A human immunodeficiency caused by mutations in the PIK3R1 gene. Whole exome sequencing. Whole exome sequencing libraries were prepared from 3
|
|
|
- Reginald Green
- 5 years ago
- Views:
Transcription
1 A human immunodeficiency caused by mutations in the PIK3R1 gene. Supplementary Methods Whole exome sequencing. Whole exome sequencing libraries were prepared from 3 µg of genomic DNA extracted from total blood after shearing with a S2 Ultrasonicator (Covaris). Exome capture was performed as recommended by the manufacturer, using the 51 Mb SureSelect Human All Exon kit V5 (Agilent Technologies). The barcoded exome library was sequenced on a HiSeq25 machine (Illumina) in HighOutput mode (48 libraries per FlowCell), which generated paired end reads. After demultiplexing, paired end sequences were mapped onto the human genome reference (NCBI build37/hg19 version) using the Burrows Wheeler Alignment software package ( bwa.sourceforge.net/). The mean depth of coverage obtained for each sample was >9X, and at least 95% of the exome was covered at least 15X. Single nucleotide polymorphism and indel calling was performed with GATK software tools. An in house software package (PolyWeb) was used to annotate and filter the variants. Sanger sequencing of genomic DNA extracted from total blood was used to confirm the splice site mutation in PI3KR1. DNA was amplified using the primers 5 AGAAAACATTTGAAGCAGATGAA 3 and 5 CAGTGACTGCTTCAGTTATCACG 3 and the PCR product was subsequently sequenced with one of the two primers. Cell culture. Peripheral blood mononuclear cells were isolated by Ficoll Paque densitygradient centrifugation and washed twice with RPMI 164 GlutaMax medium (Invitrogen). T cell blasts were obtained by stimulating 1x1 6 cells per ml in RPMI 164 GlutaMax supplemented with 1% human AB serum, penicillin and streptomycin (Invitrogen), PMA (2 ng/ml; Sigma Aldrich) and ionomycin (1 μmol/l). After 2 to 3 days of activation, viable cells were separated by Ficoll Paque density gradient
2 centrifugation, washed twice with RPMI 164 GlutaMax and then cultured in RPMI 164 GlutaMax supplemented with 1% human AB serum and 1 U/mL IL 2. Activation induced cell death. T cell blasts (3x1 5 cells) were incubated with 3µM of PI3Kδ inhibitor (IC87114, Calbiochem) for 15 minutes before and during stimulation with plate bound anti CD3 (2.5 µg/ml: OKT3, ebioscience) for 16h. The viability of CD4 and CD8 T cell blasts was determined using 7 amino actinomycin D (BD Via Probe, Bioscience) and annexin V (BD Pharmingen) staining, together with CD4 (RPA T4; Biolegend) and CD8 (BW135/8, Miltenyi Biotec) staining in accordance with the manufacturer's protocol (BD Pharmingen). Cells were measured with flow cytometry (BD FACSCalibur) and the data were analyzed with FlowJo software (Tree Star Inc.). Plasmids and transfection. The entire coding sequence of human wildtype p85α and mutated p85α Δ 434_475 was amplified using reverse transcription PCR from P1's T cell blast mrna incooperating flanking 5 BamHI site. The PCR products were subcloned into Topo TA vector (Invitrogen). After sequencing the inserts, p85α and p85α Δ 434_475 containing inserts were excised with BamHI and were inserted into BglII digested p3xflag MYC CMV 26 expression vector (Sigma) to encode FLAG p85α and FLAGp85α Δ 434_475. Transient transfection of FLAG p85 mutant or wildtype expressing plasmids into NIH3T3 cells were carried out using lipofectamine (Invitrogen) according to the manufacturer s directions. Cells were lysed 48h after transfection immunoblot analysis. Immunoblot analysis. T cell blasts (5 1 6 cells/ml) were stimulated with anti CD3 (.1 μg/ml OKT3, ebioscience) cross linked with rabbit anti mouse IgG (2 μg/ml) for 1
3 minutes. The cells were then lysed in TNE (5 mmol/l Tris at ph 8., 1% Nonidet P 4, and 2 mmol/l EDTA) buffer supplemented with protease and phosphatase inhibitors. Immunoblots were performed according to standard protocols. Antibodies against p11δ (H 219; Santa Cruz), phospho Akt (Ser473; #9271; Cell Signaling and Thr38; #9275S; Cell Signaling), Akt (#9272; Cell Signaling) PTEN (#9188S; Cell Signaling), Ku 7 (H 38; Santa Cruz) and glyceraldehyde 3 phosphate dehydrogenase (GAPDH; 6C5; Santa Cruz) were used for immunoblotting. Intracellular phospho Akt staining. After a 1 minute stimulation with anti CD3 (.1 μg/ml; OKT3) cross linked with rabbit anti mouse IgG (2 μg/ml), cells were fixed in 4% formaldehyde prior to cell membrane permeabilization with methanol. Staining was performed according to the manufacturer s instructions (Cell Signaling), with incubation with anti phospho AKT Ser 473 Alexa 647 (#475, Cell Signaling) for 1 hour in the dark at room temperature. Cells were measured by flow cytometry (BD FACSCalibur) and the data were analyzed with FlowJo software (Tree Star Inc.). For the analysis of phosphorylated S6 with anti phospho S6 Ribosomal Protein (Ser24/244) Alexa488 (#518, Cell Signaling) cells were fixed in the medium they were cultured with. Statistical analysis. Analyses were performed with PRISM software (version 4 for Macintosh, GraphPad Inc.). Statistical hypotheses were tested using 2 tailed t test. A P value less than.5 was considered significant.
4 A! B! Supplementary Figure 1. Imaging infection of upper and lower respiratory tract. (A) Thoracic computed tomography (CT) scan showing bronchitis with wall thickening of bronchial tract (arrows) and distal pneumonitis with multifocal ground glass opacities (arrow heads). (B) Facial CT scan showing right maxillary sinusitis with inflammatory thickening of the mucosal lining (star). Both CT scans were performed in P1.
5 Supplementary Figure 2. Location of the deleted amino acid residues ( ) in p85α. The figure shows the structure of human p11α (in orange) bound to the p85α regulatory component (in white) of the PI3K complex (Protein Data Bank accession code: 3HHM); the amino acid residues (encoded by exon 1 of the PIK3R1 gene) are highlighted in yellow and their backbone in green.
6 A! P1! P2! P3! p85!! p85"! p85" #434_475! Ku-7! B! FLAG! GAPDH! Supplementary Figure 3. Western blot for p85α. A) Expression of p85α and Ku 7 (as loading control) by Western blotting of cell lysates derived from T cell blasts from P1, P2 and P3 and healthy controls. B) A representative experiment showing the expression of flag tagged p85α wildtype (WT), flag tagged p85α Δ 434_475 and GAPDH by Western blotting of cell lysates derived from 293T cells 2 days after transfection with the indicated expression vector.
7 A! Non-stimulated! B! *! C! *! **! **! Non-stimulated! Non-stimulated! Supplementary Figure 4. Increased phosphorylation of Akt in patient T cell blasts detected by immunoblotting. A) Quantification of band intensity of p11δ normalized to band intensity of GAPDH from immunoblots presented in Figure 2a. B) Quantification of band intensity of phosphorylated Akt at Ser473 normalized to total Akt from immunoblots presented in Figure 2a. C) Quantification of band intensity of phosphorylated Akt at Thr38 normalized to total Akt from immunoblots presented in Figure 2a. * P=.5, ** P=.1 (unpaired t test).
