Localization-Based Super-Resolution Light Microscopy
|
|
|
- Emory Hood
- 5 years ago
- Views:
Transcription
1 Kristin A. Gabor, 1,2,3 Mudalige S. Gunewardene, 1 David Santucci, 4 and Samuel T. Hess 1,3, * 1 Department of Physics and Astronomy, 120 Bennett Hall, University of Maine, Orono, ME Department of Molecular and Biomedical Sciences, 5735 Hitchner Hall, University of Maine, Orono, ME Graduate School of Biomedical Sciences, 263 ESRB/Barrows Hall, University of Maine, Orono, ME Volen Center for Complex Systems, 415 South St., Brandeis University, Waltham, MA * sam.hess@umit.maine.edu Introduction Fluorescence microscopy is an essential and flexible tool for the study of biology, chemistry, and physics. It can provide information on a wide range of spatial and temporal scales. However, since the inception of light microscopy, diffraction has limited the size of the smallest details that could be imaged in any sample using light. Because much of biology occurs on molecular length scales, interest in circumventing the diffraction limit has been high for many years. Recently, several techniques have been introduced that can bend or break the diffraction limit. Localization-based methods introduced in 2006 have reached this goal and are now rapidly growing in popularity. Principles of Localization-Based Super-Resolution Microscopy Localization versus resolution. Diffraction blurs the details of objects imaged by a lens-based far-field microscope system. Objects closer together than R 0 = 0.61λ/NA are unresolved according to the Rayleigh criterion, where NA is the numerical aperture, λ is the wavelength of detected light, and R 0 is ~ nm for green light imaged by a high-na objective. In contrast, localization, which is the determination of the position of an object using its image, has been achieved with precision of ~1 nm [1], limited in principle only by the number of detected photons. Localization-based super-resolution microscopy [2 4] relies on both the imaging of single molecules and the stochastic activation of sparse subsets of such molecules. This is often achieved by optical control of molecular transitions between bright and dark states. When the number of emitting molecules is limited, and the distance between molecules is greater than R 0, the molecular images are distinguishable (Figure 1). By activating a subset, imaging it, photobleaching or deactivating it, and repeating this process for many subsets of molecules, coordinates of thousands of molecules can be obtained. Molecular positions are measured by fitting the molecular image with a two-dimensional Gaussian function or with the experimental point spread function (PSF). For reliable localization, no more than a few molecules per frame can be localized within any diffraction-limited area. Thus, many (for example, ) image frames are required to achieve a high density of molecules in the final rendered image. Once a sufficient number of these molecules are acquired and localized, the molecular positions are plotted to generate an image. The effective resolution of the image obtained depends Figure 1: Principle of localization-based super-resolution microscopy. By limiting the number of fluorescent molecules visible at once, the images of the individual molecules become distinguishable. (A) Molecules are initially in an inactive (non-fluorescent) state. (B) Sparse subsets of molecules are converted into a fluorescent state by the activation beam (purple). When excited by the readout laser (green), they are imaged (C) until deactivated or photobleached (D). Molecules are localized by fitting the image with a two-dimensional Gaussian. Cycles of activation (B, E), readout and localization (C, F), and photobleaching (D, G) are repeated for many subsets of molecules. Rendered images with few (H) and large number (I) of localized molecules show buildup of structural detail as density increases. (J) Conventional image with diffraction-limited resolution. 12 doi: /s July
2 on both the precision with which molecules can be localized as well as the nearest-neighbor distance between molecules [5, 6]. Fluorescence photoactivation localization microscopy (FPALM) and similar techniques have achieved a resolution approximately ten-fold better than the diffraction limit. In general, any technique that is capable of creating a sparse subset of visible molecules, imaging that subset, and sampling many subsets can be used to generate a super-resolution image. FPALM, PALM, STORM, and dstorm. Conceptually, FPALM [2], photoactivated localization microscopy (PALM) [3], stochastic optical reconstruction microscopy (STORM) [4], and direct stochastic optical reconstruction microscopy (dstorm) [7, 8] are very similar. Each technique relies on photophysical properties of probes to control the number of visible fluorescent molecules in the field of view. FPALM and PALM typically utilize photoactivatable fluorescent proteins (PA-FPs) to allow for direct optical control over the number of fluorescent molecules visible at a given time. This is achieved by adjusting the rates of photoactivation and photobleaching. Similarly, STORM uses pairs of combinations of organic fluorophores that photoswitch in the presence of thiol-containing reducing agents to control the number of molecules that are fluorescent at a given time. dstorm uses single fluorophores that photoswitch in the presence of thiols and can be used in living cells, which contain reducing agents such as glutathione at micromolar to millimolar concentrations [8]. In all of these techniques, images are subsequently reconstructed from the coordinates and intensities of the localized molecules. Probes. Localization-based techniques can be performed with either certain conventional fluorophores [8 10], PA-FPs, caged or photoswitchable organic dyes, or pairs of probes that act together as a molecular switch [11 15]. Typically, one wavelength (called the activation wavelength) is used to control the density of visible molecules while another wavelength (called the readout wavelength) is used to collect the fluorescence from the activated molecules. In some instances, one wavelength is sufficient to do both tasks; regardless, the choice of wavelength, intensity, and temporal illumination pattern enables active control over the number of visible molecules. In doing so, these methods rely on photophysical properties of the probes to control the number of molecules in the field of view at a given time [2, 16]. The photophysical properties of probes play a major role in the spatial resolution and rate of image acquisition. Resolution in localization microscopy is a function of the localization precision and density of localized molecules. As localization precision and single-molecule detection (against background) are greatly improved with more detected photons per molecule, probes that emit the maximum number of photons possible before bleaching are ideal. Desirable probes have a high rate of photon emission and a low rate of spontaneous or readout-induced activation, are small in size to minimize perturbation of cell structure or function, and have a large extinction coefficient to minimize the illumination intensity necessary for readout. Probes should also have large contrast (the ratio of brightness in active to inactive forms) so that background from inactive molecules is negligible. Probes with longer wavelength emission are beneficial to minimize autofluorescence background levels. For multicolor labeling, 2011 July optimal probe combinations have these same characteristics and are typically spectrally well separated. Fundamental limitations. One important limitation is the trade-off between spatial and temporal resolution. Sub-millisecond observation of molecular motions is limited by the intensity of the readout laser, the probe emission rate, and camera readout speed. Therefore, acquisition of super-resolution FPALM images on timescales faster than 0.1 s is currently difficult to achieve. The density of localized molecules is another limiting factor in image resolution. Because only one molecule (or at best a few molecules) can be localized per diffraction-limited area in any given frame, improved spatial resolution comes at the price of temporal resolution. Encouraging recent work using methods from astrophysics has enabled analysis of molecular images with at least ten-fold higher density, which in principle should allow imaging rates that are faster by the same factor [17]. Minimization of background is crucial both for identification of fluorescent probe molecules and for maximization of the localization precision. Thus, samples should not emit significant autofluorescence in the same spectral region as the probe being used. Use of UV-bleached water for imaging buffers, use of phenol-red-free media for cell growth, and use of a water-immersion objective beaded with bleached water can help. Additionally, if background levels are high initially, activation of probes can be delayed while the background is allowed to bleach. Many available PA-FPs have overlapping emission spectra, limiting the number of species usable in a typical multicolor imaging application. Ultimately, the continued development of photoactivatable probes will extend the capabilities of localization microscopy. The improvement of probes relies on better understanding and control of their photophysical properties. The labeling efficiency of each probe is another factor to consider, as well as the expression level of the protein to which it is fused. As probes are developed with higher peak photon emission rates, localization microscopy will be possible with higher spatial and temporal resolution. Methods and Applications Recent progress in the field of localization-based super-resolution microscopy has led to extensions of these techniques to multicolor [8, 18 20], three-dimensional [21 24], and polarization imaging [25]. For instance, using modifications to the FPALM detection path has enabled multi-channel detection on the same camera chip as well as nanoscale imaging of single molecule anisotropies. A typical experimental geometry for two-color three-dimensional imaging is shown in Figure 2. These and other advances show great promise for improved understanding of biological systems at the molecular level. Single-species localization. A typical localization microscopy setup is a fluorescence microscope with activation and readout lasers aligned so that they are collinear, a high-na objective lens, and a sensitive charge-coupled device (camera) capable of imaging single fluorescent molecules. Illumination light is focused in the center (or at the edge) of the objective back aperture to illuminate the sample by widefield (or by TIRF) over an area of ~ μm 2. 13
