HCT116 SW48 Nutlin: p53
|
|
|
- Julius Mosley
- 6 years ago
- Views:
Transcription
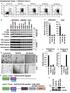
1 Figure S HCT6 SW8 Nutlin: p GAPDH Figure S. Nutlin- treatment induces p protein. HCT6 and SW8 cells were left untreated or treated for 8 hr with Nutlin- ( µm) to up-regulate p. Whole cell lysates were analyzed by immunoblotting. p protein was strongly induced following Nutlin treatment. GAPDH was used as loading control.
2 Figure S Human GCCTCAGTTGCCCCATCTGTGC------CCTGGGTAGGAATATCCTGGATCCCCTTG------GGTCT GATGGGGTAGCCGATGC Chimp GCCTCAGTTGCCCCATCTGTGC------CCTGGGTAGGAACATCCTGGACCCCCTTG------GGTCT GATGGGGTAGCCGATGC Orangutan GCCTCAGTTGCCCCATCTGTGC-nt-CCTGGGTAGGAACATCCTGGATCCCCTTG------GGTCT GATGGGGTAGCTGATGC Gibbon GCCTCAGTTGCCCCATCTGTGC------CCTGGGTAGGAACATCCTGGACCCCCTTG------GGTCT GATGGGGTAGCTGATGC Gorilla GCCTCAGTTGCTCCATCTGTGC------CCTGGGTAGGAACATCCTGGACCCCCTTG------GGTCT GATGGGGTAGCCGATGC Green GCCTCAGTTGCCCTGTCTGTGC------CCTGGGTAGGAACGTCCTGGACCCCCTTG------GGTCT AATGGGGTAGCTGATGC Rhesus GCCTCAGTTGCCCCGTCTGTGC------CCTGGGTAGGAACGTCCTGGACCCCCTTG------GGTCT GATGGGGTAGCTGATGC Crabeat_monk GCCTCAGTTGCCCCGTCTGTGC------CCTGGGTAGGAACGTCCTGGACCCCCTTG------GGTCT GATGGGGTAGCTGATGC aboon GCCTCAGTTGCCCCGTCTGTGC------CCTGGGTAGGAACGTCCTGGACCCCCTTG------GGTCT GATAGGGTAGCTGATGC Marmoset ACCTCAGTTGCTCCGCCTGTGC------CCTGGGTAGGAACATTCTGGACCCCTTTA------GGTCT GAAACCCGATGC Squirrel_monk ACCTCAGTTGCCCC-CCTGTGC------CCTGGGTAGAAACATCCTGGACCCCCTTA------GGTCT GATGGGGTGCCCGATGC Mouse GACTCAGTTTCCCCTGCTGTGA------CCT---CTGGAGCTGCCAGCAGC GGGCTCAGCCAG---AGTAGGACGGATGA--- Rat GCCTCAGTTTCCCCTGCTGTGA------CCT---CTGGAGTTGCCAGCAGTCCCCCGTGCAGTGGGCTCAGCCAG---AGTAGGAAGGACGA--- Dog GCCTCAGTCTCCCCACCTGTGA------ATCGGACAAGAACATCT-----CCCCTTG------GGGCTGACGGTGGGGGGGAGGGCGGCGGGCGG Cat GCCTCAGTTTCCTTATCTGTGA------ACCGGACAAGAACATCC-----CCCCTAG------GGGCTTCGGGGGGTGGGGGGGGGAGGTGGTG- ****** * ***** * ** ** * * Figure S. mir-89 is conserved among apes and old-world monkeys. Multiple alignment of mir-89 candidate sequence between mammalian species. Mammalian orthologous regions for the human mir-89 precursor showed a mosaic pattern of similarity wherein conserved sites are located at the end of hairpin. The rest of the precursor sequence is poorly conserved in mammals, although synteny of genomic alignments in the vicinity of this region is unambiguous. Mature human mir-89-p and -p sequences are shown in red. Positions of % sequence conservation are marked by stars.
3 Figure S Human Rhesus Squirrel Monkey Marmoset Apes OWM NWM NWM Figure S. Pre-miR-89 structure is conserved among apes and old-world monkeys. (A) Phylogenetic tree of mir-89 hairpin regions based on the multiple alignment of primate candidate sequences (apes, old-world monkey, and new-world monkey. () Secondary structure prediction for mir-89 stemloop sequence. Only apes and old-world monkey (OWM), but not new-world monkey (NWM) sequences adopt stable mirna-like structures.
4 Figure S Dox Anti-miR Figure S. mir-89-p knockdown reduces colony formation after Doxorubicin treatment. As shown in Figure, HCT6 cells were reverse transfected with Anti-miR-89-p for 8 hr, then treated with nm Dox for 8 hr and seeded for colony formation on plastic for days.
5 Fold change normalized to GAPDH si sigdf Fold change normalized to U si sigdf Figure S GDF mir-6 mir-89-p C Relative cell growth 6 sigdf Anti-miR-89-p sigdf/anti-mir-89-p mir-89-p mimic/ mir-89-p/sigdf day day day day Figure S. Comparison of mir-89 and GDF knockdowns HCT6 cells were reverse transfected with sirna targeting GDF. (A) 8 hours after transfection, GDF mrna levels were measured by SYR-Green qrt-pcr. GDF was down-regulated by over %7. () mir-89-p levels were quantitated following GDF knockdown by TaqMan qrt-pcr. The unrelated mir-6 is shown as a control. (C) HCT6 cells were reverse transfected with the indicated molecules and seeded in 96-well plates. Relative proliferation was measured over a day period using WST-8 colorimetric assays.
6 si Anti-89 Figure S6 mock dox mock dox CDK CCNA HDAC CDCA GAPDH Fold change normalized to GAPDH..... _untreated Anti-miR-89_untreated _Dox Anti-miR-89_Dox UC p MDM GADDa Figure S6 Effects of mir-89 knockdown on gene expression following DNA damage. A) HCT6-pWT or pko cells were transfected with nm Anti-miR-89 for 8 hours, then treated with nm doxorubicin for 8 hours. Whole cell lysates were probed with the indicated antibodies, GAPDH was used as a loading control. ) HCT6 cells were transfected and treated with doxorubicin as in (A), RNA was isolated and the expression of p pathway genes was measured using qrt-pcr.
7 Figure S7 Fold enrichment normalized to U6 mir-89-p IgG AGO Figure S7. mir-89-p is enriched in RISC after mimic transfection To ensure that the levels of mir-89-p incorporated into RISC after mir-89-p over-expression achieved by transient mimic transfection were physiologically reasonable, we immunoprecipitated Ago 8 hr after transfecting HCT6-pWT cells with mir-89-p mimics. RNA was isolated from the IP material and TaqMan RT-qPCR quantitation of mir-89-p in the RISC IP compared to input showed approximately -fold enrichment.
8 Figure S8 Relative absorbance 6 mir-89-p Day Day Day Day MDA-M- Relative colony formation MDA-M- mir-89-p mir-89-p Figure S8. MiR-89-p over-expression reduces the viability of MDA-M- cells. (A) The mutant p cell line MDA-M-, was transiently transfected with mir-89-p mimic or control and seeded in a 96-well plate. MTT assays were performed over days to measure proliferation. () MDA-M- cells were transfected as above, and re-seeded at cells/well in a -well plate (n=). After days, colonies were fixed with ice-cold methanol, stained with crystal violet, and counted (lower panel).
9 Figure S9.9.8 mir-89 Expression relative to U WI8 MCFa Relative cell growth mir-89 sideath day day day day Relative cell growth 6 mir-89 sideath day day day day Figure S9. mir-89-p is less potent in normal fibroblasts A) RNA isolated from the indicated normal tissues was purchased from Clontech. MiR-89 expression was measured in the tissue panel by TaqMan qrt-pcr and expression values are given as a fraction of U6 expression. () The normal fibroblast cell lines WI8 and MCFA were transfected with mir-89-p or control mimic, and their proliferation over days was measured using MTT colorimetry. The effect of mir- 89-p was much less than that of sideath.