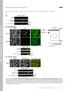
8 Ce A! B! control! P2! P3! 6 2*4/#*1$ 56$ Cell count! FL4-H: phosphoakt-s473alex647 FL4-H: phosphoakt-s473alex647 FL4-H: phosphoakt-s473alex647 4 "11$2*34/$ 16.4! 2.5! 36.6! 1µM IC87114! 1µM IC 87114! No IC 87114! 23.3! 3.5! 54.8! 1µM IC87114! 1µM IC 87114! No IC 87114! plus IC87114! FL4-H: phosphoakt-s473alex647 FL4-H: phosphoakt-s473alex647 FL4-H: phosphoakt-s473alex647 Ser473 phospho-akt!!"#$%&'$()*+()*,-./$ C! 2*4/#*1$ 56$ D! *4/#*1$ 56$!"#$%&'$ ()*+()*,-./$$ $ "11$2*34/$ phosho-s6 (Ser24/244)! phospho-s6 (Ser24/244)! Supplementary Figure 5. Hyperphosphorylation of Akt and S6 detected by flow cytometry. A) Phosphorylation of Akt at Ser473 in T cell blasts from patients and a control following activation with anti CD3 and anti CD28 antibodies for 3 minutes in the presence or absence of 1µM IC87114 inhibitor for the last 2 minutes of activation, and as measured by intracellular staining and flow cytometry. T cell blasts from patients and controls were cultured for the same period of time. B) Intracellular staining of phosphorylated Akt at Ser473 in T cell blasts from patients and a control following activation with 1µg of anti CD3 antibody in the absence (red) or presence of either 1µM (blue) or 1µM (green) IC87114 inhibitor. Numbers indicate the geometric mean fluorescence intensity. C, D) Intracellular staining of phosphorylated Akt (Ser473) and phosphorylated S6 (Ser24/244) of T cell blasts fixed immediately in medium. C) Contour plot showing phosphorylated Akt (Ser473) and phosphorylated S6 (Ser24/244) staining of non stimulated T cell blasts from patient and control D) Histogram showing the overlay of phosphorylated S6 (Ser24/244) from control (grey) and patient (black).
9 Ø! +! PTEN P1! P1! GAPDH Supplementary Figure 6. Abundance of PTEN protein in T cell blasts. Expression of PTEN and GAPDH detected by Western blotting of cell lysates derived from T cell blasts from P1 and healthy control before and after stimulation with anti CD3 antibody.
Supplementary Information
 Supplementary Information Supplementary Figure 1: Over-expression of CD300f in NIH3T3 cells enhances their capacity to phagocytize AC. (a) NIH3T3 cells were stably transduced by EV, CD300f WT or CD300f
Supplementary Information Supplementary Figure 1: Over-expression of CD300f in NIH3T3 cells enhances their capacity to phagocytize AC. (a) NIH3T3 cells were stably transduced by EV, CD300f WT or CD300f
To determine MRK-003 IC50 values, cell lines were plated in triplicate in 96-well plates at 3 x
 Supplementary methods: Cellular growth assay and cell cycle analysis To determine MRK-003 IC50 values, cell lines were plated in triplicate in 96-well plates at 3 x 10 3 cells/well (except for TALL1 and
Supplementary methods: Cellular growth assay and cell cycle analysis To determine MRK-003 IC50 values, cell lines were plated in triplicate in 96-well plates at 3 x 10 3 cells/well (except for TALL1 and
Supplementary data. sienigma. F-Enigma F-EnigmaSM. a-p53
 Supplementary data Supplemental Figure 1 A sienigma #2 sienigma sicontrol a-enigma - + ++ - - - - - - + ++ - - - - - - ++ B sienigma F-Enigma F-EnigmaSM a-flag HLK3 cells - - - + ++ + ++ - + - + + - -
Supplementary data Supplemental Figure 1 A sienigma #2 sienigma sicontrol a-enigma - + ++ - - - - - - + ++ - - - - - - ++ B sienigma F-Enigma F-EnigmaSM a-flag HLK3 cells - - - + ++ + ++ - + - + + - -
Supplementary Materials for
 advances.sciencemag.org/cgi/content/full/4/9/eaat5401/dc1 Supplementary Materials for GLK-IKKβ signaling induces dimerization and translocation of the AhR-RORγt complex in IL-17A induction and autoimmune
advances.sciencemag.org/cgi/content/full/4/9/eaat5401/dc1 Supplementary Materials for GLK-IKKβ signaling induces dimerization and translocation of the AhR-RORγt complex in IL-17A induction and autoimmune
Sarker et al. Supplementary Material. Subcellular Fractionation
 Supplementary Material Subcellular Fractionation Transfected 293T cells were harvested with phosphate buffered saline (PBS) and centrifuged at 2000 rpm (500g) for 3 min. The pellet was washed, re-centrifuged
Supplementary Material Subcellular Fractionation Transfected 293T cells were harvested with phosphate buffered saline (PBS) and centrifuged at 2000 rpm (500g) for 3 min. The pellet was washed, re-centrifuged
Cell death analysis using the high content bioimager BD PathwayTM 855 instrument (BD
 Supplemental information Materials and Methods: Cell lines, reagents and antibodies: Wild type (A3) and caspase-8 -/- (I9.2) Jurkat cells were cultured in RPMI 164 medium (Life Technologies) supplemented
Supplemental information Materials and Methods: Cell lines, reagents and antibodies: Wild type (A3) and caspase-8 -/- (I9.2) Jurkat cells were cultured in RPMI 164 medium (Life Technologies) supplemented
Supplementary Methods
 Supplementary Methods Reverse transcribed Quantitative PCR. Total RNA was isolated from bone marrow derived macrophages using RNeasy Mini Kit (Qiagen), DNase-treated (Promega RQ1), and reverse transcribed
Supplementary Methods Reverse transcribed Quantitative PCR. Total RNA was isolated from bone marrow derived macrophages using RNeasy Mini Kit (Qiagen), DNase-treated (Promega RQ1), and reverse transcribed
Cell proliferation was measured with Cell Counting Kit-8 (Dojindo Laboratories, Kumamoto, Japan).
 1 2 3 4 5 6 7 8 Supplemental Materials and Methods Cell proliferation assay Cell proliferation was measured with Cell Counting Kit-8 (Dojindo Laboratories, Kumamoto, Japan). GCs were plated at 96-well
1 2 3 4 5 6 7 8 Supplemental Materials and Methods Cell proliferation assay Cell proliferation was measured with Cell Counting Kit-8 (Dojindo Laboratories, Kumamoto, Japan). GCs were plated at 96-well
ASPP1 Fw GGTTGGGAATCCACGTGTTG ASPP1 Rv GCCATATCTTGGAGCTCTGAGAG
 Supplemental Materials and Methods Plasmids: the following plasmids were used in the supplementary data: pwzl-myc- Lats2 (Aylon et al, 2006), pretrosuper-vector and pretrosuper-shp53 (generous gift of
Supplemental Materials and Methods Plasmids: the following plasmids were used in the supplementary data: pwzl-myc- Lats2 (Aylon et al, 2006), pretrosuper-vector and pretrosuper-shp53 (generous gift of
Supporting Information
 Supporting Information Tal et al. 10.1073/pnas.0807694106 SI Materials and Methods VSV Infection and Quantification. Infection was carried out by seeding 5 10 5 MEF cells per well in a 6-well plate and
Supporting Information Tal et al. 10.1073/pnas.0807694106 SI Materials and Methods VSV Infection and Quantification. Infection was carried out by seeding 5 10 5 MEF cells per well in a 6-well plate and
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb3363 Supplementary Figure 1 Several WNTs bind to the extracellular domains of PKD1. (a) HEK293T cells were co-transfected with indicated plasmids. Flag-tagged proteins were immunoprecipiated
DOI: 10.1038/ncb3363 Supplementary Figure 1 Several WNTs bind to the extracellular domains of PKD1. (a) HEK293T cells were co-transfected with indicated plasmids. Flag-tagged proteins were immunoprecipiated
RNA was isolated using NucleoSpin RNA II (Macherey-Nagel, Bethlehem, PA) according to the
 Supplementary Methods RT-PCR and real-time PCR analysis RNA was isolated using NucleoSpin RNA II (Macherey-Nagel, Bethlehem, PA) according to the manufacturer s protocol and quantified by measuring the
Supplementary Methods RT-PCR and real-time PCR analysis RNA was isolated using NucleoSpin RNA II (Macherey-Nagel, Bethlehem, PA) according to the manufacturer s protocol and quantified by measuring the
supplementary information
 DOI: 1.138/ncb1839 a b Control 1 2 3 Control 1 2 3 Fbw7 Smad3 1 2 3 4 1 2 3 4 c d IGF-1 IGF-1Rβ IGF-1Rβ-P Control / 1 2 3 4 Real-time RT-PCR Relative quantity (IGF-1/ mrna) 2 1 IGF-1 1 2 3 4 Control /
DOI: 1.138/ncb1839 a b Control 1 2 3 Control 1 2 3 Fbw7 Smad3 1 2 3 4 1 2 3 4 c d IGF-1 IGF-1Rβ IGF-1Rβ-P Control / 1 2 3 4 Real-time RT-PCR Relative quantity (IGF-1/ mrna) 2 1 IGF-1 1 2 3 4 Control /
Supplementary Figure 1 - Characterization of rbag3 binding on macrophages cell surface.
