Supplemental Information. Kinesin 1 Drives Autolysosome Tubulation
|
|
|
- Zoe Kelley
- 5 years ago
- Views:
Transcription
1 Developmental Cell, Volume 37 Supplemental Information Kinesin 1 Drives Autolysosome Tubulation Wanqing Du, Qian Peter Su, Yang Chen, Yueyao Zhu, Dong Jiang, Yueguang Rong, Senyan Zhang, Yixiao Zhang, He Ren, Chuanmao Zhang, Xinquan Wang, Ning Gao, Yanfeng Wang, Lingfei Sun, Yujie Sun, and Li Yu
2 Supplemental Inventory Figure S1. KIF5B localizes to LAMP1-positive organelles. Related to Figure 1A-C. Figure 1A-C showed that KIF5B could rescue the defect of autolysosome reformation in kif5b knock out cells. Figure S1 showed the localization of KIF5B in support of its functions in cells.! Figure S2. WT and T92N KIF5B both localize to purified autolysosomes. Related to Figure 2H, K. Figure 2H, K showed that KIF5B WT but not KIF5B T92N could drive autolysosome tubulation in vitro. Figure S2 showed that both KIF5B WT and KIF5B T92N, but not an irrelevant protein mouse IgG localized to purified autolysosomes in vitro, which supported that tubulation required both localization and motor activity.! Figure S3. Knockdown of kinesin light chain 1 or 2 (KLC1/KLC2) does not affect autolysosome tubulation. Related to Figure 3A, B. Figure 3A, B showed that purified autolysosomes from pip5k1b knockdown cells cannot be tubulated in the in vitro reconstitution system. Figure S3 showed that interfering with other Kif5B adaptors has no effect on autolysosome tubulation, which indicates KIF5B may regulate autolysosome tubulation through PtdIns(4,5) 2. Figure S4. Determination of the lipid binding specificity of KIF5B with a protein lipid overlay assay. Related to Figure 3C, D. Figure 3C, D showed binding of KIF5B to PtdIns(4,5)P 2. Figure S4 showed the lipid binding specificity of KIF5B.! Figure S5. Mapping of the PtdIns(4,5)P 2 -binding domain of KIF5B using a liposome flotation assay. Related to Figure 3C, D. Figure 3C, D showed binding of KIF5B to PtdIns(4,5)P 2. Figure S5 showed detalied domain mapping of PtdIns(4,5)P 2 -KIF5B binding.! Fig. S6. Analysis of CRISPR/Cas knock out KIF5B cell line. Related to Figure 1. The KIF5B knock out cell lines used in Figure 1 was analyzed in Figure S6.! Movie S1.! Supplemental Experimental Procedures. The primer sequence used in cloning, the plasmids cloned in this study and the RNAi sequence used in this study were listed.
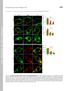
3 Figure S1 A B NRK 8h! 8h 0 C +KIF5B T92N! +KIF5B T92N! 8h! 0h +KIF5B WT! +KIF5B T92N! 0h! KIF5B WT! 8h! NRK! +KIF5B WT! 0h! % of KIF5B positive! LAMP1 structures! NRK 0h! KIF5B! KIF5B! LAMP1! Actin! Fig. S1. KIF5B localizes to LAMP1-positive organelles. Related to Figure 1A-C (A) NRK cells and kif5b knockout cells stably expressing WT KIF5B or KIF5B T92N were starved for 0h or 8h, treated with 25 μg/ml digitonin and stained with antibodies against KIF5B and LAMP1. Scale bar, 5 μm. (B) Autolysosomes from A were assessed for the presence of KIF5B puncta and quantified. n> 500 autolysosomes from three independent experiments were quantified. Error bars indicate the s.d. The result that KIF5B could rescue the phenotype of kif5b knock out cells were depicted in Figure 1A-C. (C) The KIF5B protein expression level was tested by Western Blot using anti-kif5b antibody.
4 Figure S2 MTs! Autolysosomes! KIF5B! Merge! MTs! Autolysosomes! KIF5B T92N! Merge! MTs! Autolysosomes! Mouse IgG! Merge! 2μm Fig. S2: WT and T92N KIF5B both localize to purified autolysosomes. Related to Figure 2H, K. Purified autolysosomes were stained by CM-DiI (yellow), incubated with a mixture of nonfluorescent labeled KIF5B and Atto488 labeled KIF5B WT or incubated with Atto488 labeled KIF5B T92N, and transferred into flow chamber channels coated with Hilyte Fluor 647-labeled microtubules (gray). Purified autolysosomes were stained by CM-DiI (yellow), incubated with a mixture of nonfluorescent labeled KIF5B and Atto488 mouse IgG as negative control. The images were collected with a Nikon TIRF microscope. Scale bar, 2μm. Figure 2H, K showed that KIF5B WT but not KIF5B T92N could drive autolysosome tubulation in vitro.
5 Figure S3 A NS! 8h KLC2! RNAi! 8h NS! 8h! KLC1! RNAi! 8h! LAMP1 B % Cells with tubular! autolysosomes C % KLC2 mrna level D! % Cells with tubular! autolysosomes! 80! 80! 60! 60! 40! 40! 20! 20! 0! 0! NS!KLC1! % KLC1 mrna level! E! 120! 80! 80! 40! 40! 0! 0! Fig. S3. Knockdown of kinesin light chain 1 or 2 (KLC1/KLC2) does not affect autolysosome tubulation. Related to Figure 3A, B. (A) NRK cells stably expressing LAMP1-mCherry (Red) were transfected with nonspecific (NS)- or klc1/2-rnai and starved for 8 h. Scale bar, 5 μm. (B)(D) Cells from A were assessed for tubular autolysosomes after 8 h starvation and quantified. n=100 cells from three independent experiments. Error bars indicate the s.d. (C)(E) Cells were transfected with non-specific (NS) RNAi or RNAi against klc1/2. After 60 h of transfection, mrna levels were measured by qpcr. qpcr was performed in triplicate and the data represent the mean ± s.d. This experiment was repeated 3 times. (D) mrna levels of KLC1 and KLC3 in NRK cells were determined by RNAseq. Beta-actin was used as the control. This result supplement Figure 3A, B which showed that lacking pip5k1b interfered with autolysosome tubulation in vitro. Figure S3 showed that interfered with other Kif5B adaptors has no effect on autolysosome tubulation.
6 Figure S4 Triglyceride DAG PA PS PE PC PG Cardiolipin PI PtdIns4P PtdIns(4,5)P 2 PtdIns(3,4,5)P 3 Cholesterol Sphingomyelin Sulfatide Blue Blank Fig. S4. Determination of the lipid binding specificity of KIF5B with a protein lipid overlay assay. Related to Figure 3C, D. The PIP strip was incubated with 2 μg/ ml purified KIF5B in 20 mm Tris-HCl, ph 7.4, 150 mm NaCl, supplemented with 10% (w/v) milk powder for 1 h at room temperature. Binding was detected with an anti- KIF5B antibody and an HRP-conjugated secondary antibody. The binding of KIF5B to PtdIns(4,5)P 2 was carefully analyzed in Figure 3C, D.
7 Figure S5 GST-! ! Protein+Liposome! Protein only! 55! 35! GST-! Cargo domain! Protein+Liposome! Protein only! 70! 0%! His-1-813! His-! Motor domain! Protein+Liposome! Protein only! Protein+Liposome! Protein only! 100! 70! 20%! 25%! 30%! GST! Protein+Liposome! 25! 1 Protein only! 555! 560! 814! 963! Motor domain! Cargo-binding domain! Fig. S5. Mapping of the PtdIns(4,5)P 2 -binding domain of KIF5B using a liposome flotation assay. Related to Figure 3C,D. Main panel: Retention of different domains of KIF5B by 25% PtdIns(4,5)P 2 liposomes in a sucrose gradient. The GST protein serves as a negative control for detecting nonspecific binding. Each fraction of the total sample was visualized by Coomassie staining. When a protein binds to liposomes, it is found in the top fractions. In contrast, if there are no interactions, the protein is localized in the bottom four fractions. Right panel: Schematic representation of sucrose gradient. Bottom panel: Schematic representation of different KIF5B domains used in this experiment. The binding of KIF5B to PtdIns(4,5)P 2 was carefully analyzed in Figure 3C, D.
