SUPPLEMENTARY INFORMATION
|
|
|
- Eunice Powers
- 5 years ago
- Views:
Transcription
1 ARTICLE NUMBER: 0 DOI: 0.0/NMICROBIOL.0. The host protein CLUH participates in the subnuclear transport of influenza virus ribonucleoprotein complexes Tomomi Ando,, Seiya Yamayoshi, Yuriko Tomita,, Shinji Watanabe,,a, Tokiko Watanabe,, Yoshihiro Kawaoka,, Division of Virology, Department of Microbiology and Immunology, Institute of Medical Science, University of Tokyo, Minato-ku, Tokyo 0-9, Japan Exploratory Research for Advanced Technology Infection-Induced Host Responses Project, Japan Science and Technology Agency, Kawaguchi, Saitama -00, Japan Department of Pathobiological Sciences, School of Veterinary Medicine, University of Wisconsin-Madison, Madison, WI, USA a Current address: Gakuen --, Musashimurayama-shi, Tokyo 0-00, Japan Correspondence should be addressed to S.Y. (yamayo@ims.u-tokyo.ac.jp) and Y.K. (kawaoka@ims.u-tokyo.ac.jp) This PDF file includes Supplementary Figures to NATURE MICROBIOLOGY
2 Supplementary Figures DOI: 0.0/NMICROBIOL.0. a input NFLAG- PB PB PA IP : Normal IgG PB PB PA IP : α-flag PB PB PA b WSN infection ( hpi) input IP IgG α-pb α-cluh α-cluh α-flag α-pb c CLUH/βactin mrna levels (% of sicontrol) sicluh. sicluh. d sicluh. sicluh. α-cluh α-βactin e Cell viability mrna (% (% of of sicontrol) Control) sicluh. sicluh. Virus Titer (log 0 PFU/ml) f MDCK cells sicluh. 0 9 ** ** ** ** P < 0.0 Times (hpi) g sicluh. sicluh. WT sirna resistant c-myc-cluh α-cluh α-c-myc α-βactin h Virus Titer Virus (log titer 0 (log PFU/ml) 0 0 P < 0. ** sicluh. sicluh. WT sirna resistant c-myc-cluh i Virus Titer (log 0 PFU/ml) ** sicluh. siviral NP Virus Titer (log 0 PFU/ml) ** sicluh. siviral NP Virus Titer (log 0 PFU/ml) ** P < 0.0 ** sicluh. siviral NP P HN pdm HN Influenza B Virus Supplementary Fig.. Identification of CLUH as a host protein that contributes to influenza virus replication. (a) Co-immunoprecipitation of endogenous CLUH with NFLAG-PB. NATURE MICROBIOLOGY
3 DOI: 0.0/NMICROBIOL NFLAG-tagged PB (NFLAG-PB), PB (NFLAG-PB), and PA (NFLAG-PA)-expressing plasmids were transfected individually into HEK9 cells and immunoprecipitated by using an anti-flag antibody. (b) Interaction of PB with CLUH in infected cells as assessed by use of co-immunoprecipitation. HEK9 cells were infected with WSN at an MOI of 0, and immunoprecipitated by using an anti-pb antibody. (c, d) Down-regulation of mrna and protein expression of CLUH by treatment with an sirna against CLUH. Cells were transfected with non-targeting control sirna (sicontrol) or sirnas against CLUH (sicluh. or sicluh.). The mrnas encoding CLUH and β-actin were measured by RT-qPCR (c), and CLUH was detected by use of western blotting (d). (e) Cell viability of CLUH-depleted cells. Cells were transfected with sicontrol, sicluh., or sicluh., and then subjected to a cell viability assay. (f) Virus growth in CLUH-depleted MDCK cells, infected with WSN at an MOI of **P < 0.0 (two-way ANOVA with repeated measures followed by Tukey s tests). (g) Protein expression in CLUH-expressing cells. c-myc-cluh(wt)- or c-myc-cluh(sirna-resistant)-stably-expressing cells were transfected with sicontrol or sicluh.. Protein expression was analyzed by use of western blotting. (h) Virus growth in cells expressing sirna-resistant CLUH and treated with sirna against CLUH. c-myc-cluh(wt) or c-myc-cluh(sirna-resistant)-stably-expressing cells were transfected with sicontrol or sicluh., and infected with WSN at an MOI of Virus titers were determined at hpi. **P < 0.0 (one-way ANOVA followed by Tukey s tests). (i) Growth of clinical isolates in CLUH-depleted cells. Cells were transfected with sicontrol, sicluh., or siviral NP, and infected with A/California/0/009 (HNpdm, left), A/Yokohama/UT-KA/0 (HN, middle), and B/Yokohama/UT-KA/0 (Influenza B Virus, right) at an MOI of 0.0. Virus titers were determined at hpi. Since the control siviral NP was designed for influenza A virus, the virus titer in the siviral NP-treated cells infected with NATURE MICROBIOLOGY
4 DOI: 0.0/NMICROBIOL.0. B/Yokohama/UT-KA/0 was comparable to that in the sicontrol-transfected cells. **P < 0.0 (one-way ANOVA followed by Tukey s tests). (c, e, f, h, i) The values represent the means of triplicate experiments ± SD. (a i) The results shown are representative of three independent experiments. NATURE MICROBIOLOGY
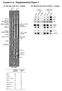
5 i i i DOI: 0.0/NMICROBIOL.0. a α-cluh α-crm α-pb α-np α-ha α-m α-ns α-βactin sicluh hpi CRM (% of sicontrol) hpi PB (% of sicontrol) hpi NP (% of sicontrol) hpi HA (% of sicontrol) hpi M (% of sicontrol) hpi NS (% of sicontrol) hpi b vrna copies (x0 ) sicluh. 0 0 Time (hpi) Supplementary Fig.. The effect of CLUH depletion on the early steps of virus propagation. (a) The expression of viral proteins in CLUH-depleted cells. Cells were transfected with sicontrol or sicluh., and were infected with WSN at an MOI of 0. The cells were then harvested at the indicating times after infection. Proteins were detected by western blotting and then quantitated (normalized to the expression of β-actin). The results shown are representative of three independent experiments. Quantification of the expression level of each protein is indicated at the bottom of the graphs. Error bars represent means ± SD from three independent NATURE MICROBIOLOGY
