Table S1. Primers used in RT-PCR studies (all in 5 to 3 direction)
|
|
|
- Scarlett Gordon
- 6 years ago
- Views:
Transcription
1 Table S1. Primers used in RT-PCR studies (all in 5 to 3 direction) Epo Fw CTGTATCATGGACCACCTCGG Epo Rw TGAAGCACAGAAGCTCTTCGG Jak2 Fw ATCTGACCTTTCCATCTGGGG Jak2 Rw TGGTTGGGTGGATACCAGATC Stat5A Fw TTACTGAAGATCAAGCTGGGG Stat5A Rw TCATTGTACAGAATGTGCCGG Stat5B Fw CATTTTCCCATTGAGGTGCG Stat5B Rw GGGTGGCCTTAATGTTCTCC GAPDH Fw ACGGATTTGGTCGTATTGGG GAPDH Rw TGATTTTGGAGGGATCTCGC ALAS-E Fw CAACATCTCAGGCACCAGTA ALAS-E Rw CTCCACTGTTACGGATACCT Fech Fw ATCCAGCAGCTGGAGGGTCT Fech Rw TGAATCTTGGGGGTTCGGCG GATA1 Fw AGTTTGTGGATCCTGCTCTG GATA1 Rw GCAATGGGTACACCTGAAAG GATA2 Fw AGCGTCTCCAGCCTCATCTTCCGCG GATA2 Rw CGAGTCTTGCTGCGCCTGCTT EKLF Fw CCTGTTTGGTGGTCTCTTCACA EKLF Rw AGGGTCCATTCGTGGGAAA NF-E2p45 Fw AGTGTCAGCTCAGGCTCAGC NF-E2p45 Rw GCAGCTCGGTGATGGACATG p53 Fw CTGTCATCTTCTGTCCCTTC p53 Rw TGGAATCAACCCACAGCTGCA p21 Fw GTCACTGTCTTGTACCCTTGTG p21 Rw CGGCGTTTGGAGTGGTAGAAA p27 Rw AAGCGACCTGCAACCGACGATTCTT p27 Fw GCTCCACAGAACCGGCATTT alpha globin Fw GAGGCCCTGGAGAGGATGTTCC alpha globin Rw ACAGCGCGTTGGGCATGTCGTC beta globin Fw TACATTTGCTTCTGACACAAC beta globin Rw ACAGATCCCCAAAGGAC gamma globin Fw CTTCAAGCTCCTGGGAAATGT gamma globin Rw GCAGAATAAAGCCTACCTTGAAAG epsilon globin Fw GCCTGTGGAGCAAGATGAAT epsilon globin Rw GCGGGCTTGAGGTTGT SCL/Tal1 Fw GTTCTTTGGGGAGCCGGATG SCL/Tal1 Rw ACATTCTGCTGCCGCCATCG Beta-actin Fw CATCACCATTGGCAATGAGC Beta-actin Rw CATACTCCTGCTTGCTGATC GPIIB Fw CTCCACAACAATGGCCCTGG GPIIB Rw CTTGAGAGGGTTGACAGGAG Bcl-xL Fw GAATGACCACCTAGAGCCTTGG
2 Bcl-xL Rw Bcl-2 Fw Bcl-2 Rw IGF-1 Fw IGF-1 Rw Sox6 Fw Endogenous Sox6 Rw Transduced Sox6 Rw (designed on Flag epitope) SOCS3 Fw SOCS3 Rw TGTTCCCATAGAGTTCCACAAAAG ATGTGTGTGGAGAGCGTCAACC TGAGCAGAGTCTTCAGAGACAGCC ATGCTCTTCAGTTCGTGTGTG GCACTCCCTCTACTTGCGTTC GAGGCAGTTCTTTACTGTGG CCGCCATCTGTCTTCATAC CTTATCGTCGTCATCCTTGTA GGAGACTTCGATTCGGGACC GAAACTTGCTGTGGGTGACC
3 Table S2. Primer used to amplify of human genomic regions, and their cloning strategies SOCS3 promoter: Fw:5 ACGTGTCGACACGTGGTACCGCTCTCCCGAAGCGGCGCC 3 ; Rev:5 ACGTAAGCTTCAAGTCGGAGCCGCCGCGG 3, containing a SalI and HindIII restriction site (underlined), respectively, for further cloning into XhoI-HindIII of the pgl2 luciferase reporter vector (Promega, Fitchburg, WI). SOCS3 enhancer: The following oligonucleotides were annealed and then cloned upstream to SOCS3 promoter (SacI-NheI restriction sites, underlined). Mutations are shown in small letters. WT Fw 5 ACGTGAGCTCACTTGACTTGTGTCAGAGCATTGTAATTTACAAAGCACTTTCTCAT CCATGCTAGCACGT3 WT Rev: 5 ACGTGCTAGCATGGATGAGAAAGTGCTTTGTAAATTACAATGCTCTGACACAAGTC AAGTGAGCTCACGT 3 Mut Fw: 5 ACGTGAGCTCACTTGACTTGTGTCAGAGCAgTGTgATTTgCAggGCACTTTCTCATCC ATGCTAGCACGT 3 Mut Rev: 5 ACGTGCTAGCATGGATGAGAAAGTGCccTGcAAATcACAcTGCTCTGACACAAGTCA AGTGAGCTCACGT 3
4 Table S3. Primers used in chromatin immunoprecipitation Human ε-globin promoter Fw: 5 GTTGCAGATAGATGAGGAGCC 3 ; Rw 5 GTCAAGGCTGACCTGTGTCC 3 Human γ-globin promoter A Fw: 5 CCAAGGTCATGGATCGAGTT 3 ; Rw 5 ACACTGTGACAGCTGGGATG 3 Human γ-globin promoter B Fw 5 AAACGGTCCCTGGCTAAACT 3 ; Rw 5 GCTGAAGGGTGCTTCCTTTT 3 The two primer pairs covering 1kb region upstream to the γ transcriptional start site gave the same result. SOCS3 enhancer Fw 5 CTCTCGGCAGAGGTTTATGG 3 ; Rw 5 TCAAGGAATAGCCCTTGAGG 3 GAPDH locus Fw: 5 CGGAGTCAACGGATTTGGTCGTAT 3 ; Rw: 5 AGCCTTCTCCATGGTGGTGAAGAC 3 EMSA oligonucleotide probes: SOCS3 WT probe: Fw: 5 GTCAGAGCATTGTAATTTACAAAGCACTTTCTC3 Rw: 5 GAGAAAGTGCTTTGTAAATTACAATGCTCTGAC3 SOCS3 Mut probe: Fw: 5 GTCAGAGCAGTGTGATTTGCAGGGCACTTTCTC3 Rev: 5 GAGAAAGTGCCCTGCAAATCACACTGCTCTGAC3 Col2a1 probe Fw: 5 CTGTGAATCGGGCTCTGTATGCGCTTGAGAAAAGCCCCATTCATGAGA3 Rw: 5 TCTCATGAATGGGGCTTTTCTCAAGCGCATACAGAGCCCGATTCACAG3
5 Figure S1 (A) Real Time quantitation of Sox6 overexpression in K562cells, CD34 + -derived primary erythroblasts and HEL cells, relative to GAPDH expression (set equal to 100%). (B) Expression of endogenous Sox6 and of transduced Sox6 (Sox6FLAG) in primary erythroid cells. Histograms show the relative levels of expression (mean+ SEM of at least 3 independent experiments) compared to GAPDH, considered as 100%. Figure S2 (A) Real Time quantitation of erythroid genes expression in K562 cells overexpressing Sox6. The increased expression of erythroid genes globins, heme biosynthesis pathway is accompanied by a downregulation of the megacaryocitic gene GPIIB. Histograms show the fold increase in the expression levels induced by Sox6 (mean+ SEM of at least 3 independent experiments) using GAPDH as internal standard. (B) All globins chains normally expressed in K562 are upregulated in Sox6-K562 cells but the ratio of their expression level change significantly. Upper panel: fold increase of α/ζ (3.68) and γ/ε (2.19) ratios in Sox6-K562 versus EV-K562 (set equal to 1). Lower panel: fold decrease of ε/α and γ/α ratios in Sox6-K562 versus EV-K562 (set equal to 1). The increased γ/ε ratio indicates that ε-globin is less strongly induced than γ, in agreement with the known repressor role of Sox6 on the ε-globin gene. 13 However, ε/α and γ/α ratios are both decreased in Sox6-K562 vs EV-K562, suggesting a Sox6 repressive effect also on γ-globin, although less evident than that on ε (the ε/α ratio is almost 10 fold reduced, while the γ/α ratio is reduced by 5 fold. (C D) ChIP analysis confirms the ability of Sox6 to bind the human ε and γ-globin promoters. (C) Sequence comparison of the double Sox6 binding sites within mouse εy and human ε-globin promoters. Nucleotides positions are relative to the start site. (D) The anti-flag antibody or rabbit IgG were used to immunoprecipitate chromatin from EV-K562 or Sox6-K562 cells. Lanes 1 and 2: input chromatins. Lanes 3 and 4: normal rabbit IgG. Lane 5 and 6: anti-flag antibody (that recognizes the Sox6FLAG transduced protein) Lane 7: water. 40 : PCR cycles number. Figure S3 (A) Sox6 overexpression anticipates and induces the expression of the α, β, and γ but not of the ε globin genes in cord blood derived primary erythroid culture. Real time PCR quantification of globins mrna expression level on samples taken at d8, d10, d12, d14 of the culture. Histograms represent the mean expression level (relative to GAPDH); standard deviations refer to 3 independent experiments. Please note the different scale on the y axis: ε expression is marginal and does not increase upon Sox6 overexpression. (B) The ε/α and γ/α ratios (set equal to1 in untransduced cells) are decreased in Sox6-transduced cells. Figure S4. SOCS3 overexpression in K562 cells (A) a SOCS3-IRES-GFP plasmid was transfected in K562 cells and 48 hours after transfection GFP + cells were FACS sorted and seeded in parallel with GFP cells, and the increased expression of SOCS3 in GFP + cells was tested by Real Time PCR. (B) GFP + K562 (SOCS3- overexpressing) cells stop proliferating with kinetics similar to that observed upon Sox6 overexpression (Fig. 1F), y axis: number of cells, x axis: days after sorting. (C) The increased SOCS3 is not associated with K562 differentiation, as shown by Real-Time PCR on α, γ and ε- globin transcripts.
