Cell Stem Cell, volume 9 Supplemental Information
|
|
|
- Julius Lucas
- 6 years ago
- Views:
Transcription
1 Cell Stem Cell, volume 9 Supplemental Information Large-Scale Analysis Reveals Acquisition of Lineage-Specific Chromosomal Aberrations in Human Adult Stem Cells Uri Ben-David, Yoav Mayshar, and Nissim Benvenisty 1
2 2
3 3
4 Legend to Supplementary Figures and Tables Supplementary Figure 1: Further identification of chromosomal aberrations in human stem cells. (a-b) Identification of chromosomal aberrations in human mesenchymal stem cells: bone marrow-derived MSC line, A18, demonstrates monosomy 6q (a, red line, p=8xe-5) and trisomy 19 (b, red line, p=6xe-20). Another MSC cell line from the same study, A23, is presented as control (blue line). (c) Identification of chromosomal aberrations in human neural stem cells: hesc-derived NSC line, H9 hes cell derived d17 definitive neural epithelial (pool #3), demonstrates trisomy 19 (red line, p=10xe-15). Another NSC sample from the same study, H9 hes cell derived d17 definitive neural epithelial (pool #2), is presented as control (blue line). (d) Identification of chromosomal aberrations in human hematopoietic stem/progenitor cells: bone marrow-derived CD34+ sample from MDS patient27 demonstrates trisomy 8 (red line, p=1xe- 15). 17 healthy donors-derived CD34+ samples from the same study are presented as controls (blue lines). Related to Figure 1. Supplementary Figure 2: Ideogram of the common chromosomal aberrations in various types of stem cells. Each chromosome is colored by the type/s of stem cells in which it was found to be aberrant in the current study. Stem cells are color-coded, so that red represents PSCs, green represents MSCs, and blue represents NSCs. Asterisks indicate common aberrations, found here or reported elsewhere in at least two independent studies. For instance, monosomy 6q was found in two independent studies of MSCs, and is therefore marked with a green asterisk. Related to Figure 2. Supplementary Table 1: A complete list of chromosomal aberrations identified in PSCs, MSCs, NSCs, and HSPCs. All stem cell samples that passed quality control were analyzed by the CGH-explorer software. Whenever aberrations were detected, the samples were subjected to a second bioinformatic test of chromosomal location enrichment, using the Expander tool (see Supplemental Experimental Procedures). Results of both analyses are listed in the table. Passages are mentioned wherever data were available. Previous reports on the chromosomal integrity of the cell lines are also mentioned where data were available. Related to Figures 1 and 2. 4
5 Supplemental Experimental Procedures Gene expression profiles database Gene expression profiles from studies which involved human embryonic stem cells (hescs), human induced pluripotent stem cells (hipscs), neural stem/progenitor cells (NSCs), mesenchymal stem cells (MSCs) and CD34+ hematopoietic stem/progenitor cells (HSPCs), and which conducted DNA microarray analysis using HG_U95A,HG-U133plus2, HG-ST1.0 (Affymetrix, CA) or HumanRef-8v3.0 (Illumina, CA) microarray platforms, were obtained from the GEO ( and from EMBL-EBI ( databases. Raw.CEL files for all samples were analyzed using MAS5 (for HG-U133plus2 arrays) or RMA (for HG-ST1.0 arrays) probeset condensation algorithm using Expression Console (Affymetrix, CA). Arrays were analyzed for quality control and outliers removed. Further outliers were removed following hierarchical clustering analysis. Normalized files were obtained for HG_U95A and HumanRef-8v3.0 arrays. Thus, the final dataset consisted of 208 samples of pluripotent stem cells, 144 samples of MSCs, 97 samples of NSCs and 177 samples of HSPCs, from 58 independent studies (See Table S1). For each cell type, in the HG-U133plus2 and HG_U95A platforms, probes absent in over 20% of the samples were discarded. In the HG- ST1.0 and HumanRef-8v3.0 platforms, probes whose expression was lower than 5.0 or 50 (respectively) in over 20% of the samples were discarded. In the case of multiple probesets for any given gene, multiple instances were discarded, so that each gene would be represented by one probeset only. Whenever possible, probesets ending with _at were used for the analysis (otherwise probesets were randomly selected). Probesets without documented chromosomal location were also removed. Thus, separate datasets containing a single probeset for each expressed gene were generated for each stem cell group. In order to reduce bias due to low expression levels, values under 50 (for HG-U133plus2, HG_U95A, and HumanRef-8v3.0) or 6.0 (for HG-ST1.0) were collectively raised to this level. In order to further reduce noise, the sum of squares of the relative expression values was calculated for each gene and highly variable genes (SSQ>150 for MSCs and HSPCs, SSQ>10 for NSCs) were removed as well. CGH-PCF over-expression analysis For each sample, the expression value of each gene was divided by the median of the same gene across the entire dataset, in order to obtain a comparative value. In order to reduce possible bias from any given experiment, groups of similar samples with highly similar gene expression profiles (as judged by hierarchical clustering) were averaged for the sake of calculating the grand population median. This median then served as the baseline for examining expression bias. The data was then processed using a freely available comparative genomic hybridization (CGH) analysis software program, CGH-Explorer ( Papers/CGH). Gene expression regional bias was detected using the program s piecewise constant fit (PCF) algorithm, using a set of parameters as follows: Least allowed deviation = 5
6 ; Least allowed aberration size = 50-80; Winsorize at quantile = 0.001; Penalty = 10-12; Threshold = Deletions in chromosome 19 could not be accurately detected, and were thus removed from the analysis. Moving-average plots of lines and regions of interest were drawn using the moving-average fit tool. Location enrichment analysis For each sample in which an aberration was detected, a list of genes that are over-expressed (>1.5 fold, for trisomies) or under-expressed (<1.5 fold, for monosomies) relative to the median expression of that gene, was comprised. In order to test for enrichment of whole chromosomes or chromosome arms, this list was subjected to location enrichment analysis, using the Expander software ( Significance was determined as Bonferroni corrected p-values lower than 1.0E-4, which is the default value of the Expander program. Cancer database analysis Data from the Mitelman Database of Chromosome Aberrations and Gene Fusions in Cancer were derived using the National Cancer Institute Recurrent Chromosome Aberrations in Cancer Database Searcher ( For each tumor of interest, the number of cases with numerical aberrations in each chromosome was calculated. Unbalanced Chromosomal Abnormalities were added to the Numerical Chromosomal Trisomy Abnormalities when they involved gains or isochromosomes, and to the Numerical Chromosomal Monosomy Abnormalities when they involved deletions. The relative frequency of the aberrations in each chromosome was then calculated, separately for trisomies/gains/isochromosomes and for monosomies/deletions. Statistical analysis Hierarchical clustering was performed using Partek Genomics Suite version 6.3 (Partek, MO; RB1 expression values were compared between the diploid and the aneuploid lines using Student s t-test. The sensitivity of the analysis was estimated using known aberrations from the dataset, revealing a false negative rate of ~5% (34/36 aberrations correctly identified). In order to test the specificity of our methodology, randomized data were generated for each cell line using its own gene expression data. This was performed five times for each of the stem cell types, and no false positive aberrations were found. The chromosomal aberrations were tested for uniform distribution using Pearson s chi-square goodness-of-fit test (with simulations). Fisher s exact test (with simulations) was used to test the dependence between the type of aberration (i.e. gain or deletion) and the type of stem cell, and 6
