Table I. Primers used in quantitative real-time PCR for detecting gene expressions in mouse liver tissues
|
|
|
- Dortha Beasley
- 6 years ago
- Views:
Transcription
1 Table I. Primers used in quantitative real-time PCR for detecting gene expressions in mouse liver tissues Gene name Forward primer 5' to 3' Gene name Reverse primer 5' to 3' CC_F TGCCCTGTTGGGGTTG CC_R TGTGGCGGGGCTTG CC_F CTTGGGCCCGCTGTC CC_R TGCCCTTTGCCTCCG lbumin_f GTGCGGGCGGCTTG lbumin_r GCCCTTCCTGGTCCTC FSN_F GGCTCTTGGGCCTCCTT FSN_R GCTGCGCCGCCTCTCT GPDH_F CTTTGGCTTGTGGGG GPDH_R GGTGCGGGTGTGTTCT HMGCR_F CTTTCGCGCTGTGCTCC HMGCR_R CTGTGGGTGTGGCTGT HMGCS1_F TTTGTGCGCTGTTTGGG HMGCS1_R CCCCTGTGGTCTGGCTT HMGCS2_F CCCCTGGGTTCCG HMGCS2_R TGCTCTCTCCCTCGTTC HNF1a_F GCCCGGCCCCGTGCC HNF1a_R GGCTTCCCCTCGCTCCCG HNRNP-D_F GTCGGGGTGTGTGGTC HNRNP-D_R GGCCCTTTTGGTCTGCTT HNRNP-I_F GCCTGGCGTGC HNRNP-I_R GCGCCCCGTGTTGTGT KSRP_F CTGGGCCCTGGTCTGT KSRP_R CGTTGTCGTGCTGTCCT LDLR_F CCTGCCGCCTGTGTTC LDLR_R GCGTCTGTTCCGGTCC Luc2_F CGCCTTCGGGTGGC Luc2_R GCGCTTTCTCGCTGCC PCSK9_F TTGCGCGCTGGGCTT PCSK9_R CCGCTGTGTGCCTCTGG SREP1c_F CGGCCTCGCTCTCCG SREP1c_R CCCCTTCGGGTTTCTGC SREP2_ F CCGGGGGGGCGG SREP2_ R CGCCGCTTGTGCTCTTG
2 Table II. mirn microarray analysis to identify R regulated micro RNs in HepG2 cells with target sites in LDLR 3 UTR. Fold change (R 4 h vs. DMSO 4 h) Fold change (R 8 h vs. DMSO 8 h) Fold change (R 24 h vs. DMSO 24 h ) Column # Column ID mml-mir-19a hsa-mir-130b hsa-mir-615-5p hsa-mir-671-5p hsa-mir hsa-mir has-mir-1244* hsa-mir-1979* hsa-mir crm-mir * mirn target site is not present in LDLR 3 UTR
3 Supplemental Figure I Relative Luc ctivity WT * * RE1-mu RE2-mu RE3-mu RE1+2-mu RE1+2+3-mu R fold induction WT RE1-mu C RE2-mu Normalized Luc ctivity RE3-mu pluc RE1+2-mu pluc-ldlrutr1 RE1+2+3-mu pluc-re1 pluc-re2 pluc-re3 Supplemental Figure I: Identification of three destabilizing RE motifs in mouse LDLR 3 UTR. () HepG2 cells were transfected with individual luciferase-utr reporter plasmid along with prl-sv40 as the normalizing transfection vector for 48 hours. R was added to cells for 8 hours before cell lysis. The stimulatory activity of R on the wild-type UTR reporter was defined as %, and activities of R on mutated vectors were plotted relative to that value. The data shown are mean ± SEM of three separate transfections in which triplicate wells were used for each reporter. () Comparison of human and mouse LDLR mrn 3 UTR sequences. Segments of RE containing sequences are shown. Mouse nucleotide sequences that differ from those of the human sequence are boxed. The sequences of REs are in red color and the UUU motifs are underlined. (C) LDLR3 UTR reporter plasmids or control vector pluc were cotransfected with prl-sv40 into Hepa 1-6 cells. Two days post-transfection, cell lysates were prepared. Dual luciferase activities were measured and expressed as normalized luciferase activity. Each value represents the mean ± SEM of four wells per condition. p < 01 as compared to pluc. The graph shown is representative of three separate experiments.
4 Supplemental Figure II p42/40 Liver MPH Hepa 1-6 HNRNPD /actin: /actin: shrn: CGGGGTGTGTGGTC human HNRN PD V1: TCCGGGCGTCGGGGTTTTGGCTTTGTGCTTTTGTCGGGGTGTGTGGTCTGGTCG mouse HNR N PD V1: TCCGGGCGTCGGGGTTTTGGCTTTGTGCTTTTGGTCGGGGTGTGTGGTCTGGTCGG mouse HNR NPD V2: TCCGGGCGTCGGGGTTTTGGCTTTGTGCTTTTGGTCGGGGTGTGTGGTCTGGTCGG mouse HNR N PD V3: TCCGGGCGTCGGGGTTTTGGCTTTGTGCTTTTGGTCGGGGTGTGTGGTCTGGTCGG mouse HNR N PD V4: TCCGGGCGTCGGGGTTTTGGCTTTGTGCTTTTGGTCGGGGTGTGTGGTCTGGTCGG Supplemental Figure II: Characterization of mouse hnrnp D protein isoforms and hnrnpd shrn sequence alignment with hnrnpd cdns. () Western blot analysis of hnrnpd isoforms in mouse liver tissue, mouse primary hepatocytes (MPH) and hepatoma cells. Protein abundances of hnrnp D / were quantified using the lpha View Software with normalization by signals of. Values are average of two samples. The relative signal intensity of hnrnp D in liver is expressed as 1. () hnrnpd shrn shows % alignment with all mouse hnrnp D isoforms and human hnrnp D V1.
5 Supplemental Figure III Normalized Luc ctivity 1.4 Hepa HepG mock p40 p42 C Normalized Luc ctivity (Fold of DMSO) mock p40 p42 mock HNRNPD DMSO R (40uM) p40 control CMV-Flag (mock) Flag- Flag-p40 Flag-p42 Flag- p42/40 Supplemental Figure III: Exogenous expression of hnrnp D isoforms reduced LDLR3 UTR reporter activity in hepatic cells. () Hepa 1-6 and HepG2 cells were cotransfected with LDLR3 UTR luciferase reporter and indicated hnrnp D isoform expression plasmids or the control vector (mock). prl-sv40 was cotransfected in each condition for normalization of transfection efficiency. In each cell line, the normalized luciferase activity in mock-transfected cells is expressed as 1. p<01 as compared to the mock transfection. The graph is representative of two independent transfection experiments. () Huh7 cells were transfected with flag-tagged hnrnp D isoforms or the empty vector. Cell lysates from transfected or untransfected control cells were analyzed for isoform expression using anti-flag antibody. (C) HepG2 cells were cotransfected with LDLR3 UTR luciferase reporter and indicated hnrnp D isoform expression plasmids or the control vector (mock). One day post transfection, cells were treated with R or DMSO for 24 h before cell lysis. prl-sv40 was cotransfected in each condition for normalization of transfection efficiency. The normalized luciferase activity in mock-transfected cells without R treatment is expressed as 1. The graph is representative of two independent transfection experiments in which six wells were used for each condition.
6 Supplemental Figure IV Relative LDLRmRN levels C h Relative LDLR mrn levels 4h DMSO h R control anti-mir-130b anti-mir h DMSO 8h R 24h DMSO anti-mir-615-5p anti-mir-671-5p anti-mir-760 anti-mir-1201 anti-mir-1826 anti-mir-223 anti-mir h R D Normalized luciferase activity anti-mir control anti-mir-130b LDLR LDLR pluc pluc-ldlr3'utr anti-mir-1979 anti-mir-615-5p anti-mir-671-5p anti-mir-760 anti-mir-1201 anti-mir-1826 anti-mir-223 anti-mir-1244 anti-mir control anti-mir-130b anti-mir-1979 anti-mir-615-5p anti-mir-671-5p anti-mir-760 anti-mir-1201 anti-mir-1826 anti-mir-223 anti-mir-1244 SSY #1 SSY #2 Supplemental Figure IV: Evaluation of mirns in LDLR3 UTR reporter activity, LDLR mrn and protein expression in HepG2 cells () HepG2 cells were treated with R for indicated times. Total RN was isolated for qpcr analysis of LDLR and GPDH mrn expressions. fter normalization with GPDH mrn levels, the relative levels are presented, and the results are means ± SE of triplicate measurements for each cdn sample. p < 01 compared to DMSO. () HepG2 cells were cotransfected with pluc control vector or LDLR3 UTR luciferase reporter and indicated anti-mirn oligos or the control oligo (mock). prl-sv40 was cotransfected in each condition for normalization of transfection efficiency. The graph is representative of two independent transfection experiments in which six wells were used for each condition. (C-D) HepG2 cells were transfected with indicated anti-mirn synthetic oligonucleotides for three days before isolation of total RN or toaly cell lysates for analyses of LDLR mrn and protein expression levels.
