Inhibition of HSV-1 Replication by Gene Editing Strategy. Running Title: Targeting HSV-1 Infection by CRISPR/Cas9
|
|
|
- Victor Neal
- 5 years ago
- Views:
Transcription
1 SUPPLEMENTARY MATERIALS 2/4/16 Inhibition of HSV-1 Replication by Gene Editing Strategy Running Title: Targeting HSV-1 Infection by CRISPR/Cas9 Pamela C. Roehm 1,2,3*, Masoud Shekarabi 1, Hassen Wollebo 1, Anna Bellizzi 1, Lifan He 1, Julian Salkind 1, Kamel Khalili 1* 1. Department of Neuroscience, Center for Neurovirology; 2. Department of Otolaryngology/Head and Neck Surgery; 3. Department of Neurosurgery; Temple University School of Medicine, 3500 N. Broad Street, Philadelphia, PA Keywords: CRISPR/Cas9, herpes simplex type 1, DNA editing, ICP0, PML Nuclear Body M.S. and H.W. contributed equally to this work. *Corresponding authors: Kamel Khalili (senior corresponding author) Pamela Roehm, Phone: /Fax:
2 Table S1: Primers (HSV-1) Primer Sequence P1 P2 P3 P4 P5 5'-GATGTCTGGGTGTTTCCCT-3' 5'-CACGATCGGGATGGTGCTGAACGACC-3' 5'- TGCTGATTGACGCGGGAAATC-3' 5'- CGATCTCATCCGTGCACACG-3' 5'-CCATCCACGCCCTGCGGCCCCAGC-3' 2
3 Table S2: Predicted ICP0 grnas and their off-target numbers (100% match) grna sequence grna 2A 1. TCCCTGCGACCGAGACCTGCCGG grna 2B 2. ACACGGAACTGTTCGAGACGGGG grna 2C 3.GCATCCCGTGCATGAAAACCTGG The genomic sequence of HSV-1 ICPO (NC_ ) was used to search for Cas9/gRNA target sites containing a 20-bp guide sequence plus the protospacer adjacent motif sequence (NGG) using Jack Lin's CRISPER/Cas9 finder tool( NGG(AGG,TGG,GGG,CGG) was blasted against available human genomic and transcript sequence with 1000 aligned sequences being displayed. After pressing Control+F, copy/paste the Target sequences (1-23 through 9-23 nucleotides) and find the number of genomic targets with 100% match. The number of off-targets for each searching was divided by 3 because of repeated genome library. The number shown indicates the sum of four search (NGG). The red nubmer indicates the grnas target sequences farthest from NGG 3
4 Table S3: Primers (cellular genes) Primer off-1 (G32) off-2 (G42) off-3 (G22) off-4 (G12) off-5 (G6) Sequence 5'-CACGTCTGTGGGTGTTCAGA-3' 5'-AAAGCTCCTAGGGCTGTTGG-3 5'-GGAACCCCCTCGCTTTCTTT-3' 5'-CGTATGTGTCCCATCGGCAT-3' 5'-CACTGCCTTGTGCCCATAGA-3' 5'-TTGAAGCTGCACGTGTTGGA-3' 5'-ACTGACGGGATCTGGGATCA-3' 5'-TTCACTGGGAGACGTCAAGC-3' 5'-GTGAGGCCCCACACATGATT-3' 5'-AAGGATGAGAGATGGGGAAGC-3' 4
5 Table S4: Sequence of guide RNAs targeting HSV genes Target genes ICP4 m1 ICP27 m1 Sequence 5'-TTCTACGCGCGCTATCGCGACGG-3' 5'-GGAGTGTTCCTCGTCGGACGAGG-3' 5
6 SUPPLEMENTARY FIGURE LEGENDS Figure S1. A. Chromatograms showing the detection of breakpoints within exon II as a result of cleavage of DNA by Cas9 and grnas 2A and 2C, and insertion of a single nucleotide A (top) and G (bottom) at the site of the breakpoint. B. Recombination of grna 2B and 2C cuts within exon II generating a new template for amplification of a 251 bp DNA. C. SURVEYOR assay showing InDel mutation in exon II at target A causing a new DNA fragment of 180 bp after cleavage with Cel1 nuclease. Figure S2. A. Production of grna 2A in L7 cells (control) expressing Cas9 and Cas9 plus grna 2A by RT-PCR. β-actin transcripts served as a loading control. B. Expression of flag-cas9 in TC620 cells were verified by Western blot analysis. Figure S3. A. TC620, a human olidodendroglioma cell line, were infected with HSV-1/GFP at MOI of 0.1, 1 or 2, and 48 hours later, expression of GFP, indicative for HSV-1 expression and replication, was assessed by fluorescence microscopy. B. Western blot analysis for evaluating expression of ICP0 in TC620 cells infected with HSV-1 at 48 hours post-infection. C and D. Detection of ICP8 and glycoprotein C in infected TC620 cells at MOI=1, 48 hours post-infection by Western blot analysis. Figure S4. A. Cell cycle assay was used to investigate the impact of Cas9/gRNA on the cell cycle of TC620 control cells has no expression of Cas9 or any grnas, whereas subclones of Cas9 and Cas9/gRNA 2A clones 6 and 10 express either Cas9 alone or Cas9 plus grna2a as indicated. As seen, no drastic differences in cell cycle progression through G1, S and G2/M phases were detected in these cell lines. B. Apoptotic assay in the clonal cell lines as shown in Panel A showed no significant changes in annexin stained cell population (ranging from 1.0-6
7 2.1%). C. Cell viability assay using propidium iodide staining assay showed changes in the viability of the cells with Cas9 or Cas9/gRNA expression in the various clones. Figure S5. A. DNA gel analysis of SURVEYOR assay illustrating no detectable bands below the expected DNA fragment from PCR amplification that can be attributed to grna 2A expression. Control is provided by the supplier of the SURVEYOR assay kit (Integrated DNA Technologies, Coralville, IA). B. Results from PCR amplification and Sanger sequencing of the cellular genes that contained potential off-targets (as shown in Panel A) revealed no InDel mutations in the amplicons. Arrows demonstrate the positions of the primers and their distance from the potential off-targets. Red indicates the PAM sequence and the block illustrate the potential target sequence for each cellular gene. The size of each amplicon is indicated. Figure S6. Epifluorescent assessment of HSV-1/GFP replication in TC260 cells after treatment with lentivirus vector (LV) expressing Cas9 (Panel A) or Cas9 plus various grnas as shown above each panel according to the method described for Fig. 4E. Images are representative of duplicate plates for each condition. 7
8
9
10
11
12
13
Simple protocol for gene editing using GenCrisprTM Cas9 nuclease
 Simple protocol for gene editing using GenCrisprTM Cas9 nuclease Contents Protocol Step 1: Choose the target DNA sequence Step 2: Design sgrna Step 3: Preparation for sgrna 3.1 In vitro transcription of
Simple protocol for gene editing using GenCrisprTM Cas9 nuclease Contents Protocol Step 1: Choose the target DNA sequence Step 2: Design sgrna Step 3: Preparation for sgrna 3.1 In vitro transcription of
Transfection of CRISPR/Cas9 Nuclease NLS ribonucleoprotein (RNP) into adherent mammalian cells using Lipofectamine RNAiMAX
 Transfection of CRISPR/Cas9 Nuclease NLS ribonucleoprotein (RNP) into adherent mammalian cells using Lipofectamine RNAiMAX INTRODUCTION The CRISPR/Cas genome editing system consists of a single guide RNA
Transfection of CRISPR/Cas9 Nuclease NLS ribonucleoprotein (RNP) into adherent mammalian cells using Lipofectamine RNAiMAX INTRODUCTION The CRISPR/Cas genome editing system consists of a single guide RNA
High-throughput genotyping of CRISPR/Cas9-mediated mutants using fluorescent