10 KO WT Figure S mimic GAPDH AX PUMA Figure S. mir-89 mimic transfection induces p-associated mediators of apoptosis HCT6-pWT or pko cells were transfected with nm mir-89 mimic for 8 hours. Whole cell lysates were probed with the indicated antibodies, GAPDH was used as a loading control.
11 C Figure S Sub G G-M S G. nm nm nm nm Cell (%) 8 6 Relative cell growth mir-89 mimic sicelldeath Fold chance normalized to GAPDH nm nm nm nm Figure S. Dose-dependence of mir-89 mimic transfection In order to test the effects of different concentrations of mir-89 mimic, we transfected HCT6-pWT cells with,,, or nm mir-89-p mimic or nm mimic. (A) After 8 hours, cells were fixed and stained with PI, and cell cycle was analyzed by flow cytometry. () RNA was harvested 8 hours after transfection, and the expression of mir-89 targets was measured by SYR-Green qrt-pcr. SDHA is shown as a negative control. (C) Cells were reverse transfected and seeded in 96 well plates, and relative growth after a further 7 hours was measured by WST-8 assay.
ASPP1 Fw GGTTGGGAATCCACGTGTTG ASPP1 Rv GCCATATCTTGGAGCTCTGAGAG
 Supplemental Materials and Methods Plasmids: the following plasmids were used in the supplementary data: pwzl-myc- Lats2 (Aylon et al, 2006), pretrosuper-vector and pretrosuper-shp53 (generous gift of
Supplemental Materials and Methods Plasmids: the following plasmids were used in the supplementary data: pwzl-myc- Lats2 (Aylon et al, 2006), pretrosuper-vector and pretrosuper-shp53 (generous gift of
Cell proliferation was measured with Cell Counting Kit-8 (Dojindo Laboratories, Kumamoto, Japan).
 1 2 3 4 5 6 7 8 Supplemental Materials and Methods Cell proliferation assay Cell proliferation was measured with Cell Counting Kit-8 (Dojindo Laboratories, Kumamoto, Japan). GCs were plated at 96-well
1 2 3 4 5 6 7 8 Supplemental Materials and Methods Cell proliferation assay Cell proliferation was measured with Cell Counting Kit-8 (Dojindo Laboratories, Kumamoto, Japan). GCs were plated at 96-well
SUPPLEMENTAL MATERIAL. Carvajal et al. Supplemental Table 1. List of quantitative PCR primers used for cdna analyses and
 UPPLEMEAL MATERIAL Carvajal et al. upplemental Table 1. List of quantitative PCR primers used for cdna analyses and chromatin immunoprecipitation assays. Figure 1. DNA damage-induced transcriptional repression
UPPLEMEAL MATERIAL Carvajal et al. upplemental Table 1. List of quantitative PCR primers used for cdna analyses and chromatin immunoprecipitation assays. Figure 1. DNA damage-induced transcriptional repression
SUPPLEMENTARY INFORMATION
 doi:.38/nature899 Supplementary Figure Suzuki et al. a c p7 -/- / WT ratio (+)/(-) p7 -/- / WT ratio Log X 3. Fold change by treatment ( (+)/(-)) Log X.5 3-3. -. b Fold change by treatment ( (+)/(-)) 8
doi:.38/nature899 Supplementary Figure Suzuki et al. a c p7 -/- / WT ratio (+)/(-) p7 -/- / WT ratio Log X 3. Fold change by treatment ( (+)/(-)) Log X.5 3-3. -. b Fold change by treatment ( (+)/(-)) 8
Supplementary Fig. 1 Proteomic analysis of ATR-interacting proteins. ATR, ARID1A and
 Supplementary Figure Legend: Supplementary Fig. 1 Proteomic analysis of ATR-interacting proteins. ATR, ARID1A and ATRIP protein peptides identified from our mass spectrum analysis were shown. Supplementary
Supplementary Figure Legend: Supplementary Fig. 1 Proteomic analysis of ATR-interacting proteins. ATR, ARID1A and ATRIP protein peptides identified from our mass spectrum analysis were shown. Supplementary
T H E J O U R N A L O F C E L L B I O L O G Y
 T H E J O U R N A L O F C E L L B I O L O G Y Supplemental material Han et al., http://www.jcb.org/cgi/content/full/jcb.201311007/dc1 Figure S1. SIVA1 interacts with PCNA. (A) HEK293T cells were transiently
T H E J O U R N A L O F C E L L B I O L O G Y Supplemental material Han et al., http://www.jcb.org/cgi/content/full/jcb.201311007/dc1 Figure S1. SIVA1 interacts with PCNA. (A) HEK293T cells were transiently
Khaled_Fig. S MITF TYROSINASE R²= MITF PDE4D R²= Variance from mean mrna expression
 Khaled_Fig. S Variance from mean mrna expression.8 2 MITF.6 TYROSINASE.4 R²=.73.2.8.6.4.2.8.6.4.2.8.6.4.2 MALME3M SKMEL28 UACC257 MITF PDE4D R²=.63 4.5 5 MITF 3.5 4 PDE4B 2.5 3 R²=.2.5 2.5.8.6.4.2.8.6.4.2
Khaled_Fig. S Variance from mean mrna expression.8 2 MITF.6 TYROSINASE.4 R²=.73.2.8.6.4.2.8.6.4.2.8.6.4.2 MALME3M SKMEL28 UACC257 MITF PDE4D R²=.63 4.5 5 MITF 3.5 4 PDE4B 2.5 3 R²=.2.5 2.5.8.6.4.2.8.6.4.2
Supplementary Figure 1. GST pull-down analysis of the interaction of GST-cIAP1 (A, B), GSTcIAP1
 Legends Supplementary Figure 1. GST pull-down analysis of the interaction of GST- (A, B), GST mutants (B) or GST- (C) with indicated proteins. A, B, Cell lysate from untransfected HeLa cells were loaded
Legends Supplementary Figure 1. GST pull-down analysis of the interaction of GST- (A, B), GST mutants (B) or GST- (C) with indicated proteins. A, B, Cell lysate from untransfected HeLa cells were loaded
Supplementary Table 1. Sequences for BTG2 and BRCA1 sirnas.
