T H E J O U R N A L O F C E L L B I O L O G Y
|
|
|
- Eugenia Reeves
- 5 years ago
- Views:
Transcription
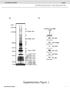
1 T H E J O U R N A L O F C E L L B I O L O G Y Supplemental material Kawai and Amano, Figure S1. Regulation of mirna expression by BRCA1. (A) Confirmation of BRCA1 overexpression. The expression level of BRCA1 was examined by qrt- PCR analysis in BRCA1-transfected or adenovirus-infected HeLa cells. (B) Expression of pri-, pre-, and mature (mat)-mirna by BRCA1 transfection in HeLa cells. (C) Expression of pri- and pre-mirna by BRCA1 transfection in HEK293 cells. (D) Expression of pri- and pre-mirna by BRCA1 transfection in MG63 cells. (E) Confirmation of BRCA1 expression after sibrca1 knockdown and in BRCA1 knockout ES cells (KO). (F) Expression of pri-, pre-, and mat-mirna by sibrca1 knockdown in HeLa cells. (G) Expression of cancer up-regulated mirna by BRCA1 overexpression in mouse BRCA1-transfected HeLa cells examined by qrt-pcr analysis. (H) Expression of non cancer-regulated mirna by BRCA1 overexpression. Error bars represent standard deviation. The broken horizontal lines represent the value of 1 used as a standard/criterion. S1
2 Figure S2. Control of mirna by BRCA1. (A) System used for in vivo monitoring of pri-mirna processing. (B) Validation of system used for in vivo monitoring of pri-mirna processing. (C) The association of BRCA1 with DROSHA was increased in a dose-dependent manner. Numbers indicate the amount of transfected DNA (in micrograms). (D) Endogenous associations of BRCA1, DROSHA, and DDX. (E) RIP analysis of BRCA1 knockdown cells. (F) RIP analysis of endogenous expression levels. Error bars represent standard deviation. The broken horizontal and vertical lines represent the value of 1 used as a standard/criterion. S2
3 Figure S3. Modulation of mirna by BRCA1 and DHX9. (A) Mouse Brca1 Ring (1 448), DBD (449 1,048), SQ (1049 1,493), and BRCT (1,494 1,811) domains were fused into the GST expression vector pgex6p1. (B) Expression of GST fusion Brca1 proteins in E. coli BL21. Arrows indicate bands of the expected proteins. (C) Human DDX5 (p68, 614 aa), DDX17 (p72, 729 aa), and DHX9 (1270 aa) were aligned, and the region around DEAD/DEIH is shown (bold). Identical amino acids are marked with asterisks, and similar amino acids with colons or periods. (D and E) Relative mrna expression of overexpressed DHX9 and sidhx9 knockdown. sidhx9 1, 9 2, and 9 3 are indicated in three sets of stealth RNAi. (F) Expression of pri-, pre-, and mat-mirna by sidhx9 knockdown. Error bars represent standard deviation. The broken horizontal lines represent the value of 1 used as a standard/criterion. S3
4 Table S1. The oligonucleotide sequences Criteria Gene Orientation Primer sequence (5-3 ) Source Cloning mp53 a Forward CACCATGACTGCCATGGAGGAGTCACAG NM_ TCATCAGTCTGAGTCAGGCCCCACTTT msmad3 a Forward CACCATGTCGTCCATCCTGCCCTTCACC NM_ CTACTAAGACACACTGGAACAGCGGAT GST-Brca1 RING Forward CGGTCGACTCATGGATTTATCTGCCGTCCAAATT NM_ GCGTCGACTTATGGTTTGGAGAAGTCTCTTCCACG GST-Brca1 DBD Forward GCGAATTCGTAGAGGATAATATCAGTGATAAA NM_ GCGTCGACTTACTTAGGCCCTCTGTTTCTACCTAG GST-Brca1 SQ Forward GCGAATTCGTGAACACTGTGCCTCCATTAGAT NM_ GCGTCGACTTACGCCTCTGATCCAGCAGGCTGGAG GST-Brca1 BRCT Forward GCGAATTCTCATCTGAGCCACACAATTCAACA NM_ GCGCGGCCGCTTAATCATTGGAGTCTTGTGGCTC sirna duplex sibrca1 Forward GGAACCUGUCUCCACAAAGdTdT Wilson and Stern, 2008 CUUUGUGGAGACAGGUUCCdTdT sidhx9-1 Forward AACAACCAAAAGCCUCCAUGGCUGCC This paper GGCAGCCAUGGAGGCUUUGGUUGUU sidhx9-2 Forward AUAAGGUUGAUACCAGCAGCUGAGG This paper CCUCAGCUGCUGGUAUCAACCUUAU sidhx9-3 Forward AAUAGCCGCCACCUCCUCUUCCCUG This paper CAGGGAAGAGGAGGUGGCGGCUAUU qrt-pcr (human) pri-let-7a-1 Forward CCTGGATGTTCTCTTCACTG Jiang et al., 2005 GCCTGGATGCAGACTTTTCT pre-let-7a-1 Forward AGGTAGTAGGTTGTATAGTTTTAGG Jiang et al., 2005 pri-mir-16-1 Forward GCAATTACAGTATTTTAAGAGATGAT Suzuki et al., 2009 CATACTCTACAGTTGTGTTTTAATGT pre-mir-16-1 Forward GCAGCACGTAAATATTGGCGT Jiang et al., 2005 CAGCAGCACAGTTAATACTGGAGA pri-mir-145 Forward TGGATTTGCCTCCTTCCCA Suzuki et al., 2009 TTGAACCCTCATCCTGTGAGCC pre-mir-145 Forward CTTGTCCTCACGGTCCAGTT This paper pri-mir-34a Forward CCTCCAAGCCAGCTCAGTTG Suzuki et al., 2009 TGACTTTGGTCCAATTCCTGTTG pre-mir-34a Forward TGGCAGTGTCTTAGCTGGTTG Jiang et al., 2005 hgapdh b Forward GTGGTCTCCTCTGACTTCAACAG NM_ CTGTAGCCAAATTCGTTGTCATAC hrnu6-1 b Forward CTCGCTTCGGCAGCACA NR_ AACGCTTCACGAATTTGCGT mbrca1 a Forward TCAAAGGACTCCTGACAACATAAA NM_ CTTTCTCAATGATTCAGTTGGATG hdhx9 b Forward TTGATCCAGTACCAGTTGGAGTAA NM_ CGTTTATGGTAATGCTTGTTTCAG pspt18 let-7a-1 Forward GCGCTAGCTCACACAGGAAACCAGGATTACCG NR_ GCGTCGACGCTGCACTACATCTCTTTAAGACA mir-16-1 Forward GCGCTAGCTGAAAAGGTGCAGGCCATATTGTG NR_ GCGTCGACTAAAAATAACAAGATTATCAATAA mir-145 Forward GCGCTAGCGCACCCCACCCTGGCTGCTACAGA NR_ GCGTCGACCTCCAGGGACAGCCTTCTTCTTGA mir-34a Forward GCGCTAGCAAGCCTGCACTACCTGCTCGCCCC NR_ GCGTCGACCTTTCCCAGCCGCCCTCACCACCC pmirglo let-7a-1 Forward GCGAATTCCTTTTCACCATTCACCCTGGATGT GCGTCGACATCAGACCGCCTGGATGCAGACTT mir-16-1 Forward GCGAATTCTCTTTTTATTCATAGCTCTTATGA GCGTCGACATATACATTAAAACACAACTGTAG mir-145 Forward GCGAATTCGGCGGCCTTGGCGCTGAAGGCCAC GCGTCGACTGGGGTGGGAAGGAGGCAAATCCA S4
5 Table S1. The oligonucleotide sequences (Continued) Criteria Gene Orientation Primer sequence (5-3 ) Source mir-34a Forward GCGAATTCCCCTCCGGATGCCGTGGACCGGCC GCGTCGACCTCGCTTCATCTTCCCTCTTGGGC RIP pri-let-7a-1 Forward CCTGGATGTTCTCTTCACTG GCCTGGATGCAGACTTTTCT pri-mir-16-1 Forward GCAATTACAGTATTTTAAGAGATGAT CATACTCTACAGTTGTGTTTTAATGT pri-mir-145 Forward TGGATTTGCCTCCTTCCCA TTGAACCCTCATCCTGTGAGCC pri-mir-34a Forward CCTCCAAGCCAGCTCAGTTG TGACTTTGGTCCAATTCCTGTTG Pull-down assay pre-let-7a-1 Forward AGGTAGTAGGTTGTATAGTTTTAGG pre-mir-16-1 Forward GCAGCACGTAAATATTGGCGT CAGCAGCACAGTTAATACTGGAGA pre-mir-145 Forward CTTGTCCTCACGGTCCAGTT pre-mir-34a Forward TGGCAGTGTCTTAGCTGGTTG EMSA c branch RNA Forward AUAGCAAUGUCAGCAGUGCCUUAGC (mir-16 based) FAM-GCUAAGGCACUGCUGACCAAAGACA linear RNA Forward UAGCAGCACGUAAAUAUUGG (mir-16 based) FAM-CCAAUAUUUACGUGCUGCUA qrt-pcr (mouse) pri-let-7a-1 Forward CCTGGATGTTCTCTTCACTG This paper CCATGAATGCAGACTTTTCC pre-let-7a-1 Forward AGGTAGTAGGTTGTATAGTTTTAGG Jiang et al., 2005 pri-mir-16-1 Forward TGGTAATGCAGCTCAGTTGG This paper GACTTGCTTTATGGCCAAGG pre-mir-16-1 Forward GCAGCACGTAAATATTGGCGT This paper CAGCAGCACAGTCAATACTGGAGG pri-mir-145 Forward TTGGGAGCTCTCTTCTGACC This paper CTCCCTCCTGGGGATGTTAG pre-mir-145 Forward CTTGTCCTCACGGTCCAGTT This paper pri-mir-34a Forward AAGGTCAGAGGTCAGCAACG This paper CCAGAAGTGCTCACACTCCA pre-mir-34a Forward TGGCAGTGTCTTAGCTGGTTG Jiang et al., 2005 Underlines show matched complementary sequence. These oligonucleotides were partially annealed and used for the experiment. a mp53, msmad3, and mbrca1 represent mouse homologue of p53, Smad3, and Brca1. b hgapdh, hrnu6-1, and hdhx9 represent human homologue of GAPDH, RNU6-1, and DHX9. c These oligonucleotides were synthesized based on mir-16. References Jiang, J., E.J. Lee, Y. Gusev, and T.D. Schmittgen Real-time expression profiling of microrna precursors in human cancer cell lines. Nucleic Acids Res. 33: Suzuki, H.I., K. Yamagata, K. Sugimoto, T. Iwamoto, S. Kato, and K. Miyazono Modulation of microrna processing by p53. Nature. 