3 Figure 2: Experimental geometry for two-color bi-plane FPALM. (Left) A set of lasers is aligned into a collinear path to allow a choice of one activation and one readout illumination wavelength. These two beams are directed through a series of mirrors into a focusing lens positioned at one focal length from the back aperture of a microscope objective within a fluorescence microscope (here shown as inverted). (Right) The sample is illuminated on the microscope stage, and emitted fluorescence from single molecules is separated from illumination by a dichroic (DM1) and sent through the tube lens and out the microscope side port. Next, the fluorescence image is magnified by a telescope and then split by a second dichroic (DM2) into two detection channels. Each detection channel can then be passed through a 50:50 beamsplitter to form separate images on each of two EMCCD cameras. Note that the two longer arms of the red and green paths should have the same distance between sample and camera, as should the two shorter arms of the red and green paths. For illustrative purposes, the figure does not show the optics arranged in this way. Also shown are an optional spectrometer, for confirmation of molecular species, and a pulsed laser for multi-photon activation and/or readout. Activation laser intensity is regulated, either by pulsing with progressively higher frequency or gradually increasing intensity as the pool of inactive molecules is depleted, to maintain a small number of activated molecules within the field of view. Readout laser intensity is controlled such that the active fluorescent molecules are clearly visible above the background. Typically, hundreds to thousands of frames are acquired and then analyzed to localize visible molecules. Images are rendered by plotting each molecule as a Gaussian spot with a weight and size proportional to the number of photons detected and the localization precision, respectively. An example of single-color imaging of dendritic spines is shown in Figure 3. The first published example of localization microscopy in living cells used single-color FPALM of the viral surface protein hemagglutinin (HA) to distinguish between several competing models of membrane organization [26]. A wide variety of biological applications have been presented [27 30]; localization microscopy is applicable to virtually any subcellular structure, as long as the structure can be labeled and the molecular millieu is compatible with imaging and localizing single molecules. In addition to molecular coordinates, a direct count of the numbers of localized molecules (and thus, with careful analysis, the densities of molecules) is obtained, and sub-populations of molecules can be identified based on brightness, or other spectroscopic properties, which would otherwise be averaged out in ensemble imaging methods. Correlation of localization microscopy results with other methods, such as confocal or widefield fluorescence, atomic-force microcsopy, or electron microscopy, is extremely valuable as a way to provide context for the molecular positions. Multicolor imaging. Multicolor super-resolution imaging is invaluable for the direct visualization of molecular interactions at the nanoscale level [8, 18 20]. Multicolor imaging and analysis has been performed as an extension of standard FPALM using a detection scheme similar to that reported previously for multicolor single molecule imaging [19]. With a detection path divided by a dichroic into two wavelength ranges [19], camera frames containing two spatially distinct (but simultaneous) images representing the two emission wavelength ranges are obtained. The two images are correlated and superimposed for localization, while the intensity ratio of one channel over the sum of both channels is used to distinguish the multiple molecular species. Both PALM and STORM have also reported multicolor super-resolution images obtained using combinations of cyanine dyes or PA-FPs [18, 20]. Advantages of PA-FPs for localization microscopy include the ability to perform live-cell imaging (that is, without thiol or oxygen-scavenging reagents), Figure 3: FPALM image of dendrites of fixed rat hippocampal neurons in a culture labeled with Dendra2-PSD95. Green: convention fluorescence. Orange: FPALM image. Credit: D. Santucci, T. Gould, T. Newpher, A. Mabb, M. Ehlers, J. Lisman, and S. Hess July
4 the use of relatively non-invasive labeling procedures (that is, transfection), and high specificity due to genetic encoding of PA-FP-protein chimeras. Time-lapse imaging. Temporally-resolved super-resolution imaging is useful for live or dynamic specimens. Time resolution depends on desired resolution: 50-nm resolution requires a nearest-neighbor distance of nm, or approximately molecules per diffraction-limited area, which requires (for one rendered image) at least frames of acquisition, or seconds per acquisition at 400 Hz. Such frame rates are achievable using a number of commercially available EMCCD cameras, such as the Andor ixon+ DU-897. Time resolution of 25 nm requires nearest-neighbor distance of nm, or frames, requiring seconds per rendered image. Intensity should be adjusted to bleach molecules within approximately one frame. Note that the structure of interest should not rearrange significantly during this time period, although the individual molecules can move within the structure as long as their motion samples or is meaningfully representative of the structure overall. For example, the dynamics of actin structures at the cell leading edge (motion toward the cell periphery at 0.1 to 2 μm/min) [31] is compatible with live-cell FPALM. The dynamics of membrane protein clusters is also well-suited to time-lapse imaging [26]. Of course, control experiments must always be performed to confirm that addition of labels to the sample does not perturb structure or function. Figures 4 and 5 show examples of two-color live-cell and fixed-cell FPALM, respectively. Molecular trajectories. Imaging conditions can be adjusted to allow probe molecules to remain fluorescent for several frames before photobleaching. Thus, several consecutive images of the same molecule can be analyzed to construct a short trajectory, which can be equivalent to a few milliseconds or as much as a few seconds. A trade-off results between trajectory length and localization precision per step from the limited photon budget per molecule: for a thousand photons detected from a single molecule before bleaching, 10 steps with 100 photons per frame results in a localization precision of ~20 30 nm at best, whereas 40 steps would yield nm localization precision. On the other hand, trajectories may be obtained for hundreds of thousands of molecules, allowing higher density sampling of structures [32]. Clearly, such methods are not well-suited to situations where long-term tracking of the same molecule(s) is important. However, for many applications, such as tracking of membrane proteins and lipids within small clusters, or measurement of the local velocity of cytoskeletal structures, trajectory imaging can be quite useful. Probes that are particularly resistant to photobleaching, allowing detection of >10 3 photons, are highly desirable. Three-dimensional imaging. Because most biological structures are three-dimensional (3D), Figure 4: Multicolor live-cell FPALM. (A) Pseudocolor image of individual PA-FP molecules (30 ms exposure with Andor ixon+ EMCCD) illustrating FPALM with simultaneous two-channel detection. (B) Two-color FPALM image of Dendra2-HA (green) and PAmCherry actin (red) in a live HAb2 fibroblast cell at 37 C (8000 frames total). (C) Dendra2-HA only. (D) PAmCherry actin only. Credit: M. Gudheti, M. Siyath Gunewardene, T. Gould, and S. Hess. methods that can image beyond the diffraction limit in the axial direction (z) are essential for advancement of our understanding of biological systems. However, the diffraction-limited resolution in z is typically worse than the lateral resolution. Advances in 3D localization microscopy have been demonstrated in several incarnations, including biplane detection [21], optical astigmatism [22], interferometry [23], and double-helix point-spread function [24]. Figure 5: Multicolor FPALM image showing colocalization of PAmCherry-caveolin-1 (red) with Dendra2-interferon receptor molecules (green) in fixed ZFL cells. Credit: K. Gabor, T. Gould, C. Kim, and S. Hess July 15