6 DOI: 0.0/NMICROBIOL.0. experiments. (b) The numbers of vrna copies in CLUH-depleted cells. Cells were transfected with sicontrol or sicluh., and infected with WSN at an MOI of 0. The number of vrna copies was determined by strand-specific real-time RT-qPCR (see Supplementary Methods). The results shown are representative of three independent experiments. The values represent the means of triplicate experiments and are expressed as the means ± SD. NATURE MICROBIOLOGY
7 DOI: 0.0/NMICROBIOL.0. vrnp WSN infection ( hpi) NS Hoechst sicluh. M CLUH Hoechst sicluh. sicluh. HA CLUH Hoechst Supplementary Fig.. The localization of NS, M, and HA in CLUH-depleted cells. Cells were transfected with sicontrol or sicluh., and infected with WSN at an MOI of 0. At hpi, the cells were fixed with % paraformaldehyde, permeabilized with 0.% Triton X-00, and subjected to immunofluorescence analysis. All images were obtained by using confocal microscopy. The results shown are representative of three independent experiments. NATURE MICROBIOLOGY
8 DOI: 0.0/NMICROBIOL.0. MitoTracker CLUH Hoechst 9 hpi Mock Supplementary Fig.. The localization of CLUH and mitochondria in uninfected or infected cells. Cells were infected with WSN at an MOI of 0 or left uninfected. At 9 hpi, the cells were stained by using Mitotracker, fixed with % paraformaldehyde, permeabilized with 0.% Triton X-00, and subjected to immunofluorescence analysis. Bar: 0 µm. All images were obtained by using confocal microscopy. The results shown are representative of three independent experiments. 9 0 NATURE MICROBIOLOGY
9 DOI: 0.0/NMICROBIOL.0. hole cells nuclear matrix CLUH Hoechst CLUH Hoechst LMB + LMB Supplementary Fig.. Intracellular localization of CLUH in uninfected cells. Cells were treated without (top) or with (bottom) Leptomycin B (LMB), an inhibitor of the CRM-dependent nuclear export pathway, for h, and were then fixed with % paraformaldehyde before (whole cells) or after treatment with DNase I and high-salt buffer (nuclear matrix). White lines in the Hoechst channel indicate the nuclei. Bar: 0 µm. All images were obtained by using confocal microscopy. The results shown are representative of three independent experiments. 9 0 NATURE MICROBIOLOGY 9
10 DOI: 0.0/NMICROBIOL.0. a whole cells PB transfection PB CLUH Hoechst PB transfection Cytoplasm Nucleoplasm/Cytoplasm Nucleoplasm n=0 PB / M co-transfection PB / M co-transfection n= CLUH localization in transfected cells (Cell population (%) ) b nuclear matrix M transfection M CLUH Hoechst M transfection fluorescense ( ) fluorescense (+) n=0 PB / M co-transfection PB / M co-transfection CLUH signal in nuclear matrix of transfected cells (Cell population (%) ) n= 9 0 Supplementary Fig.. Intracellular localization of CLUH in PB and/or M over-expressing cells. (a) Intracellular localization of CLUH in PB, or PB and M, over-expressing cells was analyzed by using an anti-pb antibody and an anti-cluh antibody. Percentages of cells in which CLUH was detected in the cytoplasm only, the nucleoplasm only, or both are shown for cells transfected with a PB-expression plasmid alone or with plasmids expressing PB and M. (b) CLUH localization in the nuclear matrix of M, or M and PB, over-expressing cells was analyzed by using an anti-m antibody and an anti-cluh antibody. The numbers of cells positive for CLUH were determined and are expressed as a percentage of the total number of cells. (a, b) The total number of cells counted is indicated to the right of the graph. All images were obtained by using confocal microscopy. Bar: 0 µm. (a, b) The results shown are representative of three independent experiments. 0 NATURE MICROBIOLOGY
11 DOI: 0.0/NMICROBIOL.0. FLAG whole cells CLUH Hoechst FLAG nuclear matrix CLUH Hoechst NFLAG-NS transfection NFLAG-NS transfection NFLAG-M transfection NFLAG-NA transfection NFLAG-HA transfection NFLAG-NP transfection NFLAG-PA transfection NFLAG-PB transfection Supplementary Fig.. The intracellular localization of CLUH in viral protein-overexpressing cells. Cells were transfected with the indicated plasmid for the NATURE MICROBIOLOGY
12 DOI: 0.0/NMICROBIOL.0. expression of N-terminally FLAG-tagged (NFLAG-) viral protein, and then fixed with % paraformaldehyde before (whole cells) or after treatment with DNase I and high-salt buffer (nuclear matrix), and subjected to immunofluorescence analysis by using an anti-flag antibody and an anti-cluh antibody. White lines in the Hoechst channel indicate nuclei. Bar: 0 µm. All images were obtained by using confocal microscopy. The results shown are representative of three independent experiments. NATURE MICROBIOLOGY
13 DOI: 0.0/NMICROBIOL.0. M WSN infection ( hpi) nuclear matrix PML Hoechst HPα CLUH Hoechst HKme CLUH Hoechst 9 Supplementary Fig.. The localization of CLUH and marker proteins of the nuclear compartments in virus-infected cells. Cells were infected with WSN at an MOI of 0. At hpi, the cells were fixed after treatment with DNase I and high-salt buffer (nuclear matrix), and subjected to immunofluorescence analysis by using an anti-cluh or an anti-m antibody and an anti-hpα antibody, an anti-trimethyl Histone H (Lys) (HKme) antibody, or an anti-pml antibody. White lines in the Hoechst channel indicate the nuclei. Bar: 0 µm. All images were obtained by using confocal microscopy. The results shown are representative of three independent experiments. 0 NATURE MICROBIOLOGY