 Supplementary Table 1. Sequences for BTG2 and BRCA1 sirnas. Target Gene Non-target / Control BTG2 BRCA1 NFE2L2 Target Sequence ON-TARGET plus Non-targeting sirna # 1 (Cat# D-001810-01-05) sirna1: GAACCGACAUGCUCCCGGA
Supplementary Table 1. Sequences for BTG2 and BRCA1 sirnas. Target Gene Non-target / Control BTG2 BRCA1 NFE2L2 Target Sequence ON-TARGET plus Non-targeting sirna # 1 (Cat# D-001810-01-05) sirna1: GAACCGACAUGCUCCCGGA
Supplementary figures
 Relative intensity Relative intensity Relative intensity Supplementary figures a None Caffeine None Caffeine c None Caffeine 6 6 6 ISG 6 6 6 UBE1L 6 6 6 UBCH8 6 6 6 EFP 1 1 DOX (h) 1 1 CPT (h) 1 1 UV (h)
Relative intensity Relative intensity Relative intensity Supplementary figures a None Caffeine None Caffeine c None Caffeine 6 6 6 ISG 6 6 6 UBE1L 6 6 6 UBCH8 6 6 6 EFP 1 1 DOX (h) 1 1 CPT (h) 1 1 UV (h)
Emanuela Tumini, Sonia Barroso, Carmen Pérez Calero and Andrés Aguilera
 SUPPLEMENTARY INFORMATION Roles of human POLD1 and POLD3 in genome stability Emanuela Tumini, Sonia Barroso, Carmen Pérez Calero and Andrés Aguilera SUPPLEMENTARY METHODS Cell proliferation After sirna
SUPPLEMENTARY INFORMATION Roles of human POLD1 and POLD3 in genome stability Emanuela Tumini, Sonia Barroso, Carmen Pérez Calero and Andrés Aguilera SUPPLEMENTARY METHODS Cell proliferation After sirna
Supplementary Figure 1. Expressions of stem cell markers decreased in TRCs on 2D plastic. TRCs were cultured on plastic for 1, 3, 5, or 7 days,
 Supplementary Figure 1. Expressions of stem cell markers decreased in TRCs on 2D plastic. TRCs were cultured on plastic for 1, 3, 5, or 7 days, respectively, and their mrnas were quantified by real time
Supplementary Figure 1. Expressions of stem cell markers decreased in TRCs on 2D plastic. TRCs were cultured on plastic for 1, 3, 5, or 7 days, respectively, and their mrnas were quantified by real time
Supplementary Fig S1 Nutlin-3a treatment does not affect cell cycle progression in the absence
 Supplementary Figure Legends Supplementary Fig S1 Nutlin-3a treatment does not affect cell cycle progression in the absence of p53 or p21. HCT116 cells which were null for either p53 (A) or p21 (B) were
Supplementary Figure Legends Supplementary Fig S1 Nutlin-3a treatment does not affect cell cycle progression in the absence of p53 or p21. HCT116 cells which were null for either p53 (A) or p21 (B) were
Coleman et al., Supplementary Figure 1
 Coleman et al., Supplementary Figure 1 BrdU Merge G1 Early S Mid S Supplementary Figure 1. Sequential destruction of CRL4 Cdt2 targets during the G1/S transition. HCT116 cells were synchronized by sequential
Coleman et al., Supplementary Figure 1 BrdU Merge G1 Early S Mid S Supplementary Figure 1. Sequential destruction of CRL4 Cdt2 targets during the G1/S transition. HCT116 cells were synchronized by sequential
Supplementary Figure 1. TSA (10 nmol/l), non-class-selective HDAC inhibitor, potentiates
 Supplementary Figure 1. TSA (10 nmol/l), non-class-selective HDAC inhibitor, potentiates vascular calcification (VC). (a) Von Kossa staining shows that TSA potentiated the Pi-induced VC. Scale bar, 100
Supplementary Figure 1. TSA (10 nmol/l), non-class-selective HDAC inhibitor, potentiates vascular calcification (VC). (a) Von Kossa staining shows that TSA potentiated the Pi-induced VC. Scale bar, 100
Supplementary Information. ATM and MET kinases are synthetic lethal with. non-genotoxic activation of p53
 Supplementary Information ATM and MET kinases are synthetic lethal with non-genotoxic activation of p53 Kelly D. Sullivan 1, Nuria Padilla-Just 1, Ryan E. Henry 1, Christopher C. Porter 2, Jihye Kim 3,
Supplementary Information ATM and MET kinases are synthetic lethal with non-genotoxic activation of p53 Kelly D. Sullivan 1, Nuria Padilla-Just 1, Ryan E. Henry 1, Christopher C. Porter 2, Jihye Kim 3,
SureSilencing sirna Array Technology Overview
 SureSilencing sirna Array Technology Overview Pathway-Focused sirna-based RNA Interference Topics to be Covered Who is SuperArray? Brief Introduction to RNA Interference Challenges Facing RNA Interference
SureSilencing sirna Array Technology Overview Pathway-Focused sirna-based RNA Interference Topics to be Covered Who is SuperArray? Brief Introduction to RNA Interference Challenges Facing RNA Interference
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb3240 Supplementary Figure 1 GBM cell lines display similar levels of p100 to p52 processing but respond differentially to TWEAK-induced TERT expression according to TERT promoter mutation
DOI: 10.1038/ncb3240 Supplementary Figure 1 GBM cell lines display similar levels of p100 to p52 processing but respond differentially to TWEAK-induced TERT expression according to TERT promoter mutation
(a) Immunoblotting to show the migration position of Flag-tagged MAVS
 Supplementary Figure 1 Characterization of six MAVS isoforms. (a) Immunoblotting to show the migration position of Flag-tagged MAVS isoforms. HEK293T Mavs -/- cells were transfected with constructs expressing
Supplementary Figure 1 Characterization of six MAVS isoforms. (a) Immunoblotting to show the migration position of Flag-tagged MAVS isoforms. HEK293T Mavs -/- cells were transfected with constructs expressing
Supplementary Table 1. Primers used to construct full-length or various truncated mutants of ISG12b2.
 Supplementary Table 1. Primers used to construct full-length or various truncated mutants of ISG12b2. Construct name ISG12b2 (No tag) HA-ISG12b2 (N-HA) ISG12b2-HA (C-HA; FL-HA) 94-283-HA (FL-GFP) 93-GFP
Supplementary Table 1. Primers used to construct full-length or various truncated mutants of ISG12b2. Construct name ISG12b2 (No tag) HA-ISG12b2 (N-HA) ISG12b2-HA (C-HA; FL-HA) 94-283-HA (FL-GFP) 93-GFP
1. Primers for PCR to amplify hairpin stem-loop precursor mir-145 plus different flanking sequence from human genomic DNA.
 Supplemental data: 1. Primers for PCR to amplify hairpin stem-loop precursor mir-145 plus different flanking sequence from human genomic DNA. Strategy#1: 20nt at both sides: #1_BglII-Fd primer : 5 -gga
Supplemental data: 1. Primers for PCR to amplify hairpin stem-loop precursor mir-145 plus different flanking sequence from human genomic DNA. Strategy#1: 20nt at both sides: #1_BglII-Fd primer : 5 -gga
OmicsLink shrna Clones guaranteed knockdown even in difficult-to-transfect cells
 OmicsLink shrna Clones guaranteed knockdown even in difficult-to-transfect cells OmicsLink shrna clone collections consist of lentiviral, and other mammalian expression vector based small hairpin RNA (shrna)
OmicsLink shrna Clones guaranteed knockdown even in difficult-to-transfect cells OmicsLink shrna clone collections consist of lentiviral, and other mammalian expression vector based small hairpin RNA (shrna)
Supplemental Figure Legends:
 Supplemental Figure Legends: Fig S1. GFP-ABRO1 localization. U2OS cells were infected with retrovirus expressing GFP- ABRO1. The cells were fixed with 3.6% formaldehyde and stained with antibodies against
Supplemental Figure Legends: Fig S1. GFP-ABRO1 localization. U2OS cells were infected with retrovirus expressing GFP- ABRO1. The cells were fixed with 3.6% formaldehyde and stained with antibodies against
transcription and the promoter occupancy of Smad proteins. (A) HepG2 cells were co-transfected with the wwp-luc reporter, and FLAG-tagged FHL1,
 Supplementary Data Supplementary Figure Legends Supplementary Figure 1 FHL-mediated TGFβ-responsive reporter transcription and the promoter occupancy of Smad proteins. (A) HepG2 cells were co-transfected
Supplementary Data Supplementary Figure Legends Supplementary Figure 1 FHL-mediated TGFβ-responsive reporter transcription and the promoter occupancy of Smad proteins. (A) HepG2 cells were co-transfected
IKK is a therapeutic target in KRAS-induced lung cancer with disrupted p53 activity
 IKK is a therapeutic target in KRAS-induced lung cancer with disrupted p5 activity H6 5 5 H58 A59 H6 H58 A59 anti-ikkα anti-ikkβ anti-panras anti-gapdh anti-ikkα anti-ikkβ anti-panras anti-gapdh anti-ikkα
IKK is a therapeutic target in KRAS-induced lung cancer with disrupted p5 activity H6 5 5 H58 A59 H6 H58 A59 anti-ikkα anti-ikkβ anti-panras anti-gapdh anti-ikkα anti-ikkβ anti-panras anti-gapdh anti-ikkα
Nature Biotechnology: doi: /nbt Supplementary Figure 1
 Supplementary Figure 1 Schematic and results of screening the combinatorial antibody library for Sox2 replacement activity. A single batch of MEFs were plated and transduced with doxycycline inducible
Supplementary Figure 1 Schematic and results of screening the combinatorial antibody library for Sox2 replacement activity. A single batch of MEFs were plated and transduced with doxycycline inducible
Supplementary Figure 1.