5 Biplane FPALM (BPFPALM) is an extension of FPALM to 3D, which works by splitting the detected fluorescence into two separate paths with different overall lengths. The two images simultaneously formed on separate regions of the camera correspond to two different focal planes and are later fitted with the measured PSF to determine the x, y, and z coordinates of each molecule. BPFPALM of samples was demonstrated with an axial resolution of ~50 75 nm [21, 33], approximately 10-fold improved over conventional fluorescence microscopy. Optical astigmatism is used to determine both axial and lateral positions of single molecules in 3D STORM [22]. By inserting a cylindrical lens into the detection path of the microscope, two slightly different focal planes result for the x and y dimensions of each molecular image. As a result, fluorophores appear elliptical in x or y depending on their axial position, allowing that axial position to be determined from the molecular image. Interferometric PALM (ipalm) resolves 3D molecular coordinates of individual PA-FP tagged proteins with ~10 nm axial resolution and ~20 nm lateral resolution using a 4Pi (two-objective) geometry [23]. The detected fluorescence is divided by a three-way beam splitter and imaged with three cameras. The relative intensities of the three images of each molecule are then analyzed to determine the axial position of the molecule. ipalm was recently used to deduce the structure of the focal adhesion complex [34]. In the double-helix point spread function (DH-PSF) method, a phase modulation pattern in the detection path yields a detected PSF with two lobes whose orientation rotates as a function of the axial position of the detected molecule. DH-PSF has a demonstrated resolution of ~10 nm laterally and 20 nm axially over a sample depth of ~2 µm [24]. Polarization FPALM: imaging molecular orientations at the nanoscale level. The localization-based imaging techniques described above are crucial for studies of biological structures, but they do not provide information about the orientation or rotational freedom of single molecules. Polarization FPALM (PolPALM) [25] has extended FPALM to image single molecule anisotropies by simultaneously detecting two spatially separated images of the fluorescence, polarized parallel and perpendicular to a specific axis at the sample. To measure the anisotropy for each localized molecule, PolPALM analyzes the relative intensities of molecules in the two images. The demonstrated resolution of ~17 nm using PolPALM allows quantification of structure on length scales inaccessible to diffraction-limited techniques. The anisotropy obtained reflects the range of accessible probe orientations and can also be used to quantify binding between molecules. Conclusion Localization microscopy provides nanoscale spatial resolution in living fluorescent specimens using far-field optics. Such capabilities suggest that a multitude of fundamental but previously inaccessible biological questions can now be addressed. Current limitations such as number of simultaneously distinguishable species and the time resolution of imaging are likely to improve with refined experimental methods and improved probes. Furthermore, using existing localization microscopy methods, the number of compatible biological systems is continuously increasing. New biological insights hopefully are imminent. Acknowledgments The authors thank M. V. Gudheti and T. J. Gould for FPALM images included within the article; S.-R. Yin and Vladislav Verkhusha for constructs; and A. Gosse, J. Lake, and J. Gosse for support. This work was funded by NIH Career Award K25 AI65459, NIH R15 GM094713, NIH R01 GM094713, NSF MRI CHE , Maine Technology Institute MTAF 1106 and 2061, UMaine V.P. for Research, and the Maine Economic Improvement Fund (MEIF). References [1] A Yildiz et al., Science 300 (2003) [2] ST Hess et al., Biophys J 91 (2006) [3] E Betzig et al., Science 313 (2006) [4] MJ Rust et al., Nature Methods 3 (2006) [5] H Shroff et al., Nature Methods 5 (2008) [6] TJ Gould et al., Nat Protoc 4 (2009) [7] T Klein et al., Nature Methods 8 (2011) 7 9. [8] M Heilemann et al., Angew Chem Int Ed Engl 47 (2008) [9] JS Biteen et al., Nature Methods 5 (2008) [10] SF Lee et al., Biophys J 100 (2011) L [11] GH Patterson and J Lippincott-Schwartz, Science 297 (2002) [12] FV Subach et al., Nature Methods 6 (2009) [13] VV Verkhusha and A Sorkin, Chem Biol 12 (2005) [14] J Wiedenmann et al., Proc Natl Acad Sci USA 101 (2004) [15] DM Chudakov et al., Biotechniques 42 (2007) 553, 555, 557 passim. [16] TJ Gould et al., Nature Protoc 4 (2009) [17] SJ Holden et al., Nature Methods 8 (2011) [18] M Bates et al., Science 317 (2007) [19] M Bossi et al., Nano Lett 8 (2008) [20] H Shroff et al., Proc Natl Acad Sci USA 104 (2007) [21] MF Juette et al., Nature Methods 5 (2008) [22] B Huang et al., Science 319 (2008) [23] G Shtengel et al., Proc Natl Acad Sci USA 106 (2009) [24] SRP Pavani et al., Proc Natl Acad Sci USA 106 (2009) [25] TJ Gould et al., Nature Methods 5 (2008) [26] ST Hess et al., Proc Natl Acad Sci USA 104 (2007) [27] ST Hess et al., Methods Mol Biol 544 (2009) [28] TJ Gould and ST Hess, Biophysical Tools for Biologists, Vol 2: In Vivo Techniques 89 (2008) [29] G Patterson et al., Annu Rev Phys Chem 61 (2010) [30] XW Zhuang, Nature Photonics 3 (2009) [31] CM Waterman-Storer and G Danuser, Curr Biol 12 (2002) R [32] S Manley et al., Nature Methods 5 (2008) [33] MJ Mlodzianoski et al., Opt Express 17 (2009) [34] P Kanchanawong et al., Nature 468 (2010) July
STORM/PALM. Super Resolution Microscopy 10/31/2011. Looking into microscopic world of life
 Super Resolution Microscopy STORM/PALM Bo Huang Department of Pharmaceutical Chemistry, UCSF CSHL Quantitative Microscopy, 1/31/211 Looking into microscopic world of life 1 µm 1 µm 1 nm 1 nm 1 nm 1 Å Naked
Super Resolution Microscopy STORM/PALM Bo Huang Department of Pharmaceutical Chemistry, UCSF CSHL Quantitative Microscopy, 1/31/211 Looking into microscopic world of life 1 µm 1 µm 1 nm 1 nm 1 nm 1 Å Naked
Fluorescence Nanoscopy
 Fluorescence Nanoscopy Keith A. Lidke University of New Mexico panda3.phys.unm.edu/~klidke/index.html Optical Microscopy http://en.wikipedia.org/wiki/k%c3%b6hler_illumination 30 µm Fluorescent Probes Michalet
Fluorescence Nanoscopy Keith A. Lidke University of New Mexico panda3.phys.unm.edu/~klidke/index.html Optical Microscopy http://en.wikipedia.org/wiki/k%c3%b6hler_illumination 30 µm Fluorescent Probes Michalet
PALM/STORM, BALM, STED
 PALM/STORM, BALM, STED Last class 2-photon Intro to PALM/STORM Cyanine dyes/dronpa This class Finish localization super-res BALM STED Localization microscopy Intensity Bins = pixels xx 2 = ss2 + aa 2 /12
PALM/STORM, BALM, STED Last class 2-photon Intro to PALM/STORM Cyanine dyes/dronpa This class Finish localization super-res BALM STED Localization microscopy Intensity Bins = pixels xx 2 = ss2 + aa 2 /12
Super-resolution Microscopy
 Semr oc kwhi t epaperser i es : 1. Introduction Super-resolution Microscopy Fluorescence microscopy has revolutionized the study of biological samples. Ever since the invention of fluorescence microscopy
Semr oc kwhi t epaperser i es : 1. Introduction Super-resolution Microscopy Fluorescence microscopy has revolutionized the study of biological samples. Ever since the invention of fluorescence microscopy
Bi177 - Lecture 13 Microscopy Outside the Box. Fluorescence Nanoscopy TIRF 4-pi STED STORM/PALM
 Bi177 - Lecture 13 Microscopy Outside the Box Fluorescence Nanoscopy TIRF 4-pi STED STORM/PALM The diffraction limit: Abbe s law The Problem Diffraction limit 100x larger than molecular scale! Green Fluorescent
Bi177 - Lecture 13 Microscopy Outside the Box Fluorescence Nanoscopy TIRF 4-pi STED STORM/PALM The diffraction limit: Abbe s law The Problem Diffraction limit 100x larger than molecular scale! Green Fluorescent
SIM SSIM. nanoscopy RESOLFT. Super-resolution STORM GSDIM. dstorm PALMIRA FPALM PALM PAINT SPRAIPAINT SOFI BALM CALM. Bo Huang
 STEDGSD STORM SOFI nanoscopy GSDIM PALMIRA SMACM BBB PAINT SPRAIPAINT CALM RESOLFT BALM SIM SSIM Super-resolution Bo Huang 2013.08.01 dstorm FPALM PALM 50 years to extend the resolution Confocal microscopy
STEDGSD STORM SOFI nanoscopy GSDIM PALMIRA SMACM BBB PAINT SPRAIPAINT CALM RESOLFT BALM SIM SSIM Super-resolution Bo Huang 2013.08.01 dstorm FPALM PALM 50 years to extend the resolution Confocal microscopy
Localization Microscopy
 Localization Microscopy Theory, Sample Prep & Practical Considerations Patrina Pellett & Ann McEvoy Applications Scientist GE Healthcare, Cell Technologies May 27 th, 2015 Localization Microscopy Talk
Localization Microscopy Theory, Sample Prep & Practical Considerations Patrina Pellett & Ann McEvoy Applications Scientist GE Healthcare, Cell Technologies May 27 th, 2015 Localization Microscopy Talk
Tracking sub-microscopic protein organisation at the plasma membrane of live cells using triple-colour superresolution
 Tracking sub-microscopic protein organisation at the plasma membrane of live cells using triple-colour superresolution microscopy Jacob Piehler Division of Biophysics, University of Osnabrück, Germany
Tracking sub-microscopic protein organisation at the plasma membrane of live cells using triple-colour superresolution microscopy Jacob Piehler Division of Biophysics, University of Osnabrück, Germany
Introduction CHAPTER 1
 CHAPTER 1 Introduction 1.1 Light Microscopy The light microscope is one of the significant inventions in the history of humankind that, along with the telescope, played a central role in the Scientific
CHAPTER 1 Introduction 1.1 Light Microscopy The light microscope is one of the significant inventions in the history of humankind that, along with the telescope, played a central role in the Scientific
Nanoscale measurement examples in biology: Sub-diffractive imaging & the fly brain challenge
 Nanoscale measurement examples in biology: Sub-diffractive imaging & the fly brain challenge Harald Hess, HHMI Janelia Farm Introduction to HHMI Janelia Farms Sample: Nerves, Worm Brains, Fly Brains, Rodent
