Chemical hijacking of auxin signaling with an engineered auxin-tir1
|
|
|
- Shanon Lee
- 6 years ago
- Views:
Transcription
1 1 SUPPLEMENTARY INFORMATION Chemical hijacking of auxin signaling with an engineered auxin-tir1 pair Naoyuki Uchida 1,2*, Koji Takahashi 2*, Rie Iwasaki 1, Ryotaro Yamada 2, Masahiko Yoshimura 2, Takaho A. Endo 3, Seisuke Kimura 4, Hua Zhang 1, Mika Nomoto 2, Yasuomi Tada 5, Toshinori Kinoshita 1,2, Kenichiro Itami 1,2, Shinya Hagihara 2,6, and Keiko U. Torii 1,2,7,8, 1 Institute of Transformative Bio-Molecules (WPI-ITbM), Nagoya University, Chikusa, Nagoya, , Japan 2 Graduate School of Science, Nagoya University, Chikusa, Nagoya, , Japan 3 Laboratory for Integrative Genomics, RIKEN Center for Integrative Medical Sciences, Yokohama , Japan 4 Department of Bioresource and Environmental Sciences, Kyoto Sangyo University, Kyoto, , Japan 5 Center for Gene Research, Nagoya University, Chikusa, Nagoya, , Japan 6 PRESTO, Japan Science and Technology Agency, Kawaguchi, Saitama, , Japan 7 Howard Hughes Medical Institute, University of Washington, Seattle, WA , USA 8 Department of Biology, University of Washington, Seattle, WA , USA * Co-first authors Lead contact: ktorii@u.washington.edu Correspondence: hagi@itbm.nagoya-u.ac.jp and ktorii@u.washington.edu
2 2 Table of Contents Supplementary Figures 1 to 14 Supplementary Tables 1 to 3 Supplementary Dataset 1: Differentially expressed genes in wild type, 35S::TIR1 and 35S::ccvTIR1 by IAA and cvxiaa treatments Supplementary Note 1: Synthetic Procedures NMR Spectra of Indole-3-Acetic Acid Derivatives Synthesized in This Study
3 3 Supplementary Fig. 1. Additional yeast-two hybrid analysis of cvxiaa-ccvtir1 pairs. (a) The DNA binding domain (LexA)-fused TIR1 and the activation-domain-fused IAA3 (DI and DII domains) was tested in the presence of serial dilutions of IAA. The blue color indicates protein-protein interactions. (b) TIR1 or TIR1 F79G fused with the LexA DNA binding domain and the activation-domain-fused IAA3 (DI and DII domains) were tested in the presence of 100 µm IAA, 2,4-D or 1-NAA. (c) TIR1 F79G fused with the LexA DNA binding domain and the activation-domain-fused IAA3 (DI and DII domains) were tested in the presence of indicted concentrations of IAA and 5-(3-MeOPh)-IAA (cvxiaa). (d) Chemical structure of 4-phenyl-indole-3-acetic acid (4-Ph-IAA), which was chosen as a candidate ligand for TIR1 engineered at the F82 position. (e) Association of three different site-directed mutagenized TIR1s at the F82 position fused with the LexA DNA binding domain and the activation-domain-fused IAA3 (DI and DII domains) were tested in the presence of serial dilutions of IAA or 4-Ph-IAA.
4 4 Supplementary Fig. 2. Original gel blot for dose-response assays of TIR1 and ccvtir1 association with the IAA7 DII peptide and quantitative analysis of signal intensity. (a) The original gel blots of three independent pull-down assays for TIR1 and ccvtir1 by the IAA7 DII peptide. Concentration of IAA or cvxiaa are on the top of each lane. Input is an aliquot of control TIR1 or ccvtir1 solution used for pull down. For uncropped blot data, see Supplementary Fig. 13. (b) Quantitative analysis of signal intensity. Dots, mean values at each concentration. IAA-TIR1 pair, green; cvxiaa-ccvtir pair, pink. Error bars, standard deviation. n=5. For individual data points and fitted dose-response curve, see Fig. 2b, c.
5 5 Supplementary Fig. 3. Protein levels of the recombinant FLAG-TIR1 in the seedlings. (a) Shown are representative blots of protein extracts from light-grown Arabidopsis wild-type (WT) and transgenic seedlings expressing 35S::TIR1 (line A) and 35S::ccvTIR1 (line B). TIR1 and ccvtir1 proteins were detected using anti-flag antibody (FLAG-TIR1). Asterisk, a nonspecific signal. As a control, transferred membrane was stained with Panceau-S as well as subjected to immune-detection with anti-h + -ATPase antibody. Independent experiments were performed three times. For uncropped blot data, see Supplementary Fig. 14. (b) qrt-pcr analysis of the light-grown Arabidopsis wild-type (WT) and transgenic seedlings expressing 35S::TIR1 (line A) and 35S::ccvTIR1 (line B), confirming that the transgenes are expressed at a similar level. Bars, mean values. Error bars, standard deviation. The expression levels with three biological replicates were normalized to that of ACT2, and the normalized mean value of 35S::TIR1 was set at 1. Dots, the exact data points of individual samples. nd, not detected.
6 6 Supplementary Fig. 4. Auxin output reporter GFP expression in roots from wild-type, 35S::TIR1, and 35S::ccvTIR1 transgenic lines. A complete set of data for Fig. 3a. DR5::GFP signals in roots in response to IAA or cvxiaa treatment. Shown are the fluorescent micrographs of 4-day-old roots from Arabidopsis DR5::GFP seedlings (wild-type (WT), two independent lines (Line [Ln] A and B) of 35S::TIR1, and two independent lines of 35S::ccvTIR1) treated with mock, indicated concentrations of IAA and cvxiaa for 24 hours. In the absence of auxin treatment (mock), GFP signals are detected only in the root tips (asterisks). Strong GFP signals are detected in WT and 35S::TIR1 only under the IAA treatment. In contrast, both IAA and cvxiaa treatments elicit strong GFP signals in 35S::ccvTIR1 roots. Dotted line, root outline. Scale bars, 300 m.
7 7 Supplementary Fig. 5. cvxiaa treatment does not substantially affect root gravitropic response. (a) Shown are DR5::GFP seedlings grown for five days in a control media plate (DMSO), a plate containing cvxiaa (0.1 M and 0.3 M) and NPA at the condition used for polar auxin transport inhibition assays (10 M), then subjected to a gravistimulus by rotating agar plates 90 for 48 hours. Three independent experiments were performed. Images are taken at the same magnification. Scale bar, 1 cm. (b) Quantitative analysis of gravitropic response. Mean values and s.e.m of the angle (degree) of final root growth direction with respect to gravity are indicated. DMSO, n=28; 0.1 M cvxiaa, n=27; 0.3 M cvxiaa, n=26; NPA, n=28. p values, Welch's unpaired T-test comapared to the DMSO control. NS, not significant. (c) DR5::GFP signals at the root tip of seedlings grown and subjected to gravistimulus as described in (a). The cvxiaa (0.1 M and 0.3 M) treatments confer negligible effects on auxin maxima (GFP signals). Scale bar, 100 m
8 8 Supplementary Fig. 6. cvxiaa induces an IAA-like transcriptional response only in the presence of ccvtir1. Shown are scatter plots of transcriptional response to IAA or cvxiaa treatments by RNA-seq analyses. Differentially expressed genes (DEGs) for each genotype/treatment over a mock treatment of the corresponding genotype are plotted. (a) Transcriptional response between 35S::TIR1 seedlings treated with cvxiaa and 35S::ccvTIR1 seedlings treated with cvxiaa. No correlation is detected (r=0.024). (b) Transcriptional response between 35S::TIR1 seedlings treated with IAA and 35S::ccvTIR1 seedlings treated with IAA. High correlation is detected (r=0.863), indicating that overexpression of ccvtir1 does not interfere with the IAA perception by endogenous TIR1 and AFB proteins. (c) Transcriptional response between 35S::TIR1 seedlings treated with cvxiaa and 35S::ccvTIR1 seedlings treated with IAA. No correlation is detected (r=0.065), indicating that cvxiaa treatment to the wild-type TIR1 overexpressor does not trigger auxin response. (d) Transcriptional response between 35S::TIR1 seedlings treated with IAA and 35S::ccvTIR1 seedlings treated with cvxiaa. High correlation is detected (r=0.787), indicating that cvxiaa treatment to ccvtir1 overexpressor triggers auxin transcriptional response.
