Nature Methods: doi: /nmeth Supplementary Figure 1
|
|
|
- Jessica Lorena Moore
- 5 years ago
- Views:
Transcription
1 Supplementary Figure 1 ATP1A1 variants with in-frame deletions are enriched in ouabain-resistant cell populations. (a) Total editing efficacy along with spectrum and frequency of individual indels as determined by the TIDE assay in K562 cells treated with ATP1A1 G2 sgrna. Analysis was performed on genomic DNA samples shown in Figure 1b. (b) Same as (a) but for ATP1A1 G4 sgrna. (c) Same as (a) but for ATP1A1 G3 sgrna.
2
3 Supplementary Figure 2 HDR-driven editing at ATP1A1 induces cellular resistance to ouabain. (a) Schematic representation of the intronic SpCas9 target site G3 and partial sequences of single-stranded oligodeoxynucleotides (ssodns) donors used to introduce the Q118D/R and N129D/R mutations. Annotated are novel restriction sites to monitor the insertion of ssodn-specified mutations. (b) K562 cells stably expressing SpCas9 were co-transfected with an ATP1A1 G3 sgrna expression vector (500ng) along with the indicated ssodns (10pmol). Genomic DNA was harvested 10 days post-transfection and a PvuI RFLP assay was used to determine the frequency of HDR at the cleavage site, indicated as the % HDR at the base of each lane. Where indicated, cells were treated with 0.5 M ouabain for 7 days starting 3 days post transfection. An expression vector encoding EGFP (-) was used as a negative control. (c) Same as in (b) but using BmgBI. (d) Same as in (b) but using ClaI. (e) Same as in (b) but using Hpy188I. (f) Same as in (e) but cells initially selected at 0.5 M ouabain were cultivated in the presence of increasing concentrations of the drug for a week. (g) Uncropped gel images for panels (e) and (f). A naturally occurring Hpy188I (TCNGA cleavage site) is present in the amplicon creating the strong RFLP signal indicated by the red arrows. The HDR-specific signal (black arrow; predicted at 77bp) is running below the 100bp marker and is best viewed when the gels are overexposed making it is impossible to quantify this signal using RFLP-based assays.
4
5 Supplementary Figure 3 Ouabain-based selection is portable to various cell lines and compatible with CRISPR AsCpf1-driven editing. (a) U2OS cells were treated and analyzed as in Figure 1b. (b) htert-rpe-1 cells were treated and analyzed as in Figure 1b and Supplementary Figure 2d. (c) Schematic representation of AsCpf1 target sites surrounding DNA encoding the first extracellular loop of human ATP1A1. Annotated are the positions of residues Q118 and N129, exon/intron boundary, protospacer adjacent motifs (PAM) and five potential AsCpf1 target sequences (Targets 1-5). Also shown are partial sequences of single-stranded oligodeoxynucleotides (ssodns) donors used to introduce the Q118R and N129D mutations. Annotated are novel restriction sites to monitor the insertion of ssodn-specified mutations. (d) K562 cells stably expressing AsCpf1 were transfected with ATP1A1 crrna expression vectors (G1-G5) (500 ng) and the Surveyor assay was used to determine the frequency of AsCpf1-induced insertions and deletions, indicated as the % Indels at the base of each lane. Where indicated, cells were treated with 0.5 M ouabain for 7 days starting 3 days post transfection. An expression vector encoding EGFP (-) was used as a negative control. (e) K562 cells were transfected with a vector expressing espcas9(1.1) and the ATP1A1 G3 sgrna (500ng each) or with a vector expressing AsCpf1 and the ATP1A1 G5 crrna (500ng each) (see Supplementary Figure 12 for details on the vectors) along with the indicated ssodns. Cells were treated as in (d) and assayed using BmgBI and EcoRI RFLP assays.
6
7 Supplementary Figure 4 Selection for CRISPR Cas9-driven targeted mutagenesis by coediting ATP1A1 via NHEJ. (a) Experimental strategy for the co-enrichment of CRISPR-driven editing at a second locus. GOI, gene of interest. (b) Typical selection process. Timing can vary according to initial modification rates at ATP1A1. (c) Schematic of the dual espcas9(1.1) and tandem U6- driven sgrnas expression vector (see also Supplementary Figure 12). (d) Surveyor assays to determine on-target and off-target activity for the EMX1 sgrna on samples reported in Table 1. (e) Same as (d) but for co-targeting AAVS1. (f) Surveyor assays on samples from K562 cells transiently co-transfected with WT SpCas9 and sgrna expression vectors targeting AAVS1 (0.5 g and 1 g of each vector).
8
9 Supplementary Figure 5 Selection for CRISPR AsCpf1-driven targeted mutagenesis by coediting ATP1A1 via NHEJ. (a) Schematic of the dual AsCpf1 and U6-driven crrna array expression vector (see also Supplementary Figure 12). (b) K562 cells were transfected with 500ng of the vectors shown in (a), treated and assayed for indels as shown in Supplementary Figure 4d. (c) Same as in (a). (d) Same as in (b) but using 1 g of the vector. (e) HEK293 cells were treated and analyzed as in (b) but the TIDE assay was used to determine the frequency of indels. Value for ATP1A1 relates to the co-selection performed with the DNMT1 crrna. (f) Same as in (e) but using the Surveyor assay.
10 Supplementary Figure 6 Off-target analysis in K562 and HEK293 cells coselected with the AsCpf1 crrna array vector targeting DNMT1. (a) Schematic of the dual AsCpf1 and U6-driven crrna array expression vector. (b) Surveyor assays to determine on-target and offtarget activity performed on samples reported in Table 1 and Supplementary Table 1. To facilitate the interpretation of the results, the gels for on-target activity presented in Supplementary Figure 5b,f have been reproduced here. Off-target activity at DNMT1-OT1 was assayed using two different sets of primers since it is found within a repetitive element making its detection more challenging. (c) Ontarget activity at DNMT1 was also determined using the TIDE assay in both co-selected and transiently transfected cells. (d) Surveyor assays to determine on-target and off-target activity in K562 cells transiently transfected with 1 g and 2 g of the vector shown in (a).
11
12 Supplementary Figure 7 Selection for CRISPR-driven targeted integration at LMNA by coediting ATP1A1 via HDR. (a) Targeting scheme for the integration of Clover and mruby2 to the N-terminus of Lamin A/C. (b) Fluorescence imaging of ouabaintreated cells expressing the Lamin A/C-Clover fusion. Scale bar, 25 m (c) A ClaI RFLP assay was used to determine the frequency of SpCas9-induced HDR at the ATP1A1 locus in samples shown in Figure 2. % HDR is indicated at the base of each lane. (d) FACSbased quantification of Clover targeting to the N-terminus of Lamin A/C at various doses of WT SpCas9 and espcas9(1.1) in presence or absence of co-selection with ouabain. (e) Same as in (d) but in cells transfected with both Clover and mruby2 donors. (f) Same as in (d) but targeting LMNA using an AsCpf1 crrna array. Transfection conditions are indicated in Supplementary Table 2.
13 Supplementary Figure 8 Sequencing of non-hdr alleles from LMNA coselection experiments. (a) Schematic of a PCR-based assay (out-out PCR) used to detect targeted integration of the Clover or mruby2 sequences at the N- terminus of Lamin A/C. Primers are located outside of the homology arms and are designed to yield a longer PCR product if the fluorescent protein is inserted. The upper band representing the targeted integration of either Clover or mruby2 was TOPO cloned and sequenced to confirm the accurate integration of the fluorescent proteins via HDR. (b) The lower band from (a) representing nontargeted LMNA alleles was TOPO cloned, sequenced and analyzed using the web tool CrispRVariantsLite. (c) ATP1A1 alleles from the same experiment were also sequenced and non-hdr alleles were analyzed as in (b). Samples are from the experiment shown in Figure 2. Ref: Lindsay, H. et al. CrispRVariants charts the mutation spectrum of genome engineering experiments. Nat Biotechnol. 34, (2016)
14
15 Supplementary Figure 9 Selection for CRISPR-driven targeted integration at endogenous loci by coediting ATP1A1 via HDR. (a) Targeting scheme for the integration of mag1 to the C-terminus of H2BK. (b) K562 cells were transfected with a vector expressing espcas9(1.1) and tandem U6-driven sgrnas targeting ATP1A1 and HIST1H2BK along with ATP1A1 ssodn RD and a plasmid donor containing a mag1 cassette. In addition, cells were transfected with a wild-type SpCas9 expression vector and two independent sgrna vectors along with donors. Cells were treated or not with 0.5 M ouabain for 14 days starting 3 days post-transfection and flow cytometry was used to determine the % of mag1 positive cells in each population. (c) Fluorescence imaging of ouabain-treated cells expressing the H2BK-mAG1 fusion. Scale bar, 10 m (d) FACS-based quantification of targeted integration of a PGK1-EGFP-pA expression cassette at HPRT1 following co-selection using WT SpCas9 and various amounts of sgrnas. (e) FACS-based quantification of targeted integration of a SA-2A-EGFP-pA gene trap cassette at AAVS1 following co-selection using an all-in-one vector expressing ATP1A1 and AAVS1 sgrnas in addition to espcas9(1.1). (f) Same as in (e) but using a PGK1-TurboGFP-pA cassette. Transfection conditions are indicated in Supplementary Table 2.
16 Supplementary Figure 10 Endogenous tagging of the NuA4 TIP60 complex in coselected HEK293 cells. (a) Schematic representation of the experiment. (b) Gene tagging scheme. (c) Western blot analysis on whole cell extracts from HEK293 cells to detect the expression of EPC1-tag and EP400-tag proteins. The FLAG M2 antibody was used to detect tagged proteins and the tubulin antibody was used as a loading control.