Nanoscale measurement examples in biology: Sub-diffractive imaging & the fly brain challenge Harald Hess, HHMI Janelia Farm Introduction to HHMI Janelia Farms Sample: Nerves, Worm Brains, Fly Brains, Rodent
Stochastic Optical Reconstruction Microscopy (STORM): A Method for Superresolution Fluorescence Imaging
 Topic Introduction Stochastic Optical Reconstruction Microscopy (STORM): A Method for Superresolution Fluorescence Imaging Mark Bates, Sara A. Jones, and Xiaowei Zhuang The relatively low spatial resolution
Topic Introduction Stochastic Optical Reconstruction Microscopy (STORM): A Method for Superresolution Fluorescence Imaging Mark Bates, Sara A. Jones, and Xiaowei Zhuang The relatively low spatial resolution
Q&A: Single-molecule localization microscopy for biological imaging
 Q U E S T I O N & ANSWER Q&A: Single-molecule localization microscopy for biological imaging Ann L McEvoy 1, Derek Greenfield 1,2,5, Mark Bates 3 and Jan Liphardt 1,2,4 * Open Access Why is it important
Q U E S T I O N & ANSWER Q&A: Single-molecule localization microscopy for biological imaging Ann L McEvoy 1, Derek Greenfield 1,2,5, Mark Bates 3 and Jan Liphardt 1,2,4 * Open Access Why is it important
Confocal Microscopy & Imaging Technology. Yan Wu
 Confocal Microscopy & Imaging Technology Yan Wu Dec. 05, 2014 Cells under the microscope What we use to see the details of the cell? Light and Electron Microscopy - Bright light / fluorescence microscopy
Confocal Microscopy & Imaging Technology Yan Wu Dec. 05, 2014 Cells under the microscope What we use to see the details of the cell? Light and Electron Microscopy - Bright light / fluorescence microscopy
Super-resolution microscopy at a glance
 ARTICLE SERIES: Imaging Cell Science at a Glance 1607 Super-resolution microscopy at a glance Catherine G. Galbraith 1, * and James A. Galbraith 2, * 1 National Institute of Health, 1 NICHD and 2 NINDS,
ARTICLE SERIES: Imaging Cell Science at a Glance 1607 Super-resolution microscopy at a glance Catherine G. Galbraith 1, * and James A. Galbraith 2, * 1 National Institute of Health, 1 NICHD and 2 NINDS,
Introduction to N-STORM
 Introduction to N-STORM Dan Metcalf Advanced Imaging Manager Outline Introduction Principles of STORM Applications N-STORM overview Biological Scale Mitochondrion Microtubule Amino Acid 1Å Kinesin 1nm
Introduction to N-STORM Dan Metcalf Advanced Imaging Manager Outline Introduction Principles of STORM Applications N-STORM overview Biological Scale Mitochondrion Microtubule Amino Acid 1Å Kinesin 1nm
Super Resolution Imaging Solution Provider. Imaging Future
 Super Resolution Imaging Solution Provider Imaging Future Imaging Solution More Than Equipment NanoBioImaging(NBI) is the Industrial Partner of HKUST Super Resolution Imaging Center (SRIC). NBI aims to
Super Resolution Imaging Solution Provider Imaging Future Imaging Solution More Than Equipment NanoBioImaging(NBI) is the Industrial Partner of HKUST Super Resolution Imaging Center (SRIC). NBI aims to
Lab 5: Optical trapping and single molecule fluorescence
 Lab 5: Optical trapping and single molecule fluorescence PI: Matt Lang Lab Instructor: Jorge Ferrer Summary Optical tweezers are an excellent experimental tool to study the biophysics of single molecule
Lab 5: Optical trapping and single molecule fluorescence PI: Matt Lang Lab Instructor: Jorge Ferrer Summary Optical tweezers are an excellent experimental tool to study the biophysics of single molecule
Fluorescence Nanoscopy 高甫仁 ) Institute of Biophotonics, National Yang Ming University. Outline
 Fluorescence Nanoscopy 高甫仁 ) Fu-Jen Kao ( 高甫仁 Institute of Biophotonics, National Yang Ming University Outline The Abbe s (diffraction) limit and nanoscopy Fundamentals and opportunities of FLIM/FRET Visualizing
Fluorescence Nanoscopy 高甫仁 ) Fu-Jen Kao ( 高甫仁 Institute of Biophotonics, National Yang Ming University Outline The Abbe s (diffraction) limit and nanoscopy Fundamentals and opportunities of FLIM/FRET Visualizing
Fast, three-dimensional super-resolution imaging of live cells
 Nature Methods Fast, three-dimensional super-resolution imaging of live cells Sara A Jones, Sang-Hee Shim, Jiang He & Xiaowei Zhuang Supplementary Figure 1 Supplementary Figure 2 Supplementary Figure 3
Nature Methods Fast, three-dimensional super-resolution imaging of live cells Sara A Jones, Sang-Hee Shim, Jiang He & Xiaowei Zhuang Supplementary Figure 1 Supplementary Figure 2 Supplementary Figure 3
A Brief History of Light Microscopy And How It Transformed Biomedical Research
 A Brief History of Light Microscopy And How It Transformed Biomedical Research Suewei Lin Office: Interdisciplinary Research Building 8A08 Email: sueweilin@gate.sinica.edu.tw TEL: 2789-9315 Microscope
A Brief History of Light Microscopy And How It Transformed Biomedical Research Suewei Lin Office: Interdisciplinary Research Building 8A08 Email: sueweilin@gate.sinica.edu.tw TEL: 2789-9315 Microscope
PEER REVIEW FILE. Reviewers' comments: Reviewer #1 (Remarks to the Author):
 PEER REVIEW FILE Reviewers' comments: Reviewer #1 (Remarks to the Author): General In its beginnings in the mid 1990s, localization microscopy based on optical isolation of fluorescent point targets was
PEER REVIEW FILE Reviewers' comments: Reviewer #1 (Remarks to the Author): General In its beginnings in the mid 1990s, localization microscopy based on optical isolation of fluorescent point targets was
PALM Microscope User Manual
 PALM Microscope User Manual Stephan Uphoff, Mathew Stracy, Federico Garza de Leon Microscope components: The laser excitation is delivered from a Toptica Multi Laser Engine with 405 nm, 488 nm, 561 nm,
PALM Microscope User Manual Stephan Uphoff, Mathew Stracy, Federico Garza de Leon Microscope components: The laser excitation is delivered from a Toptica Multi Laser Engine with 405 nm, 488 nm, 561 nm,
Supplementary Table 1. Components of an FCS setup (1PE and 2PE)
 Supplementary Table 1. Components of an FCS setup (1PE and 2PE) Component and function Laser source Excitation of fluorophores Microscope with xy-translation stage mounted on vibration isolated optical
Supplementary Table 1. Components of an FCS setup (1PE and 2PE) Component and function Laser source Excitation of fluorophores Microscope with xy-translation stage mounted on vibration isolated optical
HHS Public Access Author manuscript Phys Rev Lett. Author manuscript; available in PMC 2014 March 18.
 Photo-imprint Photoacoustic Microscopy for Three-dimensional Label-free Sub-diffraction Imaging Junjie Yao, Lidai Wang, Chiye Li, Chi Zhang, and Lihong V. Wang * Optical Imaging Laboratory, Department
Photo-imprint Photoacoustic Microscopy for Three-dimensional Label-free Sub-diffraction Imaging Junjie Yao, Lidai Wang, Chiye Li, Chi Zhang, and Lihong V. Wang * Optical Imaging Laboratory, Department
Measure of surface protein mobility with u-paint technique
 Measure of surface protein mobility with u-paint technique How dynamic image can solve the situation? Random distribution or cluster? Why live super-resolution microscopy can solve the situation With mobility
Measure of surface protein mobility with u-paint technique How dynamic image can solve the situation? Random distribution or cluster? Why live super-resolution microscopy can solve the situation With mobility
Super-resolution imaging: early days w/ Video-enhanced DIC, TIRF, PALM, STORM, etc.
 15/05/2012 Super-resolution imaging: early days w/ Video-enhanced DIC, TIRF, PALM, STORM, etc. Prof. Dr. Rainer Duden duden@bio.uni-luebeck.de 1 Using conventional light microscopy resolution is limited
15/05/2012 Super-resolution imaging: early days w/ Video-enhanced DIC, TIRF, PALM, STORM, etc. Prof. Dr. Rainer Duden duden@bio.uni-luebeck.de 1 Using conventional light microscopy resolution is limited
Fluorescence Light Microscopy for Cell Biology
 Fluorescence Light Microscopy for Cell Biology Why use light microscopy? Traditional questions that light microscopy has addressed: Structure within a cell Locations of specific molecules within a cell
Fluorescence Light Microscopy for Cell Biology Why use light microscopy? Traditional questions that light microscopy has addressed: Structure within a cell Locations of specific molecules within a cell
Direct stochastic optical reconstruction microscopy with standard fluorescent probes
 Direct stochastic optical reconstruction microscopy with standard fluorescent probes Sebastian van de Linde, Anna Löschberger, Teresa Klein, Meike Heidbreder, Steve Wolter, Mike Heilemann & Markus Sauer
Direct stochastic optical reconstruction microscopy with standard fluorescent probes Sebastian van de Linde, Anna Löschberger, Teresa Klein, Meike Heidbreder, Steve Wolter, Mike Heilemann & Markus Sauer
Two-Photon Microscopy for Deep Tissue Imaging of Living Specimens
 for Deep Tissue Imaging of Living Specimens Tilman Franke* and Sebastian Rhode TILL Photonics GmbH, an FEI company, Lochhamer Schlag 21, D-82166 Gräfelfing, Germany *tilman.franke@fei.com Introduction
for Deep Tissue Imaging of Living Specimens Tilman Franke* and Sebastian Rhode TILL Photonics GmbH, an FEI company, Lochhamer Schlag 21, D-82166 Gräfelfing, Germany *tilman.franke@fei.com Introduction
Microscopy from Carl Zeiss. DirectFRAP. News from the Cell. The New Class of Laser Manipulation for the Analysis of Cell Dynamics
 Microscopy from Carl Zeiss DirectFRAP News from the Cell The New Class of Laser Manipulation for the Analysis of Cell Dynamics DirectFRAP. New Insights into Cell Dynamics. Fluorescence breaks new ground:
Microscopy from Carl Zeiss DirectFRAP News from the Cell The New Class of Laser Manipulation for the Analysis of Cell Dynamics DirectFRAP. New Insights into Cell Dynamics. Fluorescence breaks new ground:
Nature Neuroscience: doi: /nn Supplementary Figure 1. Comprehensive opto-mechanical design of the dual-axis microscope.