14 DOI: 0.0/NMICROBIOL.0. PB WSN infection ( hpi) nuclear matrix CLUH Hoechst x 9 Supplementary Fig. 9. The co-localization of CLUH and PB in the nuclear matrix of infected cells. Cells were infected with WSN at an MOI of 0. At hpi, the cells were fixed after treatment with DNase I and high-salt buffer (nuclear matrix), and subjected to immunofluorescence analysis by using an anti-cluh and an anti-pb antibody. White lines in the Hoechst channel indicate the nuclei. Bar: 0 µm. The image was obtained by using confocal microscopy. This figure is excerpted from Figure c. The right image is an enlargement of the cell enclosed in the white box. Some of the co-localized PB and CLUH are indicated by the arrowheads. 0 NATURE MICROBIOLOGY
15 DOI: 0.0/NMICROBIOL.0. input M transfection IgG IP α-cluh α-cluh α-m Supplementary Fig. 0. Co-immunoprecipitation of endogenous CLUH with M. M-expressing plasmid was transfected into HEK9 cells and the interaction of M with CLUH was analyzed by co-immunoprecipitation using an anti-cluh antibody. The results shown are representative of three independent experiments. NATURE MICROBIOLOGY
16 DOI: 0.0/NMICROBIOL.0. Mean Intensity of CLUH at the nuclear periphery ** P < 0.0 LMB ( ) " LMB (+) 9 0 Supplementary Fig.. CLUH is enriched at the nuclear periphery upon LMB treatment. Localization of vrnp and CLUH at the nuclear matrix of cells infected with WSN at an MOI of 0 in the presence or absence of LMB was analyzed by using an anti-vrnp antibody and an anti-cluh antibody. Experiments were performed in parallel with those described in the legend to Fig. d. The mean fluorescence intensity at the nuclear periphery was analyzed from the signal in a specific confocal section. The values are expressed as means ± SD (n = 0 cells); **P < 0.0 (one-sided Mann-Whitney test). The results shown are representative of three independent experiments. NATURE MICROBIOLOGY
17 DOI: 0.0/NMICROBIOL.0. Cyt Nuc ch ch0 LMB * Cyt Nuc ch ch0 AS si AS si AS si AS si * AS : sicontrol si : sicluh. α-cluh α-mek/ α-hmgb α-tfiib α-histone H α-pb α-np α-m 9 0 Supplementary Fig.. Confirmation of subcellular fractionation using host marker proteins. Cells were infected with WSN at an MOI of, and treated with LMB from hpi (left), or were transfected with sicontrol or sicluh., and then infected with WSN at an MOI of (right). At hpi, cells were fractionated into cytoplasmic ( Cyt ), nucleoplasmic ( Nuc ), 0 mm NaCl-extractable chromatin ( ch0 ), and 00 mm NaCl-extractable chromatin ( ch00 ) fractions. Fractionation was confirmed by western blotting using the following antibodies against host marker proteins: MEK/ (cytoplasm), HMGB (cytoplasm and nucleoplasm), TFIIB (0 mm NaCl-extractable chromatin), and Histone H (00 mm NaCl-extractable chromatin). The amounts of the viral proteins, including components of the vrnp nuclear export complex such as PB, NP, and M, were not affected by CLUH depletion or LMB treatment, because excess viral proteins are present in infected cells and not all of them form vrnps. *Non-specific bands. Experiments were performed in parallel with those described in the legend to Fig. e,f. The results shown are representative of three independent experiments. NATURE MICROBIOLOGY
18 DOI: 0.0/NMICROBIOL.0. Supplementary Fig.. Full blot images of all western blots. NATURE MICROBIOLOGY
19 DOI: 0.0/NMICROBIOL.0. NATURE MICROBIOLOGY 9
20 DOI: 0.0/NMICROBIOL.0. 0 NATURE MICROBIOLOGY
21 DOI: 0.0/NMICROBIOL.0. NATURE MICROBIOLOGY
Post-translational modification
 Protein expression Western blotting, is a widely used and accepted technique to detect levels of protein expression in a cell or tissue extract. This technique measures protein levels in a biological sample
Protein expression Western blotting, is a widely used and accepted technique to detect levels of protein expression in a cell or tissue extract. This technique measures protein levels in a biological sample
SUPPLEMENTARY INFORMATION
 DOI: 1.1/ncb2918 Supplementary Figure 1 (a) Biotin-dUTP labelling does not affect S phase progression. Cells synchronized in mid-s phase were labelled with biotindutp during a 5 min hypotonic shift (red)
DOI: 1.1/ncb2918 Supplementary Figure 1 (a) Biotin-dUTP labelling does not affect S phase progression. Cells synchronized in mid-s phase were labelled with biotindutp during a 5 min hypotonic shift (red)
Sarker et al. Supplementary Material. Subcellular Fractionation
 Supplementary Material Subcellular Fractionation Transfected 293T cells were harvested with phosphate buffered saline (PBS) and centrifuged at 2000 rpm (500g) for 3 min. The pellet was washed, re-centrifuged
Supplementary Material Subcellular Fractionation Transfected 293T cells were harvested with phosphate buffered saline (PBS) and centrifuged at 2000 rpm (500g) for 3 min. The pellet was washed, re-centrifuged
T H E J O U R N A L O F C E L L B I O L O G Y
 T H E J O U R N A L O F C E L L B I O L O G Y Supplemental material Han et al., http://www.jcb.org/cgi/content/full/jcb.201311007/dc1 Figure S1. SIVA1 interacts with PCNA. (A) HEK293T cells were transiently