7 Supplemental Figure V HepG2 Hepa1-6 H9C2 7R5 Lading Y1 MPH Supplemental Figure V: Measurement of cyclin D1 mrn decay rates in various cell lines and mouse primary hepatocytes (MPH). Cells were treated with actinomycin D (5 µg/ml) for different intervals. Total RN was isolated and analyzed for the amounts of cyclin D1 mrns by qrt-pcr. The cyclin D1 mrn levels were plotted as the percentage of the mrn remaining. Decay curves were plotted versus time.
ASPP1 Fw GGTTGGGAATCCACGTGTTG ASPP1 Rv GCCATATCTTGGAGCTCTGAGAG
 Supplemental Materials and Methods Plasmids: the following plasmids were used in the supplementary data: pwzl-myc- Lats2 (Aylon et al, 2006), pretrosuper-vector and pretrosuper-shp53 (generous gift of
Supplemental Materials and Methods Plasmids: the following plasmids were used in the supplementary data: pwzl-myc- Lats2 (Aylon et al, 2006), pretrosuper-vector and pretrosuper-shp53 (generous gift of
SUPPLEMENTARY INFORMATION
 doi:.38/nature899 Supplementary Figure Suzuki et al. a c p7 -/- / WT ratio (+)/(-) p7 -/- / WT ratio Log X 3. Fold change by treatment ( (+)/(-)) Log X.5 3-3. -. b Fold change by treatment ( (+)/(-)) 8
doi:.38/nature899 Supplementary Figure Suzuki et al. a c p7 -/- / WT ratio (+)/(-) p7 -/- / WT ratio Log X 3. Fold change by treatment ( (+)/(-)) Log X.5 3-3. -. b Fold change by treatment ( (+)/(-)) 8
Transcriptional Regulation of Pro-apoptotic Protein Kinase C-delta: Implications for Oxidative Stress-induced Neuronal Cell Death
 SUPPLEMENTAL DATA Transcriptional Regulation of Pro-apoptotic Protein Kinase C-delta: Implications for Oxidative Stress-induced Neuronal Cell Death Huajun Jin 1, Arthi Kanthasamy 1, Vellareddy Anantharam
SUPPLEMENTAL DATA Transcriptional Regulation of Pro-apoptotic Protein Kinase C-delta: Implications for Oxidative Stress-induced Neuronal Cell Death Huajun Jin 1, Arthi Kanthasamy 1, Vellareddy Anantharam
Online Supplementary Information
 Online Supplementary Information NLRP4 negatively regulates type I interferon signaling by targeting TBK1 for degradation via E3 ubiquitin ligase DTX4 Jun Cui 1,4,6,7, Yinyin Li 1,5,6,7, Liang Zhu 1, Dan
Online Supplementary Information NLRP4 negatively regulates type I interferon signaling by targeting TBK1 for degradation via E3 ubiquitin ligase DTX4 Jun Cui 1,4,6,7, Yinyin Li 1,5,6,7, Liang Zhu 1, Dan
RNA oligonucleotides and 2 -O-methylated oligonucleotides were synthesized by. 5 AGACACAAACACCAUUGUCACACUCCACAGC; Rand-2 OMe,
 Materials and methods Oligonucleotides and DNA constructs RNA oligonucleotides and 2 -O-methylated oligonucleotides were synthesized by Dharmacon Inc. (Lafayette, CO). The sequences were: 122-2 OMe, 5
Materials and methods Oligonucleotides and DNA constructs RNA oligonucleotides and 2 -O-methylated oligonucleotides were synthesized by Dharmacon Inc. (Lafayette, CO). The sequences were: 122-2 OMe, 5
Nature Methods: doi: /nmeth Supplementary Figure 1. Validation of RaPID with EDEN15
 Supplementary Figure 1 Validation of RaPID with EDEN15 (a) Full Western Blot of conventional biotinylated RNA pulldown with EDEN15 and scrambled control (n=3 biologically independent experiments, representative
Supplementary Figure 1 Validation of RaPID with EDEN15 (a) Full Western Blot of conventional biotinylated RNA pulldown with EDEN15 and scrambled control (n=3 biologically independent experiments, representative
Supporting Online Material for
 www.sciencemag.org/cgi/content/full/1137999/dc1 Supporting Online Material for Disrupting the Pairing Between let-7 and Enhances Oncogenic Transformation Christine Mayr, Michael T. Hemann, David P. Bartel*
www.sciencemag.org/cgi/content/full/1137999/dc1 Supporting Online Material for Disrupting the Pairing Between let-7 and Enhances Oncogenic Transformation Christine Mayr, Michael T. Hemann, David P. Bartel*
Supplementary Figure 1. (a) The qrt-pcr for lnc-2, lnc-6 and lnc-7 RNA level in DU145, 22Rv1, wild type HCT116 and HCT116 Dicer ex5 cells transfected
 Supplementary Figure 1. (a) The qrt-pcr for lnc-2, lnc-6 and lnc-7 RNA level in DU145, 22Rv1, wild type HCT116 and HCT116 Dicer ex5 cells transfected with the sirna against lnc-2, lnc-6, lnc-7, and the
Supplementary Figure 1. (a) The qrt-pcr for lnc-2, lnc-6 and lnc-7 RNA level in DU145, 22Rv1, wild type HCT116 and HCT116 Dicer ex5 cells transfected with the sirna against lnc-2, lnc-6, lnc-7, and the
Regulation of Synaptic Structure and Function by FMRP- Associated MicroRNAs mir-125b and mir-132
 Neuron, Volume 65 Regulation of Synaptic Structure and Function by FMRP- Associated MicroRNAs mir-125b and mir-132 Dieter Edbauer, Joel R. Neilson, Kelly A. Foster, Chi-Fong Wang, Daniel P. Seeburg, Matthew
Neuron, Volume 65 Regulation of Synaptic Structure and Function by FMRP- Associated MicroRNAs mir-125b and mir-132 Dieter Edbauer, Joel R. Neilson, Kelly A. Foster, Chi-Fong Wang, Daniel P. Seeburg, Matthew
Supporting Information
 Supporting Information Horie et al. 10.1073/pnas.1008499107 SI Materials and Methods ell ulture and Reagents. THP-1 cells were obtained from the American Type ell ollection. THP-1 cells were transformed
Supporting Information Horie et al. 10.1073/pnas.1008499107 SI Materials and Methods ell ulture and Reagents. THP-1 cells were obtained from the American Type ell ollection. THP-1 cells were transformed
mir-24-mediated down-regulation of H2AX suppresses DNA repair
 Supplemental Online Material mir-24-mediated down-regulation of H2AX suppresses DNA repair in terminally differentiated blood cells Ashish Lal 1,4, Yunfeng Pan 2,4, Francisco Navarro 1,4, Derek M. Dykxhoorn
Supplemental Online Material mir-24-mediated down-regulation of H2AX suppresses DNA repair in terminally differentiated blood cells Ashish Lal 1,4, Yunfeng Pan 2,4, Francisco Navarro 1,4, Derek M. Dykxhoorn
Khaled_Fig. S MITF TYROSINASE R²= MITF PDE4D R²= Variance from mean mrna expression
 Khaled_Fig. S Variance from mean mrna expression.8 2 MITF.6 TYROSINASE.4 R²=.73.2.8.6.4.2.8.6.4.2.8.6.4.2 MALME3M SKMEL28 UACC257 MITF PDE4D R²=.63 4.5 5 MITF 3.5 4 PDE4B 2.5 3 R²=.2.5 2.5.8.6.4.2.8.6.4.2
Khaled_Fig. S Variance from mean mrna expression.8 2 MITF.6 TYROSINASE.4 R²=.73.2.8.6.4.2.8.6.4.2.8.6.4.2 MALME3M SKMEL28 UACC257 MITF PDE4D R²=.63 4.5 5 MITF 3.5 4 PDE4B 2.5 3 R²=.2.5 2.5.8.6.4.2.8.6.4.2
Supplemental Information. Mitophagy Controls the Activities. of Tumor Suppressor p53. to Regulate Hepatic Cancer Stem Cells
 Molecular Cell, Volume 68 Supplemental Information Mitophagy Controls the Activities of Tumor Suppressor to Regulate Hepatic Cancer Stem Cells Kai Liu, Jiyoung Lee, Ja Yeon Kim, Linya Wang, Yongjun Tian,
Molecular Cell, Volume 68 Supplemental Information Mitophagy Controls the Activities of Tumor Suppressor to Regulate Hepatic Cancer Stem Cells Kai Liu, Jiyoung Lee, Ja Yeon Kim, Linya Wang, Yongjun Tian,
Supplementary Fig. S1. SAMHD1c has a more potent dntpase activity than. SAMHD1c. Purified recombinant SAMHD1c and SAMHD1c proteins (with
 Supplementary Fig. S1. SAMHD1c has a more potent dntpase activity than SAMHD1c. Purified recombinant SAMHD1c and SAMHD1c proteins (with concentration of 800nM) were incubated with 1mM dgtp for the indicated
Supplementary Fig. S1. SAMHD1c has a more potent dntpase activity than SAMHD1c. Purified recombinant SAMHD1c and SAMHD1c proteins (with concentration of 800nM) were incubated with 1mM dgtp for the indicated
Supplementary information to accompany: A novel role for the DNA repair gene Rad51 in Netrin-1 signalling
 Supplementary information to accompany: A novel role for the DNA repair gene Rad51 in Netrin-1 signalling Glendining KA 1, Markie D 2, Gardner RJM 4, Franz EA 3, Robertson SP 4, Jasoni CL 1 Supplementary
Supplementary information to accompany: A novel role for the DNA repair gene Rad51 in Netrin-1 signalling Glendining KA 1, Markie D 2, Gardner RJM 4, Franz EA 3, Robertson SP 4, Jasoni CL 1 Supplementary
Nature Immunology: doi: /ni Supplementary Figure 1. Zranb1 gene targeting.