8 Figure S6 A AATTCTACTTTGTTTTCTATGTAGGGTAAACTTCATGATCCAGAAGGCATGGGGATTATTCC (Wild-type)! KIF5B - /- B AATTCTACTTTGTTTTCTATGTAGGGTAAACTTCATGATC-AGAAGGCATGGGGATTATTCC (Δ1, 3/10)! AATTCTACTTTGTTTTCTATGTAGGGTAAACTTCATGAT-CAGAAGGCATGGGGATTATTCC (Δ1, 7/10)! grna! On-target! Targeted sequence (with PAM)! Genomic Site! Locus! KIF5B! TRUE! GTAAACTTCATGATCCAGA AGG! chr17: ! NM_057202! N/A! Mean Indels (%)+SD! FALSE! GGTGGACTTGAAGATCCAGA GGG! chr1: ! NM_ ! 0! FALSE! GTTAACCTCCATCATCCAGA TAG! chr15: ! NM_ ! 0! FALSE! GGAAAATTTCATTATCTAGA AGG! chr1: ! NM_ ! 0! FALSE! GGAAAATTTCATTATCTAGA AGG! chr2: ! NM_ ! 0! FALSE! GGAAGACTTCATGATAGAGA AAG! chr1: ! NM_012693! 0! FALSE! GGTAAACATCATGATGAAAA AAG! chr4: ! NM_181381! 0! FALSE! GGTATACTTCATGGTGCAAA GGG! chr2: ! NM_ ! 0! Fig. S6. Analysis of CRISPR/Cas knock out KIF5B cell line. Related to Figure 1. (A) DNA sequencing analysis of mutated alleles in the Rat kif5b locus. Partial coding sequences of targeted genes in the genome containing the CRISPR/Cas9 grna KIF5B binding region (underlined) and sequencing analysis of the mutated alleles from a single clone of KIF5B knock out cells were shown. Primers used for the PCR reactions are listed in Supplemental Experimental Procedures, and the red nucleotides represent the PAM sequences that guide the Cas9 for DNA recognition and cleavage. (B) Off-target analysis of 7 potential off-target sites. All on-target and off-target sites were individually amplified from genomic DNA from KIF5B knock out NRK cells. Off-target sites were chosen based on analysis using the CRISPR design tool ( crispr.mit.edu/), and the seven potential off-target sites of the sgrna were selected for T7E1 assay to calculate indels with ImageJ program. The mismatch bases were highlighted in red and the PAM site is highlighted in blue. Primers used to amplify the off-target sites are listed in Supplemental Experimental Procedures. The kif5b knock out cell lines were used in Figure 1.
9 Movie S1 Movie S1. Taxol-stabilized microtubules moving over a lawn of full-length KIF5B. Related to Figure 2B. Full-length KIF5B was introduced into flow chambers, followed by Hylite Fluor 647-labled microtubules. An ATP mixture containing 20 μm ATP was subsequently injected into the chambers, and the images were recorded every 0.5 s for 55 s using a Nikon Ti-E TIRF microscope. Scale bar, 5 μm.
10 Supplemental Experimental Procedures Primer sequence: Name P1 5'KIF5B Motor_NDE1 GGAATTCCATATGGCGGACCCGGCGGAGTGCAAC P2 3'KIF5B Motor_Xho1 CCGCTCGAGTTTTAGTAATGATGCCATCATTTCAGCT P3 5'KIF5B pfast_bamh1 CGCGGATCCATGCACCACCACCACCACCACATGGCGGACCCGGCG GAG P4 3'KIF5B pfast_xba1 CTAGTCTAGATCACGCCTGCTTGCCTCCAC P5 5'KIF5B T92N CATCTGGAAAGAACCACACAATGGAGGGTAAACTT P6 3'KIF5B T92N CATTGTGTGGTTCTTTCCAGATGATGTTTGTCCA P7 5'pGEX- 4T- 1 KIF5B CGCGGATCCAGCGCTGAGGTGGACTCAG P8 3'pGEX- 4T- 1 KIF5B AAGGAAAAAAGCGGCCGCTCACGCCTGCTTGCCTCCAC P9 5'pGEX- 4T- 1 KIF5B cargo CGCGGATCCATGGCATCATTACTAAAAGACCTT P10 3'pGEX- 4T- 1 KIF5B cargo AAGGAAAAAAGCGGCCGCTCACGCCTGCTTGCCTCCAC P11 5'pCAG- RatKif5A_MLUI CGATACGCGTATGGCGGAGACCAATAACGAATG P12 3'pCAG- RatKif5A_NotI CGATGCGGCCGCTTAGCTGGCTGCCGTCTCTTGGTGG P13 5'pCAG- RatKif5C_MLUI CGATACGCGTATGGCGGATCCAGCCGAATGC P14 3'pCAG- RatKif5C_NotI CGATGCGGCCGCTTACTTCTGGTAGTGAGTGGAGTTTG P15 5'pCAG- RatKif1A_MLUI CGATACGCGTATGGCTGGGGCCTCTGTGAAGG P16 3'pCAG- RatKif1A_NotI CGATGCGGCCGCTCAGACCCGCATCTGCGCAG P17 5'pCAG- RatKif5B_MLUI CGATACGCGTATGGCGGACCCGGCGGAGTG P18 3'pCAG- RatKif5B_NotI CGATGCGGCCGCTCACGCCTGCTTGCCTCCACC P19 5'Rat KLC1_qPCR1 GCAGGGTCTTGACAATGTTC P20 3'Rat KLC1_qPCR1 GAGCTTCATCTTTCTCATTTCCT P21 5'Rat KLC2_qPCR1 GGCTCAGTCAATGGAGAGAA P22 3'Rat KLC2_qPCR1 ACTGTGGGACTGTCAACTTT P23 KIF5B- On target F ATGAGCTTGGTTAGTAAGCTGT P24 KIF5B- On target R AGAAGCTAAGAACAGACTTGGC P25 KIF5B- Off target 1F GGGACCAGAGTTTCCTTCCG P26 KIF5B- Off target 1R CCACAAGCTCGGAAGACTCA P27 KIF5B- Off target 2F GGAGTACAACGCGATTCAGC P28 KIF5B- Off target 2R GCCGCAGTCATACACCCTAT P29 KIF5B- Off target 3F TCCTTCGTACTGGAGATGAAACC P30 KIF5B- Off target 3R GCTAATGCTAACAAGTTTTTGACC P31 KIF5B- Off target 5F GTTTTCTTGCCATCCTCAATCCAT P32 KIF5B- Off target 5R AATGCTAGAGTCCAGGTGGCTAA P33 KIF5B- Off target 6F CATGGAAGCCGGTTGTGTTTC P34 KIF5B- Off target 6R CACTAGATGCAGTGGTTTCAATGG P35 KIF5B- Off target 7F TGCCTCTGCTACTTAGGCTTT P36 KIF5B- Off target 7R TTGGCACATTTTATTTCTCTGTCCC plasmid information: Name insert primer His- KIF5B motor Rat KIF5B motor domain (1-560) P1, P2 KIF5B WT pfastbac DUAL Rat KIF5B WT P3, P4 KIF5B T92N pfastbac DUAL Rat KIF5B T92N P5, P6 GST- KIF5B Rat KIF5B P7, P8 GST- KIF5B cargo ( ) Rat KIF5B cargo( ) P9, P10 pcag- KIF5B Rat KIF5B P11, P12 pcag- KIF5A Rat KIF5A P13, P14 pcag- KIF5C Rat KIF5C P15, P16 pcag- KIF1A Rat KIF1A P17, P18 RNAi sequence: PIP5K1B RNAi clathrin heavy chain RNAi Rat KLC2 Rat KLC1 CCGCTGGCTTTCCGCTACT CAGAGCCATGCTGTCTGCTAATATT TGCTATTCATGAGTCAGATCCGAAA GCATGATCCGCAAGTCCTT
Supplemental Information. Pacer Mediates the Function of Class III PI3K. and HOPS Complexes in Autophagosome. Maturation by Engaging Stx17
 Molecular Cell, Volume 65 Supplemental Information Pacer Mediates the Function of Class III PI3K and HOPS Complexes in Autophagosome Maturation by Engaging Stx17 Xiawei Cheng, Xiuling Ma, Xianming Ding,
Molecular Cell, Volume 65 Supplemental Information Pacer Mediates the Function of Class III PI3K and HOPS Complexes in Autophagosome Maturation by Engaging Stx17 Xiawei Cheng, Xiuling Ma, Xianming Ding,
Transfection of CRISPR/Cas9 Nuclease NLS ribonucleoprotein (RNP) into adherent mammalian cells using Lipofectamine RNAiMAX
 Transfection of CRISPR/Cas9 Nuclease NLS ribonucleoprotein (RNP) into adherent mammalian cells using Lipofectamine RNAiMAX INTRODUCTION The CRISPR/Cas genome editing system consists of a single guide RNA
Transfection of CRISPR/Cas9 Nuclease NLS ribonucleoprotein (RNP) into adherent mammalian cells using Lipofectamine RNAiMAX INTRODUCTION The CRISPR/Cas genome editing system consists of a single guide RNA
CRISPR RNA-guided activation of endogenous human genes
 CRISPR RNA-guided activation of endogenous human genes Morgan L Maeder, Samantha J Linder, Vincent M Cascio, Yanfang Fu, Quan H Ho, J Keith Joung Supplementary Figure 1 Comparison of VEGF activation induced
CRISPR RNA-guided activation of endogenous human genes Morgan L Maeder, Samantha J Linder, Vincent M Cascio, Yanfang Fu, Quan H Ho, J Keith Joung Supplementary Figure 1 Comparison of VEGF activation induced
Generation of App knock-in mice reveals deletion mutations protective against Alzheimer s. disease-like pathology. Nagata et al.