9 9 Supplementary Fig. 7. Inhibition of root growth of wild-type, 35S::TIR1, and 35S::ccvTIR1 transgenic lines in response to IAA or cvxiaa treatment. (a) Shown are 8-day-old seedlings (wild-type (WT), two independent lines (Line [Ln] A and B) of 35S::TIR1, and two independent lines of 35S::ccvTIR1) treated with mock or indicated concentrations of IAA and cvxiaa for 1 week. (b) Quantitative analysis of root elongation inhibition by IAA treatment. n=10. Experiments were performed three times. The mean value of mock-treatment was set at 100 in each genotype. The phenotype (root lengths) of each genotype and treatment are categorized into different classes by ANOVA followed by Tukey s HSD analysis. (c) Quantitative analysis of root elongation inhibition by cvxiaa treatment. n=10. Experiments were performed three times. The mean value of mock-treatment was set at 100 in each genotype. The phenotype (root lengths) of each genotype and treatment are categorized into different classes by ANOVA followed by Tukey s HSD analysis. For (b) and (c), box-and-whisker plots show a median (centerline), upper/lower quartiles (box limits) and maximum/minimum (upper/lower whiskers).
10 10 Supplementary Fig. 8. Lateral root density of wild-type, 35S::TIR1, and 35S::ccvTIR1 transgenic lines in response to 1-NAA or cvxiaa treatment. A complete set of data for Figure 4. (a) Representative images of eight-day-old Arabidopsis seedlings from wild type (WT), two independent lines (Line [Ln] A and B) of 35S::TIR1, and two independent lines of 35S::ccvTIR1 treated with mock or indicated concentrations of 1- NAA or cvxiaa for 3 days. (b) Representative images of 8-day-old Arabidopsis seedlings from wild type (WT), slr, two independent lines (Line [Ln] A and B) of 35S::ccvTIR1 with/without slr treated with 1 µm 1-NAA or cvxiaa for 3 days. Images were taken under the same magnification. Scale bar, 1 cm.
11 11 Supplementary Fig. 9. Inhibition of root growth of wild-type, tir1 afb2, TIR1pro::TIR1 tir1 afb2, and TIR1pro::ccvTIR1 tir1 afb2 in response to 1-NAA or cvxiaa treatment. (a) Shown are 8-day-old seedlings (wild-type (WT), tir1 afb2, two independent lines (Line [Ln] A and B) of TIR1pro::TIR1-GUS [TIR1pro::TIR1] in tir1 afb2, and two independent lines of TIR1pro::ccvTIR1-GUS [TIR1pro::ccvTIR1] in tir1 afb2) treated with mock or indicated concentrations of 1-NAA and cvxiaa for 1 week. (b) Quantitative analysis of root elongation inhibition by 1-NAA treatment. n=10. Experiments were performed three times. The mean value of mock-treatment was set at 100 in each genotype. The phenotype (root lengths) of each genotype and treatment are categorized into different classes by ANOVA followed by Tukey s HSD analysis. Note that TIR1pro::TIR1 rescues the auxin insensitivity of tir1 afb2 to the wild-type level (see categories d and e for Tukey's HSD analysis; blue). In contrast, TIR1pro::ccvTIR1 does not rescue the auxin insensitivity of tir1 afb2 (see category c for Tukey's HSD analysis; red). (c) Quantitative analysis of root elongation inhibition by cvxiaa treatment. Bars, mean root length of ten seedlings. n=10. Experiments were performed three times. The phenotype (root lengths) of each genotype and treatment are categorized into different classes by ANOVA followed by Tukey s HSD analysis. For (b) and (c), box-andwhisker plots show a median (centerline), upper/lower quartiles (box limits) and maximum/minimum (upper/lower whiskers).
12 12 Supplementary Fig. 10. cvxiaa treatment rapidly induces SAUR19 transcripts in etiolated seedlings expressing ccvtir1. Shown are qrt-pcr analysis of relative SAUR19 (a) and TUBULIN (TUB2) (b) expression levels from the hypocotyl segments of etiolated seedlings treated for 30 min with mock, IAA (magenta: 0.1 M), or cvxiaa (green: 0.1 M). Initial indicates hypocotyl segments harvested in prior to these treatments. TIR1pro::TIR1-GUS [TIR1pro::TIR1] in tir1 afb2 (Line A) and TIR1pro::ccvTIR1-GUS [TIR1pro::ccvTIR1] in tir1 afb2 (Line A) were used for the analysis. TUB2 was used as a control. Bars, mean values. Dots, the exact data points of individual samples. Error bars, standard error. The expression levels with three biological replicates were normalized to that of ACT2, and the normalized mean value of initial was set at 1. ** p < 0.001, *** p < , Student's t-test against mock treatment.
13 13 Supplementary Fig. 11. Uncropped blot images of Fig. 2a. The regions surrounded by dashed boxes are presented in Fig. 2a.
14 14 Supplementary Fig. 12. Uncropped blot images of Fig. 5c. The regions surrounded by dashed boxes are presented in Fig. 5c.
15 15 Supplementary Fig. 13. Uncropped blot images of Supplementary Fig. 2a. The regions surrounded by dashed boxes are presented in Supplementary Fig. 2a.
16 16 Supplementary Fig. 14. Uncropped blot images of Supplementary Fig. 3a. The regions surrounded by dashed boxes are presented in Supplementary Fig. 3a.
17 17 Supplementary Table 1. Mean and Standard Deviation for pull-down experiments IAA - TIR1 pair cvxiaa - ccvtir1 pair IAA (µm) Means SD n cvxiaa(µm) Means SD n
18 18 Supplementary Table 2. Plasmids used in this study
19 19 Supplementary Table 3. Oligo DNA primers used in this study
Supplemental Data. Steiner et al. Plant Cell. (2012) /tpc
 Supplemental Figure 1. SPY does not interact with free GST. Invitro pull-down assay using E. coli-expressed MBP-SPY and GST, GST-TCP14 and GST-TCP15. MBP-SPY was used as bait and incubated with equal amount
Supplemental Figure 1. SPY does not interact with free GST. Invitro pull-down assay using E. coli-expressed MBP-SPY and GST, GST-TCP14 and GST-TCP15. MBP-SPY was used as bait and incubated with equal amount
Supplementary Figure 1 qrt-pcr expression analysis of NLP8 with and without KNO 3 during germination.
 Supplementary Figure 1 qrt-pcr expression analysis of NLP8 with and without KNO 3 during germination. Seeds of Col-0 were harvested from plants grown at 16 C, stored for 2 months, imbibed for indicated
Supplementary Figure 1 qrt-pcr expression analysis of NLP8 with and without KNO 3 during germination. Seeds of Col-0 were harvested from plants grown at 16 C, stored for 2 months, imbibed for indicated
Enhancers mutations that make the original mutant phenotype more extreme. Suppressors mutations that make the original mutant phenotype less extreme
 Interactomics and Proteomics 1. Interactomics The field of interactomics is concerned with interactions between genes or proteins. They can be genetic interactions, in which two genes are involved in the
Interactomics and Proteomics 1. Interactomics The field of interactomics is concerned with interactions between genes or proteins. They can be genetic interactions, in which two genes are involved in the
A subclass of HSP70s regulate development and abiotic stress responses in Arabidopsis thaliana
 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 Journal of Plant Research A subclass of HSP70s regulate development and abiotic stress responses in Arabidopsis thaliana Linna Leng 1 Qianqian Liang
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 Journal of Plant Research A subclass of HSP70s regulate development and abiotic stress responses in Arabidopsis thaliana Linna Leng 1 Qianqian Liang
Supplementary Figure S1. Immunodetection of full-length XA21 and the XA21 C-terminal cleavage product.