 Supplementary Figure 1 Zranb1 gene targeting. (a) Schematic picture of Zranb1 gene targeting using an FRT-LoxP vector, showing the first 6 exons of Zranb1 gene (exons 7-9 are not shown). Targeted mice
Supplementary Figure 1 Zranb1 gene targeting. (a) Schematic picture of Zranb1 gene targeting using an FRT-LoxP vector, showing the first 6 exons of Zranb1 gene (exons 7-9 are not shown). Targeted mice
Supplementary Fig. 1 related to Fig. 1 Clinical relevance of lncrna candidate
 Supplementary Figure Legends Supplementary Fig. 1 related to Fig. 1 Clinical relevance of lncrna candidate BC041951 in gastric cancer. (A) The flow chart for selected candidate lncrnas in 660 up-regulated
Supplementary Figure Legends Supplementary Fig. 1 related to Fig. 1 Clinical relevance of lncrna candidate BC041951 in gastric cancer. (A) The flow chart for selected candidate lncrnas in 660 up-regulated
Figure S1. Immunoblotting showing proper expression of tagged R and AVR proteins in N. benthamiana.
 SUPPLEMENTARY FIGURES Figure S1. Immunoblotting showing proper expression of tagged R and AVR proteins in N. benthamiana. YFP:, :CFP, HA: and (A) and HA, CFP and YFPtagged and AVR1-CO39 (B) were expressed
SUPPLEMENTARY FIGURES Figure S1. Immunoblotting showing proper expression of tagged R and AVR proteins in N. benthamiana. YFP:, :CFP, HA: and (A) and HA, CFP and YFPtagged and AVR1-CO39 (B) were expressed
microrna-122 stimulates translation of Hepatitis C Virus RNA
 Supplementary information for: microrna-122 stimulates translation of Hepatitis C Virus RNA Jura Inga Henke 1,5, Dagmar Goergen 1,5, Junfeng Zheng 3, Yutong Song 4, Christian G. Schüttler 2, Carmen Fehr
Supplementary information for: microrna-122 stimulates translation of Hepatitis C Virus RNA Jura Inga Henke 1,5, Dagmar Goergen 1,5, Junfeng Zheng 3, Yutong Song 4, Christian G. Schüttler 2, Carmen Fehr
Thyroid peroxidase gene expression is induced by lipopolysaccharide involving Nuclear Factor (NF)-κB p65 subunit phosphorylation
 1 2 3 4 5 SUPPLEMENTAL DATA Thyroid peroxidase gene expression is induced by lipopolysaccharide involving Nuclear Factor (NF)-κB p65 subunit phosphorylation Magalí Nazar, Juan Pablo Nicola, María Laura
1 2 3 4 5 SUPPLEMENTAL DATA Thyroid peroxidase gene expression is induced by lipopolysaccharide involving Nuclear Factor (NF)-κB p65 subunit phosphorylation Magalí Nazar, Juan Pablo Nicola, María Laura
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1.
 Supplementary Figure 1. Characterization and high expression of Lnc-β-Catm in liver CSCs. (a) Heatmap of differently expressed lncrnas in Liver CSCs (CD13 + CD133 + ) and non-cscs (CD13 - CD133 - ) according
Supplementary Figure 1. Characterization and high expression of Lnc-β-Catm in liver CSCs. (a) Heatmap of differently expressed lncrnas in Liver CSCs (CD13 + CD133 + ) and non-cscs (CD13 - CD133 - ) according
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION Dynamic Phosphorylation of HP1 Regulates Mitotic Progression in Human Cells Supplementary Figures Supplementary Figure 1. NDR1 interacts with HP1. (a) Immunoprecipitation using
SUPPLEMENTARY INFORMATION Dynamic Phosphorylation of HP1 Regulates Mitotic Progression in Human Cells Supplementary Figures Supplementary Figure 1. NDR1 interacts with HP1. (a) Immunoprecipitation using
Parthanatos mediates AIMP2-activated age-dependent dopaminergic neuronal loss
 SUPPLEMENTARY INFORMATION Parthanatos mediates AIMP2-activated age-dependent dopaminergic neuronal loss Yunjong Lee, Senthilkumar S. Karuppagounder, Joo-Ho Shin, Yun-Il Lee, Han Seok Ko, Debbie Swing,
SUPPLEMENTARY INFORMATION Parthanatos mediates AIMP2-activated age-dependent dopaminergic neuronal loss Yunjong Lee, Senthilkumar S. Karuppagounder, Joo-Ho Shin, Yun-Il Lee, Han Seok Ko, Debbie Swing,
Table S1. Primer sequences
 Table S1. Primer sequences Primers for quantitative PCR Tgf 1 Forward Tgf 1 Reverse Tgf 2Forward Tgf 2Reverse Tgf 3 Forward Tgf 3 Reverse Tgf r1 Forward Tgf r1 Reverse Tgf r2 Forward Tgf r2 Reverse Thbs1
Table S1. Primer sequences Primers for quantitative PCR Tgf 1 Forward Tgf 1 Reverse Tgf 2Forward Tgf 2Reverse Tgf 3 Forward Tgf 3 Reverse Tgf r1 Forward Tgf r1 Reverse Tgf r2 Forward Tgf r2 Reverse Thbs1
supplementary information
 DOI: 1.138/ncb1839 a b Control 1 2 3 Control 1 2 3 Fbw7 Smad3 1 2 3 4 1 2 3 4 c d IGF-1 IGF-1Rβ IGF-1Rβ-P Control / 1 2 3 4 Real-time RT-PCR Relative quantity (IGF-1/ mrna) 2 1 IGF-1 1 2 3 4 Control /
DOI: 1.138/ncb1839 a b Control 1 2 3 Control 1 2 3 Fbw7 Smad3 1 2 3 4 1 2 3 4 c d IGF-1 IGF-1Rβ IGF-1Rβ-P Control / 1 2 3 4 Real-time RT-PCR Relative quantity (IGF-1/ mrna) 2 1 IGF-1 1 2 3 4 Control /
Technical Review. Real time PCR
 Technical Review Real time PCR Normal PCR: Analyze with agarose gel Normal PCR vs Real time PCR Real-time PCR, also known as quantitative PCR (qpcr) or kinetic PCR Key feature: Used to amplify and simultaneously
Technical Review Real time PCR Normal PCR: Analyze with agarose gel Normal PCR vs Real time PCR Real-time PCR, also known as quantitative PCR (qpcr) or kinetic PCR Key feature: Used to amplify and simultaneously
Gene Expression Technology
 Gene Expression Technology Bing Zhang Department of Biomedical Informatics Vanderbilt University bing.zhang@vanderbilt.edu Gene expression Gene expression is the process by which information from a gene
Gene Expression Technology Bing Zhang Department of Biomedical Informatics Vanderbilt University bing.zhang@vanderbilt.edu Gene expression Gene expression is the process by which information from a gene
Wnt16 smact merge VK/AB
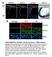 A WT Wnt6 smact merge VK/A KO ctrl IgG WT KO Wnt6 smact DAPI SUPPLEMENTAL FIGURE I: Wnt6 expression in MGP-deficient aortae. Immunostaining for Wnt6 and smooth muscle actin (smact) in aortae from 7 day
A WT Wnt6 smact merge VK/A KO ctrl IgG WT KO Wnt6 smact DAPI SUPPLEMENTAL FIGURE I: Wnt6 expression in MGP-deficient aortae. Immunostaining for Wnt6 and smooth muscle actin (smact) in aortae from 7 day
Short hairpin RNA (shrna) against MMP14. Lentiviral plasmids containing shrna
 Supplemental Materials and Methods Short hairpin RNA (shrna) against MMP14. Lentiviral plasmids containing shrna (Mission shrna, Sigma) against mouse MMP14 were transfected into HEK293 cells using FuGene6
Supplemental Materials and Methods Short hairpin RNA (shrna) against MMP14. Lentiviral plasmids containing shrna (Mission shrna, Sigma) against mouse MMP14 were transfected into HEK293 cells using FuGene6
Figure 1a RFLP from mouse tissue, splicing of mh2a1 isoforms.