T H E J O U R N A L O F C E L L B I O L O G Y Supplemental material Han et al., http://www.jcb.org/cgi/content/full/jcb.201311007/dc1 Figure S1. SIVA1 interacts with PCNA. (A) HEK293T cells were transiently
Phenotypic lentivirus screens to identify functional single domain antibodies
 ARTICLE NUMBER: 16080 DOI: 10.1038/NMICROBIOL.2016.80 Phenotypic lentivirus screens to identify functional single domain antibodies Florian I. Schmidt, Leo Hanke, Benjamin Morin, Rebeccah Brewer, Vesna
ARTICLE NUMBER: 16080 DOI: 10.1038/NMICROBIOL.2016.80 Phenotypic lentivirus screens to identify functional single domain antibodies Florian I. Schmidt, Leo Hanke, Benjamin Morin, Rebeccah Brewer, Vesna
(a) Immunoblotting to show the migration position of Flag-tagged MAVS
 Supplementary Figure 1 Characterization of six MAVS isoforms. (a) Immunoblotting to show the migration position of Flag-tagged MAVS isoforms. HEK293T Mavs -/- cells were transfected with constructs expressing
Supplementary Figure 1 Characterization of six MAVS isoforms. (a) Immunoblotting to show the migration position of Flag-tagged MAVS isoforms. HEK293T Mavs -/- cells were transfected with constructs expressing
Stargazin regulates AMPA receptor trafficking through adaptor protein. complexes during long term depression
 Supplementary Information Stargazin regulates AMPA receptor trafficking through adaptor protein complexes during long term depression Shinji Matsuda, Wataru Kakegawa, Timotheus Budisantoso, Toshihiro Nomura,
Supplementary Information Stargazin regulates AMPA receptor trafficking through adaptor protein complexes during long term depression Shinji Matsuda, Wataru Kakegawa, Timotheus Budisantoso, Toshihiro Nomura,
Nature Immunology: doi: /ni Supplementary Figure 1. Control experiments for Figure 1.
 Supplementary Figure 1 Control experiments for Figure 1. (a) Localization of SETX in untreated or PR8ΔNS1 virus infected A549 cells (4hours). Nuclear (DAPI) and Tubulin staining are shown. SETX antibody
Supplementary Figure 1 Control experiments for Figure 1. (a) Localization of SETX in untreated or PR8ΔNS1 virus infected A549 cells (4hours). Nuclear (DAPI) and Tubulin staining are shown. SETX antibody
supplementary information
 DOI: 10.1038/ncb2156 Figure S1 Depletion of p114rhogef with different sirnas. Caco-2 (a) and HCE (b) cells were transfected with individual sirnas, pools of the two sirnas or the On Target (OnT) sirna
DOI: 10.1038/ncb2156 Figure S1 Depletion of p114rhogef with different sirnas. Caco-2 (a) and HCE (b) cells were transfected with individual sirnas, pools of the two sirnas or the On Target (OnT) sirna
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION Supplementary figures Supplementary Figure 1: Suv39h1, but not Suv39h2, promotes HP1α sumoylation in vivo. In vivo HP1α sumoylation assay. Top: experimental scheme. Middle: we
SUPPLEMENTARY INFORMATION Supplementary figures Supplementary Figure 1: Suv39h1, but not Suv39h2, promotes HP1α sumoylation in vivo. In vivo HP1α sumoylation assay. Top: experimental scheme. Middle: we
Supplementary Figure 1. RAD51 and RAD51 paralogs are enriched spontaneously onto
 Supplementary Figure legends Supplementary Figure 1. and paralogs are enriched spontaneously onto the S-phase chromatin during DN replication. () Chromatin fractionation was carried out as described in
Supplementary Figure legends Supplementary Figure 1. and paralogs are enriched spontaneously onto the S-phase chromatin during DN replication. () Chromatin fractionation was carried out as described in
hnrnp C promotes APP translation by competing with FMRP for APP mrna recruitment to P bodies
 hnrnp C promotes APP translation by competing with for APP mrna recruitment to P bodies Eun Kyung Lee 1, Hyeon Ho Kim 1, Yuki Kuwano 1, Kotb Abdelmohsen 1, Subramanya Srikantan 1, Sarah S. Subaran 2, Marc
hnrnp C promotes APP translation by competing with for APP mrna recruitment to P bodies Eun Kyung Lee 1, Hyeon Ho Kim 1, Yuki Kuwano 1, Kotb Abdelmohsen 1, Subramanya Srikantan 1, Sarah S. Subaran 2, Marc
SUPPLEMENTARY INFORMATION
 The Supplementary Information (SI) Methods Cell culture and transfections H1299, U2OS, 293, HeLa cells were maintained in DMEM medium supplemented with 10% fetal bovine serum. H1299 and 293 cells were
The Supplementary Information (SI) Methods Cell culture and transfections H1299, U2OS, 293, HeLa cells were maintained in DMEM medium supplemented with 10% fetal bovine serum. H1299 and 293 cells were
Supplemental Fig. 1: PEA-15 knockdown efficiency assessed by immunohistochemistry and qpcr
 Supplemental figure legends Supplemental Fig. 1: PEA-15 knockdown efficiency assessed by immunohistochemistry and qpcr A, LβT2 cells were transfected with either scrambled or PEA-15 sirna. Cells were then
Supplemental figure legends Supplemental Fig. 1: PEA-15 knockdown efficiency assessed by immunohistochemistry and qpcr A, LβT2 cells were transfected with either scrambled or PEA-15 sirna. Cells were then
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb3209 Supplementary Figure 1 IR induces the association of FH with chromatin. a, U2OS cells synchronized by thymidine double block (2 mm) underwent no release (G1 phase) or release for 2