 Supplementary Information Supplementary Figure S1. Immunodetection of full-length XA21 and the XA21 C-terminal cleavage product. Total protein extracted from Kitaake wild type and rice plants carrying
Supplementary Information Supplementary Figure S1. Immunodetection of full-length XA21 and the XA21 C-terminal cleavage product. Total protein extracted from Kitaake wild type and rice plants carrying
pgbkt7 Anti- Myc AH109 strain (KDa) 50
 pgbkt7 (KDa) 50 37 Anti- Myc AH109 strain Supplementary Figure 1. Protein expression of CRN and TDR in yeast. To analyse the protein expression of CRNKD and TDRKD, total proteins extracted from yeast culture
pgbkt7 (KDa) 50 37 Anti- Myc AH109 strain Supplementary Figure 1. Protein expression of CRN and TDR in yeast. To analyse the protein expression of CRNKD and TDRKD, total proteins extracted from yeast culture
Supplementary Figure 1. (a) The qrt-pcr for lnc-2, lnc-6 and lnc-7 RNA level in DU145, 22Rv1, wild type HCT116 and HCT116 Dicer ex5 cells transfected
 Supplementary Figure 1. (a) The qrt-pcr for lnc-2, lnc-6 and lnc-7 RNA level in DU145, 22Rv1, wild type HCT116 and HCT116 Dicer ex5 cells transfected with the sirna against lnc-2, lnc-6, lnc-7, and the
Supplementary Figure 1. (a) The qrt-pcr for lnc-2, lnc-6 and lnc-7 RNA level in DU145, 22Rv1, wild type HCT116 and HCT116 Dicer ex5 cells transfected with the sirna against lnc-2, lnc-6, lnc-7, and the
Supplementary Information
 Supplementary Information Live imaging reveals the dynamics and regulation of mitochondrial nucleoids during the cell cycle in Fucci2-HeLa cells Taeko Sasaki 1, Yoshikatsu Sato 2, Tetsuya Higashiyama 1,2,
Supplementary Information Live imaging reveals the dynamics and regulation of mitochondrial nucleoids during the cell cycle in Fucci2-HeLa cells Taeko Sasaki 1, Yoshikatsu Sato 2, Tetsuya Higashiyama 1,2,
Jung-Nam Cho, Jee-Youn Ryu, Young-Min Jeong, Jihye Park, Ji-Joon Song, Richard M. Amasino, Bosl Noh, and Yoo-Sun Noh
 Developmental Cell, Volume 22 Supplemental Information Control of Seed Germination by Light-Induced Histone Arginine Demethylation Activity Jung-Nam Cho, Jee-Youn Ryu, Young-Min Jeong, Jihye Park, Ji-Joon
Developmental Cell, Volume 22 Supplemental Information Control of Seed Germination by Light-Induced Histone Arginine Demethylation Activity Jung-Nam Cho, Jee-Youn Ryu, Young-Min Jeong, Jihye Park, Ji-Joon
Figure S6. Detection of anti-gfp antibodies in anti-dna and normal plasma without competition DNA--9
 Supplementary Information Ultrasensitive antibody detection by agglutination-pcr (ADAP) Cheng-ting Tsai 1 *, Peter V. Robinson 1 *, Carole A. Spencer 2 and Carolyn R. Bertozzi 3,4ǂ Department of 1 Chemistry,
Supplementary Information Ultrasensitive antibody detection by agglutination-pcr (ADAP) Cheng-ting Tsai 1 *, Peter V. Robinson 1 *, Carole A. Spencer 2 and Carolyn R. Bertozzi 3,4ǂ Department of 1 Chemistry,
Hossain_Supplemental Figure 1
 Hossain_Supplemental Figure 1 GFP-PACT GFP-PACT Motif I GFP-PACT Motif II A. MG132 (1µM) GFP Tubulin GFP-PACT Pericentrin GFP-PACT GFP-PACT Pericentrin Fig. S1. Expression and localization of Orc1 PACT
Hossain_Supplemental Figure 1 GFP-PACT GFP-PACT Motif I GFP-PACT Motif II A. MG132 (1µM) GFP Tubulin GFP-PACT Pericentrin GFP-PACT GFP-PACT Pericentrin Fig. S1. Expression and localization of Orc1 PACT
Nature Neuroscience: doi: /nn Supplementary Figure 1
 Supplementary Figure 1 PCR-genotyping of the three mouse models used in this study and controls for behavioral experiments after semi-chronic Pten inhibition. a-c. DNA from App/Psen1 (a), Pten tg (b) and
Supplementary Figure 1 PCR-genotyping of the three mouse models used in this study and controls for behavioral experiments after semi-chronic Pten inhibition. a-c. DNA from App/Psen1 (a), Pten tg (b) and
Supplemental Data. Lee et al. Plant Cell. (2010) /tpc Supplemental Figure 1. Protein and Gene Structures of DWA1 and DWA2.
 Supplemental Figure 1. Protein and Gene Structures of DWA1 and DWA2. (A) Protein structures of DWA1 and DWA2. WD40 region was determined based on the NCBI conserved domain databases (B, C) Schematic representation
Supplemental Figure 1. Protein and Gene Structures of DWA1 and DWA2. (A) Protein structures of DWA1 and DWA2. WD40 region was determined based on the NCBI conserved domain databases (B, C) Schematic representation
Construction of plant complementation vector and generation of transgenic plants
 MATERIAL S AND METHODS Plant materials and growth conditions Arabidopsis ecotype Columbia (Col0) was used for this study. SALK_072009, SALK_076309, and SALK_027645 were obtained from the Arabidopsis Biological
MATERIAL S AND METHODS Plant materials and growth conditions Arabidopsis ecotype Columbia (Col0) was used for this study. SALK_072009, SALK_076309, and SALK_027645 were obtained from the Arabidopsis Biological
Roche Molecular Biochemicals Technical Note No. LC 10/2000
 Roche Molecular Biochemicals Technical Note No. LC 10/2000 LightCycler Overview of LightCycler Quantification Methods 1. General Introduction Introduction Content Definitions This Technical Note will introduce
Roche Molecular Biochemicals Technical Note No. LC 10/2000 LightCycler Overview of LightCycler Quantification Methods 1. General Introduction Introduction Content Definitions This Technical Note will introduce
Highly efficient genome engineering in flowering plants ~ Development of a rapid method to knockout genes in Arabidopsis thaliana ~
 Highly efficient genome engineering in flowering plants ~ Development of a rapid method to knockout genes in Arabidopsis thaliana ~ December 5, 2016 Plant biologists at ITbM, Nagoya University have developed
Highly efficient genome engineering in flowering plants ~ Development of a rapid method to knockout genes in Arabidopsis thaliana ~ December 5, 2016 Plant biologists at ITbM, Nagoya University have developed
Technical Review. Real time PCR
 Technical Review Real time PCR Normal PCR: Analyze with agarose gel Normal PCR vs Real time PCR Real-time PCR, also known as quantitative PCR (qpcr) or kinetic PCR Key feature: Used to amplify and simultaneously
Technical Review Real time PCR Normal PCR: Analyze with agarose gel Normal PCR vs Real time PCR Real-time PCR, also known as quantitative PCR (qpcr) or kinetic PCR Key feature: Used to amplify and simultaneously
Recombinant DNA Technology. The Role of Recombinant DNA Technology in Biotechnology. yeast. Biotechnology. Recombinant DNA technology.
 PowerPoint Lecture Presentations prepared by Mindy Miller-Kittrell, North Carolina State University C H A P T E R 8 Recombinant DNA Technology The Role of Recombinant DNA Technology in Biotechnology Biotechnology?
PowerPoint Lecture Presentations prepared by Mindy Miller-Kittrell, North Carolina State University C H A P T E R 8 Recombinant DNA Technology The Role of Recombinant DNA Technology in Biotechnology Biotechnology?
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1.