 Supplementary Figure 1 - Characterization of rbag3 binding on macrophages cell surface. (a) Human PDAC cell lines were treated as indicated in Figure 1 panel F. Cells were analyzed for FITC-rBAG3 binding
Supplementary Figure 1 - Characterization of rbag3 binding on macrophages cell surface. (a) Human PDAC cell lines were treated as indicated in Figure 1 panel F. Cells were analyzed for FITC-rBAG3 binding
Human skin punch biopsies were obtained under informed consent from normal healthy
 SUPPLEMENTAL METHODS Acquisition of human skin specimens. Human skin punch biopsies were obtained under informed consent from normal healthy volunteers (n = 30) and psoriasis patients (n = 45) under a
SUPPLEMENTAL METHODS Acquisition of human skin specimens. Human skin punch biopsies were obtained under informed consent from normal healthy volunteers (n = 30) and psoriasis patients (n = 45) under a
Fig. S1 TGF RI inhibitor SB effectively blocks phosphorylation of Smad2 induced by TGF. FET cells were treated with TGF in the presence of
 Fig. S1 TGF RI inhibitor SB525334 effectively blocks phosphorylation of Smad2 induced by TGF. FET cells were treated with TGF in the presence of different concentrations of SB525334. Cells were lysed and
Fig. S1 TGF RI inhibitor SB525334 effectively blocks phosphorylation of Smad2 induced by TGF. FET cells were treated with TGF in the presence of different concentrations of SB525334. Cells were lysed and
used at a final concentration of 5 ng/ml. Rabbit anti-bim and mouse anti-mkp2 antibodies were
 1 Supplemental Methods Reagents and chemicals: TGFβ was a generous gift from Genzyme Inc. (Cambridge, MA) and was used at a final concentration of 5 ng/ml. Rabbit anti-bim and mouse anti-mkp2 antibodies
1 Supplemental Methods Reagents and chemicals: TGFβ was a generous gift from Genzyme Inc. (Cambridge, MA) and was used at a final concentration of 5 ng/ml. Rabbit anti-bim and mouse anti-mkp2 antibodies
Description of Supplementary Files. File name: Supplementary Information Description: Supplementary figures and supplementary tables.
 Description of Supplementary Files File name: Supplementary Information Description: Supplementary figures and supplementary tables. File name: Supplementary Data 1 Description: Differential expression
Description of Supplementary Files File name: Supplementary Information Description: Supplementary figures and supplementary tables. File name: Supplementary Data 1 Description: Differential expression
Supplementary Figure 1 Generation of migg1-yf mice. (A) Targeting strategy. Upper panel: schematic organization of the murine ɣ1 immunoglobulin
 Supplementary Figure 1 Generation of migg1-yf mice. (A) Targeting strategy. Upper panel: schematic organization of the murine ɣ1 immunoglobulin locus. The EcoRI restriction site between exons M1 and M2
Supplementary Figure 1 Generation of migg1-yf mice. (A) Targeting strategy. Upper panel: schematic organization of the murine ɣ1 immunoglobulin locus. The EcoRI restriction site between exons M1 and M2
Supplemental Table 1: Sequences of real time PCR primers. Primers were intronspanning
 Symbol Accession Number Sense-primer (5-3 ) Antisense-primer (5-3 ) T a C ACTB NM_001101.3 CCAGAGGCGTACAGGGATAG CCAACCGCGAGAAGATGA 57 HSD3B2 NM_000198.3 CTTGGACAAGGCCTTCAGAC TCAAGTACAGTCAGCTTGGTCCT 60
Symbol Accession Number Sense-primer (5-3 ) Antisense-primer (5-3 ) T a C ACTB NM_001101.3 CCAGAGGCGTACAGGGATAG CCAACCGCGAGAAGATGA 57 HSD3B2 NM_000198.3 CTTGGACAAGGCCTTCAGAC TCAAGTACAGTCAGCTTGGTCCT 60
Segments of the obstructed intestinal loops were fixed in 4% paraformaldehyde
 Supplementary text Supplementary materials and methods Histopathological examination Segments of the obstructed intestinal loops were fixed in 4% paraformaldehyde (PFA) and embedded in paraffin wax with
Supplementary text Supplementary materials and methods Histopathological examination Segments of the obstructed intestinal loops were fixed in 4% paraformaldehyde (PFA) and embedded in paraffin wax with
Supplementary Figure 1
 Supplementary Figure 1 Virus infection induces RNF128 expression. (a,b) RT-PCR analysis of Rnf128 (RNF128) mrna expression in mouse peritoneal macrophages (a) and THP-1 cells (b) upon stimulation with
Supplementary Figure 1 Virus infection induces RNF128 expression. (a,b) RT-PCR analysis of Rnf128 (RNF128) mrna expression in mouse peritoneal macrophages (a) and THP-1 cells (b) upon stimulation with
Flag-Rac Vector V12 V12 N17 C40. Vector C40 pakt (T308) Akt1. Myc-DN-PAK1 (N-SP)
 a b FlagRac FlagRac V2 V2 N7 C4 V2 V2 N7 C4 p (T38) p (S99, S24) p Flag (Rac) NIH 3T3 COS c +Serum p (T38) MycDN (NSP) Mycp27 3 6 2 3 6 2 3 6 2 min p Myc ( or p27) Figure S (a) Effects of Rac mutants on
a b FlagRac FlagRac V2 V2 N7 C4 V2 V2 N7 C4 p (T38) p (S99, S24) p Flag (Rac) NIH 3T3 COS c +Serum p (T38) MycDN (NSP) Mycp27 3 6 2 3 6 2 3 6 2 min p Myc ( or p27) Figure S (a) Effects of Rac mutants on
Supplemental Table S1. RT-PCR primers used in this study
 Supplemental Table S1. RT-PCR primers used in this study -----------------------------------------------------------------------------------------------------------------------------------------------
Supplemental Table S1. RT-PCR primers used in this study -----------------------------------------------------------------------------------------------------------------------------------------------
i-stop codon positions in the mcherry gene
 Supplementary Figure 1 i-stop codon positions in the mcherry gene The grnas (green) that can potentially generate stop codons from Trp (63 th and 98 th aa, upper panel) and Gln (47 th and 114 th aa, bottom
Supplementary Figure 1 i-stop codon positions in the mcherry gene The grnas (green) that can potentially generate stop codons from Trp (63 th and 98 th aa, upper panel) and Gln (47 th and 114 th aa, bottom
T H E J O U R N A L O F C E L L B I O L O G Y
 T H E J O U R N A L O F C E L L B I O L O G Y Supplemental material Han et al., http://www.jcb.org/cgi/content/full/jcb.201311007/dc1 Figure S1. SIVA1 interacts with PCNA. (A) HEK293T cells were transiently
T H E J O U R N A L O F C E L L B I O L O G Y Supplemental material Han et al., http://www.jcb.org/cgi/content/full/jcb.201311007/dc1 Figure S1. SIVA1 interacts with PCNA. (A) HEK293T cells were transiently
Nature Medicine doi: /nm.2548
 Supplementary Table 1: Genotypes of offspring and embryos from matings of Pmm2 WT/F118L mice with Pmm2 WT/R137H mice total events Pmm2 WT/WT Pmm2 WT/R137H Pmm2 WT/F118L Pmm2 R137H/F118L offspring 117 (100%)
Supplementary Table 1: Genotypes of offspring and embryos from matings of Pmm2 WT/F118L mice with Pmm2 WT/R137H mice total events Pmm2 WT/WT Pmm2 WT/R137H Pmm2 WT/F118L Pmm2 R137H/F118L offspring 117 (100%)
Mutagenesis and generation of expression vectors Gene expression profiling Lentiviral production and infection of TF1 cells
 Mutagenesis and generation of expression vectors Human c-kit wild-type cdna encoding the short c-kit isoform was excised from vector pbshkitwt (kind gift from Dr. L Gros, France) by Sal I-Acc65 I digestion.