7 was similarly used to test the dependence between the identity of the aberration (i.e. chromosomes involved) and the type of stem cell. All of these tests were performed using the R language and environment for statistical computing ( Similar aberrations in multiple samples from the same study were considered as one aberration for the purpose of all statistical analyses. 7
Outline. Analysis of Microarray Data. Most important design question. General experimental issues
 Outline Analysis of Microarray Data Lecture 1: Experimental Design and Data Normalization Introduction to microarrays Experimental design Data normalization Other data transformation Exercises George Bell,
Outline Analysis of Microarray Data Lecture 1: Experimental Design and Data Normalization Introduction to microarrays Experimental design Data normalization Other data transformation Exercises George Bell,
Gene Expression Data Analysis (I)
 Gene Expression Data Analysis (I) Bing Zhang Department of Biomedical Informatics Vanderbilt University bing.zhang@vanderbilt.edu Bioinformatics tasks Biological question Experiment design Microarray experiment
Gene Expression Data Analysis (I) Bing Zhang Department of Biomedical Informatics Vanderbilt University bing.zhang@vanderbilt.edu Bioinformatics tasks Biological question Experiment design Microarray experiment
Philippe Hupé 1,2. The R User Conference 2009 Rennes
 A suite of R packages for the analysis of DNA copy number microarray experiments Application in cancerology Philippe Hupé 1,2 1 UMR144 Institut Curie, CNRS 2 U900 Institut Curie, INSERM, Mines Paris Tech
A suite of R packages for the analysis of DNA copy number microarray experiments Application in cancerology Philippe Hupé 1,2 1 UMR144 Institut Curie, CNRS 2 U900 Institut Curie, INSERM, Mines Paris Tech
Analysis of Microarray Data
 Analysis of Microarray Data Lecture 1: Experimental Design and Data Normalization George Bell, Ph.D. Senior Bioinformatics Scientist Bioinformatics and Research Computing Whitehead Institute Outline Introduction
Analysis of Microarray Data Lecture 1: Experimental Design and Data Normalization George Bell, Ph.D. Senior Bioinformatics Scientist Bioinformatics and Research Computing Whitehead Institute Outline Introduction
CS 5984: Application of Basic Clustering Algorithms to Find Expression Modules in Cancer
 CS 5984: Application of Basic Clustering Algorithms to Find Expression Modules in Cancer T. M. Murali January 31, 2006 Innovative Application of Hierarchical Clustering A module map showing conditional
CS 5984: Application of Basic Clustering Algorithms to Find Expression Modules in Cancer T. M. Murali January 31, 2006 Innovative Application of Hierarchical Clustering A module map showing conditional
Analysis of a Tiling Regulation Study in Partek Genomics Suite 6.6
 Analysis of a Tiling Regulation Study in Partek Genomics Suite 6.6 The example data set used in this tutorial consists of 6 technical replicates from the same human cell line, 3 are SP1 treated, and 3
Analysis of a Tiling Regulation Study in Partek Genomics Suite 6.6 The example data set used in this tutorial consists of 6 technical replicates from the same human cell line, 3 are SP1 treated, and 3
Agilent GeneSpring GX 10: Beyond. Pam Tangvoranuntakul Product Manager, GeneSpring October 1, 2008
 Agilent GeneSpring GX 10: Gene Expression and Beyond Pam Tangvoranuntakul Product Manager, GeneSpring October 1, 2008 GeneSpring GX 10 in the News Our Goals for GeneSpring GX 10 Goal 1: Bring back GeneSpring
Agilent GeneSpring GX 10: Gene Expression and Beyond Pam Tangvoranuntakul Product Manager, GeneSpring October 1, 2008 GeneSpring GX 10 in the News Our Goals for GeneSpring GX 10 Goal 1: Bring back GeneSpring
FEATURE-LEVEL EXPLORATION OF THE CHOE ET AL. AFFYMETRIX GENECHIP CONTROL DATASET
 Johns Hopkins University, Dept. of Biostatistics Working Papers 3-17-2006 FEATURE-LEVEL EXPLORATION OF THE CHOE ET AL. AFFYETRIX GENECHIP CONTROL DATASET Rafael A. Irizarry Johns Hopkins Bloomberg School
Johns Hopkins University, Dept. of Biostatistics Working Papers 3-17-2006 FEATURE-LEVEL EXPLORATION OF THE CHOE ET AL. AFFYETRIX GENECHIP CONTROL DATASET Rafael A. Irizarry Johns Hopkins Bloomberg School
Analysis of Microarray Data
 Analysis of Microarray Data Lecture 1: Experimental Design and Data Normalization George Bell, Ph.D. Senior Bioinformatics Scientist Bioinformatics and Research Computing Whitehead Institute Outline Introduction
Analysis of Microarray Data Lecture 1: Experimental Design and Data Normalization George Bell, Ph.D. Senior Bioinformatics Scientist Bioinformatics and Research Computing Whitehead Institute Outline Introduction
Detecting copy-neutral LOH in cancer using Agilent SurePrint G3 Cancer CGH+SNP Microarrays
 Detecting copy-neutral LOH in cancer using Agilent SurePrint G3 Cancer CGH+SNP Microarrays Application Note Authors Paula Costa Anniek De Witte Jayati Ghosh Agilent Technologies, Inc. Santa Clara, CA USA
Detecting copy-neutral LOH in cancer using Agilent SurePrint G3 Cancer CGH+SNP Microarrays Application Note Authors Paula Costa Anniek De Witte Jayati Ghosh Agilent Technologies, Inc. Santa Clara, CA USA
New Statistical Algorithms for Monitoring Gene Expression on GeneChip Probe Arrays
 GENE EXPRESSION MONITORING TECHNICAL NOTE New Statistical Algorithms for Monitoring Gene Expression on GeneChip Probe Arrays Introduction Affymetrix has designed new algorithms for monitoring GeneChip
GENE EXPRESSION MONITORING TECHNICAL NOTE New Statistical Algorithms for Monitoring Gene Expression on GeneChip Probe Arrays Introduction Affymetrix has designed new algorithms for monitoring GeneChip
A Distribution Free Summarization Method for Affymetrix GeneChip Arrays
 A Distribution Free Summarization Method for Affymetrix GeneChip Arrays Zhongxue Chen 1,2, Monnie McGee 1,*, Qingzhong Liu 3, and Richard Scheuermann 2 1 Department of Statistical Science, Southern Methodist
A Distribution Free Summarization Method for Affymetrix GeneChip Arrays Zhongxue Chen 1,2, Monnie McGee 1,*, Qingzhong Liu 3, and Richard Scheuermann 2 1 Department of Statistical Science, Southern Methodist
Introduction to gene expression microarray data analysis
 Introduction to gene expression microarray data analysis Outline Brief introduction: Technology and data. Statistical challenges in data analysis. Preprocessing data normalization and transformation. Useful
Introduction to gene expression microarray data analysis Outline Brief introduction: Technology and data. Statistical challenges in data analysis. Preprocessing data normalization and transformation. Useful
Effects of Multiple Probesets in Affymetrix GeneChips on Identifying Differentially Expressed Genes in ips Cells
 The Fourth International Conference on Computational Systems Biology (ISB2010) Suzhou, China, September 9 11, 2010 Copyright 2010 ORSC & APORC, pp. 187 195 Effects of Multiple Probesets in Affymetrix GeneChips
The Fourth International Conference on Computational Systems Biology (ISB2010) Suzhou, China, September 9 11, 2010 Copyright 2010 ORSC & APORC, pp. 187 195 Effects of Multiple Probesets in Affymetrix GeneChips
Supporting Online Material for
 www.sciencemag.org/cgi/content/full/313/5795/1929/dc1 Supporting Online Material for The Connectivity Map: Using Gene-Expression Signatures to Connect Small Molecules, Genes, and Disease Justin Lamb, *
www.sciencemag.org/cgi/content/full/313/5795/1929/dc1 Supporting Online Material for The Connectivity Map: Using Gene-Expression Signatures to Connect Small Molecules, Genes, and Disease Justin Lamb, *
Analysis of Microarray Data
 Analysis of Microarray Data Lecture 3: Visualization and Functional Analysis George Bell, Ph.D. Senior Bioinformatics Scientist Bioinformatics and Research Computing Whitehead Institute Outline Review
Analysis of Microarray Data Lecture 3: Visualization and Functional Analysis George Bell, Ph.D. Senior Bioinformatics Scientist Bioinformatics and Research Computing Whitehead Institute Outline Review