 Supplementary Figure 1. Quantification of western blot analysis of fibroblasts (related to Figure 1) (A-F) Quantification of western blot analysis for control and IR-Mut fibroblasts. Data are expressed
Supplementary Figure 1. Quantification of western blot analysis of fibroblasts (related to Figure 1) (A-F) Quantification of western blot analysis for control and IR-Mut fibroblasts. Data are expressed
Regulation of transcription by the MLL2 complex and MLL complex-associated AKAP95
 Supplementary Information Regulation of transcription by the complex and MLL complex-associated Hao Jiang, Xiangdong Lu, Miho Shimada, Yali Dou, Zhanyun Tang, and Robert G. Roeder Input HeLa NE IP lot:
Supplementary Information Regulation of transcription by the complex and MLL complex-associated Hao Jiang, Xiangdong Lu, Miho Shimada, Yali Dou, Zhanyun Tang, and Robert G. Roeder Input HeLa NE IP lot:
IGF-1 promotes the development and cytotoxic activity of human NK cells regulated
 Supplementary Information for IGF-1 promotes the development and cytotoxic activity of human NK cells regulated by mir-483-3p Fang Ni 1, 2, Rui Sun 1, Binqing Fu 1, Fuyan Wang 1, Chuang Guo 1, Zhigang
Supplementary Information for IGF-1 promotes the development and cytotoxic activity of human NK cells regulated by mir-483-3p Fang Ni 1, 2, Rui Sun 1, Binqing Fu 1, Fuyan Wang 1, Chuang Guo 1, Zhigang
supplementary information
 Figure S1 ZEB1 full length mrna. (a) Analysis of the ZEB1 mrna using the UCSC genome browser (http://genome.ucsc.edu) revealed truncation of the annotated Refseq sequence (NM_030751). The probable terminus
Figure S1 ZEB1 full length mrna. (a) Analysis of the ZEB1 mrna using the UCSC genome browser (http://genome.ucsc.edu) revealed truncation of the annotated Refseq sequence (NM_030751). The probable terminus
TE5 KYSE510 TE7 KYSE70 KYSE140
 TE5 KYSE5 TT KYSE7 KYSE4 Supplementary Figure. Hockey stick plots showing input normalized, rank ordered H3K7ac signals for the candidate SE-associated lncrnas in this study. Rpm Rpm Rpm Chip-seq H3K7ac
TE5 KYSE5 TT KYSE7 KYSE4 Supplementary Figure. Hockey stick plots showing input normalized, rank ordered H3K7ac signals for the candidate SE-associated lncrnas in this study. Rpm Rpm Rpm Chip-seq H3K7ac
Document S1. Supplemental Experimental Procedures and Three Figures (see next page)
 Supplemental Data Document S1. Supplemental Experimental Procedures and Three Figures (see next page) Table S1. List of Candidate Genes Identified from the Screen. Candidate genes, corresponding dsrnas
Supplemental Data Document S1. Supplemental Experimental Procedures and Three Figures (see next page) Table S1. List of Candidate Genes Identified from the Screen. Candidate genes, corresponding dsrnas
Supplementary Information. A novel human endogenous retroviral protein inhibits cell-cell fusion. Supplementary Figures:
 Supplementary Information A novel human endogenous retroviral protein inhibits cell-cell fusion Jun Sugimoto, Makiko Sugimoto, Helene Bernstein, Yoshihiro Jinno and Danny J. Schust Supplementary Figures:
Supplementary Information A novel human endogenous retroviral protein inhibits cell-cell fusion Jun Sugimoto, Makiko Sugimoto, Helene Bernstein, Yoshihiro Jinno and Danny J. Schust Supplementary Figures:
Supplementary data 1. Prmers and probes used in Taqman real-time PCR.
 Supplementary data 1. Prmers and probes used in Taqman real-time PCR. PIG-T Forward Primer: gatctgcctcacgtgcactgt Reverse Primer: aggttcgggtgaggagattgt Probe: 6FAMTGG CCG TGT GCT ATG GCT CCT TCTAMRA PIG-U
Supplementary data 1. Prmers and probes used in Taqman real-time PCR. PIG-T Forward Primer: gatctgcctcacgtgcactgt Reverse Primer: aggttcgggtgaggagattgt Probe: 6FAMTGG CCG TGT GCT ATG GCT CCT TCTAMRA PIG-U
Hairpin-it TM mirnas qpcr Quantitation Kit
 Hairpin-it TM mirnas qpcr Quantitation Kit For the detection and quantification of micrornas using real-time PCR detection instruments. Catalog No. QPM-010/ QPM-011/ QPM-012/ QPM-013 User Manual Table
Hairpin-it TM mirnas qpcr Quantitation Kit For the detection and quantification of micrornas using real-time PCR detection instruments. Catalog No. QPM-010/ QPM-011/ QPM-012/ QPM-013 User Manual Table
Total RNA was isolated using Trizol reagent (Invitrogen) and reverse transcribed using
 Supplementary Methods RNA Isolation and Quantitative RT-PCR Total RNA was isolated using Trizol reagent (Invitrogen) and reverse transcribed using random hexamers and superscript II reverse transcriptase
Supplementary Methods RNA Isolation and Quantitative RT-PCR Total RNA was isolated using Trizol reagent (Invitrogen) and reverse transcribed using random hexamers and superscript II reverse transcriptase
Supplementary Figure 1. NORAD expression in mouse (A) and dog (B). The black boxes indicate the position of the regions alignable to the 12 repeat
 Supplementary Figure 1. NORAD expression in mouse (A) and dog (B). The black boxes indicate the position of the regions alignable to the 12 repeat units in the human genome. Annotated transposable elements
Supplementary Figure 1. NORAD expression in mouse (A) and dog (B). The black boxes indicate the position of the regions alignable to the 12 repeat units in the human genome. Annotated transposable elements
MicroRNA Analysis Paired with Novel Cell Health Assays: A Complete Workflow
 MicroRNA Analysis Paired with Novel Cell Health Assays: A Complete Workflow Brad Hook, Ph.D Manager, NA Scientific Applications 2016 Presentation Outline From cells to RNA a seemingly easy, yet complex
MicroRNA Analysis Paired with Novel Cell Health Assays: A Complete Workflow Brad Hook, Ph.D Manager, NA Scientific Applications 2016 Presentation Outline From cells to RNA a seemingly easy, yet complex
MicroRNA Destabilization Enables. Dynamic Regulation of the mir-16 Family. In Response to Cell-Cycle Changes SUPPLEMENTAL INFORMATION
 Molecular Cell, Volume 43 SUPPLEMENTAL INFORMATION MicroRNA Destabilization Enables Dynamic Regulation of the mir-16 Family In Response to Cell-Cycle Changes Olivia S. Risland, Sue-Jean Hong, and David
Molecular Cell, Volume 43 SUPPLEMENTAL INFORMATION MicroRNA Destabilization Enables Dynamic Regulation of the mir-16 Family In Response to Cell-Cycle Changes Olivia S. Risland, Sue-Jean Hong, and David
Supplementary Figure 1, related to Figure 1. GAS5 is highly expressed in the cytoplasm of hescs, and positively correlates with pluripotency.