Mutagenesis and generation of expression vectors Human c-kit wild-type cdna encoding the short c-kit isoform was excised from vector pbshkitwt (kind gift from Dr. L Gros, France) by Sal I-Acc65 I digestion.
Supplementary Figure 1. Co-localization of GLUT1 and DNAL4 in BeWo cells cultured
 Supplementary Figure 1. Co-localization of GLUT1 and DNAL4 in BeWo cells cultured under static conditions. Cells were seeded in the chamber area of the device and cultured overnight without medium perfusion.
Supplementary Figure 1. Co-localization of GLUT1 and DNAL4 in BeWo cells cultured under static conditions. Cells were seeded in the chamber area of the device and cultured overnight without medium perfusion.
Antibodies used in this study Figure S1. Akt expression in neutrophils from WT and individual Akt isoform knockout mice
 ntibodies used in this study The anti β-actin monoclonal antibody (Sigma-ldrich, St. Louis, MO) was generated against a slightly modified human β-actin N-terminal peptide, c-sp-sp-sp-ile-la-la-leu-val-ile-
ntibodies used in this study The anti β-actin monoclonal antibody (Sigma-ldrich, St. Louis, MO) was generated against a slightly modified human β-actin N-terminal peptide, c-sp-sp-sp-ile-la-la-leu-val-ile-
Cell viability. Cell viability was examined by MTT assay (Sigma-Aldrich).
 Supplementary Materials Supplementary materials and methods Cell culture. Primary human dermal fibroblasts (DFs) were isolated from full-thickness skin samples. Tissue samples were dissected into small
Supplementary Materials Supplementary materials and methods Cell culture. Primary human dermal fibroblasts (DFs) were isolated from full-thickness skin samples. Tissue samples were dissected into small
Revised: RG-RV2 by Fukuhara et al.
 Supplemental Figure 1 The generation of Spns2 conditional knockout mice. (A) Schematic representation of the wild type Spns2 locus (Spns2 + ), the targeted allele, the floxed allele (Spns2 f ) and the
Supplemental Figure 1 The generation of Spns2 conditional knockout mice. (A) Schematic representation of the wild type Spns2 locus (Spns2 + ), the targeted allele, the floxed allele (Spns2 f ) and the
Apoptosis assay: Apoptotic cells were identified by Annexin V-Alexa Fluor 488 and Propidium
 Apoptosis assay: Apoptotic cells were identified by Annexin V-Alexa Fluor 488 and Propidium Iodide (Invitrogen, Carlsbad, CA) staining. Briefly, 2x10 5 cells were washed once in cold PBS and resuspended
Apoptosis assay: Apoptotic cells were identified by Annexin V-Alexa Fluor 488 and Propidium Iodide (Invitrogen, Carlsbad, CA) staining. Briefly, 2x10 5 cells were washed once in cold PBS and resuspended
Single-cell suspensions of C57BL/6J splenocytes were incubated with CD5 beads
 S S Single-cell suspensions of C57BL/6J splenocytes were incubated with CD5 beads (Miltenyi Biotech) and T cells were positively selected by magnetically activated cell sorting (MACS). 50x10 6 or greater
S S Single-cell suspensions of C57BL/6J splenocytes were incubated with CD5 beads (Miltenyi Biotech) and T cells were positively selected by magnetically activated cell sorting (MACS). 50x10 6 or greater
Supplementary Information (Ha, et. al) Supplementary Figures Supplementary Fig. S1
 Supplementary Information (Ha, et. al) Supplementary Figures Supplementary Fig. S1 a His-ORMDL3 ~ 17 His-ORMDL3 GST-ORMDL3 - + - + IPTG GST-ORMDL3 ~ b Integrated Density (ORMDL3/ -actin) 0.4 0.3 0.2 0.1
Supplementary Information (Ha, et. al) Supplementary Figures Supplementary Fig. S1 a His-ORMDL3 ~ 17 His-ORMDL3 GST-ORMDL3 - + - + IPTG GST-ORMDL3 ~ b Integrated Density (ORMDL3/ -actin) 0.4 0.3 0.2 0.1
B. C. Staurosporine BEZ235 DMSO. -actin. Suppl. Fig. 1 BEZ235 and BGT226 inhibit phosphorylation of Akt and downstream targets of PI3K and mtor.
 DMSO Staurosporine BGT226 A. B. C. BGT226 P-Akt 0 1 2 4 8 hrs. P-Akt 0 5 10 25 50 100 nm PS6 P-4E-BP1 -actin C-Caspase3 -actin PS6 P-4E-BP1 -actin Suppl. Fig. 1 and BGT226 inhibit phosphorylation of Akt
DMSO Staurosporine BGT226 A. B. C. BGT226 P-Akt 0 1 2 4 8 hrs. P-Akt 0 5 10 25 50 100 nm PS6 P-4E-BP1 -actin C-Caspase3 -actin PS6 P-4E-BP1 -actin Suppl. Fig. 1 and BGT226 inhibit phosphorylation of Akt
Spironolactone ameliorates PIT1-dependent vascular osteoinduction in klotho-hypomorphic mice
 Spironolactone ameliorates PIT1-dependent vascular osteoinduction in klotho-hypomorphic mice Supplementary Material Supplementary Methods Materials Spironolactone, aldosterone and β-glycerophosphate were
Spironolactone ameliorates PIT1-dependent vascular osteoinduction in klotho-hypomorphic mice Supplementary Material Supplementary Methods Materials Spironolactone, aldosterone and β-glycerophosphate were
Fig. S1. Effect of p120-catenin overexpression on the interaction of SCUBE2 with E-cadherin. The expression plasmid encoding FLAG.