Comparative Genomic Hybridization
 Comparative Genomic Hybridization Srikesh G. Arunajadai Division of Biostatistics University of California Berkeley PH 296 Presentation Fall 2002 December 9 th 2002 OUTLINE CGH Introduction Methodology,
Comparative Genomic Hybridization Srikesh G. Arunajadai Division of Biostatistics University of California Berkeley PH 296 Presentation Fall 2002 December 9 th 2002 OUTLINE CGH Introduction Methodology,
Introduction to Microarray Analysis
 Introduction to Microarray Analysis Methods Course: Gene Expression Data Analysis -Day One Rainer Spang Microarrays Highly parallel measurement devices for gene expression levels 1. How does the microarray
Introduction to Microarray Analysis Methods Course: Gene Expression Data Analysis -Day One Rainer Spang Microarrays Highly parallel measurement devices for gene expression levels 1. How does the microarray
Humboldt Universität zu Berlin. Grundlagen der Bioinformatik SS Microarrays. Lecture
 Humboldt Universität zu Berlin Microarrays Grundlagen der Bioinformatik SS 2017 Lecture 6 09.06.2017 Agenda 1.mRNA: Genomic background 2.Overview: Microarray 3.Data-analysis: Quality control & normalization
Humboldt Universität zu Berlin Microarrays Grundlagen der Bioinformatik SS 2017 Lecture 6 09.06.2017 Agenda 1.mRNA: Genomic background 2.Overview: Microarray 3.Data-analysis: Quality control & normalization
Background Correction and Normalization. Lecture 3 Computational and Statistical Aspects of Microarray Analysis June 21, 2005 Bressanone, Italy
 Background Correction and Normalization Lecture 3 Computational and Statistical Aspects of Microarray Analysis June 21, 2005 Bressanone, Italy Feature Level Data Outline Affymetrix GeneChip arrays Two
Background Correction and Normalization Lecture 3 Computational and Statistical Aspects of Microarray Analysis June 21, 2005 Bressanone, Italy Feature Level Data Outline Affymetrix GeneChip arrays Two
V3 differential gene expression analysis
 differential gene expression analysis - What is measured by microarrays? - Microarray normalization - Differential gene expression (DE) analysis based on microarray data - Detection of outliers - RNAseq
differential gene expression analysis - What is measured by microarrays? - Microarray normalization - Differential gene expression (DE) analysis based on microarray data - Detection of outliers - RNAseq
Background and Normalization:
 Background and Normalization: Investigating the effects of preprocessing on gene expression estimates Ben Bolstad Group in Biostatistics University of California, Berkeley bolstad@stat.berkeley.edu http://www.stat.berkeley.edu/~bolstad
Background and Normalization: Investigating the effects of preprocessing on gene expression estimates Ben Bolstad Group in Biostatistics University of California, Berkeley bolstad@stat.berkeley.edu http://www.stat.berkeley.edu/~bolstad
Characterization of Allele-Specific Copy Number in Tumor Genomes
 Characterization of Allele-Specific Copy Number in Tumor Genomes Hao Chen 2 Haipeng Xing 1 Nancy R. Zhang 2 1 Department of Statistics Stonybrook University of New York 2 Department of Statistics Stanford
Characterization of Allele-Specific Copy Number in Tumor Genomes Hao Chen 2 Haipeng Xing 1 Nancy R. Zhang 2 1 Department of Statistics Stonybrook University of New York 2 Department of Statistics Stanford
Copy Number Analysis in Partek Genomics Suite 6.6
 Copy Number Analysis in Partek Genomics Suite 6.6 Introduction At a simple level, copy number analysis can be thought of as a gene dosage question: where are the regions of the genome that have altered
Copy Number Analysis in Partek Genomics Suite 6.6 Introduction At a simple level, copy number analysis can be thought of as a gene dosage question: where are the regions of the genome that have altered
Analyzing Affymetrix GeneChip SNP 6 Copy Number Data in Partek
 Analyzing Affymetrix GeneChip SNP 6 Copy Number Data in Partek This example data set consists of 20 selected HapMap samples, representing 10 females and 10 males, drawn from a mixed ethnic population of
Analyzing Affymetrix GeneChip SNP 6 Copy Number Data in Partek This example data set consists of 20 selected HapMap samples, representing 10 females and 10 males, drawn from a mixed ethnic population of
High-Resolution Oligonucleotide- Based acgh Analysis of Single Cells in Under 24 Hours
 High-Resolution Oligonucleotide- Based acgh Analysis of Single Cells in Under 24 Hours Application Note Authors Paula Costa and Anniek De Witte Agilent Technologies, Inc. Santa Clara, CA USA Abstract As
High-Resolution Oligonucleotide- Based acgh Analysis of Single Cells in Under 24 Hours Application Note Authors Paula Costa and Anniek De Witte Agilent Technologies, Inc. Santa Clara, CA USA Abstract As
Mutations, Genetic Testing and Engineering
 Mutations, Genetic Testing and Engineering Objectives Describe how techniques such as DNA fingerprinting, genetic modifications, and chromosomal analysis are used to study the genomes of organisms (TEKS
Mutations, Genetic Testing and Engineering Objectives Describe how techniques such as DNA fingerprinting, genetic modifications, and chromosomal analysis are used to study the genomes of organisms (TEKS
Gene expression: Microarray data analysis. Copyright notice. Outline: microarray data analysis. Schedule
 Gene expression: Microarray data analysis Copyright notice Many of the images in this powerpoint presentation are from Bioinformatics and Functional Genomics by Jonathan Pevsner (ISBN -47-4-8). Copyright
Gene expression: Microarray data analysis Copyright notice Many of the images in this powerpoint presentation are from Bioinformatics and Functional Genomics by Jonathan Pevsner (ISBN -47-4-8). Copyright
Gene-Level Analysis of Exon Array Data using Partek Genomics Suite 6.6
 Gene-Level Analysis of Exon Array Data using Partek Genomics Suite 6.6 Overview This tutorial will demonstrate how to: Summarize core exon-level data to produce gene-level data Perform exploratory analysis
Gene-Level Analysis of Exon Array Data using Partek Genomics Suite 6.6 Overview This tutorial will demonstrate how to: Summarize core exon-level data to produce gene-level data Perform exploratory analysis
Chang Xu Mohammad R Nezami Ranjbar Zhong Wu John DiCarlo Yexun Wang
 Supplementary Materials for: Detecting very low allele fraction variants using targeted DNA sequencing and a novel molecular barcode-aware variant caller Chang Xu Mohammad R Nezami Ranjbar Zhong Wu John
Supplementary Materials for: Detecting very low allele fraction variants using targeted DNA sequencing and a novel molecular barcode-aware variant caller Chang Xu Mohammad R Nezami Ranjbar Zhong Wu John
Chromosome-scale scaffolding of de novo genome assemblies based on chromatin interactions. Supplementary Material
 Chromosome-scale scaffolding of de novo genome assemblies based on chromatin interactions Joshua N. Burton 1, Andrew Adey 1, Rupali P. Patwardhan 1, Ruolan Qiu 1, Jacob O. Kitzman 1, Jay Shendure 1 1 Department
Chromosome-scale scaffolding of de novo genome assemblies based on chromatin interactions Joshua N. Burton 1, Andrew Adey 1, Rupali P. Patwardhan 1, Ruolan Qiu 1, Jacob O. Kitzman 1, Jay Shendure 1 1 Department
Pre- Processing Methodologies for Microarray Gene Data for Cancer Detection
 IJSRD - International Journal for Scientific Research & Development Vol. 3, Issue 06, 205 ISSN (online): 232-063 Pre- Processing Methodologies for Microarray Gene Data for Cancer Detection Shashank K S
IJSRD - International Journal for Scientific Research & Development Vol. 3, Issue 06, 205 ISSN (online): 232-063 Pre- Processing Methodologies for Microarray Gene Data for Cancer Detection Shashank K S
Estimation problems in high throughput SNP platforms
 Estimation problems in high throughput SNP platforms Rob Scharpf Department of Biostatistics Johns Hopkins Bloomberg School of Public Health November, 8 Outline Introduction Introduction What is a SNP?