 Supplementary Figure 1. Characterization and high expression of Lnc-β-Catm in liver CSCs. (a) Heatmap of differently expressed lncrnas in Liver CSCs (CD13 + CD133 + ) and non-cscs (CD13 - CD133 - ) according
Supplementary Figure 1. Characterization and high expression of Lnc-β-Catm in liver CSCs. (a) Heatmap of differently expressed lncrnas in Liver CSCs (CD13 + CD133 + ) and non-cscs (CD13 - CD133 - ) according
42 fl organelles = 34.5 fl (1) 3.5X X 0.93 = 78,000 (2)
 SUPPLEMENTAL DATA Supplementary Experimental Procedures Fluorescence Microscopy - A Zeiss Axiovert 200M microscope equipped with a Zeiss 100x Plan- Apochromat (1.40 NA) DIC objective and Hamamatsu Orca
SUPPLEMENTAL DATA Supplementary Experimental Procedures Fluorescence Microscopy - A Zeiss Axiovert 200M microscope equipped with a Zeiss 100x Plan- Apochromat (1.40 NA) DIC objective and Hamamatsu Orca
Coleman et al., Supplementary Figure 1
 Coleman et al., Supplementary Figure 1 BrdU Merge G1 Early S Mid S Supplementary Figure 1. Sequential destruction of CRL4 Cdt2 targets during the G1/S transition. HCT116 cells were synchronized by sequential
Coleman et al., Supplementary Figure 1 BrdU Merge G1 Early S Mid S Supplementary Figure 1. Sequential destruction of CRL4 Cdt2 targets during the G1/S transition. HCT116 cells were synchronized by sequential
Description: Nuclear morphology and dynamics in nontargeting sirna transfected cells. HeLa Kyoto
 Title of file for HTML: Supplementary Information Description: Supplementary Figures and Supplementary Tables Title of file for HTML: Supplementary Movie 1 Description: Nuclear morphology and dynamics
Title of file for HTML: Supplementary Information Description: Supplementary Figures and Supplementary Tables Title of file for HTML: Supplementary Movie 1 Description: Nuclear morphology and dynamics
GTTCGGGTTCC TTTTGAGCAG
 Supplementary Figures Splice variants of the SIP1 transcripts play a role in nodule organogenesis in Lotus japonicus. Wang C, Zhu H, Jin L, Chen T, Wang L, Kang H, Hong Z, Zhang Z. 5 UTR CDS 3 UTR TCTCAACCATCCTTTGTCTGCTTCCGCCGCATGGGTGAGGTCATTTTGTCTAGATGACGTGCAATTTACAATGA
Supplementary Figures Splice variants of the SIP1 transcripts play a role in nodule organogenesis in Lotus japonicus. Wang C, Zhu H, Jin L, Chen T, Wang L, Kang H, Hong Z, Zhang Z. 5 UTR CDS 3 UTR TCTCAACCATCCTTTGTCTGCTTCCGCCGCATGGGTGAGGTCATTTTGTCTAGATGACGTGCAATTTACAATGA
Supplementary Information
 Supplementary Information Supplementary Figure 1. ZBTB20 expression in the developing DRG. ZBTB20 expression in the developing DRG was detected by immunohistochemistry using anti-zbtb20 antibody 9A10 on
Supplementary Information Supplementary Figure 1. ZBTB20 expression in the developing DRG. ZBTB20 expression in the developing DRG was detected by immunohistochemistry using anti-zbtb20 antibody 9A10 on
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION Legends for Supplementary Tables. Supplementary Table 1. An excel file containing primary screen data. Worksheet 1, Normalized quantification data from a duplicated screen: valid
SUPPLEMENTARY INFORMATION Legends for Supplementary Tables. Supplementary Table 1. An excel file containing primary screen data. Worksheet 1, Normalized quantification data from a duplicated screen: valid
Supplementary Fig. S1. SAMHD1c has a more potent dntpase activity than. SAMHD1c. Purified recombinant SAMHD1c and SAMHD1c proteins (with
 Supplementary Fig. S1. SAMHD1c has a more potent dntpase activity than SAMHD1c. Purified recombinant SAMHD1c and SAMHD1c proteins (with concentration of 800nM) were incubated with 1mM dgtp for the indicated
Supplementary Fig. S1. SAMHD1c has a more potent dntpase activity than SAMHD1c. Purified recombinant SAMHD1c and SAMHD1c proteins (with concentration of 800nM) were incubated with 1mM dgtp for the indicated
SUPPLEMENTARY INFORMATION
 doi:.38/nature899 Supplementary Figure Suzuki et al. a c p7 -/- / WT ratio (+)/(-) p7 -/- / WT ratio Log X 3. Fold change by treatment ( (+)/(-)) Log X.5 3-3. -. b Fold change by treatment ( (+)/(-)) 8
doi:.38/nature899 Supplementary Figure Suzuki et al. a c p7 -/- / WT ratio (+)/(-) p7 -/- / WT ratio Log X 3. Fold change by treatment ( (+)/(-)) Log X.5 3-3. -. b Fold change by treatment ( (+)/(-)) 8
Fatchiyah
 Fatchiyah Email: fatchiya@yahoo.co.id RNAs: mrna trna rrna RNAi DNAs: Protein: genome DNA cdna mikro-makro mono-poly single-multi Analysis: Identification human and animal disease Finger printing Sexing
Fatchiyah Email: fatchiya@yahoo.co.id RNAs: mrna trna rrna RNAi DNAs: Protein: genome DNA cdna mikro-makro mono-poly single-multi Analysis: Identification human and animal disease Finger printing Sexing
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Origin use and efficiency are similar among WT, rrm3, pif1-m2, and pif1-m2; rrm3 strains. A. Analysis of fork progression around confirmed and likely origins (from cerevisiae.oridb.org).
Supplementary Figure 1 Origin use and efficiency are similar among WT, rrm3, pif1-m2, and pif1-m2; rrm3 strains. A. Analysis of fork progression around confirmed and likely origins (from cerevisiae.oridb.org).
% Viability. isw2 ino isw2 ino isw2 ino isw2 ino mM HU 4-NQO CPT
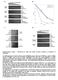 a Drug concentration b 1.3% MMS nhp1 nhp1 8 nhp1 mag1.5% MMS.3% MMS nhp1 nhp1 ino8 9 ino8 9 % Viability 4.5% MMS ino8 9 ino8 9 2.5.1.15 % MMS c d nhp1 nhp1 nhp1 nhp1 nhp1 nhp1 Control (YPD) γ IR (1 gy)
a Drug concentration b 1.3% MMS nhp1 nhp1 8 nhp1 mag1.5% MMS.3% MMS nhp1 nhp1 ino8 9 ino8 9 % Viability 4.5% MMS ino8 9 ino8 9 2.5.1.15 % MMS c d nhp1 nhp1 nhp1 nhp1 nhp1 nhp1 Control (YPD) γ IR (1 gy)
Supplementary Fig. 1 related to Fig. 1 Clinical relevance of lncrna candidate
 Supplementary Figure Legends Supplementary Fig. 1 related to Fig. 1 Clinical relevance of lncrna candidate BC041951 in gastric cancer. (A) The flow chart for selected candidate lncrnas in 660 up-regulated
Supplementary Figure Legends Supplementary Fig. 1 related to Fig. 1 Clinical relevance of lncrna candidate BC041951 in gastric cancer. (A) The flow chart for selected candidate lncrnas in 660 up-regulated
RNA oligonucleotides and 2 -O-methylated oligonucleotides were synthesized by. 5 AGACACAAACACCAUUGUCACACUCCACAGC; Rand-2 OMe,
 Materials and methods Oligonucleotides and DNA constructs RNA oligonucleotides and 2 -O-methylated oligonucleotides were synthesized by Dharmacon Inc. (Lafayette, CO). The sequences were: 122-2 OMe, 5
Materials and methods Oligonucleotides and DNA constructs RNA oligonucleotides and 2 -O-methylated oligonucleotides were synthesized by Dharmacon Inc. (Lafayette, CO). The sequences were: 122-2 OMe, 5
Parthanatos mediates AIMP2-activated age-dependent dopaminergic neuronal loss
 SUPPLEMENTARY INFORMATION Parthanatos mediates AIMP2-activated age-dependent dopaminergic neuronal loss Yunjong Lee, Senthilkumar S. Karuppagounder, Joo-Ho Shin, Yun-Il Lee, Han Seok Ko, Debbie Swing,
SUPPLEMENTARY INFORMATION Parthanatos mediates AIMP2-activated age-dependent dopaminergic neuronal loss Yunjong Lee, Senthilkumar S. Karuppagounder, Joo-Ho Shin, Yun-Il Lee, Han Seok Ko, Debbie Swing,
MIT Department of Biology 7.013: Introductory Biology - Spring 2005 Instructors: Professor Hazel Sive, Professor Tyler Jacks, Dr.
 MIT Department of Biology 7.01: Introductory Biology - Spring 2005 Instructors: Professor Hazel Sive, Professor Tyler Jacks, Dr. Claudette Gardel iv) Would Xba I be useful for cloning? Why or why not?
MIT Department of Biology 7.01: Introductory Biology - Spring 2005 Instructors: Professor Hazel Sive, Professor Tyler Jacks, Dr. Claudette Gardel iv) Would Xba I be useful for cloning? Why or why not?