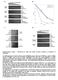
6 Figure S5. Stat5 phosphorylation does not change upon Sox6 overexpression (A) STAT5 phosphorylation was monitored by FACS analysis at 1h, 3h and 6h (and 72h, not shown) after Sox6 transduction in K562, in parallel with GFP expression. x axis: Mean Fluorescence Intensity, y axis: Phospho STAT5. Left panels: EV-K562; Right panels, Sox6- K562. Gray peaks: unstained cells. Black peaks: uninfected K562. Red peaks: GFP cells, not expressing the transduced vectors. Green peaks: GFP + cells, expressing the transduced vector (note that GFP positivity coincides with Sox6 expression thanks to the bicistronic Sox6-IRES- GFP cassette). (B) Semiquantitative PCR reveals non major changes in the level of EPOR, JAK2 and STAT5A/B transcripts in K562 cells overexpressing Sox6. GAPDH is used as internal standard. PCR cycles are indicated below the figure.
7 Supplementary Fig. 1 A Expression relative to GAPDH (%) Sox6FLAG expression Sox6-K562 Sox6-Erythroblasts Sox6-HEL B Expression relative to GAPDH (%) CB-derived Erythroblasts Endogenous 1 Sox6 Sox6FLAG 2
8
9
10
11
SUPPLEMENTARY INFORMATION
 doi:.38/nature899 Supplementary Figure Suzuki et al. a c p7 -/- / WT ratio (+)/(-) p7 -/- / WT ratio Log X 3. Fold change by treatment ( (+)/(-)) Log X.5 3-3. -. b Fold change by treatment ( (+)/(-)) 8
doi:.38/nature899 Supplementary Figure Suzuki et al. a c p7 -/- / WT ratio (+)/(-) p7 -/- / WT ratio Log X 3. Fold change by treatment ( (+)/(-)) Log X.5 3-3. -. b Fold change by treatment ( (+)/(-)) 8
supplementary information
 DOI: 1.138/ncb1839 a b Control 1 2 3 Control 1 2 3 Fbw7 Smad3 1 2 3 4 1 2 3 4 c d IGF-1 IGF-1Rβ IGF-1Rβ-P Control / 1 2 3 4 Real-time RT-PCR Relative quantity (IGF-1/ mrna) 2 1 IGF-1 1 2 3 4 Control /
DOI: 1.138/ncb1839 a b Control 1 2 3 Control 1 2 3 Fbw7 Smad3 1 2 3 4 1 2 3 4 c d IGF-1 IGF-1Rβ IGF-1Rβ-P Control / 1 2 3 4 Real-time RT-PCR Relative quantity (IGF-1/ mrna) 2 1 IGF-1 1 2 3 4 Control /
Supplementary Table 1. The Q-PCR primer sequence is summarized in the following table.
 Supplementary Table 1. The Q-PCR primer sequence is summarized in the following table. Name Sequence (5-3 ) Application Flag-u ggactacaaggacgacgatgac Shared upstream primer for all the amplifications of
Supplementary Table 1. The Q-PCR primer sequence is summarized in the following table. Name Sequence (5-3 ) Application Flag-u ggactacaaggacgacgatgac Shared upstream primer for all the amplifications of
Transcriptional Regulation of Pro-apoptotic Protein Kinase C-delta: Implications for Oxidative Stress-induced Neuronal Cell Death
 SUPPLEMENTAL DATA Transcriptional Regulation of Pro-apoptotic Protein Kinase C-delta: Implications for Oxidative Stress-induced Neuronal Cell Death Huajun Jin 1, Arthi Kanthasamy 1, Vellareddy Anantharam
SUPPLEMENTAL DATA Transcriptional Regulation of Pro-apoptotic Protein Kinase C-delta: Implications for Oxidative Stress-induced Neuronal Cell Death Huajun Jin 1, Arthi Kanthasamy 1, Vellareddy Anantharam
Thyroid peroxidase gene expression is induced by lipopolysaccharide involving Nuclear Factor (NF)-κB p65 subunit phosphorylation
 1 2 3 4 5 SUPPLEMENTAL DATA Thyroid peroxidase gene expression is induced by lipopolysaccharide involving Nuclear Factor (NF)-κB p65 subunit phosphorylation Magalí Nazar, Juan Pablo Nicola, María Laura
1 2 3 4 5 SUPPLEMENTAL DATA Thyroid peroxidase gene expression is induced by lipopolysaccharide involving Nuclear Factor (NF)-κB p65 subunit phosphorylation Magalí Nazar, Juan Pablo Nicola, María Laura
Supplemental figures Supplemental Figure 1: Fluorescence recovery for FRAP experiments depicted in Figure 1.
 Supplemental figures Supplemental Figure 1: Fluorescence recovery for FRAP experiments depicted in Figure 1. Percent of original fluorescence was plotted as a function of time following photobleaching
Supplemental figures Supplemental Figure 1: Fluorescence recovery for FRAP experiments depicted in Figure 1. Percent of original fluorescence was plotted as a function of time following photobleaching
Supplementary information to accompany: A novel role for the DNA repair gene Rad51 in Netrin-1 signalling
 Supplementary information to accompany: A novel role for the DNA repair gene Rad51 in Netrin-1 signalling Glendining KA 1, Markie D 2, Gardner RJM 4, Franz EA 3, Robertson SP 4, Jasoni CL 1 Supplementary
Supplementary information to accompany: A novel role for the DNA repair gene Rad51 in Netrin-1 signalling Glendining KA 1, Markie D 2, Gardner RJM 4, Franz EA 3, Robertson SP 4, Jasoni CL 1 Supplementary
Supplementary Figure 1. Isolation of GFPHigh cells.
 Supplementary Figure 1. Isolation of GFP High cells. (A) Schematic diagram of cell isolation based on Wnt signaling activity. Colorectal cancer (CRC) cell lines were stably transduced with lentivirus encoding
Supplementary Figure 1. Isolation of GFP High cells. (A) Schematic diagram of cell isolation based on Wnt signaling activity. Colorectal cancer (CRC) cell lines were stably transduced with lentivirus encoding
Nature Immunology: doi: /ni Supplementary Figure 1. Zranb1 gene targeting.
