The primers used to amplify exon 2 and exon 10 of the mefv gene are
FMF NIRCA PROTOCOL STEP 1. After you have isolated patient s DNA and DNA from a healthy donor (wild type), you perform a nested PCR. The primers used to amplify exon 2 and exon 10 of the mefv gene are