 Fig. S1. Effect of p120-catenin overexpression on the interaction of SCUBE2 with E-cadherin. The expression plasmid encoding FLAG.SCUBE2, E-cadherin.Myc, or HA.p120-catenin was transfected in a combination
Fig. S1. Effect of p120-catenin overexpression on the interaction of SCUBE2 with E-cadherin. The expression plasmid encoding FLAG.SCUBE2, E-cadherin.Myc, or HA.p120-catenin was transfected in a combination
Supplementary Information
 Journal : Nature Biotechnology Supplementary Information Targeted genome engineering in human cells with RNA-guided endonucleases Seung Woo Cho, Sojung Kim, Jong Min Kim, and Jin-Soo Kim* National Creative
Journal : Nature Biotechnology Supplementary Information Targeted genome engineering in human cells with RNA-guided endonucleases Seung Woo Cho, Sojung Kim, Jong Min Kim, and Jin-Soo Kim* National Creative
Supplementary information to accompany: A novel role for the DNA repair gene Rad51 in Netrin-1 signalling
 Supplementary information to accompany: A novel role for the DNA repair gene Rad51 in Netrin-1 signalling Glendining KA 1, Markie D 2, Gardner RJM 4, Franz EA 3, Robertson SP 4, Jasoni CL 1 Supplementary
Supplementary information to accompany: A novel role for the DNA repair gene Rad51 in Netrin-1 signalling Glendining KA 1, Markie D 2, Gardner RJM 4, Franz EA 3, Robertson SP 4, Jasoni CL 1 Supplementary
RNA oligonucleotides and 2 -O-methylated oligonucleotides were synthesized by. 5 AGACACAAACACCAUUGUCACACUCCACAGC; Rand-2 OMe,
 Materials and methods Oligonucleotides and DNA constructs RNA oligonucleotides and 2 -O-methylated oligonucleotides were synthesized by Dharmacon Inc. (Lafayette, CO). The sequences were: 122-2 OMe, 5
Materials and methods Oligonucleotides and DNA constructs RNA oligonucleotides and 2 -O-methylated oligonucleotides were synthesized by Dharmacon Inc. (Lafayette, CO). The sequences were: 122-2 OMe, 5
Supplementary Information: Materials and Methods. Immunoblot and immunoprecipitation. Cells were washed in phosphate buffered
 Supplementary Information: Materials and Methods Immunoblot and immunoprecipitation. Cells were washed in phosphate buffered saline (PBS) and lysed in TNN lysis buffer (50mM Tris at ph 8.0, 120mM NaCl
Supplementary Information: Materials and Methods Immunoblot and immunoprecipitation. Cells were washed in phosphate buffered saline (PBS) and lysed in TNN lysis buffer (50mM Tris at ph 8.0, 120mM NaCl
Cell extracts and western blotting RNA isolation and real-time PCR Chromatin immunoprecipitation (ChIP)
 Cell extracts and western blotting Cells were washed with ice-cold phosphate-buffered saline (PBS) and lysed with lysis buffer. 1 Total cell extracts were separated by SDS-PAGE and transferred to nitrocellulose
Cell extracts and western blotting Cells were washed with ice-cold phosphate-buffered saline (PBS) and lysed with lysis buffer. 1 Total cell extracts were separated by SDS-PAGE and transferred to nitrocellulose
transcription and the promoter occupancy of Smad proteins. (A) HepG2 cells were co-transfected with the wwp-luc reporter, and FLAG-tagged FHL1,
 Supplementary Data Supplementary Figure Legends Supplementary Figure 1 FHL-mediated TGFβ-responsive reporter transcription and the promoter occupancy of Smad proteins. (A) HepG2 cells were co-transfected
Supplementary Data Supplementary Figure Legends Supplementary Figure 1 FHL-mediated TGFβ-responsive reporter transcription and the promoter occupancy of Smad proteins. (A) HepG2 cells were co-transfected
Supplemental Materials and Methods
 Supplemental Materials and Methods Co-immunoprecipitation (Co-IP) assay Cells were lysed with NETN buffer (20 mm Tris-HCl, ph 8.0, 0 mm NaCl, 1 mm EDT, 0.5% Nonidet P-40) containing 50 mm β-glycerophosphate,
Supplemental Materials and Methods Co-immunoprecipitation (Co-IP) assay Cells were lysed with NETN buffer (20 mm Tris-HCl, ph 8.0, 0 mm NaCl, 1 mm EDT, 0.5% Nonidet P-40) containing 50 mm β-glycerophosphate,
supplementary information
 DOI: 10.1038/ncb1862 Figure S1 Identification of SCAI as a Dia1 associating factor. (a) Identification of SCAI (Riken cdna 930041I02) as a Dia1-FH3 interacting protein. Mouse brain lysate was incubated
DOI: 10.1038/ncb1862 Figure S1 Identification of SCAI as a Dia1 associating factor. (a) Identification of SCAI (Riken cdna 930041I02) as a Dia1-FH3 interacting protein. Mouse brain lysate was incubated
SUPPLEMENTARY INFORMATION
 Secondary mutations as a mechanism of cisplatin resistance in BRCA2-mutated cancers Wataru Sakai, Elizabeth M. Swisher, Beth Y. Karlan, Mukesh K. Agarwal, Jake Higgins, Cynthia Friedman, Emily Villegas,
Secondary mutations as a mechanism of cisplatin resistance in BRCA2-mutated cancers Wataru Sakai, Elizabeth M. Swisher, Beth Y. Karlan, Mukesh K. Agarwal, Jake Higgins, Cynthia Friedman, Emily Villegas,
Supplementary Figure 1 Pfn1, but not other Pfn isoforms are expressed in
 Supplementary Figure 1 Pfn1, but not other Pfn isoforms are expressed in platelets. (a) RT-PCR of Pfn isoforms in control mouse platelets, Pfn1 -/- platelets and control heart. Expected band size for Pfn1
Supplementary Figure 1 Pfn1, but not other Pfn isoforms are expressed in platelets. (a) RT-PCR of Pfn isoforms in control mouse platelets, Pfn1 -/- platelets and control heart. Expected band size for Pfn1
Supplementary Material
 Supplementary Material Supplementary Methods Cell synchronization. For synchronized cell growth, thymidine was added to 30% confluent U2OS cells to a final concentration of 2.5mM. Cells were incubated
Supplementary Material Supplementary Methods Cell synchronization. For synchronized cell growth, thymidine was added to 30% confluent U2OS cells to a final concentration of 2.5mM. Cells were incubated
The RT-qPCR analysis of selected type-i IFNs related genes, IRF7 and Oas3. The
 SUPPLEMENTARY MATERIALS AND METHODS Real time quantitative PCR The RT-qPCR analysis of selected type-i IFNs related genes, IRF7 and Oas3. The RT-qPCR was performed on the Applied Biosystems StepOne TM
SUPPLEMENTARY MATERIALS AND METHODS Real time quantitative PCR The RT-qPCR analysis of selected type-i IFNs related genes, IRF7 and Oas3. The RT-qPCR was performed on the Applied Biosystems StepOne TM
Supplementary Fig. 1 Proteomic analysis of ATR-interacting proteins. ATR, ARID1A and
 Supplementary Figure Legend: Supplementary Fig. 1 Proteomic analysis of ATR-interacting proteins. ATR, ARID1A and ATRIP protein peptides identified from our mass spectrum analysis were shown. Supplementary
Supplementary Figure Legend: Supplementary Fig. 1 Proteomic analysis of ATR-interacting proteins. ATR, ARID1A and ATRIP protein peptides identified from our mass spectrum analysis were shown. Supplementary
No wash 2 Washes 2 Days ** ** IgG-bead phagocytosis (%)
 Supplementary Figures Supplementary Figure 1. No wash 2 Washes 2 Days Tat Control ** ** 2 4 6 8 IgG-bead phagocytosis (%) Supplementary Figure 1. Reversibility of phagocytosis inhibition by Tat. Human
Supplementary Figures Supplementary Figure 1. No wash 2 Washes 2 Days Tat Control ** ** 2 4 6 8 IgG-bead phagocytosis (%) Supplementary Figure 1. Reversibility of phagocytosis inhibition by Tat. Human
FIGURE S1. Representative images to illustrate RAD51 foci induction in FANCD2 wild-type (wt) and
 Supplementary Figures FIGURE S1. Representative images to illustrate RAD51 foci induction in FANCD2 wild-type (wt) and mutant (mut) cells. Higher magnification is shown in Figure 1A. Arrows indicate cisplatin-induced
Supplementary Figures FIGURE S1. Representative images to illustrate RAD51 foci induction in FANCD2 wild-type (wt) and mutant (mut) cells. Higher magnification is shown in Figure 1A. Arrows indicate cisplatin-induced
We performed RT-PCR, cloning, sequencing and qrt-pcr in murine melanoma. cell lines and melanocytic tumors from RET-mice in accordance with the method
 Supplementary Material and Methods Quantitative RT-PCR (qrt-pcr) We performed RT-PCR, cloning, sequencing and qrt-pcr in murine melanoma cell lines and melanocytic tumors from RET-mice in accordance with
Supplementary Material and Methods Quantitative RT-PCR (qrt-pcr) We performed RT-PCR, cloning, sequencing and qrt-pcr in murine melanoma cell lines and melanocytic tumors from RET-mice in accordance with
Supplementary Figure S1
 Supplementary Figure S1 Supplementary Figure S1. CD11b expression on different B cell subsets in anti-snrnp Ig Tg mice. (a) Highly purified follicular (FO) and marginal zone (MZ) B cells were sorted from
Supplementary Figure S1 Supplementary Figure S1. CD11b expression on different B cell subsets in anti-snrnp Ig Tg mice. (a) Highly purified follicular (FO) and marginal zone (MZ) B cells were sorted from
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION Supplementary Figure S1. Efficacy of STIM1 knockdown by electroporation of STIM1 sirnas. (a) Western blot of FDB muscles from 4 mice treated with either STIM1-specific sirna (KD)
SUPPLEMENTARY INFORMATION Supplementary Figure S1. Efficacy of STIM1 knockdown by electroporation of STIM1 sirnas. (a) Western blot of FDB muscles from 4 mice treated with either STIM1-specific sirna (KD)
Supporting Information
 Supporting Information He et al. 10.1073/pnas.1116302108 SI Methods Cell Culture. Mouse J774A.1 and RAW 264.7 macrophages were obtained from ATCC and were cultured in MEM supplemented with 10% FS (Sigma)
Supporting Information He et al. 10.1073/pnas.1116302108 SI Methods Cell Culture. Mouse J774A.1 and RAW 264.7 macrophages were obtained from ATCC and were cultured in MEM supplemented with 10% FS (Sigma)
Supplemental material
 Supplemental material THE JOURNAL OF CELL BIOLOGY Taylor et al., http://www.jcb.org/cgi/content/full/jcb.201403021/dc1 Figure S1. Representative images of Cav 1a -YFP mutants with and without LMB treatment.