460: dx.doi.org/ /nature08199 Wilson, K.A., and D.F. Stern NFBD1/MDC1, 53BP1 and BRCA1 have both redundant and unique roles in the ATM pathway. Cell Cycle. 7: dx.doi.org/ /cc S5
SUPPLEMENTARY INFORMATION
 doi:.38/nature899 Supplementary Figure Suzuki et al. a c p7 -/- / WT ratio (+)/(-) p7 -/- / WT ratio Log X 3. Fold change by treatment ( (+)/(-)) Log X.5 3-3. -. b Fold change by treatment ( (+)/(-)) 8
doi:.38/nature899 Supplementary Figure Suzuki et al. a c p7 -/- / WT ratio (+)/(-) p7 -/- / WT ratio Log X 3. Fold change by treatment ( (+)/(-)) Log X.5 3-3. -. b Fold change by treatment ( (+)/(-)) 8
Supplementary Figure 1, related to Figure 1. GAS5 is highly expressed in the cytoplasm of hescs, and positively correlates with pluripotency.
 Supplementary Figure 1, related to Figure 1. GAS5 is highly expressed in the cytoplasm of hescs, and positively correlates with pluripotency. (a) Transfection of different concentration of GAS5-overexpressing
Supplementary Figure 1, related to Figure 1. GAS5 is highly expressed in the cytoplasm of hescs, and positively correlates with pluripotency. (a) Transfection of different concentration of GAS5-overexpressing
Cell proliferation was measured with Cell Counting Kit-8 (Dojindo Laboratories, Kumamoto, Japan).
 1 2 3 4 5 6 7 8 Supplemental Materials and Methods Cell proliferation assay Cell proliferation was measured with Cell Counting Kit-8 (Dojindo Laboratories, Kumamoto, Japan). GCs were plated at 96-well
1 2 3 4 5 6 7 8 Supplemental Materials and Methods Cell proliferation assay Cell proliferation was measured with Cell Counting Kit-8 (Dojindo Laboratories, Kumamoto, Japan). GCs were plated at 96-well
Nature Structural and Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Distribution of mirnas between lncrna and protein-coding genes. Pie chart showing distribution of human mirna between protein coding and lncrna genes. To the right, lncrna mirna
Supplementary Figure 1 Distribution of mirnas between lncrna and protein-coding genes. Pie chart showing distribution of human mirna between protein coding and lncrna genes. To the right, lncrna mirna
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature12119 SUPPLEMENTARY FIGURES AND LEGENDS pre-let-7a- 1 +14U pre-let-7a- 1 Ddx3x Dhx30 Dis3l2 Elavl1 Ggt5 Hnrnph 2 Osbpl5 Puf60 Rnpc3 Rpl7 Sf3b3 Sf3b4 Tia1 Triobp U2af1 U2af2 1 6 2 4 3
doi:10.1038/nature12119 SUPPLEMENTARY FIGURES AND LEGENDS pre-let-7a- 1 +14U pre-let-7a- 1 Ddx3x Dhx30 Dis3l2 Elavl1 Ggt5 Hnrnph 2 Osbpl5 Puf60 Rnpc3 Rpl7 Sf3b3 Sf3b4 Tia1 Triobp U2af1 U2af2 1 6 2 4 3
MeCP2. MeCP2/α-tubulin. GFP mir1-1 mir132
 Conservation Figure S1. Schematic showing 3 UTR (top; thick black line), mir132 MRE (arrow) and nucleotide sequence conservation (vertical black lines; http://genome.ucsc.edu). a GFP mir1-1 mir132 b GFP
Conservation Figure S1. Schematic showing 3 UTR (top; thick black line), mir132 MRE (arrow) and nucleotide sequence conservation (vertical black lines; http://genome.ucsc.edu). a GFP mir1-1 mir132 b GFP
Total RNA was isolated with Trizol reagent (Invitrogen Corp.) according to the
 Supplemental Materials and Methods Isolation and analysis of RNA Total RNA was isolated with Trizol reagent (Invitrogen Corp.) according to the manufacturer s instructions. For isolation of nuclear and
Supplemental Materials and Methods Isolation and analysis of RNA Total RNA was isolated with Trizol reagent (Invitrogen Corp.) according to the manufacturer s instructions. For isolation of nuclear and
Supplemental Table/Figure Legends
 MiR-26a is required for skeletal muscle differentiation and regeneration in mice Bijan K. Dey, Jeffrey Gagan, Zhen Yan #, Anindya Dutta Supplemental Table/Figure Legends Suppl. Table 1: Effect of overexpression
MiR-26a is required for skeletal muscle differentiation and regeneration in mice Bijan K. Dey, Jeffrey Gagan, Zhen Yan #, Anindya Dutta Supplemental Table/Figure Legends Suppl. Table 1: Effect of overexpression
Supporting Online Material for
 www.sciencemag.org/cgi/content/full/1154040/dc1 Supporting Online Material for Selective Blockade of MicroRNA Processing by Lin-28 Srinivas R. Viswanathan, George Q. Daley,* Richard I. Gregory* *To whom
www.sciencemag.org/cgi/content/full/1154040/dc1 Supporting Online Material for Selective Blockade of MicroRNA Processing by Lin-28 Srinivas R. Viswanathan, George Q. Daley,* Richard I. Gregory* *To whom
Thermo Scientific Dharmacon SMARTvector 2.0 Lentiviral shrna Particles
 Thermo Scientific Dharmacon SMARTvector 2.0 Lentiviral shrna Particles Long-term gene silencing shrna-specific design algorithm High titer, purified particles Thermo Scientific Dharmacon SMARTvector shrna
Thermo Scientific Dharmacon SMARTvector 2.0 Lentiviral shrna Particles Long-term gene silencing shrna-specific design algorithm High titer, purified particles Thermo Scientific Dharmacon SMARTvector shrna
Investigating the regulation of mirna biogenesis and Argonaute2 by RNA binding proteins
 Investigating the regulation of mirna biogenesis and Argonaute2 by RNA binding proteins Patrick Peter Connerty Supervisor: Gyorgy Hutvagner Thesis Submitted for the Degree of Doctor of Philosophy (Science)
Investigating the regulation of mirna biogenesis and Argonaute2 by RNA binding proteins Patrick Peter Connerty Supervisor: Gyorgy Hutvagner Thesis Submitted for the Degree of Doctor of Philosophy (Science)
SUPPLEMENTARY INFORMATION
 (Supplementary Methods and Materials) GST pull-down assay GST-fusion proteins Fe65 365-533, and Fe65 538-700 were expressed in BL21 bacterial cells and purified with glutathione-agarose beads (Sigma).