 Figure 1a RFLP from mouse tissue, splicing of mh2a1 isoforms. Samples order in the frame: Control 1, control 2, Skeletal muscle, liver, brain, testis, kidney, spleen, small intestin, big intestin, lung.
Figure 1a RFLP from mouse tissue, splicing of mh2a1 isoforms. Samples order in the frame: Control 1, control 2, Skeletal muscle, liver, brain, testis, kidney, spleen, small intestin, big intestin, lung.
X2-C/X1-Y X2-C/VCAM-Y. FRET efficiency. Ratio YFP/CFP
 FRET efficiency.7.6..4.3.2 X2-C/X1-Y X2-C/VCAM-Y.1 1 2 3 Ratio YFP/CFP Supplemental Data 1. Analysis of / heterodimers in live cells using FRET. FRET saturation curves were obtained using cells transiently
FRET efficiency.7.6..4.3.2 X2-C/X1-Y X2-C/VCAM-Y.1 1 2 3 Ratio YFP/CFP Supplemental Data 1. Analysis of / heterodimers in live cells using FRET. FRET saturation curves were obtained using cells transiently
Percent survival. Supplementary fig. S3 A.
 Supplementary fig. S3 A. B. 100 Percent survival 80 60 40 20 Ml 0 0 100 C. Fig. S3 Comparison of leukaemia incidence rate in the triple targeted chimaeric mice and germline-transmission translocator mice
Supplementary fig. S3 A. B. 100 Percent survival 80 60 40 20 Ml 0 0 100 C. Fig. S3 Comparison of leukaemia incidence rate in the triple targeted chimaeric mice and germline-transmission translocator mice
Supplementary Material
 Fig S1. Interactions of OsDof24 with the OsC4PPDK promoter in rice protoplasts (a) Loss-of-function analysis of the OsC4PPDK promoter Effects of OsDof24 overexpression and the mapping of OsDof24-binding
Fig S1. Interactions of OsDof24 with the OsC4PPDK promoter in rice protoplasts (a) Loss-of-function analysis of the OsC4PPDK promoter Effects of OsDof24 overexpression and the mapping of OsDof24-binding
Supplemental Information. The TRAIL-Induced Cancer Secretome. Promotes a Tumor-Supportive Immune. Microenvironment via CCR2
 Molecular Cell, Volume 65 Supplemental Information The TRAIL-Induced Cancer Secretome Promotes a Tumor-Supportive Immune Microenvironment via CCR2 Torsten Hartwig, Antonella Montinaro, Silvia von Karstedt,
Molecular Cell, Volume 65 Supplemental Information The TRAIL-Induced Cancer Secretome Promotes a Tumor-Supportive Immune Microenvironment via CCR2 Torsten Hartwig, Antonella Montinaro, Silvia von Karstedt,
Supplemental Information
 Supplemental Information Itemized List Materials and Methods, Related to Supplemental Figures S5A-C and S6. Supplemental Figure S1, Related to Figures 1 and 2. Supplemental Figure S2, Related to Figure
Supplemental Information Itemized List Materials and Methods, Related to Supplemental Figures S5A-C and S6. Supplemental Figure S1, Related to Figures 1 and 2. Supplemental Figure S2, Related to Figure
Supplemental Information. Loss of MicroRNA-7 Regulation Leads. to a-synuclein Accumulation and. Dopaminergic Neuronal Loss In Vivo
 YMTHE, Volume 25 Supplemental Information Loss of MicroRNA-7 Regulation Leads to a-synuclein Accumulation and Dopaminergic Neuronal Loss In Vivo Kirsty J. McMillan, Tracey K. Murray, Nora Bengoa-Vergniory,
YMTHE, Volume 25 Supplemental Information Loss of MicroRNA-7 Regulation Leads to a-synuclein Accumulation and Dopaminergic Neuronal Loss In Vivo Kirsty J. McMillan, Tracey K. Murray, Nora Bengoa-Vergniory,
RNA-Guided Gene Activation by CRISPR-Cas9-Based Transcription Factors
 Supplementary Information RNA-Guided Gene Activation by CRISPR-Cas9-Based Transcription Factors Pablo Perez-Pinera 1, Daniel D. Kocak 1, Christopher M. Vockley 2,3, Andrew F. Adler 1, Ami M. Kabadi 1,
Supplementary Information RNA-Guided Gene Activation by CRISPR-Cas9-Based Transcription Factors Pablo Perez-Pinera 1, Daniel D. Kocak 1, Christopher M. Vockley 2,3, Andrew F. Adler 1, Ami M. Kabadi 1,
Supplemental Information. HEXIM1 and NEAT1 Long Non-coding RNA Form. a Multi-subunit Complex that Regulates. DNA-Mediated Innate Immune Response
 Molecular Cell, Volume 67 Supplemental Information HEXIM1 and NEAT1 Long Non-coding RNA Form a Multi-subunit Complex that Regulates DNA-Mediated Innate Immune Response Mehdi Morchikh, Alexandra Cribier,
Molecular Cell, Volume 67 Supplemental Information HEXIM1 and NEAT1 Long Non-coding RNA Form a Multi-subunit Complex that Regulates DNA-Mediated Innate Immune Response Mehdi Morchikh, Alexandra Cribier,
SUPPLEMENTAL MATERIALS
 SUPPLEMENL MERILS Eh-seq: RISPR epitope tagging hip-seq of DN-binding proteins Daniel Savic, E. hristopher Partridge, Kimberly M. Newberry, Sophia. Smith, Sarah K. Meadows, rian S. Roberts, Mark Mackiewicz,
SUPPLEMENL MERILS Eh-seq: RISPR epitope tagging hip-seq of DN-binding proteins Daniel Savic, E. hristopher Partridge, Kimberly M. Newberry, Sophia. Smith, Sarah K. Meadows, rian S. Roberts, Mark Mackiewicz,
Supplemental figures Supplemental Figure 1: Fluorescence recovery for FRAP experiments depicted in Figure 1.
 Supplemental figures Supplemental Figure 1: Fluorescence recovery for FRAP experiments depicted in Figure 1. Percent of original fluorescence was plotted as a function of time following photobleaching
Supplemental figures Supplemental Figure 1: Fluorescence recovery for FRAP experiments depicted in Figure 1. Percent of original fluorescence was plotted as a function of time following photobleaching
Supplementary Information
 Supplementary Information MLL histone methylases regulate expression of HDLR- in presence of estrogen and control plasma cholesterol in vivo Khairul I. Ansari 1, Sahba Kasiri 1, Imran Hussain 1, Samara
Supplementary Information MLL histone methylases regulate expression of HDLR- in presence of estrogen and control plasma cholesterol in vivo Khairul I. Ansari 1, Sahba Kasiri 1, Imran Hussain 1, Samara
SANTA CRUZ BIOTECHNOLOGY, INC.