 High-throughput genotyping of CRISPR/Cas9-mediated mutants using fluorescent PCR-capillary gel electrophoresis Muhammad Khairul RAMLEE, Tingdong YAN, Alice M. S. CHEUNG, Charles CHUAH, Shang LI Figure
High-throughput genotyping of CRISPR/Cas9-mediated mutants using fluorescent PCR-capillary gel electrophoresis Muhammad Khairul RAMLEE, Tingdong YAN, Alice M. S. CHEUNG, Charles CHUAH, Shang LI Figure
Nature Immunology: doi: /ni Supplementary Figure 1
 Supplementary Figure 1 Sox2 localizes in neutrophils of mouse spleen. (a) Microscopy analysis of Sox2 and MPO in wild-type mouse spleen. Nucl, nucleus. (b) Immunohistochemistry staining of Sox2 and MPO
Supplementary Figure 1 Sox2 localizes in neutrophils of mouse spleen. (a) Microscopy analysis of Sox2 and MPO in wild-type mouse spleen. Nucl, nucleus. (b) Immunohistochemistry staining of Sox2 and MPO
XactEdit Cas9 Nuclease with NLS User Manual
 XactEdit Cas9 Nuclease with NLS User Manual An RNA-guided recombinant endonuclease for efficient targeted DNA cleavage Catalog Numbers CE1000-50K, CE1000-50, CE1000-250, CE1001-250, CE1001-1000 Table of
XactEdit Cas9 Nuclease with NLS User Manual An RNA-guided recombinant endonuclease for efficient targeted DNA cleavage Catalog Numbers CE1000-50K, CE1000-50, CE1000-250, CE1001-250, CE1001-1000 Table of
Genome-wide genetic screening with chemically-mutagenized haploid embryonic stem cells
 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 Supplementary Information Genome-wide genetic screening with chemically-mutagenized haploid embryonic stem cells Josep V. Forment 1,2, Mareike Herzog
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 Supplementary Information Genome-wide genetic screening with chemically-mutagenized haploid embryonic stem cells Josep V. Forment 1,2, Mareike Herzog
A Guide to CRISPR/Cas9
 Genome editing and beyond freepik A Guide to CRISPR/Cas9 The latest advance in genomic DNA editing is the Clustered Regularly Interspaced Short Palindromic Repeat (CRISPR)/Cas9 system. This simple-touse
Genome editing and beyond freepik A Guide to CRISPR/Cas9 The latest advance in genomic DNA editing is the Clustered Regularly Interspaced Short Palindromic Repeat (CRISPR)/Cas9 system. This simple-touse
CRISPR/Cas9-mediated viral interference in plants
 CRISPR/Cas9mediated viral interference in plants Item Type Article Authors Ali, Zahir; Abulfaraj, Aala A.; Idris, Ali; Ali, Shawkat; Tashkandi, Manal; Mahfouz, Magdy M. Citation CRISPR/Cas9mediated viral
CRISPR/Cas9mediated viral interference in plants Item Type Article Authors Ali, Zahir; Abulfaraj, Aala A.; Idris, Ali; Ali, Shawkat; Tashkandi, Manal; Mahfouz, Magdy M. Citation CRISPR/Cas9mediated viral
Molecular Cell Biology - Problem Drill 11: Recombinant DNA
 Molecular Cell Biology - Problem Drill 11: Recombinant DNA Question No. 1 of 10 1. Which of the following statements about the sources of DNA used for molecular cloning is correct? Question #1 (A) cdna
Molecular Cell Biology - Problem Drill 11: Recombinant DNA Question No. 1 of 10 1. Which of the following statements about the sources of DNA used for molecular cloning is correct? Question #1 (A) cdna
Supplementary Information
 Supplementary Information Super-resolution imaging of fluorescently labeled, endogenous RNA Polymerase II in living cells with CRISPR/Cas9-mediated gene editing Won-Ki Cho 1, Namrata Jayanth 1, Susan Mullen
Supplementary Information Super-resolution imaging of fluorescently labeled, endogenous RNA Polymerase II in living cells with CRISPR/Cas9-mediated gene editing Won-Ki Cho 1, Namrata Jayanth 1, Susan Mullen
CRISPR Ribonucleoprotein (RNP) User Manual
 CRISPR Ribonucleoprotein (RNP) User Manual Description This user manual describes how to use GenScript s CRISPR Ribonucleoprotein (RNP) products for targeted genome editing. The RNP system is comprised
CRISPR Ribonucleoprotein (RNP) User Manual Description This user manual describes how to use GenScript s CRISPR Ribonucleoprotein (RNP) products for targeted genome editing. The RNP system is comprised
SUPPLEMENTARY INFORMATION
 Gene replacements and insertions in rice by intron targeting using CRISPR Cas9 Table of Contents Supplementary Figure 1. sgrna-induced targeted mutations in the OsEPSPS gene in rice protoplasts. Supplementary
Gene replacements and insertions in rice by intron targeting using CRISPR Cas9 Table of Contents Supplementary Figure 1. sgrna-induced targeted mutations in the OsEPSPS gene in rice protoplasts. Supplementary
Introduction to CRISPR/Cas9 Background DNA Cleavage and Repair (NHEJ and HDR) Alternative Cas9 Variants Delivery of Cas9 and sgrna Library Products
 Introduction to CRISPR/Cas9 Background DNA Cleavage and Repair (NHEJ and HDR) Alternative Cas9 Variants Delivery of Cas9 and sgrna Library Products which one is right for you? CRISPR Workflow abm s Toolbox
Introduction to CRISPR/Cas9 Background DNA Cleavage and Repair (NHEJ and HDR) Alternative Cas9 Variants Delivery of Cas9 and sgrna Library Products which one is right for you? CRISPR Workflow abm s Toolbox
Chapter 6 - Molecular Genetic Techniques
 Chapter 6 - Molecular Genetic Techniques Two objects of molecular & genetic technologies For analysis For generation Molecular genetic technologies! For analysis DNA gel electrophoresis Southern blotting
Chapter 6 - Molecular Genetic Techniques Two objects of molecular & genetic technologies For analysis For generation Molecular genetic technologies! For analysis DNA gel electrophoresis Southern blotting
CRISPR RNA-guided activation of endogenous human genes
 CRISPR RNA-guided activation of endogenous human genes Morgan L Maeder, Samantha J Linder, Vincent M Cascio, Yanfang Fu, Quan H Ho, J Keith Joung Supplementary Figure 1 Comparison of VEGF activation induced
CRISPR RNA-guided activation of endogenous human genes Morgan L Maeder, Samantha J Linder, Vincent M Cascio, Yanfang Fu, Quan H Ho, J Keith Joung Supplementary Figure 1 Comparison of VEGF activation induced
Supplementary Information
 Supplementary Information Vidigal and Ventura a wt locus 5 region 3 region CCTCTGCCACTGCGAGGGCGTCCAATGGTGCTTG(...)AACAGGTGGAATATCCCTACTCTA predicted deletion clone 1 clone 2 clone 3 CCTCTGCCACTGCGAGGGCGTC-AGGTGGAATATCCCTACTCTA
Supplementary Information Vidigal and Ventura a wt locus 5 region 3 region CCTCTGCCACTGCGAGGGCGTCCAATGGTGCTTG(...)AACAGGTGGAATATCCCTACTCTA predicted deletion clone 1 clone 2 clone 3 CCTCTGCCACTGCGAGGGCGTC-AGGTGGAATATCCCTACTCTA
User Instructions:Transfection-ready CRISPR/Cas9 Reagents. Target DNA. NHEJ repair pathway. Nucleotide deletion. Nucleotide insertion Gene disruption
 User Instructions:Transfection-ready CRISPR/Cas9 Reagents Background Introduction to CRISPR/Cas9 genome editing In bacteria and archaea, clustered regularly interspaced short palindromic repeats (CRISPR)