17
18 Supplementary Figure 11 Introduction of the sickle mutation in primary human cord blood (CB) CD34 + cells by coediting ATP1A1 via HDR. (a) Typical selection process. Timing can vary according to initial modification rates at ATP1A1. (b) Schematic representation of SpCas9 target site in ATP1A1 and predicted HDR outcome dictated by the ssodn donor used to introduce the Q118R and N129D mutations. Annotated are novel restriction sites to monitor the insertion of ssodn-specified mutations. (c) Cultured CD34 + cells were electroporated with ATP1A1 and HBB RNPs along with HBB ssodn #1 and ATP1A1 G4 RD ssodn and treated as shown in (a). Genomic DNA was harvested at indicated time points and a BmgBI RFLP assay was used to determine the frequency of SpCas9- induced HDR at ATP1A1, which is indicated as the % HDR at the base of each lane. Recombinant Cas9 was used as a negative control (-). (d) Same as in (c) but using the Surveyor assay to determine the total frequency of edited alleles at ATP1A1 (NHEJ + HDR). (e) Schematic representation of SpCas9 target site in HBB and predicted HDR outcome dictated by ssodns donor #2 used to introduce the E6V mutation. Annotated are the positions of E6 residue, 5 UTR, protospacer adjacent motifs (PAM) and novel restriction site to monitor the insertion of ssodn-specified mutations. (f) Same as in (c) but using HBB ssodn #2 and a PstI RFLP assay to determine the frequency of SpCas9-induced HDR at HBB. (g) Same as in (f) but using the Surveyor assay to determine the total frequency of HBB edited alleles (NHEJ + HDR). (h) Same as in (f) but using a BmgBI RFLP assay to determine the frequency of SpCas9-induced HDR at ATP1A1. (i) Same as in (g) but using the Surveyor assay to determine the total frequency of ATP1A1 edited alleles (NHEJ + HDR).
19 Supplementary Figure 12 espcas9(1.1) and AsCpf1 vectors for targeting ATP1A1, available from Addgene. Ref for py036: Engineered Cpf1 Enzymes with Altered PAM Specificities. Linyi Gao, David B.T. Cox, Winston X Yan, John Manteiga, Martin Schneider, Takashi Yamano, Hiroshi Nishimasu, Osamu Nureki, Feng Zhang. biorxiv ; doi:
20 Supplementary Note 1 Several approaches have been implemented to capture and isolate cells expressing high amounts of nucleases. First, co-transfection of fluorescent proteins combined with fluorescence-activated cell sorting (FACS) can be used to isolate nuclease-expressing cells 1. Second, direct coupling of expression of fluorescent proteins and nucleases via 2A peptide sequences allows for efficient isolation of cell populations with increasingly higher nuclease expression levels, which translates into increasingly higher genome editing rates 2. Fluorescence-based surrogate target gene reporters have also been successfully used to enrich for cells with high nuclease activity 3, 4. A limitation of these methods is that single-cell FACS enrichment is not suitable for more sensitive cell lines. In addition, multi-dsbs arising from cleavage of the episomal reporter plasmids are likely to induce a gene-independent antiproliferative response as recently described for CRISPR-Cas9-induced DNA breaks 5, 6. SUPPLEMENTARY REFERENCES 1. Whyte, J.J. et al. Gene targeting with zinc finger nucleases to produce cloned egfp knockout pigs. Mol Reprod Dev 78, 2 (2011). 2. Duda, K. et al. High-efficiency genome editing via 2A-coupled co-expression of fluorescent proteins and zinc finger nucleases or CRISPR/Cas9 nickase pairs. Nucleic Acids Res 42, e84 (2014). 3. Certo, M.T. et al. Tracking genome engineering outcome at individual DNA breakpoints. Nat Methods 8, (2011). 4. Kim, H. et al. Surrogate reporters for enrichment of cells with nuclease-induced mutations. Nat Methods 8, (2011). 5. Aguirre, A.J. et al. Genomic Copy Number Dictates a Gene-Independent Cell Response to CRISPR/Cas9 Targeting. Cancer Discov 6, (2016). 6. Munoz, D.M. et al. CRISPR Screens Provide a Comprehensive Assessment of Cancer Vulnerabilities but Generate False-Positive Hits for Highly Amplified Genomic Regions. Cancer Discov 6, (2016). Nature Methods doi: /nmeth.4265
21 Supplementary Data 1 Total editing efficacy along with spectrum and frequency of individual indels as determined by the TIDE assay for Table 1 and Supplementary Table 1. espcas9(1.1) tandem U6-driven sgrnas EMX1 + ATP1A1 (K562-50ng) EMX1 untreated EMX1 ouabain ATP1A1 untreated ATP1A1 ouabain espcas9 (1.1) tandem U6-driven sgrnas EMX1 + ATP1A1 (K ng) EMX1 untreated EMX1 ouabain ATP1A1 untreated ATP1A1 ouabain Nature Methods doi: /nmeth.4265
22 espcas9 (1.1) tandem U6-driven sgrnas AAVS1 + ATP1A1 (K562 50ng) AAVS1 untreated AAVS1 ouabain ATP1A1 untreated ATP1A1 ouabain espcas9 (1.1) tandem U6-driven sgrnas AAVS1 + ATP1A1 (K ng) AAVS1 untreated AAVS1 ouabain ATP1A1 untreated ATP1A1 ouabain Nature Methods doi: /nmeth.4265
23 AsCpf1 crrna array DMNT1 + ATP1A1 (K ng) DNMT1 untreated DNMT1 ouabain ATP1A1 untreated ATP1A1 ouabain AsCpf1 crrna array EMX1 + ATP1A1 (K ng) EMX1 untreated EMX1 ouabain ATP1A1 untreated ATP1A1 ouabain Nature Methods doi: /nmeth.4265
24 AsCpf1 crrna array VEGFA + ATP1A1 (K ng) VEGFA untreated VEGFA ouabain ATP1A1 untreated ATP1A1 ouabain AsCpf1 crrna array GRIN2b + ATP1A1 (K ng) GRIN2B untreated GRIN2B ouabain ATP1A1 untreated ATP1A1 ouabain Nature Methods doi: /nmeth.4265
25 AsCpf1 crrna array DNMT1 + EMX1 + VEGFA + GRIN2b + ATP1A1 (K562-1µg) DNMT1 untreated DNMT1 ouabain EMX1 untreated EMX1 ouabain VEGFA untreated VEGFA ouabain GRIN2B untreated GRIN2B ouabain ATP1A1 untreated ATP1A1 ouabain Nature Methods doi: /nmeth.4265
26 AsCpf1 crrna array DNMT1 + ATP1A1 (HEK293LTV - 500ng) DNMT1 untreated DNMT1 ouabain ATP1A1 untreated ATP1A1 ouabain AsCpf1 crrna array EMX1 + ATP1A1 (HEK293LTV - 500ng) EMX1 untreated EMX1 ouabain ATP1A1 untreated ATP1A1 ouabain Nature Methods doi: /nmeth.4265
27 AsCpf1 crrna array VEGFA + ATP1A1 (HEK293LTV - 500ng) VEGFA untreated VEGFA ouabain ATP1A1 untreated ATP1A1 ouabain AsCpf1 crrna array GRIN2b + ATP1A1 (HEK293LTV - 500ng) GRIN2B untreated GRIN2B ouabain ATP1A1 untreated ATP1A1 ouabain Nature Methods doi: /nmeth.4265