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION Dynamic Phosphorylation of HP1 Regulates Mitotic Progression in Human Cells Supplementary Figures Supplementary Figure 1. NDR1 interacts with HP1. (a) Immunoprecipitation using
SUPPLEMENTARY INFORMATION Dynamic Phosphorylation of HP1 Regulates Mitotic Progression in Human Cells Supplementary Figures Supplementary Figure 1. NDR1 interacts with HP1. (a) Immunoprecipitation using
sides of the aleurone (Al) but it is excluded from the basal endosperm transfer layer
 Supplemental Data. Gómez et al. (2009). The maize transcription factor MRP-1 (Myb-Related-Protein-1) is a key regulator of the differentiation of transfer cells. Supplemental Figure 1. Expression analyses
Supplemental Data. Gómez et al. (2009). The maize transcription factor MRP-1 (Myb-Related-Protein-1) is a key regulator of the differentiation of transfer cells. Supplemental Figure 1. Expression analyses
Control of secondary cell wall patterning involves xylan deacetylation by a GDSL esterase
 In the format provided by the authors and unedited. SUPPLEMENTARY INFORMATION VOLUME: 3 ARTICLE NUMBER: 17017 Control of secondary cell wall patterning involves xylan deacetylation by a GDSL esterase Baocai
In the format provided by the authors and unedited. SUPPLEMENTARY INFORMATION VOLUME: 3 ARTICLE NUMBER: 17017 Control of secondary cell wall patterning involves xylan deacetylation by a GDSL esterase Baocai
ASPP1 Fw GGTTGGGAATCCACGTGTTG ASPP1 Rv GCCATATCTTGGAGCTCTGAGAG
 Supplemental Materials and Methods Plasmids: the following plasmids were used in the supplementary data: pwzl-myc- Lats2 (Aylon et al, 2006), pretrosuper-vector and pretrosuper-shp53 (generous gift of
Supplemental Materials and Methods Plasmids: the following plasmids were used in the supplementary data: pwzl-myc- Lats2 (Aylon et al, 2006), pretrosuper-vector and pretrosuper-shp53 (generous gift of
2 Gene Technologies in Our Lives
 CHAPTER 15 2 Gene Technologies in Our Lives SECTION Gene Technologies and Human Applications KEY IDEAS As you read this section, keep these questions in mind: For what purposes are genes and proteins manipulated?
CHAPTER 15 2 Gene Technologies in Our Lives SECTION Gene Technologies and Human Applications KEY IDEAS As you read this section, keep these questions in mind: For what purposes are genes and proteins manipulated?
sirna Transfection Into Primary Neurons Using Fuse-It-siRNA
 sirna Transfection Into Primary Neurons Using Fuse-It-siRNA This Application Note describes a protocol for sirna transfection into sensitive, primary cortical neurons using Fuse-It-siRNA. This innovative
sirna Transfection Into Primary Neurons Using Fuse-It-siRNA This Application Note describes a protocol for sirna transfection into sensitive, primary cortical neurons using Fuse-It-siRNA. This innovative
Molecular Cell Biology - Problem Drill 11: Recombinant DNA
 Molecular Cell Biology - Problem Drill 11: Recombinant DNA Question No. 1 of 10 1. Which of the following statements about the sources of DNA used for molecular cloning is correct? Question #1 (A) cdna
Molecular Cell Biology - Problem Drill 11: Recombinant DNA Question No. 1 of 10 1. Which of the following statements about the sources of DNA used for molecular cloning is correct? Question #1 (A) cdna
Nature Methods: doi: /nmeth Supplementary Figure 1. Retention of RNA with LabelX.
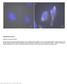 Supplementary Figure 1 Retention of RNA with LabelX. (a) Epi-fluorescence image of single molecule FISH (smfish) against GAPDH on HeLa cells expanded without LabelX treatment. (b) Epi-fluorescence image
Supplementary Figure 1 Retention of RNA with LabelX. (a) Epi-fluorescence image of single molecule FISH (smfish) against GAPDH on HeLa cells expanded without LabelX treatment. (b) Epi-fluorescence image
Human IL10RB ELISA Pair Set ( CRFB4 )
 Human IL10RB ELISA Pair Set ( CRFB4 ) Catalog Number : SEK10945 To achieve the best assay results, this manual must be read carefully before using this product and the assay is run as summarized in the
Human IL10RB ELISA Pair Set ( CRFB4 ) Catalog Number : SEK10945 To achieve the best assay results, this manual must be read carefully before using this product and the assay is run as summarized in the
Supplementary Figures Montero et al._supplementary Figure 1
 Montero et al_suppl. Info 1 Supplementary Figures Montero et al._supplementary Figure 1 Montero et al_suppl. Info 2 Supplementary Figure 1. Transcripts arising from the structurally conserved subtelomeres
Montero et al_suppl. Info 1 Supplementary Figures Montero et al._supplementary Figure 1 Montero et al_suppl. Info 2 Supplementary Figure 1. Transcripts arising from the structurally conserved subtelomeres
Supplemental Data. Aung et al. (2011). Plant Cell /tpc
 35S pro:pmd1-yfp 10 µm Supplemental Figure 1. -terminal YFP fusion of PMD1 (PMD1-YFP, in green) is localized to the cytosol. grobacterium cells harboring 35S pro :PMD1-YFP were infiltrated into tobacco
35S pro:pmd1-yfp 10 µm Supplemental Figure 1. -terminal YFP fusion of PMD1 (PMD1-YFP, in green) is localized to the cytosol. grobacterium cells harboring 35S pro :PMD1-YFP were infiltrated into tobacco
Figure 1: TDP-43 is subject to lysine acetylation within the RNA-binding domain a) QBI-293 cells were transfected with TDP-43 in the presence or
 Figure 1: TDP-43 is subject to lysine acetylation within the RNA-binding domain a) QBI-293 cells were transfected with TDP-43 in the presence or absence of the acetyltransferase CBP and acetylated TDP-43
Figure 1: TDP-43 is subject to lysine acetylation within the RNA-binding domain a) QBI-293 cells were transfected with TDP-43 in the presence or absence of the acetyltransferase CBP and acetylated TDP-43
Gene Expression Technology
 Gene Expression Technology Bing Zhang Department of Biomedical Informatics Vanderbilt University bing.zhang@vanderbilt.edu Gene expression Gene expression is the process by which information from a gene
Gene Expression Technology Bing Zhang Department of Biomedical Informatics Vanderbilt University bing.zhang@vanderbilt.edu Gene expression Gene expression is the process by which information from a gene
Supplemental Data. mir156-regulated SPL Transcription. Factors Define an Endogenous Flowering. Pathway in Arabidopsis thaliana
 Cell, Volume 138 Supplemental Data mir156-regulated SPL Transcription Factors Define an Endogenous Flowering Pathway in Arabidopsis thaliana Jia-Wei Wang, Benjamin Czech, and Detlef Weigel Table S1. Interaction
Cell, Volume 138 Supplemental Data mir156-regulated SPL Transcription Factors Define an Endogenous Flowering Pathway in Arabidopsis thaliana Jia-Wei Wang, Benjamin Czech, and Detlef Weigel Table S1. Interaction
qpcr Quantitative PCR or Real-time PCR Gives a measurement of PCR product at end of each cycle real time
 qpcr qpcr Quantitative PCR or Real-time PCR Gives a measurement of PCR product at end of each cycle real time Differs from endpoint PCR gel on last cycle Used to determines relative amount of template
qpcr qpcr Quantitative PCR or Real-time PCR Gives a measurement of PCR product at end of each cycle real time Differs from endpoint PCR gel on last cycle Used to determines relative amount of template
FIG S1: Calibration curves of standards for HPLC detection of Dopachrome (Dopac), Dopamine (DA) and Homovanillic acid (HVA), showing area of peak vs
 FIG S1: Calibration curves of standards for HPLC detection of Dopachrome (Dopac), Dopamine (DA) and Homovanillic acid (HVA), showing area of peak vs amount of standard analysed. Each point represents the
FIG S1: Calibration curves of standards for HPLC detection of Dopachrome (Dopac), Dopamine (DA) and Homovanillic acid (HVA), showing area of peak vs amount of standard analysed. Each point represents the
SUPPLEMENTARY INFORMATION
 a before amputation regeneration regenerated limb DERMIS SKELETON MUSCLE SCHWANN CELLS EPIDERMIS DERMIS SKELETON MUSCLE SCHWANN CELLS EPIDERMIS developmental origin: lateral plate mesoderm presomitic mesoderm
a before amputation regeneration regenerated limb DERMIS SKELETON MUSCLE SCHWANN CELLS EPIDERMIS DERMIS SKELETON MUSCLE SCHWANN CELLS EPIDERMIS developmental origin: lateral plate mesoderm presomitic mesoderm
Supplementary Figure 1. Bone density was decreased in osteoclast-lineage cell specific Gna13 deficient mice. (a-c) PCR genotyping of mice by mouse
 Supplementary Figure 1. Bone density was decreased in osteoclast-lineage cell specific Gna13 deficient mice. (a-c) PCR genotyping of mice by mouse tail DNA. Primers were designed to detect Gna13-WT/f (~400bp/470bp)
Supplementary Figure 1. Bone density was decreased in osteoclast-lineage cell specific Gna13 deficient mice. (a-c) PCR genotyping of mice by mouse tail DNA. Primers were designed to detect Gna13-WT/f (~400bp/470bp)
Supplementary Table 1. The Q-PCR primer sequence is summarized in the following table.