 Supplementary Figure 1, related to Figure 1. GAS5 is highly expressed in the cytoplasm of hescs, and positively correlates with pluripotency. (a) Transfection of different concentration of GAS5-overexpressing
Supplementary Figure 1, related to Figure 1. GAS5 is highly expressed in the cytoplasm of hescs, and positively correlates with pluripotency. (a) Transfection of different concentration of GAS5-overexpressing
supplementary information
 DOI: 10.1038/ncb2116 Figure S1 CDK phosphorylation of EZH2 in cells. (a) Comparison of candidate CDK phosphorylation sites on EZH2 with known CDK substrates by multiple sequence alignments. (b) CDK1 and
DOI: 10.1038/ncb2116 Figure S1 CDK phosphorylation of EZH2 in cells. (a) Comparison of candidate CDK phosphorylation sites on EZH2 with known CDK substrates by multiple sequence alignments. (b) CDK1 and
Supplementary Figures
 Supplementary Figures Supplementary Figure 1 Experimental schema for the identification of circular RNAs in six normal tissues and seven cancerous tissues. Supplementary Fiure 2 Comparison of human circrnas
Supplementary Figures Supplementary Figure 1 Experimental schema for the identification of circular RNAs in six normal tissues and seven cancerous tissues. Supplementary Fiure 2 Comparison of human circrnas
Supporting Online Material for
 www.sciencemag.org/cgi/content/full/1154040/dc1 Supporting Online Material for Selective Blockade of MicroRNA Processing by Lin-28 Srinivas R. Viswanathan, George Q. Daley,* Richard I. Gregory* *To whom
www.sciencemag.org/cgi/content/full/1154040/dc1 Supporting Online Material for Selective Blockade of MicroRNA Processing by Lin-28 Srinivas R. Viswanathan, George Q. Daley,* Richard I. Gregory* *To whom
SUPPLEMENTAL MATERIALS SIRTUIN 1 PROMOTES HYPEROXIA-INDUCED LUNG EPITHELIAL DEATH INDEPENDENT OF NRF2 ACTIVATION
 SUPPLEMENTAL MATERIALS SIRTUIN PROMOTES HYPEROXIA-INDUCED LUNG EPITHELIAL DEATH INDEPENDENT OF NRF ACTIVATION Haranatha R. Potteti*, Subbiah Rajasekaran*, Senthilkumar B. Rajamohan*, Chandramohan R. Tamatam,
SUPPLEMENTAL MATERIALS SIRTUIN PROMOTES HYPEROXIA-INDUCED LUNG EPITHELIAL DEATH INDEPENDENT OF NRF ACTIVATION Haranatha R. Potteti*, Subbiah Rajasekaran*, Senthilkumar B. Rajamohan*, Chandramohan R. Tamatam,
Fig. S1 TGF RI inhibitor SB effectively blocks phosphorylation of Smad2 induced by TGF. FET cells were treated with TGF in the presence of
 Fig. S1 TGF RI inhibitor SB525334 effectively blocks phosphorylation of Smad2 induced by TGF. FET cells were treated with TGF in the presence of different concentrations of SB525334. Cells were lysed and
Fig. S1 TGF RI inhibitor SB525334 effectively blocks phosphorylation of Smad2 induced by TGF. FET cells were treated with TGF in the presence of different concentrations of SB525334. Cells were lysed and
Supplementary information for: Mutant p53 gain-of-function induces epithelial-mesenchymal transition. through modulation of the mir-130b-zeb1 axis
 Supplementary information for: Mutant p53 gain-of-function induces epithelial-mesenchymal transition through modulation of the mir-3b-zeb axis AUTHORS: Peixin Dong, Mihriban Karaayvaz, Nan Jia, Masanori
Supplementary information for: Mutant p53 gain-of-function induces epithelial-mesenchymal transition through modulation of the mir-3b-zeb axis AUTHORS: Peixin Dong, Mihriban Karaayvaz, Nan Jia, Masanori
TOOLS sirna and mirna. User guide
 TOOLS sirna and mirna User guide Introduction RNA interference (RNAi) is a powerful tool for suppression gene expression by causing the destruction of specific mrna molecules. Small Interfering RNAs (sirnas)
TOOLS sirna and mirna User guide Introduction RNA interference (RNAi) is a powerful tool for suppression gene expression by causing the destruction of specific mrna molecules. Small Interfering RNAs (sirnas)
Sarker et al. Supplementary Material. Subcellular Fractionation
 Supplementary Material Subcellular Fractionation Transfected 293T cells were harvested with phosphate buffered saline (PBS) and centrifuged at 2000 rpm (500g) for 3 min. The pellet was washed, re-centrifuged
Supplementary Material Subcellular Fractionation Transfected 293T cells were harvested with phosphate buffered saline (PBS) and centrifuged at 2000 rpm (500g) for 3 min. The pellet was washed, re-centrifuged
sirna Transfection Into Primary Neurons Using Fuse-It-siRNA
 sirna Transfection Into Primary Neurons Using Fuse-It-siRNA This Application Note describes a protocol for sirna transfection into sensitive, primary cortical neurons using Fuse-It-siRNA. This innovative
sirna Transfection Into Primary Neurons Using Fuse-It-siRNA This Application Note describes a protocol for sirna transfection into sensitive, primary cortical neurons using Fuse-It-siRNA. This innovative
Supplemental Materials and Methods
 Supplemental Materials and Methods Antibodies: Anti-SRF (cat# Sc-335) and anti-igf1r (sc-712) (Santa Cruz Biotech), and anti- ADAM-10 (14-6211) were from e-bioscience, anti-ku70 (cat# MS-329-P) (Labvision),
Supplemental Materials and Methods Antibodies: Anti-SRF (cat# Sc-335) and anti-igf1r (sc-712) (Santa Cruz Biotech), and anti- ADAM-10 (14-6211) were from e-bioscience, anti-ku70 (cat# MS-329-P) (Labvision),
Comparative Analysis of Argonaute-Dependent Small RNA Pathways in Drosophila
 Molecular Cell, Volume 32 Supplemental Data Comparative Analysis of Argonaute-Dependent Small RNA Pathways in Drosophila Rui Zhou, Ikuko Hotta, Ahmet M. Denli, Pengyu Hong, Norbert Perrimon, and Gregory
Molecular Cell, Volume 32 Supplemental Data Comparative Analysis of Argonaute-Dependent Small RNA Pathways in Drosophila Rui Zhou, Ikuko Hotta, Ahmet M. Denli, Pengyu Hong, Norbert Perrimon, and Gregory
MeCP2. MeCP2/α-tubulin. GFP mir1-1 mir132
 Conservation Figure S1. Schematic showing 3 UTR (top; thick black line), mir132 MRE (arrow) and nucleotide sequence conservation (vertical black lines; http://genome.ucsc.edu). a GFP mir1-1 mir132 b GFP
Conservation Figure S1. Schematic showing 3 UTR (top; thick black line), mir132 MRE (arrow) and nucleotide sequence conservation (vertical black lines; http://genome.ucsc.edu). a GFP mir1-1 mir132 b GFP
Supplementary information. Supplementary Figures
 Supplementary information Supplementary Figures Supplementary Figure 1. A. i. HA-JMY expressing U2OS cells were treated with SAHA (6h). DAPI was used to visualise nuclei. ii. U2OS cells stably expressing
Supplementary information Supplementary Figures Supplementary Figure 1. A. i. HA-JMY expressing U2OS cells were treated with SAHA (6h). DAPI was used to visualise nuclei. ii. U2OS cells stably expressing
mir-24-mediated down-regulation of H2AX suppresses DNA repair
 Supplemental Online Material mir-24-mediated down-regulation of H2AX suppresses DNA repair in terminally differentiated blood cells Ashish Lal 1,4, Yunfeng Pan 2,4, Francisco Navarro 1,4, Derek M. Dykxhoorn
Supplemental Online Material mir-24-mediated down-regulation of H2AX suppresses DNA repair in terminally differentiated blood cells Ashish Lal 1,4, Yunfeng Pan 2,4, Francisco Navarro 1,4, Derek M. Dykxhoorn
Electromagnetic Fields Mediate Efficient Cell Reprogramming Into a Pluripotent State
 Electromagnetic Fields Mediate Efficient Cell Reprogramming Into a Pluripotent State Soonbong Baek, 1 Xiaoyuan Quan, 1 Soochan Kim, 2 Christopher Lengner, 3 Jung- Keug Park, 4 and Jongpil Kim 1* 1 Lab
Electromagnetic Fields Mediate Efficient Cell Reprogramming Into a Pluripotent State Soonbong Baek, 1 Xiaoyuan Quan, 1 Soochan Kim, 2 Christopher Lengner, 3 Jung- Keug Park, 4 and Jongpil Kim 1* 1 Lab
Gene Forward Primer Reverse Primer GAPDH ATCATCCCTGCCTCTACTGG GTCAGGTCCACCACTGACAC SSB1 AACTTCAGTGAGCCAAACCC GTTCTCAGAGGCTGGAGAGG
 Supplemental Data EXPERIMENTAL PROCEDURES Plasmids and Antibodies- Full length cdna of INT11 or INT12 were cloned into ps- Flag-SBP vector respectively. Anti-RNA pol II (RPB1) was purchased from Santa
Supplemental Data EXPERIMENTAL PROCEDURES Plasmids and Antibodies- Full length cdna of INT11 or INT12 were cloned into ps- Flag-SBP vector respectively. Anti-RNA pol II (RPB1) was purchased from Santa
Supplementary Fig. 1. (A) Working model. The pluripotency transcription factor OCT4
 SUPPLEMENTARY FIGURE LEGENDS Supplementary Fig. 1. (A) Working model. The pluripotency transcription factor OCT4 directly up-regulates the expression of NIPP1 and CCNF that together inhibit protein phosphatase
SUPPLEMENTARY FIGURE LEGENDS Supplementary Fig. 1. (A) Working model. The pluripotency transcription factor OCT4 directly up-regulates the expression of NIPP1 and CCNF that together inhibit protein phosphatase
Transcriptional Regulation (Gene Regulation)
 Experimental Techniques in Biomedical Sciences 의생명과학실험기법 Transcriptional Regulation (Gene Regulation) 4/17/13 Jeong Hoon Kim (jeongkim@skku.edu) Department of Health Sciences and Technology, SKKU Graduate
Experimental Techniques in Biomedical Sciences 의생명과학실험기법 Transcriptional Regulation (Gene Regulation) 4/17/13 Jeong Hoon Kim (jeongkim@skku.edu) Department of Health Sciences and Technology, SKKU Graduate
Galina Gabriely, Ph.D. BWH/HMS
 Galina Gabriely, Ph.D. BWH/HMS Email: ggabriely@rics.bwh.harvard.edu Outline: microrna overview microrna expression analysis microrna functional analysis microrna (mirna) Characteristics mirnas discovered
Galina Gabriely, Ph.D. BWH/HMS Email: ggabriely@rics.bwh.harvard.edu Outline: microrna overview microrna expression analysis microrna functional analysis microrna (mirna) Characteristics mirnas discovered
Table 1. Primers, annealing temperatures, and product sizes for PCR amplification.