DOI: 10.1038/ncb3209 Supplementary Figure 1 IR induces the association of FH with chromatin. a, U2OS cells synchronized by thymidine double block (2 mm) underwent no release (G1 phase) or release for 2
The Human Protein PRR14 Tethers Heterochromatin to the Nuclear Lamina During Interphase and Mitotic Exit
 Cell Reports, Volume 5 Supplemental Information The Human Protein PRR14 Tethers Heterochromatin to the Nuclear Lamina During Interphase and Mitotic Exit Andrey Poleshko, Katelyn M. Mansfield, Caroline
Cell Reports, Volume 5 Supplemental Information The Human Protein PRR14 Tethers Heterochromatin to the Nuclear Lamina During Interphase and Mitotic Exit Andrey Poleshko, Katelyn M. Mansfield, Caroline
ASPP1 Fw GGTTGGGAATCCACGTGTTG ASPP1 Rv GCCATATCTTGGAGCTCTGAGAG
 Supplemental Materials and Methods Plasmids: the following plasmids were used in the supplementary data: pwzl-myc- Lats2 (Aylon et al, 2006), pretrosuper-vector and pretrosuper-shp53 (generous gift of
Supplemental Materials and Methods Plasmids: the following plasmids were used in the supplementary data: pwzl-myc- Lats2 (Aylon et al, 2006), pretrosuper-vector and pretrosuper-shp53 (generous gift of
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb3363 Supplementary Figure 1 Several WNTs bind to the extracellular domains of PKD1. (a) HEK293T cells were co-transfected with indicated plasmids. Flag-tagged proteins were immunoprecipiated
DOI: 10.1038/ncb3363 Supplementary Figure 1 Several WNTs bind to the extracellular domains of PKD1. (a) HEK293T cells were co-transfected with indicated plasmids. Flag-tagged proteins were immunoprecipiated
Supplemental Materials and Methods
 Supplemental Materials and Methods Co-immunoprecipitation (Co-IP) assay Cells were lysed with NETN buffer (20 mm Tris-HCl, ph 8.0, 0 mm NaCl, 1 mm EDT, 0.5% Nonidet P-40) containing 50 mm β-glycerophosphate,
Supplemental Materials and Methods Co-immunoprecipitation (Co-IP) assay Cells were lysed with NETN buffer (20 mm Tris-HCl, ph 8.0, 0 mm NaCl, 1 mm EDT, 0.5% Nonidet P-40) containing 50 mm β-glycerophosphate,
Supplementary Figure 1 PARP1 is involved in regulating the stability of mrnas from pro-inflammatory cytokine/chemokine mediators.
 Supplementary Figure 1 PARP1 is involved in regulating the stability of mrnas from pro-inflammatory cytokine/chemokine mediators. (a) A graphic depiction of the approach to determining the stability of
Supplementary Figure 1 PARP1 is involved in regulating the stability of mrnas from pro-inflammatory cytokine/chemokine mediators. (a) A graphic depiction of the approach to determining the stability of
Supplementary Material
 Supplementary Material Supplementary Methods Cell synchronization. For synchronized cell growth, thymidine was added to 30% confluent U2OS cells to a final concentration of 2.5mM. Cells were incubated
Supplementary Material Supplementary Methods Cell synchronization. For synchronized cell growth, thymidine was added to 30% confluent U2OS cells to a final concentration of 2.5mM. Cells were incubated
Transcriptional regulation of BRCA1 expression by a metabolic switch: Di, Fernandez, De Siervi, Longo, and Gardner. H3K4Me3
 ChIP H3K4Me3 enrichment.25.2.15.1.5 H3K4Me3 H3K4Me3 ctrl H3K4Me3 + E2 NS + E2 1. kb kb +82 kb Figure S1. Estrogen promotes entry of MCF-7 into the cell cycle but does not significantly change activation-associated
ChIP H3K4Me3 enrichment.25.2.15.1.5 H3K4Me3 H3K4Me3 ctrl H3K4Me3 + E2 NS + E2 1. kb kb +82 kb Figure S1. Estrogen promotes entry of MCF-7 into the cell cycle but does not significantly change activation-associated
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1. Analyses of ECTRs by C-circle and T-circle assays.
 Supplementary Figure 1 Analyses of ECTRs by C-circle and T-circle assays. (a) C-circle and (b) T-circle amplification reactions using genomic DNA from different cell lines in the presence (+) or absence
Supplementary Figure 1 Analyses of ECTRs by C-circle and T-circle assays. (a) C-circle and (b) T-circle amplification reactions using genomic DNA from different cell lines in the presence (+) or absence
Nature Structural & Molecular Biology: doi: /nsmb.1583
 Acetylation by GCN5 regulates CDC6 phosphorylation in the S-phase of the cell cycle Roberta Paolinelli 1,2, Ramiro Mendoza-Maldonado 2, Anna Cereseto 1 and Mauro Giacca 2 1 Molecular Biology Laboratory,
Acetylation by GCN5 regulates CDC6 phosphorylation in the S-phase of the cell cycle Roberta Paolinelli 1,2, Ramiro Mendoza-Maldonado 2, Anna Cereseto 1 and Mauro Giacca 2 1 Molecular Biology Laboratory,
Supplementary Figure 1. GST pull-down analysis of the interaction of GST-cIAP1 (A, B), GSTcIAP1
 Legends Supplementary Figure 1. GST pull-down analysis of the interaction of GST- (A, B), GST mutants (B) or GST- (C) with indicated proteins. A, B, Cell lysate from untransfected HeLa cells were loaded
Legends Supplementary Figure 1. GST pull-down analysis of the interaction of GST- (A, B), GST mutants (B) or GST- (C) with indicated proteins. A, B, Cell lysate from untransfected HeLa cells were loaded
SUPPLEMENTARY INFORMATION
 (Supplementary Methods and Materials) GST pull-down assay GST-fusion proteins Fe65 365-533, and Fe65 538-700 were expressed in BL21 bacterial cells and purified with glutathione-agarose beads (Sigma).