Estimation problems in high throughput SNP platforms Rob Scharpf Department of Biostatistics Johns Hopkins Bloomberg School of Public Health November, 8 Outline Introduction Introduction What is a SNP?
Oligonucleotide microarray data are not normally distributed
 Oligonucleotide microarray data are not normally distributed Johanna Hardin Jason Wilson John Kloke Abstract Novel techniques for analyzing microarray data are constantly being developed. Though many of
Oligonucleotide microarray data are not normally distributed Johanna Hardin Jason Wilson John Kloke Abstract Novel techniques for analyzing microarray data are constantly being developed. Though many of
Expression summarization
 Expression Quantification: Affy Affymetrix Genechip is an oligonucleotide array consisting of a several perfect match (PM) and their corresponding mismatch (MM) probes that interrogate for a single gene.
Expression Quantification: Affy Affymetrix Genechip is an oligonucleotide array consisting of a several perfect match (PM) and their corresponding mismatch (MM) probes that interrogate for a single gene.
Using the Association Workflow in Partek Genomics Suite
 Using the Association Workflow in Partek Genomics Suite This user guide will illustrate the use of the Association workflow in Partek Genomics Suite (PGS) and discuss the basic functions available within
Using the Association Workflow in Partek Genomics Suite This user guide will illustrate the use of the Association workflow in Partek Genomics Suite (PGS) and discuss the basic functions available within
Whole Transcriptome Analysis of Illumina RNA- Seq Data. Ryan Peters Field Application Specialist
 Whole Transcriptome Analysis of Illumina RNA- Seq Data Ryan Peters Field Application Specialist Partek GS in your NGS Pipeline Your Start-to-Finish Solution for Analysis of Next Generation Sequencing Data
Whole Transcriptome Analysis of Illumina RNA- Seq Data Ryan Peters Field Application Specialist Partek GS in your NGS Pipeline Your Start-to-Finish Solution for Analysis of Next Generation Sequencing Data
Quality Control Assessment in Genotyping Console
 Quality Control Assessment in Genotyping Console Introduction Prior to the release of Genotyping Console (GTC) 2.1, quality control (QC) assessment of the SNP Array 6.0 assay was performed using the Dynamic
Quality Control Assessment in Genotyping Console Introduction Prior to the release of Genotyping Console (GTC) 2.1, quality control (QC) assessment of the SNP Array 6.0 assay was performed using the Dynamic
DNA Microarray Data Oligonucleotide Arrays
 DNA Microarray Data Oligonucleotide Arrays Sandrine Dudoit, Robert Gentleman, Rafael Irizarry, and Yee Hwa Yang Bioconductor Short Course 2003 Copyright 2002, all rights reserved Biological question Experimental
DNA Microarray Data Oligonucleotide Arrays Sandrine Dudoit, Robert Gentleman, Rafael Irizarry, and Yee Hwa Yang Bioconductor Short Course 2003 Copyright 2002, all rights reserved Biological question Experimental
Preprocessing Affymetrix GeneChip Data. Affymetrix GeneChip Design. Terminology TGTGATGGTGGGGAATGGGTCAGAAGGCCTCCGATGCGCCGATTGAGAAT
 Preprocessing Affymetrix GeneChip Data Credit for some of today s materials: Ben Bolstad, Leslie Cope, Laurent Gautier, Terry Speed and Zhijin Wu Affymetrix GeneChip Design 5 3 Reference sequence TGTGATGGTGGGGAATGGGTCAGAAGGCCTCCGATGCGCCGATTGAGAAT
Preprocessing Affymetrix GeneChip Data Credit for some of today s materials: Ben Bolstad, Leslie Cope, Laurent Gautier, Terry Speed and Zhijin Wu Affymetrix GeneChip Design 5 3 Reference sequence TGTGATGGTGGGGAATGGGTCAGAAGGCCTCCGATGCGCCGATTGAGAAT
Microarray Technique. Some background. M. Nath
 Microarray Technique Some background M. Nath Outline Introduction Spotting Array Technique GeneChip Technique Data analysis Applications Conclusion Now Blind Guess? Functional Pathway Microarray Technique
Microarray Technique Some background M. Nath Outline Introduction Spotting Array Technique GeneChip Technique Data analysis Applications Conclusion Now Blind Guess? Functional Pathway Microarray Technique
massir: MicroArray Sample Sex Identifier
 massir: MicroArray Sample Sex Identifier Sam Buckberry October 30, 2017 Contents 1 The Problem 2 2 Importing data and beginning the analysis 2 3 Extracting the Y chromosome probe data 3 4 Predicting the
massir: MicroArray Sample Sex Identifier Sam Buckberry October 30, 2017 Contents 1 The Problem 2 2 Importing data and beginning the analysis 2 3 Extracting the Y chromosome probe data 3 4 Predicting the
Genetics - Problem Drill 19: Dissection of Gene Function: Mutational Analysis of Model Organisms
 Genetics - Problem Drill 19: Dissection of Gene Function: Mutational Analysis of Model Organisms No. 1 of 10 1. The mouse gene knockout is based on. (A) Homologous recombination (B) Site-specific recombination
Genetics - Problem Drill 19: Dissection of Gene Function: Mutational Analysis of Model Organisms No. 1 of 10 1. The mouse gene knockout is based on. (A) Homologous recombination (B) Site-specific recombination
Package citbcmst. February 19, 2015
 Package citbcmst February 19, 2015 Version 1.0.4 Date 2014-01-08 Title CIT Breast Cancer Molecular SubTypes Prediction This package implements the approach to assign tumor gene expression dataset to the
Package citbcmst February 19, 2015 Version 1.0.4 Date 2014-01-08 Title CIT Breast Cancer Molecular SubTypes Prediction This package implements the approach to assign tumor gene expression dataset to the
SUPPLEMENTARY INFORMATION
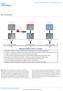 DOI: 10.1038/ncb2239 Hepatocytes (Endoderm) Foreskin fibroblasts (Mesoderm) Melanocytes (Ectoderm) +Dox rtta TetO CMV O,S,M,K & N H1 hes H7 hes H9 hes H1 hes-derived EBs H7 hes-derived EBs H9 hes-derived
DOI: 10.1038/ncb2239 Hepatocytes (Endoderm) Foreskin fibroblasts (Mesoderm) Melanocytes (Ectoderm) +Dox rtta TetO CMV O,S,M,K & N H1 hes H7 hes H9 hes H1 hes-derived EBs H7 hes-derived EBs H9 hes-derived
Probe-Level Data Normalisation: RMA and GC-RMA Sam Robson Images courtesy of Neil Ward, European Application Engineer, Agilent Technologies.