 Supplementary Figure 1 Zranb1 gene targeting. (a) Schematic picture of Zranb1 gene targeting using an FRT-LoxP vector, showing the first 6 exons of Zranb1 gene (exons 7-9 are not shown). Targeted mice
Supplementary Figure 1 Zranb1 gene targeting. (a) Schematic picture of Zranb1 gene targeting using an FRT-LoxP vector, showing the first 6 exons of Zranb1 gene (exons 7-9 are not shown). Targeted mice
ASPP1 Fw GGTTGGGAATCCACGTGTTG ASPP1 Rv GCCATATCTTGGAGCTCTGAGAG
 Supplemental Materials and Methods Plasmids: the following plasmids were used in the supplementary data: pwzl-myc- Lats2 (Aylon et al, 2006), pretrosuper-vector and pretrosuper-shp53 (generous gift of
Supplemental Materials and Methods Plasmids: the following plasmids were used in the supplementary data: pwzl-myc- Lats2 (Aylon et al, 2006), pretrosuper-vector and pretrosuper-shp53 (generous gift of
RNA-Guided Gene Activation by CRISPR-Cas9-Based Transcription Factors
 Supplementary Information RNA-Guided Gene Activation by CRISPR-Cas9-Based Transcription Factors Pablo Perez-Pinera 1, Daniel D. Kocak 1, Christopher M. Vockley 2,3, Andrew F. Adler 1, Ami M. Kabadi 1,
Supplementary Information RNA-Guided Gene Activation by CRISPR-Cas9-Based Transcription Factors Pablo Perez-Pinera 1, Daniel D. Kocak 1, Christopher M. Vockley 2,3, Andrew F. Adler 1, Ami M. Kabadi 1,
Figure S1: Insert sequences for GR1, GR2, Qc3c_AgeI and GR3 vectors
 Figure S1: Insert sequences for GR1, GR2, Qc3c_AgeI and GR3 vectors A Vector GR1 SacI BamHI CTCTGGCTAACTAGGC Insert 5/7nt - G TCGAGAGACCGATTGATCCG Insert 5/7nt - CCTAG 1 G CAACAACAACAACAACAACAACAACAACAACAACAACAACAACAACAACAACAACAACAACAAC
Figure S1: Insert sequences for GR1, GR2, Qc3c_AgeI and GR3 vectors A Vector GR1 SacI BamHI CTCTGGCTAACTAGGC Insert 5/7nt - G TCGAGAGACCGATTGATCCG Insert 5/7nt - CCTAG 1 G CAACAACAACAACAACAACAACAACAACAACAACAACAACAACAACAACAACAACAACAACAAC
Supplementary Figures
 Supplementary Figures Supplementary Figure 1. Description of the observed lymphatic metastases in two different SIX1-induced MCF7 metastasis models (Nude and NOD/SCID). Supplementary Figure 2. MCF7-SIX1
Supplementary Figures Supplementary Figure 1. Description of the observed lymphatic metastases in two different SIX1-induced MCF7 metastasis models (Nude and NOD/SCID). Supplementary Figure 2. MCF7-SIX1
The lineage-defining factors T-bet and Bcl-6 collaborate to regulate Th1 gene expression patterns
 The lineage-defining factors T-bet and Bcl-6 collaborate to regulate Th1 gene expression patterns Kenneth J. Oestreich, 1 Albert C. Huang, 1,2 and Amy S. Weinmann 1,2 1 Department of Immunology and 2 Molecular
The lineage-defining factors T-bet and Bcl-6 collaborate to regulate Th1 gene expression patterns Kenneth J. Oestreich, 1 Albert C. Huang, 1,2 and Amy S. Weinmann 1,2 1 Department of Immunology and 2 Molecular
A novel two-step genome editing strategy with CRISPR-Cas9 provides new insights into telomerase action and TERT gene expression
 Xi et al. Genome Biology (2015) 16:231 DOI 10.1186/s13059-015-0791-1 RESEARCH A novel two-step genome editing strategy with CRISPR-Cas9 provides new insights into telomerase action and TERT gene expression
Xi et al. Genome Biology (2015) 16:231 DOI 10.1186/s13059-015-0791-1 RESEARCH A novel two-step genome editing strategy with CRISPR-Cas9 provides new insights into telomerase action and TERT gene expression
Supplementary Figure 1
 Supplementary Figure 1 Supplementary Figure 1: Vector maps of TRMPV and TRMPVIR variants. Many derivatives of TRMPV have been generated and tested. Unless otherwise noted, experiments in this paper use
Supplementary Figure 1 Supplementary Figure 1: Vector maps of TRMPV and TRMPVIR variants. Many derivatives of TRMPV have been generated and tested. Unless otherwise noted, experiments in this paper use
Supplemental Data. LMO4 Controls the Balance between Excitatory. and Inhibitory Spinal V2 Interneurons
 Neuron, Volume 61 Supplemental Data LMO4 Controls the Balance between Excitatory and Inhibitory Spinal V2 Interneurons Kaumudi Joshi, Seunghee Lee, Bora Lee, Jae W. Lee, and Soo-Kyung Lee Supplemental
Neuron, Volume 61 Supplemental Data LMO4 Controls the Balance between Excitatory and Inhibitory Spinal V2 Interneurons Kaumudi Joshi, Seunghee Lee, Bora Lee, Jae W. Lee, and Soo-Kyung Lee Supplemental
Alternative Cleavage and Polyadenylation of RNA
 Developmental Cell 18 Supplemental Information The Spen Family Protein FPA Controls Alternative Cleavage and Polyadenylation of RNA Csaba Hornyik, Lionel C. Terzi, and Gordon G. Simpson Figure S1, related
Developmental Cell 18 Supplemental Information The Spen Family Protein FPA Controls Alternative Cleavage and Polyadenylation of RNA Csaba Hornyik, Lionel C. Terzi, and Gordon G. Simpson Figure S1, related
Khaled_Fig. S MITF TYROSINASE R²= MITF PDE4D R²= Variance from mean mrna expression
 Khaled_Fig. S Variance from mean mrna expression.8 2 MITF.6 TYROSINASE.4 R²=.73.2.8.6.4.2.8.6.4.2.8.6.4.2 MALME3M SKMEL28 UACC257 MITF PDE4D R²=.63 4.5 5 MITF 3.5 4 PDE4B 2.5 3 R²=.2.5 2.5.8.6.4.2.8.6.4.2
Khaled_Fig. S Variance from mean mrna expression.8 2 MITF.6 TYROSINASE.4 R²=.73.2.8.6.4.2.8.6.4.2.8.6.4.2 MALME3M SKMEL28 UACC257 MITF PDE4D R²=.63 4.5 5 MITF 3.5 4 PDE4B 2.5 3 R²=.2.5 2.5.8.6.4.2.8.6.4.2
Table S1. Primer sequences
 Table S1. Primer sequences Primers for quantitative PCR Tgf 1 Forward Tgf 1 Reverse Tgf 2Forward Tgf 2Reverse Tgf 3 Forward Tgf 3 Reverse Tgf r1 Forward Tgf r1 Reverse Tgf r2 Forward Tgf r2 Reverse Thbs1
Table S1. Primer sequences Primers for quantitative PCR Tgf 1 Forward Tgf 1 Reverse Tgf 2Forward Tgf 2Reverse Tgf 3 Forward Tgf 3 Reverse Tgf r1 Forward Tgf r1 Reverse Tgf r2 Forward Tgf r2 Reverse Thbs1
Supplemental Table 1 Gene Symbol FDR corrected p-value PLOD1 CSRP2 PFKP ADFP ADM C10orf10 GPI LOX PLEKHA2 WIPF1
 Supplemental Table 1 Gene Symbol FDR corrected p-value PLOD1 4.52E-18 PDK1 6.77E-18 CSRP2 4.42E-17 PFKP 1.23E-14 MSH2 3.79E-13 NARF_A 5.56E-13 ADFP 5.56E-13 FAM13A1 1.56E-12 FAM29A_A 1.22E-11 CA9 1.54E-11
Supplemental Table 1 Gene Symbol FDR corrected p-value PLOD1 4.52E-18 PDK1 6.77E-18 CSRP2 4.42E-17 PFKP 1.23E-14 MSH2 3.79E-13 NARF_A 5.56E-13 ADFP 5.56E-13 FAM13A1 1.56E-12 FAM29A_A 1.22E-11 CA9 1.54E-11
Fig. S1. Nature Medicine: doi: /nm HoxA9 expression levels BM MOZ-TIF2 AML BM. Sca-1-H. c-kit-h CSF1R-H CD16/32-H. Mac1-H.