(Supplementary Methods and Materials) GST pull-down assay GST-fusion proteins Fe65 365-533, and Fe65 538-700 were expressed in BL21 bacterial cells and purified with glutathione-agarose beads (Sigma).
Comparative Analysis of Argonaute-Dependent Small RNA Pathways in Drosophila
 Molecular Cell, Volume 32 Supplemental Data Comparative Analysis of Argonaute-Dependent Small RNA Pathways in Drosophila Rui Zhou, Ikuko Hotta, Ahmet M. Denli, Pengyu Hong, Norbert Perrimon, and Gregory
Molecular Cell, Volume 32 Supplemental Data Comparative Analysis of Argonaute-Dependent Small RNA Pathways in Drosophila Rui Zhou, Ikuko Hotta, Ahmet M. Denli, Pengyu Hong, Norbert Perrimon, and Gregory
Supplementary
 Supplementary information Supplementary Material and Methods Plasmid construction The transposable element vectors for inducible expression of RFP-FUS wt and EGFP-FUS R521C and EGFP-FUS P525L were derived
Supplementary information Supplementary Material and Methods Plasmid construction The transposable element vectors for inducible expression of RFP-FUS wt and EGFP-FUS R521C and EGFP-FUS P525L were derived
SUPPLEMENTARY INFORMATION
 Supplementary Figure a T m ( C) Seq. '-3' uguuugugguaacagugugaggu L 62 AttGtcAcaCtcC L2 6 ccattgtcacactcc L3 66 attgtcacactcc 7 ccattgtcacactcca L 73 ccattgtcacactcc L6 74 AttGTcaCaCtCC L7 7 attgtcacactcc
Supplementary Figure a T m ( C) Seq. '-3' uguuugugguaacagugugaggu L 62 AttGtcAcaCtcC L2 6 ccattgtcacactcc L3 66 attgtcacactcc 7 ccattgtcacactcca L 73 ccattgtcacactcc L6 74 AttGTcaCaCtCC L7 7 attgtcacactcc
TOOLS sirna and mirna. User guide
 TOOLS sirna and mirna User guide Introduction RNA interference (RNAi) is a powerful tool for suppression gene expression by causing the destruction of specific mrna molecules. Small Interfering RNAs (sirnas)
TOOLS sirna and mirna User guide Introduction RNA interference (RNAi) is a powerful tool for suppression gene expression by causing the destruction of specific mrna molecules. Small Interfering RNAs (sirnas)
Table 1. Primers, annealing temperatures, and product sizes for PCR amplification.
 Table 1. Primers, annealing temperatures, and product sizes for PCR amplification. Gene Direction Primer sequence (5 3 ) Annealing Temperature Size (bp) BRCA1 Forward TTGCGGGAGGAAAATGGGTAGTTA 50 o C 292
Table 1. Primers, annealing temperatures, and product sizes for PCR amplification. Gene Direction Primer sequence (5 3 ) Annealing Temperature Size (bp) BRCA1 Forward TTGCGGGAGGAAAATGGGTAGTTA 50 o C 292
Supplementary Materials
 Supplementary Materials Table S1. Oligonucleotide sequences and PCR conditions used to amplify the indicated genes. TA = annealing temperature; gdna = genomic DNA; cdna = complementary DNA; c = concentration.
Supplementary Materials Table S1. Oligonucleotide sequences and PCR conditions used to amplify the indicated genes. TA = annealing temperature; gdna = genomic DNA; cdna = complementary DNA; c = concentration.
MISSION shrna Library: Next Generation RNA Interference
 Page 1 of 6 Page 1 of 6 Return to Web Version MISSION shrna Library: Next Generation RNA Interference By: Stephanie Uder, Henry George, Betsy Boedeker, LSI Volume 6 Article 2 Introduction The technology
Page 1 of 6 Page 1 of 6 Return to Web Version MISSION shrna Library: Next Generation RNA Interference By: Stephanie Uder, Henry George, Betsy Boedeker, LSI Volume 6 Article 2 Introduction The technology
Technology Overview. Figure 1. asirna structure
 BMT, Inc. Technology Overview Small interfering RNAs (sirnas) are short, double-stranded RNAs (dsrnas) that mediate efficient gene silencing in a sequence-specific manner. The specific cleavage of mrna
BMT, Inc. Technology Overview Small interfering RNAs (sirnas) are short, double-stranded RNAs (dsrnas) that mediate efficient gene silencing in a sequence-specific manner. The specific cleavage of mrna
sherwood - UltramiR shrna Collections
 sherwood - UltramiR shrna Collections Incorporating advances in shrna design and processing for superior potency and specificity sherwood - UltramiR shrna Collections Enabling Discovery Across the Genome
sherwood - UltramiR shrna Collections Incorporating advances in shrna design and processing for superior potency and specificity sherwood - UltramiR shrna Collections Enabling Discovery Across the Genome
Genomic DNA fragments identified by ChIP-chip assay for MR [28] corresponding to two
![Genomic DNA fragments identified by ChIP-chip assay for MR [28] corresponding to two Genomic DNA fragments identified by ChIP-chip assay for MR [28] corresponding to two](/thumbs/89/98549522.jpg) Supplemental Materials and Methods Genomic DNA fragments identified by ChIP-chip assay for MR [28] corresponding to two upstream regions of the human Klf9 gene (-5139 to -5771 bp and -3875 to -4211 bp)
Supplemental Materials and Methods Genomic DNA fragments identified by ChIP-chip assay for MR [28] corresponding to two upstream regions of the human Klf9 gene (-5139 to -5771 bp and -3875 to -4211 bp)
Regulation of Synaptic Structure and Function by FMRP- Associated MicroRNAs mir-125b and mir-132
 Neuron, Volume 65 Regulation of Synaptic Structure and Function by FMRP- Associated MicroRNAs mir-125b and mir-132 Dieter Edbauer, Joel R. Neilson, Kelly A. Foster, Chi-Fong Wang, Daniel P. Seeburg, Matthew
Neuron, Volume 65 Regulation of Synaptic Structure and Function by FMRP- Associated MicroRNAs mir-125b and mir-132 Dieter Edbauer, Joel R. Neilson, Kelly A. Foster, Chi-Fong Wang, Daniel P. Seeburg, Matthew
HCT116 SW48 Nutlin: p53
 Figure S HCT6 SW8 Nutlin: - + - + p GAPDH Figure S. Nutlin- treatment induces p protein. HCT6 and SW8 cells were left untreated or treated for 8 hr with Nutlin- ( µm) to up-regulate p. Whole cell lysates
Figure S HCT6 SW8 Nutlin: - + - + p GAPDH Figure S. Nutlin- treatment induces p protein. HCT6 and SW8 cells were left untreated or treated for 8 hr with Nutlin- ( µm) to up-regulate p. Whole cell lysates
Regulation of transcription by the MLL2 complex and MLL complex-associated AKAP95
 Supplementary Information Regulation of transcription by the complex and MLL complex-associated Hao Jiang, Xiangdong Lu, Miho Shimada, Yali Dou, Zhanyun Tang, and Robert G. Roeder Input HeLa NE IP lot:
Supplementary Information Regulation of transcription by the complex and MLL complex-associated Hao Jiang, Xiangdong Lu, Miho Shimada, Yali Dou, Zhanyun Tang, and Robert G. Roeder Input HeLa NE IP lot:
Document S1. Supplemental Experimental Procedures and Three Figures (see next page)
 Supplemental Data Document S1. Supplemental Experimental Procedures and Three Figures (see next page) Table S1. List of Candidate Genes Identified from the Screen. Candidate genes, corresponding dsrnas
Supplemental Data Document S1. Supplemental Experimental Procedures and Three Figures (see next page) Table S1. List of Candidate Genes Identified from the Screen. Candidate genes, corresponding dsrnas
Supporting Information
 Supporting Information SI Materials and Methods RT-qPCR The 25 µl qrt-pcr reaction mixture included 1 µl of cdna or DNA, 12.5 µl of 2X SYBER Green Master Mix (Applied Biosystems ), 5 µm of primers and
Supporting Information SI Materials and Methods RT-qPCR The 25 µl qrt-pcr reaction mixture included 1 µl of cdna or DNA, 12.5 µl of 2X SYBER Green Master Mix (Applied Biosystems ), 5 µm of primers and
supplementary information
 Figure S1 ZEB1 full length mrna. (a) Analysis of the ZEB1 mrna using the UCSC genome browser (http://genome.ucsc.edu) revealed truncation of the annotated Refseq sequence (NM_030751). The probable terminus
Figure S1 ZEB1 full length mrna. (a) Analysis of the ZEB1 mrna using the UCSC genome browser (http://genome.ucsc.edu) revealed truncation of the annotated Refseq sequence (NM_030751). The probable terminus
1. Primers for PCR to amplify hairpin stem-loop precursor mir-145 plus different flanking sequence from human genomic DNA.