Supplemental material THE JOURNAL OF CELL BIOLOGY Taylor et al., http://www.jcb.org/cgi/content/full/jcb.201403021/dc1 Figure S1. Representative images of Cav 1a -YFP mutants with and without LMB treatment.
A novel tool for monitoring endogenous alpha-synuclein transcription by NanoLuciferase
 A novel tool for monitoring endogenous alpha-synuclein transcription by NanoLuciferase tag insertion at the 3 end using CRISPR-Cas9 genome editing technique Sambuddha Basu 1, 3, Levi Adams 1, 3, Subhrangshu
A novel tool for monitoring endogenous alpha-synuclein transcription by NanoLuciferase tag insertion at the 3 end using CRISPR-Cas9 genome editing technique Sambuddha Basu 1, 3, Levi Adams 1, 3, Subhrangshu
Flow-cytometry assessment of parasitemia in cell parasite co-culture assay using hydroethidine staining Western blotting RT-PCR
 Flow-cytometry assessment of parasitemia in cell parasite co-culture assay using hydroethidine staining The capability of cells, either PBMC or purified T- cell lines, to inhibit parasite growth or merozoite
Flow-cytometry assessment of parasitemia in cell parasite co-culture assay using hydroethidine staining The capability of cells, either PBMC or purified T- cell lines, to inhibit parasite growth or merozoite
Supplementary Table 1. Primers used to construct full-length or various truncated mutants of ISG12b2.
 Supplementary Table 1. Primers used to construct full-length or various truncated mutants of ISG12b2. Construct name ISG12b2 (No tag) HA-ISG12b2 (N-HA) ISG12b2-HA (C-HA; FL-HA) 94-283-HA (FL-GFP) 93-GFP
Supplementary Table 1. Primers used to construct full-length or various truncated mutants of ISG12b2. Construct name ISG12b2 (No tag) HA-ISG12b2 (N-HA) ISG12b2-HA (C-HA; FL-HA) 94-283-HA (FL-GFP) 93-GFP
Supplementary Table 1. Sequences for BTG2 and BRCA1 sirnas.
 Supplementary Table 1. Sequences for BTG2 and BRCA1 sirnas. Target Gene Non-target / Control BTG2 BRCA1 NFE2L2 Target Sequence ON-TARGET plus Non-targeting sirna # 1 (Cat# D-001810-01-05) sirna1: GAACCGACAUGCUCCCGGA
Supplementary Table 1. Sequences for BTG2 and BRCA1 sirnas. Target Gene Non-target / Control BTG2 BRCA1 NFE2L2 Target Sequence ON-TARGET plus Non-targeting sirna # 1 (Cat# D-001810-01-05) sirna1: GAACCGACAUGCUCCCGGA
A novel therapeutic strategy to rescue the immune effector function of proteolyticallyinactivated
 A novel therapeutic strategy to rescue the immune effector function of proteolyticallyinactivated cancer therapeutic antibodies Supplemental Data File in this Data Supplement: Supplementary Figure 1-4
A novel therapeutic strategy to rescue the immune effector function of proteolyticallyinactivated cancer therapeutic antibodies Supplemental Data File in this Data Supplement: Supplementary Figure 1-4
SANTA CRUZ BIOTECHNOLOGY, INC.
 TECHNICAL SERVICE GUIDE: Western Blotting 2. What size bands were expected and what size bands were detected? 3. Was the blot blank or was a dark background or non-specific bands seen? 4. Did this same
TECHNICAL SERVICE GUIDE: Western Blotting 2. What size bands were expected and what size bands were detected? 3. Was the blot blank or was a dark background or non-specific bands seen? 4. Did this same
Four different active promoter genes were chosen, ATXN7L2, PSRC1, CELSR2 and
 SUPPLEMENTARY MATERIALS AND METHODS Chromatin Immunoprecipitation for qpcr analysis Four different active promoter genes were chosen, ATXN7L2, PSRC1, CELSR2 and IL24, all located on chromosome 1. Primer
SUPPLEMENTARY MATERIALS AND METHODS Chromatin Immunoprecipitation for qpcr analysis Four different active promoter genes were chosen, ATXN7L2, PSRC1, CELSR2 and IL24, all located on chromosome 1. Primer
Supplementary Figure 1
 Supplementary Figure 1 Supplementary Fig. 1 shrna mediated knockdown of ZRSR2 in K562 and 293T cells. (a) ZRSR2 transcript levels in stably transduced K562 cells were determined using qrt-pcr. GAPDH was
Supplementary Figure 1 Supplementary Fig. 1 shrna mediated knockdown of ZRSR2 in K562 and 293T cells. (a) ZRSR2 transcript levels in stably transduced K562 cells were determined using qrt-pcr. GAPDH was
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb3240 Supplementary Figure 1 GBM cell lines display similar levels of p100 to p52 processing but respond differentially to TWEAK-induced TERT expression according to TERT promoter mutation
DOI: 10.1038/ncb3240 Supplementary Figure 1 GBM cell lines display similar levels of p100 to p52 processing but respond differentially to TWEAK-induced TERT expression according to TERT promoter mutation
SUPPLEMENTARY INFORMATION
 DOI:.38/ncb327 a b Sequence coverage (%) 4 3 2 IP: -GFP isoform IP: GFP IP: -GFP IP: GFP Sequence coverage (%) 4 3 2 IP: -GFP IP: GFP 33 52 58 isoform 2 33 49 47 IP: Control IP: Peptide Sequence Start
DOI:.38/ncb327 a b Sequence coverage (%) 4 3 2 IP: -GFP isoform IP: GFP IP: -GFP IP: GFP Sequence coverage (%) 4 3 2 IP: -GFP IP: GFP 33 52 58 isoform 2 33 49 47 IP: Control IP: Peptide Sequence Start
SUPPLEMENTARY INFORMATION
 The Supplementary Information (SI) Methods Cell culture and transfections H1299, U2OS, 293, HeLa cells were maintained in DMEM medium supplemented with 10% fetal bovine serum. H1299 and 293 cells were
The Supplementary Information (SI) Methods Cell culture and transfections H1299, U2OS, 293, HeLa cells were maintained in DMEM medium supplemented with 10% fetal bovine serum. H1299 and 293 cells were
Supporting Information
 Supporting Information Casson et al. 10.1073/pnas.1421699112 Sl Materials and Methods Macrophage Infections. In experiments where macrophages were primed with LPS, cells were pretreated with 0.5 μg/ml
Supporting Information Casson et al. 10.1073/pnas.1421699112 Sl Materials and Methods Macrophage Infections. In experiments where macrophages were primed with LPS, cells were pretreated with 0.5 μg/ml