 TECHNICAL SERVICE GUIDE: Western Blotting 2. What size bands were expected and what size bands were detected? 3. Was the blot blank or was a dark background or non-specific bands seen? 4. Did this same
TECHNICAL SERVICE GUIDE: Western Blotting 2. What size bands were expected and what size bands were detected? 3. Was the blot blank or was a dark background or non-specific bands seen? 4. Did this same
Molecular analysis and biological implications of STAT3 signal transduction Schuringa, Jan Jacob
 University of Groningen Molecular analysis and biological implications of STAT3 signal transduction Schuringa, Jan Jacob IMPORTANT NOTE: You are advised to consult the publisher's version (publisher's
University of Groningen Molecular analysis and biological implications of STAT3 signal transduction Schuringa, Jan Jacob IMPORTANT NOTE: You are advised to consult the publisher's version (publisher's
Quantitative Real Time PCR USING SYBR GREEN
 Quantitative Real Time PCR USING SYBR GREEN SYBR Green SYBR Green is a cyanine dye that binds to double stranded DNA. When it is bound to D.S. DNA it has a much greater fluorescence than when bound to
Quantitative Real Time PCR USING SYBR GREEN SYBR Green SYBR Green is a cyanine dye that binds to double stranded DNA. When it is bound to D.S. DNA it has a much greater fluorescence than when bound to
CASE-STUDY- VALIDATION of PCR based methodology. Beata Surmacz-Cordle Senior Analytical Development Scientist
 CASE-STUDY- VALIDATION of PCR based methodology Beata Surmacz-Cordle Senior Analytical Development Scientist UK RMP Pluripotent Stem Cell Platform Validation Workshop 2 nd June 2016 RT-qPCR assay for detection
CASE-STUDY- VALIDATION of PCR based methodology Beata Surmacz-Cordle Senior Analytical Development Scientist UK RMP Pluripotent Stem Cell Platform Validation Workshop 2 nd June 2016 RT-qPCR assay for detection
GUGAUAAUGGAGCGAGAUUUUCUGUUGUGCUUGAUCUAACCAUGUGCUUGCGAGGUAUGA GAAAAACAUGGUUCCGUCAAGCACCAUGGAACGUCACGCAGCUUUCUACA
 Precursor mmu-mir-8- GUGAUAAUGGAGCGAGAUUUUCUGUUGUGCUUGAUCUAACCAUGUGCUUGCGAGGUAUGA GAAAAACAUGGUUCCGUCAAGCACCAUGGAACGUCACGCAGCUUUCUACA Precursor mmu-mir-8- GACCAGUUGCCGCGGGGCUUUCCUUUGUGCUUGAUCUAACCAUGUGGUGGAACGAUGGAA
Precursor mmu-mir-8- GUGAUAAUGGAGCGAGAUUUUCUGUUGUGCUUGAUCUAACCAUGUGCUUGCGAGGUAUGA GAAAAACAUGGUUCCGUCAAGCACCAUGGAACGUCACGCAGCUUUCUACA Precursor mmu-mir-8- GACCAGUUGCCGCGGGGCUUUCCUUUGUGCUUGAUCUAACCAUGUGGUGGAACGAUGGAA
The MAP Kinase Family
 The MAP Kinase Family Extracellular stimuli Classical MAP kinases Atypical MAP kinases MAPKKK MLK1/2/3/7; LZK RAF-1/A/B TAK1; TPL2 c-mos MEKK1-4; DLK ASK1/2; MLTK TAO1/2 ASK1 TAK1 MEKK1-4 MEKK2/3 TPL2???
The MAP Kinase Family Extracellular stimuli Classical MAP kinases Atypical MAP kinases MAPKKK MLK1/2/3/7; LZK RAF-1/A/B TAK1; TPL2 c-mos MEKK1-4; DLK ASK1/2; MLTK TAO1/2 ASK1 TAK1 MEKK1-4 MEKK2/3 TPL2???
Galina Gabriely, Ph.D. BWH/HMS
 Galina Gabriely, Ph.D. BWH/HMS Email: ggabriely@rics.bwh.harvard.edu Outline: microrna overview microrna expression analysis microrna functional analysis microrna (mirna) Characteristics mirnas discovered
Galina Gabriely, Ph.D. BWH/HMS Email: ggabriely@rics.bwh.harvard.edu Outline: microrna overview microrna expression analysis microrna functional analysis microrna (mirna) Characteristics mirnas discovered
Therapeutic RNA editing by Axiomer editing oligonucleotides
 Therapeutic RN editing by xiomer editing oligonucleotides Forward looking statements This presentation contains forward-looking statements that involve substantial risks and uncertainties. ll statements,
Therapeutic RN editing by xiomer editing oligonucleotides Forward looking statements This presentation contains forward-looking statements that involve substantial risks and uncertainties. ll statements,
Supplemental figure 1
 Supplemental figure 1 0.3 Copy number ratio relative to beta actin 0.25 0.2 0.15 0.1 0.05 0 Series1 Series2 Series3 0 1 PC3 BP 2 DU145 BP 3 PC3 CMV 4 DU145 CMV Supplemental figure 1. qpcr to quantify the
Supplemental figure 1 0.3 Copy number ratio relative to beta actin 0.25 0.2 0.15 0.1 0.05 0 Series1 Series2 Series3 0 1 PC3 BP 2 DU145 BP 3 PC3 CMV 4 DU145 CMV Supplemental figure 1. qpcr to quantify the
Supplementary Figure Legend
 Supplementary Figure Legend Supplementary Figure S1. Effects of MMP-1 silencing on HEp3-hi/diss cell proliferation in 2D and 3D culture conditions. (A) Downregulation of MMP-1 expression in HEp3-hi/diss
Supplementary Figure Legend Supplementary Figure S1. Effects of MMP-1 silencing on HEp3-hi/diss cell proliferation in 2D and 3D culture conditions. (A) Downregulation of MMP-1 expression in HEp3-hi/diss
Supplementary Information Design of small molecule-responsive micrornas based on structural requirements for Drosha processing
 Supplementary Information Design of small molecule-responsive micrornas based on structural requirements for Drosha processing Chase L. Beisel, Yvonne Y. Chen, Stephanie J. Culler, Kevin G. Hoff, & Christina
Supplementary Information Design of small molecule-responsive micrornas based on structural requirements for Drosha processing Chase L. Beisel, Yvonne Y. Chen, Stephanie J. Culler, Kevin G. Hoff, & Christina
HCT116 SW48 Nutlin: p53
 Figure S HCT6 SW8 Nutlin: - + - + p GAPDH Figure S. Nutlin- treatment induces p protein. HCT6 and SW8 cells were left untreated or treated for 8 hr with Nutlin- ( µm) to up-regulate p. Whole cell lysates
Figure S HCT6 SW8 Nutlin: - + - + p GAPDH Figure S. Nutlin- treatment induces p protein. HCT6 and SW8 cells were left untreated or treated for 8 hr with Nutlin- ( µm) to up-regulate p. Whole cell lysates
The microtubule-associated tau protein has intrinsic acetyltransferase activity. Todd J. Cohen, Dave Friedmann, Andrew W. Hwang, Ronen Marmorstein and
 SUPPLEMENTARY INFORMATION: The microtubule-associated tau protein has intrinsic acetyltransferase activity Todd J. Cohen, Dave Friedmann, Andrew W. Hwang, Ronen Marmorstein and Virginia M.Y. Lee Cohen
SUPPLEMENTARY INFORMATION: The microtubule-associated tau protein has intrinsic acetyltransferase activity Todd J. Cohen, Dave Friedmann, Andrew W. Hwang, Ronen Marmorstein and Virginia M.Y. Lee Cohen
ENCODE RBP Antibody Characterization Guidelines
 ENCODE RBP Antibody Characterization Guidelines Approved on November 18, 2016 Background An integral part of the ENCODE Project is to characterize the antibodies used in the experiments. This document
ENCODE RBP Antibody Characterization Guidelines Approved on November 18, 2016 Background An integral part of the ENCODE Project is to characterize the antibodies used in the experiments. This document
Supplemental Data. Steiner et al. Plant Cell. (2012) /tpc
 Supplemental Figure 1. SPY does not interact with free GST. Invitro pull-down assay using E. coli-expressed MBP-SPY and GST, GST-TCP14 and GST-TCP15. MBP-SPY was used as bait and incubated with equal amount
Supplemental Figure 1. SPY does not interact with free GST. Invitro pull-down assay using E. coli-expressed MBP-SPY and GST, GST-TCP14 and GST-TCP15. MBP-SPY was used as bait and incubated with equal amount
Figure S1. Figure S2. Figure S3 HB Anti-FSP27 (COOH-terminal peptide) Ab. Anti-GST-FSP27(45-127) Ab.