 Generation of App knock-in mice reveals deletion mutations protective against Alzheimer s disease-like pathology Nagata et al. Supplementary Fig 1. Previous App knock-in model did not show Aβ accumulation
Generation of App knock-in mice reveals deletion mutations protective against Alzheimer s disease-like pathology Nagata et al. Supplementary Fig 1. Previous App knock-in model did not show Aβ accumulation
A CRISPR/Cas9 Vector System for Tissue-Specific Gene Disruption in Zebrafish
 Developmental Cell Supplemental Information A CRISPR/Cas9 Vector System for Tissue-Specific Gene Disruption in Zebrafish Julien Ablain, Ellen M. Durand, Song Yang, Yi Zhou, and Leonard I. Zon % larvae
Developmental Cell Supplemental Information A CRISPR/Cas9 Vector System for Tissue-Specific Gene Disruption in Zebrafish Julien Ablain, Ellen M. Durand, Song Yang, Yi Zhou, and Leonard I. Zon % larvae
Supplementary data. sienigma. F-Enigma F-EnigmaSM. a-p53
 Supplementary data Supplemental Figure 1 A sienigma #2 sienigma sicontrol a-enigma - + ++ - - - - - - + ++ - - - - - - ++ B sienigma F-Enigma F-EnigmaSM a-flag HLK3 cells - - - + ++ + ++ - + - + + - -
Supplementary data Supplemental Figure 1 A sienigma #2 sienigma sicontrol a-enigma - + ++ - - - - - - + ++ - - - - - - ++ B sienigma F-Enigma F-EnigmaSM a-flag HLK3 cells - - - + ++ + ++ - + - + + - -
T H E J O U R N A L O F C E L L B I O L O G Y
 T H E J O U R N A L O F C E L L B I O L O G Y Supplemental material Han et al., http://www.jcb.org/cgi/content/full/jcb.201311007/dc1 Figure S1. SIVA1 interacts with PCNA. (A) HEK293T cells were transiently
T H E J O U R N A L O F C E L L B I O L O G Y Supplemental material Han et al., http://www.jcb.org/cgi/content/full/jcb.201311007/dc1 Figure S1. SIVA1 interacts with PCNA. (A) HEK293T cells were transiently
Nature Biotechnology: doi: /nbt.4166
 Supplementary Figure 1 Validation of correct targeting at targeted locus. (a) by immunofluorescence staining of 2C-HR-CRISPR microinjected embryos cultured to the blastocyst stage. Embryos were stained
Supplementary Figure 1 Validation of correct targeting at targeted locus. (a) by immunofluorescence staining of 2C-HR-CRISPR microinjected embryos cultured to the blastocyst stage. Embryos were stained
JCB. Supplemental material THE JOURNAL OF CELL BIOLOGY. Hong et al.,
 Supplemental material JCB Hong et al., http://www.jcb.org/cgi/content/full/jcb.201412127/dc1 THE JOURNAL OF CELL BIOLOGY Figure S1. Analysis of purified proteins by SDS-PAGE and pull-down assays. (A) Coomassie-stained
Supplemental material JCB Hong et al., http://www.jcb.org/cgi/content/full/jcb.201412127/dc1 THE JOURNAL OF CELL BIOLOGY Figure S1. Analysis of purified proteins by SDS-PAGE and pull-down assays. (A) Coomassie-stained
THE JOURNAL OF CELL BIOLOGY
 Supplemental Material THE JOURNAL OF CELL BIOLOGY Yeung et al., http://www.jcb.org/cgi/content/full/jcb.200903020/dc1 Figure S1. Assessment of the surface charge of maturing phagosomes. (A C and F H) RAW
Supplemental Material THE JOURNAL OF CELL BIOLOGY Yeung et al., http://www.jcb.org/cgi/content/full/jcb.200903020/dc1 Figure S1. Assessment of the surface charge of maturing phagosomes. (A C and F H) RAW
Supplementary Figure S1. The tetracycline-inducible CRISPR system. A) Hela cells stably
 Supplementary Information Supplementary Figure S1. The tetracycline-inducible CRISPR system. A) Hela cells stably expressing shrna sequences against TRF2 were examined by western blotting. shcon, shrna
Supplementary Information Supplementary Figure S1. The tetracycline-inducible CRISPR system. A) Hela cells stably expressing shrna sequences against TRF2 were examined by western blotting. shcon, shrna
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb3240 Supplementary Figure 1 GBM cell lines display similar levels of p100 to p52 processing but respond differentially to TWEAK-induced TERT expression according to TERT promoter mutation
DOI: 10.1038/ncb3240 Supplementary Figure 1 GBM cell lines display similar levels of p100 to p52 processing but respond differentially to TWEAK-induced TERT expression according to TERT promoter mutation
Fig. S1. Endocytosis of extracellular cargo does not depend on dia. (A) To visualize endocytosis, fluorescently labelled wheat germ agglutinin
 Fig. S1. Endocytosis of extracellular cargo does not depend on dia. (A) To visualize endocytosis, fluorescently labelled wheat germ agglutinin (WGA-Alexa555) was injected into the extracellular perivitteline
Fig. S1. Endocytosis of extracellular cargo does not depend on dia. (A) To visualize endocytosis, fluorescently labelled wheat germ agglutinin (WGA-Alexa555) was injected into the extracellular perivitteline
Supplementary Figure 1. Nature Structural & Molecular Biology: doi: /nsmb.3494
 Supplementary Figure 1 Pol structure-function analysis (a) Inactivating polymerase and helicase mutations do not alter the stability of Pol. Flag epitopes were introduced using CRISPR/Cas9 gene targeting
Supplementary Figure 1 Pol structure-function analysis (a) Inactivating polymerase and helicase mutations do not alter the stability of Pol. Flag epitopes were introduced using CRISPR/Cas9 gene targeting
GFP CCD2 GFP IP:GFP
 D1 D2 1 75 95 148 178 492 GFP CCD1 CCD2 CCD2 GFP D1 D2 GFP D1 D2 Beclin 1 IB:GFP IP:GFP Supplementary Figure 1: Mapping domains required for binding to HEK293T cells are transfected with EGFP-tagged mutant
D1 D2 1 75 95 148 178 492 GFP CCD1 CCD2 CCD2 GFP D1 D2 GFP D1 D2 Beclin 1 IB:GFP IP:GFP Supplementary Figure 1: Mapping domains required for binding to HEK293T cells are transfected with EGFP-tagged mutant
File name: Supplementary Information Description: Supplementary figures and supplementary tables. File name: Peer review file Description:
 File name: Supplementary Information Description: Supplementary figures and supplementary tables. File name: Peer review file Description: Supplementary Figure 1. dcas9-mq1 fusion protein induces de novo
File name: Supplementary Information Description: Supplementary figures and supplementary tables. File name: Peer review file Description: Supplementary Figure 1. dcas9-mq1 fusion protein induces de novo
Nature Biotechnology: doi: /nbt Supplementary Figure 1. In vitro validation of OTC sgrnas and donor template.