28 Supplementary Table 1 Co-selection for CRISPR-induced indels in HEK293 cells. TIDE Assay (% indels 1 ) Surveyor Assay (% indels 1 ) Target Nuclease Dose (ng) Untreated Ouabain Untreated Ouabain DNMT1 AsCpf EMX1 " VEGFA " GRIN2b " ATP1A1 3 " Rounded to the nearest one. Undetectable set to 0. 2 U6-driven crrna array (2 guides) coupled to AsCpf1 expression from a single vector. 3 Value taken from DNMT1 co-selection experiment. Supplementary Table 2 Transfection mixes for HDR-based co-selections Figure 2 SpCas9_LMNA_G2 (px330) 100 ng Clover_LMNA_Plasmid_donor 250 ng mruby2_lmna_plasmid_donor 250 ng ATP1A1_G3_pUC19_sgRNA 100 ng ssodn_atp1a1_rd 5 pmol Supplementary Figure 7d espcas9(1.1)_no_flag_atp1a1_g3_dual_lmna_g2 50 ng 100 ng 500 ng Clover_LMNA_Plasmid_donor 250 ng 250 ng 250 ng ssodn_atp1a1_rd 5 pmol 5 pmol 5 pmol SpCas9_LMNA_G2 (px330) Clover_LMNA_Plasmid_donor ATP1A1_G3_pUC19_sgRNA ssodn_atp1a1_rd 100 ng 250 ng 100 ng 5 pmol Supplementary Figure 7e espcas9(1.1)_no_flag_atp1a1_g3_dual_lmna_g2 100 ng Clover_LMNA_Plasmid_donor 250 ng mruby2_lmna_plasmid_donor 250 ng ssodn_atp1a1_rd 5 pmol SpCas9_LMNA_G2 (px330) Clover_LMNA_Plasmid_donor mruby2_lmna_plasmid_donor ATP1A1_G3_pUC19_sgRNA ssodn_atp1a1_rd 100 ng 250 ng 250 ng 100 ng 5 pmol Nature Methods doi: /nmeth.4265
29 Supplementary Figure 7f py036_atp1a1_g5_array_lmna Clover_LMNA_Plasmid_donor ssodn_atp1a1_rd2 Supplementary Figure 9b espcas9(1.1)_no_flag_atp1a1_g3_dual_h2bk H2BK_mAG1_Plasmid_donor ssodn_atp1a1_rd 1000 ng 250 ng 5 pmol 100 ng 250 ng 5 pmol Transfection for WT SpCas9 was done with 1E6 cells in an amaxa cuvette hcas9 (WT) 500 ng H2BK_mAG1_Plasmid_donor 1250 ng ATP1A1_G3_pUC19_sgRNA 500 ng HIST1H2BK_pUC19_sgRNA 500 ng ssodn_atp1a1_rd 10 pmol Supplementary Figure 9d Transfection was done with 1E6 cells in an amaxa cuvette hcas9 (WT) 500 ng 500 ng hpgk1_egfp_plasmid_donor 1000 ng 1000 ng ATP1A1_G3_pUC19_sgRNA 50 ng 500 ng HPRT1_pUC19_sgRNA 50 ng 50 ng ssodn_atp1a1_rd 10 pmol 10 pmol Supplementary Figure 9e espcas9(1.1)_no_flag_aavs1 AAVS1_SA_2A_EGFP_Plasmid_donor ATP1A1_G3_pUC19_sgRNA ssodn_atp1a1_rd 100 ng 250 ng 100 ng 5 pmol Supplementary Figure 9f espcas9(1.1)_no_flag_atp1a1_g3_dual_aavs1 50 ng AAVS1_PGK_TurboGFP_Plasmid_donor 500 ng ATP1A1_RD_Plasmid_donor 5 pmol Figure 3 and Supplementary Figure 10 espcas9(1.1)_no_flag_atp1a1_g3 P400_FS_Plasmid_donor P400_sgRNA_pUC19_sgRNA ssodn_atp1a1_g3_rd 100 ng 250 ng 100 ng 5 pmol Nature Methods doi: /nmeth.4265
30 espcas9(1.1)_no_flag_atp1a1_g3 EPC1_FS_Plasmid_donor EPC1_sgRNA_pUC19_sgRNA ssodn_atp1a1_g3_rd 100 ng 250 ng 100 ng 5 pmol Supplementary Table 3 Sequences for SpCas9 guides Target Sequence ATP1A1 G1 GAACTCACATTATCGTTTTG ATP1A1 G2 GATCCAAGCTGCTACAGAAG ATP1A1 G3 GAGTTCTGTAATTCAGCATA ATP1A1 G4 GTTCCTCTTCTGTAGCAGCT AAVS1 GGGGCCACTAGGGACAGGAT LMNA G2 GCCATGGAGACCCCGTCCCAG EMX1 GAGTCCGAGCAGAAGAAGAA EPC1 GTGACGTAGCTTCCTCCGAG EP400 GGCCCTGACTACTGGCACGG HPRT1 GATGTGATGAAGGAGATGGG HIST1H2BK GGGGCTTTAAGACGCTTACT Supplementary Table 4 Sequences for AsCpf1 guides Target Sequence ATP1A1 G1 TTTCTTGGCTTATAGCATCC ATP1A1 G2 TTGGCTTATAGCATCCAAGC ATP1A1 G3 AGGTTCCTCTTCTGTAGCAG ATP1A1 G4 GAGGTTCCTCTTCTGTAGCA ATP1A1 G5 TAGTACACATCAGATATCTT ATP1A1 G3 (23bp for crrna arrays) AGGTTCCTCTTCTGTAGCAGCTT ATP1A1 G5 (23bp for crrna arrays) TAGTACACATCAGATATCTTCTC DNMT1 CTGATGGTCCATGTCTGTTACTC EMX1 TGGTTGCCCACCCTAGTCATTGG GRIN2b GTGCTCAATGAAAGGAGATAAGG VEGFA CTAGGAATATTGAAGGGGGCAGG LMNA CGGGACCCCTGCCCCGCGGGCAG Nature Methods doi: /nmeth.4265
31 Supplementary Table 5 Sequences for ssodns ssodn Sequence ATP1A1 DR CAATGTTACTGTGGATTGGAGCGATTCTTTGTTTCTTGGCTT ATAGCATCGATGCTGCTACAGAAGAGGAACCTCAAAACGAT CGTGTGAGTTCTGTAATTCAGCATATCGATTTGTAGTACACA TCAGATATCTT ATP1A1 RD CAATGTTACTGTGGATTGGAGCGATTCTTTGTTTCTTGGCTT ATAGCATCAGAGCTGCTACAGAAGAGGAACCTCAAAACGAT GACGTGAGTTCTGTAATTCAGCATATCGATTTGTAGTACACA TCAGATATCTT ATP1A1 RD2 TCTTTGTTTCTTGGCTTATAGCATCAGAGCTGCTACAGAAGA GGAACCTCAAAACGATGACGTGAGTTCTGTAATTCAGCATA TGAATTCGTAGTACACATCAGATATCTTCTCCGTCTTTGTCT CCCACTTCTTCTCAATTACCACTCATTACTTAATGGTTAT ATP1A1 G4 RD AAGATATCTGATGTGTACTACAAATCCATATGCTGAATTAC AGAACTCACGTCATCGTTTTGAGGTTCCTCTTCTGTAGCAGC TCTGATGCTATAAGCCAAGAAACAAAGAATCGCTCCAATCC ACAGTAACATTG HBB #1 TCTGACACAACTGTGTTCACTAGCAACCTCAAACAGACACC ATGGTGCATCTGACTCCTGTCGAGAAGTCTGCAGTTACTGCC CTGTGGGGCAAGGTGAACGTGGATGAAGTTGGTGGTGAGGC CCTGGGCAG HBB #2 TCTGACACAACTGTGTTCACTAGCAACCTCAAACAGACACC ATGGTGCATCTGACTCCTGTGGAGAAGTCTGCAGTTACTGCC CTGTGGGGCAAGGTGAACGTGGATGAAGTTGGTGGTGAGGC CCTGGGCAG Supplementary Table 6 PCR primers used for Surveyor assays and expected size of cleavage products. Target Primer Size (bp) Surveyor Signal (bp) ATP1A1 SpCas9 G2 Forward GGATTAACATCTGCTCGTGCAGC ATP1A1 SpCas9 G2 Reverse CACTTGTAAGAGCATCTACAACG /238 ATP1A1 SpCas9 G3 Forward GGATTAACATCTGCTCGTGCAGC ATP1A1 SpCas9 G3 Reverse CACTTGTAAGAGCATCTACAACG /196 ATP1A1 SpCas9 G4 Forward GGATTAACATCTGCTCGTGCAGC ATP1A1 SpCas9 G4 Reverse CACTTGTAAGAGCATCTACAACG /246 ATP1A1 AsCpf1 G2 Forward GGATTAACATCTGCTCGTGCAGC ATP1A1 AsCpf1 G2 Reverse CACTTGTAAGAGCATCTACAACG /247 ATP1A1 AsCpf1 G3 Forward GGATTAACATCTGCTCGTGCAGC ATP1A1 AsCpf1 G3 Reverse CACTTGTAAGAGCATCTACAACG /246 ATP1A1 AsCpf1 G5 Forward GGATTAACATCTGCTCGTGCAGC ATP1A1 AsCpf1 G5 Reverse CACTTGTAAGAGCATCTACAACG /166 AAVS1 Forward TTCGGGTCACCTCTCACTCC AAVS1 Reverse GGCTCCATCGTAAGCAAACC /181 Nature Methods doi: /nmeth.4265
32 DNMT1 site 3 Forward DNMT1 site 3 Reverse EMX1 Forward EMX1 Reverse VEGFA Forward VEGFA Reverse GRIN2b-7 Forward GRIN2b-7 Reverse HBB Forward HBB Reverse DNMT1 site 3 OT1 Forward 1 DNMT1 site 3 OT1 Reverse 1 DNMT1 site 3 OT1 Forward 2 DNMT1 site 3 OT1 Reverse 2 AAVS1 OT1 Forward AAVS1 OT1 Reverse EMX1 OT1 Forward EMX1 OT1 Reverse EMX1 OT2 Forward EMX1 OT2 Reverse CTGGGACTCAGGCGGGTCAC CCTCACACAACAGCTTCATGTCAGC CCATCCCCTTCTGTGAATGT GGAGATTGGAGACACGGAGA GAGAAGGCCAGGGGTCACTCCAG AGCCCGCCGCAATGAAGG GCATACTCGCATGGCTACCT CTCCCTGCAGCCCCTTTTTA GACAGGTACGGCTGTCATCA CAGCCTAAGGGTGGGAAAAT GCCCCATTAGCCTGTGTTTT GTGTGCAAATAGGAAGGTGTCT CGGAAGTATTTCTGAGAGCATGG GAAGGTGTCTAACTCTCCAATCG GGGATGAGTCATGTCCACCACAG TTACATCCCCTCTGGGCATGAGC ATCTGCACATGTATGTACAGGAGTC TCTCTCCTTCAACTCATGACCAGC CACTGCCTACCTTCCTCTACTTC TCTTCATCCTCCTCATCTGAGGACTC / / / / / / / / / /200 Supplementary Table 7 Primers used for out-out PCR assays Target Primer Size (bp) LMNA OO Forward TGAGTCACACTGATGGGCACC LMNA OO Reverse TTCTAACAGGACCCTTTCTGCC 1404 P400 OO Forward CTCTCACCCTTTTCCCAAGA P400 OO Reverse AAGACCACGAGGCATTTTTC 988 EPC1 OO Forward AAGCCTGACACAAATCTCAGT EPC1 OO Reverse CCAAGGAGTCCACAGCTACC 943 Nature Methods doi: /nmeth.4265
Nature Biotechnology: doi: /nbt Supplementary Figure 1. In vitro validation of OTC sgrnas and donor template.