 Supplementary Figure 1 Comprehensive opto-mechanical design of the dual-axis microscope. (a) The complete microscope is shown to scale. The laser beam is depicted in light red. (b) A close-up view of the
Supplementary Figure 1 Comprehensive opto-mechanical design of the dual-axis microscope. (a) The complete microscope is shown to scale. The laser beam is depicted in light red. (b) A close-up view of the
Fluorescence microscopy
 Fluorescence microscopy 1 Fluorescence microscopies basic fluorescence, fluorophores Deconvolution Confocal Two-photon/multi-photon 4Pi Light sheet Total internal reflection STED FRAP/FLIP/FCS FRET PALM/STORM/iPALM
Fluorescence microscopy 1 Fluorescence microscopies basic fluorescence, fluorophores Deconvolution Confocal Two-photon/multi-photon 4Pi Light sheet Total internal reflection STED FRAP/FLIP/FCS FRET PALM/STORM/iPALM
Fluorescent probes for superresolution imaging in living cells
 Fluorescent probes for superresolution imaging in living cells Marta Fernández-Suárez* and Alice Y. Ting* Abstract In 1873, Ernst Abbe discovered that features closer than ~200 nm cannot be resolved by
Fluorescent probes for superresolution imaging in living cells Marta Fernández-Suárez* and Alice Y. Ting* Abstract In 1873, Ernst Abbe discovered that features closer than ~200 nm cannot be resolved by
Sample region with fluorescent labeled molecules
 FLUORESCENCE IMAGING I. Fluorescence-imaging with diffraction limited spots The resolution in optical microscopy has been hampered by the smallest spot possible (~ λ/2) that can be achieved by conventional
FLUORESCENCE IMAGING I. Fluorescence-imaging with diffraction limited spots The resolution in optical microscopy has been hampered by the smallest spot possible (~ λ/2) that can be achieved by conventional
Microscopy. CS/CME/BioE/Biophys/BMI 279 Nov. 2, 2017 Ron Dror
 Microscopy CS/CME/BioE/Biophys/BMI 279 Nov. 2, 2017 Ron Dror 1 Outline Microscopy: the basics Fluorescence microscopy Resolution limits The diffraction limit Beating the diffraction limit 2 Microscopy:
Microscopy CS/CME/BioE/Biophys/BMI 279 Nov. 2, 2017 Ron Dror 1 Outline Microscopy: the basics Fluorescence microscopy Resolution limits The diffraction limit Beating the diffraction limit 2 Microscopy:
Super Resolution Microscopy - Breaking the Diffraction Limit Radiological Research Accelerator Facility
 Super Resolution Microscopy - Breaking the Diffraction Limit Radiological Research Accelerator Facility Sabrina Campelo, Dr. Andrew Harken Outline Motivation Fluorescence Microscopy -Multiphoton Imaging
Super Resolution Microscopy - Breaking the Diffraction Limit Radiological Research Accelerator Facility Sabrina Campelo, Dr. Andrew Harken Outline Motivation Fluorescence Microscopy -Multiphoton Imaging
Special Techniques 1. Mark Scott FILM Facility
 Special Techniques 1 Mark Scott FILM Facility SPECIAL TECHNIQUES Multi-photon microscopy Second Harmonic Generation FRAP FRET FLIM In-vivo imaging TWO-PHOTON MICROSCOPY Alternative to confocal and deconvolution
Special Techniques 1 Mark Scott FILM Facility SPECIAL TECHNIQUES Multi-photon microscopy Second Harmonic Generation FRAP FRET FLIM In-vivo imaging TWO-PHOTON MICROSCOPY Alternative to confocal and deconvolution
CHARACTERIZATION OF MOLECULAR ORIENTATION IN SUPER-RESOLUTION FLUORESCENCE MICROSCOPY
 Master Erasmus Mundus in Photonics Engineering, Nanophotonics and Biophotonics Europhotonics MASTER THESIS WORK CHARACTERIZATION OF MOLECULAR ORIENTATION IN SUPER-RESOLUTION FLUORESCENCE MICROSCOPY Yibing
Master Erasmus Mundus in Photonics Engineering, Nanophotonics and Biophotonics Europhotonics MASTER THESIS WORK CHARACTERIZATION OF MOLECULAR ORIENTATION IN SUPER-RESOLUTION FLUORESCENCE MICROSCOPY Yibing
Imagerie et spectroscopie de fluorescence par excitation non radiative
 Imagerie et spectroscopie de fluorescence par excitation non radiative comment s affranchir de la limite de diffraction Rodolphe Jaffiol, Cyrille Vézy, Marcelina Cardoso Dos Santos LNIO, UTT, Troyes NanoBioPhotonics
Imagerie et spectroscopie de fluorescence par excitation non radiative comment s affranchir de la limite de diffraction Rodolphe Jaffiol, Cyrille Vézy, Marcelina Cardoso Dos Santos LNIO, UTT, Troyes NanoBioPhotonics
Simultaneous multi-color, multiphoton fluorophore excitation using dual-color fiber lasers
 Multiphoton Microscopy / Fiber Laser Simultaneous multi-color, multiphoton fluorophore excitation using dual-color fiber lasers Matthias Handloser, Tim Paasch-Colberg, Bernhard Wolfring TOPTICA Photonics
Multiphoton Microscopy / Fiber Laser Simultaneous multi-color, multiphoton fluorophore excitation using dual-color fiber lasers Matthias Handloser, Tim Paasch-Colberg, Bernhard Wolfring TOPTICA Photonics
FLIM Fluorescence Lifetime IMaging
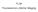 FLIM Fluorescence Lifetime IMaging Fluorescence lifetime t I(t) = F0 exp( ) τ 1 τ = k f + k nr k nr = k IC + k ISC + k bl Batiaens et al, Trends in Cell Biology, 1999 τ τ = fluorescence lifetime (~ns to
FLIM Fluorescence Lifetime IMaging Fluorescence lifetime t I(t) = F0 exp( ) τ 1 τ = k f + k nr k nr = k IC + k ISC + k bl Batiaens et al, Trends in Cell Biology, 1999 τ τ = fluorescence lifetime (~ns to
Widefield Microscopy Bleed-Through
 In widefield microscopy the excitation wavelengths which illuminate the sample, and the emission wavelengths which reach the CCD camera are selected throughout a filter cube. A filter cube consists of
In widefield microscopy the excitation wavelengths which illuminate the sample, and the emission wavelengths which reach the CCD camera are selected throughout a filter cube. A filter cube consists of
Single-Particle Tracking Photoactivated Localization Microscopy for Mapping Single-Molecule Dynamics
 CHAPTER FIVE Single-Particle Tracking Photoactivated Localization Microscopy for Mapping Single-Molecule Dynamics Suliana Manley,* Jennifer M. Gillette, and Jennifer Lippincott-Schwartz Contents 1. Introduction
CHAPTER FIVE Single-Particle Tracking Photoactivated Localization Microscopy for Mapping Single-Molecule Dynamics Suliana Manley,* Jennifer M. Gillette, and Jennifer Lippincott-Schwartz Contents 1. Introduction
THE ADVANCED IMAGING CENTER AT JANELIA RESEARCH CAMPUS. Call for Proposals
 THE ADVANCED IMAGING CENTER AT JANELIA RESEARCH CAMPUS Call for Proposals janelia.org/aic Call for Proposals THE ADVANCED IMAGING CENTER AT JANELIA RESEARCH CAMPUS We are now accepting proposals from scientists
THE ADVANCED IMAGING CENTER AT JANELIA RESEARCH CAMPUS Call for Proposals janelia.org/aic Call for Proposals THE ADVANCED IMAGING CENTER AT JANELIA RESEARCH CAMPUS We are now accepting proposals from scientists
Multiplexed 3D FRET imaging in deep tissue of live embryos Ming Zhao, Xiaoyang Wan, Yu Li, Weibin Zhou and Leilei Peng
 Scientific Reports Multiplexed 3D FRET imaging in deep tissue of live embryos Ming Zhao, Xiaoyang Wan, Yu Li, Weibin Zhou and Leilei Peng 1 Supplementary figures and notes Supplementary Figure S1 Volumetric
Scientific Reports Multiplexed 3D FRET imaging in deep tissue of live embryos Ming Zhao, Xiaoyang Wan, Yu Li, Weibin Zhou and Leilei Peng 1 Supplementary figures and notes Supplementary Figure S1 Volumetric
Super-Resolution Localization Microscopy
 APPLICATION NOTE Super-Resolution Localization Microscopy Light microscopy techniques have been vital to our understanding of biological structures and systems since their invention in the late 16 th Century.
APPLICATION NOTE Super-Resolution Localization Microscopy Light microscopy techniques have been vital to our understanding of biological structures and systems since their invention in the late 16 th Century.