 Table 1. Primers, annealing temperatures, and product sizes for PCR amplification. Gene Direction Primer sequence (5 3 ) Annealing Temperature Size (bp) BRCA1 Forward TTGCGGGAGGAAAATGGGTAGTTA 50 o C 292
Table 1. Primers, annealing temperatures, and product sizes for PCR amplification. Gene Direction Primer sequence (5 3 ) Annealing Temperature Size (bp) BRCA1 Forward TTGCGGGAGGAAAATGGGTAGTTA 50 o C 292
mmu-mir-200a-3p Real-time RT-PCR Detection and U6 Calibration Kit User Manual MyBioSource.com Catalog # MBS826230
 mmu-mir-200a-3p Real-time RT-PCR Detection and U6 Calibration Kit User Manual Catalog # MBS826230 For the detection and quantification of mirnas mmu-mir-200a-3p normalized by U6 snrna using Real-time RT-PCR
mmu-mir-200a-3p Real-time RT-PCR Detection and U6 Calibration Kit User Manual Catalog # MBS826230 For the detection and quantification of mirnas mmu-mir-200a-3p normalized by U6 snrna using Real-time RT-PCR
Supplementary Figure 1. (a) The qrt-pcr for lnc-2, lnc-6 and lnc-7 RNA level in DU145, 22Rv1, wild type HCT116 and HCT116 Dicer ex5 cells transfected
 Supplementary Figure 1. (a) The qrt-pcr for lnc-2, lnc-6 and lnc-7 RNA level in DU145, 22Rv1, wild type HCT116 and HCT116 Dicer ex5 cells transfected with the sirna against lnc-2, lnc-6, lnc-7, and the
Supplementary Figure 1. (a) The qrt-pcr for lnc-2, lnc-6 and lnc-7 RNA level in DU145, 22Rv1, wild type HCT116 and HCT116 Dicer ex5 cells transfected with the sirna against lnc-2, lnc-6, lnc-7, and the
To generate the luciferase fusion to the human 3 UTRs, we sub-cloned the 3 UTR
 Plasmids To generate the luciferase fusion to the human 3 UTRs, we sub-cloned the 3 UTR fragments downstream of firefly luciferase (luc) in pgl3 control (Promega). pgl3- CDK6 was made by amplifying a 2,886
Plasmids To generate the luciferase fusion to the human 3 UTRs, we sub-cloned the 3 UTR fragments downstream of firefly luciferase (luc) in pgl3 control (Promega). pgl3- CDK6 was made by amplifying a 2,886
Transient silencing of the CDKN2A gene was achieved using a pool of four sirna duplexes
 Supplementary Methods Small interfering RNA (sirna) knockdown of CDKN2A (p16 INK4a /p14 ARF ) Transient silencing of the CDKN2A gene was achieved using a pool of four sirna duplexes that each targeted
Supplementary Methods Small interfering RNA (sirna) knockdown of CDKN2A (p16 INK4a /p14 ARF ) Transient silencing of the CDKN2A gene was achieved using a pool of four sirna duplexes that each targeted
SANTA CRUZ BIOTECHNOLOGY, INC.
 TECHNICAL SERVICE GUIDE: Western Blotting 2. What size bands were expected and what size bands were detected? 3. Was the blot blank or was a dark background or non-specific bands seen? 4. Did this same
TECHNICAL SERVICE GUIDE: Western Blotting 2. What size bands were expected and what size bands were detected? 3. Was the blot blank or was a dark background or non-specific bands seen? 4. Did this same
Selective Inhibitors and Preliminary Biology Insight.
 Supporting Information Towards the Validation of Maternal Embryonic Leucine Zipper Kinase: Discovery, Optimization of Highly Potent and Selective Inhibitors and Preliminary Biology Insight. B. Barry Touré,*
Supporting Information Towards the Validation of Maternal Embryonic Leucine Zipper Kinase: Discovery, Optimization of Highly Potent and Selective Inhibitors and Preliminary Biology Insight. B. Barry Touré,*
Xfect Protein Transfection Reagent
 Xfect Protein Transfection Reagent Mammalian Expression Systems Rapid, high-efficiency, low-toxicity protein transfection Transfect a large amount of active protein Virtually no cytotoxicity, unlike lipofection
Xfect Protein Transfection Reagent Mammalian Expression Systems Rapid, high-efficiency, low-toxicity protein transfection Transfect a large amount of active protein Virtually no cytotoxicity, unlike lipofection
Supplemental Information. DNp63 Inhibits Oxidative Stress-Induced Cell. Death, Including Ferroptosis, and Cooperates with
 Cell Reports, Volume 21 Supplemental Information DNp63 Inhibits Oxidative Stress-Induced Cell Death, Including Ferroptosis, and Cooperates with the BCL-2 Family to Promote Clonogenic Survival Gary X. Wang,
Cell Reports, Volume 21 Supplemental Information DNp63 Inhibits Oxidative Stress-Induced Cell Death, Including Ferroptosis, and Cooperates with the BCL-2 Family to Promote Clonogenic Survival Gary X. Wang,
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature09732 Supplementary Figure 1: Depletion of Fbw7 results in elevated Mcl-1 abundance. a, Total thymocytes from 8-wk-old Lck-Cre/Fbw7 +/fl (Control) or Lck-Cre/Fbw7 fl/fl (Fbw7 KO) mice
doi:10.1038/nature09732 Supplementary Figure 1: Depletion of Fbw7 results in elevated Mcl-1 abundance. a, Total thymocytes from 8-wk-old Lck-Cre/Fbw7 +/fl (Control) or Lck-Cre/Fbw7 fl/fl (Fbw7 KO) mice
Supplementary Figure 1: Overexpression of EBV-encoded proteins Western blot analysis of the expression levels of EBV-encoded latency III proteins in
 Supplementary Figure 1: Overexpression of EBV-encoded proteins Western blot analysis of the expression levels of EBV-encoded latency III proteins in BL2 cells. The Ponceau S staining of the membranes or
Supplementary Figure 1: Overexpression of EBV-encoded proteins Western blot analysis of the expression levels of EBV-encoded latency III proteins in BL2 cells. The Ponceau S staining of the membranes or
Blimp-1/PRDM1 rabbit monoclonal antibody (C14A4) was purchased from Cell Signaling
 1 2 3 4 5 6 7 8 9 10 11 Supplementary Methods Antibodies Blimp-1/PRDM1 rabbit monoclonal antibody (C14A4) was purchased from Cell Signaling (Danvers, MA) and used at 1:1000 to detect the total PRDM1 protein
1 2 3 4 5 6 7 8 9 10 11 Supplementary Methods Antibodies Blimp-1/PRDM1 rabbit monoclonal antibody (C14A4) was purchased from Cell Signaling (Danvers, MA) and used at 1:1000 to detect the total PRDM1 protein
Supplementary Material
 Supplementary Material Supplementary Methods Cell synchronization. For synchronized cell growth, thymidine was added to 30% confluent U2OS cells to a final concentration of 2.5mM. Cells were incubated
Supplementary Material Supplementary Methods Cell synchronization. For synchronized cell growth, thymidine was added to 30% confluent U2OS cells to a final concentration of 2.5mM. Cells were incubated
Fig. S1. Effect of p120-catenin overexpression on the interaction of SCUBE2 with E-cadherin. The expression plasmid encoding FLAG.