 Supplemental data: 1. Primers for PCR to amplify hairpin stem-loop precursor mir-145 plus different flanking sequence from human genomic DNA. Strategy#1: 20nt at both sides: #1_BglII-Fd primer : 5 -gga
Supplemental data: 1. Primers for PCR to amplify hairpin stem-loop precursor mir-145 plus different flanking sequence from human genomic DNA. Strategy#1: 20nt at both sides: #1_BglII-Fd primer : 5 -gga
Supplementary Figure 1. NORAD expression in mouse (A) and dog (B). The black boxes indicate the position of the regions alignable to the 12 repeat
 Supplementary Figure 1. NORAD expression in mouse (A) and dog (B). The black boxes indicate the position of the regions alignable to the 12 repeat units in the human genome. Annotated transposable elements
Supplementary Figure 1. NORAD expression in mouse (A) and dog (B). The black boxes indicate the position of the regions alignable to the 12 repeat units in the human genome. Annotated transposable elements
Supplemental Information. Pacer Mediates the Function of Class III PI3K. and HOPS Complexes in Autophagosome. Maturation by Engaging Stx17
 Molecular Cell, Volume 65 Supplemental Information Pacer Mediates the Function of Class III PI3K and HOPS Complexes in Autophagosome Maturation by Engaging Stx17 Xiawei Cheng, Xiuling Ma, Xianming Ding,
Molecular Cell, Volume 65 Supplemental Information Pacer Mediates the Function of Class III PI3K and HOPS Complexes in Autophagosome Maturation by Engaging Stx17 Xiawei Cheng, Xiuling Ma, Xianming Ding,
mirnaselect pegp-mir Cloning and Expression Vector
 Product Data Sheet mirnaselect pegp-mir Cloning and Expression Vector CATALOG NUMBER: MIR-EXP-GP-C STORAGE: -80ºC QUANTITY: 100 µl of bacterial glycerol stock Components 1. mirnaselect pegp-mir Cloning
Product Data Sheet mirnaselect pegp-mir Cloning and Expression Vector CATALOG NUMBER: MIR-EXP-GP-C STORAGE: -80ºC QUANTITY: 100 µl of bacterial glycerol stock Components 1. mirnaselect pegp-mir Cloning
Supplementary data. sienigma. F-Enigma F-EnigmaSM. a-p53
 Supplementary data Supplemental Figure 1 A sienigma #2 sienigma sicontrol a-enigma - + ++ - - - - - - + ++ - - - - - - ++ B sienigma F-Enigma F-EnigmaSM a-flag HLK3 cells - - - + ++ + ++ - + - + + - -
Supplementary data Supplemental Figure 1 A sienigma #2 sienigma sicontrol a-enigma - + ++ - - - - - - + ++ - - - - - - ++ B sienigma F-Enigma F-EnigmaSM a-flag HLK3 cells - - - + ++ + ++ - + - + + - -
Supplementary Figures
 Supplementary Figures Supplementary Figure 1 Experimental schema for the identification of circular RNAs in six normal tissues and seven cancerous tissues. Supplementary Fiure 2 Comparison of human circrnas
Supplementary Figures Supplementary Figure 1 Experimental schema for the identification of circular RNAs in six normal tissues and seven cancerous tissues. Supplementary Fiure 2 Comparison of human circrnas
Supplemental Data. Cui et al. (2012). Plant Cell /tpc a b c d. Stem UBC32 ACTIN
 A Root Stem Leaf Flower Silique Senescence leaf B a b c d UBC32 ACTIN C * Supplemental Figure 1. Expression Pattern and Protein Sequence of UBC32 Homologues in Yeast, Human, and Arabidopsis. (A) Expression
A Root Stem Leaf Flower Silique Senescence leaf B a b c d UBC32 ACTIN C * Supplemental Figure 1. Expression Pattern and Protein Sequence of UBC32 Homologues in Yeast, Human, and Arabidopsis. (A) Expression
Supplemental Data. Na Xu et al. (2016). Plant Cell /tpc
 Supplemental Figure 1. The weak fluorescence phenotype is not caused by the mutation in At3g60240. (A) A mutation mapped to the gene At3g60240. Map-based cloning strategy was used to map the mutated site
Supplemental Figure 1. The weak fluorescence phenotype is not caused by the mutation in At3g60240. (A) A mutation mapped to the gene At3g60240. Map-based cloning strategy was used to map the mutated site
Coleman et al., Supplementary Figure 1
 Coleman et al., Supplementary Figure 1 BrdU Merge G1 Early S Mid S Supplementary Figure 1. Sequential destruction of CRL4 Cdt2 targets during the G1/S transition. HCT116 cells were synchronized by sequential
Coleman et al., Supplementary Figure 1 BrdU Merge G1 Early S Mid S Supplementary Figure 1. Sequential destruction of CRL4 Cdt2 targets during the G1/S transition. HCT116 cells were synchronized by sequential
To generate the luciferase fusion to the human 3 UTRs, we sub-cloned the 3 UTR
 Plasmids To generate the luciferase fusion to the human 3 UTRs, we sub-cloned the 3 UTR fragments downstream of firefly luciferase (luc) in pgl3 control (Promega). pgl3- CDK6 was made by amplifying a 2,886
Plasmids To generate the luciferase fusion to the human 3 UTRs, we sub-cloned the 3 UTR fragments downstream of firefly luciferase (luc) in pgl3 control (Promega). pgl3- CDK6 was made by amplifying a 2,886
Supplemental Figure 1 HDA18 has an HDAC domain and therefore has concentration dependent and TSA inhibited histone deacetylase activity.
 Supplemental Figure 1 HDA18 has an HDAC domain and therefore has concentration dependent and TSA inhibited histone deacetylase activity. (A) Amino acid alignment of HDA5, HDA15 and HDA18. The blue line
Supplemental Figure 1 HDA18 has an HDAC domain and therefore has concentration dependent and TSA inhibited histone deacetylase activity. (A) Amino acid alignment of HDA5, HDA15 and HDA18. The blue line
FCN1 (M-ficolin), which directly associates with immunoglobulin G1, is a molecular target of intravenous immunoglobulin therapy for Kawasaki disease
 FCN1 (M-ficolin), which directly associates with immunoglobulin G1, is a molecular target of intravenous immunoglobulin therapy for Kawasaki disease Daisuke Okuzaki, Kaori Ota, Shin-ichi Takatsuki, Yukari
FCN1 (M-ficolin), which directly associates with immunoglobulin G1, is a molecular target of intravenous immunoglobulin therapy for Kawasaki disease Daisuke Okuzaki, Kaori Ota, Shin-ichi Takatsuki, Yukari
Suppl. Table S1. Characteristics of DHS regions analyzed by bisulfite sequencing. No. CpGs analyzed in the amplicon. Genomic location specificity
 Suppl. Table S1. Characteristics of DHS regions analyzed by bisulfite sequencing. DHS/GRE Genomic location Tissue specificity DHS type CpG density (per 100 bp) No. CpGs analyzed in the amplicon CpG within
Suppl. Table S1. Characteristics of DHS regions analyzed by bisulfite sequencing. DHS/GRE Genomic location Tissue specificity DHS type CpG density (per 100 bp) No. CpGs analyzed in the amplicon CpG within
Fig. S1. Effect of p120-catenin overexpression on the interaction of SCUBE2 with E-cadherin. The expression plasmid encoding FLAG.