 Supplementary Figure 1 In vitro validation of OTC sgrnas and donor template. (a) In vitro validation of sgrnas targeted to OTC in the MC57G mouse cell line by transient transfection followed by 4-day puromycin
Supplementary Figure 1 In vitro validation of OTC sgrnas and donor template. (a) In vitro validation of sgrnas targeted to OTC in the MC57G mouse cell line by transient transfection followed by 4-day puromycin
CRISPR/Cas9 Gene Editing Tools
 CRISPR/Cas9 Gene Editing Tools - Guide-it Products for Successful CRISPR/Cas9 Gene Editing - Why choose Guide-it products? Optimized methods designed for speed and ease of use Complete kits that don t
CRISPR/Cas9 Gene Editing Tools - Guide-it Products for Successful CRISPR/Cas9 Gene Editing - Why choose Guide-it products? Optimized methods designed for speed and ease of use Complete kits that don t
CRISPR/Cas9 Gene Editing Tools
 CRISPR/Cas9 Gene Editing Tools - Separations Simply Spectacular INDELS Identify indels Determine if one or both copies of your gene have indels The Guide-it Genotype Confirmation Kit: Simple detection
CRISPR/Cas9 Gene Editing Tools - Separations Simply Spectacular INDELS Identify indels Determine if one or both copies of your gene have indels The Guide-it Genotype Confirmation Kit: Simple detection
CRISPR/Cas9 Genome Editing: Transfection Methods
 CRISPR/ Genome Editing: Transfection Methods For over 20 years Mirus Bio has developed and manufactured high performance transfection products and technologies. That expertise is now being applied to the
CRISPR/ Genome Editing: Transfection Methods For over 20 years Mirus Bio has developed and manufactured high performance transfection products and technologies. That expertise is now being applied to the
Optimizing CRISPR ribonucleoprotein components for precision genome editing
 Optimizing CRISPR ribonucleoprotein components for precision genome editing Justin Barr Sr. Product Manager, Integrated DNA Technologies VIB CRISPR User Meeting 18 September 2018 1 Agenda Overview of CRISPR-Cas9
Optimizing CRISPR ribonucleoprotein components for precision genome editing Justin Barr Sr. Product Manager, Integrated DNA Technologies VIB CRISPR User Meeting 18 September 2018 1 Agenda Overview of CRISPR-Cas9
Applications of Cas9 nickases for genome engineering
 application note genome editing Applications of Cas9 nickases for genome engineering Shuqi Yan, Mollie Schubert, Maureen Young, Brian Wang Integrated DNA Technologies, 17 Commercial Park, Coralville, IA,
application note genome editing Applications of Cas9 nickases for genome engineering Shuqi Yan, Mollie Schubert, Maureen Young, Brian Wang Integrated DNA Technologies, 17 Commercial Park, Coralville, IA,
Transfection of CRISPR/Cas9 Nuclease NLS ribonucleoprotein (RNP) into adherent mammalian cells using Lipofectamine RNAiMAX
 Transfection of CRISPR/Cas9 Nuclease NLS ribonucleoprotein (RNP) into adherent mammalian cells using Lipofectamine RNAiMAX INTRODUCTION The CRISPR/Cas genome editing system consists of a single guide RNA
Transfection of CRISPR/Cas9 Nuclease NLS ribonucleoprotein (RNP) into adherent mammalian cells using Lipofectamine RNAiMAX INTRODUCTION The CRISPR/Cas genome editing system consists of a single guide RNA
CRISPR RNA-guided activation of endogenous human genes
 CRISPR RNA-guided activation of endogenous human genes Morgan L Maeder, Samantha J Linder, Vincent M Cascio, Yanfang Fu, Quan H Ho, J Keith Joung Supplementary Figure 1 Comparison of VEGF activation induced
CRISPR RNA-guided activation of endogenous human genes Morgan L Maeder, Samantha J Linder, Vincent M Cascio, Yanfang Fu, Quan H Ho, J Keith Joung Supplementary Figure 1 Comparison of VEGF activation induced
domain. Bottom panel: hybrid surface/ribbon structure (PDB ID: 4UN3) of SpCas9 in complex with sgrna and target DNA. The REC3
 Supplementary Figure 1 Yeast screening for high-specificity SpCas9 variants (a) Top panel: scheme of SpCas9 domains. The REC3 domain is part of the recognition lobe. BH: bridge helix. PI: PAM-interacting
Supplementary Figure 1 Yeast screening for high-specificity SpCas9 variants (a) Top panel: scheme of SpCas9 domains. The REC3 domain is part of the recognition lobe. BH: bridge helix. PI: PAM-interacting
Return to Web Version
 Page 1 of 7 Page 1 of 7 Return to Web Version ZFN Technology Biowire Volume 10 Article 1 Have your genomic work cut out for you The genomes of several organisms, including humans, have been sequenced,
Page 1 of 7 Page 1 of 7 Return to Web Version ZFN Technology Biowire Volume 10 Article 1 Have your genomic work cut out for you The genomes of several organisms, including humans, have been sequenced,
SUPPLEMENTARY INFORMATION
 Gene replacements and insertions in rice by intron targeting using CRISPR Cas9 Table of Contents Supplementary Figure 1. sgrna-induced targeted mutations in the OsEPSPS gene in rice protoplasts. Supplementary
Gene replacements and insertions in rice by intron targeting using CRISPR Cas9 Table of Contents Supplementary Figure 1. sgrna-induced targeted mutations in the OsEPSPS gene in rice protoplasts. Supplementary
Genome Engineering with ZFNs, TALENs and CRISPR/Cas9
 Genome Engineering with ZFNs, TALENs and CRISPR/Cas9 Designer Endonucleases ZFNs (zinc finger nucleases), TALENs (transcription activator-like effector nucleases) and CRISPR/Cas9 (clustered regularly interspaced
Genome Engineering with ZFNs, TALENs and CRISPR/Cas9 Designer Endonucleases ZFNs (zinc finger nucleases), TALENs (transcription activator-like effector nucleases) and CRISPR/Cas9 (clustered regularly interspaced
IMPROVEMENT OF CRISPR GENE EDITING EFFICIENCY AND BEYONDS
 IMPROVEMENT OF CRISPR GENE EDITING EFFICIENCY AND BEYONDS YONGLUN LUO (ALUN) ALUN@BIOMED.AU.DK VIB, NOV. 21. 2017 Associate Professor, Department of Biomedicine, Aarhus University, Denmark Executive Director,
IMPROVEMENT OF CRISPR GENE EDITING EFFICIENCY AND BEYONDS YONGLUN LUO (ALUN) ALUN@BIOMED.AU.DK VIB, NOV. 21. 2017 Associate Professor, Department of Biomedicine, Aarhus University, Denmark Executive Director,
Improving CRISPR-Cas9 Gene Knockout with a Validated Guide RNA Algorithm
 Improving CRISPR-Cas9 Gene Knockout with a Validated Guide RNA Algorithm Anja Smith Director R&D Dharmacon, part of GE Healthcare Imagination at work crrna:tracrrna program Cas9 nuclease Active crrna is
Improving CRISPR-Cas9 Gene Knockout with a Validated Guide RNA Algorithm Anja Smith Director R&D Dharmacon, part of GE Healthcare Imagination at work crrna:tracrrna program Cas9 nuclease Active crrna is
User Instructions:Transfection-ready CRISPR/Cas9 Reagents. Target DNA. NHEJ repair pathway. Nucleotide deletion. Nucleotide insertion Gene disruption
 User Instructions:Transfection-ready CRISPR/Cas9 Reagents Background Introduction to CRISPR/Cas9 genome editing In bacteria and archaea, clustered regularly interspaced short palindromic repeats (CRISPR)
User Instructions:Transfection-ready CRISPR/Cas9 Reagents Background Introduction to CRISPR/Cas9 genome editing In bacteria and archaea, clustered regularly interspaced short palindromic repeats (CRISPR)
Supplementary Information
 Supplementary Information Super-resolution imaging of fluorescently labeled, endogenous RNA Polymerase II in living cells with CRISPR/Cas9-mediated gene editing Won-Ki Cho 1, Namrata Jayanth 1, Susan Mullen
Supplementary Information Super-resolution imaging of fluorescently labeled, endogenous RNA Polymerase II in living cells with CRISPR/Cas9-mediated gene editing Won-Ki Cho 1, Namrata Jayanth 1, Susan Mullen
Surrogate reporter-based enrichment of cells containing RNA-guided Cas9 nucleaseinduced
 Supplementary Data Surrogate reporter-based enrichment of cells containing RNA-guided Cas9 nucleaseinduced mutations Suresh Ramakrishna 1, Seung Woo Cho 2, Sojung Kim 2, Myungjae Song 1, Ramu Gopalappa
Supplementary Data Surrogate reporter-based enrichment of cells containing RNA-guided Cas9 nucleaseinduced mutations Suresh Ramakrishna 1, Seung Woo Cho 2, Sojung Kim 2, Myungjae Song 1, Ramu Gopalappa
Simple protocol for gene editing using GenCrisprTM Cas9 nuclease
 Simple protocol for gene editing using GenCrisprTM Cas9 nuclease Contents Protocol Step 1: Choose the target DNA sequence Step 2: Design sgrna Step 3: Preparation for sgrna 3.1 In vitro transcription of
Simple protocol for gene editing using GenCrisprTM Cas9 nuclease Contents Protocol Step 1: Choose the target DNA sequence Step 2: Design sgrna Step 3: Preparation for sgrna 3.1 In vitro transcription of
SUPPLEMENTARY INFORMATION
 VOLUME: 1 ARTICLE NUMBER: 0066 In the format provided by the authors and unedited. Engineering CRISPR-Cpf1 crrnas and mrnas to maximize genome editing efficiency Bin Li 1, Weiyu Zhao 1, Xiao Luo 1, Xinfu
VOLUME: 1 ARTICLE NUMBER: 0066 In the format provided by the authors and unedited. Engineering CRISPR-Cpf1 crrnas and mrnas to maximize genome editing efficiency Bin Li 1, Weiyu Zhao 1, Xiao Luo 1, Xinfu
Nature Methods: doi: /nmeth Supplementary Figure 1. Comparison of REC2 and BCL-xL domains and modeling domain replacement.