Yan Zhu, Masha V. Poyurovsky, Yingchun Li, Lynn Biderman, Joachim Stahl, Xavier Jacq, and Carol Prives
 Molecular Cell, Volume 35 Supplemental Data Ribosomal Protein S7 Is Both a Regulator and a Substrate of MDM2 Yan Zhu, Masha V. Poyurovsky, Yingchun Li, Lynn Biderman, Joachim Stahl, Xavier Jacq, and Carol
Molecular Cell, Volume 35 Supplemental Data Ribosomal Protein S7 Is Both a Regulator and a Substrate of MDM2 Yan Zhu, Masha V. Poyurovsky, Yingchun Li, Lynn Biderman, Joachim Stahl, Xavier Jacq, and Carol
Site directed mutagenesis, Insertional and Deletion Mutagenesis. Mitesh Shrestha
 Site directed mutagenesis, Insertional and Deletion Mutagenesis Mitesh Shrestha Mutagenesis Mutagenesis (the creation or formation of a mutation) can be used as a powerful genetic tool. By inducing mutations
Site directed mutagenesis, Insertional and Deletion Mutagenesis Mitesh Shrestha Mutagenesis Mutagenesis (the creation or formation of a mutation) can be used as a powerful genetic tool. By inducing mutations
Supplementary Information for
 Supplementary Information for Siglec-7 Engagement by GBS -protein Suppresses Pyroptotic Cell Death of Natural Killer Cells Jerry J. Fong a,b, Chih-Ming Tsai a,b, Sudeshna Saha a,b, Victor Nizet a,c,d Ajit
Supplementary Information for Siglec-7 Engagement by GBS -protein Suppresses Pyroptotic Cell Death of Natural Killer Cells Jerry J. Fong a,b, Chih-Ming Tsai a,b, Sudeshna Saha a,b, Victor Nizet a,c,d Ajit
Blimp-1/PRDM1 rabbit monoclonal antibody (C14A4) was purchased from Cell Signaling
 1 2 3 4 5 6 7 8 9 10 11 Supplementary Methods Antibodies Blimp-1/PRDM1 rabbit monoclonal antibody (C14A4) was purchased from Cell Signaling (Danvers, MA) and used at 1:1000 to detect the total PRDM1 protein
1 2 3 4 5 6 7 8 9 10 11 Supplementary Methods Antibodies Blimp-1/PRDM1 rabbit monoclonal antibody (C14A4) was purchased from Cell Signaling (Danvers, MA) and used at 1:1000 to detect the total PRDM1 protein
High throughput screening: Huh-7 cells were seeded into 96-well plate (2000
 1 SUPPLEMENTARY INFORMATION METHODS 6 7 8 9 1 11 1 1 1 1 16 17 18 19 High throughput screening: Huh-7 cells were seeded into 96-well plate ( cells/well) and infected with MOI of DENV-. One hour post-infection
1 SUPPLEMENTARY INFORMATION METHODS 6 7 8 9 1 11 1 1 1 1 16 17 18 19 High throughput screening: Huh-7 cells were seeded into 96-well plate ( cells/well) and infected with MOI of DENV-. One hour post-infection
Erk activation drives intestinal tumorigenesis in Apc min/+ mice
 correction notice Nat. Med. 16, 665 670 (2010) Erk activation drives intestinal tumorigenesis in Apc min/+ mice Sung Hee Lee, Li-Li Hu, Jose Gonzalez-Navajas, Geom Seog Seo, Carol Shen, Jonathan Brick,
correction notice Nat. Med. 16, 665 670 (2010) Erk activation drives intestinal tumorigenesis in Apc min/+ mice Sung Hee Lee, Li-Li Hu, Jose Gonzalez-Navajas, Geom Seog Seo, Carol Shen, Jonathan Brick,
Data Supplement Plasmin Triggers Cytokine Induction in Human Monocyte-Derived Macrophages
 Data Supplement Plasmin Triggers Cytokine Induction in Human Monocyte-Derived Macrophages Qun Li, Yves Laumonnier, Tatiana Syrovets, and Thomas Simmet Methods Antibodies and Reagents Antibodies for immunoblotting:
Data Supplement Plasmin Triggers Cytokine Induction in Human Monocyte-Derived Macrophages Qun Li, Yves Laumonnier, Tatiana Syrovets, and Thomas Simmet Methods Antibodies and Reagents Antibodies for immunoblotting:
Figure S1. Verification of ihog Mutation by Protein Immunoblotting Figure S2. Verification of ihog and boi
 Figure S1. Verification of ihog Mutation by Protein Immunoblotting Extracts from S2R+ cells, embryos, and adults were analyzed by immunoprecipitation and immunoblotting with anti-ihog antibody. The Ihog
Figure S1. Verification of ihog Mutation by Protein Immunoblotting Extracts from S2R+ cells, embryos, and adults were analyzed by immunoprecipitation and immunoblotting with anti-ihog antibody. The Ihog
Supplementary Materials and Methods
 Supplementary Materials and Methods Reagents Supplementary Material (ESI) for Lab on a Chip RPMI medium, FBS, HEPES buffer solution, sodium pyruvate, penicillin, and streptomycin were obtained from Biological
Supplementary Materials and Methods Reagents Supplementary Material (ESI) for Lab on a Chip RPMI medium, FBS, HEPES buffer solution, sodium pyruvate, penicillin, and streptomycin were obtained from Biological
Supplementary material and methods
 Inhibitory effect of caffeic acid on ADP-induced thrombus formation and platelet activation involves mitogen-activated protein kinases Yu Lu 1,2,3,#, Quan Li 3,4,#, Yu-Ying Liu 3,4, Kai Sun 3,4, Jing-Yu
Inhibitory effect of caffeic acid on ADP-induced thrombus formation and platelet activation involves mitogen-activated protein kinases Yu Lu 1,2,3,#, Quan Li 3,4,#, Yu-Ying Liu 3,4, Kai Sun 3,4, Jing-Yu
Figure 1: TDP-43 is subject to lysine acetylation within the RNA-binding domain a) QBI-293 cells were transfected with TDP-43 in the presence or
 Figure 1: TDP-43 is subject to lysine acetylation within the RNA-binding domain a) QBI-293 cells were transfected with TDP-43 in the presence or absence of the acetyltransferase CBP and acetylated TDP-43
Figure 1: TDP-43 is subject to lysine acetylation within the RNA-binding domain a) QBI-293 cells were transfected with TDP-43 in the presence or absence of the acetyltransferase CBP and acetylated TDP-43
Supplemental Materials and Methods
 Supplemental Materials and Methods In situ hybridization In situ hybridization analysis of HFE2 and genin mrna in rat liver tissues was performed as previously described (1). Briefly, the digoxigenin-labeled
Supplemental Materials and Methods In situ hybridization In situ hybridization analysis of HFE2 and genin mrna in rat liver tissues was performed as previously described (1). Briefly, the digoxigenin-labeled
TRIM31 is recruited to mitochondria after infection with SeV.