 A 1 4 1 4 1 4 CSF1RH 1 3 1 2 1 1 CSF1RH 1 3 1 2 1 1 CSF1RH 1 3 1 2 1 1 1 1 1 1 1 2 1 3 1 4 GFPH 1 1 1 1 1 2 1 3 1 4 Sca1H 1 1 1 1 1 2 1 3 1 4 ckith 1 4 1 4 1 4 CSF1RH 1 3 1 2 1 1 CSF1RH 1 3 1 2 1 1 CSF1RH
A 1 4 1 4 1 4 CSF1RH 1 3 1 2 1 1 CSF1RH 1 3 1 2 1 1 CSF1RH 1 3 1 2 1 1 1 1 1 1 1 2 1 3 1 4 GFPH 1 1 1 1 1 2 1 3 1 4 Sca1H 1 1 1 1 1 2 1 3 1 4 ckith 1 4 1 4 1 4 CSF1RH 1 3 1 2 1 1 CSF1RH 1 3 1 2 1 1 CSF1RH
Jung-Nam Cho, Jee-Youn Ryu, Young-Min Jeong, Jihye Park, Ji-Joon Song, Richard M. Amasino, Bosl Noh, and Yoo-Sun Noh
 Developmental Cell, Volume 22 Supplemental Information Control of Seed Germination by Light-Induced Histone Arginine Demethylation Activity Jung-Nam Cho, Jee-Youn Ryu, Young-Min Jeong, Jihye Park, Ji-Joon
Developmental Cell, Volume 22 Supplemental Information Control of Seed Germination by Light-Induced Histone Arginine Demethylation Activity Jung-Nam Cho, Jee-Youn Ryu, Young-Min Jeong, Jihye Park, Ji-Joon
Technical Review. Real time PCR
 Technical Review Real time PCR Normal PCR: Analyze with agarose gel Normal PCR vs Real time PCR Real-time PCR, also known as quantitative PCR (qpcr) or kinetic PCR Key feature: Used to amplify and simultaneously
Technical Review Real time PCR Normal PCR: Analyze with agarose gel Normal PCR vs Real time PCR Real-time PCR, also known as quantitative PCR (qpcr) or kinetic PCR Key feature: Used to amplify and simultaneously
Roche Molecular Biochemicals Technical Note No. LC 9/2000
 Roche Molecular Biochemicals Technical Note No. LC 9/2000 LightCycler Optimization Strategy Introduction Purpose of this Note Table of Contents The LightCycler system provides different detection formats
Roche Molecular Biochemicals Technical Note No. LC 9/2000 LightCycler Optimization Strategy Introduction Purpose of this Note Table of Contents The LightCycler system provides different detection formats
Peter Dy, Weihuan Wang, Pallavi Bhattaram, Qiuqing Wang, Lai Wang, R. Tracy Ballock, and Véronique Lefebvre
 Developmental Cell, Volume 22 Supplemental Information Sox9 Directs Hypertrophic Maturation and Blocks Osteoblast Differentiation of Growth Plate Chondrocytes Peter Dy, Weihuan Wang, Pallavi Bhattaram,
Developmental Cell, Volume 22 Supplemental Information Sox9 Directs Hypertrophic Maturation and Blocks Osteoblast Differentiation of Growth Plate Chondrocytes Peter Dy, Weihuan Wang, Pallavi Bhattaram,
Supplementary Fig. 1 related to Fig. 1 Clinical relevance of lncrna candidate
 Supplementary Figure Legends Supplementary Fig. 1 related to Fig. 1 Clinical relevance of lncrna candidate BC041951 in gastric cancer. (A) The flow chart for selected candidate lncrnas in 660 up-regulated
Supplementary Figure Legends Supplementary Fig. 1 related to Fig. 1 Clinical relevance of lncrna candidate BC041951 in gastric cancer. (A) The flow chart for selected candidate lncrnas in 660 up-regulated
Fatchiyah
 Fatchiyah Email: fatchiya@yahoo.co.id RNAs: mrna trna rrna RNAi DNAs: Protein: genome DNA cdna mikro-makro mono-poly single-multi Analysis: Identification human and animal disease Finger printing Sexing
Fatchiyah Email: fatchiya@yahoo.co.id RNAs: mrna trna rrna RNAi DNAs: Protein: genome DNA cdna mikro-makro mono-poly single-multi Analysis: Identification human and animal disease Finger printing Sexing
/04/$15.00/0 Molecular Endocrinology 18(3): Copyright 2004 by The Endocrine Society doi: /me
 0888-8809/04/$15.00/0 Molecular Endocrinology 18(3):588 605 Printed in U.S.A. Copyright 2004 by The Endocrine Society doi: 10.1210/me.2003-0090 Increased Cytochrome P450 17 -Hydroxylase Promoter Function
0888-8809/04/$15.00/0 Molecular Endocrinology 18(3):588 605 Printed in U.S.A. Copyright 2004 by The Endocrine Society doi: 10.1210/me.2003-0090 Increased Cytochrome P450 17 -Hydroxylase Promoter Function
Int. J. Mol. Sci. 2016, 17, 1259; doi: /ijms
 S1 of S5 Supplementary Materials: Fibroblast-Derived Extracellular Matrix Induces Chondrogenic Differentiation in Human Adipose-Derived Mesenchymal Stromal/Stem Cells in Vitro Kevin Dzobo, Taegyn Turnley,
S1 of S5 Supplementary Materials: Fibroblast-Derived Extracellular Matrix Induces Chondrogenic Differentiation in Human Adipose-Derived Mesenchymal Stromal/Stem Cells in Vitro Kevin Dzobo, Taegyn Turnley,
mir-24-mediated down-regulation of H2AX suppresses DNA repair
 Supplemental Online Material mir-24-mediated down-regulation of H2AX suppresses DNA repair in terminally differentiated blood cells Ashish Lal 1,4, Yunfeng Pan 2,4, Francisco Navarro 1,4, Derek M. Dykxhoorn
Supplemental Online Material mir-24-mediated down-regulation of H2AX suppresses DNA repair in terminally differentiated blood cells Ashish Lal 1,4, Yunfeng Pan 2,4, Francisco Navarro 1,4, Derek M. Dykxhoorn
Online Supplementary Information
 Online Supplementary Information NLRP4 negatively regulates type I interferon signaling by targeting TBK1 for degradation via E3 ubiquitin ligase DTX4 Jun Cui 1,4,6,7, Yinyin Li 1,5,6,7, Liang Zhu 1, Dan
Online Supplementary Information NLRP4 negatively regulates type I interferon signaling by targeting TBK1 for degradation via E3 ubiquitin ligase DTX4 Jun Cui 1,4,6,7, Yinyin Li 1,5,6,7, Liang Zhu 1, Dan
Supporting Information
 Supporting Information Table S1. Overview of samples used for sequencing, and the number of sequences obtained from each sample. Visit 1 is day 0, Visit 2 is day 7, Visit 3 is day 28, and Visit 4 is day
Supporting Information Table S1. Overview of samples used for sequencing, and the number of sequences obtained from each sample. Visit 1 is day 0, Visit 2 is day 7, Visit 3 is day 28, and Visit 4 is day
IGF-I is a polypeptide hormone that plays important roles
 ORIGINAL RESEARCH Growth Hormone-Activated STAT5 May Indirectly Stimulate IGF-I Gene Transcription through HNF-3 Satyanarayana Eleswarapu, Xiaomei Ge, Ying Wang, Jie Yu, and Honglin Jiang Department of
ORIGINAL RESEARCH Growth Hormone-Activated STAT5 May Indirectly Stimulate IGF-I Gene Transcription through HNF-3 Satyanarayana Eleswarapu, Xiaomei Ge, Ying Wang, Jie Yu, and Honglin Jiang Department of
Ascl2 Acts as an R-spondin/Wnt-Responsive Switch to Control Stemness in Intestinal Crypts
 Cell Stem Cell Supplemental Information Ascl2 Acts as an R-spondin/Wnt-Responsive Switch to Control Stemness in Intestinal Crypts Jurian Schuijers, Jan Philipp Junker, Michal Mokry, Pantelis Hatzis, Bon-Kyoung
Cell Stem Cell Supplemental Information Ascl2 Acts as an R-spondin/Wnt-Responsive Switch to Control Stemness in Intestinal Crypts Jurian Schuijers, Jan Philipp Junker, Michal Mokry, Pantelis Hatzis, Bon-Kyoung
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb2774 Figure S1 TRF2 dosage modulates the tumorigenicity of mouse and human tumor cells. (a) Left: immunoblotting with antibodies directed against the Myc tag of the transduced TRF2 forms
DOI: 10.1038/ncb2774 Figure S1 TRF2 dosage modulates the tumorigenicity of mouse and human tumor cells. (a) Left: immunoblotting with antibodies directed against the Myc tag of the transduced TRF2 forms
Percent survival. Supplementary fig. S3 A.