 / 36B4 mrna ratio Figure S1 * 2. 1.6 1.2.8 *.4 control TNFα BRL49653 Figure S2 Su bw AT p iw Anti- (COOH-terminal peptide) Ab Blot : Anti-GST-(45-127) Ab β-actin Figure S3 HB2 HW AT BA T Figure S4 A TAG
/ 36B4 mrna ratio Figure S1 * 2. 1.6 1.2.8 *.4 control TNFα BRL49653 Figure S2 Su bw AT p iw Anti- (COOH-terminal peptide) Ab Blot : Anti-GST-(45-127) Ab β-actin Figure S3 HB2 HW AT BA T Figure S4 A TAG
IKK is a therapeutic target in KRAS-induced lung cancer with disrupted p53 activity
 IKK is a therapeutic target in KRAS-induced lung cancer with disrupted p5 activity H6 5 5 H58 A59 H6 H58 A59 anti-ikkα anti-ikkβ anti-panras anti-gapdh anti-ikkα anti-ikkβ anti-panras anti-gapdh anti-ikkα
IKK is a therapeutic target in KRAS-induced lung cancer with disrupted p5 activity H6 5 5 H58 A59 H6 H58 A59 anti-ikkα anti-ikkβ anti-panras anti-gapdh anti-ikkα anti-ikkβ anti-panras anti-gapdh anti-ikkα
Junhong Han, Yu-ichi Tsukada, Eiji Hara, Naomi Kitamura, and Toshiaki Tanaka
 THE JOURNAL OF BIOLOGICAL CHEMISTRY Vol. 280, No. 36, Issue of September 9, pp. 31548 31556, 2005 2005 by The American Society for Biochemistry and Molecular Biology, Inc. Printed in U.S.A. Hepatocyte
THE JOURNAL OF BIOLOGICAL CHEMISTRY Vol. 280, No. 36, Issue of September 9, pp. 31548 31556, 2005 2005 by The American Society for Biochemistry and Molecular Biology, Inc. Printed in U.S.A. Hepatocyte
Alternative Cleavage and Polyadenylation of RNA
 Developmental Cell 18 Supplemental Information The Spen Family Protein FPA Controls Alternative Cleavage and Polyadenylation of RNA Csaba Hornyik, Lionel C. Terzi, and Gordon G. Simpson Figure S1, related
Developmental Cell 18 Supplemental Information The Spen Family Protein FPA Controls Alternative Cleavage and Polyadenylation of RNA Csaba Hornyik, Lionel C. Terzi, and Gordon G. Simpson Figure S1, related
reverse transcription! RT 1! RT 2! RT 3!
 Supplementary Figure! Entire workflow repeated 3 times! mirna! stock! -fold dilution series! to yield - copies! per µl in PCR reaction! reverse transcription! RT! pre-pcr! mastermix!!!! 3!!! 3! ddpcr!
Supplementary Figure! Entire workflow repeated 3 times! mirna! stock! -fold dilution series! to yield - copies! per µl in PCR reaction! reverse transcription! RT! pre-pcr! mastermix!!!! 3!!! 3! ddpcr!
Figure S1: NUN preparation yields nascent, unadenylated RNA with a different profile from Total RNA.
 Summary of Supplemental Information Figure S1: NUN preparation yields nascent, unadenylated RNA with a different profile from Total RNA. Figure S2: rrna removal procedure is effective for clearing out
Summary of Supplemental Information Figure S1: NUN preparation yields nascent, unadenylated RNA with a different profile from Total RNA. Figure S2: rrna removal procedure is effective for clearing out
- NaCr. + NaCr. α H3K4me2 α H3K4me3 α H3K9me3 α H3K27me3 α H3K36me3 H3 H2A-2B H4 H3 H2A-2B H4 H3 H2A-2B H4. α Kcr. (rabbit) α Kac.
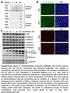 + NaCr NaCr + NaCr NaCr Peptides 10ng 50ng 250ng K α Pan (mouse) Pan (mouse) 10ng 50ng 250ng α Pan (rabbit) C 10ng 50ng 250ng α Pan (mouse) 0 1.25 2.5 5 10 20 40 (mm) NaCr 24h α Pan (rabbit) α K4me2 α
+ NaCr NaCr + NaCr NaCr Peptides 10ng 50ng 250ng K α Pan (mouse) Pan (mouse) 10ng 50ng 250ng α Pan (rabbit) C 10ng 50ng 250ng α Pan (mouse) 0 1.25 2.5 5 10 20 40 (mm) NaCr 24h α Pan (rabbit) α K4me2 α
/04/$15.00/0 Molecular Endocrinology 18(3): Copyright 2004 by The Endocrine Society doi: /me
 0888-8809/04/$15.00/0 Molecular Endocrinology 18(3):588 605 Printed in U.S.A. Copyright 2004 by The Endocrine Society doi: 10.1210/me.2003-0090 Increased Cytochrome P450 17 -Hydroxylase Promoter Function
0888-8809/04/$15.00/0 Molecular Endocrinology 18(3):588 605 Printed in U.S.A. Copyright 2004 by The Endocrine Society doi: 10.1210/me.2003-0090 Increased Cytochrome P450 17 -Hydroxylase Promoter Function
affects the development of newborn neurons Brain Mind Institute and School of Life Sciences, Ecole Polytechnique Fédérale de
 Shedding of neurexin 3β ectodomain by ADAM10 releases a soluble fragment that affects the development of newborn neurons Erika Borcel* a, Magda Palczynska* a, Marine Krzisch b, Mitko Dimitrov a, Giorgio
Shedding of neurexin 3β ectodomain by ADAM10 releases a soluble fragment that affects the development of newborn neurons Erika Borcel* a, Magda Palczynska* a, Marine Krzisch b, Mitko Dimitrov a, Giorgio
Cardiovascular disease remains the leading cause of mortality. Basic Science
 Basic Science The Critical Role of mrna Destabilizing Protein Heterogeneous Nuclear Ribonucleoprotein D in 3 Untranslated Region Mediated Decay of Low-Density Lipoprotein Receptor mrna in Liver Tissue
Basic Science The Critical Role of mrna Destabilizing Protein Heterogeneous Nuclear Ribonucleoprotein D in 3 Untranslated Region Mediated Decay of Low-Density Lipoprotein Receptor mrna in Liver Tissue
Supplementary information
 Supplementary information Inhibition of mitochondrial dysfunction and neuronal cell death in models of Huntington s disease and in HD patients-derived cells Xing Guo 1,#, Marie-Helene Disatnik 3,#, Marie
Supplementary information Inhibition of mitochondrial dysfunction and neuronal cell death in models of Huntington s disease and in HD patients-derived cells Xing Guo 1,#, Marie-Helene Disatnik 3,#, Marie
Protocol for induction of expression and cell lysate production
 Protocol for induction of expression and cell lysate production AV-04 Doxycyclin induction and cell lysate 1.0 Introduction / Description This method is intended for the treatment of the previously transfected
Protocol for induction of expression and cell lysate production AV-04 Doxycyclin induction and cell lysate 1.0 Introduction / Description This method is intended for the treatment of the previously transfected
Fatchiyah
 Fatchiyah Email: fatchiya@yahoo.co.id RNAs: mrna trna rrna RNAi DNAs: Protein: genome DNA cdna mikro-makro mono-poly single-multi Analysis: Identification human and animal disease Finger printing Sexing
Fatchiyah Email: fatchiya@yahoo.co.id RNAs: mrna trna rrna RNAi DNAs: Protein: genome DNA cdna mikro-makro mono-poly single-multi Analysis: Identification human and animal disease Finger printing Sexing
Supplementary Table 1. The Q-PCR primer sequence is summarized in the following table.