 Probe-Level Data Normalisation: RMA and GC-RMA Sam Robson Images courtesy of Neil Ward, European Application Engineer, Agilent Technologies. References Summaries of Affymetrix Genechip Probe Level Data,
Probe-Level Data Normalisation: RMA and GC-RMA Sam Robson Images courtesy of Neil Ward, European Application Engineer, Agilent Technologies. References Summaries of Affymetrix Genechip Probe Level Data,
MulCom: a Multiple Comparison statistical test for microarray data in Bioconductor.
 MulCom: a Multiple Comparison statistical test for microarray data in Bioconductor. Claudio Isella, Tommaso Renzulli, Davide Corà and Enzo Medico May 3, 2016 Abstract Many microarray experiments compare
MulCom: a Multiple Comparison statistical test for microarray data in Bioconductor. Claudio Isella, Tommaso Renzulli, Davide Corà and Enzo Medico May 3, 2016 Abstract Many microarray experiments compare
Introduction to Genome Wide Association Studies 2015 Sydney Brenner Institute for Molecular Bioscience Shaun Aron
 Introduction to Genome Wide Association Studies 2015 Sydney Brenner Institute for Molecular Bioscience Shaun Aron Many sources of technical bias in a genotyping experiment DNA sample quality and handling
Introduction to Genome Wide Association Studies 2015 Sydney Brenner Institute for Molecular Bioscience Shaun Aron Many sources of technical bias in a genotyping experiment DNA sample quality and handling
Compendium of Immune Signatures Identifies Conserved and Species-Specific Biology in Response to Inflammation
 Immunity Supplemental Information Compendium of Immune Signatures Identifies Conserved and Species-Specific Biology in Response to Inflammation Jernej Godec, Yan Tan, Arthur Liberzon, Pablo Tamayo, Sanchita
Immunity Supplemental Information Compendium of Immune Signatures Identifies Conserved and Species-Specific Biology in Response to Inflammation Jernej Godec, Yan Tan, Arthur Liberzon, Pablo Tamayo, Sanchita
Copynumber: Efficient algorithms for singleand multi-track copy number segmentation
 Nilsen et al. BMC Genomics 2012, 13:591 SOFTWARE Open Access Copynumber: Efficient algorithms for singleand multi-track copy number segmentation Gro Nilsen 1,2,KnutLiestøl 1,2, Peter Van Loo 3,4, Hans
Nilsen et al. BMC Genomics 2012, 13:591 SOFTWARE Open Access Copynumber: Efficient algorithms for singleand multi-track copy number segmentation Gro Nilsen 1,2,KnutLiestøl 1,2, Peter Van Loo 3,4, Hans
Introduction to Genome Wide Association Studies 2014 Sydney Brenner Institute for Molecular Bioscience/Wits Bioinformatics Shaun Aron
 Introduction to Genome Wide Association Studies 2014 Sydney Brenner Institute for Molecular Bioscience/Wits Bioinformatics Shaun Aron Genotype calling Genotyping methods for Affymetrix arrays Genotyping
Introduction to Genome Wide Association Studies 2014 Sydney Brenner Institute for Molecular Bioscience/Wits Bioinformatics Shaun Aron Genotype calling Genotyping methods for Affymetrix arrays Genotyping
Introduction to Bioinformatics. Fabian Hoti 6.10.
 Introduction to Bioinformatics Fabian Hoti 6.10. Analysis of Microarray Data Introduction Different types of microarrays Experiment Design Data Normalization Feature selection/extraction Clustering Introduction
Introduction to Bioinformatics Fabian Hoti 6.10. Analysis of Microarray Data Introduction Different types of microarrays Experiment Design Data Normalization Feature selection/extraction Clustering Introduction
Comp/Phys/APSc 715. Example Videos. Administrative 4/17/2014. Bioinformatics Visualization. Vis 2013, Schindler. Matlab bioinformatics toolbox
 Comp/Phys/APSc 715 Bioinformatics Visualization Example Videos Vis 2013, Schindler Lagrangian coherent structures in flow Matlab bioinformatics toolbox http://www.mathworks.com/videos/bioinformatic s-toolbox-overview-61196.html
Comp/Phys/APSc 715 Bioinformatics Visualization Example Videos Vis 2013, Schindler Lagrangian coherent structures in flow Matlab bioinformatics toolbox http://www.mathworks.com/videos/bioinformatic s-toolbox-overview-61196.html
STATC 141 Spring 2005, April 5 th Lecture notes on Affymetrix arrays. Materials are from
 STATC 141 Spring 2005, April 5 th Lecture notes on Affymetrix arrays Materials are from http://www.ohsu.edu/gmsr/amc/amc_technology.html The GeneChip high-density oligonucleotide arrays are fabricated
STATC 141 Spring 2005, April 5 th Lecture notes on Affymetrix arrays Materials are from http://www.ohsu.edu/gmsr/amc/amc_technology.html The GeneChip high-density oligonucleotide arrays are fabricated
Detection and Restoration of Hybridization Problems in Affymetrix GeneChip Data by Parametric Scanning
 100 Genome Informatics 17(2): 100-109 (2006) Detection and Restoration of Hybridization Problems in Affymetrix GeneChip Data by Parametric Scanning Tomokazu Konishi konishi@akita-pu.ac.jp Faculty of Bioresource
100 Genome Informatics 17(2): 100-109 (2006) Detection and Restoration of Hybridization Problems in Affymetrix GeneChip Data by Parametric Scanning Tomokazu Konishi konishi@akita-pu.ac.jp Faculty of Bioresource
Mixed effects model for assessing RNA degradation in Affymetrix GeneChip experiments
 Mixed effects model for assessing RNA degradation in Affymetrix GeneChip experiments Kellie J. Archer, Ph.D. Suresh E. Joel Viswanathan Ramakrishnan,, Ph.D. Department of Biostatistics Virginia Commonwealth
Mixed effects model for assessing RNA degradation in Affymetrix GeneChip experiments Kellie J. Archer, Ph.D. Suresh E. Joel Viswanathan Ramakrishnan,, Ph.D. Department of Biostatistics Virginia Commonwealth
What we ll do today. Types of stem cells. Do engineered ips and ES cells have. What genes are special in stem cells?