 Supplementary Figure 1 In vitro validation of OTC sgrnas and donor template. (a) In vitro validation of sgrnas targeted to OTC in the MC57G mouse cell line by transient transfection followed by 4-day puromycin
Supplementary Figure 1 In vitro validation of OTC sgrnas and donor template. (a) In vitro validation of sgrnas targeted to OTC in the MC57G mouse cell line by transient transfection followed by 4-day puromycin
i-stop codon positions in the mcherry gene
 Supplementary Figure 1 i-stop codon positions in the mcherry gene The grnas (green) that can potentially generate stop codons from Trp (63 th and 98 th aa, upper panel) and Gln (47 th and 114 th aa, bottom
Supplementary Figure 1 i-stop codon positions in the mcherry gene The grnas (green) that can potentially generate stop codons from Trp (63 th and 98 th aa, upper panel) and Gln (47 th and 114 th aa, bottom
Supplementary Figure 1. Espn-1 knockout characterization. (a) The predicted recombinant Espn-1 -/- allele was detected by PCR of the left (5 )
 Supplementary Figure 1. Espn-1 knockout characterization. (a) The predicted recombinant Espn-1 -/- allele was detected by PCR of the left (5 ) homologous recombination arm (left) and of the right (3 )
Supplementary Figure 1. Espn-1 knockout characterization. (a) The predicted recombinant Espn-1 -/- allele was detected by PCR of the left (5 ) homologous recombination arm (left) and of the right (3 )
Allele-specific locus binding and genome editing by CRISPR at the
 Supplementary Information Allele-specific locus binding and genome editing by CRISPR at the p6ink4a locus Toshitsugu Fujita, Miyuki Yuno, and Hodaka Fujii Supplementary Figure Legends Supplementary Figure
Supplementary Information Allele-specific locus binding and genome editing by CRISPR at the p6ink4a locus Toshitsugu Fujita, Miyuki Yuno, and Hodaka Fujii Supplementary Figure Legends Supplementary Figure
Figure legends for supplement
 Figure legends for supplement Supplemental Figure 1 Characterization of purified and recombinant proteins Relevant fractions related the final stage of the purification protocol(bingham et al., 1998; Toba
Figure legends for supplement Supplemental Figure 1 Characterization of purified and recombinant proteins Relevant fractions related the final stage of the purification protocol(bingham et al., 1998; Toba
supplementary information
 DOI: 10.1038/ncb2172 Figure S1 p53 regulates cellular NADPH and lipid levels via inhibition of G6PD. (a) U2OS cells stably expressing p53 shrna or a control shrna were transfected with control sirna or
DOI: 10.1038/ncb2172 Figure S1 p53 regulates cellular NADPH and lipid levels via inhibition of G6PD. (a) U2OS cells stably expressing p53 shrna or a control shrna were transfected with control sirna or
Supplementary Figure S1. Binding of HSA mutants to hfcrn. (a) The levels of titrated amounts of HSA
 Supplementary Figure S1. Binding of HSA mutants to hfcrn. (a) The levels of titrated amounts of HSA variants (5.0-0.002 μg/ml) directly coated in the wells at ph 6.0 were controlled using a horseradish
Supplementary Figure S1. Binding of HSA mutants to hfcrn. (a) The levels of titrated amounts of HSA variants (5.0-0.002 μg/ml) directly coated in the wells at ph 6.0 were controlled using a horseradish
Supplementary Figure 1. jmj30-2 and jmj32-1 produce null mutants. (a) Schematic drawing of JMJ30 and JMJ32 genome structure showing regions amplified
 Supplementary Figure 1. jmj30-2 and jmj32-1 produce null mutants. (a) Schematic drawing of JMJ30 and JMJ32 genome structure showing regions amplified by primers used for mrna expression analysis. Gray
Supplementary Figure 1. jmj30-2 and jmj32-1 produce null mutants. (a) Schematic drawing of JMJ30 and JMJ32 genome structure showing regions amplified by primers used for mrna expression analysis. Gray
CRISPR/Cas9 Gene Editing Tools
 CRISPR/Cas9 Gene Editing Tools - Guide-it Products for Successful CRISPR/Cas9 Gene Editing - Why choose Guide-it products? Optimized methods designed for speed and ease of use Complete kits that don t
CRISPR/Cas9 Gene Editing Tools - Guide-it Products for Successful CRISPR/Cas9 Gene Editing - Why choose Guide-it products? Optimized methods designed for speed and ease of use Complete kits that don t
CRISPR/Cas9 Gene Editing Tools
 CRISPR/Cas9 Gene Editing Tools - Separations Simply Spectacular INDELS Identify indels Determine if one or both copies of your gene have indels The Guide-it Genotype Confirmation Kit: Simple detection
CRISPR/Cas9 Gene Editing Tools - Separations Simply Spectacular INDELS Identify indels Determine if one or both copies of your gene have indels The Guide-it Genotype Confirmation Kit: Simple detection
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/3/146/ra80/dc1 Supplementary Materials for DNMT1 Stability Is Regulated by Proteins Coordinating Deubiquitination and Acetylation-Driven Ubiquitination Zhanwen
www.sciencesignaling.org/cgi/content/full/3/146/ra80/dc1 Supplementary Materials for DNMT1 Stability Is Regulated by Proteins Coordinating Deubiquitination and Acetylation-Driven Ubiquitination Zhanwen
MeCP2. MeCP2/α-tubulin. GFP mir1-1 mir132
 Conservation Figure S1. Schematic showing 3 UTR (top; thick black line), mir132 MRE (arrow) and nucleotide sequence conservation (vertical black lines; http://genome.ucsc.edu). a GFP mir1-1 mir132 b GFP
Conservation Figure S1. Schematic showing 3 UTR (top; thick black line), mir132 MRE (arrow) and nucleotide sequence conservation (vertical black lines; http://genome.ucsc.edu). a GFP mir1-1 mir132 b GFP
(phosphatase tensin) domain is shown in dark gray, the FH1 domain in black, and the
 Supplemental Figure 1. Predicted Domain Organization of the AFH14 Protein. (A) Schematic representation of the predicted domain organization of AFH14. The PTEN (phosphatase tensin) domain is shown in dark
Supplemental Figure 1. Predicted Domain Organization of the AFH14 Protein. (A) Schematic representation of the predicted domain organization of AFH14. The PTEN (phosphatase tensin) domain is shown in dark
Supplementary Materials. China
 Supplementary Materials An Efficient Genotyping Method for Genome-modified Animals and Human Cells Generated with CRISPR/Cas9 System Xiaoxiao Zhu 1,2 *, Yajie Xu 1 *, Shanshan Yu 1 *, Lu Lu 1,2, Mingqin
Supplementary Materials An Efficient Genotyping Method for Genome-modified Animals and Human Cells Generated with CRISPR/Cas9 System Xiaoxiao Zhu 1,2 *, Yajie Xu 1 *, Shanshan Yu 1 *, Lu Lu 1,2, Mingqin
SUPPLEMENTAL FIGURES AND TABLES
 SUPPLEMENTAL FIGURES AND TABLES A B Flag-ALDH1A1 IP: α-ac HEK293T WT 91R 128R 252Q 367R 41/ 419R 435R 495R 412R C Flag-ALDH1A1 NAM IP: HEK293T + + - + D NAM - + + E Relative ALDH1A1 activity 1..8.6.4.2
SUPPLEMENTAL FIGURES AND TABLES A B Flag-ALDH1A1 IP: α-ac HEK293T WT 91R 128R 252Q 367R 41/ 419R 435R 495R 412R C Flag-ALDH1A1 NAM IP: HEK293T + + - + D NAM - + + E Relative ALDH1A1 activity 1..8.6.4.2
Figure S1. Figure S2. Figure S3 HB Anti-FSP27 (COOH-terminal peptide) Ab. Anti-GST-FSP27(45-127) Ab.
 / 36B4 mrna ratio Figure S1 * 2. 1.6 1.2.8 *.4 control TNFα BRL49653 Figure S2 Su bw AT p iw Anti- (COOH-terminal peptide) Ab Blot : Anti-GST-(45-127) Ab β-actin Figure S3 HB2 HW AT BA T Figure S4 A TAG
/ 36B4 mrna ratio Figure S1 * 2. 1.6 1.2.8 *.4 control TNFα BRL49653 Figure S2 Su bw AT p iw Anti- (COOH-terminal peptide) Ab Blot : Anti-GST-(45-127) Ab β-actin Figure S3 HB2 HW AT BA T Figure S4 A TAG
T H E J O U R N A L O F C E L L B I O L O G Y
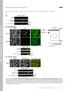 T H E J O U R N A L O F C E L L B I O L O G Y Supplemental material Nakajima and Tanoue, http://www.jcb.org/cgi/content/full/jcb.201104118/dc1 Figure S1. DLD-1 cells exhibit the characteristic morphology
T H E J O U R N A L O F C E L L B I O L O G Y Supplemental material Nakajima and Tanoue, http://www.jcb.org/cgi/content/full/jcb.201104118/dc1 Figure S1. DLD-1 cells exhibit the characteristic morphology
Supplementary Figure 1. Localization of MST1 in RPE cells. Proliferating or ciliated HA- MST1 expressing RPE cells (see Fig. 5b for establishment of
 Supplementary Figure 1. Localization of MST1 in RPE cells. Proliferating or ciliated HA- MST1 expressing RPE cells (see Fig. 5b for establishment of the cell line) were immunostained for HA, acetylated
Supplementary Figure 1. Localization of MST1 in RPE cells. Proliferating or ciliated HA- MST1 expressing RPE cells (see Fig. 5b for establishment of the cell line) were immunostained for HA, acetylated
Revised: RG-RV2 by Fukuhara et al.
 Supplemental Figure 1 The generation of Spns2 conditional knockout mice. (A) Schematic representation of the wild type Spns2 locus (Spns2 + ), the targeted allele, the floxed allele (Spns2 f ) and the
Supplemental Figure 1 The generation of Spns2 conditional knockout mice. (A) Schematic representation of the wild type Spns2 locus (Spns2 + ), the targeted allele, the floxed allele (Spns2 f ) and the
Confocal immunofluorescence microscopy
 Confocal immunofluorescence microscopy HL-6 and cells were cultured and cytospun onto glass slides. The cells were double immunofluorescence stained for Mt NPM1 and fibrillarin (nucleolar marker). Briefly,
Confocal immunofluorescence microscopy HL-6 and cells were cultured and cytospun onto glass slides. The cells were double immunofluorescence stained for Mt NPM1 and fibrillarin (nucleolar marker). Briefly,
XactEdit Cas9 Nuclease with NLS User Manual
 XactEdit Cas9 Nuclease with NLS User Manual An RNA-guided recombinant endonuclease for efficient targeted DNA cleavage Catalog Numbers CE1000-50K, CE1000-50, CE1000-250, CE1001-250, CE1001-1000 Table of
XactEdit Cas9 Nuclease with NLS User Manual An RNA-guided recombinant endonuclease for efficient targeted DNA cleavage Catalog Numbers CE1000-50K, CE1000-50, CE1000-250, CE1001-250, CE1001-1000 Table of
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/8/404/ra120/dc1 Supplementary Materials for The subcellular localization and activity of cortactin is regulated by acetylation and interaction with Keap1 Akihiro
www.sciencesignaling.org/cgi/content/full/8/404/ra120/dc1 Supplementary Materials for The subcellular localization and activity of cortactin is regulated by acetylation and interaction with Keap1 Akihiro
Cell proliferation was measured with Cell Counting Kit-8 (Dojindo Laboratories, Kumamoto, Japan).