 Fig. S1. Effect of p120-catenin overexpression on the interaction of SCUBE2 with E-cadherin. The expression plasmid encoding FLAG.SCUBE2, E-cadherin.Myc, or HA.p120-catenin was transfected in a combination
Fig. S1. Effect of p120-catenin overexpression on the interaction of SCUBE2 with E-cadherin. The expression plasmid encoding FLAG.SCUBE2, E-cadherin.Myc, or HA.p120-catenin was transfected in a combination
Supplementary Figure 1 Characterization of sirna-onv stability. (a) Fluorescence recovery curves of SQ-siRNA-ONV and SQ-ds-siRNA in 1 TAMg buffer
 Supplementary Figure 1 Characterization of sirna-onv stability. (a) Fluorescence recovery curves of SQ-siRNA-ONV and SQ-ds-siRNA in 1 TAMg buffer containing 10% serum The data error bars indicate means
Supplementary Figure 1 Characterization of sirna-onv stability. (a) Fluorescence recovery curves of SQ-siRNA-ONV and SQ-ds-siRNA in 1 TAMg buffer containing 10% serum The data error bars indicate means
Alteration of microrna expression correlates to fatty acidmediated. insulin resistance in mouse myoblasts
 Alteration of microrna expression correlates to fatty acidmediated insulin resistance in mouse myoblasts Supplemental Data Correlation coefficient matrix con PA PA 0.9425 1.0000 PA+OA 0.9407 0.9626 con
Alteration of microrna expression correlates to fatty acidmediated insulin resistance in mouse myoblasts Supplemental Data Correlation coefficient matrix con PA PA 0.9425 1.0000 PA+OA 0.9407 0.9626 con
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Endogenous gene tagging to study subcellular localization and chromatin binding. a, b, Schematic of experimental set-up to endogenously tag RNAi factors using the CRISPR Cas9 technology,
Supplementary Figure 1 Endogenous gene tagging to study subcellular localization and chromatin binding. a, b, Schematic of experimental set-up to endogenously tag RNAi factors using the CRISPR Cas9 technology,
(a) Immunoblotting to show the migration position of Flag-tagged MAVS
 Supplementary Figure 1 Characterization of six MAVS isoforms. (a) Immunoblotting to show the migration position of Flag-tagged MAVS isoforms. HEK293T Mavs -/- cells were transfected with constructs expressing
Supplementary Figure 1 Characterization of six MAVS isoforms. (a) Immunoblotting to show the migration position of Flag-tagged MAVS isoforms. HEK293T Mavs -/- cells were transfected with constructs expressing
Genome Sequence Assembly
 Genome Sequence Assembly Learning Goals: Introduce the field of bioinformatics Familiarize the student with performing sequence alignments Understand the assembly process in genome sequencing Introduction:
Genome Sequence Assembly Learning Goals: Introduce the field of bioinformatics Familiarize the student with performing sequence alignments Understand the assembly process in genome sequencing Introduction:
supplementary information
 DOI: 10.1038/ncb2116 Figure S1 CDK phosphorylation of EZH2 in cells. (a) Comparison of candidate CDK phosphorylation sites on EZH2 with known CDK substrates by multiple sequence alignments. (b) CDK1 and
DOI: 10.1038/ncb2116 Figure S1 CDK phosphorylation of EZH2 in cells. (a) Comparison of candidate CDK phosphorylation sites on EZH2 with known CDK substrates by multiple sequence alignments. (b) CDK1 and
RNA Interference and the World of Small RNAs
 RNA Interference and the World of Small RNAs O, I die, Horatio; The potent poison quite o'er-crows my spirit: I cannot live to hear the news from England; But I do prophesy the election lights On Fortinbras:
RNA Interference and the World of Small RNAs O, I die, Horatio; The potent poison quite o'er-crows my spirit: I cannot live to hear the news from England; But I do prophesy the election lights On Fortinbras:
CRISPR RNA-guided activation of endogenous human genes
 CRISPR RNA-guided activation of endogenous human genes Morgan L Maeder, Samantha J Linder, Vincent M Cascio, Yanfang Fu, Quan H Ho, J Keith Joung Supplementary Figure 1 Comparison of VEGF activation induced
CRISPR RNA-guided activation of endogenous human genes Morgan L Maeder, Samantha J Linder, Vincent M Cascio, Yanfang Fu, Quan H Ho, J Keith Joung Supplementary Figure 1 Comparison of VEGF activation induced
Supplementary Information
 Supplementary Information Negative regulation of initial steps in skeletal myogenesis by mtor and other kinases Raphael A. Wilson 1*, Jing Liu 1*, Lin Xu 1, James Annis 2, Sara Helmig 1, Gregory Moore
Supplementary Information Negative regulation of initial steps in skeletal myogenesis by mtor and other kinases Raphael A. Wilson 1*, Jing Liu 1*, Lin Xu 1, James Annis 2, Sara Helmig 1, Gregory Moore
Suplementary Materials Epub: No 2016_1339 Vol. 63, Regular paper
 Suplementary Materials Epub: No 2016_1339 Vol. 63, 2016 https://doi.org/10.18388/abp.2016_1339 Regular paper How short RNAs impact the human ribonuclease Dicer activity: putative regulatory feedback-loops
Suplementary Materials Epub: No 2016_1339 Vol. 63, 2016 https://doi.org/10.18388/abp.2016_1339 Regular paper How short RNAs impact the human ribonuclease Dicer activity: putative regulatory feedback-loops
Transcriptional Regulation (Gene Regulation)
 Experimental Techniques in Biomedical Sciences 의생명과학실험기법 Transcriptional Regulation (Gene Regulation) 4/17/13 Jeong Hoon Kim (jeongkim@skku.edu) Department of Health Sciences and Technology, SKKU Graduate
Experimental Techniques in Biomedical Sciences 의생명과학실험기법 Transcriptional Regulation (Gene Regulation) 4/17/13 Jeong Hoon Kim (jeongkim@skku.edu) Department of Health Sciences and Technology, SKKU Graduate
Supplementary Information Design of small molecule-responsive micrornas based on structural requirements for Drosha processing
 Supplementary Information Design of small molecule-responsive micrornas based on structural requirements for Drosha processing Chase L. Beisel, Yvonne Y. Chen, Stephanie J. Culler, Kevin G. Hoff, & Christina
Supplementary Information Design of small molecule-responsive micrornas based on structural requirements for Drosha processing Chase L. Beisel, Yvonne Y. Chen, Stephanie J. Culler, Kevin G. Hoff, & Christina
Long-term, efficient inhibition of microrna function in mice using raav vectors
 Nature Methods Long-term, efficient inhibition of microrna function in mice using raav vectors Jun Xie, Stefan L Ameres, Randall Friedline, Jui-Hung Hung, Yu Zhang, Qing Xie, Li Zhong, Qin Su, Ran He,
Nature Methods Long-term, efficient inhibition of microrna function in mice using raav vectors Jun Xie, Stefan L Ameres, Randall Friedline, Jui-Hung Hung, Yu Zhang, Qing Xie, Li Zhong, Qin Su, Ran He,
Galina Gabriely, Ph.D. BWH/HMS
 Galina Gabriely, Ph.D. BWH/HMS Email: ggabriely@rics.bwh.harvard.edu Outline: microrna overview microrna expression analysis microrna functional analysis microrna (mirna) Characteristics mirnas discovered
Galina Gabriely, Ph.D. BWH/HMS Email: ggabriely@rics.bwh.harvard.edu Outline: microrna overview microrna expression analysis microrna functional analysis microrna (mirna) Characteristics mirnas discovered
8Br-cAMP was purchased from Sigma (St. Louis, MO). Silencer Negative Control sirna #1 and
 1 Supplemental information 2 3 Materials and Methods 4 Reagents and animals 5 8Br-cAMP was purchased from Sigma (St. Louis, MO). Silencer Negative Control sirna #1 and 6 Silencer Select Pre-designed sirna