Confocal Microscopy Analyzes Cells
 Choosing Filters for Fluorescence A Laurin Publication Photonic Solutions for Biotechnology and Medicine November 2002 Confocal Microscopy Analyzes Cells Reprinted from the November 2002 issue of Biophotonics
Choosing Filters for Fluorescence A Laurin Publication Photonic Solutions for Biotechnology and Medicine November 2002 Confocal Microscopy Analyzes Cells Reprinted from the November 2002 issue of Biophotonics
SUPER-RESOLUTION MICROSCOPY. Dr. Nathalie Garin
 SUPER-RESOLUTION MICROSCOPY Dr. Nathalie Garin Content Motivation for superresolution Superresolution, nanoscopy, : definition Structured Illumination Microscopy (SIM) Localization microscopy STimulated
SUPER-RESOLUTION MICROSCOPY Dr. Nathalie Garin Content Motivation for superresolution Superresolution, nanoscopy, : definition Structured Illumination Microscopy (SIM) Localization microscopy STimulated
Macromolecular environments influence proteins
 Research & Development Protein Dynamics Macromolecular environments influence proteins 6 www.q-more.com/en/ q&more 01.16 Studying proteins in the presence of high concentrations of macromolecules ( molecular
Research & Development Protein Dynamics Macromolecular environments influence proteins 6 www.q-more.com/en/ q&more 01.16 Studying proteins in the presence of high concentrations of macromolecules ( molecular
Fs- Using Ultrafast Lasers to Add New Functionality to Glass
 An IMI Video Reproduction of Invited Lectures from the 17th University Glass Conference Fs- Using Ultrafast Lasers to Add New Functionality to Glass Denise M. Krol University of California, Davis 17th
An IMI Video Reproduction of Invited Lectures from the 17th University Glass Conference Fs- Using Ultrafast Lasers to Add New Functionality to Glass Denise M. Krol University of California, Davis 17th
Three Dimensional Orientation of Anisotropic. Plasmonic Aggregates at Intracellular Nuclear. Indentation Sites by Integrated Light Sheet Super-
 Supporting Information Three Dimensional Orientation of Anisotropic Plasmonic Aggregates at Intracellular Nuclear Indentation Sites y Integrated Light Sheet Super- Resolution Microscopy Suresh Kumar Chakkarapani,
Supporting Information Three Dimensional Orientation of Anisotropic Plasmonic Aggregates at Intracellular Nuclear Indentation Sites y Integrated Light Sheet Super- Resolution Microscopy Suresh Kumar Chakkarapani,
HYPERSPECTRAL MICROSCOPE PLATFORM FOR HIGHLY MULTIPLEX BIOLOGICAL IMAGING. Marc Verhaegen
 HYPERSPECTRAL MICROSCOPE PLATFORM FOR HIGHLY MULTIPLEX BIOLOGICAL IMAGING Marc Verhaegen CMCS, MONTREAL, MAY 11 th, 2017 OVERVIEW Hyperspectral Imaging Multiplex Biological Imaging Multiplex Single Particle
HYPERSPECTRAL MICROSCOPE PLATFORM FOR HIGHLY MULTIPLEX BIOLOGICAL IMAGING Marc Verhaegen CMCS, MONTREAL, MAY 11 th, 2017 OVERVIEW Hyperspectral Imaging Multiplex Biological Imaging Multiplex Single Particle
High Throughput Whole Organ Imaging Based on Multifocal Multiphoton Microscope
 High Throughput Whole Organ Imaging Based on Multifocal Multiphoton Microscope LBRC researchers: Peter So, Jae Won Cha, Elijah Yew, Vijay Singh External technology collaborators: Prof. Hanry Yu (University
High Throughput Whole Organ Imaging Based on Multifocal Multiphoton Microscope LBRC researchers: Peter So, Jae Won Cha, Elijah Yew, Vijay Singh External technology collaborators: Prof. Hanry Yu (University
Active delivery of single DNA molecules into a plasmonic nanopore for. label-free optical sensing
 Supporting Information: Active delivery of single DNA molecules into a plasmonic nanopore for label-free optical sensing Xin Shi 1,2, Daniel V Verschueren 1, and Cees Dekker 1* 1. Department of Bionanoscience,
Supporting Information: Active delivery of single DNA molecules into a plasmonic nanopore for label-free optical sensing Xin Shi 1,2, Daniel V Verschueren 1, and Cees Dekker 1* 1. Department of Bionanoscience,
MicroTime 200 STED. Super-resolution add-on for the confocal time-resolved microscopy platform
 MicroTime 200 STED Super-resolution add-on for the confocal time-resolved microscopy platform confocal STED 2 Vision The MicroTime 200... The MicroTime 200 is a high-end confocal fluorescence lifetime
MicroTime 200 STED Super-resolution add-on for the confocal time-resolved microscopy platform confocal STED 2 Vision The MicroTime 200... The MicroTime 200 is a high-end confocal fluorescence lifetime
SUPPLEMENTARY INFORMATION
 Measuring subwavelength spatial coherence with plasmonic interferometry Drew Morrill, Dongfang Li, and Domenico Pacifici School of Engineering, Brown University, Providence, RI 02912, United States List
Measuring subwavelength spatial coherence with plasmonic interferometry Drew Morrill, Dongfang Li, and Domenico Pacifici School of Engineering, Brown University, Providence, RI 02912, United States List
Photoactivatable mcherry for high-resolution two-color fluorescence microscopy
 Photoactivatable mcherry for high-resolution two-color fluorescence microscopy Fedor V Subach 1 3, George H Patterson,3, Suliana Manley, Jennifer M Gillette, Jennifer Lippincott-Schwartz & Vladislav V
Photoactivatable mcherry for high-resolution two-color fluorescence microscopy Fedor V Subach 1 3, George H Patterson,3, Suliana Manley, Jennifer M Gillette, Jennifer Lippincott-Schwartz & Vladislav V
Spectral Separation of Multifluorescence Labels with the LSM 510 META
 Microscopy from Carl Zeiss Spectral Separation of Multifluorescence Labels with the LSM 510 META Indians living in the South American rain forest can distinguish between almost 200 hues of green in their
Microscopy from Carl Zeiss Spectral Separation of Multifluorescence Labels with the LSM 510 META Indians living in the South American rain forest can distinguish between almost 200 hues of green in their
Imaging facilities at WUR
 Imaging facilities at WUR Advanced light microscopy facilities at Wageningen UR Programme Thursday 13 June 2013 Lunch meeting organized by Cat-Agro Food 12.00 Welcome and sandwich lunch 12.10 Introduction
Imaging facilities at WUR Advanced light microscopy facilities at Wageningen UR Programme Thursday 13 June 2013 Lunch meeting organized by Cat-Agro Food 12.00 Welcome and sandwich lunch 12.10 Introduction
Introduction to Computational Fluorescence Microscopy!
 Introduction to Computational Fluorescence Microscopy! EE367/CS448I: Computational Imaging and Display! stanford.edu/class/ee367! Lecture 13! Gordon Wetzstein! Stanford University! Midterm! Tuesday, Feb
Introduction to Computational Fluorescence Microscopy! EE367/CS448I: Computational Imaging and Display! stanford.edu/class/ee367! Lecture 13! Gordon Wetzstein! Stanford University! Midterm! Tuesday, Feb
Breaking the Diffraction Barrier: Super-Resolution Imaging of Cells
 Leading Edge Primer Breaking the Diffraction Barrier: Super-Resolution Imaging of Cells Bo Huang, 1 Hazen Babcock, 2 and Xiaowei Zhuang 2,3, * 1 Department of Pharmaceutical Chemistry and Department of
Leading Edge Primer Breaking the Diffraction Barrier: Super-Resolution Imaging of Cells Bo Huang, 1 Hazen Babcock, 2 and Xiaowei Zhuang 2,3, * 1 Department of Pharmaceutical Chemistry and Department of
Supporting Information
 Supporting Information Azido Push Pull Fluorogens Photoactivate to Produce Bright Fluorescent Labels Samuel J. Lord, Hsiao-lu D. Lee, Reichel Samuel, Ryan Weber, Na Liu, Nicholas R. Conley, Michael A.
Supporting Information Azido Push Pull Fluorogens Photoactivate to Produce Bright Fluorescent Labels Samuel J. Lord, Hsiao-lu D. Lee, Reichel Samuel, Ryan Weber, Na Liu, Nicholas R. Conley, Michael A.
Building An Ultrafast Photon-Induced Near-field Transmission Electron Microscope
 Building An Ultrafast Photon-Induced Near-field Transmission Electron Microscope Dr. Tom T.A. Lummen École Polytechnique Fédérale de Lausanne -- LUMES Photonic Instruments 2013 September 11, 2013 Zürich,
Building An Ultrafast Photon-Induced Near-field Transmission Electron Microscope Dr. Tom T.A. Lummen École Polytechnique Fédérale de Lausanne -- LUMES Photonic Instruments 2013 September 11, 2013 Zürich,
Nanoscale Spatial Organization of Prokaryotic Cells Studied by Super-Resolution Optical Microscopy. Andrea Lynn McEvoy
 Nanoscale Spatial Organization of Prokaryotic Cells Studied by Super-Resolution Optical Microscopy by Andrea Lynn McEvoy A dissertation submitted in partial satisfaction of the requirements for the degree
Nanoscale Spatial Organization of Prokaryotic Cells Studied by Super-Resolution Optical Microscopy by Andrea Lynn McEvoy A dissertation submitted in partial satisfaction of the requirements for the degree
Digital resolution enhancement in surface plasmon microscopy
 Digital resolution enhancement in surface plasmon microscopy I.I. Smolyaninov 1) *, J. Elliott 2), G. Wurtz 2), A.V. Zayats 2), C.C. Davis 1) 1) Department of Electrical and Computer Engineering, University
Digital resolution enhancement in surface plasmon microscopy I.I. Smolyaninov 1) *, J. Elliott 2), G. Wurtz 2), A.V. Zayats 2), C.C. Davis 1) 1) Department of Electrical and Computer Engineering, University
Supporting Information for. Electrical control of Förster energy transfer.