 Supplementary Figure 1 Comparison of REC2 and BCL-xL domains and modeling domain replacement. (a) The REC2 domain of Cas9 and (b) BCL-xL display similar globular structures and volumes. (c, d) Modeling
Supplementary Figure 1 Comparison of REC2 and BCL-xL domains and modeling domain replacement. (a) The REC2 domain of Cas9 and (b) BCL-xL display similar globular structures and volumes. (c, d) Modeling
Nature Biotechnology: doi: /nbt.4166
 Supplementary Figure 1 Validation of correct targeting at targeted locus. (a) by immunofluorescence staining of 2C-HR-CRISPR microinjected embryos cultured to the blastocyst stage. Embryos were stained
Supplementary Figure 1 Validation of correct targeting at targeted locus. (a) by immunofluorescence staining of 2C-HR-CRISPR microinjected embryos cultured to the blastocyst stage. Embryos were stained
Construct Design and Cloning Guide for Cas9-triggered homologous recombination
 Construct Design and Cloning Guide for Cas9-triggered homologous recombination Written by Dan Dickinson (ddickins@live.unc.edu) and last updated December 2013. Reference: Dickinson DJ, Ward JD, Reiner
Construct Design and Cloning Guide for Cas9-triggered homologous recombination Written by Dan Dickinson (ddickins@live.unc.edu) and last updated December 2013. Reference: Dickinson DJ, Ward JD, Reiner
Genome-wide genetic screening with chemically-mutagenized haploid embryonic stem cells
 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 Supplementary Information Genome-wide genetic screening with chemically-mutagenized haploid embryonic stem cells Josep V. Forment 1,2, Mareike Herzog
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 Supplementary Information Genome-wide genetic screening with chemically-mutagenized haploid embryonic stem cells Josep V. Forment 1,2, Mareike Herzog
Introduction to CRISPR/Cas9 Background DNA Cleavage and Repair (NHEJ and HDR) Alternative Cas9 Variants Delivery of Cas9 and sgrna Library Products
 Introduction to CRISPR/Cas9 Background DNA Cleavage and Repair (NHEJ and HDR) Alternative Cas9 Variants Delivery of Cas9 and sgrna Library Products which one is right for you? CRISPR Workflow abm s Toolbox
Introduction to CRISPR/Cas9 Background DNA Cleavage and Repair (NHEJ and HDR) Alternative Cas9 Variants Delivery of Cas9 and sgrna Library Products which one is right for you? CRISPR Workflow abm s Toolbox
CRISPR Ribonucleoprotein (RNP) User Manual
 CRISPR Ribonucleoprotein (RNP) User Manual Description This user manual describes how to use GenScript s CRISPR Ribonucleoprotein (RNP) products for targeted genome editing. The RNP system is comprised
CRISPR Ribonucleoprotein (RNP) User Manual Description This user manual describes how to use GenScript s CRISPR Ribonucleoprotein (RNP) products for targeted genome editing. The RNP system is comprised
Supplementary Information. Optimization of the production of knock-in alleles by CRISPR/Cas9 microinjection into the mouse zygote
 Supplementary Information Supplementary Figures S1 to S5 and Tables S1 to S3. Optimization of the production of knock-in alleles by CRISPR/Cas9 microinjection into the mouse zygote Aurélien Raveux, Sandrine
Supplementary Information Supplementary Figures S1 to S5 and Tables S1 to S3. Optimization of the production of knock-in alleles by CRISPR/Cas9 microinjection into the mouse zygote Aurélien Raveux, Sandrine
Development of High Quality CRISPR/Cas9 Agents
 Development of High Quality CRISPR/Cas9 Agents TIDES May 7 th, 2018 Terence Ta editasmedicine.com 1 Agenda Overview of CRISPR and Editas platform Development of NGS-based method for guide RNA QC Covalently-coupled
Development of High Quality CRISPR/Cas9 Agents TIDES May 7 th, 2018 Terence Ta editasmedicine.com 1 Agenda Overview of CRISPR and Editas platform Development of NGS-based method for guide RNA QC Covalently-coupled
Genome editing. Knock-ins
 Genome editing Knock-ins Experiment design? Should we even do it? In mouse or rat, the HR-mediated knock-in of homologous fragments derived from a donor vector functions well. However, HR-dependent knock-in
Genome editing Knock-ins Experiment design? Should we even do it? In mouse or rat, the HR-mediated knock-in of homologous fragments derived from a donor vector functions well. However, HR-dependent knock-in
Supplementary Figure 1. Nature Structural & Molecular Biology: doi: /nsmb.3494
 Supplementary Figure 1 Pol structure-function analysis (a) Inactivating polymerase and helicase mutations do not alter the stability of Pol. Flag epitopes were introduced using CRISPR/Cas9 gene targeting
Supplementary Figure 1 Pol structure-function analysis (a) Inactivating polymerase and helicase mutations do not alter the stability of Pol. Flag epitopes were introduced using CRISPR/Cas9 gene targeting
Allele-specific locus binding and genome editing by CRISPR at the
 Supplementary Information Allele-specific locus binding and genome editing by CRISPR at the p6ink4a locus Toshitsugu Fujita, Miyuki Yuno, and Hodaka Fujii Supplementary Figure Legends Supplementary Figure
Supplementary Information Allele-specific locus binding and genome editing by CRISPR at the p6ink4a locus Toshitsugu Fujita, Miyuki Yuno, and Hodaka Fujii Supplementary Figure Legends Supplementary Figure
File name: Supplementary Information Description: Supplementary figures and supplementary tables. File name: Peer review file Description:
 File name: Supplementary Information Description: Supplementary figures and supplementary tables. File name: Peer review file Description: Supplementary Figure 1. dcas9-mq1 fusion protein induces de novo
File name: Supplementary Information Description: Supplementary figures and supplementary tables. File name: Peer review file Description: Supplementary Figure 1. dcas9-mq1 fusion protein induces de novo
Supplementary Information
 Supplementary Information Vidigal and Ventura a wt locus 5 region 3 region CCTCTGCCACTGCGAGGGCGTCCAATGGTGCTTG(...)AACAGGTGGAATATCCCTACTCTA predicted deletion clone 1 clone 2 clone 3 CCTCTGCCACTGCGAGGGCGTC-AGGTGGAATATCCCTACTCTA
Supplementary Information Vidigal and Ventura a wt locus 5 region 3 region CCTCTGCCACTGCGAGGGCGTCCAATGGTGCTTG(...)AACAGGTGGAATATCCCTACTCTA predicted deletion clone 1 clone 2 clone 3 CCTCTGCCACTGCGAGGGCGTC-AGGTGGAATATCCCTACTCTA
A universal chassis plasmid pwh34 was first constructed to facilitate the
 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 File name: Supplementary method Plasmids construction A universal chassis plasmid pwh34 was first constructed to facilitate the construction
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 File name: Supplementary method Plasmids construction A universal chassis plasmid pwh34 was first constructed to facilitate the construction
Protocol CRISPR Genome Editing In Cell Lines
 Protocol CRISPR Genome Editing In Cell Lines Protocol 2: HDR donor plasmid applications (gene knockout, gene mutagenesis, gene tagging, Safe Harbor ORF knock-in) Notes: 1. sgrna validation: GeneCopoeia
Protocol CRISPR Genome Editing In Cell Lines Protocol 2: HDR donor plasmid applications (gene knockout, gene mutagenesis, gene tagging, Safe Harbor ORF knock-in) Notes: 1. sgrna validation: GeneCopoeia
Using CRISPR for genetic alteration
 Using CRISPR for genetic alteration Joffrey Mianné. j.mianne@har.mrc.ac.uk Mary Lyon Centre, MRC Harwell. CRISPR/Cas origins Origin of the CRISPR/Cas system: Clustered-Regularly Interspaced Short Palindromic
Using CRISPR for genetic alteration Joffrey Mianné. j.mianne@har.mrc.ac.uk Mary Lyon Centre, MRC Harwell. CRISPR/Cas origins Origin of the CRISPR/Cas system: Clustered-Regularly Interspaced Short Palindromic
Easi CRISPR for conditional and insertional alleles
 Easi CRISPR for conditional and insertional alleles C.B Gurumurthy, University Of Nebraska Medical Center Omaha, NE cgurumurthy@unmc.edu Types of Genome edits Gene disruption/inactivation Types of Genome
Easi CRISPR for conditional and insertional alleles C.B Gurumurthy, University Of Nebraska Medical Center Omaha, NE cgurumurthy@unmc.edu Types of Genome edits Gene disruption/inactivation Types of Genome
i-stop codon positions in the mcherry gene
 Supplementary Figure 1 i-stop codon positions in the mcherry gene The grnas (green) that can potentially generate stop codons from Trp (63 th and 98 th aa, upper panel) and Gln (47 th and 114 th aa, bottom
Supplementary Figure 1 i-stop codon positions in the mcherry gene The grnas (green) that can potentially generate stop codons from Trp (63 th and 98 th aa, upper panel) and Gln (47 th and 114 th aa, bottom
Nature Biotechnology: doi: /nbt Supplementary Figure 1
 Supplementary Figure 1 Negative selection analysis of sgrnas targeting all Brd4 exons comparing day 2 to day 10 time points. Systematic evaluation of 64 Brd4 sgrnas in negative selection experiments, targeting
Supplementary Figure 1 Negative selection analysis of sgrnas targeting all Brd4 exons comparing day 2 to day 10 time points. Systematic evaluation of 64 Brd4 sgrnas in negative selection experiments, targeting
Protocol for cloning SEC-based repair templates using Gibson assembly and ccdb negative selection
 Protocol for cloning SEC-based repair templates using Gibson assembly and ccdb negative selection Written by Dan Dickinson (daniel.dickinson@austin.utexas.edu) and last updated January 2018. A version
Protocol for cloning SEC-based repair templates using Gibson assembly and ccdb negative selection Written by Dan Dickinson (daniel.dickinson@austin.utexas.edu) and last updated January 2018. A version
Effect of Different CRISPR/Cas9 Variants on Repair Pathway Choice
 Effect of Different CRISPR/Cas9 Variants on Repair Pathway Choice Cecilia Cotta-Ramusino www.editasmedicine.com 1 Cas9 Stimulates the Endogenous Repair Pathways WT Cas9 DSB C-NHEJ Resection 3 Locus Unaltered