 Fig. S1. Effect of p120-catenin overexpression on the interaction of SCUBE2 with E-cadherin. The expression plasmid encoding FLAG.SCUBE2, E-cadherin.Myc, or HA.p120-catenin was transfected in a combination
Fig. S1. Effect of p120-catenin overexpression on the interaction of SCUBE2 with E-cadherin. The expression plasmid encoding FLAG.SCUBE2, E-cadherin.Myc, or HA.p120-catenin was transfected in a combination
Supplementary Figure Legends
 Supplementary Figure Legends Figure S1 gene targeting strategy for disruption of chicken gene, related to Figure 1 (f)-(i). (a) The locus and the targeting constructs showing HpaI restriction sites. The
Supplementary Figure Legends Figure S1 gene targeting strategy for disruption of chicken gene, related to Figure 1 (f)-(i). (a) The locus and the targeting constructs showing HpaI restriction sites. The
Investigating the regulation of mirna biogenesis and Argonaute2 by RNA binding proteins
 Investigating the regulation of mirna biogenesis and Argonaute2 by RNA binding proteins Patrick Peter Connerty Supervisor: Gyorgy Hutvagner Thesis Submitted for the Degree of Doctor of Philosophy (Science)
Investigating the regulation of mirna biogenesis and Argonaute2 by RNA binding proteins Patrick Peter Connerty Supervisor: Gyorgy Hutvagner Thesis Submitted for the Degree of Doctor of Philosophy (Science)
Supplementary Figure 1. Soft fibrin gels promote growth and organized mesodermal differentiation. Representative images of single OGTR1 ESCs cultured
 Supplementary Figure 1. Soft fibrin gels promote growth and organized mesodermal differentiation. Representative images of single OGTR1 ESCs cultured in 90-Pa 3D fibrin gels for 5 days in the presence
Supplementary Figure 1. Soft fibrin gels promote growth and organized mesodermal differentiation. Representative images of single OGTR1 ESCs cultured in 90-Pa 3D fibrin gels for 5 days in the presence
ChampionChIP Quick, High Throughput Chromatin Immunoprecipitation Assay System
 ChampionChIP Quick, High Throughput Chromatin Immunoprecipitation Assay System Liyan Pang, Ph.D. Application Scientist 1 Topics to be Covered Introduction What is ChIP-qPCR? Challenges Facing Biological
ChampionChIP Quick, High Throughput Chromatin Immunoprecipitation Assay System Liyan Pang, Ph.D. Application Scientist 1 Topics to be Covered Introduction What is ChIP-qPCR? Challenges Facing Biological
Supplementary Figure 1
 Supplementary Figure 1 Supplementary Fig. 1 shrna mediated knockdown of ZRSR2 in K562 and 293T cells. (a) ZRSR2 transcript levels in stably transduced K562 cells were determined using qrt-pcr. GAPDH was
Supplementary Figure 1 Supplementary Fig. 1 shrna mediated knockdown of ZRSR2 in K562 and 293T cells. (a) ZRSR2 transcript levels in stably transduced K562 cells were determined using qrt-pcr. GAPDH was
monoclonal antibody. (a) The specificity of the anti-rhbdd1 monoclonal antibody was examined in
 Supplementary information Supplementary figures Supplementary Figure 1 Determination of the s pecificity of in-house anti-rhbdd1 mouse monoclonal antibody. (a) The specificity of the anti-rhbdd1 monoclonal
Supplementary information Supplementary figures Supplementary Figure 1 Determination of the s pecificity of in-house anti-rhbdd1 mouse monoclonal antibody. (a) The specificity of the anti-rhbdd1 monoclonal
Supplementary methods Shoc2 In Vitro Ubiquitination Assay
 Supplementary methods Shoc2 In Vitro Ubiquitination Assay 35 S-labelled Shoc2 was prepared using a TNT quick Coupled transcription/ translation System (Promega) as recommended by manufacturer. For the
Supplementary methods Shoc2 In Vitro Ubiquitination Assay 35 S-labelled Shoc2 was prepared using a TNT quick Coupled transcription/ translation System (Promega) as recommended by manufacturer. For the
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/10/496/eaam6291/dc1 Supplementary Materials for Regulation of autophagy, NF-κB signaling, and cell viability by mir-124 in KRAS mutant mesenchymal-like NSCLC cells
www.sciencesignaling.org/cgi/content/full/10/496/eaam6291/dc1 Supplementary Materials for Regulation of autophagy, NF-κB signaling, and cell viability by mir-124 in KRAS mutant mesenchymal-like NSCLC cells
Figure S1. Optimization of reverse-transfection protocols using GAPDH sirnas. MDA- MB-231 cells were seeded in 96-well plates at a density of 5000
 Figure S1. Optimization of reverse-transfection protocols using GAPDH sirnas. MDA- MB-231 cells were seeded in 96-well plates at a density of 5000 cells per well, and reverse transfected with the indicated
Figure S1. Optimization of reverse-transfection protocols using GAPDH sirnas. MDA- MB-231 cells were seeded in 96-well plates at a density of 5000 cells per well, and reverse transfected with the indicated
T H E J O U R N A L O F C E L L B I O L O G Y
 T H E J O U R N A L O F C E L L B I O L O G Y Supplemental material Rainero et al., http://www.jcb.org/cgi/content/full/jcb.201109112/dc1 Figure S1. The expression of DGK- is reduced upon transfection
T H E J O U R N A L O F C E L L B I O L O G Y Supplemental material Rainero et al., http://www.jcb.org/cgi/content/full/jcb.201109112/dc1 Figure S1. The expression of DGK- is reduced upon transfection
Rajesh Pandey et al. Alu-miRNA interactions modulate transcript isoform diversity in stress response and reveal signatures of positive selection
 Rajesh Pandey et al. Alu-miRNA interactions modulate transcript isoform diversity in stress response and reveal signatures of positive selection Rajesh Pandey 1, Aniket Bhattacharya 2,3#, Vivek Bhardwaj
Rajesh Pandey et al. Alu-miRNA interactions modulate transcript isoform diversity in stress response and reveal signatures of positive selection Rajesh Pandey 1, Aniket Bhattacharya 2,3#, Vivek Bhardwaj
Thermo Scientific DharmaFECT Transfection Reagents sirna Transfection Protocol
 Protocol Thermo Scientific Transfection Reagents sirna Transfection Protocol The following is a general protocol for use of Thermo Scientific transfection reagents to deliver sirna into cultured mammalian
Protocol Thermo Scientific Transfection Reagents sirna Transfection Protocol The following is a general protocol for use of Thermo Scientific transfection reagents to deliver sirna into cultured mammalian
Supplementary Fig. 1
 a FL (1-2266) NL (1-1190) CL (1191-2266) HA-ICE1: - HA-ICE1: - - - FLAG-ICE2: + + + + FLAG-ELL: + + + + + + IP: anti-ha FLAG-ICE2 HA-ICE1-FL HA-ICE1-NL HA-ICE1-CL FLAG-ICE2 b IP: anti-ha FL (1-2266) NL
a FL (1-2266) NL (1-1190) CL (1191-2266) HA-ICE1: - HA-ICE1: - - - FLAG-ICE2: + + + + FLAG-ELL: + + + + + + IP: anti-ha FLAG-ICE2 HA-ICE1-FL HA-ICE1-NL HA-ICE1-CL FLAG-ICE2 b IP: anti-ha FL (1-2266) NL