 Supplementary fig. S3 A. B. 100 Percent survival 80 60 40 20 Ml 0 0 100 C. Fig. S3 Comparison of leukaemia incidence rate in the triple targeted chimaeric mice and germline-transmission translocator mice
Supplementary fig. S3 A. B. 100 Percent survival 80 60 40 20 Ml 0 0 100 C. Fig. S3 Comparison of leukaemia incidence rate in the triple targeted chimaeric mice and germline-transmission translocator mice
Supplementary Figure 1. RAD51 and RAD51 paralogs are enriched spontaneously onto
 Supplementary Figure legends Supplementary Figure 1. and paralogs are enriched spontaneously onto the S-phase chromatin during DN replication. () Chromatin fractionation was carried out as described in
Supplementary Figure legends Supplementary Figure 1. and paralogs are enriched spontaneously onto the S-phase chromatin during DN replication. () Chromatin fractionation was carried out as described in
Supplemental Information
 Supplemental Information Itemized List Materials and Methods, Related to Supplemental Figures S5A-C and S6. Supplemental Figure S1, Related to Figures 1 and 2. Supplemental Figure S2, Related to Figure
Supplemental Information Itemized List Materials and Methods, Related to Supplemental Figures S5A-C and S6. Supplemental Figure S1, Related to Figures 1 and 2. Supplemental Figure S2, Related to Figure
Contents... vii. List of Figures... xii. List of Tables... xiv. Abbreviatons... xv. Summary... xvii. 1. Introduction In vitro evolution...
 vii Contents Contents... vii List of Figures... xii List of Tables... xiv Abbreviatons... xv Summary... xvii 1. Introduction...1 1.1 In vitro evolution... 1 1.2 Phage Display Technology... 3 1.3 Cell surface
vii Contents Contents... vii List of Figures... xii List of Tables... xiv Abbreviatons... xv Summary... xvii 1. Introduction...1 1.1 In vitro evolution... 1 1.2 Phage Display Technology... 3 1.3 Cell surface
Transcriptional Regulation in Eukaryotes
 Transcriptional Regulation in Eukaryotes Concepts, Strategies, and Techniques Michael Carey Stephen T. Smale COLD SPRING HARBOR LABORATORY PRESS NEW YORK 2000 Cold Spring Harbor Laboratory Press, 0-87969-537-4
Transcriptional Regulation in Eukaryotes Concepts, Strategies, and Techniques Michael Carey Stephen T. Smale COLD SPRING HARBOR LABORATORY PRESS NEW YORK 2000 Cold Spring Harbor Laboratory Press, 0-87969-537-4
Supplementary Figure 1: Expression of RNF8, HERC2 and NEURL4 in the cerebellum and knockdown of RNF8 by RNAi (a) Lysates of the cerebellum from rat
 Supplementary Figure 1: Expression of RNF8, HERC2 and NEURL4 in the cerebellum and knockdown of RNF8 by RNAi (a) Lysates of the cerebellum from rat pups at P6, P14, P22, P30 and adult (A) rats were subjected
Supplementary Figure 1: Expression of RNF8, HERC2 and NEURL4 in the cerebellum and knockdown of RNF8 by RNAi (a) Lysates of the cerebellum from rat pups at P6, P14, P22, P30 and adult (A) rats were subjected
SANTA CRUZ BIOTECHNOLOGY, INC.
 TECHNICAL SERVICE GUIDE: Western Blotting 2. What size bands were expected and what size bands were detected? 3. Was the blot blank or was a dark background or non-specific bands seen? 4. Did this same
TECHNICAL SERVICE GUIDE: Western Blotting 2. What size bands were expected and what size bands were detected? 3. Was the blot blank or was a dark background or non-specific bands seen? 4. Did this same
Supplemental Materials and Methods
 Supplemental Materials and Methods 125 I-CXCL12 binding assay KG1 cells (2 10 6 ) were preincubated on ice with cold CXCL12 (1.6µg/mL corresponding to 200nM), CXCL11 (1.66µg/mL corresponding to 200nM),
Supplemental Materials and Methods 125 I-CXCL12 binding assay KG1 cells (2 10 6 ) were preincubated on ice with cold CXCL12 (1.6µg/mL corresponding to 200nM), CXCL11 (1.66µg/mL corresponding to 200nM),
Pre-made Lentiviral Particles for nuclear permeant CRE recombinase expression
 Pre-made Lentiviral Particles for nuclear permeant CRE recombinase expression Cat# Product Name Amounts* LVP336 NLS-CRE (Bsd) LVP336-PBS NLS-CRE (Bsd), in vivo ready LVP339 NLS-CRE (Puro) LVP339-PBS NLS-CRE
Pre-made Lentiviral Particles for nuclear permeant CRE recombinase expression Cat# Product Name Amounts* LVP336 NLS-CRE (Bsd) LVP336-PBS NLS-CRE (Bsd), in vivo ready LVP339 NLS-CRE (Puro) LVP339-PBS NLS-CRE
Supporting Information
 Supporting Information Liu et al. 10.1073/pnas.0901216106 SI Materials and Methods Reagents. Thioglycolate and LPS from Escherichia coli 0111:B4 were from Sigma-Aldrich. PAM3CSK4 and poly(i:c) were from
Supporting Information Liu et al. 10.1073/pnas.0901216106 SI Materials and Methods Reagents. Thioglycolate and LPS from Escherichia coli 0111:B4 were from Sigma-Aldrich. PAM3CSK4 and poly(i:c) were from
Regulation of Synaptic Structure and Function by FMRP- Associated MicroRNAs mir-125b and mir-132
 Neuron, Volume 65 Regulation of Synaptic Structure and Function by FMRP- Associated MicroRNAs mir-125b and mir-132 Dieter Edbauer, Joel R. Neilson, Kelly A. Foster, Chi-Fong Wang, Daniel P. Seeburg, Matthew
Neuron, Volume 65 Regulation of Synaptic Structure and Function by FMRP- Associated MicroRNAs mir-125b and mir-132 Dieter Edbauer, Joel R. Neilson, Kelly A. Foster, Chi-Fong Wang, Daniel P. Seeburg, Matthew
Supplementary Material. TRIB3 inhibits proliferation and promotes osteogenesis in hbmscs by regulating the. ERK1/2 signaling pathway
 Supplementary Material TRIB3 inhibits proliferation and promotes osteogenesis in hbmscs by regulating the ERK1/2 signaling pathway Cui Zhang 1, Fan-Fan Hong 1, Cui-Cui Wang 1, Liang Li 1, Jian-Ling Chen
Supplementary Material TRIB3 inhibits proliferation and promotes osteogenesis in hbmscs by regulating the ERK1/2 signaling pathway Cui Zhang 1, Fan-Fan Hong 1, Cui-Cui Wang 1, Liang Li 1, Jian-Ling Chen
% Viability. isw2 ino isw2 ino isw2 ino isw2 ino mM HU 4-NQO CPT
 a Drug concentration b 1.3% MMS nhp1 nhp1 8 nhp1 mag1.5% MMS.3% MMS nhp1 nhp1 ino8 9 ino8 9 % Viability 4.5% MMS ino8 9 ino8 9 2.5.1.15 % MMS c d nhp1 nhp1 nhp1 nhp1 nhp1 nhp1 Control (YPD) γ IR (1 gy)
a Drug concentration b 1.3% MMS nhp1 nhp1 8 nhp1 mag1.5% MMS.3% MMS nhp1 nhp1 ino8 9 ino8 9 % Viability 4.5% MMS ino8 9 ino8 9 2.5.1.15 % MMS c d nhp1 nhp1 nhp1 nhp1 nhp1 nhp1 Control (YPD) γ IR (1 gy)
Methods of Biomaterials Testing Lesson 3-5. Biochemical Methods - Molecular Biology -
 Methods of Biomaterials Testing Lesson 3-5 Biochemical Methods - Molecular Biology - Chromosomes in the Cell Nucleus DNA in the Chromosome Deoxyribonucleic Acid (DNA) DNA has double-helix structure The
Methods of Biomaterials Testing Lesson 3-5 Biochemical Methods - Molecular Biology - Chromosomes in the Cell Nucleus DNA in the Chromosome Deoxyribonucleic Acid (DNA) DNA has double-helix structure The
Problem Set 8. Answer Key
 MCB 102 University of California, Berkeley August 11, 2009 Isabelle Philipp Online Document Problem Set 8 Answer Key 1. The Genetic Code (a) Are all amino acids encoded by the same number of codons? no
MCB 102 University of California, Berkeley August 11, 2009 Isabelle Philipp Online Document Problem Set 8 Answer Key 1. The Genetic Code (a) Are all amino acids encoded by the same number of codons? no
Supplementary Figures Montero et al._supplementary Figure 1
 Montero et al_suppl. Info 1 Supplementary Figures Montero et al._supplementary Figure 1 Montero et al_suppl. Info 2 Supplementary Figure 1. Transcripts arising from the structurally conserved subtelomeres
Montero et al_suppl. Info 1 Supplementary Figures Montero et al._supplementary Figure 1 Montero et al_suppl. Info 2 Supplementary Figure 1. Transcripts arising from the structurally conserved subtelomeres
GFP CCD2 GFP IP:GFP
 D1 D2 1 75 95 148 178 492 GFP CCD1 CCD2 CCD2 GFP D1 D2 GFP D1 D2 Beclin 1 IB:GFP IP:GFP Supplementary Figure 1: Mapping domains required for binding to HEK293T cells are transfected with EGFP-tagged mutant
D1 D2 1 75 95 148 178 492 GFP CCD1 CCD2 CCD2 GFP D1 D2 GFP D1 D2 Beclin 1 IB:GFP IP:GFP Supplementary Figure 1: Mapping domains required for binding to HEK293T cells are transfected with EGFP-tagged mutant
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1.