User Instructions:Transfection-ready CRISPR/Cas9 Reagents Background Introduction to CRISPR/Cas9 genome editing In bacteria and archaea, clustered regularly interspaced short palindromic repeats (CRISPR)
SUPPLEMENTAL MATERIALS
 SUPPLEMENL MERILS Eh-seq: RISPR epitope tagging hip-seq of DN-binding proteins Daniel Savic, E. hristopher Partridge, Kimberly M. Newberry, Sophia. Smith, Sarah K. Meadows, rian S. Roberts, Mark Mackiewicz,
SUPPLEMENL MERILS Eh-seq: RISPR epitope tagging hip-seq of DN-binding proteins Daniel Savic, E. hristopher Partridge, Kimberly M. Newberry, Sophia. Smith, Sarah K. Meadows, rian S. Roberts, Mark Mackiewicz,
Application Note: Generating GFP-Tagged Human CD81 Tetraspanin Protein Using SBI s PrecisionX SmartNuclease System And HR Tagging Vectors
 Application Note: Generating GFP-Tagged Human CD81 Tetraspanin Protein Using SBI s PrecisionX SmartNuclease System And HR Tagging Vectors Table of Contents: I. Background Pg. 1 II. Analysis of Gene Target
Application Note: Generating GFP-Tagged Human CD81 Tetraspanin Protein Using SBI s PrecisionX SmartNuclease System And HR Tagging Vectors Table of Contents: I. Background Pg. 1 II. Analysis of Gene Target
Supplementary Information. Optimization of the production of knock-in alleles by CRISPR/Cas9 microinjection into the mouse zygote
 Supplementary Information Supplementary Figures S1 to S5 and Tables S1 to S3. Optimization of the production of knock-in alleles by CRISPR/Cas9 microinjection into the mouse zygote Aurélien Raveux, Sandrine
Supplementary Information Supplementary Figures S1 to S5 and Tables S1 to S3. Optimization of the production of knock-in alleles by CRISPR/Cas9 microinjection into the mouse zygote Aurélien Raveux, Sandrine
Design. Construction. Characterization
 Design Construction Characterization DNA mrna (messenger) A C C transcription translation C A C protein His A T G C T A C G Plasmids replicon copy number incompatibility selection marker origin of replication
Design Construction Characterization DNA mrna (messenger) A C C transcription translation C A C protein His A T G C T A C G Plasmids replicon copy number incompatibility selection marker origin of replication
Guide-it sgrna In Vitro Transcription and Screening Systems User Manual
 Guide-it sgrna In Vitro Transcription and Screening Systems User Manual Cat. Nos. 632638, 632639, 632635, 632636, 632637 (040618) 1290 Terra Bella Avenue, Mountain View, CA 94043, USA U.S. Technical Support:
Guide-it sgrna In Vitro Transcription and Screening Systems User Manual Cat. Nos. 632638, 632639, 632635, 632636, 632637 (040618) 1290 Terra Bella Avenue, Mountain View, CA 94043, USA U.S. Technical Support:
Nature Biotechnology: doi: /nbt Supplementary Figure 1. In vitro validation of OTC sgrnas and donor template.
 Supplementary Figure 1 In vitro validation of OTC sgrnas and donor template. (a) In vitro validation of sgrnas targeted to OTC in the MC57G mouse cell line by transient transfection followed by 4-day puromycin
Supplementary Figure 1 In vitro validation of OTC sgrnas and donor template. (a) In vitro validation of sgrnas targeted to OTC in the MC57G mouse cell line by transient transfection followed by 4-day puromycin
CRISPR/Cas9 Gene Editing Tools
 CRISPR/Cas9 Gene Editing Tools - Guide-it Products for Successful CRISPR/Cas9 Gene Editing - Why choose Guide-it products? Optimized methods designed for speed and ease of use Complete kits that don t
CRISPR/Cas9 Gene Editing Tools - Guide-it Products for Successful CRISPR/Cas9 Gene Editing - Why choose Guide-it products? Optimized methods designed for speed and ease of use Complete kits that don t
Protocols for cell lines using CRISPR/CAS
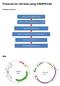 Protocols for cell lines using CRISPR/CAS Procedure overview Map Preparation of CRISPR/CAS plasmids Expression vectors for guide RNA (U6-gRNA) and Cas9 gene (CMV-p-Cas9) are ampicillin-resist ant and stable
Protocols for cell lines using CRISPR/CAS Procedure overview Map Preparation of CRISPR/CAS plasmids Expression vectors for guide RNA (U6-gRNA) and Cas9 gene (CMV-p-Cas9) are ampicillin-resist ant and stable
Development of High Quality CRISPR/Cas9 Agents
 Development of High Quality CRISPR/Cas9 Agents TIDES May 7 th, 2018 Terence Ta editasmedicine.com 1 Agenda Overview of CRISPR and Editas platform Development of NGS-based method for guide RNA QC Covalently-coupled
Development of High Quality CRISPR/Cas9 Agents TIDES May 7 th, 2018 Terence Ta editasmedicine.com 1 Agenda Overview of CRISPR and Editas platform Development of NGS-based method for guide RNA QC Covalently-coupled
Protocol for Genome Editing via the RNA-guided Cas9 Nuclease in. Zebrafish Embryos 1
 Protocol for Genome Editing via the RNA-guided Cas9 Nuclease in Zebrafish Embryos 1 1. In vitro synthesis of capped Cas9 mrna The full length of humanized Cas9 cdnas with double NLS were cloned into pxt7
Protocol for Genome Editing via the RNA-guided Cas9 Nuclease in Zebrafish Embryos 1 1. In vitro synthesis of capped Cas9 mrna The full length of humanized Cas9 cdnas with double NLS were cloned into pxt7
CRISPR/Cas9 Gene Editing Tools
 CRISPR/Cas9 Gene Editing Tools - Separations Simply Spectacular INDELS Identify indels Determine if one or both copies of your gene have indels The Guide-it Genotype Confirmation Kit: Simple detection
CRISPR/Cas9 Gene Editing Tools - Separations Simply Spectacular INDELS Identify indels Determine if one or both copies of your gene have indels The Guide-it Genotype Confirmation Kit: Simple detection
Supplementary Methods
 Supplementary Methods Reverse transcribed Quantitative PCR. Total RNA was isolated from bone marrow derived macrophages using RNeasy Mini Kit (Qiagen), DNase-treated (Promega RQ1), and reverse transcribed
Supplementary Methods Reverse transcribed Quantitative PCR. Total RNA was isolated from bone marrow derived macrophages using RNeasy Mini Kit (Qiagen), DNase-treated (Promega RQ1), and reverse transcribed
CELLTECHGEN For Research Only. Construction of sgrna expression vector for Lenti-virus system (Example: Lenti-U6 sgrna-ef1 -Puro vector)
 Construction of sgrna expression vector for Lenti-virus system (Example: Lenti-U6 sgrna-ef1 -Puro vector) Catalog number: CTG-CAS9-11 Introduction The vector Lenti-U6 sgrna-ef1 -Puro is designed for expression
Construction of sgrna expression vector for Lenti-virus system (Example: Lenti-U6 sgrna-ef1 -Puro vector) Catalog number: CTG-CAS9-11 Introduction The vector Lenti-U6 sgrna-ef1 -Puro is designed for expression
File name: Supplementary Information Description: Supplementary figures and supplementary tables. File name: Peer review file Description:
 File name: Supplementary Information Description: Supplementary figures and supplementary tables. File name: Peer review file Description: Supplementary Figure 1. dcas9-mq1 fusion protein induces de novo
File name: Supplementary Information Description: Supplementary figures and supplementary tables. File name: Peer review file Description: Supplementary Figure 1. dcas9-mq1 fusion protein induces de novo
Guide-it Indel Identification Kit User Manual
 Clontech Laboratories, Inc. Guide-it Indel Identification Kit User Manual Cat. No. 631444 (120114) Clontech Laboratories, Inc. A Takara Bio Company 1290 Terra Bella Avenue, Mountain View, CA 94043, USA
Clontech Laboratories, Inc. Guide-it Indel Identification Kit User Manual Cat. No. 631444 (120114) Clontech Laboratories, Inc. A Takara Bio Company 1290 Terra Bella Avenue, Mountain View, CA 94043, USA
CELLTECHGEN For Research Only. Construction of all-in-one vector for Lenti-virus system (Example: Lenti-EF1 -Cas9-EGFP-U6 sgrna vector)
 Construction of all-in-one vector for Lenti-virus system (Example: Lenti-EF1 -Cas9-EGFP-U6 sgrna vector) Catalog number: CTG-CAS9-18 Introduction The vector Lenti-EF1 -Cas9-EGFP-U6 sgrna is designed for
Construction of all-in-one vector for Lenti-virus system (Example: Lenti-EF1 -Cas9-EGFP-U6 sgrna vector) Catalog number: CTG-CAS9-18 Introduction The vector Lenti-EF1 -Cas9-EGFP-U6 sgrna is designed for
Genome Sequence Assembly
 Genome Sequence Assembly Learning Goals: Introduce the field of bioinformatics Familiarize the student with performing sequence alignments Understand the assembly process in genome sequencing Introduction:
Genome Sequence Assembly Learning Goals: Introduce the field of bioinformatics Familiarize the student with performing sequence alignments Understand the assembly process in genome sequencing Introduction:
Guide-it sgrna In Vitro Transcription and Screening Systems User Manual
 Clontech Laboratories, Inc. Guide-it sgrna In Vitro Transcription and Screening Systems User Manual Cat. Nos. 631438, 631439 & 631440 (042114) Clontech Laboratories, Inc. A Takara Bio Company 1290 Terra
Clontech Laboratories, Inc. Guide-it sgrna In Vitro Transcription and Screening Systems User Manual Cat. Nos. 631438, 631439 & 631440 (042114) Clontech Laboratories, Inc. A Takara Bio Company 1290 Terra
Allele-specific locus binding and genome editing by CRISPR at the
 Supplementary Information Allele-specific locus binding and genome editing by CRISPR at the p6ink4a locus Toshitsugu Fujita, Miyuki Yuno, and Hodaka Fujii Supplementary Figure Legends Supplementary Figure
Supplementary Information Allele-specific locus binding and genome editing by CRISPR at the p6ink4a locus Toshitsugu Fujita, Miyuki Yuno, and Hodaka Fujii Supplementary Figure Legends Supplementary Figure
Supplemental figure 1
 Supplemental figure 1 0.3 Copy number ratio relative to beta actin 0.25 0.2 0.15 0.1 0.05 0 Series1 Series2 Series3 0 1 PC3 BP 2 DU145 BP 3 PC3 CMV 4 DU145 CMV Supplemental figure 1. qpcr to quantify the
Supplemental figure 1 0.3 Copy number ratio relative to beta actin 0.25 0.2 0.15 0.1 0.05 0 Series1 Series2 Series3 0 1 PC3 BP 2 DU145 BP 3 PC3 CMV 4 DU145 CMV Supplemental figure 1. qpcr to quantify the
PrecisionX Multiplex grna Cloning Kit. Cat. # CAS9-GRNA-KIT. User Manual
 PrecisionX Multiplex grna Cloning Kit Store at -20 C upon receipt A limited-use label license covers this product. By use of this product, you accept the terms and conditions outlined in the Licensing
PrecisionX Multiplex grna Cloning Kit Store at -20 C upon receipt A limited-use label license covers this product. By use of this product, you accept the terms and conditions outlined in the Licensing
A 5 P degradation hot spot influences molecular farming of anticancerogenic nuclease TBN1 in tobacco cells
 Plant Cell, Tissue and Organ Culture (PCTOC) A 5 P degradation hot spot influences molecular farming of anticancerogenic nuclease TBN1 in tobacco cells Anna Týcová a,b, Rajen J. J. Piernikarczyk c, Michael
Plant Cell, Tissue and Organ Culture (PCTOC) A 5 P degradation hot spot influences molecular farming of anticancerogenic nuclease TBN1 in tobacco cells Anna Týcová a,b, Rajen J. J. Piernikarczyk c, Michael
Supplementary Fig.1. Over-expression of RNase L in stable polyclonal cell line
 Supplemental Data mrna Protein A kda 75 40 NEO/vector NEO/RNase L RNASE L β-actin RNASE L β-actin B % of control (neo vector), normalized 240 ** 200 160 120 80 40 0 Neo Neo/RNase L RNase L Protein Supplementary
Supplemental Data mrna Protein A kda 75 40 NEO/vector NEO/RNase L RNASE L β-actin RNASE L β-actin B % of control (neo vector), normalized 240 ** 200 160 120 80 40 0 Neo Neo/RNase L RNase L Protein Supplementary
Supplementary Figure 1. Nature Structural & Molecular Biology: doi: /nsmb.3494
 Supplementary Figure 1 Pol structure-function analysis (a) Inactivating polymerase and helicase mutations do not alter the stability of Pol. Flag epitopes were introduced using CRISPR/Cas9 gene targeting
Supplementary Figure 1 Pol structure-function analysis (a) Inactivating polymerase and helicase mutations do not alter the stability of Pol. Flag epitopes were introduced using CRISPR/Cas9 gene targeting
Supplementary Figure 1
 Supplementary Figure 1 Product purity of cytosine base editing for the wheat genomic loci tested. Product distributions and indels frequencies at four representative wheat genomic DNA sites in wheat protoplasts
Supplementary Figure 1 Product purity of cytosine base editing for the wheat genomic loci tested. Product distributions and indels frequencies at four representative wheat genomic DNA sites in wheat protoplasts
i-stop codon positions in the mcherry gene
 Supplementary Figure 1 i-stop codon positions in the mcherry gene The grnas (green) that can potentially generate stop codons from Trp (63 th and 98 th aa, upper panel) and Gln (47 th and 114 th aa, bottom
Supplementary Figure 1 i-stop codon positions in the mcherry gene The grnas (green) that can potentially generate stop codons from Trp (63 th and 98 th aa, upper panel) and Gln (47 th and 114 th aa, bottom
Construction of plant complementation vector and generation of transgenic plants
 MATERIAL S AND METHODS Plant materials and growth conditions Arabidopsis ecotype Columbia (Col0) was used for this study. SALK_072009, SALK_076309, and SALK_027645 were obtained from the Arabidopsis Biological
MATERIAL S AND METHODS Plant materials and growth conditions Arabidopsis ecotype Columbia (Col0) was used for this study. SALK_072009, SALK_076309, and SALK_027645 were obtained from the Arabidopsis Biological