 Do engineered ips and ES cells have similar molecular signatures? What we ll do today Research questions in stem cell biology Comparing expression and epigenetics in stem cells asuring gene expression
Do engineered ips and ES cells have similar molecular signatures? What we ll do today Research questions in stem cell biology Comparing expression and epigenetics in stem cells asuring gene expression
Description of Logit-t: Detecting Differentially Expressed Genes Using Probe-Level Data
 Description of Logit-t: Detecting Differentially Expressed Genes Using Probe-Level Data Tobias Guennel October 22, 2008 Contents 1 Introduction 2 2 What s new in this version 3 3 Preparing data for use
Description of Logit-t: Detecting Differentially Expressed Genes Using Probe-Level Data Tobias Guennel October 22, 2008 Contents 1 Introduction 2 2 What s new in this version 3 3 Preparing data for use
BABELOMICS: Microarray Data Analysis
 BABELOMICS: Microarray Data Analysis Madrid, 21 June 2010 Martina Marbà mmarba@cipf.es Bioinformatics and Genomics Department Centro de Investigación Príncipe Felipe (CIPF) (Valencia, Spain) DNA Microarrays
BABELOMICS: Microarray Data Analysis Madrid, 21 June 2010 Martina Marbà mmarba@cipf.es Bioinformatics and Genomics Department Centro de Investigación Príncipe Felipe (CIPF) (Valencia, Spain) DNA Microarrays
Affymetrix GeneChip Arrays. Lecture 3 (continued) Computational and Statistical Aspects of Microarray Analysis June 21, 2005 Bressanone, Italy
 Affymetrix GeneChip Arrays Lecture 3 (continued) Computational and Statistical Aspects of Microarray Analysis June 21, 2005 Bressanone, Italy Affymetrix GeneChip Design 5 3 Reference sequence TGTGATGGTGGGGAATGGGTCAGAAGGCCTCCGATGCGCCGATTGAGAAT
Affymetrix GeneChip Arrays Lecture 3 (continued) Computational and Statistical Aspects of Microarray Analysis June 21, 2005 Bressanone, Italy Affymetrix GeneChip Design 5 3 Reference sequence TGTGATGGTGGGGAATGGGTCAGAAGGCCTCCGATGCGCCGATTGAGAAT
Reliable classification of two-class cancer data using evolutionary algorithms
 BioSystems 72 (23) 111 129 Reliable classification of two-class cancer data using evolutionary algorithms Kalyanmoy Deb, A. Raji Reddy Kanpur Genetic Algorithms Laboratory (KanGAL), Indian Institute of
BioSystems 72 (23) 111 129 Reliable classification of two-class cancer data using evolutionary algorithms Kalyanmoy Deb, A. Raji Reddy Kanpur Genetic Algorithms Laboratory (KanGAL), Indian Institute of
Do engineered ips and ES cells have similar molecular signatures?
 Do engineered ips and ES cells have similar molecular signatures? Comparing expression and epigenetics in stem cells George Bell, Ph.D. Bioinformatics and Research Computing 2012 Spring Lecture Series
Do engineered ips and ES cells have similar molecular signatures? Comparing expression and epigenetics in stem cells George Bell, Ph.D. Bioinformatics and Research Computing 2012 Spring Lecture Series
ImmGen microarray gene expression data: Data Generation and Quality Control pipeline.
 ImmGen microarray gene expression data: Data Generation and Quality Control pipeline. Jeff Ericson*, Scott Davis*, Jon Lesh #, Melissa Howard #, Diane Mathis and Christophe Benoist Division of Immunology,
ImmGen microarray gene expression data: Data Generation and Quality Control pipeline. Jeff Ericson*, Scott Davis*, Jon Lesh #, Melissa Howard #, Diane Mathis and Christophe Benoist Division of Immunology,
CS273B: Deep Learning in Genomics and Biomedicine. Recitation 1 30/9/2016
 CS273B: Deep Learning in Genomics and Biomedicine. Recitation 1 30/9/2016 Topics Genetic variation Population structure Linkage disequilibrium Natural disease variants Genome Wide Association Studies Gene
CS273B: Deep Learning in Genomics and Biomedicine. Recitation 1 30/9/2016 Topics Genetic variation Population structure Linkage disequilibrium Natural disease variants Genome Wide Association Studies Gene
Rapidly regulated genes are intron poor
 Supplementary material Rapidly regulated genes are intron poor Daniel C. Jeffares 1 *, Christopher J. Penkett 1,2 * and Jürg Bähler 1,3 1 Wellcome Trust Sanger Institute, Hinxton, Cambridge CB10 1SA, UK
Supplementary material Rapidly regulated genes are intron poor Daniel C. Jeffares 1 *, Christopher J. Penkett 1,2 * and Jürg Bähler 1,3 1 Wellcome Trust Sanger Institute, Hinxton, Cambridge CB10 1SA, UK
Gene Signal Estimates from Exon Arrays
 Gene Signal Estimates from Exon Arrays I. Introduction: With exon arrays like the GeneChip Human Exon 1.0 ST Array, researchers can examine the transcriptional profile of an entire gene (Figure 1). Being
Gene Signal Estimates from Exon Arrays I. Introduction: With exon arrays like the GeneChip Human Exon 1.0 ST Array, researchers can examine the transcriptional profile of an entire gene (Figure 1). Being
Finding Recurrent Copy Number Alteration Regions: A Review of Methods
 Current Bioinformatics, 2010, 5, 1-17 1 Finding Recurrent Copy Number Alteration Regions: A Review of Methods Oscar M. Rueda*,1,2 and Ramon Diaz-Uriarte*,1 1 Structural and Computational Biology Programme,
Current Bioinformatics, 2010, 5, 1-17 1 Finding Recurrent Copy Number Alteration Regions: A Review of Methods Oscar M. Rueda*,1,2 and Ramon Diaz-Uriarte*,1 1 Structural and Computational Biology Programme,
Gene Regulation Solutions. Microarrays and Next-Generation Sequencing
 Gene Regulation Solutions Microarrays and Next-Generation Sequencing Gene Regulation Solutions The Microarrays Advantage Microarrays Lead the Industry in: Comprehensive Content SurePrint G3 Human Gene
Gene Regulation Solutions Microarrays and Next-Generation Sequencing Gene Regulation Solutions The Microarrays Advantage Microarrays Lead the Industry in: Comprehensive Content SurePrint G3 Human Gene
A Comparative Study of Microarray Data Analysis for Cancer Classification
 A Comparative Study of Microarray Data Analysis for Cancer Classification Kshipra Chitode Research Student Government College of Engineering Aurangabad, India Meghana Nagori Asst. Professor, CSE Dept Government
A Comparative Study of Microarray Data Analysis for Cancer Classification Kshipra Chitode Research Student Government College of Engineering Aurangabad, India Meghana Nagori Asst. Professor, CSE Dept Government