1 Supplemental information 2 3 Materials and Methods 4 Reagents and animals 5 8Br-cAMP was purchased from Sigma (St. Louis, MO). Silencer Negative Control sirna #1 and 6 Silencer Select Pre-designed sirna
Supplementary Materials for
 advances.sciencemag.org/cgi/content/full/3/11/eaao1799/dc1 Supplementary Materials for Measuring quantitative effects of methylation on transcription factor DNA binding affinity Zheng Zuo, Basab Roy, Yiming
advances.sciencemag.org/cgi/content/full/3/11/eaao1799/dc1 Supplementary Materials for Measuring quantitative effects of methylation on transcription factor DNA binding affinity Zheng Zuo, Basab Roy, Yiming
QPCR ASSAYS FOR MIRNA EXPRESSION PROFILING
 TECH NOTE 4320 Forest Park Ave Suite 303 Saint Louis, MO 63108 +1 (314) 833-9764 mirna qpcr ASSAYS - powered by NAWGEN Our mirna qpcr Assays were developed by mirna experts at Nawgen to improve upon previously
TECH NOTE 4320 Forest Park Ave Suite 303 Saint Louis, MO 63108 +1 (314) 833-9764 mirna qpcr ASSAYS - powered by NAWGEN Our mirna qpcr Assays were developed by mirna experts at Nawgen to improve upon previously
sirna Transfection Into Primary Neurons Using Fuse-It-siRNA
 sirna Transfection Into Primary Neurons Using Fuse-It-siRNA This Application Note describes a protocol for sirna transfection into sensitive, primary cortical neurons using Fuse-It-siRNA. This innovative
sirna Transfection Into Primary Neurons Using Fuse-It-siRNA This Application Note describes a protocol for sirna transfection into sensitive, primary cortical neurons using Fuse-It-siRNA. This innovative
CRISPR-dCas9 mediated TET1 targeting for selective DNA demethylation at BRCA1 promoter
 CRISPR-dCas9 mediated TET1 targeting for selective DNA demethylation at BRCA1 promoter SUPPLEMENTARY DATA See Supplementary Sequence File: 1 Supplementary Figure S1: The total protein was extracted from
CRISPR-dCas9 mediated TET1 targeting for selective DNA demethylation at BRCA1 promoter SUPPLEMENTARY DATA See Supplementary Sequence File: 1 Supplementary Figure S1: The total protein was extracted from
transcription and the promoter occupancy of Smad proteins. (A) HepG2 cells were co-transfected with the wwp-luc reporter, and FLAG-tagged FHL1,
 Supplementary Data Supplementary Figure Legends Supplementary Figure 1 FHL-mediated TGFβ-responsive reporter transcription and the promoter occupancy of Smad proteins. (A) HepG2 cells were co-transfected
Supplementary Data Supplementary Figure Legends Supplementary Figure 1 FHL-mediated TGFβ-responsive reporter transcription and the promoter occupancy of Smad proteins. (A) HepG2 cells were co-transfected
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION Supplementary figure 1: List of primers/oligonucleotides used in this study. 1 Supplementary figure 2: Sequences and mirna-targets of i) mcherry expresses in transgenic fish used
SUPPLEMENTARY INFORMATION Supplementary figure 1: List of primers/oligonucleotides used in this study. 1 Supplementary figure 2: Sequences and mirna-targets of i) mcherry expresses in transgenic fish used
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/4/198/ra74/dc1 Supplementary Materials for Short RNA Duplexes Elicit RIG-I Mediated Apoptosis in a Cell Type and Length-Dependent Manner Osamu Ishibashi, Md. Moksed
www.sciencesignaling.org/cgi/content/full/4/198/ra74/dc1 Supplementary Materials for Short RNA Duplexes Elicit RIG-I Mediated Apoptosis in a Cell Type and Length-Dependent Manner Osamu Ishibashi, Md. Moksed
Revision Checklist for Science Signaling Research Manuscripts: Data Requirements and Style Guidelines
 Revision Checklist for Science Signaling Research Manuscripts: Data Requirements and Style Guidelines Further information can be found at: http://stke.sciencemag.org/sites/default/files/researcharticlerevmsinstructions_0.pdf.
Revision Checklist for Science Signaling Research Manuscripts: Data Requirements and Style Guidelines Further information can be found at: http://stke.sciencemag.org/sites/default/files/researcharticlerevmsinstructions_0.pdf.
mir-24-mediated down-regulation of H2AX suppresses DNA repair
 Supplemental Online Material mir-24-mediated down-regulation of H2AX suppresses DNA repair in terminally differentiated blood cells Ashish Lal 1,4, Yunfeng Pan 2,4, Francisco Navarro 1,4, Derek M. Dykxhoorn
Supplemental Online Material mir-24-mediated down-regulation of H2AX suppresses DNA repair in terminally differentiated blood cells Ashish Lal 1,4, Yunfeng Pan 2,4, Francisco Navarro 1,4, Derek M. Dykxhoorn
Adenoviral Expression Systems. Lentivirus is not the only choice for gene delivery. Adeno-X
 Adenoviral Expression Systems Lentivirus is not the only choice for gene delivery 3 Adeno-X Why choose adenoviral gene delivery? Table I: Adenoviral vs. Lentiviral Gene Delivery Lentivirus Adenovirus Infects
Adenoviral Expression Systems Lentivirus is not the only choice for gene delivery 3 Adeno-X Why choose adenoviral gene delivery? Table I: Adenoviral vs. Lentiviral Gene Delivery Lentivirus Adenovirus Infects
Supplementary Figure 1. (A) Cell proliferative ability and (B) the invasiveness of
 LEGEND FOR SUPPLEMENTARY FIGURES Supplementary Figure 1. (A) Cell proliferative ability and (B) the invasiveness of LLC-1, LLC-3, and LLC-5 cell line series were quantified by BrdU assay after 72 h of
LEGEND FOR SUPPLEMENTARY FIGURES Supplementary Figure 1. (A) Cell proliferative ability and (B) the invasiveness of LLC-1, LLC-3, and LLC-5 cell line series were quantified by BrdU assay after 72 h of
Concepts and Methods in Developmental Biology
 Biology 4361 Developmental Biology Concepts and Methods in Developmental Biology June 16, 2009 Conceptual and Methodological Tools Concepts Genomic equivalence Differential gene expression Differentiation/de-differentiation
Biology 4361 Developmental Biology Concepts and Methods in Developmental Biology June 16, 2009 Conceptual and Methodological Tools Concepts Genomic equivalence Differential gene expression Differentiation/de-differentiation
Cambridge University Press
 Figure 1.1. Model of RNAi pathway in C. elegans. Transmembrane protein SID-1 allows dsrna to enter the cell. In the cytoplasm,dsrna gets processed by DCR-1,existing in a complex with RDE-4,RDE-1 and DRH-1.
Figure 1.1. Model of RNAi pathway in C. elegans. Transmembrane protein SID-1 allows dsrna to enter the cell. In the cytoplasm,dsrna gets processed by DCR-1,existing in a complex with RDE-4,RDE-1 and DRH-1.
Exploring of microrna markers for body fluid identification using NGS
 Exploring of microrna markers for body fluid identification using NGS Zheng Wang, Yiping Hou Institute of Forensic Medicine Sichuan University, China Barcelona May, 11, 2016 Outline Introduction of Institute
Exploring of microrna markers for body fluid identification using NGS Zheng Wang, Yiping Hou Institute of Forensic Medicine Sichuan University, China Barcelona May, 11, 2016 Outline Introduction of Institute
Supporting Online Material for
 www.sciencemag.org/cgi/content/full/1137999/dc1 Supporting Online Material for Disrupting the Pairing Between let-7 and Enhances Oncogenic Transformation Christine Mayr, Michael T. Hemann, David P. Bartel*
www.sciencemag.org/cgi/content/full/1137999/dc1 Supporting Online Material for Disrupting the Pairing Between let-7 and Enhances Oncogenic Transformation Christine Mayr, Michael T. Hemann, David P. Bartel*
Pattern Formation via Small RNA Mobility SUPPLEMENTAL FIGURE 1 SUPPLEMENTAL INFORMATION. Daniel H. Chitwood et al.