Optimized, chemically-modified crrna:tracrrna complexes for CRISPR gene editing
 Optimized, chemically-modified crrna:tracrrna complexes for CRISPR gene editing Mark Behlke MD, PhD Chief Scientific Officer February 24, 2016 1 Implementing CRISPR/Cas9 gene editing 2 To focus on RNA
Optimized, chemically-modified crrna:tracrrna complexes for CRISPR gene editing Mark Behlke MD, PhD Chief Scientific Officer February 24, 2016 1 Implementing CRISPR/Cas9 gene editing 2 To focus on RNA
Schematic representation of the endogenous PALB2 locus and gene-disruption constructs
 Supplementary Figures Supplementary Figure 1. Generation of PALB2 -/- and BRCA2 -/- /PALB2 -/- DT40 cells. (A) Schematic representation of the endogenous PALB2 locus and gene-disruption constructs carrying
Supplementary Figures Supplementary Figure 1. Generation of PALB2 -/- and BRCA2 -/- /PALB2 -/- DT40 cells. (A) Schematic representation of the endogenous PALB2 locus and gene-disruption constructs carrying
The Biotechnology Toolbox
 Chapter 15 The Biotechnology Toolbox Cutting and Pasting DNA Cutting DNA Restriction endonuclease or restriction enzymes Cellular protection mechanism for infected foreign DNA Recognition and cutting specific
Chapter 15 The Biotechnology Toolbox Cutting and Pasting DNA Cutting DNA Restriction endonuclease or restriction enzymes Cellular protection mechanism for infected foreign DNA Recognition and cutting specific
SUPPLEMENTARY INFORMATION
 Secondary mutations as a mechanism of cisplatin resistance in BRCA2-mutated cancers Wataru Sakai, Elizabeth M. Swisher, Beth Y. Karlan, Mukesh K. Agarwal, Jake Higgins, Cynthia Friedman, Emily Villegas,
Secondary mutations as a mechanism of cisplatin resistance in BRCA2-mutated cancers Wataru Sakai, Elizabeth M. Swisher, Beth Y. Karlan, Mukesh K. Agarwal, Jake Higgins, Cynthia Friedman, Emily Villegas,
Improving CRISPR-Cas9 Gene Knockout with a Validated Guide RNA Algorithm
 Improving CRISPR-Cas9 Gene Knockout with a Validated Guide RNA Algorithm Anja Smith Director R&D Dharmacon, part of GE Healthcare Imagination at work crrna:tracrrna program Cas9 nuclease Active crrna is
Improving CRISPR-Cas9 Gene Knockout with a Validated Guide RNA Algorithm Anja Smith Director R&D Dharmacon, part of GE Healthcare Imagination at work crrna:tracrrna program Cas9 nuclease Active crrna is
Supplementary Information
 Supplementary Information Generation of inheritable and transgene clean targeted genome-modified rice in later generations using the CRISPR/Cas9 system. Rong-Fang Xu, Hao Li, Rui-Ying Qin, Juan Li, Chun-Hong
Supplementary Information Generation of inheritable and transgene clean targeted genome-modified rice in later generations using the CRISPR/Cas9 system. Rong-Fang Xu, Hao Li, Rui-Ying Qin, Juan Li, Chun-Hong
Incorporating SeqStudio Genetic Analyzer and Sanger sequencing into genome editing workflows
 Incorporating SeqStudio Genetic Analyzer and Sanger sequencing into genome editing workflows Stephen Jackson, Ph.D 27 May 2017 The world leader in serving science Key Applications for Genome Editing Research
Incorporating SeqStudio Genetic Analyzer and Sanger sequencing into genome editing workflows Stephen Jackson, Ph.D 27 May 2017 The world leader in serving science Key Applications for Genome Editing Research
Supplementary Information
 Supplementary Information This Supplementary Information file contains the following contents: Supplementary Methods, Supplementary Discussion, Supplementary Figures (S1~S9), Supplementary Tables (1~3)
Supplementary Information This Supplementary Information file contains the following contents: Supplementary Methods, Supplementary Discussion, Supplementary Figures (S1~S9), Supplementary Tables (1~3)
UTR Reporter Vectors and Viruses
 UTR Reporter Vectors and Viruses 3 UTR, 5 UTR, Promoter Reporter (Version 1) Applied Biological Materials Inc. #1-3671 Viking Way Richmond, BC V6V 2J5 Canada Notice to Purchaser All abm products are for
UTR Reporter Vectors and Viruses 3 UTR, 5 UTR, Promoter Reporter (Version 1) Applied Biological Materials Inc. #1-3671 Viking Way Richmond, BC V6V 2J5 Canada Notice to Purchaser All abm products are for
Reprogramming MHC specificity by CRISPR-Cas9-assisted cassette exchange
 1 Supplementary Materials Reprogramming MHC specificity by CRISPR-Cas9-assisted cassette exchange 3 4 5 6 Authors William Kelton 1, Ann Cathrin Waindok 1, Theresa Pesch 1, Mark Pogson 1, Kyle Ford 1, Cristina
1 Supplementary Materials Reprogramming MHC specificity by CRISPR-Cas9-assisted cassette exchange 3 4 5 6 Authors William Kelton 1, Ann Cathrin Waindok 1, Theresa Pesch 1, Mark Pogson 1, Kyle Ford 1, Cristina
Cell proliferation was measured with Cell Counting Kit-8 (Dojindo Laboratories, Kumamoto, Japan).
 1 2 3 4 5 6 7 8 Supplemental Materials and Methods Cell proliferation assay Cell proliferation was measured with Cell Counting Kit-8 (Dojindo Laboratories, Kumamoto, Japan). GCs were plated at 96-well
1 2 3 4 5 6 7 8 Supplemental Materials and Methods Cell proliferation assay Cell proliferation was measured with Cell Counting Kit-8 (Dojindo Laboratories, Kumamoto, Japan). GCs were plated at 96-well
Supplementary data. sienigma. F-Enigma F-EnigmaSM. a-p53
 Supplementary data Supplemental Figure 1 A sienigma #2 sienigma sicontrol a-enigma - + ++ - - - - - - + ++ - - - - - - ++ B sienigma F-Enigma F-EnigmaSM a-flag HLK3 cells - - - + ++ + ++ - + - + + - -
Supplementary data Supplemental Figure 1 A sienigma #2 sienigma sicontrol a-enigma - + ++ - - - - - - + ++ - - - - - - ++ B sienigma F-Enigma F-EnigmaSM a-flag HLK3 cells - - - + ++ + ++ - + - + + - -
Supplementary Figure S1. The tetracycline-inducible CRISPR system. A) Hela cells stably
 Supplementary Information Supplementary Figure S1. The tetracycline-inducible CRISPR system. A) Hela cells stably expressing shrna sequences against TRF2 were examined by western blotting. shcon, shrna
Supplementary Information Supplementary Figure S1. The tetracycline-inducible CRISPR system. A) Hela cells stably expressing shrna sequences against TRF2 were examined by western blotting. shcon, shrna
Hop Latent Viroid (HLVd) End-Point RT-PCR Kit Product# EP38700
 3430 Schmon Parkway Thorold, ON, Canada L2V 4Y6 Phone: 866-667-4362 (905) 227-8848 Fax: (905) 227-1061 Email: techsupport@norgenbiotek.com Hop Latent Viroid (HLVd) End-Point RT-PCR Kit Product# EP38700
3430 Schmon Parkway Thorold, ON, Canada L2V 4Y6 Phone: 866-667-4362 (905) 227-8848 Fax: (905) 227-1061 Email: techsupport@norgenbiotek.com Hop Latent Viroid (HLVd) End-Point RT-PCR Kit Product# EP38700
Supplementary Figure 1. Quantitative RT-PCR experimental validation of CRISPR/Cas9 and sgrnas expression in HEK293A transfected cells.