Identifying Candidate Informative Genes for Biomarker Prediction of Liver Cancer
 Identifying Candidate Informative Genes for Biomarker Prediction of Liver Cancer Nagwan M. Abdel Samee 1, Nahed H. Solouma 2, Mahmoud Elhefnawy 3, Abdalla S. Ahmed 4, Yasser M. Kadah 5 1 Computer Engineering
Identifying Candidate Informative Genes for Biomarker Prediction of Liver Cancer Nagwan M. Abdel Samee 1, Nahed H. Solouma 2, Mahmoud Elhefnawy 3, Abdalla S. Ahmed 4, Yasser M. Kadah 5 1 Computer Engineering
Emma Huxley. Principal Clinical Scientist West Midlands Regional Genetics Laboratory
 Genetic Analysis Using a High Density SNP Array in Myelodysplastic Syndrome: Clinical Utility and Comparative Analysis Study Compared to Metaphase Chromosome Analysis. Emma Huxley Principal Clinical Scientist
Genetic Analysis Using a High Density SNP Array in Myelodysplastic Syndrome: Clinical Utility and Comparative Analysis Study Compared to Metaphase Chromosome Analysis. Emma Huxley Principal Clinical Scientist
Nature Methods: doi: /nmeth.4396
 Supplementary Figure 1 Comparison of technical replicate consistency between and across the standard ATAC-seq method, DNase-seq, and Omni-ATAC. (a) Heatmap-based representation of ATAC-seq quality control
Supplementary Figure 1 Comparison of technical replicate consistency between and across the standard ATAC-seq method, DNase-seq, and Omni-ATAC. (a) Heatmap-based representation of ATAC-seq quality control
Chromosome Analysis Suite 3.0 (ChAS 3.0)
 Chromosome Analysis Suite 3.0 (ChAS 3.0) FAQs related to CytoScan Cytogenetics Suite 1. What is required for processing and viewing CytoScan array CEL files? CytoScan array CEL files are processed and
Chromosome Analysis Suite 3.0 (ChAS 3.0) FAQs related to CytoScan Cytogenetics Suite 1. What is required for processing and viewing CytoScan array CEL files? CytoScan array CEL files are processed and
6. GENE EXPRESSION ANALYSIS MICROARRAYS
 6. GENE EXPRESSION ANALYSIS MICROARRAYS BIOINFORMATICS COURSE MTAT.03.239 16.10.2013 GENE EXPRESSION ANALYSIS MICROARRAYS Slides adapted from Konstantin Tretyakov s 2011/2012 and Priit Adlers 2010/2011
6. GENE EXPRESSION ANALYSIS MICROARRAYS BIOINFORMATICS COURSE MTAT.03.239 16.10.2013 GENE EXPRESSION ANALYSIS MICROARRAYS Slides adapted from Konstantin Tretyakov s 2011/2012 and Priit Adlers 2010/2011
RNA Degradation and NUSE Plots. Austin Bowles STAT 5570/6570 April 22, 2011
 RNA Degradation and NUSE Plots Austin Bowles STAT 5570/6570 April 22, 2011 References Sections 3.4 and 3.5.1 of Bioinformatics and Computational Biology Solutions Using R and Bioconductor (Gentleman et
RNA Degradation and NUSE Plots Austin Bowles STAT 5570/6570 April 22, 2011 References Sections 3.4 and 3.5.1 of Bioinformatics and Computational Biology Solutions Using R and Bioconductor (Gentleman et
John Gurdon was testing the hypothesis of genomic equivalence or that when cells divide they retain a full genomic compliment.
 1. (15 pts) John Gurdon won the 2012 Nobel Prize in Physiology or Medicine for work he did in the 1960 s. What was the major developmental hypothesis he set out to test? What techniques did he development
1. (15 pts) John Gurdon won the 2012 Nobel Prize in Physiology or Medicine for work he did in the 1960 s. What was the major developmental hypothesis he set out to test? What techniques did he development
% Viability. isw2 ino isw2 ino isw2 ino isw2 ino mM HU 4-NQO CPT
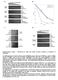 a Drug concentration b 1.3% MMS nhp1 nhp1 8 nhp1 mag1.5% MMS.3% MMS nhp1 nhp1 ino8 9 ino8 9 % Viability 4.5% MMS ino8 9 ino8 9 2.5.1.15 % MMS c d nhp1 nhp1 nhp1 nhp1 nhp1 nhp1 Control (YPD) γ IR (1 gy)
a Drug concentration b 1.3% MMS nhp1 nhp1 8 nhp1 mag1.5% MMS.3% MMS nhp1 nhp1 ino8 9 ino8 9 % Viability 4.5% MMS ino8 9 ino8 9 2.5.1.15 % MMS c d nhp1 nhp1 nhp1 nhp1 nhp1 nhp1 Control (YPD) γ IR (1 gy)
Nucleofector Technology in. Somatic Stem Cell Research. gene transfer begins here
 Nucleofector Technology in Somatic Stem Cell Research The Nucleofector technology The first highly efficient method for non-viral gene transfer into primary cells and difficult-to-transfect cell lines
Nucleofector Technology in Somatic Stem Cell Research The Nucleofector technology The first highly efficient method for non-viral gene transfer into primary cells and difficult-to-transfect cell lines
Life Sciences Reporting Summary
 Life Sciences Reporting Summary Corresponding author(s): George Q. Daley Initial submission Revised version Final submission Nature Research wishes to improve the reproducibility of the work that we publish.
Life Sciences Reporting Summary Corresponding author(s): George Q. Daley Initial submission Revised version Final submission Nature Research wishes to improve the reproducibility of the work that we publish.
PRIME-XV Cell Therapy Products by
 PRIME-XV Cell Therapy Products by Product List 2017 TRINOVA BIOCHEM now distributes the PRIME-XV Cell Therapy line by Irvine Scientific in Germany, Austria and Switzerland. Irvine Scientific is a worldwide
PRIME-XV Cell Therapy Products by Product List 2017 TRINOVA BIOCHEM now distributes the PRIME-XV Cell Therapy line by Irvine Scientific in Germany, Austria and Switzerland. Irvine Scientific is a worldwide
Chapter 20: Biotechnology
 Name Period The AP Biology exam has reached into this chapter for essay questions on a regular basis over the past 15 years. Student responses show that biotechnology is a difficult topic. This chapter
Name Period The AP Biology exam has reached into this chapter for essay questions on a regular basis over the past 15 years. Student responses show that biotechnology is a difficult topic. This chapter
affy: Built-in Processing Methods
 affy: Built-in Processing Methods Ben Bolstad October 30, 2017 Contents 1 Introduction 2 2 Background methods 2 2.1 none...................................... 2 2.2 rma/rma2...................................
affy: Built-in Processing Methods Ben Bolstad October 30, 2017 Contents 1 Introduction 2 2 Background methods 2 2.1 none...................................... 2 2.2 rma/rma2...................................