 Supplementary Table 1. The Q-PCR primer sequence is summarized in the following table. Name Sequence (5-3 ) Application Flag-u ggactacaaggacgacgatgac Shared upstream primer for all the amplifications of
Supplementary Table 1. The Q-PCR primer sequence is summarized in the following table. Name Sequence (5-3 ) Application Flag-u ggactacaaggacgacgatgac Shared upstream primer for all the amplifications of
Biotechnology. Chapter 20. Biology Eighth Edition Neil Campbell and Jane Reece. PowerPoint Lecture Presentations for
 Chapter 20 Biotechnology PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley with contributions from Joan Sharp Copyright
Chapter 20 Biotechnology PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley with contributions from Joan Sharp Copyright
Contents... vii. List of Figures... xii. List of Tables... xiv. Abbreviatons... xv. Summary... xvii. 1. Introduction In vitro evolution...
 vii Contents Contents... vii List of Figures... xii List of Tables... xiv Abbreviatons... xv Summary... xvii 1. Introduction...1 1.1 In vitro evolution... 1 1.2 Phage Display Technology... 3 1.3 Cell surface
vii Contents Contents... vii List of Figures... xii List of Tables... xiv Abbreviatons... xv Summary... xvii 1. Introduction...1 1.1 In vitro evolution... 1 1.2 Phage Display Technology... 3 1.3 Cell surface
GFP CCD2 GFP IP:GFP
 D1 D2 1 75 95 148 178 492 GFP CCD1 CCD2 CCD2 GFP D1 D2 GFP D1 D2 Beclin 1 IB:GFP IP:GFP Supplementary Figure 1: Mapping domains required for binding to HEK293T cells are transfected with EGFP-tagged mutant
D1 D2 1 75 95 148 178 492 GFP CCD1 CCD2 CCD2 GFP D1 D2 GFP D1 D2 Beclin 1 IB:GFP IP:GFP Supplementary Figure 1: Mapping domains required for binding to HEK293T cells are transfected with EGFP-tagged mutant
Supplemental Information. Boundary Formation through a Direct. Threshold-Based Readout. of Mobile Small RNA Gradients
 Developmental Cell, Volume 43 Supplemental Information Boundary Formation through a Direct Threshold-Based Readout of Mobile Small RNA Gradients Damianos S. Skopelitis, Anna H. Benkovics, Aman Y. Husbands,
Developmental Cell, Volume 43 Supplemental Information Boundary Formation through a Direct Threshold-Based Readout of Mobile Small RNA Gradients Damianos S. Skopelitis, Anna H. Benkovics, Aman Y. Husbands,
Mouse ICAM-1 / CD54 ELISA Pair Set
 Mouse ICAM-1 / CD54 ELISA Pair Set Catalog Number : SEK50440 To achieve the best assay results, this manual must be read carefully before using this product and the assay is run as summarized in the General
Mouse ICAM-1 / CD54 ELISA Pair Set Catalog Number : SEK50440 To achieve the best assay results, this manual must be read carefully before using this product and the assay is run as summarized in the General
BIO 315 Lab Exam I. Section #: Name:
 Section #: Name: Also provide this information on the computer grid sheet given to you. (Section # in special code box) BIO 315 Lab Exam I 1. In labeling the parts of a standard compound light microscope
Section #: Name: Also provide this information on the computer grid sheet given to you. (Section # in special code box) BIO 315 Lab Exam I 1. In labeling the parts of a standard compound light microscope
Recent technology allow production of microarrays composed of 70-mers (essentially a hybrid of the two techniques)
 Microarrays and Transcript Profiling Gene expression patterns are traditionally studied using Northern blots (DNA-RNA hybridization assays). This approach involves separation of total or polya + RNA on
Microarrays and Transcript Profiling Gene expression patterns are traditionally studied using Northern blots (DNA-RNA hybridization assays). This approach involves separation of total or polya + RNA on
Nature Structural and Molecular Biology: doi: /nsmb.2937
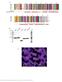 Supplementary Figure 1 Multiple sequence alignment of the CtIP N-terminal domain, purified CtIP protein constructs and details of the 2F o F c electron density map of CtIP-NTD. (a) Multiple sequence alignment,
Supplementary Figure 1 Multiple sequence alignment of the CtIP N-terminal domain, purified CtIP protein constructs and details of the 2F o F c electron density map of CtIP-NTD. (a) Multiple sequence alignment,
CASE-STUDY- VALIDATION of PCR based methodology. Beata Surmacz-Cordle Senior Analytical Development Scientist
 CASE-STUDY- VALIDATION of PCR based methodology Beata Surmacz-Cordle Senior Analytical Development Scientist UK RMP Pluripotent Stem Cell Platform Validation Workshop 2 nd June 2016 RT-qPCR assay for detection
CASE-STUDY- VALIDATION of PCR based methodology Beata Surmacz-Cordle Senior Analytical Development Scientist UK RMP Pluripotent Stem Cell Platform Validation Workshop 2 nd June 2016 RT-qPCR assay for detection
Supplemental Data Supplemental Figure 1.
 Supplemental Data Supplemental Figure 1. Silique arrangement in the wild-type, jhs, and complemented lines. Wild-type (WT) (A), the jhs1 mutant (B,C), and the jhs1 mutant complemented with JHS1 (Com) (D)
Supplemental Data Supplemental Figure 1. Silique arrangement in the wild-type, jhs, and complemented lines. Wild-type (WT) (A), the jhs1 mutant (B,C), and the jhs1 mutant complemented with JHS1 (Com) (D)
Chapter 15 Gene Technologies and Human Applications
 Chapter Outline Chapter 15 Gene Technologies and Human Applications Section 1: The Human Genome KEY IDEAS > Why is the Human Genome Project so important? > How do genomics and gene technologies affect
Chapter Outline Chapter 15 Gene Technologies and Human Applications Section 1: The Human Genome KEY IDEAS > Why is the Human Genome Project so important? > How do genomics and gene technologies affect
Supplementary Figure 1
 Supplementary Figure 1 Supplementary Figure 1: Vector maps of TRMPV and TRMPVIR variants. Many derivatives of TRMPV have been generated and tested. Unless otherwise noted, experiments in this paper use
Supplementary Figure 1 Supplementary Figure 1: Vector maps of TRMPV and TRMPVIR variants. Many derivatives of TRMPV have been generated and tested. Unless otherwise noted, experiments in this paper use
2054, Chap. 14, page 1
 2054, Chap. 14, page 1 I. Recombinant DNA technology (Chapter 14) A. recombinant DNA technology = collection of methods used to perform genetic engineering 1. genetic engineering = deliberate modification
2054, Chap. 14, page 1 I. Recombinant DNA technology (Chapter 14) A. recombinant DNA technology = collection of methods used to perform genetic engineering 1. genetic engineering = deliberate modification
A) B) Ladder. Supplementary Figure 1. Recombinant calpain 14 purification analysis. A) The general domain structure of classical
 A) B) Ladder C) r4 r4 Nt- -Ct 78 kda Supplementary Figure 1. Recombinant calpain 14 purification analysis. A) The general domain structure of classical calpains is shown. The protease core consists of
A) B) Ladder C) r4 r4 Nt- -Ct 78 kda Supplementary Figure 1. Recombinant calpain 14 purification analysis. A) The general domain structure of classical calpains is shown. The protease core consists of
Supplementary Figure 1. Isolation of GFPHigh cells.