Effect of Different CRISPR/Cas9 Variants on Repair Pathway Choice Cecilia Cotta-Ramusino www.editasmedicine.com 1 Cas9 Stimulates the Endogenous Repair Pathways WT Cas9 DSB C-NHEJ Resection 3 Locus Unaltered
Supplementary Figure S1. The tetracycline-inducible CRISPR system. A) Hela cells stably
 Supplementary Information Supplementary Figure S1. The tetracycline-inducible CRISPR system. A) Hela cells stably expressing shrna sequences against TRF2 were examined by western blotting. shcon, shrna
Supplementary Information Supplementary Figure S1. The tetracycline-inducible CRISPR system. A) Hela cells stably expressing shrna sequences against TRF2 were examined by western blotting. shcon, shrna
Homology-directed repair with Dharmacon Edit-R CRISPR-Cas9 reagents and single-stranded DNA oligos
 APPLICATION NOTE Homology-directed repair with Dharmacon Edit-R CRISPR- reagents and single-stranded DNA oligos John A. Schiel, Eldon Chou, Maren Mayer, Emily M. Anderson, and Anja van Brabant Smith Horizon
APPLICATION NOTE Homology-directed repair with Dharmacon Edit-R CRISPR- reagents and single-stranded DNA oligos John A. Schiel, Eldon Chou, Maren Mayer, Emily M. Anderson, and Anja van Brabant Smith Horizon
Genome edi3ng with the CRISPR-Cas9 system
 CRISPR-Cas9 Genome Edi3ng Bootcamp AHA Council on Func3onal Genomics and Transla3onal Biology Narrated video link: hfps://youtu.be/h18hmftybnq Genome edi3ng with the CRISPR-Cas9 system Kiran Musunuru,
CRISPR-Cas9 Genome Edi3ng Bootcamp AHA Council on Func3onal Genomics and Transla3onal Biology Narrated video link: hfps://youtu.be/h18hmftybnq Genome edi3ng with the CRISPR-Cas9 system Kiran Musunuru,
Amplicons, Heteroduplexes and Enzymes - Proper Processing Elevates Detection of CRISPR Gene Editing Events
 Amplicons, Heteroduplexes and Enzymes - Proper Processing Elevates Detection of CRISPR Gene Editing Events Steve Siembieda, MS MBA VP Commercialization ABRF Conference February 2015 What Is CRISPR? Clustered
Amplicons, Heteroduplexes and Enzymes - Proper Processing Elevates Detection of CRISPR Gene Editing Events Steve Siembieda, MS MBA VP Commercialization ABRF Conference February 2015 What Is CRISPR? Clustered
Genome Editing: Considerations for Therapeutic Applications Cecilia Fernandez TIDES,
 Genome Editing: Considerations for Therapeutic Applications Cecilia Fernandez TIDES, 2017 www.editasmedicine.com 1 The Potential to Repair Any Broken Gene Transformative new category of medicines for genetically-defined
Genome Editing: Considerations for Therapeutic Applications Cecilia Fernandez TIDES, 2017 www.editasmedicine.com 1 The Potential to Repair Any Broken Gene Transformative new category of medicines for genetically-defined
Optimized, chemically-modified crrna:tracrrna complexes for CRISPR gene editing
 Optimized, chemically-modified crrna:tracrrna complexes for CRISPR gene editing Mark Behlke MD, PhD Chief Scientific Officer February 24, 2016 1 Implementing CRISPR/Cas9 gene editing 2 To focus on RNA
Optimized, chemically-modified crrna:tracrrna complexes for CRISPR gene editing Mark Behlke MD, PhD Chief Scientific Officer February 24, 2016 1 Implementing CRISPR/Cas9 gene editing 2 To focus on RNA
Cationic lipid-mediated delivery of proteins enables efficient proteinbased genome editing in vitro and in vivo
 correction notice Nat. Biotechnol. 33, 73 8 (215) Cationic lipid-mediated delivery of proteins enables efficient proteinbased genome editing in vitro and in vivo John A Zuris, David B Thompson, Yilai Shu,
correction notice Nat. Biotechnol. 33, 73 8 (215) Cationic lipid-mediated delivery of proteins enables efficient proteinbased genome editing in vitro and in vivo John A Zuris, David B Thompson, Yilai Shu,
Getting started with CRISPR: A review of gene knockout and homology-directed repair
 Getting started with CRISPR: A review of gene knockout and homology-directed repair Justin Barr Product Manager, Functional Genomics Integrated DNA Technologies 1 Agenda: Getting started with CRISPR Background
Getting started with CRISPR: A review of gene knockout and homology-directed repair Justin Barr Product Manager, Functional Genomics Integrated DNA Technologies 1 Agenda: Getting started with CRISPR Background
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/3/146/ra80/dc1 Supplementary Materials for DNMT1 Stability Is Regulated by Proteins Coordinating Deubiquitination and Acetylation-Driven Ubiquitination Zhanwen
www.sciencesignaling.org/cgi/content/full/3/146/ra80/dc1 Supplementary Materials for DNMT1 Stability Is Regulated by Proteins Coordinating Deubiquitination and Acetylation-Driven Ubiquitination Zhanwen
CRISPR Genome Editing: Considerations for Therapeutic Applications
 CRISPR Genome Editing: Considerations for Therapeutic Applications November 9, 2017 Cecilia Fernandez editasmedicine.com 1 Medicines that Aim to Repair Any Broken Gene Potential to create the next major
CRISPR Genome Editing: Considerations for Therapeutic Applications November 9, 2017 Cecilia Fernandez editasmedicine.com 1 Medicines that Aim to Repair Any Broken Gene Potential to create the next major
Application Note: Generating GFP-Tagged Human CD81 Tetraspanin Protein Using SBI s PrecisionX SmartNuclease System And HR Tagging Vectors
 Application Note: Generating GFP-Tagged Human CD81 Tetraspanin Protein Using SBI s PrecisionX SmartNuclease System And HR Tagging Vectors Table of Contents: I. Background Pg. 1 II. Analysis of Gene Target
Application Note: Generating GFP-Tagged Human CD81 Tetraspanin Protein Using SBI s PrecisionX SmartNuclease System And HR Tagging Vectors Table of Contents: I. Background Pg. 1 II. Analysis of Gene Target
Supplementary Figure 1. Supporting data for Figure 1
 Supplementary Figure 1 Supporting data for Figure 1 (A) Schematic of BLI assay used to measure dissociation. 5 monobiotinylated substrate DNA (identical to λ1, Figure 2) is associated with streptavidin-coated
Supplementary Figure 1 Supporting data for Figure 1 (A) Schematic of BLI assay used to measure dissociation. 5 monobiotinylated substrate DNA (identical to λ1, Figure 2) is associated with streptavidin-coated
Supplementary Figure 1. Quantitative RT-PCR experimental validation of CRISPR/Cas9 and sgrnas expression in HEK293A transfected cells.
 Supplementary Figure 1. Quantitative RT-PCR experimental validation of CRISPR/Cas9 and sgrnas expression in HEK293A transfected cells. HEK293A cells were transfected with the indicated combinations of
Supplementary Figure 1. Quantitative RT-PCR experimental validation of CRISPR/Cas9 and sgrnas expression in HEK293A transfected cells. HEK293A cells were transfected with the indicated combinations of
Designing CRISPR mediated Gene disruptions with gblocks Gene Fragments
 Designing CRISPR mediated Gene disruptions with gblocks Gene Fragments Adam Clore, PhD Manager, Synthetic Biology Design Integrated DNA technology Typical CRISPR timeline in Mammalian cell lines Design
Designing CRISPR mediated Gene disruptions with gblocks Gene Fragments Adam Clore, PhD Manager, Synthetic Biology Design Integrated DNA technology Typical CRISPR timeline in Mammalian cell lines Design
Incorporating SeqStudio Genetic Analyzer and Sanger sequencing into genome editing workflows
 Incorporating SeqStudio Genetic Analyzer and Sanger sequencing into genome editing workflows Stephen Jackson, Ph.D 27 May 2017 The world leader in serving science Key Applications for Genome Editing Research
Incorporating SeqStudio Genetic Analyzer and Sanger sequencing into genome editing workflows Stephen Jackson, Ph.D 27 May 2017 The world leader in serving science Key Applications for Genome Editing Research
CRISPR/Cas9: Tools and Applications for Eukaryotic Genome Editing
 CRISPR/Cas9: Tools and Applications for Eukaryotic Genome Editing Fei Ann Ran Broad Institute Cambridge, Massachusetts ran@fas.harvard.edu I will provide some background on the CRISPR/Cas9 technology,
CRISPR/Cas9: Tools and Applications for Eukaryotic Genome Editing Fei Ann Ran Broad Institute Cambridge, Massachusetts ran@fas.harvard.edu I will provide some background on the CRISPR/Cas9 technology,
Targeted complete next generation sequencing and quality control of transgenes and integration sites in CHO cell line development
 Targeted Locus Amplification Technology Targeted complete next generation sequencing and quality control of transgenes and integration sites in CHO cell line development Cergentis B.V. Yalelaan 62 3584
Targeted Locus Amplification Technology Targeted complete next generation sequencing and quality control of transgenes and integration sites in CHO cell line development Cergentis B.V. Yalelaan 62 3584
ISU CBC CRISPR Workshop. Michael Collingwood & Mollie Schubert
 ISU CBC CRISPR Workshop Michael Collingwood & Mollie Schubert 1 Genome editing (Cas12a) 2 Alt-R CRISPR System complete workflow Design Cut Repair Analyze Cas9/Cas12a + Guide RNA (grna) 3 CRISPR-Cas9 genome
ISU CBC CRISPR Workshop Michael Collingwood & Mollie Schubert 1 Genome editing (Cas12a) 2 Alt-R CRISPR System complete workflow Design Cut Repair Analyze Cas9/Cas12a + Guide RNA (grna) 3 CRISPR-Cas9 genome
Schematic representation of the endogenous PALB2 locus and gene-disruption constructs
 Supplementary Figures Supplementary Figure 1. Generation of PALB2 -/- and BRCA2 -/- /PALB2 -/- DT40 cells. (A) Schematic representation of the endogenous PALB2 locus and gene-disruption constructs carrying
Supplementary Figures Supplementary Figure 1. Generation of PALB2 -/- and BRCA2 -/- /PALB2 -/- DT40 cells. (A) Schematic representation of the endogenous PALB2 locus and gene-disruption constructs carrying
Supplementary Figure 1. Isolation of GFPHigh cells.