 Supplementary Figure 1. Characterization and high expression of Lnc-β-Catm in liver CSCs. (a) Heatmap of differently expressed lncrnas in Liver CSCs (CD13 + CD133 + ) and non-cscs (CD13 - CD133 - ) according
Supplementary Figure 1. Characterization and high expression of Lnc-β-Catm in liver CSCs. (a) Heatmap of differently expressed lncrnas in Liver CSCs (CD13 + CD133 + ) and non-cscs (CD13 - CD133 - ) according
Supplementary Figure 1: sgrna library generation and the length of sgrnas for the functional screen. (a) A diagram of the retroviral vector for sgrna
 Supplementary Figure 1: sgrna library generation and the length of sgrnas for the functional screen. (a) A diagram of the retroviral vector for sgrna expression. It contains a U6-promoter-driven sgrna
Supplementary Figure 1: sgrna library generation and the length of sgrnas for the functional screen. (a) A diagram of the retroviral vector for sgrna expression. It contains a U6-promoter-driven sgrna
7.06 Problem Set #3, Spring 2005
 7.06 Problem Set #3, Spring 2005 1. The Drosophila compound eye is composed of about 800 units called ommatidia. Each ommatidium contains eight photoreceptor neurons (R1 through R8), which develop in a
7.06 Problem Set #3, Spring 2005 1. The Drosophila compound eye is composed of about 800 units called ommatidia. Each ommatidium contains eight photoreceptor neurons (R1 through R8), which develop in a
Supplemental Materials. Matrix Proteases Contribute to Progression of Pelvic Organ Prolapse in Mice and Humans
 Supplemental Materials Matrix Proteases Contribute to Progression of Pelvic Organ Prolapse in Mice and Humans Madhusudhan Budatha, Shayzreen Roshanravan, Qian Zheng, Cecilia Weislander, Shelby L. Chapman,
Supplemental Materials Matrix Proteases Contribute to Progression of Pelvic Organ Prolapse in Mice and Humans Madhusudhan Budatha, Shayzreen Roshanravan, Qian Zheng, Cecilia Weislander, Shelby L. Chapman,
Construction of plant complementation vector and generation of transgenic plants
 MATERIAL S AND METHODS Plant materials and growth conditions Arabidopsis ecotype Columbia (Col0) was used for this study. SALK_072009, SALK_076309, and SALK_027645 were obtained from the Arabidopsis Biological
MATERIAL S AND METHODS Plant materials and growth conditions Arabidopsis ecotype Columbia (Col0) was used for this study. SALK_072009, SALK_076309, and SALK_027645 were obtained from the Arabidopsis Biological
Supplementary Figure 1
 number of cells, normalized number of cells, normalized number of cells, normalized Supplementary Figure CD CD53 Cd3e fluorescence intensity fluorescence intensity fluorescence intensity Supplementary
number of cells, normalized number of cells, normalized number of cells, normalized Supplementary Figure CD CD53 Cd3e fluorescence intensity fluorescence intensity fluorescence intensity Supplementary
Supplementary Figure 1
 Supplementary Figure A Name Location Start position End position Oct4P Chr 3 29433673 29435 Annotation Mus musculus predicted gene 572 (Gm572), noncoding RNA RefSeq accession number NR_33594. Oct4P2 Chr
Supplementary Figure A Name Location Start position End position Oct4P Chr 3 29433673 29435 Annotation Mus musculus predicted gene 572 (Gm572), noncoding RNA RefSeq accession number NR_33594. Oct4P2 Chr
A tool kit for rapid cloning and expression of. recombinant antibodies
 A tool kit for rapid cloning and expression of recombinant antibodies Tihomir S Dodev 1,4, Panagiotis Karagiannis 1,2, Amy E Gilbert 1,2, Debra H Josephs 1,2,3, Holly Bowen 1,4, Louisa K James 4, Heather
A tool kit for rapid cloning and expression of recombinant antibodies Tihomir S Dodev 1,4, Panagiotis Karagiannis 1,2, Amy E Gilbert 1,2, Debra H Josephs 1,2,3, Holly Bowen 1,4, Louisa K James 4, Heather
TransAM Kits are DNA-binding ELISAs that facilitate the study of transcription factor activation in mammalian tissue and cell culture extracts.
 Transcription Factor ELISAs TransAM sensitive quantitative transcription factor ELISAs TransAM Kits are DNA-binding ELISAs that facilitate the study of transcription factor activation in mammalian tissue
Transcription Factor ELISAs TransAM sensitive quantitative transcription factor ELISAs TransAM Kits are DNA-binding ELISAs that facilitate the study of transcription factor activation in mammalian tissue
Cycles of vascular plexus formation within the nephrogenic zone of the developing mouse kidney
 1 Supplementary text and data for: 2 3 4 5 Cycles of vascular plexus formation within the nephrogenic zone of the developing mouse kidney Authors: David A. D. Munro 1*, Peter Hohenstein 2, and Jamie A.
1 Supplementary text and data for: 2 3 4 5 Cycles of vascular plexus formation within the nephrogenic zone of the developing mouse kidney Authors: David A. D. Munro 1*, Peter Hohenstein 2, and Jamie A.
Supplementary Figure 1 Activated B cells are subdivided into three groups
 Supplementary Figure 1 Activated B cells are subdivided into three groups according to mitochondrial status (a) Flow cytometric analysis of mitochondrial status monitored by MitoTracker staining or differentiation
Supplementary Figure 1 Activated B cells are subdivided into three groups according to mitochondrial status (a) Flow cytometric analysis of mitochondrial status monitored by MitoTracker staining or differentiation
Influence of the polymorphisms of the α-major regulatory element HS-40 on in vitro gene expression
 Volume 42 (9) 776-869 September 2009 Braz J Med Biol Res, September 2009, Volume 42(9) 783-786 Influence of the polymorphisms of the α-major regulatory element HS-40 on in vitro gene expression D.M. Ribeiro,
Volume 42 (9) 776-869 September 2009 Braz J Med Biol Res, September 2009, Volume 42(9) 783-786 Influence of the polymorphisms of the α-major regulatory element HS-40 on in vitro gene expression D.M. Ribeiro,
Figure S1. Immunoblotting showing proper expression of tagged R and AVR proteins in N. benthamiana.