 Supplementary Figure 1. Isolation of GFP High cells. (A) Schematic diagram of cell isolation based on Wnt signaling activity. Colorectal cancer (CRC) cell lines were stably transduced with lentivirus encoding
Supplementary Figure 1. Isolation of GFP High cells. (A) Schematic diagram of cell isolation based on Wnt signaling activity. Colorectal cancer (CRC) cell lines were stably transduced with lentivirus encoding
BIOTECHNOLOGY. Course Syllabus. Section A: Engineering Mathematics. Subject Code: BT. Course Structure. Engineering Mathematics. General Biotechnology
 BIOTECHNOLOGY Subject Code: BT Course Structure Sections/Units Section A Section B Unit 1 Unit 2 Unit 3 Unit 4 Unit 5 Unit 6 Unit 7 Section C Section D Section E Topics Engineering Mathematics General
BIOTECHNOLOGY Subject Code: BT Course Structure Sections/Units Section A Section B Unit 1 Unit 2 Unit 3 Unit 4 Unit 5 Unit 6 Unit 7 Section C Section D Section E Topics Engineering Mathematics General
Supplementary Figure 1 Two distinct conformational states of the HNH domain in crystal structures. a
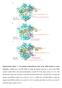 Supplementary Figure 1 Two distinct conformational states of the HNH domain in crystal structures. a HNH-state 1 in PDB 4OO8, in which the distance from the C atom of the HNH catalytic residue 840 to the
Supplementary Figure 1 Two distinct conformational states of the HNH domain in crystal structures. a HNH-state 1 in PDB 4OO8, in which the distance from the C atom of the HNH catalytic residue 840 to the
Figure S1. Immunoblotting showing proper expression of tagged R and AVR proteins in N. benthamiana.
 SUPPLEMENTARY FIGURES Figure S1. Immunoblotting showing proper expression of tagged R and AVR proteins in N. benthamiana. YFP:, :CFP, HA: and (A) and HA, CFP and YFPtagged and AVR1-CO39 (B) were expressed
SUPPLEMENTARY FIGURES Figure S1. Immunoblotting showing proper expression of tagged R and AVR proteins in N. benthamiana. YFP:, :CFP, HA: and (A) and HA, CFP and YFPtagged and AVR1-CO39 (B) were expressed
Molecular Genetics Techniques. BIT 220 Chapter 20
 Molecular Genetics Techniques BIT 220 Chapter 20 What is Cloning? Recombinant DNA technologies 1. Producing Recombinant DNA molecule Incorporate gene of interest into plasmid (cloning vector) 2. Recombinant
Molecular Genetics Techniques BIT 220 Chapter 20 What is Cloning? Recombinant DNA technologies 1. Producing Recombinant DNA molecule Incorporate gene of interest into plasmid (cloning vector) 2. Recombinant
Roche Molecular Biochemicals Technical Note No. LC 9/2000
 Roche Molecular Biochemicals Technical Note No. LC 9/2000 LightCycler Optimization Strategy Introduction Purpose of this Note Table of Contents The LightCycler system provides different detection formats
Roche Molecular Biochemicals Technical Note No. LC 9/2000 LightCycler Optimization Strategy Introduction Purpose of this Note Table of Contents The LightCycler system provides different detection formats
Chapter 20: Biotechnology
 Name Period The AP Biology exam has reached into this chapter for essay questions on a regular basis over the past 15 years. Student responses show that biotechnology is a difficult topic. This chapter
Name Period The AP Biology exam has reached into this chapter for essay questions on a regular basis over the past 15 years. Student responses show that biotechnology is a difficult topic. This chapter
GM130 Is Required for Compartmental Organization of Dendritic Golgi Outposts
 Current Biology, Volume 24 Supplemental Information GM130 Is Required for Compartmental Organization of Dendritic Golgi Outposts Wei Zhou, Jin Chang, Xin Wang, Masha G. Savelieff, Yinyin Zhao, Shanshan
Current Biology, Volume 24 Supplemental Information GM130 Is Required for Compartmental Organization of Dendritic Golgi Outposts Wei Zhou, Jin Chang, Xin Wang, Masha G. Savelieff, Yinyin Zhao, Shanshan
Chapter 17: Immunization & Immune Testing. 1. Immunization 2. Diagnostic Immunology
 Chapter 17: Immunization & Immune Testing 1. Immunization 2. Diagnostic Immunology 1. Immunization Chapter Reading pp. 505-511 What is Immunization? A method of inducing artificial immunity by exposing
Chapter 17: Immunization & Immune Testing 1. Immunization 2. Diagnostic Immunology 1. Immunization Chapter Reading pp. 505-511 What is Immunization? A method of inducing artificial immunity by exposing
reverse transcription! RT 1! RT 2! RT 3!
 Supplementary Figure! Entire workflow repeated 3 times! mirna! stock! -fold dilution series! to yield - copies! per µl in PCR reaction! reverse transcription! RT! pre-pcr! mastermix!!!! 3!!! 3! ddpcr!
Supplementary Figure! Entire workflow repeated 3 times! mirna! stock! -fold dilution series! to yield - copies! per µl in PCR reaction! reverse transcription! RT! pre-pcr! mastermix!!!! 3!!! 3! ddpcr!
Supporting Information
 Supporting Information Yuan et al. 10.1073/pnas.0906869106 Fig. S1. Heat map showing that Populus ICS is coregulated with orthologs of Arabidopsis genes involved in PhQ biosynthesis and PSI function, but
Supporting Information Yuan et al. 10.1073/pnas.0906869106 Fig. S1. Heat map showing that Populus ICS is coregulated with orthologs of Arabidopsis genes involved in PhQ biosynthesis and PSI function, but
RNA-Guided Gene Activation by CRISPR-Cas9-Based Transcription Factors
 Supplementary Information RNA-Guided Gene Activation by CRISPR-Cas9-Based Transcription Factors Pablo Perez-Pinera 1, Daniel D. Kocak 1, Christopher M. Vockley 2,3, Andrew F. Adler 1, Ami M. Kabadi 1,
Supplementary Information RNA-Guided Gene Activation by CRISPR-Cas9-Based Transcription Factors Pablo Perez-Pinera 1, Daniel D. Kocak 1, Christopher M. Vockley 2,3, Andrew F. Adler 1, Ami M. Kabadi 1,
Supplementary Information Design of small molecule-responsive micrornas based on structural requirements for Drosha processing
 Supplementary Information Design of small molecule-responsive micrornas based on structural requirements for Drosha processing Chase L. Beisel, Yvonne Y. Chen, Stephanie J. Culler, Kevin G. Hoff, & Christina
Supplementary Information Design of small molecule-responsive micrornas based on structural requirements for Drosha processing Chase L. Beisel, Yvonne Y. Chen, Stephanie J. Culler, Kevin G. Hoff, & Christina
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION Conserved arginines on the rim of Hfq catalyze base pair formation and exchange Subrata Panja and Sarah A. Woodson T.C. Jenkins Department of Biophysics, Johns Hopkins University,
SUPPLEMENTARY INFORMATION Conserved arginines on the rim of Hfq catalyze base pair formation and exchange Subrata Panja and Sarah A. Woodson T.C. Jenkins Department of Biophysics, Johns Hopkins University,
Supporting Information
 Supporting Information Wiley-VCH 2006 69451 Weinheim, Germany RNA ligands that distinguish metabolite-induced conformations in the TPP riboswitch Günter Mayer, Marie-Sophie L. Raddatz, Julia D. Grunwald,
Supporting Information Wiley-VCH 2006 69451 Weinheim, Germany RNA ligands that distinguish metabolite-induced conformations in the TPP riboswitch Günter Mayer, Marie-Sophie L. Raddatz, Julia D. Grunwald,
Figure S1: NUN preparation yields nascent, unadenylated RNA with a different profile from Total RNA.