 Supplementary Figure 1. Isolation of GFP High cells. (A) Schematic diagram of cell isolation based on Wnt signaling activity. Colorectal cancer (CRC) cell lines were stably transduced with lentivirus encoding
Supplementary Figure 1. Isolation of GFP High cells. (A) Schematic diagram of cell isolation based on Wnt signaling activity. Colorectal cancer (CRC) cell lines were stably transduced with lentivirus encoding
Supplementary Materials for
 advances.sciencemag.org/cgi/content/full/3/4/e1602814/dc1 Supplementary Materials for CRISPR-Cpf1 correction of muscular dystrophy mutations in human cardiomyocytes and mice Yu Zhang, Chengzu Long, Hui
advances.sciencemag.org/cgi/content/full/3/4/e1602814/dc1 Supplementary Materials for CRISPR-Cpf1 correction of muscular dystrophy mutations in human cardiomyocytes and mice Yu Zhang, Chengzu Long, Hui
A novel tool for monitoring endogenous alpha-synuclein transcription by NanoLuciferase
 A novel tool for monitoring endogenous alpha-synuclein transcription by NanoLuciferase tag insertion at the 3 end using CRISPR-Cas9 genome editing technique Sambuddha Basu 1, 3, Levi Adams 1, 3, Subhrangshu
A novel tool for monitoring endogenous alpha-synuclein transcription by NanoLuciferase tag insertion at the 3 end using CRISPR-Cas9 genome editing technique Sambuddha Basu 1, 3, Levi Adams 1, 3, Subhrangshu
Webinar: Genome Editing using Nucleofector Technology
 Pharma&Biotech BioResearch Webinar: Genome Editing using Nucleofector Technology Efficient Transfection of ZFN, TALEN and CRISPR/Cas In collaboration with: Lonza Walkersville, Inc., Walkersville, MD 21793
Pharma&Biotech BioResearch Webinar: Genome Editing using Nucleofector Technology Efficient Transfection of ZFN, TALEN and CRISPR/Cas In collaboration with: Lonza Walkersville, Inc., Walkersville, MD 21793
SURVEY AND SUMMARY How to create state-of-the-art genetic model systems: strategies for optimal CRISPR-mediated genome editing
 Published online 28 June 2018 Nucleic Acids Research, 2018, Vol. 46, No. 13 6435 6454 doi: 10.1093/nar/gky571 SURVEY AND SUMMARY How to create state-of-the-art genetic model systems: strategies for optimal
Published online 28 June 2018 Nucleic Acids Research, 2018, Vol. 46, No. 13 6435 6454 doi: 10.1093/nar/gky571 SURVEY AND SUMMARY How to create state-of-the-art genetic model systems: strategies for optimal
Donor DNA Utilization during Gene Targeting with Zinc- finger Nucleases
 Donor DNA Utilization during Gene Targeting with Zinc- finger Nucleases Kelly J. Beumer, Jonathan K. Trautman, Kusumika Mukherjee and Dana Carroll Department of Biochemistry University of Utah School of
Donor DNA Utilization during Gene Targeting with Zinc- finger Nucleases Kelly J. Beumer, Jonathan K. Trautman, Kusumika Mukherjee and Dana Carroll Department of Biochemistry University of Utah School of
CRISPR: A Simple Tool for Answering Complex Questions
 CRISPR: A Simple Tool for Answering Complex Questions www.bcm.edu/cbass c_bass@bcm.edu 713-798-8987 CRISPR-Cas systems CRISPRs: Clustered Regularly Interspaced Short Palindromic Repeats Cas: The accompanying
CRISPR: A Simple Tool for Answering Complex Questions www.bcm.edu/cbass c_bass@bcm.edu 713-798-8987 CRISPR-Cas systems CRISPRs: Clustered Regularly Interspaced Short Palindromic Repeats Cas: The accompanying
Genome Editing: Cas9 Stable Cell Lines for CRISPR sgrna Validation, Library Screening, and More. Ed Davis, Ph.D.
 TECHNICAL NOTE Genome Editing: Cas9 Stable Cell Lines for CRISPR sgrna Validation, Library Screening, and More Introduction Ed Davis, Ph.D. The CRISPR-Cas9 system has become greatly popular for genome
TECHNICAL NOTE Genome Editing: Cas9 Stable Cell Lines for CRISPR sgrna Validation, Library Screening, and More Introduction Ed Davis, Ph.D. The CRISPR-Cas9 system has become greatly popular for genome
Supplementary Figure 1
 Supplementary Figure 1 Product purity of cytosine base editing for the wheat genomic loci tested. Product distributions and indels frequencies at four representative wheat genomic DNA sites in wheat protoplasts
Supplementary Figure 1 Product purity of cytosine base editing for the wheat genomic loci tested. Product distributions and indels frequencies at four representative wheat genomic DNA sites in wheat protoplasts
Protocols for cell lines using CRISPR/CAS
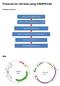 Protocols for cell lines using CRISPR/CAS Procedure overview Map Preparation of CRISPR/CAS plasmids Expression vectors for guide RNA (U6-gRNA) and Cas9 gene (CMV-p-Cas9) are ampicillin-resist ant and stable
Protocols for cell lines using CRISPR/CAS Procedure overview Map Preparation of CRISPR/CAS plasmids Expression vectors for guide RNA (U6-gRNA) and Cas9 gene (CMV-p-Cas9) are ampicillin-resist ant and stable
A novel two-step genome editing strategy with CRISPR-Cas9 provides new insights into telomerase action and TERT gene expression
 Xi et al. Genome Biology (2015) 16:231 DOI 10.1186/s13059-015-0791-1 RESEARCH A novel two-step genome editing strategy with CRISPR-Cas9 provides new insights into telomerase action and TERT gene expression
Xi et al. Genome Biology (2015) 16:231 DOI 10.1186/s13059-015-0791-1 RESEARCH A novel two-step genome editing strategy with CRISPR-Cas9 provides new insights into telomerase action and TERT gene expression
A Guide to CRISPR/Cas9
 Genome editing and beyond freepik A Guide to CRISPR/Cas9 The latest advance in genomic DNA editing is the Clustered Regularly Interspaced Short Palindromic Repeat (CRISPR)/Cas9 system. This simple-touse
Genome editing and beyond freepik A Guide to CRISPR/Cas9 The latest advance in genomic DNA editing is the Clustered Regularly Interspaced Short Palindromic Repeat (CRISPR)/Cas9 system. This simple-touse
XactEdit Cas9 Nuclease with NLS User Manual
 XactEdit Cas9 Nuclease with NLS User Manual An RNA-guided recombinant endonuclease for efficient targeted DNA cleavage Catalog Numbers CE1000-50K, CE1000-50, CE1000-250, CE1001-250, CE1001-1000 Table of
XactEdit Cas9 Nuclease with NLS User Manual An RNA-guided recombinant endonuclease for efficient targeted DNA cleavage Catalog Numbers CE1000-50K, CE1000-50, CE1000-250, CE1001-250, CE1001-1000 Table of
Eric Paul Bennett, MSc, Dr.med. Cellular Indel Profiles and Dynamics CRISPR/Cas9 delivery formats: Applying in vitro methodologies ex vivo
 Copenhagen Center for Glycomics/Department of Odontology Eric Paul Bennett, MSc, Dr.med. Cellular Indel Profiles and Dynamics CRISPR/Cas9 delivery formats: Applying in vitro methodologies ex vivo Eric
Copenhagen Center for Glycomics/Department of Odontology Eric Paul Bennett, MSc, Dr.med. Cellular Indel Profiles and Dynamics CRISPR/Cas9 delivery formats: Applying in vitro methodologies ex vivo Eric
Generating stable cell lines expressing various Cas9 proteins from AAVS1 locus
 IGI Protocol prepared by Chong Park 1 Generating stable cell lines expressing various Cas9 proteins from AAVS1 locus This is modified from original protocol from the Conklin laboratory (Mandegar et al.
IGI Protocol prepared by Chong Park 1 Generating stable cell lines expressing various Cas9 proteins from AAVS1 locus This is modified from original protocol from the Conklin laboratory (Mandegar et al.
Isolation of single-base genome-edited human ips cells without
 Nature Methods Isolation of single-base genome-edited human ips cells without antibiotic selection Yuichiro Miyaoka, Amanda H. Chan, Luke M. Judge, Jennie Yoo, Miller Huang, Trieu D. Nguyen, Paweena P.
Nature Methods Isolation of single-base genome-edited human ips cells without antibiotic selection Yuichiro Miyaoka, Amanda H. Chan, Luke M. Judge, Jennie Yoo, Miller Huang, Trieu D. Nguyen, Paweena P.
Characterization and Optimization of the CRISPR/Cas System for Applications in Genome Engineering
 Characterization and Optimization of the CRISPR/Cas System for Applications in Genome Engineering The Harvard community has made this article openly available. Please share how this access benefits you.
Characterization and Optimization of the CRISPR/Cas System for Applications in Genome Engineering The Harvard community has made this article openly available. Please share how this access benefits you.