 SUPPLEMENTAL INFORMATION Pattern Formation via Small RNA Mobility Daniel H. Chitwood et al. SUPPLEMENTAL FIGURE 1 Supplemental Figure 1. The ARF3 promoter drives expression throughout leaves. (A, B) Expression
SUPPLEMENTAL INFORMATION Pattern Formation via Small RNA Mobility Daniel H. Chitwood et al. SUPPLEMENTAL FIGURE 1 Supplemental Figure 1. The ARF3 promoter drives expression throughout leaves. (A, B) Expression
Clontech Product Selection Guide
 New New Clontech Product Selection Guide PCR THigh Yield - TITANIUM Taq High Yield & Fidelity - Advantage 2 polymerase High Fidelity CloneAmp HiFi Direct PCR- Terra PCR Direct polymearase Transfection
New New Clontech Product Selection Guide PCR THigh Yield - TITANIUM Taq High Yield & Fidelity - Advantage 2 polymerase High Fidelity CloneAmp HiFi Direct PCR- Terra PCR Direct polymearase Transfection
Fig. S1. eif6 expression in HEK293 transfected with shrna against eif6 or pcmv-eif6 vector.
 Fig. S1. eif6 expression in HEK293 transfected with shrna against eif6 or pcmv-eif6 vector. (a) Western blotting analysis and (b) qpcr analysis of eif6 expression in HEK293 T cells transfected with either
Fig. S1. eif6 expression in HEK293 transfected with shrna against eif6 or pcmv-eif6 vector. (a) Western blotting analysis and (b) qpcr analysis of eif6 expression in HEK293 T cells transfected with either
3 UTR (untranslated region) Reporter Clone and its vector, pmirtarget. Application Guide. OriGene Technologies, Inc
 3 UTR (untranslated region) Reporter Clone and its vector, pmirtarget Application Guide OriGene Technologies, Inc Package Contents and Storage Conditions 3 UTR reporter clone as 10ug lyophilized plasmid
3 UTR (untranslated region) Reporter Clone and its vector, pmirtarget Application Guide OriGene Technologies, Inc Package Contents and Storage Conditions 3 UTR reporter clone as 10ug lyophilized plasmid
SUPPLEMENTAL MATERIAL. Carvajal et al. Supplemental Table 1. List of quantitative PCR primers used for cdna analyses and
 UPPLEMEAL MATERIAL Carvajal et al. upplemental Table 1. List of quantitative PCR primers used for cdna analyses and chromatin immunoprecipitation assays. Figure 1. DNA damage-induced transcriptional repression
UPPLEMEAL MATERIAL Carvajal et al. upplemental Table 1. List of quantitative PCR primers used for cdna analyses and chromatin immunoprecipitation assays. Figure 1. DNA damage-induced transcriptional repression
B. Transgenic plants with strong phenotype (%)
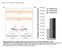 A. TCTAGTTGTTGTTGTTATGGTCTAGTTGTTGTTGTTATGGTCTAATTT AAATATGGTCTAAAGAAGAAGAATATGGTCTAAAGAAGAAGAATATGG 2XP35S STTM165 5 GGGGGATGAAGctaCCTGGTCCGA3 3 CCCCCUACUUC---GGACCAGGCU5 mir165 HindIII mir165 96 nt GTTGTTGTTGTTATGGTCTAGTTGTTGTTGTTATGGTCTAATTT
A. TCTAGTTGTTGTTGTTATGGTCTAGTTGTTGTTGTTATGGTCTAATTT AAATATGGTCTAAAGAAGAAGAATATGGTCTAAAGAAGAAGAATATGG 2XP35S STTM165 5 GGGGGATGAAGctaCCTGGTCCGA3 3 CCCCCUACUUC---GGACCAGGCU5 mir165 HindIII mir165 96 nt GTTGTTGTTGTTATGGTCTAGTTGTTGTTGTTATGGTCTAATTT
Total RNA was isolated using Trizol reagent (Invitrogen) and reverse transcribed using
 Supplementary Methods RNA Isolation and Quantitative RT-PCR Total RNA was isolated using Trizol reagent (Invitrogen) and reverse transcribed using random hexamers and superscript II reverse transcriptase
Supplementary Methods RNA Isolation and Quantitative RT-PCR Total RNA was isolated using Trizol reagent (Invitrogen) and reverse transcribed using random hexamers and superscript II reverse transcriptase
SUPPLEMENTARY INFORMATION
 doi: 10.1038/nature05841 SUPPLEMENTARY INFORMATION www.nature.com/nature 1 www.nature.com/nature 2 www.nature.com/nature 3 www.nature.com/nature 4 a Hours of Development: eif-6(rnai): 4 wildtype 30 4 lin-4
doi: 10.1038/nature05841 SUPPLEMENTARY INFORMATION www.nature.com/nature 1 www.nature.com/nature 2 www.nature.com/nature 3 www.nature.com/nature 4 a Hours of Development: eif-6(rnai): 4 wildtype 30 4 lin-4
Supplementary Figure 1. Reintroduction of HDAC1 in HDAC1-/- ES cells reverts the
 SUPPLEMENTARY FIGURE LEGENDS Supplementary Figure 1. Reintroduction of HDAC1 in HDAC1-/- ES cells reverts the phenotype of HDAC1-/- teratomas. 3x10 6 HDAC1 reintroduced (HDAC1-/-re) and empty vector infected
SUPPLEMENTARY FIGURE LEGENDS Supplementary Figure 1. Reintroduction of HDAC1 in HDAC1-/- ES cells reverts the phenotype of HDAC1-/- teratomas. 3x10 6 HDAC1 reintroduced (HDAC1-/-re) and empty vector infected
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb2880 Supplementary Figure 1 Sequence alignment of Deup1 and Cep63. The protein sequence alignment was generated by the Clustal X 2.0 multiple sequence alignment program using default parameters.
DOI: 10.1038/ncb2880 Supplementary Figure 1 Sequence alignment of Deup1 and Cep63. The protein sequence alignment was generated by the Clustal X 2.0 multiple sequence alignment program using default parameters.
Supplementary Figure 1: Overexpression of EBV-encoded proteins Western blot analysis of the expression levels of EBV-encoded latency III proteins in
 Supplementary Figure 1: Overexpression of EBV-encoded proteins Western blot analysis of the expression levels of EBV-encoded latency III proteins in BL2 cells. The Ponceau S staining of the membranes or
Supplementary Figure 1: Overexpression of EBV-encoded proteins Western blot analysis of the expression levels of EBV-encoded latency III proteins in BL2 cells. The Ponceau S staining of the membranes or
[KCl] Heated. No SS III. No NE. unspliced spliced. b kda
![[KCl] Heated. No SS III. No NE. unspliced spliced. b kda [KCl] Heated. No SS III. No NE. unspliced spliced. b kda](/thumbs/86/93796483.jpg) rrna level a 1 MDQGYGGYGA WSAGPANTQG AYGTGVASWQ GYENYNYYGA QNTSVTTGAT YSYGPASWEA 61 AKANDGGLAA GAPAMHMASY GPEPCTDNSD SLIAKINQRL DMMSKEGGRG GSGGGGEGIQ 121 DRESSFRFQP FESYDSRPCL PEHNPYRPSY SYDYEFDLGS DRNGSFGGQY
rrna level a 1 MDQGYGGYGA WSAGPANTQG AYGTGVASWQ GYENYNYYGA QNTSVTTGAT YSYGPASWEA 61 AKANDGGLAA GAPAMHMASY GPEPCTDNSD SLIAKINQRL DMMSKEGGRG GSGGGGEGIQ 121 DRESSFRFQP FESYDSRPCL PEHNPYRPSY SYDYEFDLGS DRNGSFGGQY
RNA oligonucleotides and 2 -O-methylated oligonucleotides were synthesized by. 5 AGACACAAACACCAUUGUCACACUCCACAGC; Rand-2 OMe,
 Materials and methods Oligonucleotides and DNA constructs RNA oligonucleotides and 2 -O-methylated oligonucleotides were synthesized by Dharmacon Inc. (Lafayette, CO). The sequences were: 122-2 OMe, 5
Materials and methods Oligonucleotides and DNA constructs RNA oligonucleotides and 2 -O-methylated oligonucleotides were synthesized by Dharmacon Inc. (Lafayette, CO). The sequences were: 122-2 OMe, 5
Return to Web Version
 Page 1 of 7 Page 1 of 7 Return to Web Version ZFN Technology Biowire Volume 10 Article 1 Have your genomic work cut out for you The genomes of several organisms, including humans, have been sequenced,
Page 1 of 7 Page 1 of 7 Return to Web Version ZFN Technology Biowire Volume 10 Article 1 Have your genomic work cut out for you The genomes of several organisms, including humans, have been sequenced,
Site-Directed Mutagenesis. Mutations in four Smad4 sites of mouse Gat1 promoter
 Supplement Supporting Materials and Methods Site-Directed Mutagenesis. Mutations in four Smad4 sites of mouse Gat1 promoter were independently generated using a two-step PCR method. The Smad4 binding site
Supplement Supporting Materials and Methods Site-Directed Mutagenesis. Mutations in four Smad4 sites of mouse Gat1 promoter were independently generated using a two-step PCR method. The Smad4 binding site
Supplementary Fig. 1. Schematic structure of TRAIP and RAP80. The prey line below TRAIP indicates bait and the two lines above RAP80 highlight the
 Supplementary Fig. 1. Schematic structure of TRAIP and RAP80. The prey line below TRAIP indicates bait and the two lines above RAP80 highlight the prey clones identified in the yeast two hybrid screen.