 Summary of Supplemental Information Figure S1: NUN preparation yields nascent, unadenylated RNA with a different profile from Total RNA. Figure S2: rrna removal procedure is effective for clearing out
Summary of Supplemental Information Figure S1: NUN preparation yields nascent, unadenylated RNA with a different profile from Total RNA. Figure S2: rrna removal procedure is effective for clearing out
Quantitative Real Time PCR USING SYBR GREEN
 Quantitative Real Time PCR USING SYBR GREEN SYBR Green SYBR Green is a cyanine dye that binds to double stranded DNA. When it is bound to D.S. DNA it has a much greater fluorescence than when bound to
Quantitative Real Time PCR USING SYBR GREEN SYBR Green SYBR Green is a cyanine dye that binds to double stranded DNA. When it is bound to D.S. DNA it has a much greater fluorescence than when bound to
Unit 6: Molecular Genetics & DNA Technology Guided Reading Questions (100 pts total)
 Name: AP Biology Biology, Campbell and Reece, 7th Edition Adapted from chapter reading guides originally created by Lynn Miriello Chapter 16 The Molecular Basis of Inheritance Unit 6: Molecular Genetics
Name: AP Biology Biology, Campbell and Reece, 7th Edition Adapted from chapter reading guides originally created by Lynn Miriello Chapter 16 The Molecular Basis of Inheritance Unit 6: Molecular Genetics
Supplementary information to accompany: A novel role for the DNA repair gene Rad51 in Netrin-1 signalling
 Supplementary information to accompany: A novel role for the DNA repair gene Rad51 in Netrin-1 signalling Glendining KA 1, Markie D 2, Gardner RJM 4, Franz EA 3, Robertson SP 4, Jasoni CL 1 Supplementary
Supplementary information to accompany: A novel role for the DNA repair gene Rad51 in Netrin-1 signalling Glendining KA 1, Markie D 2, Gardner RJM 4, Franz EA 3, Robertson SP 4, Jasoni CL 1 Supplementary
Activation of a Floral Homeotic Gene in Arabidopsis
 Activation of a Floral Homeotic Gene in Arabidopsis By Maximiliam A. Busch, Kirsten Bomblies, and Detlef Weigel Presentation by Lis Garrett and Andrea Stevenson http://ucsdnews.ucsd.edu/archive/graphics/images/image5.jpg
Activation of a Floral Homeotic Gene in Arabidopsis By Maximiliam A. Busch, Kirsten Bomblies, and Detlef Weigel Presentation by Lis Garrett and Andrea Stevenson http://ucsdnews.ucsd.edu/archive/graphics/images/image5.jpg
CHAPTER 9 DNA Technologies
 CHAPTER 9 DNA Technologies Recombinant DNA Artificially created DNA that combines sequences that do not occur together in the nature Basis of much of the modern molecular biology Molecular cloning of genes
CHAPTER 9 DNA Technologies Recombinant DNA Artificially created DNA that combines sequences that do not occur together in the nature Basis of much of the modern molecular biology Molecular cloning of genes
Supplementary Material
 Fig S1. Interactions of OsDof24 with the OsC4PPDK promoter in rice protoplasts (a) Loss-of-function analysis of the OsC4PPDK promoter Effects of OsDof24 overexpression and the mapping of OsDof24-binding
Fig S1. Interactions of OsDof24 with the OsC4PPDK promoter in rice protoplasts (a) Loss-of-function analysis of the OsC4PPDK promoter Effects of OsDof24 overexpression and the mapping of OsDof24-binding
over time using live cell microscopy. The time post infection is indicated in the lower left corner.
 Title of file for HTML: Supplementary Information Description: Supplementary Figures and Supplementary Table Title of file for HTML: Supplementary Movie 1 Description: Fusion of NBs. BSR cells were infected
Title of file for HTML: Supplementary Information Description: Supplementary Figures and Supplementary Table Title of file for HTML: Supplementary Movie 1 Description: Fusion of NBs. BSR cells were infected
Supplemental Figure 1. Mutation in NLA Causes Increased Pi Uptake Activity and
 Supplemental Figure 1. Mutation in NLA Causes Increased Pi Uptake Activity and PHT1 Protein Amounts. (A) Shoot morphology of 19-day-old nla mutants under Pi-sufficient conditions. (B) [ 33 P]Pi uptake
Supplemental Figure 1. Mutation in NLA Causes Increased Pi Uptake Activity and PHT1 Protein Amounts. (A) Shoot morphology of 19-day-old nla mutants under Pi-sufficient conditions. (B) [ 33 P]Pi uptake
Human Granulin / GRN / Progranulin ELISA Pair Set
 Human Granulin / GRN / Progranulin ELISA Pair Set Catalog Number : SEKA10826 To achieve the best assay results, this manual must be read carefully before using this product and the assay is run as summarized
Human Granulin / GRN / Progranulin ELISA Pair Set Catalog Number : SEKA10826 To achieve the best assay results, this manual must be read carefully before using this product and the assay is run as summarized
mmu-mir-200a-3p Real-time RT-PCR Detection and U6 Calibration Kit User Manual MyBioSource.com Catalog # MBS826230
 mmu-mir-200a-3p Real-time RT-PCR Detection and U6 Calibration Kit User Manual Catalog # MBS826230 For the detection and quantification of mirnas mmu-mir-200a-3p normalized by U6 snrna using Real-time RT-PCR
mmu-mir-200a-3p Real-time RT-PCR Detection and U6 Calibration Kit User Manual Catalog # MBS826230 For the detection and quantification of mirnas mmu-mir-200a-3p normalized by U6 snrna using Real-time RT-PCR
SUPPLEMENTARY INFORMATION
 b Supplementary Figure 1. Analysis of Vkg-Dpp interactions. a, In vitro pulldown showing binding of Dpp-HA to the GST-VkgC fusion protein. No interaction is detected between denatured Dpp-HA and GST-Vkg.
b Supplementary Figure 1. Analysis of Vkg-Dpp interactions. a, In vitro pulldown showing binding of Dpp-HA to the GST-VkgC fusion protein. No interaction is detected between denatured Dpp-HA and GST-Vkg.
Supporting Online Material for
 www.sciencemag.org/cgi/content/full/1137999/dc1 Supporting Online Material for Disrupting the Pairing Between let-7 and Enhances Oncogenic Transformation Christine Mayr, Michael T. Hemann, David P. Bartel*
www.sciencemag.org/cgi/content/full/1137999/dc1 Supporting Online Material for Disrupting the Pairing Between let-7 and Enhances Oncogenic Transformation Christine Mayr, Michael T. Hemann, David P. Bartel*
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:10.1038/nature09937 a Name Position Primersets 1a 1b 2 3 4 b2 Phenotype Genotype b Primerset 1a D T C R I E 10000 8000 6000 5000 4000 3000 2500 2000 1500 1000 800 Donor (D)
SUPPLEMENTARY INFORMATION doi:10.1038/nature09937 a Name Position Primersets 1a 1b 2 3 4 b2 Phenotype Genotype b Primerset 1a D T C R I E 10000 8000 6000 5000 4000 3000 2500 2000 1500 1000 800 Donor (D)
SUPPLEMENTARY MATERIAL
 SUPPLEMENTARY MATERIAL Materials and Methods Circular dichroism (CD) spectroscopy. Far ultraviolet (UV) CD spectra of apo- and holo- CaM and the CaM mutants were recorded on a Jasco J-715 spectropolarimeter
SUPPLEMENTARY MATERIAL Materials and Methods Circular dichroism (CD) spectroscopy. Far ultraviolet (UV) CD spectra of apo- and holo- CaM and the CaM mutants were recorded on a Jasco J-715 spectropolarimeter
Quantitative analysis of recombination in YFP and CFP gene of FRET biosensor induced by lentiviral or retroviral gene transfer.
 Supplementary Information: Quantitative analysis of recombination in and gene of FRET biosensor induced by lentiviral or retroviral gene transfer. Akira T. Komatsubara, Michiyuki Matsuda,, and Kazuhiro
Supplementary Information: Quantitative analysis of recombination in and gene of FRET biosensor induced by lentiviral or retroviral gene transfer. Akira T. Komatsubara, Michiyuki Matsuda,, and Kazuhiro
Supplemental Information. Natural RNA Polymerase Aptamers. Regulate Transcription in E. coli
 Molecular Cell, Volume 67 Supplemental Information Natural RNA Polymerase Aptamers Regulate Transcription in E. coli Nadezda Sedlyarova, Philipp Rescheneder, Andrés Magán, Niko Popitsch, Natascha Rziha,
Molecular Cell, Volume 67 Supplemental Information Natural RNA Polymerase Aptamers Regulate Transcription in E. coli Nadezda Sedlyarova, Philipp Rescheneder, Andrés Magán, Niko Popitsch, Natascha Rziha,
Online Supplementary Information
 Online Supplementary Information NLRP4 negatively regulates type I interferon signaling by targeting TBK1 for degradation via E3 ubiquitin ligase DTX4 Jun Cui 1,4,6,7, Yinyin Li 1,5,6,7, Liang Zhu 1, Dan
Online Supplementary Information NLRP4 negatively regulates type I interferon signaling by targeting TBK1 for degradation via E3 ubiquitin ligase DTX4 Jun Cui 1,4,6,7, Yinyin Li 1,5,6,7, Liang Zhu 1, Dan