(Supplementary Methods and Materials) GST pull-down assay GST-fusion proteins Fe65 365-533, and Fe65 538-700 were expressed in BL21 bacterial cells and purified with glutathione-agarose beads (Sigma).
Supplementary Figure 1, related to Figure 1. GAS5 is highly expressed in the cytoplasm of hescs, and positively correlates with pluripotency.
 Supplementary Figure 1, related to Figure 1. GAS5 is highly expressed in the cytoplasm of hescs, and positively correlates with pluripotency. (a) Transfection of different concentration of GAS5-overexpressing
Supplementary Figure 1, related to Figure 1. GAS5 is highly expressed in the cytoplasm of hescs, and positively correlates with pluripotency. (a) Transfection of different concentration of GAS5-overexpressing
T H E J O U R N A L O F C E L L B I O L O G Y
 T H E J O U R N A L O F C E L L B I O L O G Y Supplemental material Rainero et al., http://www.jcb.org/cgi/content/full/jcb.201109112/dc1 Figure S1. The expression of DGK- is reduced upon transfection
T H E J O U R N A L O F C E L L B I O L O G Y Supplemental material Rainero et al., http://www.jcb.org/cgi/content/full/jcb.201109112/dc1 Figure S1. The expression of DGK- is reduced upon transfection
Supplementary Figure 1. Drawing of spinal cord open-book preparations and DiI tracing. Nature Neuroscience: doi: /nn.3893
 Supplementary Figure 1 Drawing of spinal cord open-book preparations and DiI tracing. Supplementary Figure 2 In ovo electroporation of dominant-negative PlexinA1 in commissural neurons induces midline
Supplementary Figure 1 Drawing of spinal cord open-book preparations and DiI tracing. Supplementary Figure 2 In ovo electroporation of dominant-negative PlexinA1 in commissural neurons induces midline
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb2271 Supplementary Figure a! WM266.4 mock WM266.4 #7 sirna WM266.4 #10 sirna SKMEL28 mock SKMEL28 #7 sirna SKMEL28 #10 sirna WM1361 mock WM1361 #7 sirna WM1361 #10 sirna 9 WM266. WM136
DOI: 10.1038/ncb2271 Supplementary Figure a! WM266.4 mock WM266.4 #7 sirna WM266.4 #10 sirna SKMEL28 mock SKMEL28 #7 sirna SKMEL28 #10 sirna WM1361 mock WM1361 #7 sirna WM1361 #10 sirna 9 WM266. WM136
Supporting Information
 Supporting Information Stavru et al. 0.073/pnas.357840 SI Materials and Methods Immunofluorescence. For immunofluorescence, cells were fixed for 0 min in 4% (wt/vol) paraformaldehyde (Electron Microscopy
Supporting Information Stavru et al. 0.073/pnas.357840 SI Materials and Methods Immunofluorescence. For immunofluorescence, cells were fixed for 0 min in 4% (wt/vol) paraformaldehyde (Electron Microscopy
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Endogenous gene tagging to study subcellular localization and chromatin binding. a, b, Schematic of experimental set-up to endogenously tag RNAi factors using the CRISPR Cas9 technology,
Supplementary Figure 1 Endogenous gene tagging to study subcellular localization and chromatin binding. a, b, Schematic of experimental set-up to endogenously tag RNAi factors using the CRISPR Cas9 technology,
Short hairpin RNA (shrna) against MMP14. Lentiviral plasmids containing shrna
 Supplemental Materials and Methods Short hairpin RNA (shrna) against MMP14. Lentiviral plasmids containing shrna (Mission shrna, Sigma) against mouse MMP14 were transfected into HEK293 cells using FuGene6
Supplemental Materials and Methods Short hairpin RNA (shrna) against MMP14. Lentiviral plasmids containing shrna (Mission shrna, Sigma) against mouse MMP14 were transfected into HEK293 cells using FuGene6
TRIM31 is recruited to mitochondria after infection with SeV.
 Supplementary Figure 1 TRIM31 is recruited to mitochondria after infection with SeV. (a) Confocal microscopy of TRIM31-GFP transfected into HEK293T cells for 24 h followed with SeV infection for 6 h. MitoTracker
Supplementary Figure 1 TRIM31 is recruited to mitochondria after infection with SeV. (a) Confocal microscopy of TRIM31-GFP transfected into HEK293T cells for 24 h followed with SeV infection for 6 h. MitoTracker
Supplementary Fig. 1 Identification of Nedd4 as an IRS-2-associated protein in camp-treated FRTL-5 cells.
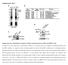 Supplementary Fig. 1 Supplementary Fig. 1 Identification of Nedd4 as an IRS-2-associated protein in camp-treated FRTL-5 cells. (a) FRTL-5 cells were treated with 1 mm dibutyryl camp for 24 h, and the lysates
Supplementary Fig. 1 Supplementary Fig. 1 Identification of Nedd4 as an IRS-2-associated protein in camp-treated FRTL-5 cells. (a) FRTL-5 cells were treated with 1 mm dibutyryl camp for 24 h, and the lysates
Supplementary Figure 1. α-synuclein is truncated in PD and LBD brains. Nature Structural & Molecular Biology: doi: /nsmb.