 Supplementary Table 1. The Q-PCR primer sequence is summarized in the following table. Name Sequence (5-3 ) Application Flag-u ggactacaaggacgacgatgac Shared upstream primer for all the amplifications of
Supplementary Table 1. The Q-PCR primer sequence is summarized in the following table. Name Sequence (5-3 ) Application Flag-u ggactacaaggacgacgatgac Shared upstream primer for all the amplifications of
Figure 1: TDP-43 is subject to lysine acetylation within the RNA-binding domain a) QBI-293 cells were transfected with TDP-43 in the presence or
 Figure 1: TDP-43 is subject to lysine acetylation within the RNA-binding domain a) QBI-293 cells were transfected with TDP-43 in the presence or absence of the acetyltransferase CBP and acetylated TDP-43
Figure 1: TDP-43 is subject to lysine acetylation within the RNA-binding domain a) QBI-293 cells were transfected with TDP-43 in the presence or absence of the acetyltransferase CBP and acetylated TDP-43
Jae Myoung Suh, Daniel Zeve, Renee McKay, Jin Seo, Zack Salo, Robert Li, Michael Wang, and Jonathan M. Graff
 Cell Metabolism, Volume 6 Supplemental Data Adipose Is a Conserved, Dosage-Sensitive Antiobesity Gene Jae Myoung Suh, Daniel Zeve, Renee McKay, Jin Seo, Zack Salo, Robert Li, Michael Wang, and Jonathan
Cell Metabolism, Volume 6 Supplemental Data Adipose Is a Conserved, Dosage-Sensitive Antiobesity Gene Jae Myoung Suh, Daniel Zeve, Renee McKay, Jin Seo, Zack Salo, Robert Li, Michael Wang, and Jonathan
Microarrays: since we use probes we obviously must know the sequences we are looking at!
 These background are needed: 1. - Basic Molecular Biology & Genetics DNA replication Transcription Post-transcriptional RNA processing Translation Post-translational protein modification Gene expression
These background are needed: 1. - Basic Molecular Biology & Genetics DNA replication Transcription Post-transcriptional RNA processing Translation Post-translational protein modification Gene expression
Increased Expression of Hepatocyte Nuclear Factor 6 Stimulates Hepatocyte Proliferation During Mouse Liver Regeneration
 GASTROENTEROLOGY 2006;130:1283 1300 Increased Expression of Hepatocyte Nuclear Factor 6 Stimulates Hepatocyte Proliferation During Mouse Liver Regeneration YONGJUN TAN, YUICHI YOSHIDA, DOUGLAS E. HUGHES,
GASTROENTEROLOGY 2006;130:1283 1300 Increased Expression of Hepatocyte Nuclear Factor 6 Stimulates Hepatocyte Proliferation During Mouse Liver Regeneration YONGJUN TAN, YUICHI YOSHIDA, DOUGLAS E. HUGHES,
Supplemental Data. LMO4 Controls the Balance between Excitatory. and Inhibitory Spinal V2 Interneurons
 Neuron, Volume 61 Supplemental Data LMO4 Controls the Balance between Excitatory and Inhibitory Spinal V2 Interneurons Kaumudi Joshi, Seunghee Lee, Bora Lee, Jae W. Lee, and Soo-Kyung Lee Supplemental
Neuron, Volume 61 Supplemental Data LMO4 Controls the Balance between Excitatory and Inhibitory Spinal V2 Interneurons Kaumudi Joshi, Seunghee Lee, Bora Lee, Jae W. Lee, and Soo-Kyung Lee Supplemental
GFP CCD2 GFP IP:GFP
 D1 D2 1 75 95 148 178 492 GFP CCD1 CCD2 CCD2 GFP D1 D2 GFP D1 D2 Beclin 1 IB:GFP IP:GFP Supplementary Figure 1: Mapping domains required for binding to HEK293T cells are transfected with EGFP-tagged mutant
D1 D2 1 75 95 148 178 492 GFP CCD1 CCD2 CCD2 GFP D1 D2 GFP D1 D2 Beclin 1 IB:GFP IP:GFP Supplementary Figure 1: Mapping domains required for binding to HEK293T cells are transfected with EGFP-tagged mutant
Supporting Information
 Supporting Information Su et al. 10.1073/pnas.1211604110 SI Materials and Methods Cell Culture and Plasmids. Tera-1 and Tera-2 cells (ATCC: HTB- 105/106) were maintained in McCoy s 5A medium with 15% FBS
Supporting Information Su et al. 10.1073/pnas.1211604110 SI Materials and Methods Cell Culture and Plasmids. Tera-1 and Tera-2 cells (ATCC: HTB- 105/106) were maintained in McCoy s 5A medium with 15% FBS
Roche Molecular Biochemicals Technical Note No. LC 10/2000
 Roche Molecular Biochemicals Technical Note No. LC 10/2000 LightCycler Overview of LightCycler Quantification Methods 1. General Introduction Introduction Content Definitions This Technical Note will introduce
Roche Molecular Biochemicals Technical Note No. LC 10/2000 LightCycler Overview of LightCycler Quantification Methods 1. General Introduction Introduction Content Definitions This Technical Note will introduce
TaqMan Advanced mirna Assays
 PRODUCT BULLETIN TaqMan Advanced mirna Assays TaqMan Advanced mirna Assays Key features Universal reverse transcription (RT) one RT step for all Applied Biosystems TaqMan Advanced mirna Assays Sensitive
PRODUCT BULLETIN TaqMan Advanced mirna Assays TaqMan Advanced mirna Assays Key features Universal reverse transcription (RT) one RT step for all Applied Biosystems TaqMan Advanced mirna Assays Sensitive
Supplementary Figure 1: MYCER protein expressed from the transgene can enhance
 Relative luciferase activity Relative luciferase activity MYC is a critical target FBXW7 MYC Supplementary is a critical Figures target 1-7. FBXW7 Supplementary Material A E-box sequences 1 2 3 4 5 6 HSV-TK
Relative luciferase activity Relative luciferase activity MYC is a critical target FBXW7 MYC Supplementary is a critical Figures target 1-7. FBXW7 Supplementary Material A E-box sequences 1 2 3 4 5 6 HSV-TK
FIG S1: Calibration curves of standards for HPLC detection of Dopachrome (Dopac), Dopamine (DA) and Homovanillic acid (HVA), showing area of peak vs
 FIG S1: Calibration curves of standards for HPLC detection of Dopachrome (Dopac), Dopamine (DA) and Homovanillic acid (HVA), showing area of peak vs amount of standard analysed. Each point represents the
FIG S1: Calibration curves of standards for HPLC detection of Dopachrome (Dopac), Dopamine (DA) and Homovanillic acid (HVA), showing area of peak vs amount of standard analysed. Each point represents the
Mir-X mirna First-Strand Synthesis and SYBR qrt-pcr
 User Manual Mir-X mirna First-Strand Synthesis and SYBR qrt-pcr User Manual United States/Canada 800.662.2566 Asia Pacific +1.650.919.7300 Europe +33.(0)1.3904.6880 Japan +81.(0)77.543.6116 Clontech Laboratories,
User Manual Mir-X mirna First-Strand Synthesis and SYBR qrt-pcr User Manual United States/Canada 800.662.2566 Asia Pacific +1.650.919.7300 Europe +33.(0)1.3904.6880 Japan +81.(0)77.543.6116 Clontech Laboratories,
Supplementary Figure 1: Expression of RNF8, HERC2 and NEURL4 in the cerebellum and knockdown of RNF8 by RNAi (a) Lysates of the cerebellum from rat
 Supplementary Figure 1: Expression of RNF8, HERC2 and NEURL4 in the cerebellum and knockdown of RNF8 by RNAi (a) Lysates of the cerebellum from rat pups at P6, P14, P22, P30 and adult (A) rats were subjected
Supplementary Figure 1: Expression of RNF8, HERC2 and NEURL4 in the cerebellum and knockdown of RNF8 by RNAi (a) Lysates of the cerebellum from rat pups at P6, P14, P22, P30 and adult (A) rats were subjected
Transfection of CRISPR/Cas9 Nuclease NLS ribonucleoprotein (RNP) into adherent mammalian cells using Lipofectamine RNAiMAX
 Transfection of CRISPR/Cas9 Nuclease NLS ribonucleoprotein (RNP) into adherent mammalian cells using Lipofectamine RNAiMAX INTRODUCTION The CRISPR/Cas genome editing system consists of a single guide RNA
Transfection of CRISPR/Cas9 Nuclease NLS ribonucleoprotein (RNP) into adherent mammalian cells using Lipofectamine RNAiMAX INTRODUCTION The CRISPR/Cas genome editing system consists of a single guide RNA
Lecture #1. Introduction to microarray technology
 Lecture #1 Introduction to microarray technology Outline General purpose Microarray assay concept Basic microarray experimental process cdna/two channel arrays Oligonucleotide arrays Exon arrays Comparing
Lecture #1 Introduction to microarray technology Outline General purpose Microarray assay concept Basic microarray experimental process cdna/two channel arrays Oligonucleotide arrays Exon arrays Comparing
Generation of ips-derived model cells for analyses of hair shaft differentiation
 Supplementary Material Generation of ips-derived model cells for analyses of hair shaft differentiation Takumi Kido, Tomoatsu Horigome, Minori Uda, Naoki Adachi, Yohei Hirai Department of Biomedical Chemistry,
Supplementary Material Generation of ips-derived model cells for analyses of hair shaft differentiation Takumi Kido, Tomoatsu Horigome, Minori Uda, Naoki Adachi, Yohei Hirai Department of Biomedical Chemistry,
Supplementary Figure S1. Immunodetection of full-length XA21 and the XA21 C-terminal cleavage product.