 SUPPLEMENTARY FIGURES Figure S1. Immunoblotting showing proper expression of tagged R and AVR proteins in N. benthamiana. YFP:, :CFP, HA: and (A) and HA, CFP and YFPtagged and AVR1-CO39 (B) were expressed
SUPPLEMENTARY FIGURES Figure S1. Immunoblotting showing proper expression of tagged R and AVR proteins in N. benthamiana. YFP:, :CFP, HA: and (A) and HA, CFP and YFPtagged and AVR1-CO39 (B) were expressed
PPAR-γ - Peroxisome proliferator-ac4vated receptor gamma
 PPAR-γ - Peroxisome proliferator-ac4vated receptor gamma PPAR-γ - Peroxisome proliferator-ac4vated receptor gamma C/EBPalpha - CCAAT/enhancer-binding protein alpha C/EBPα contains: - a func)onally related
PPAR-γ - Peroxisome proliferator-ac4vated receptor gamma PPAR-γ - Peroxisome proliferator-ac4vated receptor gamma C/EBPalpha - CCAAT/enhancer-binding protein alpha C/EBPα contains: - a func)onally related
ZFP36L1 Negatively Regulates Erythroid Differentiation of CD34 Hematopoietic Stem Cells by Interfering with the Stat5b Pathway
 Molecular Biology of the Cell Vol. 21, 3340 3351, October 1, 2010 ZFP36L1 Negatively Regulates Erythroid Differentiation of CD34 Hematopoietic Stem Cells by Interfering with the Stat5b Pathway Tatiana
Molecular Biology of the Cell Vol. 21, 3340 3351, October 1, 2010 ZFP36L1 Negatively Regulates Erythroid Differentiation of CD34 Hematopoietic Stem Cells by Interfering with the Stat5b Pathway Tatiana
Supplemental Information. HEXIM1 and NEAT1 Long Non-coding RNA Form. a Multi-subunit Complex that Regulates. DNA-Mediated Innate Immune Response
 Molecular Cell, Volume 67 Supplemental Information HEXIM1 and NEAT1 Long Non-coding RNA Form a Multi-subunit Complex that Regulates DNA-Mediated Innate Immune Response Mehdi Morchikh, Alexandra Cribier,
Molecular Cell, Volume 67 Supplemental Information HEXIM1 and NEAT1 Long Non-coding RNA Form a Multi-subunit Complex that Regulates DNA-Mediated Innate Immune Response Mehdi Morchikh, Alexandra Cribier,
TECH NOTE SMARTer T-cell receptor profiling in single cells
 TECH NOTE SMARTer T-cell receptor profiling in single cells Flexible workflow: Illumina-ready libraries from FACS or manually sorted single cells Ease of use: Optimized indexing allows for pooling 96 cells
TECH NOTE SMARTer T-cell receptor profiling in single cells Flexible workflow: Illumina-ready libraries from FACS or manually sorted single cells Ease of use: Optimized indexing allows for pooling 96 cells
Molecular Genetics Techniques. BIT 220 Chapter 20
 Molecular Genetics Techniques BIT 220 Chapter 20 What is Cloning? Recombinant DNA technologies 1. Producing Recombinant DNA molecule Incorporate gene of interest into plasmid (cloning vector) 2. Recombinant
Molecular Genetics Techniques BIT 220 Chapter 20 What is Cloning? Recombinant DNA technologies 1. Producing Recombinant DNA molecule Incorporate gene of interest into plasmid (cloning vector) 2. Recombinant
Figure S2. Response of mouse ES cells to GSK3 inhibition. Mentioned in discussion
 Stem Cell Reports, Volume 1 Supplemental Information Robust Self-Renewal of Rat Embryonic Stem Cells Requires Fine-Tuning of Glycogen Synthase Kinase-3 Inhibition Yaoyao Chen, Kathryn Blair, and Austin
Stem Cell Reports, Volume 1 Supplemental Information Robust Self-Renewal of Rat Embryonic Stem Cells Requires Fine-Tuning of Glycogen Synthase Kinase-3 Inhibition Yaoyao Chen, Kathryn Blair, and Austin
Figure 1: TDP-43 is subject to lysine acetylation within the RNA-binding domain a) QBI-293 cells were transfected with TDP-43 in the presence or
 Figure 1: TDP-43 is subject to lysine acetylation within the RNA-binding domain a) QBI-293 cells were transfected with TDP-43 in the presence or absence of the acetyltransferase CBP and acetylated TDP-43
Figure 1: TDP-43 is subject to lysine acetylation within the RNA-binding domain a) QBI-293 cells were transfected with TDP-43 in the presence or absence of the acetyltransferase CBP and acetylated TDP-43
IKK is a therapeutic target in KRAS-induced lung cancer with disrupted p53 activity
 IKK is a therapeutic target in KRAS-induced lung cancer with disrupted p5 activity H6 5 5 H58 A59 H6 H58 A59 anti-ikkα anti-ikkβ anti-panras anti-gapdh anti-ikkα anti-ikkβ anti-panras anti-gapdh anti-ikkα
IKK is a therapeutic target in KRAS-induced lung cancer with disrupted p5 activity H6 5 5 H58 A59 H6 H58 A59 anti-ikkα anti-ikkβ anti-panras anti-gapdh anti-ikkα anti-ikkβ anti-panras anti-gapdh anti-ikkα
ENCODE RBP Antibody Characterization Guidelines
 ENCODE RBP Antibody Characterization Guidelines Approved on November 18, 2016 Background An integral part of the ENCODE Project is to characterize the antibodies used in the experiments. This document
ENCODE RBP Antibody Characterization Guidelines Approved on November 18, 2016 Background An integral part of the ENCODE Project is to characterize the antibodies used in the experiments. This document
466 Asn (N) to Ala (A) Generate beta dimer Interface
 Table S1: Amino acid changes to the HexA α-subunit to convert the dimer interface from α to β and to introduce the putative GM2A binding surface from β- onto the α- subunit Residue position (α-numbering)
Table S1: Amino acid changes to the HexA α-subunit to convert the dimer interface from α to β and to introduce the putative GM2A binding surface from β- onto the α- subunit Residue position (α-numbering)
HCT116 SW48 Nutlin: p53
 Figure S HCT6 SW8 Nutlin: - + - + p GAPDH Figure S. Nutlin- treatment induces p protein. HCT6 and SW8 cells were left untreated or treated for 8 hr with Nutlin- ( µm) to up-regulate p. Whole cell lysates
Figure S HCT6 SW8 Nutlin: - + - + p GAPDH Figure S. Nutlin- treatment induces p protein. HCT6 and SW8 cells were left untreated or treated for 8 hr with Nutlin- ( µm) to up-regulate p. Whole cell lysates
DNA Microarray Technology
 CHAPTER 1 DNA Microarray Technology All living organisms are composed of cells. As a functional unit, each cell can make copies of itself, and this process depends on a proper replication of the genetic
CHAPTER 1 DNA Microarray Technology All living organisms are composed of cells. As a functional unit, each cell can make copies of itself, and this process depends on a proper replication of the genetic
Engineering Nanomedical Systems. Assessing the efficacy of nanomedical therapies
 BME 626 October 9, 2014 Engineering Nanomedical Systems Lecture 12 Assessing the efficacy of nanomedical therapies James F. Leary, Ph.D. SVM Endowed Professor of Nanomedicine Professor of Basic Medical
BME 626 October 9, 2014 Engineering Nanomedical Systems Lecture 12 Assessing the efficacy of nanomedical therapies James F. Leary, Ph.D. SVM Endowed Professor of Nanomedicine Professor of Basic Medical
Biology 201 (Genetics) Exam #3 120 points 20 November Read the question carefully before answering. Think before you write.
 Name KEY Section Biology 201 (Genetics) Exam #3 120 points 20 November 2006 Read the question carefully before answering. Think before you write. You will have up to 50 minutes to take this exam. After
Name KEY Section Biology 201 (Genetics) Exam #3 120 points 20 November 2006 Read the question carefully before answering. Think before you write. You will have up to 50 minutes to take this exam. After
FEBRUARY 12, 2010 VOLUME 285 NUMBER 7 JOURNAL OF BIOLOGICAL CHEMISTRY 4847
 THE JOURNAL OF BIOLOGICAL CHEMISTRY VOL. 285, NO. 7, pp. 4847 4858, February 12, 2010 2010 by The American Society for Biochemistry and Molecular Biology, Inc. Printed in the U.S.A. The Tumor Suppressor
THE JOURNAL OF BIOLOGICAL CHEMISTRY VOL. 285, NO. 7, pp. 4847 4858, February 12, 2010 2010 by The American Society for Biochemistry and Molecular Biology, Inc. Printed in the U.S.A. The Tumor Suppressor
Supplementary Fig. S1. SAMHD1c has a more potent dntpase activity than. SAMHD1c. Purified recombinant SAMHD1c and SAMHD1c proteins (with
 Supplementary Fig. S1. SAMHD1c has a more potent dntpase activity than SAMHD1c. Purified recombinant SAMHD1c and SAMHD1c proteins (with concentration of 800nM) were incubated with 1mM dgtp for the indicated
Supplementary Fig. S1. SAMHD1c has a more potent dntpase activity than SAMHD1c. Purified recombinant SAMHD1c and SAMHD1c proteins (with concentration of 800nM) were incubated with 1mM dgtp for the indicated
Figure S6. Detection of anti-gfp antibodies in anti-dna and normal plasma without competition DNA--9
 Supplementary Information Ultrasensitive antibody detection by agglutination-pcr (ADAP) Cheng-ting Tsai 1 *, Peter V. Robinson 1 *, Carole A. Spencer 2 and Carolyn R. Bertozzi 3,4ǂ Department of 1 Chemistry,
Supplementary Information Ultrasensitive antibody detection by agglutination-pcr (ADAP) Cheng-ting Tsai 1 *, Peter V. Robinson 1 *, Carole A. Spencer 2 and Carolyn R. Bertozzi 3,4ǂ Department of 1 Chemistry,
Galina Gabriely, Ph.D. BWH/HMS
 Galina Gabriely, Ph.D. BWH/HMS Email: ggabriely@rics.bwh.harvard.edu Outline: microrna overview microrna expression analysis microrna functional analysis microrna (mirna) Characteristics mirnas discovered
Galina Gabriely, Ph.D. BWH/HMS Email: ggabriely@rics.bwh.harvard.edu Outline: microrna overview microrna expression analysis microrna functional analysis microrna (mirna) Characteristics mirnas discovered
File name: Supplementary Information Description: Supplementary figures and supplementary tables. File name: Peer review file Description:
 File name: Supplementary Information Description: Supplementary figures and supplementary tables. File name: Peer review file Description: Supplementary figure 1 Flow diagram summarizing the overall experimental
File name: Supplementary Information Description: Supplementary figures and supplementary tables. File name: Peer review file Description: Supplementary figure 1 Flow diagram summarizing the overall experimental
Pre-made Lentiviral Particles for Nuclear Permeant CRE Recombinase Expression
 Pre-made Lentiviral Particles for Nuclear Permeant CRE Recombinase Expression LVP336 LVP336-PBS LVP339 LVP339-PBS LVP297 LVP297-PBS LVP013 LVP013-PBS LVP338 LVP338-PBS LVP027 LVP027-PBS LVP337 LVP337-PBS
Pre-made Lentiviral Particles for Nuclear Permeant CRE Recombinase Expression LVP336 LVP336-PBS LVP339 LVP339-PBS LVP297 LVP297-PBS LVP013 LVP013-PBS LVP338 LVP338-PBS LVP027 LVP027-PBS LVP337 LVP337-PBS
Assay Design Considerations, Optimization and Validation
 Assay Design Considerations, Optimization and Validation Ray Meng, Ph.D. International Field Applications Specialist Gene Expression Division Bio-Rad Laboratories, Inc. Assay Design Considerations Experiment
Assay Design Considerations, Optimization and Validation Ray Meng, Ph.D. International Field Applications Specialist Gene Expression Division Bio-Rad Laboratories, Inc. Assay Design Considerations Experiment
Supplementary Information Design of small molecule-responsive micrornas based on structural requirements for Drosha processing
 Supplementary Information Design of small molecule-responsive micrornas based on structural requirements for Drosha processing Chase L. Beisel, Yvonne Y. Chen, Stephanie J. Culler, Kevin G. Hoff, & Christina
Supplementary Information Design of small molecule-responsive micrornas based on structural requirements for Drosha processing Chase L. Beisel, Yvonne Y. Chen, Stephanie J. Culler, Kevin G. Hoff, & Christina
Bronchial epithelium and its associated tissues act as a
 The Journal of Immunology A JNK-Independent Signaling Pathway Regulates TNF -Stimulated, c-jun-driven FRA-1 Protooncogene Transcription in Pulmonary Epithelial Cells 1 Pavan Adiseshaiah,* Dhananjaya V.