 Supplementary Figure 1 α-synuclein is truncated in PD and LBD brains. (a) Specificity of anti-n103 antibody. Anti-N103 antibody was coated on an ELISA plate and different concentrations of full-length
Supplementary Figure 1 α-synuclein is truncated in PD and LBD brains. (a) Specificity of anti-n103 antibody. Anti-N103 antibody was coated on an ELISA plate and different concentrations of full-length
Four different active promoter genes were chosen, ATXN7L2, PSRC1, CELSR2 and
 SUPPLEMENTARY MATERIALS AND METHODS Chromatin Immunoprecipitation for qpcr analysis Four different active promoter genes were chosen, ATXN7L2, PSRC1, CELSR2 and IL24, all located on chromosome 1. Primer
SUPPLEMENTARY MATERIALS AND METHODS Chromatin Immunoprecipitation for qpcr analysis Four different active promoter genes were chosen, ATXN7L2, PSRC1, CELSR2 and IL24, all located on chromosome 1. Primer
Flag-Rac Vector V12 V12 N17 C40. Vector C40 pakt (T308) Akt1. Myc-DN-PAK1 (N-SP)
 a b FlagRac FlagRac V2 V2 N7 C4 V2 V2 N7 C4 p (T38) p (S99, S24) p Flag (Rac) NIH 3T3 COS c +Serum p (T38) MycDN (NSP) Mycp27 3 6 2 3 6 2 3 6 2 min p Myc ( or p27) Figure S (a) Effects of Rac mutants on
a b FlagRac FlagRac V2 V2 N7 C4 V2 V2 N7 C4 p (T38) p (S99, S24) p Flag (Rac) NIH 3T3 COS c +Serum p (T38) MycDN (NSP) Mycp27 3 6 2 3 6 2 3 6 2 min p Myc ( or p27) Figure S (a) Effects of Rac mutants on
SUPPLEMENTAL MATERIAL. Carvajal et al. Supplemental Table 1. List of quantitative PCR primers used for cdna analyses and
 UPPLEMEAL MATERIAL Carvajal et al. upplemental Table 1. List of quantitative PCR primers used for cdna analyses and chromatin immunoprecipitation assays. Figure 1. DNA damage-induced transcriptional repression
UPPLEMEAL MATERIAL Carvajal et al. upplemental Table 1. List of quantitative PCR primers used for cdna analyses and chromatin immunoprecipitation assays. Figure 1. DNA damage-induced transcriptional repression
Supplementary Figure 1. Intracellular distribution of the EPE peptide. HeLa cells were serum-starved (16 h, 0.1%), and treated with EPE peptide,
 Supplementary Figure 1. Intracellular distribution of the EPE peptide. HeLa cells were serum-starved (16 h, 0.1%), and treated with EPE peptide, conjugated with either TAT or Myristic acid and biotin for
Supplementary Figure 1. Intracellular distribution of the EPE peptide. HeLa cells were serum-starved (16 h, 0.1%), and treated with EPE peptide, conjugated with either TAT or Myristic acid and biotin for
SUPPLEMENTAL FIGURES AND TABLES
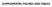 SUPPLEMENTAL FIGURES AND TABLES A B Flag-ALDH1A1 IP: α-ac HEK293T WT 91R 128R 252Q 367R 41/ 419R 435R 495R 412R C Flag-ALDH1A1 NAM IP: HEK293T + + - + D NAM - + + E Relative ALDH1A1 activity 1..8.6.4.2
SUPPLEMENTAL FIGURES AND TABLES A B Flag-ALDH1A1 IP: α-ac HEK293T WT 91R 128R 252Q 367R 41/ 419R 435R 495R 412R C Flag-ALDH1A1 NAM IP: HEK293T + + - + D NAM - + + E Relative ALDH1A1 activity 1..8.6.4.2
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION Dynamic Phosphorylation of HP1 Regulates Mitotic Progression in Human Cells Supplementary Figures Supplementary Figure 1. NDR1 interacts with HP1. (a) Immunoprecipitation using
SUPPLEMENTARY INFORMATION Dynamic Phosphorylation of HP1 Regulates Mitotic Progression in Human Cells Supplementary Figures Supplementary Figure 1. NDR1 interacts with HP1. (a) Immunoprecipitation using
This is the author's accepted version of the manuscript.