Supplementary Fig. 1. Schematic structure of TRAIP and RAP80. The prey line below TRAIP indicates bait and the two lines above RAP80 highlight the prey clones identified in the yeast two hybrid screen.
Time allowed: 2 hours Answer ALL questions in Section A, ALL PARTS of the question in Section B and ONE question from Section C.
 UNIVERSITY OF EAST ANGLIA School of Biological Sciences Main Series UG Examination 2013-2014 MOLECULAR BIOLOGY BIO-2B02 Time allowed: 2 hours Answer ALL questions in Section A, ALL PARTS of the question
UNIVERSITY OF EAST ANGLIA School of Biological Sciences Main Series UG Examination 2013-2014 MOLECULAR BIOLOGY BIO-2B02 Time allowed: 2 hours Answer ALL questions in Section A, ALL PARTS of the question
SUPPLEMENTARY INFORMATION
 Figure S1: Activation of the ATM pathway by I-PpoI. A. HEK293T cells were either untransfected, vector transfected, transfected with an I-PpoI expression vector, or subjected to 2Gy γ-irradiation. 24 hrs
Figure S1: Activation of the ATM pathway by I-PpoI. A. HEK293T cells were either untransfected, vector transfected, transfected with an I-PpoI expression vector, or subjected to 2Gy γ-irradiation. 24 hrs
Supplementary Information for. Regulation of Rev1 by the Fanconi Anemia Core Complex
 Supplementary Information for Regulation of Rev1 by the Fanconi Anemia Core Complex Hyungjin Kim, Kailin Yang, Donniphat Dejsuphong, Alan D. D Andrea* *Corresponding Author: Alan D. D Andrea, M.D. Alan_dandrea@dfci.harvard.edu
Supplementary Information for Regulation of Rev1 by the Fanconi Anemia Core Complex Hyungjin Kim, Kailin Yang, Donniphat Dejsuphong, Alan D. D Andrea* *Corresponding Author: Alan D. D Andrea, M.D. Alan_dandrea@dfci.harvard.edu
Somatic Primary pirna Biogenesis Driven by cis-acting RNA Elements and Trans-Acting Yb
 Cell Reports Supplemental Information Somatic Primary pirna Biogenesis Driven by cis-acting RNA Elements and Trans-Acting Yb Hirotsugu Ishizu, Yuka W. Iwasaki, Shigeki Hirakata, Haruka Ozaki, Wataru Iwasaki,
Cell Reports Supplemental Information Somatic Primary pirna Biogenesis Driven by cis-acting RNA Elements and Trans-Acting Yb Hirotsugu Ishizu, Yuka W. Iwasaki, Shigeki Hirakata, Haruka Ozaki, Wataru Iwasaki,
Online Supplementary Information
 Online Supplementary Information NLRP4 negatively regulates type I interferon signaling by targeting TBK1 for degradation via E3 ubiquitin ligase DTX4 Jun Cui 1,4,6,7, Yinyin Li 1,5,6,7, Liang Zhu 1, Dan
Online Supplementary Information NLRP4 negatively regulates type I interferon signaling by targeting TBK1 for degradation via E3 ubiquitin ligase DTX4 Jun Cui 1,4,6,7, Yinyin Li 1,5,6,7, Liang Zhu 1, Dan
Non-Organic-Based Isolation of Mammalian microrna using Norgen s microrna Purification Kit
 Application Note 13 RNA Sample Preparation Non-Organic-Based Isolation of Mammalian microrna using Norgen s microrna Purification Kit B. Lam, PhD 1, P. Roberts, MSc 1 Y. Haj-Ahmad, M.Sc., Ph.D 1,2 1 Norgen
Application Note 13 RNA Sample Preparation Non-Organic-Based Isolation of Mammalian microrna using Norgen s microrna Purification Kit B. Lam, PhD 1, P. Roberts, MSc 1 Y. Haj-Ahmad, M.Sc., Ph.D 1,2 1 Norgen
Experimental genetics - 2 Partha Roy
 Partha Roy Experimental genetics - 2 Making genetically altered animal 1) Gene knock-out k from: a) the entire animal b) selected cell-type/ tissue c) selected cell-type/tissue at certain time 2) Transgenic
Partha Roy Experimental genetics - 2 Making genetically altered animal 1) Gene knock-out k from: a) the entire animal b) selected cell-type/ tissue c) selected cell-type/tissue at certain time 2) Transgenic
OmicsLink shrna Clones guaranteed knockdown even in difficult-to-transfect cells
 OmicsLink shrna Clones guaranteed knockdown even in difficult-to-transfect cells OmicsLink shrna clone collections consist of lentiviral, and other mammalian expression vector based small hairpin RNA (shrna)
OmicsLink shrna Clones guaranteed knockdown even in difficult-to-transfect cells OmicsLink shrna clone collections consist of lentiviral, and other mammalian expression vector based small hairpin RNA (shrna)
JCB. Supplemental material THE JOURNAL OF CELL BIOLOGY. Kimura et al.,
 Supplemental material JCB Kimura et al., http://www.jcb.org/cgi/content/full/jcb.201503023/dc1 THE JOURNAL OF CELL BIOLOGY Figure S1. TRIMs regulate IFN-γ induced autophagy. (A and B) HC image analysis
Supplemental material JCB Kimura et al., http://www.jcb.org/cgi/content/full/jcb.201503023/dc1 THE JOURNAL OF CELL BIOLOGY Figure S1. TRIMs regulate IFN-γ induced autophagy. (A and B) HC image analysis
Supplementary Figure 1. Nature Structural & Molecular Biology: doi: /nsmb.3494
 Supplementary Figure 1 Pol structure-function analysis (a) Inactivating polymerase and helicase mutations do not alter the stability of Pol. Flag epitopes were introduced using CRISPR/Cas9 gene targeting
Supplementary Figure 1 Pol structure-function analysis (a) Inactivating polymerase and helicase mutations do not alter the stability of Pol. Flag epitopes were introduced using CRISPR/Cas9 gene targeting
It s All in the Details (or Small RNA): Simplified and Improved mirna Purification from Tissue
 It s All in the Details (or Small RNA): Simplified and Improved mirna Purification from Tissue Douglas Horejsh January 2015 Discussion Outline Non-coding RNA General Overview mirna What is it? The Future
It s All in the Details (or Small RNA): Simplified and Improved mirna Purification from Tissue Douglas Horejsh January 2015 Discussion Outline Non-coding RNA General Overview mirna What is it? The Future
SUPPLEMENTARY INFORMATION
 Materials and Methods Transgenic Plant Materials and DNA Constructs. VEX1::H2B-GFP, ACA3::H2B-GFP, KRP6::H2B-GFP, KRP6::mock21ts-GFP, KRP6::TE21ts- GFP and KRP6::miR161ts-GFP constructs were generated
Materials and Methods Transgenic Plant Materials and DNA Constructs. VEX1::H2B-GFP, ACA3::H2B-GFP, KRP6::H2B-GFP, KRP6::mock21ts-GFP, KRP6::TE21ts- GFP and KRP6::miR161ts-GFP constructs were generated