 1 2 3 4 5 6 7 8 Supplemental Materials and Methods Cell proliferation assay Cell proliferation was measured with Cell Counting Kit-8 (Dojindo Laboratories, Kumamoto, Japan). GCs were plated at 96-well
1 2 3 4 5 6 7 8 Supplemental Materials and Methods Cell proliferation assay Cell proliferation was measured with Cell Counting Kit-8 (Dojindo Laboratories, Kumamoto, Japan). GCs were plated at 96-well
Engineering splicing factors with designed specificities
 nature methods Engineering splicing factors with designed specificities Yang Wang, Cheom-Gil Cheong, Traci M Tanaka Hall & Zefeng Wang Supplementary figures and text: Supplementary Figure 1 Supplementary
nature methods Engineering splicing factors with designed specificities Yang Wang, Cheom-Gil Cheong, Traci M Tanaka Hall & Zefeng Wang Supplementary figures and text: Supplementary Figure 1 Supplementary
Sarker et al. Supplementary Material. Subcellular Fractionation
 Supplementary Material Subcellular Fractionation Transfected 293T cells were harvested with phosphate buffered saline (PBS) and centrifuged at 2000 rpm (500g) for 3 min. The pellet was washed, re-centrifuged
Supplementary Material Subcellular Fractionation Transfected 293T cells were harvested with phosphate buffered saline (PBS) and centrifuged at 2000 rpm (500g) for 3 min. The pellet was washed, re-centrifuged
Supplementary Information
 Supplementary Information Generation of inheritable and transgene clean targeted genome-modified rice in later generations using the CRISPR/Cas9 system. Rong-Fang Xu, Hao Li, Rui-Ying Qin, Juan Li, Chun-Hong
Supplementary Information Generation of inheritable and transgene clean targeted genome-modified rice in later generations using the CRISPR/Cas9 system. Rong-Fang Xu, Hao Li, Rui-Ying Qin, Juan Li, Chun-Hong
Supplementary Figure 1. Quantitative RT-PCR experimental validation of CRISPR/Cas9 and sgrnas expression in HEK293A transfected cells.
 Supplementary Figure 1. Quantitative RT-PCR experimental validation of CRISPR/Cas9 and sgrnas expression in HEK293A transfected cells. HEK293A cells were transfected with the indicated combinations of
Supplementary Figure 1. Quantitative RT-PCR experimental validation of CRISPR/Cas9 and sgrnas expression in HEK293A transfected cells. HEK293A cells were transfected with the indicated combinations of
Simple protocol for gene editing using GenCrisprTM Cas9 nuclease
 Simple protocol for gene editing using GenCrisprTM Cas9 nuclease Contents Protocol Step 1: Choose the target DNA sequence Step 2: Design sgrna Step 3: Preparation for sgrna 3.1 In vitro transcription of
Simple protocol for gene editing using GenCrisprTM Cas9 nuclease Contents Protocol Step 1: Choose the target DNA sequence Step 2: Design sgrna Step 3: Preparation for sgrna 3.1 In vitro transcription of
Xu et al., Supplementary Figures 1-7
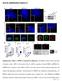 Xu et al., Supplementary Figures 1-7 Supplementary Figure 1. PIPKI is required for ciliogenesis. (a) PIPKI localizes at the basal body of primary cilium. RPE-1 cells treated with two sirnas targeting to
Xu et al., Supplementary Figures 1-7 Supplementary Figure 1. PIPKI is required for ciliogenesis. (a) PIPKI localizes at the basal body of primary cilium. RPE-1 cells treated with two sirnas targeting to
Supplemental Materials and Methods
 Supplemental Materials and Methods Co-immunoprecipitation (Co-IP) assay Cells were lysed with NETN buffer (20 mm Tris-HCl, ph 8.0, 0 mm NaCl, 1 mm EDT, 0.5% Nonidet P-40) containing 50 mm β-glycerophosphate,
Supplemental Materials and Methods Co-immunoprecipitation (Co-IP) assay Cells were lysed with NETN buffer (20 mm Tris-HCl, ph 8.0, 0 mm NaCl, 1 mm EDT, 0.5% Nonidet P-40) containing 50 mm β-glycerophosphate,
Nature Genetics: doi: /ng Supplementary Figure 1
 Supplementary Figure 1 Ihh interacts preferentially with its upstream neighboring gene Nhej1. Genes are indicated by gray lines, and Ihh and Nhej1 are highlighted in blue. 4C seq performed in E14.5 limbs
Supplementary Figure 1 Ihh interacts preferentially with its upstream neighboring gene Nhej1. Genes are indicated by gray lines, and Ihh and Nhej1 are highlighted in blue. 4C seq performed in E14.5 limbs
Integrated Omics Study Delineates the Dynamics of Lipid Droplets in Rhodococcus Opacus PD630
 School of Natural Sciences and Mathematics 2013-10-22 Integrated Omics Study Delineates the Dynamics of Lipid Droplets in Rhodococcus Opacus PD630 UTD AUTHOR(S): Michael Qiwei Zhang 2013 The Authors This
School of Natural Sciences and Mathematics 2013-10-22 Integrated Omics Study Delineates the Dynamics of Lipid Droplets in Rhodococcus Opacus PD630 UTD AUTHOR(S): Michael Qiwei Zhang 2013 The Authors This
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb2271 Supplementary Figure a! WM266.4 mock WM266.4 #7 sirna WM266.4 #10 sirna SKMEL28 mock SKMEL28 #7 sirna SKMEL28 #10 sirna WM1361 mock WM1361 #7 sirna WM1361 #10 sirna 9 WM266. WM136
DOI: 10.1038/ncb2271 Supplementary Figure a! WM266.4 mock WM266.4 #7 sirna WM266.4 #10 sirna SKMEL28 mock SKMEL28 #7 sirna SKMEL28 #10 sirna WM1361 mock WM1361 #7 sirna WM1361 #10 sirna 9 WM266. WM136
HPV E6 oncoprotein targets histone methyltransferases for modulating specific. Chih-Hung Hsu, Kai-Lin Peng, Hua-Ci Jhang, Chia-Hui Lin, Shwu-Yuan Wu,
 1 HPV E oncoprotein targets histone methyltransferases for modulating specific gene transcription 3 5 Chih-Hung Hsu, Kai-Lin Peng, Hua-Ci Jhang, Chia-Hui Lin, Shwu-Yuan Wu, Cheng-Ming Chiang, Sheng-Chung
1 HPV E oncoprotein targets histone methyltransferases for modulating specific gene transcription 3 5 Chih-Hung Hsu, Kai-Lin Peng, Hua-Ci Jhang, Chia-Hui Lin, Shwu-Yuan Wu, Cheng-Ming Chiang, Sheng-Chung
Jung-Nam Cho, Jee-Youn Ryu, Young-Min Jeong, Jihye Park, Ji-Joon Song, Richard M. Amasino, Bosl Noh, and Yoo-Sun Noh
 Developmental Cell, Volume 22 Supplemental Information Control of Seed Germination by Light-Induced Histone Arginine Demethylation Activity Jung-Nam Cho, Jee-Youn Ryu, Young-Min Jeong, Jihye Park, Ji-Joon
Developmental Cell, Volume 22 Supplemental Information Control of Seed Germination by Light-Induced Histone Arginine Demethylation Activity Jung-Nam Cho, Jee-Youn Ryu, Young-Min Jeong, Jihye Park, Ji-Joon
Nature Biotechnology: doi: /nbt Supplementary Figure 1. Design and sequence of system 1 for targeted demethylation.