The Journal of Immunology A JNK-Independent Signaling Pathway Regulates TNF -Stimulated, c-jun-driven FRA-1 Protooncogene Transcription in Pulmonary Epithelial Cells 1 Pavan Adiseshaiah,* Dhananjaya V.
Supporting Online Material for
 www.sciencemag.org/cgi/content/full/1137999/dc1 Supporting Online Material for Disrupting the Pairing Between let-7 and Enhances Oncogenic Transformation Christine Mayr, Michael T. Hemann, David P. Bartel*
www.sciencemag.org/cgi/content/full/1137999/dc1 Supporting Online Material for Disrupting the Pairing Between let-7 and Enhances Oncogenic Transformation Christine Mayr, Michael T. Hemann, David P. Bartel*
Genome Sequence Assembly
 Genome Sequence Assembly Learning Goals: Introduce the field of bioinformatics Familiarize the student with performing sequence alignments Understand the assembly process in genome sequencing Introduction:
Genome Sequence Assembly Learning Goals: Introduce the field of bioinformatics Familiarize the student with performing sequence alignments Understand the assembly process in genome sequencing Introduction:
Supplementary Materials
 Supplementary Materials Construction of Synthetic Nucleoli in Human Cells Reveals How a Major Functional Nuclear Domain is Formed and Propagated Through Cell Divisision Authors: Alice Grob, Christine Colleran
Supplementary Materials Construction of Synthetic Nucleoli in Human Cells Reveals How a Major Functional Nuclear Domain is Formed and Propagated Through Cell Divisision Authors: Alice Grob, Christine Colleran
Supplementary Information
 Journal : Nature Biotechnology Supplementary Information Targeted genome engineering in human cells with RNA-guided endonucleases Seung Woo Cho, Sojung Kim, Jong Min Kim, and Jin-Soo Kim* National Creative
Journal : Nature Biotechnology Supplementary Information Targeted genome engineering in human cells with RNA-guided endonucleases Seung Woo Cho, Sojung Kim, Jong Min Kim, and Jin-Soo Kim* National Creative
Pros and Cons of Gene Therapy in ß-Thalassemia
 Pros and Cons of Gene Therapy in ß-Thalassemia ß-Thalassemia: established and potential new therapeutic approaches Many ß-thalassemic patients are treated with blood transfusion and iron chelation. However,
Pros and Cons of Gene Therapy in ß-Thalassemia ß-Thalassemia: established and potential new therapeutic approaches Many ß-thalassemic patients are treated with blood transfusion and iron chelation. However,
Supplementary Information
 Supplementary Information Supplementary Figure 1 A B Tetracycline HEK293-eGFP-671 HEK293-eGFP-769 mir-671 mir-671 / RNU48 0.3 0.2 0.1 mir-769 mir-15b 0 HEK293 mir-671- mir-671+3d mir-671+40d 4.28 1 6.17
Supplementary Information Supplementary Figure 1 A B Tetracycline HEK293-eGFP-671 HEK293-eGFP-769 mir-671 mir-671 / RNU48 0.3 0.2 0.1 mir-769 mir-15b 0 HEK293 mir-671- mir-671+3d mir-671+40d 4.28 1 6.17
CHAPTER 9 DNA Technologies
 CHAPTER 9 DNA Technologies Recombinant DNA Artificially created DNA that combines sequences that do not occur together in the nature Basis of much of the modern molecular biology Molecular cloning of genes
CHAPTER 9 DNA Technologies Recombinant DNA Artificially created DNA that combines sequences that do not occur together in the nature Basis of much of the modern molecular biology Molecular cloning of genes
MIT Department of Biology 7.013: Introductory Biology - Spring 2005 Instructors: Professor Hazel Sive, Professor Tyler Jacks, Dr.
 MIT Department of Biology 7.01: Introductory Biology - Spring 2005 Instructors: Professor Hazel Sive, Professor Tyler Jacks, Dr. Claudette Gardel iv) Would Xba I be useful for cloning? Why or why not?
MIT Department of Biology 7.01: Introductory Biology - Spring 2005 Instructors: Professor Hazel Sive, Professor Tyler Jacks, Dr. Claudette Gardel iv) Would Xba I be useful for cloning? Why or why not?
Candidate region (0.74 Mb) ATC TCT GGG ACT CAT GAG CAG GAG GCT AGC ATC TCT GGG ACT CAT TAG CAG GAG GCT AGC
 A idm-3 idm-3 B Physical distance (Mb) 4.6 4.86.6 8.4 C Chr.3 Recom. Rate (%) ATG 3.9.9.9 9.74 Candidate region (.74 Mb) n=4 TAA D idm-3 G3T(E4) G4A(W988) WT idm-3 ATC TCT GGG ACT CAT GAG CAG GAG GCT AGC
A idm-3 idm-3 B Physical distance (Mb) 4.6 4.86.6 8.4 C Chr.3 Recom. Rate (%) ATG 3.9.9.9 9.74 Candidate region (.74 Mb) n=4 TAA D idm-3 G3T(E4) G4A(W988) WT idm-3 ATC TCT GGG ACT CAT GAG CAG GAG GCT AGC
Coleman et al., Supplementary Figure 1
 Coleman et al., Supplementary Figure 1 BrdU Merge G1 Early S Mid S Supplementary Figure 1. Sequential destruction of CRL4 Cdt2 targets during the G1/S transition. HCT116 cells were synchronized by sequential
Coleman et al., Supplementary Figure 1 BrdU Merge G1 Early S Mid S Supplementary Figure 1. Sequential destruction of CRL4 Cdt2 targets during the G1/S transition. HCT116 cells were synchronized by sequential
Parthanatos mediates AIMP2-activated age-dependent dopaminergic neuronal loss
 SUPPLEMENTARY INFORMATION Parthanatos mediates AIMP2-activated age-dependent dopaminergic neuronal loss Yunjong Lee, Senthilkumar S. Karuppagounder, Joo-Ho Shin, Yun-Il Lee, Han Seok Ko, Debbie Swing,
SUPPLEMENTARY INFORMATION Parthanatos mediates AIMP2-activated age-dependent dopaminergic neuronal loss Yunjong Lee, Senthilkumar S. Karuppagounder, Joo-Ho Shin, Yun-Il Lee, Han Seok Ko, Debbie Swing,