 Supplementary Figure 1 TRIM31 is recruited to mitochondria after infection with SeV. (a) Confocal microscopy of TRIM31-GFP transfected into HEK293T cells for 24 h followed with SeV infection for 6 h. MitoTracker
Supplementary Figure 1 TRIM31 is recruited to mitochondria after infection with SeV. (a) Confocal microscopy of TRIM31-GFP transfected into HEK293T cells for 24 h followed with SeV infection for 6 h. MitoTracker
Supplemental Information
 Supplemental Information Intrinsic protein-protein interaction mediated and chaperonin assisted sequential assembly of a stable Bardet Biedl syndome protein complex, the BBSome * Qihong Zhang 1#, Dahai
Supplemental Information Intrinsic protein-protein interaction mediated and chaperonin assisted sequential assembly of a stable Bardet Biedl syndome protein complex, the BBSome * Qihong Zhang 1#, Dahai
Supplementary Figure 1 a
 3 min PMA 45 min PMA AnnexinV-FITC Supplementary Figure 1 5 min PMA 15 min PMA a 9 min PMA 12 min PMA 5 min FGF7 15 min FGF7 3 min FGF7 6 min FGF7 9 min FGF7 12 min FGF7 5 min control 3 min control 6 min
3 min PMA 45 min PMA AnnexinV-FITC Supplementary Figure 1 5 min PMA 15 min PMA a 9 min PMA 12 min PMA 5 min FGF7 15 min FGF7 3 min FGF7 6 min FGF7 9 min FGF7 12 min FGF7 5 min control 3 min control 6 min
Arf6 Activation Assay Kit
 A helping hand for your research Product Manual Configuration-specific Monoclonal Antibody Based Arf6 Activation Assay Kit Catalog Number: 82401 20 assays NewEast Biosciences 1 Table of Content Product
A helping hand for your research Product Manual Configuration-specific Monoclonal Antibody Based Arf6 Activation Assay Kit Catalog Number: 82401 20 assays NewEast Biosciences 1 Table of Content Product
To examine the silencing effects of Celsr3 shrna, we co transfected 293T cells with expression
 Supplemental figures Supplemental Figure. 1. Silencing expression of Celsr3 by shrna. To examine the silencing effects of Celsr3 shrna, we co transfected 293T cells with expression plasmids for the shrna
Supplemental figures Supplemental Figure. 1. Silencing expression of Celsr3 by shrna. To examine the silencing effects of Celsr3 shrna, we co transfected 293T cells with expression plasmids for the shrna
Supplementary Information
 Supplementary Information Supplementary Figures Supplementary Figure 1. MLK1-4 phosphorylate MEK in the presence of RAF inhibitors. (a) H157 cells were transiently transfected with Flag- or HA-tagged MLK1-4
Supplementary Information Supplementary Figures Supplementary Figure 1. MLK1-4 phosphorylate MEK in the presence of RAF inhibitors. (a) H157 cells were transiently transfected with Flag- or HA-tagged MLK1-4
T H E J O U R N A L O F C E L L B I O L O G Y
 T H E J O U R N A L O F C E L L B I O L O G Y Supplemental material Nakajima and Tanoue, http://www.jcb.org/cgi/content/full/jcb.201104118/dc1 Figure S1. DLD-1 cells exhibit the characteristic morphology
T H E J O U R N A L O F C E L L B I O L O G Y Supplemental material Nakajima and Tanoue, http://www.jcb.org/cgi/content/full/jcb.201104118/dc1 Figure S1. DLD-1 cells exhibit the characteristic morphology
Generation of mouse BMDCs Vectors Short hairpin RNA constructs cdna constructs and generation of stable cell lines
 Generation of mouse BMDCs Mouse BMDCs were differentiated from BM cells isolated from femur and tibiae of the hind legs. Cell suspensions were filtered through a 40 µm cell strainer and incubated for 2
Generation of mouse BMDCs Mouse BMDCs were differentiated from BM cells isolated from femur and tibiae of the hind legs. Cell suspensions were filtered through a 40 µm cell strainer and incubated for 2
Cytotoxicity of Botulinum Neurotoxins Reveals a Direct Role of
 Supplementary Information Cytotoxicity of Botulinum Neurotoxins Reveals a Direct Role of Syntaxin 1 and SNAP-25 in Neuron Survival Lisheng Peng, Huisheng Liu, Hongyu Ruan, William H. Tepp, William H. Stoothoff,
Supplementary Information Cytotoxicity of Botulinum Neurotoxins Reveals a Direct Role of Syntaxin 1 and SNAP-25 in Neuron Survival Lisheng Peng, Huisheng Liu, Hongyu Ruan, William H. Tepp, William H. Stoothoff,
Genome-wide CRISPR screen reveals novel host factors required for Staphylococcus aureus α-hemolysin-mediated toxicity
 Genome-wide CRISPR screen reveals novel host factors required for Staphylococcus aureus α-hemolysin-mediated toxicity Sebastian Virreira Winter, Arturo Zychlinsky and Bart W. Bardoel Department of Cellular
Genome-wide CRISPR screen reveals novel host factors required for Staphylococcus aureus α-hemolysin-mediated toxicity Sebastian Virreira Winter, Arturo Zychlinsky and Bart W. Bardoel Department of Cellular
Impairment of Alveolar Macrophage Transcription in. Idiopathic Pulmonary Fibrosis. Online Data Supplement
 Impairment of Alveolar Macrophage Transcription in Idiopathic Pulmonary Fibrosis Online Data Supplement Ping Ren, Ivan O. Rosas, Sandra D. MacDonald, Hai-Ping Wu, Eric M. Billings, and Bernadette R. Gochuico
Impairment of Alveolar Macrophage Transcription in Idiopathic Pulmonary Fibrosis Online Data Supplement Ping Ren, Ivan O. Rosas, Sandra D. MacDonald, Hai-Ping Wu, Eric M. Billings, and Bernadette R. Gochuico
Cdc42 Activation Assay Kit
 A helping hand for your research Product Manual Configuration-specific Monoclonal Antibody Based Cdc42 Activation Assay Kit Catalog Number: 80701 20 assays 1 Table of Content Product Description 3 Assay
A helping hand for your research Product Manual Configuration-specific Monoclonal Antibody Based Cdc42 Activation Assay Kit Catalog Number: 80701 20 assays 1 Table of Content Product Description 3 Assay
Supplementary Figure 1. jmj30-2 and jmj32-1 produce null mutants. (a) Schematic drawing of JMJ30 and JMJ32 genome structure showing regions amplified
 Supplementary Figure 1. jmj30-2 and jmj32-1 produce null mutants. (a) Schematic drawing of JMJ30 and JMJ32 genome structure showing regions amplified by primers used for mrna expression analysis. Gray
Supplementary Figure 1. jmj30-2 and jmj32-1 produce null mutants. (a) Schematic drawing of JMJ30 and JMJ32 genome structure showing regions amplified by primers used for mrna expression analysis. Gray
RayBio Phospho- Akt (Ser473) ELISA Kit
 RayBio Phospho- Akt (Ser473) ELISA Kit For Measuring Phosphorylated Akt (Ser473) in Human, Mouse and Rat Cell Lysates User Manual (Revised Mar 1, 2012) RayBio Akt (Ser473) ELISA Kit Protocol (Cat#: PEL-Akt-S473-001)
RayBio Phospho- Akt (Ser473) ELISA Kit For Measuring Phosphorylated Akt (Ser473) in Human, Mouse and Rat Cell Lysates User Manual (Revised Mar 1, 2012) RayBio Akt (Ser473) ELISA Kit Protocol (Cat#: PEL-Akt-S473-001)
Supplementary Figure 1. Intracellular distribution of the EPE peptide. HeLa cells were serum-starved (16 h, 0.1%), and treated with EPE peptide,
 Supplementary Figure 1. Intracellular distribution of the EPE peptide. HeLa cells were serum-starved (16 h, 0.1%), and treated with EPE peptide, conjugated with either TAT or Myristic acid and biotin for
Supplementary Figure 1. Intracellular distribution of the EPE peptide. HeLa cells were serum-starved (16 h, 0.1%), and treated with EPE peptide, conjugated with either TAT or Myristic acid and biotin for
Supplementary Figure S1. The tetracycline-inducible CRISPR system. A) Hela cells stably
 Supplementary Information Supplementary Figure S1. The tetracycline-inducible CRISPR system. A) Hela cells stably expressing shrna sequences against TRF2 were examined by western blotting. shcon, shrna
Supplementary Information Supplementary Figure S1. The tetracycline-inducible CRISPR system. A) Hela cells stably expressing shrna sequences against TRF2 were examined by western blotting. shcon, shrna
ab Hypoxic Response Human Flow Cytometry Kit
 ab126585 Hypoxic Response Human Flow Cytometry Kit Instructions for Use For measuring protein levels by flow cytometry: hypoxia-inducible factor 1-alpha (HIF1A) and BCL2/adenovirus E1B 19 kda proteininteracting
ab126585 Hypoxic Response Human Flow Cytometry Kit Instructions for Use For measuring protein levels by flow cytometry: hypoxia-inducible factor 1-alpha (HIF1A) and BCL2/adenovirus E1B 19 kda proteininteracting
We used HEK293T cells for lentiviral supernatant production. HEK293T cells were
 Lentiviral transduction of CD34 + cells and cell lines We used HEK293T cells for lentiviral supernatant production. HEK293T cells were maintained in DMEM supplemented with 1% FCS, 1 U/ml penicillin-streptomycin,
Lentiviral transduction of CD34 + cells and cell lines We used HEK293T cells for lentiviral supernatant production. HEK293T cells were maintained in DMEM supplemented with 1% FCS, 1 U/ml penicillin-streptomycin,