Cpf1 proteins excise CRISPR RNAs from mrna transcripts in mammalian cells
 SUPPLEMENTARY INFORMATION Cpf1 proteins exise CRISPR RNAs from mrna transripts in mammalian ells Guoai Zhong 1, Haimin Wang 1, Yujun Li 1, 2, Mai H. Tran 1, Mihael Farzan 1 1 Department of Immunology and
SUPPLEMENTARY INFORMATION Cpf1 proteins exise CRISPR RNAs from mrna transripts in mammalian ells Guoai Zhong 1, Haimin Wang 1, Yujun Li 1, 2, Mai H. Tran 1, Mihael Farzan 1 1 Department of Immunology and
Somatic Primary pirna Biogenesis Driven by cis-acting RNA Elements and Trans-Acting Yb
 Cell Reports Supplemental Information Somatic Primary pirna Biogenesis Driven by cis-acting RNA Elements and Trans-Acting Yb Hirotsugu Ishizu, Yuka W. Iwasaki, Shigeki Hirakata, Haruka Ozaki, Wataru Iwasaki,
Cell Reports Supplemental Information Somatic Primary pirna Biogenesis Driven by cis-acting RNA Elements and Trans-Acting Yb Hirotsugu Ishizu, Yuka W. Iwasaki, Shigeki Hirakata, Haruka Ozaki, Wataru Iwasaki,
Inhibition of HSV-1 Replication by Gene Editing Strategy. Running Title: Targeting HSV-1 Infection by CRISPR/Cas9
 SUPPLEMENTARY MATERIALS 2/4/16 Inhibition of HSV-1 Replication by Gene Editing Strategy Running Title: Targeting HSV-1 Infection by CRISPR/Cas9 Pamela C. Roehm 1,2,3*, Masoud Shekarabi 1, Hassen Wollebo
SUPPLEMENTARY MATERIALS 2/4/16 Inhibition of HSV-1 Replication by Gene Editing Strategy Running Title: Targeting HSV-1 Infection by CRISPR/Cas9 Pamela C. Roehm 1,2,3*, Masoud Shekarabi 1, Hassen Wollebo
Supplementary Figures Montero et al._supplementary Figure 1
 Montero et al_suppl. Info 1 Supplementary Figures Montero et al._supplementary Figure 1 Montero et al_suppl. Info 2 Supplementary Figure 1. Transcripts arising from the structurally conserved subtelomeres
Montero et al_suppl. Info 1 Supplementary Figures Montero et al._supplementary Figure 1 Montero et al_suppl. Info 2 Supplementary Figure 1. Transcripts arising from the structurally conserved subtelomeres
Supplementary Information
 Supplementary Information Supplementary Figure 1: Synthetic annealed poly(ra):poly(dt) hybrid induces type I interferon response. a) Bone marrow derived macrophages (BMDM) were transfected with or without
Supplementary Information Supplementary Figure 1: Synthetic annealed poly(ra):poly(dt) hybrid induces type I interferon response. a) Bone marrow derived macrophages (BMDM) were transfected with or without
A CRISPR/Cas9 Vector System for Tissue-Specific Gene Disruption in Zebrafish
 Developmental Cell Supplemental Information A CRISPR/Cas9 Vector System for Tissue-Specific Gene Disruption in Zebrafish Julien Ablain, Ellen M. Durand, Song Yang, Yi Zhou, and Leonard I. Zon % larvae
Developmental Cell Supplemental Information A CRISPR/Cas9 Vector System for Tissue-Specific Gene Disruption in Zebrafish Julien Ablain, Ellen M. Durand, Song Yang, Yi Zhou, and Leonard I. Zon % larvae
CRISPR 101: Optimizing Your Gene Editing Experiments
 CRISPR 101: Optimizing Your Gene Editing Experiments PRESENTER Michele Auldridge, Ph.D. Senior Scientist, R&D MODERATOR Beth Frey Product Manager Agenda 1 Technology Overview 2 Experimental Decisions 3
CRISPR 101: Optimizing Your Gene Editing Experiments PRESENTER Michele Auldridge, Ph.D. Senior Scientist, R&D MODERATOR Beth Frey Product Manager Agenda 1 Technology Overview 2 Experimental Decisions 3
Guide-it sgrna In Vitro Transcription and Screening Systems User Manual
 Guide-it sgrna In Vitro Transcription and Screening Systems User Manual Cat. Nos. 632638, 632639, 632635, 632636, 632637 (040618) 1290 Terra Bella Avenue, Mountain View, CA 94043, USA U.S. Technical Support:
Guide-it sgrna In Vitro Transcription and Screening Systems User Manual Cat. Nos. 632638, 632639, 632635, 632636, 632637 (040618) 1290 Terra Bella Avenue, Mountain View, CA 94043, USA U.S. Technical Support:
CRISPR GENOMIC SERVICES PRODUCT CATALOG
 CRISPR GENOMIC SERVICES PRODUCT CATALOG DESIGN BUILD ANALYZE The experts at Desktop Genetics can help you design, prepare and manufacture all of the components needed for your CRISPR screen. We provide
CRISPR GENOMIC SERVICES PRODUCT CATALOG DESIGN BUILD ANALYZE The experts at Desktop Genetics can help you design, prepare and manufacture all of the components needed for your CRISPR screen. We provide
TECHNOLOGIES. 3/19/18 Kayla Nygaard. https://i.ytimg.com/vi/pxw5yya-kh0/maxresdefault.jpg
 TECHNOLOGIES 3/19/18 Kayla Nygaard https://i.ytimg.com/vi/pxw5yya-kh0/maxresdefault.jpg CRISPR IN THE NEWS CRISPR in 2018: Coming to a Human Near You Sickle-cell treatment clinical trials CRISPR Therapeutics
TECHNOLOGIES 3/19/18 Kayla Nygaard https://i.ytimg.com/vi/pxw5yya-kh0/maxresdefault.jpg CRISPR IN THE NEWS CRISPR in 2018: Coming to a Human Near You Sickle-cell treatment clinical trials CRISPR Therapeutics
Supplementary Figures and Figure legends
 Supplementary Figures and Figure legends 3 Supplementary Figure 1. Conditional targeting construct for the murine Satb1 locus with a modified FLEX switch. Schematic of the wild type Satb1 locus; the conditional
Supplementary Figures and Figure legends 3 Supplementary Figure 1. Conditional targeting construct for the murine Satb1 locus with a modified FLEX switch. Schematic of the wild type Satb1 locus; the conditional
SUPPLEMENTAL MATERIALS
 SUPPLEMENL MERILS Eh-seq: RISPR epitope tagging hip-seq of DN-binding proteins Daniel Savic, E. hristopher Partridge, Kimberly M. Newberry, Sophia. Smith, Sarah K. Meadows, rian S. Roberts, Mark Mackiewicz,
SUPPLEMENL MERILS Eh-seq: RISPR epitope tagging hip-seq of DN-binding proteins Daniel Savic, E. hristopher Partridge, Kimberly M. Newberry, Sophia. Smith, Sarah K. Meadows, rian S. Roberts, Mark Mackiewicz,
Introduction and History of Genome Modification. Adam Clore, PhD Director, Synthetic Biology Design
 Introduction and History of Genome Modification Adam Clore, PhD Director, Synthetic Biology Design Early Non-site Directed Genome Modification Homologous recombination in yeast TARGET GENE 5 Arm URA3 3
Introduction and History of Genome Modification Adam Clore, PhD Director, Synthetic Biology Design Early Non-site Directed Genome Modification Homologous recombination in yeast TARGET GENE 5 Arm URA3 3
Reprogramming MHC specificity by CRISPR-Cas9-assisted cassette exchange
 1 Supplementary Materials Reprogramming MHC specificity by CRISPR-Cas9-assisted cassette exchange 3 4 5 6 Authors William Kelton 1, Ann Cathrin Waindok 1, Theresa Pesch 1, Mark Pogson 1, Kyle Ford 1, Cristina
1 Supplementary Materials Reprogramming MHC specificity by CRISPR-Cas9-assisted cassette exchange 3 4 5 6 Authors William Kelton 1, Ann Cathrin Waindok 1, Theresa Pesch 1, Mark Pogson 1, Kyle Ford 1, Cristina
Mouse Engineering Technology. Musculoskeletal Research Center 2016 Summer Educational Series David M. Ornitz Department of Developmental Biology
 Mouse Engineering Technology Musculoskeletal Research Center 2016 Summer Educational Series David M. Ornitz Department of Developmental Biology Core service and new technologies Mouse ES core Discussions
Mouse Engineering Technology Musculoskeletal Research Center 2016 Summer Educational Series David M. Ornitz Department of Developmental Biology Core service and new technologies Mouse ES core Discussions
Targeted Protein Degradation
 The PITCh dtag vectors for rapid protein degradation of endogenous targets The dissection of complex biological systems requires target-specific control of protein function or abundance. Genetic perturbations
The PITCh dtag vectors for rapid protein degradation of endogenous targets The dissection of complex biological systems requires target-specific control of protein function or abundance. Genetic perturbations
Rescue of high-specificity Cas9 variants using sgrnas with matched 5 nucleotides
 Kim et al. Genome Biology (2017) 18:218 DOI 10.1186/s13059-017-1355-3 METHOD Rescue of high-specificity Cas9 variants using sgrnas with matched 5 nucleotides Sojung Kim 1, Taegeun Bae 1,2, Jaewoong Hwang
Kim et al. Genome Biology (2017) 18:218 DOI 10.1186/s13059-017-1355-3 METHOD Rescue of high-specificity Cas9 variants using sgrnas with matched 5 nucleotides Sojung Kim 1, Taegeun Bae 1,2, Jaewoong Hwang
Supplementary Fig. 1 Proteomic analysis of ATR-interacting proteins. ATR, ARID1A and
 Supplementary Figure Legend: Supplementary Fig. 1 Proteomic analysis of ATR-interacting proteins. ATR, ARID1A and ATRIP protein peptides identified from our mass spectrum analysis were shown. Supplementary
Supplementary Figure Legend: Supplementary Fig. 1 Proteomic analysis of ATR-interacting proteins. ATR, ARID1A and ATRIP protein peptides identified from our mass spectrum analysis were shown. Supplementary
CRISPR: hot, hot, hot
 CRISPR: hot, hot, hot 166 CRISPR is the latest technique for genome engineering and is generating tons of excitement due to its versatility, high specificity, and ease of use. CRISPR stands for clustered
CRISPR: hot, hot, hot 166 CRISPR is the latest technique for genome engineering and is generating tons of excitement due to its versatility, high specificity, and ease of use. CRISPR stands for clustered
Nature Biotechnology: doi: /nbt.4198
 Supplementary Figure 1 Effect of DNA methylation on the base editing efficiency induced by BE3. (a) DNMT3-induced DNA methylation leads to the decreased base editing efficiency induced by BE3. Top panels,
Supplementary Figure 1 Effect of DNA methylation on the base editing efficiency induced by BE3. (a) DNMT3-induced DNA methylation leads to the decreased base editing efficiency induced by BE3. Top panels,
CRISPR-dCas9 mediated TET1 targeting for selective DNA demethylation at BRCA1 promoter
 CRISPR-dCas9 mediated TET1 targeting for selective DNA demethylation at BRCA1 promoter SUPPLEMENTARY DATA See Supplementary Sequence File: 1 Supplementary Figure S1: The total protein was extracted from
CRISPR-dCas9 mediated TET1 targeting for selective DNA demethylation at BRCA1 promoter SUPPLEMENTARY DATA See Supplementary Sequence File: 1 Supplementary Figure S1: The total protein was extracted from
High-throughput genotyping of CRISPR/Cas9-mediated mutants using fluorescent
 High-throughput genotyping of CRISPR/Cas9-mediated mutants using fluorescent PCR-capillary gel electrophoresis Muhammad Khairul RAMLEE, Tingdong YAN, Alice M. S. CHEUNG, Charles CHUAH, Shang LI Figure
High-throughput genotyping of CRISPR/Cas9-mediated mutants using fluorescent PCR-capillary gel electrophoresis Muhammad Khairul RAMLEE, Tingdong YAN, Alice M. S. CHEUNG, Charles CHUAH, Shang LI Figure
Applications For CRISPR-Cas9 Stable Cell Lines
 Applications For CRISPR-Cas9 Stable Cell Lines Presenter: March 22, 2017 Ed Davis, Ph.D. Senior Application Scientist GeneCopoeia, Inc. GeneCopoeia products & services Functional Genomics & Cell Biology
Applications For CRISPR-Cas9 Stable Cell Lines Presenter: March 22, 2017 Ed Davis, Ph.D. Senior Application Scientist GeneCopoeia, Inc. GeneCopoeia products & services Functional Genomics & Cell Biology
Supplementary Materials for
 www.advances.sciencemag.org/cgi/content/full/1/7/e1500454/dc1 Supplementary Materials for CRISPR-Cas9 delivery to hard-to-transfect cells via membrane deformation Xin Han, Zongbin Liu, Myeong chan Jo,
www.advances.sciencemag.org/cgi/content/full/1/7/e1500454/dc1 Supplementary Materials for CRISPR-Cas9 delivery to hard-to-transfect cells via membrane deformation Xin Han, Zongbin Liu, Myeong chan Jo,