 1 Supporting Information for Electrical control of Förster energy transfer. Klaus Becker 1, John M. Lupton 1*, Josef Müller 1, Andrey. L. Rogach 1, Dmitri V. Talapin, Horst Weller & Jochen Feldmann 1 1
1 Supporting Information for Electrical control of Förster energy transfer. Klaus Becker 1, John M. Lupton 1*, Josef Müller 1, Andrey. L. Rogach 1, Dmitri V. Talapin, Horst Weller & Jochen Feldmann 1 1
New developments in STED Microscopy
 New developments in STED Microscopy Arnold Giske*, Jochen Sieber, Hilmar Gugel, Marcus Dyba, Volker Seyfried, Dietmar Gnass Leica Microsystems CMS, Am Friedensplatz 3, 68126 Mannheim, Germany ABSTRACT
New developments in STED Microscopy Arnold Giske*, Jochen Sieber, Hilmar Gugel, Marcus Dyba, Volker Seyfried, Dietmar Gnass Leica Microsystems CMS, Am Friedensplatz 3, 68126 Mannheim, Germany ABSTRACT
Supporting Information. Two-Photon Luminescence of Single Colloidal Gold NanoRods: Revealing the Origin of Plasmon Relaxation in Small Nanocrystals
 Supporting Information Two-Photon Luminescence of Single Colloidal Gold NanoRods: Revealing the Origin of Plasmon Relaxation in Small Nanocrystals Céline Molinaro 1, Yara El Harfouch 1, Etienne Palleau
Supporting Information Two-Photon Luminescence of Single Colloidal Gold NanoRods: Revealing the Origin of Plasmon Relaxation in Small Nanocrystals Céline Molinaro 1, Yara El Harfouch 1, Etienne Palleau
F* techniques: FRAP, FLIP, FRET, FLIM,
 F* techniques: FRAP, FLIP, FRET, FLIM, FCS Antonia Göhler March 2015 Fluorescence explained in the Bohr model Absorption of light (blue) causes an electron to move to a higher energy orbit. After a particular
F* techniques: FRAP, FLIP, FRET, FLIM, FCS Antonia Göhler March 2015 Fluorescence explained in the Bohr model Absorption of light (blue) causes an electron to move to a higher energy orbit. After a particular
BIO 315 Lab Exam I. Section #: Name:
 Section #: Name: Also provide this information on the computer grid sheet given to you. (Section # in special code box) BIO 315 Lab Exam I 1. In labeling the parts of a standard compound light microscope
Section #: Name: Also provide this information on the computer grid sheet given to you. (Section # in special code box) BIO 315 Lab Exam I 1. In labeling the parts of a standard compound light microscope
Supplementary Information
 Supplementary Information STED nanoscopy combined with optical tweezers reveals protein dynamics on densely covered DNA Iddo Heller, Gerrit Sitters, Onno D. Broekmans, Géraldine Farge, Carolin Menges,
Supplementary Information STED nanoscopy combined with optical tweezers reveals protein dynamics on densely covered DNA Iddo Heller, Gerrit Sitters, Onno D. Broekmans, Géraldine Farge, Carolin Menges,
Femtosecond micromachining in polymers
 Femtosecond micromachining in polymers Prof. Dr Cleber R. Mendonca Daniel S. Corrêa Prakriti Tayalia Dr. Tobias Voss Dr. Tommaso Baldacchini Prof. Dr. Eric Mazur fs-micromachining focus laser beam inside
Femtosecond micromachining in polymers Prof. Dr Cleber R. Mendonca Daniel S. Corrêa Prakriti Tayalia Dr. Tobias Voss Dr. Tommaso Baldacchini Prof. Dr. Eric Mazur fs-micromachining focus laser beam inside
STED microscopy with single light source. TeodoraŞcheul
 STED microscopy with single light source TeodoraŞcheul Dr. Iréne Wang, Dr. Jean-Claude Vial LIPhy, Grenoble, France Summary I. Introduction to STED microscopy II. STED with one laser source 1. Two-photon
STED microscopy with single light source TeodoraŞcheul Dr. Iréne Wang, Dr. Jean-Claude Vial LIPhy, Grenoble, France Summary I. Introduction to STED microscopy II. STED with one laser source 1. Two-photon
a) JOURNAL OF BIOLOGICAL CHEMISTRY b) PNAS c) NATURE
 a) JOURNAL OF BIOLOGICAL CHEMISTRY b) c) d) ........................ JOURNAL OF BIOLOGICAL CHEMISTRY MOLECULAR PHARMACOLOGY TRENDS IN PHARMACOLOGICAL S AMERICAN JOURNAL OF PHYSIOLOGY-HEART AND CIRCULATORY
a) JOURNAL OF BIOLOGICAL CHEMISTRY b) c) d) ........................ JOURNAL OF BIOLOGICAL CHEMISTRY MOLECULAR PHARMACOLOGY TRENDS IN PHARMACOLOGICAL S AMERICAN JOURNAL OF PHYSIOLOGY-HEART AND CIRCULATORY
Lab 1: Ensemble Fluorescence Basics
 Lab 1: Ensemble Fluorescence Basics This laboratory module is divided into two sections. The first one is on organic fluorophores, and the second one is on ensemble measurement of FRET (Fluorescence Resonance
Lab 1: Ensemble Fluorescence Basics This laboratory module is divided into two sections. The first one is on organic fluorophores, and the second one is on ensemble measurement of FRET (Fluorescence Resonance
Fluorescence Microscopy. Terms and concepts to know: 10/11/2011. Visible spectrum (of light) and energy
 Fluorescence Microscopy Louisiana Tech University Ruston, Louisiana Microscopy Workshop Dr. Mark DeCoster Associate Professor Biomedical Engineering 1 Terms and concepts to know: Signal to Noise Excitation
Fluorescence Microscopy Louisiana Tech University Ruston, Louisiana Microscopy Workshop Dr. Mark DeCoster Associate Professor Biomedical Engineering 1 Terms and concepts to know: Signal to Noise Excitation
Contact Details. Dr Alexander Galkin. Office: MBC Room 186. Tel: (028) Frequency and wavelength.
 Contact Details The electromagnetic spectrum Biological Spectroscopy Dr Alexander Galkin Email: a.galkin@qub.ac.uk Dr Alexander Galkin MSc Biomolecular Function - BBC8045 Office: MBC Room 186 Tel: (028)
Contact Details The electromagnetic spectrum Biological Spectroscopy Dr Alexander Galkin Email: a.galkin@qub.ac.uk Dr Alexander Galkin MSc Biomolecular Function - BBC8045 Office: MBC Room 186 Tel: (028)
Dino-Lite knowledge & education. Fluorescence Microscopes
 Dino-Lite knowledge & education Fluorescence Microscopes Dino-Lite Fluorescence models Smallest fluorescence microscope in the world Revolution to biomedical and educational applications Flexible Easy
Dino-Lite knowledge & education Fluorescence Microscopes Dino-Lite Fluorescence models Smallest fluorescence microscope in the world Revolution to biomedical and educational applications Flexible Easy
Peter D. Dahlberg 1, Annina M. Sartor 1, Jiarui Wang 1,2, Saumya Saurabh 2, Lucy Shapiro 2, W.E. Moerner 1
 Peter D. Dahlberg 1, Annina M. Sartor 1, Jiarui Wang 1,2, Saumya Saurabh 2, Lucy Shapiro 2, W.E. Moerner 1 1 Department of Chemistry, Stanford University, Stanford, CA 94305, United States 2 Department
Peter D. Dahlberg 1, Annina M. Sartor 1, Jiarui Wang 1,2, Saumya Saurabh 2, Lucy Shapiro 2, W.E. Moerner 1 1 Department of Chemistry, Stanford University, Stanford, CA 94305, United States 2 Department
Evaluation of fluorophores for optimal performance in localization-based super-resolution imaging
 Evaluation of fluorophores for optimal performance in localization-based super-resolution imaging Graham T Dempsey 1,6, Joshua C Vaughan 2,3,6, Kok Hao Chen 3,6, Mark Bates 4 & Xiaowei Zhuang 2,3,5 211
Evaluation of fluorophores for optimal performance in localization-based super-resolution imaging Graham T Dempsey 1,6, Joshua C Vaughan 2,3,6, Kok Hao Chen 3,6, Mark Bates 4 & Xiaowei Zhuang 2,3,5 211
Introduction to Fluorescence Jablonski Diagram
 ntroduction to Fluorescence Jablonski Diagram Excited Singlet Manifold S1 internal conversion S2 k -isc k isc Excited riplet Manifold 1 S0 k nr k k' f nr fluorescence k p phosphorescence Singlet round
ntroduction to Fluorescence Jablonski Diagram Excited Singlet Manifold S1 internal conversion S2 k -isc k isc Excited riplet Manifold 1 S0 k nr k k' f nr fluorescence k p phosphorescence Singlet round
FLUORESCENCE. Matyas Molnar and Dirk Pacholsky
 FLUORESCENCE Matyas Molnar and Dirk Pacholsky 1 Information This lecture contains images and information from the following internet homepages http://micro.magnet.fsu.edu/primer/index.html http://www.microscopyu.com/
FLUORESCENCE Matyas Molnar and Dirk Pacholsky 1 Information This lecture contains images and information from the following internet homepages http://micro.magnet.fsu.edu/primer/index.html http://www.microscopyu.com/
Live cell microscopy
 Live cell microscopy 1. Why do live cell microscopy? 2. Maintaining living cells on a microscope stage. 3. Considerations for imaging living cells. 4. Fluorescence labeling of living cells. 5. Imaging
Live cell microscopy 1. Why do live cell microscopy? 2. Maintaining living cells on a microscope stage. 3. Considerations for imaging living cells. 4. Fluorescence labeling of living cells. 5. Imaging