 Supplementary Figure 1 Design and sequence of system 1 for targeted demethylation. (a) Design of system 1 for targeted demethylation. TET1CD was fused to a catalytic inactive Cas9 nuclease (dcas9) and
Supplementary Figure 1 Design and sequence of system 1 for targeted demethylation. (a) Design of system 1 for targeted demethylation. TET1CD was fused to a catalytic inactive Cas9 nuclease (dcas9) and
Supplementary Figure 1 An overview of pirna biogenesis during fetal mouse reprogramming. (a) (b)
 Supplementary Figure 1 An overview of pirna biogenesis during fetal mouse reprogramming. (a) A schematic overview of the production and amplification of a single pirna from a transposon transcript. The
Supplementary Figure 1 An overview of pirna biogenesis during fetal mouse reprogramming. (a) A schematic overview of the production and amplification of a single pirna from a transposon transcript. The
A Nucleus-Encoded Chloroplast Protein YL1 Is Involved in Chloroplast. Development and Efficient Biogenesis of Chloroplast ATP Synthase in Rice
 A Nucleus-Encoded Chloroplast Protein YL1 Is Involved in Chloroplast Development and Efficient Biogenesis of Chloroplast ATP Synthase in Rice Fei Chen 1,*, Guojun Dong 2,*, Limin Wu 1, Fang Wang 3, Xingzheng
A Nucleus-Encoded Chloroplast Protein YL1 Is Involved in Chloroplast Development and Efficient Biogenesis of Chloroplast ATP Synthase in Rice Fei Chen 1,*, Guojun Dong 2,*, Limin Wu 1, Fang Wang 3, Xingzheng
A novel tool for monitoring endogenous alpha-synuclein transcription by NanoLuciferase
 A novel tool for monitoring endogenous alpha-synuclein transcription by NanoLuciferase tag insertion at the 3 end using CRISPR-Cas9 genome editing technique Sambuddha Basu 1, 3, Levi Adams 1, 3, Subhrangshu
A novel tool for monitoring endogenous alpha-synuclein transcription by NanoLuciferase tag insertion at the 3 end using CRISPR-Cas9 genome editing technique Sambuddha Basu 1, 3, Levi Adams 1, 3, Subhrangshu
Supplementary Figure 1
 Supplementary Figure 1 Product purity of cytosine base editing for the wheat genomic loci tested. Product distributions and indels frequencies at four representative wheat genomic DNA sites in wheat protoplasts
Supplementary Figure 1 Product purity of cytosine base editing for the wheat genomic loci tested. Product distributions and indels frequencies at four representative wheat genomic DNA sites in wheat protoplasts
Supplemental Data. Liu et al. (2013). Plant Cell /tpc
 Supplemental Figure 1. The GFP Tag Does Not Disturb the Physiological Functions of WDL3. (A) RT-PCR analysis of WDL3 expression in wild-type, WDL3-GFP, and WDL3 (without the GFP tag) transgenic seedlings.
Supplemental Figure 1. The GFP Tag Does Not Disturb the Physiological Functions of WDL3. (A) RT-PCR analysis of WDL3 expression in wild-type, WDL3-GFP, and WDL3 (without the GFP tag) transgenic seedlings.
Supplementary Materials for
 www.advances.sciencemag.org/cgi/content/full/1/7/e1500454/dc1 Supplementary Materials for CRISPR-Cas9 delivery to hard-to-transfect cells via membrane deformation Xin Han, Zongbin Liu, Myeong chan Jo,
www.advances.sciencemag.org/cgi/content/full/1/7/e1500454/dc1 Supplementary Materials for CRISPR-Cas9 delivery to hard-to-transfect cells via membrane deformation Xin Han, Zongbin Liu, Myeong chan Jo,
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Endogenous gene tagging to study subcellular localization and chromatin binding. a, b, Schematic of experimental set-up to endogenously tag RNAi factors using the CRISPR Cas9 technology,
Supplementary Figure 1 Endogenous gene tagging to study subcellular localization and chromatin binding. a, b, Schematic of experimental set-up to endogenously tag RNAi factors using the CRISPR Cas9 technology,
Supplementary Information
 Journal : Nature Biotechnology Supplementary Information Targeted genome engineering in human cells with RNA-guided endonucleases Seung Woo Cho, Sojung Kim, Jong Min Kim, and Jin-Soo Kim* National Creative
Journal : Nature Biotechnology Supplementary Information Targeted genome engineering in human cells with RNA-guided endonucleases Seung Woo Cho, Sojung Kim, Jong Min Kim, and Jin-Soo Kim* National Creative
Supplementary Fig. 1
 a FL (1-2266) NL (1-1190) CL (1191-2266) HA-ICE1: - HA-ICE1: - - - FLAG-ICE2: + + + + FLAG-ELL: + + + + + + IP: anti-ha FLAG-ICE2 HA-ICE1-FL HA-ICE1-NL HA-ICE1-CL FLAG-ICE2 b IP: anti-ha FL (1-2266) NL
a FL (1-2266) NL (1-1190) CL (1191-2266) HA-ICE1: - HA-ICE1: - - - FLAG-ICE2: + + + + FLAG-ELL: + + + + + + IP: anti-ha FLAG-ICE2 HA-ICE1-FL HA-ICE1-NL HA-ICE1-CL FLAG-ICE2 b IP: anti-ha FL (1-2266) NL
Supplementary Figure 1. Confirmation of sirna in PC3 and H1299 cells PC3 (a) and H1299 (b) cells were transfected with sirna oligonucleotides
 Supplementary Figure 1. Confirmation of sirna in PC3 and H1299 cells PC3 (a) and H1299 (b) cells were transfected with sirna oligonucleotides targeting RCP (SMARTPool (RCP) or two individual oligos (RCP#1
Supplementary Figure 1. Confirmation of sirna in PC3 and H1299 cells PC3 (a) and H1299 (b) cells were transfected with sirna oligonucleotides targeting RCP (SMARTPool (RCP) or two individual oligos (RCP#1
Supplementary Material
 Supplementary Material Supplementary Methods Cell synchronization. For synchronized cell growth, thymidine was added to 30% confluent U2OS cells to a final concentration of 2.5mM. Cells were incubated
Supplementary Material Supplementary Methods Cell synchronization. For synchronized cell growth, thymidine was added to 30% confluent U2OS cells to a final concentration of 2.5mM. Cells were incubated
Supplementary Information
 Supplementary Information Super-resolution imaging of fluorescently labeled, endogenous RNA Polymerase II in living cells with CRISPR/Cas9-mediated gene editing Won-Ki Cho 1, Namrata Jayanth 1, Susan Mullen
Supplementary Information Super-resolution imaging of fluorescently labeled, endogenous RNA Polymerase II in living cells with CRISPR/Cas9-mediated gene editing Won-Ki Cho 1, Namrata Jayanth 1, Susan Mullen
SUPPLEMENTARY INFORMATION. Small molecule activation of the TRAIL receptor DR5 in human cancer cells
 SUPPLEMENTARY INFORMATION Small molecule activation of the TRAIL receptor DR5 in human cancer cells Gelin Wang 1*, Xiaoming Wang 2, Hong Yu 1, Shuguang Wei 1, Noelle Williams 1, Daniel L. Holmes 1, Randal
SUPPLEMENTARY INFORMATION Small molecule activation of the TRAIL receptor DR5 in human cancer cells Gelin Wang 1*, Xiaoming Wang 2, Hong Yu 1, Shuguang Wei 1, Noelle Williams 1, Daniel L. Holmes 1, Randal
Supplementary Materials
 Supplementary Materials Supplementary Figure 1. PKM2 interacts with MLC2 in cytokinesis. a, U87, U87/EGFRvIII, and HeLa cells in cytokinesis were immunostained with DAPI and an anti-pkm2 antibody. Thirty
Supplementary Materials Supplementary Figure 1. PKM2 interacts with MLC2 in cytokinesis. a, U87, U87/EGFRvIII, and HeLa cells in cytokinesis were immunostained with DAPI and an anti-pkm2 antibody. Thirty
Supplementary Figure 1. APP cleavage assay. HEK293 cells were transfected with various
 Supplementary Figure 1. APP cleavage assay. HEK293 cells were transfected with various GST-tagged N-terminal truncated APP fragments including GST-APP full-length (FL), APP (123-695), APP (189-695), or
Supplementary Figure 1. APP cleavage assay. HEK293 cells were transfected with various GST-tagged N-terminal truncated APP fragments including GST-APP full-length (FL), APP (123-695), APP (189-695), or
At E17.5, the embryos were rinsed in phosphate-buffered saline (PBS) and immersed in
 Supplementary Materials and Methods Barrier function assays At E17.5, the embryos were rinsed in phosphate-buffered saline (PBS) and immersed in acidic X-gal mix (100 mm phosphate buffer at ph4.3, 3 mm
Supplementary Materials and Methods Barrier function assays At E17.5, the embryos were rinsed in phosphate-buffered saline (PBS) and immersed in acidic X-gal mix (100 mm phosphate buffer at ph4.3, 3 mm
B. ADM: C. D. Apoptosis: 1.68% 2.99% 1.31% Figure.S1,Li et al. number. invaded cells. HuH7 BxPC-3 DLD-1.
 A. - Figure.S1,Li et al. B. : - + - + - + E-cadherin CK19 α-sma vimentin β -actin C. D. Apoptosis: 1.68% 2.99% 1.31% - : - + - + - + Apoptosis: 48.33% 45.32% 44.59% E. invaded cells number 400 300 200
A. - Figure.S1,Li et al. B. : - + - + - + E-cadherin CK19 α-sma vimentin β -actin C. D. Apoptosis: 1.68% 2.99% 1.31% - : - + - + - + Apoptosis: 48.33% 45.32% 44.59% E. invaded cells number 400 300 200
T H E J O U R N A L O F C E L L B I O L O G Y
 Supplemental material Wang et al., http://www.jcb.org/cgi/content/full/jcb.201405026/dc1 T H E J O U R N A L O F C E L L B I O L O G Y Figure S1. Generation and characterization of unc-40 alleles. (A and
Supplemental material Wang et al., http://www.jcb.org/cgi/content/full/jcb.201405026/dc1 T H E J O U R N A L O F C E L L B I O L O G Y Figure S1. Generation and characterization of unc-40 alleles. (A and
S156AT168AY175A (AAA) were purified as GST-fusion proteins and incubated with GSTfused
 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 Supplemental Materials Supplemental Figure S1 (a) Phenotype of the wild type and grik1-2 grik2-1 plants after 8 days in darkness.