 Supplementary Figure 1. Quantitative RT-PCR experimental validation of CRISPR/Cas9 and sgrnas expression in HEK293A transfected cells. HEK293A cells were transfected with the indicated combinations of
Supplementary Figure 1. Quantitative RT-PCR experimental validation of CRISPR/Cas9 and sgrnas expression in HEK293A transfected cells. HEK293A cells were transfected with the indicated combinations of
CFTR-null wt CFTR-null 1.0. Probe: Neo R. Figure S1
 A. B. 4.0 3.0 2.0 1.0 4.0 3.0 2.0 1.0 1 2 3 4 5 6 7 8 9 10 11 12 13 14 Probe: Neo R CFTR-null wt CFTR-null Figure S1 A. 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 10kb 8kb CFTR-null wt B. Probe: CFTR
A. B. 4.0 3.0 2.0 1.0 4.0 3.0 2.0 1.0 1 2 3 4 5 6 7 8 9 10 11 12 13 14 Probe: Neo R CFTR-null wt CFTR-null Figure S1 A. 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 10kb 8kb CFTR-null wt B. Probe: CFTR
SUPPLEMENTAL MATERIAL
 SUPPLEMENTAL MATERIAL MATERIALS AND METHODS Generation of TSPOΔ/Δ murine embryonic fibroblasts Embryos were harvested from 13.5-day pregnant TSPOfl/fl mice. After dissection to eviscerate and remove the
SUPPLEMENTAL MATERIAL MATERIALS AND METHODS Generation of TSPOΔ/Δ murine embryonic fibroblasts Embryos were harvested from 13.5-day pregnant TSPOfl/fl mice. After dissection to eviscerate and remove the
Alternative Cleavage and Polyadenylation of RNA
 Developmental Cell 18 Supplemental Information The Spen Family Protein FPA Controls Alternative Cleavage and Polyadenylation of RNA Csaba Hornyik, Lionel C. Terzi, and Gordon G. Simpson Figure S1, related
Developmental Cell 18 Supplemental Information The Spen Family Protein FPA Controls Alternative Cleavage and Polyadenylation of RNA Csaba Hornyik, Lionel C. Terzi, and Gordon G. Simpson Figure S1, related
Supplemental Data. Na Xu et al. (2016). Plant Cell /tpc
 Supplemental Figure 1. The weak fluorescence phenotype is not caused by the mutation in At3g60240. (A) A mutation mapped to the gene At3g60240. Map-based cloning strategy was used to map the mutated site
Supplemental Figure 1. The weak fluorescence phenotype is not caused by the mutation in At3g60240. (A) A mutation mapped to the gene At3g60240. Map-based cloning strategy was used to map the mutated site
Antisense RNA Targeting the First Periplasmic Domain of YidC in Escherichia coli Appears to Induce Filamentation but Does Not Affect Cell Viability
 Antisense RNA Targeting the First Periplasmic Domain of YidC in Escherichia coli Appears to Induce Filamentation but Does Not Affect Cell Viability Riaaz Lalani, Nathaniel Susilo, Elisa Xiao, Andrea Xu
Antisense RNA Targeting the First Periplasmic Domain of YidC in Escherichia coli Appears to Induce Filamentation but Does Not Affect Cell Viability Riaaz Lalani, Nathaniel Susilo, Elisa Xiao, Andrea Xu
Supporting Information
 Supporting Information Park et al. 10.1073/pnas.1410555111 5 -TCAAGTCCATCTACATGGCC-3 5 -CAGCTGCCCGGCTACTACTA-3 5 -TGCAGCTGCCCGGCTACTAC-3 5 -AAGCTGGACATCACCTCCCA-3 5 -TGACAGGAACACCTACAAGT-3 5 -AAGGCACCTTTCTGTCTCCA-3
Supporting Information Park et al. 10.1073/pnas.1410555111 5 -TCAAGTCCATCTACATGGCC-3 5 -CAGCTGCCCGGCTACTACTA-3 5 -TGCAGCTGCCCGGCTACTAC-3 5 -AAGCTGGACATCACCTCCCA-3 5 -TGACAGGAACACCTACAAGT-3 5 -AAGGCACCTTTCTGTCTCCA-3
Lentiviral CRISPR guild RNA Cloning Kit for constructing CRISPR targeting grna lentivectors
 Lentiviral CRISPR guild RNA Cloning Kit for constructing CRISPR targeting grna lentivectors Cat# Product Name Amount Application grna-h1-gb grna-h1-gp grna-h1-rb grna-h1-rp grna-h1-puro grna-h1-bsd grna-u6-gb
Lentiviral CRISPR guild RNA Cloning Kit for constructing CRISPR targeting grna lentivectors Cat# Product Name Amount Application grna-h1-gb grna-h1-gp grna-h1-rb grna-h1-rp grna-h1-puro grna-h1-bsd grna-u6-gb
SUPPLEMETARY INFORMATION. Cdk7 mediates RPB1-driven mrna synthesis in Toxoplasma gondii. Abhijit S. Deshmukh 1 *, Pallabi Mitra 2 and Mulaka Maruthi 3
 SUPPLEMETARY INFORMATION Cdk7 mediates RPB1-driven mrna synthesis in Toxoplasma gondii Abhijit S. Deshmukh 1 *, Pallabi Mitra 2 and Mulaka Maruthi 3 1 National Institute of Animal Biotechnology, Hyderabad,
SUPPLEMETARY INFORMATION Cdk7 mediates RPB1-driven mrna synthesis in Toxoplasma gondii Abhijit S. Deshmukh 1 *, Pallabi Mitra 2 and Mulaka Maruthi 3 1 National Institute of Animal Biotechnology, Hyderabad,
Designing CRISPR mediated Gene disruptions with gblocks Gene Fragments
 Designing CRISPR mediated Gene disruptions with gblocks Gene Fragments Adam Clore, PhD Manager, Synthetic Biology Design Integrated DNA technology Typical CRISPR timeline in Mammalian cell lines Design
Designing CRISPR mediated Gene disruptions with gblocks Gene Fragments Adam Clore, PhD Manager, Synthetic Biology Design Integrated DNA technology Typical CRISPR timeline in Mammalian cell lines Design
Supplementary Information
 Supplementary Information MED18 interaction with distinct transcription factors regulates plant immunity, flowering time and responses to hormones Supplementary Figure 1. Diagram showing T-DNA insertion
Supplementary Information MED18 interaction with distinct transcription factors regulates plant immunity, flowering time and responses to hormones Supplementary Figure 1. Diagram showing T-DNA insertion
Virus-induced gene complementation reveals a transcription factor network in modulation of tomato fruit ripening
 Supplementary Information Virus-induced gene complementation reveals a transcription factor network in modulation of tomato fruit ripening Tao Zhou 2,3, Hang Zhang 2,4, Tongfei Lai 1, Cheng Qin 1, Nongnong
Supplementary Information Virus-induced gene complementation reveals a transcription factor network in modulation of tomato fruit ripening Tao Zhou 2,3, Hang Zhang 2,4, Tongfei Lai 1, Cheng Qin 1, Nongnong
Bootcamp: Molecular Biology Techniques and Interpretation
 Bootcamp: Molecular Biology Techniques and Interpretation Bi8 Winter 2016 Today s outline Detecting and quantifying nucleic acids and proteins: Basic nucleic acid properties Hybridization PCR and Designing
Bootcamp: Molecular Biology Techniques and Interpretation Bi8 Winter 2016 Today s outline Detecting and quantifying nucleic acids and proteins: Basic nucleic acid properties Hybridization PCR and Designing
A protocol for the assembly of 2 or 3 sgrnas. Table of contents
 A protocol for the assembly of 2 or 3 sgrnas Table of contents Simplified protocol... 2 Table 1 Nomenclature of PCR products/primers... 3 Table 2 Structure features of primers... 3 Sequence of DT1T2-PCR
A protocol for the assembly of 2 or 3 sgrnas Table of contents Simplified protocol... 2 Table 1 Nomenclature of PCR products/primers... 3 Table 2 Structure features of primers... 3 Sequence of DT1T2-PCR
Contents... vii. List of Figures... xii. List of Tables... xiv. Abbreviatons... xv. Summary... xvii. 1. Introduction In vitro evolution...