 Supplementary Information Supplementary Figure S1. Immunodetection of full-length XA21 and the XA21 C-terminal cleavage product. Total protein extracted from Kitaake wild type and rice plants carrying
Supplementary Information Supplementary Figure S1. Immunodetection of full-length XA21 and the XA21 C-terminal cleavage product. Total protein extracted from Kitaake wild type and rice plants carrying
of Medicine, Zhejiang University, Hangzhou, Zhejiang , China Michigan, 4424B MS-1, 1301 Catherine Street, Ann Arbor, MI 48109, USA.
 Supplemental figure legends: Neddylation inhibitor MLN4924 suppresses growth and migration of human gastric cancer cells Huiyin Lan 1,2#, Zaiming Tang 1#, Hongchuan Jin 2, and Yi Sun 1,3,4* 1 Institute
Supplemental figure legends: Neddylation inhibitor MLN4924 suppresses growth and migration of human gastric cancer cells Huiyin Lan 1,2#, Zaiming Tang 1#, Hongchuan Jin 2, and Yi Sun 1,3,4* 1 Institute
Convoy TM Transfection Reagent
 Convoy TM Transfection Reagent Catalog No.11103 0.25ml (40-80 transfections in 35mm dishes) Catalog No.11105 0.5 ml (80-165 transfections in 35mm dishes) Catalog No.11110 1.0 ml (165-330 transfections
Convoy TM Transfection Reagent Catalog No.11103 0.25ml (40-80 transfections in 35mm dishes) Catalog No.11105 0.5 ml (80-165 transfections in 35mm dishes) Catalog No.11110 1.0 ml (165-330 transfections
Identification of mrna binding proteins that regulate the stability of LDL receptor mrna through AU-rich elements
 Identification of mrna binding proteins that regulate the stability of LDL receptor mrna through AU-rich elements Hai Li, Wei Chen, Yue Zhou, Parveen Abidi, Orr Sharpe, William H. Robinson, Fredric B.
Identification of mrna binding proteins that regulate the stability of LDL receptor mrna through AU-rich elements Hai Li, Wei Chen, Yue Zhou, Parveen Abidi, Orr Sharpe, William H. Robinson, Fredric B.
Supplemental Online Material. The mouse embryonic fibroblast cell line #10 derived from β-arrestin1 -/- -β-arrestin2 -/-
 #1074683s 1 Supplemental Online Material Materials and Methods Cell lines and tissue culture The mouse embryonic fibroblast cell line #10 derived from β-arrestin1 -/- -β-arrestin2 -/- knock-out animals
#1074683s 1 Supplemental Online Material Materials and Methods Cell lines and tissue culture The mouse embryonic fibroblast cell line #10 derived from β-arrestin1 -/- -β-arrestin2 -/- knock-out animals
Isolation, culture, and transfection of primary mammary epithelial organoids
 Supplementary Experimental Procedures Isolation, culture, and transfection of primary mammary epithelial organoids Primary mammary epithelial organoids were prepared from 8-week-old CD1 mice (Charles River)
Supplementary Experimental Procedures Isolation, culture, and transfection of primary mammary epithelial organoids Primary mammary epithelial organoids were prepared from 8-week-old CD1 mice (Charles River)
of signal transduction pathways?
 CELL BIOLOGY Signal Transduction + How do you study activation of signal transduction pathways? + IDENTIFICATION OF PATHWAY-SPECIFIC TRANSCRIPTION ACTIVATION + EASIER, FASTER THAN BLOTTING OR GEL-SHIFT
CELL BIOLOGY Signal Transduction + How do you study activation of signal transduction pathways? + IDENTIFICATION OF PATHWAY-SPECIFIC TRANSCRIPTION ACTIVATION + EASIER, FASTER THAN BLOTTING OR GEL-SHIFT
NTM486-04, NTM174-04,
 Transfection of transformed human trabecular meshwork TM5, and primary human NTM210-05, NTM486-04, NTM174-04, and NTM153-00 cells with Metafectene Easy Adnan Dibas1A,C, Ming Jiang1A,C, Thomas Yorio1A,C.
Transfection of transformed human trabecular meshwork TM5, and primary human NTM210-05, NTM486-04, NTM174-04, and NTM153-00 cells with Metafectene Easy Adnan Dibas1A,C, Ming Jiang1A,C, Thomas Yorio1A,C.
Comparison of Commercial Transfection Reagents: Cell line optimized transfection kits for in vitro cancer research.
 Comparison of Commercial Transfection Reagents: Cell line optimized transfection kits for in vitro cancer research. by Altogen Labs, 11200 Manchaca Road, Suite 203 Austin TX 78748 USA Tel. (512) 433-6177
Comparison of Commercial Transfection Reagents: Cell line optimized transfection kits for in vitro cancer research. by Altogen Labs, 11200 Manchaca Road, Suite 203 Austin TX 78748 USA Tel. (512) 433-6177
Supplemental Materials and Methods
 Supplemental Materials and Methods 125 I-CXCL12 binding assay KG1 cells (2 10 6 ) were preincubated on ice with cold CXCL12 (1.6µg/mL corresponding to 200nM), CXCL11 (1.66µg/mL corresponding to 200nM),
Supplemental Materials and Methods 125 I-CXCL12 binding assay KG1 cells (2 10 6 ) were preincubated on ice with cold CXCL12 (1.6µg/mL corresponding to 200nM), CXCL11 (1.66µg/mL corresponding to 200nM),
SUPPORTING INFORMATION. Optical Control of CRISPR/Cas9 Gene Editing
 SUPPORTING INFORMATION Optical Control of CRISPR/ Gene Editing James Hemphill 1, Erin K. Borchardt 2, Kalyn Brown 1, Aravind Asokan 2, and Alexander Deiters 1 * 1 University of Pittsburgh, Department of
SUPPORTING INFORMATION Optical Control of CRISPR/ Gene Editing James Hemphill 1, Erin K. Borchardt 2, Kalyn Brown 1, Aravind Asokan 2, and Alexander Deiters 1 * 1 University of Pittsburgh, Department of
Western-GUARANTEED Antibody Service FAQ
 Western-GUARANTEED Antibody Service FAQ Content Q 1: When do I need a Western GUARANTEED Peptide Antibody Package?...2 Q 2: Can GenScript provide a Western blot guaranteed antibody?...2 Q 3: Does GenScript
Western-GUARANTEED Antibody Service FAQ Content Q 1: When do I need a Western GUARANTEED Peptide Antibody Package?...2 Q 2: Can GenScript provide a Western blot guaranteed antibody?...2 Q 3: Does GenScript
Supporting Online Material for
 www.sciencemag.org/cgi/content/full/323/5910/124/dc1 Supporting Online Material for Regulation of Neuronal Survival Factor MEF2D by Chaperone-Mediated Autophagy Qian Yang, Hua She, Marla Gearing, Emanuela
www.sciencemag.org/cgi/content/full/323/5910/124/dc1 Supporting Online Material for Regulation of Neuronal Survival Factor MEF2D by Chaperone-Mediated Autophagy Qian Yang, Hua She, Marla Gearing, Emanuela
Nature Neuroscience: doi: /nn Supplementary Figure 1
 Supplementary Figure 1 PCR-genotyping of the three mouse models used in this study and controls for behavioral experiments after semi-chronic Pten inhibition. a-c. DNA from App/Psen1 (a), Pten tg (b) and
Supplementary Figure 1 PCR-genotyping of the three mouse models used in this study and controls for behavioral experiments after semi-chronic Pten inhibition. a-c. DNA from App/Psen1 (a), Pten tg (b) and