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 Supplemental Materials Supplemental Figure S1 (a) Phenotype of the wild type and grik1-2 grik2-1 plants after 8 days in darkness.
Supplementary
 Supplementary information Supplementary Material and Methods Plasmid construction The transposable element vectors for inducible expression of RFP-FUS wt and EGFP-FUS R521C and EGFP-FUS P525L were derived
Supplementary information Supplementary Material and Methods Plasmid construction The transposable element vectors for inducible expression of RFP-FUS wt and EGFP-FUS R521C and EGFP-FUS P525L were derived
Supplemental Figure 1 HDA18 has an HDAC domain and therefore has concentration dependent and TSA inhibited histone deacetylase activity.
 Supplemental Figure 1 HDA18 has an HDAC domain and therefore has concentration dependent and TSA inhibited histone deacetylase activity. (A) Amino acid alignment of HDA5, HDA15 and HDA18. The blue line
Supplemental Figure 1 HDA18 has an HDAC domain and therefore has concentration dependent and TSA inhibited histone deacetylase activity. (A) Amino acid alignment of HDA5, HDA15 and HDA18. The blue line
MEFs were treated with the indicated concentrations of LLOMe for three hours, washed
 Supplementary Materials and Methods Cell Fractionation MEFs were treated with the indicated concentrations of LLOMe for three hours, washed with ice-cold PBS, collected by centrifugation, and then homogenized
Supplementary Materials and Methods Cell Fractionation MEFs were treated with the indicated concentrations of LLOMe for three hours, washed with ice-cold PBS, collected by centrifugation, and then homogenized
Supplemental Table 1. Sequences of the primers used for RT-PCR and. Sequence (5-prime to 3-prime)
 Supplemental Table 1. Sequences of the primers used for RT-PCR and qrt-pcr analysis Gene Primer Sequence (5-prime to 3-prime) fdx1 F agtcaaaccagagcgtttcc R cagctacaggcatatctgcaa fdx1b F gagcagcgtatttgtcacaga
Supplemental Table 1. Sequences of the primers used for RT-PCR and qrt-pcr analysis Gene Primer Sequence (5-prime to 3-prime) fdx1 F agtcaaaccagagcgtttcc R cagctacaggcatatctgcaa fdx1b F gagcagcgtatttgtcacaga
Supplemental Data. Lee et al. Plant Cell. (2010) /tpc Supplemental Figure 1. Protein and Gene Structures of DWA1 and DWA2.
 Supplemental Figure 1. Protein and Gene Structures of DWA1 and DWA2. (A) Protein structures of DWA1 and DWA2. WD40 region was determined based on the NCBI conserved domain databases (B, C) Schematic representation
Supplemental Figure 1. Protein and Gene Structures of DWA1 and DWA2. (A) Protein structures of DWA1 and DWA2. WD40 region was determined based on the NCBI conserved domain databases (B, C) Schematic representation
Supplementary Figure 1. Reintroduction of HDAC1 in HDAC1-/- ES cells reverts the
 SUPPLEMENTARY FIGURE LEGENDS Supplementary Figure 1. Reintroduction of HDAC1 in HDAC1-/- ES cells reverts the phenotype of HDAC1-/- teratomas. 3x10 6 HDAC1 reintroduced (HDAC1-/-re) and empty vector infected
SUPPLEMENTARY FIGURE LEGENDS Supplementary Figure 1. Reintroduction of HDAC1 in HDAC1-/- ES cells reverts the phenotype of HDAC1-/- teratomas. 3x10 6 HDAC1 reintroduced (HDAC1-/-re) and empty vector infected
Supplementary Fig. 1 Proteomic analysis of ATR-interacting proteins. ATR, ARID1A and
 Supplementary Figure Legend: Supplementary Fig. 1 Proteomic analysis of ATR-interacting proteins. ATR, ARID1A and ATRIP protein peptides identified from our mass spectrum analysis were shown. Supplementary
Supplementary Figure Legend: Supplementary Fig. 1 Proteomic analysis of ATR-interacting proteins. ATR, ARID1A and ATRIP protein peptides identified from our mass spectrum analysis were shown. Supplementary
Supplementary Materials and Methods
 Supplementary Materials and Methods sirna sequences used in this study The sequences of Stealth Select RNAi for ALK and FLOT-1 were as follows: ALK sense no.1 (ALK): 5 -AAUACUGACAGCCACAGGCAAUGUC-3 ; ALK
Supplementary Materials and Methods sirna sequences used in this study The sequences of Stealth Select RNAi for ALK and FLOT-1 were as follows: ALK sense no.1 (ALK): 5 -AAUACUGACAGCCACAGGCAAUGUC-3 ; ALK
USP19 modulates autophagy and antiviral immune responses by. deubiquitinating Beclin-1
 USP19 modulates autophagy and antiviral immune responses by deubiquitinating Beclin-1 Shouheng Jin 1,2,, Shuo Tian 1,, Yamei Chen 1,, Chuanxia Zhang 1,2, Weihong Xie, 1 Xiaojun Xia 3,4, Jun Cui 1,3* &
USP19 modulates autophagy and antiviral immune responses by deubiquitinating Beclin-1 Shouheng Jin 1,2,, Shuo Tian 1,, Yamei Chen 1,, Chuanxia Zhang 1,2, Weihong Xie, 1 Xiaojun Xia 3,4, Jun Cui 1,3* &
Nature Immunology: doi: /ni Supplementary Figure 1
 Supplementary Figure 1 Sox2 localizes in neutrophils of mouse spleen. (a) Microscopy analysis of Sox2 and MPO in wild-type mouse spleen. Nucl, nucleus. (b) Immunohistochemistry staining of Sox2 and MPO
Supplementary Figure 1 Sox2 localizes in neutrophils of mouse spleen. (a) Microscopy analysis of Sox2 and MPO in wild-type mouse spleen. Nucl, nucleus. (b) Immunohistochemistry staining of Sox2 and MPO
SUPPLEMENTARY INFORMATION
 Ca 2+ /calmodulin Regulates Salicylic Acid-mediated Plant Immunity Liqun Du, Gul S. Ali, Kayla A. Simons, Jingguo Hou, Tianbao Yang, A.S.N. Reddy and B. W. Poovaiah * *To whom correspondence should be
Ca 2+ /calmodulin Regulates Salicylic Acid-mediated Plant Immunity Liqun Du, Gul S. Ali, Kayla A. Simons, Jingguo Hou, Tianbao Yang, A.S.N. Reddy and B. W. Poovaiah * *To whom correspondence should be
Supplementary Figure 1 a
 3 min PMA 45 min PMA AnnexinV-FITC Supplementary Figure 1 5 min PMA 15 min PMA a 9 min PMA 12 min PMA 5 min FGF7 15 min FGF7 3 min FGF7 6 min FGF7 9 min FGF7 12 min FGF7 5 min control 3 min control 6 min
3 min PMA 45 min PMA AnnexinV-FITC Supplementary Figure 1 5 min PMA 15 min PMA a 9 min PMA 12 min PMA 5 min FGF7 15 min FGF7 3 min FGF7 6 min FGF7 9 min FGF7 12 min FGF7 5 min control 3 min control 6 min
pt7ht vector and over-expressed in E. coli as inclusion bodies. Cells were lysed in 6 M
 Supplementary Methods MIG6 production, purification, inhibition, and kinase assays MIG6 segment 1 (30mer, residues 334 364) peptide was synthesized using standard solid-phase peptide synthesis as described
Supplementary Methods MIG6 production, purification, inhibition, and kinase assays MIG6 segment 1 (30mer, residues 334 364) peptide was synthesized using standard solid-phase peptide synthesis as described
Fig. S1. Effect of p120-catenin overexpression on the interaction of SCUBE2 with E-cadherin. The expression plasmid encoding FLAG.