 vii Contents Contents... vii List of Figures... xii List of Tables... xiv Abbreviatons... xv Summary... xvii 1. Introduction...1 1.1 In vitro evolution... 1 1.2 Phage Display Technology... 3 1.3 Cell surface
vii Contents Contents... vii List of Figures... xii List of Tables... xiv Abbreviatons... xv Summary... xvii 1. Introduction...1 1.1 In vitro evolution... 1 1.2 Phage Display Technology... 3 1.3 Cell surface
Amplicons, Heteroduplexes and Enzymes - Proper Processing Elevates Detection of CRISPR Gene Editing Events
 Amplicons, Heteroduplexes and Enzymes - Proper Processing Elevates Detection of CRISPR Gene Editing Events Steve Siembieda, MS MBA VP Commercialization ABRF Conference February 2015 What Is CRISPR? Clustered
Amplicons, Heteroduplexes and Enzymes - Proper Processing Elevates Detection of CRISPR Gene Editing Events Steve Siembieda, MS MBA VP Commercialization ABRF Conference February 2015 What Is CRISPR? Clustered
Supplementary Figures Montero et al._supplementary Figure 1
 Montero et al_suppl. Info 1 Supplementary Figures Montero et al._supplementary Figure 1 Montero et al_suppl. Info 2 Supplementary Figure 1. Transcripts arising from the structurally conserved subtelomeres
Montero et al_suppl. Info 1 Supplementary Figures Montero et al._supplementary Figure 1 Montero et al_suppl. Info 2 Supplementary Figure 1. Transcripts arising from the structurally conserved subtelomeres
b alternative classical none
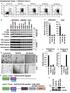 Supplementary Figure. 1: Related to Figure.1 a d e b alternative classical none NIK P-IkBa Total IkBa Tubulin P52 (Lighter) P52 (Darker) RelB (Lighter) RelB (Darker) HDAC1 Control-Sh RelB-Sh NF-kB2-Sh
Supplementary Figure. 1: Related to Figure.1 a d e b alternative classical none NIK P-IkBa Total IkBa Tubulin P52 (Lighter) P52 (Darker) RelB (Lighter) RelB (Darker) HDAC1 Control-Sh RelB-Sh NF-kB2-Sh
Nature Methods: doi: /nmeth Supplementary Figure 1
 Supplementary Figure 1 ATP1A1 variants with in-frame deletions are enriched in ouabain-resistant cell populations. (a) Total editing efficacy along with spectrum and frequency of individual indels as determined
Supplementary Figure 1 ATP1A1 variants with in-frame deletions are enriched in ouabain-resistant cell populations. (a) Total editing efficacy along with spectrum and frequency of individual indels as determined
Supplementary Table 1. Sequences for BTG2 and BRCA1 sirnas.
 Supplementary Table 1. Sequences for BTG2 and BRCA1 sirnas. Target Gene Non-target / Control BTG2 BRCA1 NFE2L2 Target Sequence ON-TARGET plus Non-targeting sirna # 1 (Cat# D-001810-01-05) sirna1: GAACCGACAUGCUCCCGGA
Supplementary Table 1. Sequences for BTG2 and BRCA1 sirnas. Target Gene Non-target / Control BTG2 BRCA1 NFE2L2 Target Sequence ON-TARGET plus Non-targeting sirna # 1 (Cat# D-001810-01-05) sirna1: GAACCGACAUGCUCCCGGA
Chapter 20 Biotechnology
 Chapter 20 Biotechnology Manipulation of DNA In 2007, the first entire human genome had been sequenced. The ability to sequence an organisms genomes were made possible by advances in biotechnology, (the
Chapter 20 Biotechnology Manipulation of DNA In 2007, the first entire human genome had been sequenced. The ability to sequence an organisms genomes were made possible by advances in biotechnology, (the
Genome edi3ng with the CRISPR-Cas9 system
 CRISPR-Cas9 Genome Edi3ng Bootcamp AHA Council on Func3onal Genomics and Transla3onal Biology Narrated video link: hfps://youtu.be/h18hmftybnq Genome edi3ng with the CRISPR-Cas9 system Kiran Musunuru,
CRISPR-Cas9 Genome Edi3ng Bootcamp AHA Council on Func3onal Genomics and Transla3onal Biology Narrated video link: hfps://youtu.be/h18hmftybnq Genome edi3ng with the CRISPR-Cas9 system Kiran Musunuru,
Fatchiyah
 Fatchiyah Email: fatchiya@yahoo.co.id RNAs: mrna trna rrna RNAi DNAs: Protein: genome DNA cdna mikro-makro mono-poly single-multi Analysis: Identification human and animal disease Finger printing Sexing
Fatchiyah Email: fatchiya@yahoo.co.id RNAs: mrna trna rrna RNAi DNAs: Protein: genome DNA cdna mikro-makro mono-poly single-multi Analysis: Identification human and animal disease Finger printing Sexing
Polymerase Chain Reaction
 Polymerase Chain Reaction Variations of PCR in the Diagnostic Lab The most common variations of standard PCR used in the diagnostic laboratory are: Reverse Transcriptase PCR (RT-PCR) Nested PCR (n-pcr)
Polymerase Chain Reaction Variations of PCR in the Diagnostic Lab The most common variations of standard PCR used in the diagnostic laboratory are: Reverse Transcriptase PCR (RT-PCR) Nested PCR (n-pcr)
Combining Techniques to Answer Molecular Questions
 Combining Techniques to Answer Molecular Questions UNIT FM02 How to cite this article: Curr. Protoc. Essential Lab. Tech. 9:FM02.1-FM02.5. doi: 10.1002/9780470089941.etfm02s9 INTRODUCTION This manual is
Combining Techniques to Answer Molecular Questions UNIT FM02 How to cite this article: Curr. Protoc. Essential Lab. Tech. 9:FM02.1-FM02.5. doi: 10.1002/9780470089941.etfm02s9 INTRODUCTION This manual is
Supplementary Methods pcfd5 cloning protocol
 Supplementary Methods cloning protocol vermilion trna grna trna grna U6:3 Terminator AmpR attb is a vector for expressing one or multiple trna-flanked Cas9 grnas under the control of the strong, ubiquitous
Supplementary Methods cloning protocol vermilion trna grna trna grna U6:3 Terminator AmpR attb is a vector for expressing one or multiple trna-flanked Cas9 grnas under the control of the strong, ubiquitous
Alt-Ring the genome with CRISPR- Cas9
 Alt-Ring the genome with CRISPR- Cas9 Genome editing done easily and fast CHIA Jin Ngee Regional Application Specialist 1 Integrated DNA Technologies The Custom Biology Company 2 IDT was founded by a scientist
Alt-Ring the genome with CRISPR- Cas9 Genome editing done easily and fast CHIA Jin Ngee Regional Application Specialist 1 Integrated DNA Technologies The Custom Biology Company 2 IDT was founded by a scientist
Construct Design and Cloning Guide for Cas9-triggered homologous recombination
 Construct Design and Cloning Guide for Cas9-triggered homologous recombination Written by Dan Dickinson (ddickins@live.unc.edu) and last updated December 2013. Reference: Dickinson DJ, Ward JD, Reiner
Construct Design and Cloning Guide for Cas9-triggered homologous recombination Written by Dan Dickinson (ddickins@live.unc.edu) and last updated December 2013. Reference: Dickinson DJ, Ward JD, Reiner
Molecular Genetics Techniques. BIT 220 Chapter 20
 Molecular Genetics Techniques BIT 220 Chapter 20 What is Cloning? Recombinant DNA technologies 1. Producing Recombinant DNA molecule Incorporate gene of interest into plasmid (cloning vector) 2. Recombinant
Molecular Genetics Techniques BIT 220 Chapter 20 What is Cloning? Recombinant DNA technologies 1. Producing Recombinant DNA molecule Incorporate gene of interest into plasmid (cloning vector) 2. Recombinant
Supplementary Figure 1 An overview of pirna biogenesis during fetal mouse reprogramming. (a) (b)
 Supplementary Figure 1 An overview of pirna biogenesis during fetal mouse reprogramming. (a) A schematic overview of the production and amplification of a single pirna from a transposon transcript. The
Supplementary Figure 1 An overview of pirna biogenesis during fetal mouse reprogramming. (a) A schematic overview of the production and amplification of a single pirna from a transposon transcript. The
GENETICS EXAM 3 FALL a) is a technique that allows you to separate nucleic acids (DNA or RNA) by size.