Supplementary Information. HBx-upregulated lncrna UCA1 promotes cell growth and tumorigenesis
 Supplementary Information HBx-upregulated lncrna UCA1 promotes cell growth and tumorigenesis by recruiting EZH2 and repressing p27kip1/cdk2 signaling Jiao-Jiao Hu 1, Wei Song 1, Shao-Dan Zhang 1, Xiao-Hui
Supplementary Information HBx-upregulated lncrna UCA1 promotes cell growth and tumorigenesis by recruiting EZH2 and repressing p27kip1/cdk2 signaling Jiao-Jiao Hu 1, Wei Song 1, Shao-Dan Zhang 1, Xiao-Hui
Supplementary Figure 1, Wiel et al
 Supplementary Figure 1, Wiel et al Supplementary Figure 1 ITPR2 increases in benign tumors and decreases in aggressive ones (a-b) According to the Oncomine database, expression of ITPR2 increases in renal
Supplementary Figure 1, Wiel et al Supplementary Figure 1 ITPR2 increases in benign tumors and decreases in aggressive ones (a-b) According to the Oncomine database, expression of ITPR2 increases in renal
mmu-mir-34a Real-time RT-PCR Detection Kit User Manual
 mmu-mir-34a Real-time RT-PCR Detection Kit User Manual Catalog # CPK1272 For the detection and quantification of mirna mmu-mir-34a using Real-Time RT-PCR detection instruments. For research use only. Not
mmu-mir-34a Real-time RT-PCR Detection Kit User Manual Catalog # CPK1272 For the detection and quantification of mirna mmu-mir-34a using Real-Time RT-PCR detection instruments. For research use only. Not
Supplementary Figure S1 Evi/Wls and Wnt3 are overexpressed in epithelial cells in colon cancers. (a) Validation of the specificity of the Wnt3
 Supplementary Figure S1 Evi/Wls and Wnt3 are overexpressed in epithelial cells in colon cancers. (a) Validation of the specificity of the Wnt3 antibody. HCT116 cells were reverse transfected with sicontrol
Supplementary Figure S1 Evi/Wls and Wnt3 are overexpressed in epithelial cells in colon cancers. (a) Validation of the specificity of the Wnt3 antibody. HCT116 cells were reverse transfected with sicontrol
To determine MRK-003 IC50 values, cell lines were plated in triplicate in 96-well plates at 3 x
 Supplementary methods: Cellular growth assay and cell cycle analysis To determine MRK-003 IC50 values, cell lines were plated in triplicate in 96-well plates at 3 x 10 3 cells/well (except for TALL1 and
Supplementary methods: Cellular growth assay and cell cycle analysis To determine MRK-003 IC50 values, cell lines were plated in triplicate in 96-well plates at 3 x 10 3 cells/well (except for TALL1 and
b alternative classical none
 Supplementary Figure. 1: Related to Figure.1 a d e b alternative classical none NIK P-IkBa Total IkBa Tubulin P52 (Lighter) P52 (Darker) RelB (Lighter) RelB (Darker) HDAC1 Control-Sh RelB-Sh NF-kB2-Sh
Supplementary Figure. 1: Related to Figure.1 a d e b alternative classical none NIK P-IkBa Total IkBa Tubulin P52 (Lighter) P52 (Darker) RelB (Lighter) RelB (Darker) HDAC1 Control-Sh RelB-Sh NF-kB2-Sh
a KYSE270-CON KYSE270-Id1
 a KYSE27-CON KYSE27- shcon shcon sh b Human Mouse CD31 Relative MVD 3.5 3 2.5 2 1.5 1.5 *** *** c KYSE15 KYSE27 sirna (nm) 5 1 Id2 Id2 sirna 5 1 sirna (nm) 5 1 Id2 sirna 5 1 Id2 [h] (pg per ml) d 3 2 1
a KYSE27-CON KYSE27- shcon shcon sh b Human Mouse CD31 Relative MVD 3.5 3 2.5 2 1.5 1.5 *** *** c KYSE15 KYSE27 sirna (nm) 5 1 Id2 Id2 sirna 5 1 sirna (nm) 5 1 Id2 sirna 5 1 Id2 [h] (pg per ml) d 3 2 1
Chemically defined conditions for human ipsc derivation and culture
 Nature Methods Chemically defined conditions for human ipsc derivation and culture Guokai Chen, Daniel R Gulbranson, Zhonggang Hou, Jennifer M Bolin, Victor Ruotti, Mitchell D Probasco, Kimberly Smuga-Otto,
Nature Methods Chemically defined conditions for human ipsc derivation and culture Guokai Chen, Daniel R Gulbranson, Zhonggang Hou, Jennifer M Bolin, Victor Ruotti, Mitchell D Probasco, Kimberly Smuga-Otto,
Supplementary Information
 Supplementary Information promotes cancer cell invasion and proliferation by receptor-mediated endocytosis-dependent and -independent mechanisms, respectively Kensaku Shojima, Akira Sato, Hideaki Hanaki,
Supplementary Information promotes cancer cell invasion and proliferation by receptor-mediated endocytosis-dependent and -independent mechanisms, respectively Kensaku Shojima, Akira Sato, Hideaki Hanaki,
SUPPLEMENTAL EXPERIMENTAL PROCEDURES
 SUPPLEMENTAL EXPERIMENTAL PROCEDURES Luciferase Assays Cells were seeded on 24well plates and grown to 7% confluency. Cells were then transfected with ng of reporter constructs and 1 ng of the renilla
SUPPLEMENTAL EXPERIMENTAL PROCEDURES Luciferase Assays Cells were seeded on 24well plates and grown to 7% confluency. Cells were then transfected with ng of reporter constructs and 1 ng of the renilla
Lullaby sirna Transfection Reagent - Results
 sirna Transfection Reagent - Results OZ Biosciences is delighted to announce the launching of a new sirna transfection reagent: -sirna. This lipid based transfection reagent is specifically designed for
sirna Transfection Reagent - Results OZ Biosciences is delighted to announce the launching of a new sirna transfection reagent: -sirna. This lipid based transfection reagent is specifically designed for
pdsipher and pdsipher -GFP shrna Vector User s Guide
 pdsipher and pdsipher -GFP shrna Vector User s Guide NOTE: PLEASE READ THE ENTIRE PROTOCOL CAREFULLY BEFORE USE Page 1. Introduction... 1 2. Vector Overview... 1 3. Vector Maps 2 4. Materials Provided...
pdsipher and pdsipher -GFP shrna Vector User s Guide NOTE: PLEASE READ THE ENTIRE PROTOCOL CAREFULLY BEFORE USE Page 1. Introduction... 1 2. Vector Overview... 1 3. Vector Maps 2 4. Materials Provided...
RPL27A is a target of mir-595 and may contribute to the myelodysplastic phenotype through ribosomal dysgenesis
 www.impactjournals.com/oncotarget/ Oncotarget, Supplementary Materials 2016 RPL27A is a target of mir-595 and may contribute to the myelodysplastic phenotype through ribosomal dysgenesis Supplementary
www.impactjournals.com/oncotarget/ Oncotarget, Supplementary Materials 2016 RPL27A is a target of mir-595 and may contribute to the myelodysplastic phenotype through ribosomal dysgenesis Supplementary
SideStep Lysis and Stabilization Buffer
 SideStep Lysis and Stabilization Buffer INSTRUCTION MANUAL Catalog #400900 Revision B.0 For Research Use Only. Not for use in diagnostic procedures. 400900-12 LIMITED PRODUCT WARRANTY This warranty limits
SideStep Lysis and Stabilization Buffer INSTRUCTION MANUAL Catalog #400900 Revision B.0 For Research Use Only. Not for use in diagnostic procedures. 400900-12 LIMITED PRODUCT WARRANTY This warranty limits