 Fig. S1. Effect of p120-catenin overexpression on the interaction of SCUBE2 with E-cadherin. The expression plasmid encoding FLAG.SCUBE2, E-cadherin.Myc, or HA.p120-catenin was transfected in a combination
Fig. S1. Effect of p120-catenin overexpression on the interaction of SCUBE2 with E-cadherin. The expression plasmid encoding FLAG.SCUBE2, E-cadherin.Myc, or HA.p120-catenin was transfected in a combination
Supplemental Information
 Supplemental Information ATP-dependent unwinding of U4/U6 snrnas by the Brr2 helicase requires the C-terminus of Prp8 Corina Maeder 1,3, Alan K. Kutach 1,2,3, and Christine Guthrie 1 1 Department of Biochemistry
Supplemental Information ATP-dependent unwinding of U4/U6 snrnas by the Brr2 helicase requires the C-terminus of Prp8 Corina Maeder 1,3, Alan K. Kutach 1,2,3, and Christine Guthrie 1 1 Department of Biochemistry
M X 500 µl. M X 1000 µl
 GeneGlide TM sirna Transfection Reagent (Catalog # M1081-300, -500, -1000; Store at 4 C) I. Introduction: BioVision s GeneGlide TM sirna Transfection reagent is a cationic proprietary polymer/lipid formulation,
GeneGlide TM sirna Transfection Reagent (Catalog # M1081-300, -500, -1000; Store at 4 C) I. Introduction: BioVision s GeneGlide TM sirna Transfection reagent is a cationic proprietary polymer/lipid formulation,
Improving CRISPR-Cas9 Gene Knockout with a Validated Guide RNA Algorithm
 Improving CRISPR-Cas9 Gene Knockout with a Validated Guide RNA Algorithm Anja Smith Director R&D Dharmacon, part of GE Healthcare Imagination at work crrna:tracrrna program Cas9 nuclease Active crrna is
Improving CRISPR-Cas9 Gene Knockout with a Validated Guide RNA Algorithm Anja Smith Director R&D Dharmacon, part of GE Healthcare Imagination at work crrna:tracrrna program Cas9 nuclease Active crrna is
Supplemental Data. Wu et al. (2). Plant Cell..5/tpc RGLG Hormonal treatment H2O B RGLG µm ABA µm ACC µm GA Time (hours) µm µm MJ µm IA
 Supplemental Data. Wu et al. (2). Plant Cell..5/tpc..4. A B Supplemental Figure. Immunoblot analysis verifies the expression of the AD-PP2C and BD-RGLG proteins in the Y2H assay. Total proteins were extracted
Supplemental Data. Wu et al. (2). Plant Cell..5/tpc..4. A B Supplemental Figure. Immunoblot analysis verifies the expression of the AD-PP2C and BD-RGLG proteins in the Y2H assay. Total proteins were extracted
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb3209 Supplementary Figure 1 IR induces the association of FH with chromatin. a, U2OS cells synchronized by thymidine double block (2 mm) underwent no release (G1 phase) or release for 2
DOI: 10.1038/ncb3209 Supplementary Figure 1 IR induces the association of FH with chromatin. a, U2OS cells synchronized by thymidine double block (2 mm) underwent no release (G1 phase) or release for 2
DDA3 associates with microtubule plus ends and orchestrates microtubule dynamics and directional cell migration
 DDA3 associates with microtubule plus ends and orchestrates microtubule dynamics and directional cell migration Liangyu Zhang 1,2,4, Hengyi Shao 1,4, Tongge Zhu, Peng Xia 1, Zhikai Wang 1,2, Lifang Liu
DDA3 associates with microtubule plus ends and orchestrates microtubule dynamics and directional cell migration Liangyu Zhang 1,2,4, Hengyi Shao 1,4, Tongge Zhu, Peng Xia 1, Zhikai Wang 1,2, Lifang Liu
Supplementary Information. Optimization of the production of knock-in alleles by CRISPR/Cas9 microinjection into the mouse zygote
 Supplementary Information Supplementary Figures S1 to S5 and Tables S1 to S3. Optimization of the production of knock-in alleles by CRISPR/Cas9 microinjection into the mouse zygote Aurélien Raveux, Sandrine
Supplementary Information Supplementary Figures S1 to S5 and Tables S1 to S3. Optimization of the production of knock-in alleles by CRISPR/Cas9 microinjection into the mouse zygote Aurélien Raveux, Sandrine
p53 increases MHC class I expression by upregulating the endoplasmic reticulum aminopeptidase ERAP1
 Supplementary Information p53 increases MHC class I expression by upregulating the endoplasmic reticulum aminopeptidase ERAP1 Bei Wang 1, Dandan Niu 1, Liyun Lai 1, & Ee Chee Ren 1,2 1 Singapore Immunology
Supplementary Information p53 increases MHC class I expression by upregulating the endoplasmic reticulum aminopeptidase ERAP1 Bei Wang 1, Dandan Niu 1, Liyun Lai 1, & Ee Chee Ren 1,2 1 Singapore Immunology
p Kinesin light chain Kinesin heavy chain Kinesin heavy chain Kinesin light chain + - BCMA
 p.959-970 Kinesin light chain Kinesin heavy chain Kinesin heavy chain Kinesin light chain + - BCMA - 2011-05 1 tetratricopeptide repeat Zona di possibile interazione con cargo BCMA - 2011-05 2 Verhey et
p.959-970 Kinesin light chain Kinesin heavy chain Kinesin heavy chain Kinesin light chain + - BCMA - 2011-05 1 tetratricopeptide repeat Zona di possibile interazione con cargo BCMA - 2011-05 2 Verhey et
Supplementary Information
 Supplementary Information stability is regulated by CK2-dependent interaction with R2TP complex Patrick von Morgen 1,2, Kamila Burdova 1, Thomas G. Flower 3, Nicola J. O'Reilly 4, Simon J. Boulton 5, Stephen
Supplementary Information stability is regulated by CK2-dependent interaction with R2TP complex Patrick von Morgen 1,2, Kamila Burdova 1, Thomas G. Flower 3, Nicola J. O'Reilly 4, Simon J. Boulton 5, Stephen
Zhang et al., RepID facilitates replication Initiation. Supplemental Information:
 Supplemental Information: a b 1 Supplementary Figure 1 (a) DNA sequence of all the oligonucleotides used in this study. Only one strand is shown. The unshaded nucleotide sequences show changes from the
Supplemental Information: a b 1 Supplementary Figure 1 (a) DNA sequence of all the oligonucleotides used in this study. Only one strand is shown. The unshaded nucleotide sequences show changes from the
OmicsLink shrna Clones guaranteed knockdown even in difficult-to-transfect cells
 OmicsLink shrna Clones guaranteed knockdown even in difficult-to-transfect cells OmicsLink shrna clone collections consist of lentiviral, and other mammalian expression vector based small hairpin RNA (shrna)
OmicsLink shrna Clones guaranteed knockdown even in difficult-to-transfect cells OmicsLink shrna clone collections consist of lentiviral, and other mammalian expression vector based small hairpin RNA (shrna)
Khaled_Fig. S MITF TYROSINASE R²= MITF PDE4D R²= Variance from mean mrna expression
 Khaled_Fig. S Variance from mean mrna expression.8 2 MITF.6 TYROSINASE.4 R²=.73.2.8.6.4.2.8.6.4.2.8.6.4.2 MALME3M SKMEL28 UACC257 MITF PDE4D R²=.63 4.5 5 MITF 3.5 4 PDE4B 2.5 3 R²=.2.5 2.5.8.6.4.2.8.6.4.2
Khaled_Fig. S Variance from mean mrna expression.8 2 MITF.6 TYROSINASE.4 R²=.73.2.8.6.4.2.8.6.4.2.8.6.4.2 MALME3M SKMEL28 UACC257 MITF PDE4D R²=.63 4.5 5 MITF 3.5 4 PDE4B 2.5 3 R²=.2.5 2.5.8.6.4.2.8.6.4.2
Schematic representation of the endogenous PALB2 locus and gene-disruption constructs
 Supplementary Figures Supplementary Figure 1. Generation of PALB2 -/- and BRCA2 -/- /PALB2 -/- DT40 cells. (A) Schematic representation of the endogenous PALB2 locus and gene-disruption constructs carrying
Supplementary Figures Supplementary Figure 1. Generation of PALB2 -/- and BRCA2 -/- /PALB2 -/- DT40 cells. (A) Schematic representation of the endogenous PALB2 locus and gene-disruption constructs carrying
pdsipher and pdsipher -GFP shrna Vector User s Guide
 pdsipher and pdsipher -GFP shrna Vector User s Guide NOTE: PLEASE READ THE ENTIRE PROTOCOL CAREFULLY BEFORE USE Page 1. Introduction... 1 2. Vector Overview... 1 3. Vector Maps 2 4. Materials Provided...
pdsipher and pdsipher -GFP shrna Vector User s Guide NOTE: PLEASE READ THE ENTIRE PROTOCOL CAREFULLY BEFORE USE Page 1. Introduction... 1 2. Vector Overview... 1 3. Vector Maps 2 4. Materials Provided...