Fundamental properties of Stem Cells
 Stem cells Learning Goals: Define what a stem cell is and describe its general properties, using hematopoietic stem cells as an example. Describe to a non-scientist the current progress of human stem cell
Stem cells Learning Goals: Define what a stem cell is and describe its general properties, using hematopoietic stem cells as an example. Describe to a non-scientist the current progress of human stem cell
Estoril Education Day
 Estoril Education Day -Experimental design in Proteomics October 23rd, 2010 Peter James Note Taking All the Powerpoint slides from the Talks are available for download from: http://www.immun.lth.se/education/
Estoril Education Day -Experimental design in Proteomics October 23rd, 2010 Peter James Note Taking All the Powerpoint slides from the Talks are available for download from: http://www.immun.lth.se/education/
A Technical Guide to Aneuploidy Calling with VeriSeq PGS
 A Technical Guide to Aneuploidy Calling with VeriSeq PGS Introduction 3 Typical Cases 4 Examples of Calling Challenging Profiles 7 Mosaic Samples 10 Segmental Copy Number Changes 11 Differences Between
A Technical Guide to Aneuploidy Calling with VeriSeq PGS Introduction 3 Typical Cases 4 Examples of Calling Challenging Profiles 7 Mosaic Samples 10 Segmental Copy Number Changes 11 Differences Between
DB3230 Midterm 1 11/15/2013 Name:
 1. (15 pts) Nuclear cloning by John Gurdon was rarely successful in producing fertile adults. Why not? Explain why serial transplantation improves the success rate. What else could you do to improve the
1. (15 pts) Nuclear cloning by John Gurdon was rarely successful in producing fertile adults. Why not? Explain why serial transplantation improves the success rate. What else could you do to improve the
Next-Generation Sequencing Gene Expression Analysis Using Agilent GeneSpring GX
 Next-Generation Sequencing Gene Expression Analysis Using Agilent GeneSpring GX Technical Overview Introduction RNA Sequencing (RNA-Seq) is one of the most commonly used next-generation sequencing (NGS)
Next-Generation Sequencing Gene Expression Analysis Using Agilent GeneSpring GX Technical Overview Introduction RNA Sequencing (RNA-Seq) is one of the most commonly used next-generation sequencing (NGS)
Nature Methods: doi: /nmeth.3732
 Supplementary Figure 1 GenomeSpace allows the biologist user to conduct complex analysis. 1a. Flow chart of our analytic approach to dissecting a stem cell-like gene regulatory network in human cancers.
Supplementary Figure 1 GenomeSpace allows the biologist user to conduct complex analysis. 1a. Flow chart of our analytic approach to dissecting a stem cell-like gene regulatory network in human cancers.
Characteristics. Capable of dividing and renewing themselves for long periods of time (proliferation and renewal)
 STEM CELLS (p 2-8) Overview The body is made up of about 200 different kinds of specialised cells, such as muscle cells, nerve cells, fat cells and skin cells. Cells with the capacity to give rise to the
STEM CELLS (p 2-8) Overview The body is made up of about 200 different kinds of specialised cells, such as muscle cells, nerve cells, fat cells and skin cells. Cells with the capacity to give rise to the
At the conclusion of this lesson you should be able to:
 Learning Objectives At the conclusion of this lesson you should be able to: Understand the key terms and definitions regarding stem cells Differentiate between the adult and embryonic stem cells Differentiate
Learning Objectives At the conclusion of this lesson you should be able to: Understand the key terms and definitions regarding stem cells Differentiate between the adult and embryonic stem cells Differentiate
Runs of Homozygosity Analysis Tutorial
 Runs of Homozygosity Analysis Tutorial Release 8.7.0 Golden Helix, Inc. March 22, 2017 Contents 1. Overview of the Project 2 2. Identify Runs of Homozygosity 6 Illustrative Example...............................................
Runs of Homozygosity Analysis Tutorial Release 8.7.0 Golden Helix, Inc. March 22, 2017 Contents 1. Overview of the Project 2 2. Identify Runs of Homozygosity 6 Illustrative Example...............................................
Microarray Analysis of Gene Expression in Huntington's Disease Peripheral Blood - a Platform Comparison. CodeLink compatible
 Microarray Analysis of Gene Expression in Huntington's Disease Peripheral Blood - a Platform Comparison CodeLink compatible Microarray Analysis of Gene Expression in Huntington's Disease Peripheral Blood
Microarray Analysis of Gene Expression in Huntington's Disease Peripheral Blood - a Platform Comparison CodeLink compatible Microarray Analysis of Gene Expression in Huntington's Disease Peripheral Blood
Getting high-quality cytogenetic data is a SNP.
 Getting high-quality cytogenetic data is a SNP. SNP data. Increased insight. Cytogenetics is at the forefront of the study of cancer and congenital disorders. And we put you at the forefront of cytogenetics.
Getting high-quality cytogenetic data is a SNP. SNP data. Increased insight. Cytogenetics is at the forefront of the study of cancer and congenital disorders. And we put you at the forefront of cytogenetics.
Introduction to ChIP Seq data analyses. Acknowledgement: slides taken from Dr. H
 Introduction to ChIP Seq data analyses Acknowledgement: slides taken from Dr. H Wu @Emory ChIP seq: Chromatin ImmunoPrecipitation it ti + sequencing Same biological motivation as ChIP chip: measure specific
Introduction to ChIP Seq data analyses Acknowledgement: slides taken from Dr. H Wu @Emory ChIP seq: Chromatin ImmunoPrecipitation it ti + sequencing Same biological motivation as ChIP chip: measure specific
Cancer Genetics Solutions
 Cancer Genetics Solutions Cancer Genetics Solutions Pushing the Boundaries in Cancer Genetics Cancer is a formidable foe that presents significant challenges. The complexity of this disease can be daunting
Cancer Genetics Solutions Cancer Genetics Solutions Pushing the Boundaries in Cancer Genetics Cancer is a formidable foe that presents significant challenges. The complexity of this disease can be daunting
Nature Methods: doi: /nmeth Supplementary Figure 1
 Supplementary Figure 1 Workflow for multimodal analysis using sc-gem on a programmable microfluidic device (Fluidigm). 1) Cells are captured and lysed, 2) RNA from lysed single cell is reverse-transcribed
Supplementary Figure 1 Workflow for multimodal analysis using sc-gem on a programmable microfluidic device (Fluidigm). 1) Cells are captured and lysed, 2) RNA from lysed single cell is reverse-transcribed
Exploration, Normalization, Summaries, and Software for Affymetrix Probe Level Data
 Exploration, Normalization, Summaries, and Software for Affymetrix Probe Level Data Rafael A. Irizarry Department of Biostatistics, JHU March 12, 2003 Outline Review of technology Why study probe level
Exploration, Normalization, Summaries, and Software for Affymetrix Probe Level Data Rafael A. Irizarry Department of Biostatistics, JHU March 12, 2003 Outline Review of technology Why study probe level
PRIME-XV Cell Therapy Products by
 PRIME-XV Cell Therapy Products by Product List 2017 TRINOVA BIOCHEM now distributes the PRIME-XV Cell Therapy line by Irvine Scientific in Germany, Austria and Switzerland. Irvine Scientific is a worldwide
PRIME-XV Cell Therapy Products by Product List 2017 TRINOVA BIOCHEM now distributes the PRIME-XV Cell Therapy line by Irvine Scientific in Germany, Austria and Switzerland. Irvine Scientific is a worldwide
Lecture #1. Introduction to microarray technology
 Lecture #1 Introduction to microarray technology Outline General purpose Microarray assay concept Basic microarray experimental process cdna/two channel arrays Oligonucleotide arrays Exon arrays Comparing
Lecture #1 Introduction to microarray technology Outline General purpose Microarray assay concept Basic microarray experimental process cdna/two channel arrays Oligonucleotide arrays Exon arrays Comparing