 Student Name: All questions are worth 5 pts. each. GENETICS EXAM 3 FALL 2004 1. a) is a technique that allows you to separate nucleic acids (DNA or RNA) by size. b) Name one of the materials (of the two
Student Name: All questions are worth 5 pts. each. GENETICS EXAM 3 FALL 2004 1. a) is a technique that allows you to separate nucleic acids (DNA or RNA) by size. b) Name one of the materials (of the two
A novel tool for monitoring endogenous alpha-synuclein transcription by NanoLuciferase
 A novel tool for monitoring endogenous alpha-synuclein transcription by NanoLuciferase tag insertion at the 3 end using CRISPR-Cas9 genome editing technique Sambuddha Basu 1, 3, Levi Adams 1, 3, Subhrangshu
A novel tool for monitoring endogenous alpha-synuclein transcription by NanoLuciferase tag insertion at the 3 end using CRISPR-Cas9 genome editing technique Sambuddha Basu 1, 3, Levi Adams 1, 3, Subhrangshu
General Product Insert. End-Point PCR/RT-PCR Kit Product# EPxxxxx
 3430 Schmon Parkway Thorold, ON, Canada L2V 4Y6 Phone: 866-667-4362 (905) 227-8848 Fax: (905) 227-1061 Email: techsupport@norgenbiotek.com End-Point PCR/RT-PCR Kit Product# EPxxxxx General Product Insert
3430 Schmon Parkway Thorold, ON, Canada L2V 4Y6 Phone: 866-667-4362 (905) 227-8848 Fax: (905) 227-1061 Email: techsupport@norgenbiotek.com End-Point PCR/RT-PCR Kit Product# EPxxxxx General Product Insert
Supplementary Materials. China
 Supplementary Materials An Efficient Genotyping Method for Genome-modified Animals and Human Cells Generated with CRISPR/Cas9 System Xiaoxiao Zhu 1,2 *, Yajie Xu 1 *, Shanshan Yu 1 *, Lu Lu 1,2, Mingqin
Supplementary Materials An Efficient Genotyping Method for Genome-modified Animals and Human Cells Generated with CRISPR/Cas9 System Xiaoxiao Zhu 1,2 *, Yajie Xu 1 *, Shanshan Yu 1 *, Lu Lu 1,2, Mingqin
BIOLOGY - CLUTCH CH.20 - BIOTECHNOLOGY.
 !! www.clutchprep.com CONCEPT: DNA CLONING DNA cloning is a technique that inserts a foreign gene into a living host to replicate the gene and produce gene products. Transformation the process by which
!! www.clutchprep.com CONCEPT: DNA CLONING DNA cloning is a technique that inserts a foreign gene into a living host to replicate the gene and produce gene products. Transformation the process by which
Automated Electrophoresis Applications for Synthetic Biology
 Automated Electrophoresis Applications for Synthetic Biology Melissa Huang Liu, Ph.D. Product Manager, Electrophoresis Agilent Technologies 1 Outline 2100 Bioanalyzer & 4200 TapeStation Systems Automated
Automated Electrophoresis Applications for Synthetic Biology Melissa Huang Liu, Ph.D. Product Manager, Electrophoresis Agilent Technologies 1 Outline 2100 Bioanalyzer & 4200 TapeStation Systems Automated
Restriction Enzymes (endonucleases)
 In order to understand and eventually manipulate DNA (human or otherwise) an array of DNA technologies have been developed. Here are some of the tools: Restriction Enzymes (endonucleases) In order to manipulate
In order to understand and eventually manipulate DNA (human or otherwise) an array of DNA technologies have been developed. Here are some of the tools: Restriction Enzymes (endonucleases) In order to manipulate
Multiple choice questions (numbers in brackets indicate the number of correct answers)
 1 Multiple choice questions (numbers in brackets indicate the number of correct answers) February 1, 2013 1. Ribose is found in Nucleic acids Proteins Lipids RNA DNA (2) 2. Most RNA in cells is transfer
1 Multiple choice questions (numbers in brackets indicate the number of correct answers) February 1, 2013 1. Ribose is found in Nucleic acids Proteins Lipids RNA DNA (2) 2. Most RNA in cells is transfer
RNA-Guided Gene Activation by CRISPR-Cas9-Based Transcription Factors
 Supplementary Information RNA-Guided Gene Activation by CRISPR-Cas9-Based Transcription Factors Pablo Perez-Pinera 1, Daniel D. Kocak 1, Christopher M. Vockley 2,3, Andrew F. Adler 1, Ami M. Kabadi 1,
Supplementary Information RNA-Guided Gene Activation by CRISPR-Cas9-Based Transcription Factors Pablo Perez-Pinera 1, Daniel D. Kocak 1, Christopher M. Vockley 2,3, Andrew F. Adler 1, Ami M. Kabadi 1,
Applications of Cas9 nickases for genome engineering
 application note genome editing Applications of Cas9 nickases for genome engineering Shuqi Yan, Mollie Schubert, Maureen Young, Brian Wang Integrated DNA Technologies, 17 Commercial Park, Coralville, IA,
application note genome editing Applications of Cas9 nickases for genome engineering Shuqi Yan, Mollie Schubert, Maureen Young, Brian Wang Integrated DNA Technologies, 17 Commercial Park, Coralville, IA,
Somatic Primary pirna Biogenesis Driven by cis-acting RNA Elements and Trans-Acting Yb
 Cell Reports Supplemental Information Somatic Primary pirna Biogenesis Driven by cis-acting RNA Elements and Trans-Acting Yb Hirotsugu Ishizu, Yuka W. Iwasaki, Shigeki Hirakata, Haruka Ozaki, Wataru Iwasaki,
Cell Reports Supplemental Information Somatic Primary pirna Biogenesis Driven by cis-acting RNA Elements and Trans-Acting Yb Hirotsugu Ishizu, Yuka W. Iwasaki, Shigeki Hirakata, Haruka Ozaki, Wataru Iwasaki,
B. Incorrect! Ligation is also a necessary step for cloning.
 Genetics - Problem Drill 15: The Techniques in Molecular Genetics No. 1 of 10 1. Which of the following is not part of the normal process of cloning recombinant DNA in bacteria? (A) Restriction endonuclease
Genetics - Problem Drill 15: The Techniques in Molecular Genetics No. 1 of 10 1. Which of the following is not part of the normal process of cloning recombinant DNA in bacteria? (A) Restriction endonuclease