Visualisation, Sizing and Counting of Fluorescent and Fluorescently-Labelled Nanoparticles
 Visualisation, Sizing and Counting of Fluorescent and Fluorescently-Labelled Nanoparticles Introduction Fluorescent molecules have long been used to specifically label particular structures and features
Visualisation, Sizing and Counting of Fluorescent and Fluorescently-Labelled Nanoparticles Introduction Fluorescent molecules have long been used to specifically label particular structures and features
Crystallographic Characterization of GaN Nanowires by Raman Spectral Image Mapping
 Crystallographic Characterization of GaN Nanowires by Raman Spectral Image Mapping Heerad Farkhoor, Adam Schwartzberg, Jeffrey Urban August 12, 2009 Abstract Obtaining structural information of nano-structured
Crystallographic Characterization of GaN Nanowires by Raman Spectral Image Mapping Heerad Farkhoor, Adam Schwartzberg, Jeffrey Urban August 12, 2009 Abstract Obtaining structural information of nano-structured
Confocal Microscopes. Evolution of Imaging
 Confocal Microscopes and Evolution of Imaging Judi Reilly Hans Richter Massachusetts Institute of Technology Environment, Health & Safety Office Radiation Protection What is Confocal? Pinhole diaphragm
Confocal Microscopes and Evolution of Imaging Judi Reilly Hans Richter Massachusetts Institute of Technology Environment, Health & Safety Office Radiation Protection What is Confocal? Pinhole diaphragm
Abstract. Introduction
 Abstract The investigation of interactions between engineered nanostructures and biological systems is a key component in the assessment of potential environmental and health implications due to the increasing
Abstract The investigation of interactions between engineered nanostructures and biological systems is a key component in the assessment of potential environmental and health implications due to the increasing
D e c N o. 2 8
 D e c. 2 0 0 7 N o. 2 8 CONFOCAL APPLICATION LETTER resolution FRET Acceptor Photobleaching LAS AF Application Wizard FRET with Leica TCS SP5 LAS AF Version 1.7.0 Introduction Fluorescence Resonance Energy
D e c. 2 0 0 7 N o. 2 8 CONFOCAL APPLICATION LETTER resolution FRET Acceptor Photobleaching LAS AF Application Wizard FRET with Leica TCS SP5 LAS AF Version 1.7.0 Introduction Fluorescence Resonance Energy
Next-generation optical microscopy
 Next-generation optical microscopy Rahul Roy* Department of Chemical Engineering, and Molecular Biophysics Unit, Indian Institute of Science, Bangalore 560 012, India A new breed of microscopy techniques
Next-generation optical microscopy Rahul Roy* Department of Chemical Engineering, and Molecular Biophysics Unit, Indian Institute of Science, Bangalore 560 012, India A new breed of microscopy techniques
Measurement of surface concentration of fluorophores using. fluorescence fluctuation spectroscopy
 Measurement of surface concentration of fluorophores using fluorescence fluctuation spectroscopy A. Delon 1, J. Derouard 1, G. Delapierre and R. Jaffiol 1 (1) Laboratoire de Spectrométrie Physique (UMR
Measurement of surface concentration of fluorophores using fluorescence fluctuation spectroscopy A. Delon 1, J. Derouard 1, G. Delapierre and R. Jaffiol 1 (1) Laboratoire de Spectrométrie Physique (UMR
the image contrast at the atomic scale. The same method for STM image
 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 Supplementary Note 1. The interpretation of element-specific scanning tunneling microscopy (STM) images Oxygen vacancy lines
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 Supplementary Note 1. The interpretation of element-specific scanning tunneling microscopy (STM) images Oxygen vacancy lines
Cellular imaging using Nano- Materials. A Case-Study based approach Arun Murali, Srivats V
 Cellular imaging using Nano- Materials A Case-Study based approach Arun Murali, Srivats V Agenda Discuss a few papers Explain a couple of new imaging techniques and their benefits over conventional imaging
Cellular imaging using Nano- Materials A Case-Study based approach Arun Murali, Srivats V Agenda Discuss a few papers Explain a couple of new imaging techniques and their benefits over conventional imaging
Sub-micron scale patterning of fluorescent. silver nanoclusters using low-power laser
 Sub-micron scale patterning of fluorescent silver nanoclusters using low-power laser Puskal Kunwar 1,*, Jukka Hassinen 2, Godofredo Bautista 1, Robin H. A. Ras 2, and Juha Toivonen 1 1 Tampere University
Sub-micron scale patterning of fluorescent silver nanoclusters using low-power laser Puskal Kunwar 1,*, Jukka Hassinen 2, Godofredo Bautista 1, Robin H. A. Ras 2, and Juha Toivonen 1 1 Tampere University
Department of Chemistry, Center for Photochemical Sciences, Bowling Green State University,
 Supporting Information for Revealing Abrupt and Spontaneous Ruptures of Protein Native Structure under PicoNewton Compressive Force Manipulation S. Roy Chowdhury, Jin Cao, Yufan He, H. Peter Lu * Department
Supporting Information for Revealing Abrupt and Spontaneous Ruptures of Protein Native Structure under PicoNewton Compressive Force Manipulation S. Roy Chowdhury, Jin Cao, Yufan He, H. Peter Lu * Department
Workshop advanced light microscopy
 Workshop advanced light microscopy Multi-mode confocal laser scanning microscope Jan Willem Borst Laboratory of Biochemistry Biomolecular Networks www.bic.wur.nl MicroSpectroscopy Centre Wageningen Microspectroscopy
Workshop advanced light microscopy Multi-mode confocal laser scanning microscope Jan Willem Borst Laboratory of Biochemistry Biomolecular Networks www.bic.wur.nl MicroSpectroscopy Centre Wageningen Microspectroscopy
In-situ laser-induced contamination monitoring using long-distance microscopy
 In-situ laser-induced contamination monitoring using long-distance microscopy Paul Wagner a, Helmut Schröder* a, Wolfgang Riede a a German Aerospace Center (DLR), Institute of Technical Physics, Pfaffenwaldring
In-situ laser-induced contamination monitoring using long-distance microscopy Paul Wagner a, Helmut Schröder* a, Wolfgang Riede a a German Aerospace Center (DLR), Institute of Technical Physics, Pfaffenwaldring
Principles of flow cytometry: overview of flow cytometry and its uses for cell analysis and sorting. Shoreline Community College BIOL 288
 Principles of flow cytometry: overview of flow cytometry and its uses for cell analysis and sorting Shoreline Community College BIOL 288 Flow Cytometry What is Flow Cytometry? Measurement of cells or particles
Principles of flow cytometry: overview of flow cytometry and its uses for cell analysis and sorting Shoreline Community College BIOL 288 Flow Cytometry What is Flow Cytometry? Measurement of cells or particles
Supporting Information
 Digital Microarrays: Single-Molecule Readout with Interferometric Detection of Plasmonic Nanorod Labels Derin Sevenler 1, George G. Daaboul 2, Fulya Ekiz Kanik 1, Neşe Lortlar Ünlü 3 and M. Selim Ünlü
Digital Microarrays: Single-Molecule Readout with Interferometric Detection of Plasmonic Nanorod Labels Derin Sevenler 1, George G. Daaboul 2, Fulya Ekiz Kanik 1, Neşe Lortlar Ünlü 3 and M. Selim Ünlü
Multi-modality imaging of structure and function combining spectral-domain optical coherence and multiphoton microscopy
 Multi-modality imaging of structure and function combining spectral-domain optical coherence and multiphoton microscopy Claudio Vinegoni a, Tyler Ralston a,b, Wei Tan a, Wei Luo a, Daniel L. Marks a,b,
Multi-modality imaging of structure and function combining spectral-domain optical coherence and multiphoton microscopy Claudio Vinegoni a, Tyler Ralston a,b, Wei Tan a, Wei Luo a, Daniel L. Marks a,b,
Time-resolved Measurements Using the Agilent Cary Eclipse Fluorescence Spectrophotometer A Versatile Instrument for Accurate Measurements
 Time-resolved Measurements Using the Agilent Cary Eclipse Fluorescence Spectrophotometer A Versatile Instrument for Accurate Measurements Technical Overview Authors Dr. Fabian Zieschang, Katherine MacNamara,
Time-resolved Measurements Using the Agilent Cary Eclipse Fluorescence Spectrophotometer A Versatile Instrument for Accurate Measurements Technical Overview Authors Dr. Fabian Zieschang, Katherine MacNamara,