 This is the author's accepted version of the manuscript. The definitive version is published in Nature Communications Online Edition: 2015/4/16 (Japan time), doi:10.1038/ncomms7780. The final version published
This is the author's accepted version of the manuscript. The definitive version is published in Nature Communications Online Edition: 2015/4/16 (Japan time), doi:10.1038/ncomms7780. The final version published
Supplemental Data. Zhang et al. Plant Cell (2014) /tpc
 Supplemental Data. Zhang et al. Plant Cell (214) 1.115/tpc.114.134163 55 - T C N SDIRIP1-GFP 35-25 - Psb 18 - Histone H3 Supplemental Figure 1. Detection of SDIRIP1-GFP in the nuclear fraction by Western
Supplemental Data. Zhang et al. Plant Cell (214) 1.115/tpc.114.134163 55 - T C N SDIRIP1-GFP 35-25 - Psb 18 - Histone H3 Supplemental Figure 1. Detection of SDIRIP1-GFP in the nuclear fraction by Western
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb3240 Supplementary Figure 1 GBM cell lines display similar levels of p100 to p52 processing but respond differentially to TWEAK-induced TERT expression according to TERT promoter mutation
DOI: 10.1038/ncb3240 Supplementary Figure 1 GBM cell lines display similar levels of p100 to p52 processing but respond differentially to TWEAK-induced TERT expression according to TERT promoter mutation
At E17.5, the embryos were rinsed in phosphate-buffered saline (PBS) and immersed in
 Supplementary Materials and Methods Barrier function assays At E17.5, the embryos were rinsed in phosphate-buffered saline (PBS) and immersed in acidic X-gal mix (100 mm phosphate buffer at ph4.3, 3 mm
Supplementary Materials and Methods Barrier function assays At E17.5, the embryos were rinsed in phosphate-buffered saline (PBS) and immersed in acidic X-gal mix (100 mm phosphate buffer at ph4.3, 3 mm
a. Increased active HPSE (50 kda) protein expression upon infection with multiple herpes viruses: bovine
 Fold change vs uninfeced MOI 0.1 MOI 1 MOI 0.1 MOI 1 MOI 0.001 MOI 0.01 Fold change vs uninfected a 6 4 2 50 kda HPSE * ** * * BHV PRV HSV-2 333 ** * 0 - + + - + + - + + b 50 kda HPSE 3 2 ** *** mock inf
Fold change vs uninfeced MOI 0.1 MOI 1 MOI 0.1 MOI 1 MOI 0.001 MOI 0.01 Fold change vs uninfected a 6 4 2 50 kda HPSE * ** * * BHV PRV HSV-2 333 ** * 0 - + + - + + - + + b 50 kda HPSE 3 2 ** *** mock inf
B Western blot of the p150caf-1 complex
 Loyola et al., Supplementary Figure 1 A MS data of the HP1α complex RIF1 SPT6 SMARCA4 POGZ CAF-1 p150 KAP-1 HP1α 200 116 97.1 CAF-1 p150 tnasp Importin4 B Western blot of the p150caf-1 complex Input mock
Loyola et al., Supplementary Figure 1 A MS data of the HP1α complex RIF1 SPT6 SMARCA4 POGZ CAF-1 p150 KAP-1 HP1α 200 116 97.1 CAF-1 p150 tnasp Importin4 B Western blot of the p150caf-1 complex Input mock
Supplementary Table 1. The Q-PCR primer sequence is summarized in the following table.
 Supplementary Table 1. The Q-PCR primer sequence is summarized in the following table. Name Sequence (5-3 ) Application Flag-u ggactacaaggacgacgatgac Shared upstream primer for all the amplifications of
Supplementary Table 1. The Q-PCR primer sequence is summarized in the following table. Name Sequence (5-3 ) Application Flag-u ggactacaaggacgacgatgac Shared upstream primer for all the amplifications of
Supplementary Figure S1. Immunodetection of full-length XA21 and the XA21 C-terminal cleavage product.
 Supplementary Information Supplementary Figure S1. Immunodetection of full-length XA21 and the XA21 C-terminal cleavage product. Total protein extracted from Kitaake wild type and rice plants carrying
Supplementary Information Supplementary Figure S1. Immunodetection of full-length XA21 and the XA21 C-terminal cleavage product. Total protein extracted from Kitaake wild type and rice plants carrying
Transcription Factor Runx3 Is Induced by Influenza A Virus and Double-Strand RNA and
 Transcription Factor Is Induced by Influenza A Virus and Double-Strand RNA and Mediates Airway Epithelial Cell Apoptosis Huachen Gan 1, Qin Hao 1, Steven Idell 1,2 & Hua Tang 1 1 Department of Cellular
Transcription Factor Is Induced by Influenza A Virus and Double-Strand RNA and Mediates Airway Epithelial Cell Apoptosis Huachen Gan 1, Qin Hao 1, Steven Idell 1,2 & Hua Tang 1 1 Department of Cellular
Supplementary Figure 1
 Supplementary Figure 1 Virus infection induces RNF128 expression. (a,b) RT-PCR analysis of Rnf128 (RNF128) mrna expression in mouse peritoneal macrophages (a) and THP-1 cells (b) upon stimulation with
Supplementary Figure 1 Virus infection induces RNF128 expression. (a,b) RT-PCR analysis of Rnf128 (RNF128) mrna expression in mouse peritoneal macrophages (a) and THP-1 cells (b) upon stimulation with
supplementary information
 DOI: 1.138/ncb1839 a b Control 1 2 3 Control 1 2 3 Fbw7 Smad3 1 2 3 4 1 2 3 4 c d IGF-1 IGF-1Rβ IGF-1Rβ-P Control / 1 2 3 4 Real-time RT-PCR Relative quantity (IGF-1/ mrna) 2 1 IGF-1 1 2 3 4 Control /
DOI: 1.138/ncb1839 a b Control 1 2 3 Control 1 2 3 Fbw7 Smad3 1 2 3 4 1 2 3 4 c d IGF-1 IGF-1Rβ IGF-1Rβ-P Control / 1 2 3 4 Real-time RT-PCR Relative quantity (IGF-1/ mrna) 2 1 IGF-1 1 2 3 4 Control /
Supplementary Figure S1. N-terminal fragments of LRRK1 bind to Grb2.
 Myc- HA-Grb2 Mr(K) 105 IP HA 75 25 105 1-1163 1-595 - + - + - + 1164-1989 Blot Myc HA total lysate 75 25 Myc HA Supplementary Figure S1. N-terminal fragments of bind to Grb2. COS7 cells were cotransfected
Myc- HA-Grb2 Mr(K) 105 IP HA 75 25 105 1-1163 1-595 - + - + - + 1164-1989 Blot Myc HA total lysate 75 25 Myc HA Supplementary Figure S1. N-terminal fragments of bind to Grb2. COS7 cells were cotransfected
Regulation of transcription by the MLL2 complex and MLL complex-associated AKAP95
 Supplementary Information Regulation of transcription by the complex and MLL complex-associated Hao Jiang, Xiangdong Lu, Miho Shimada, Yali Dou, Zhanyun Tang, and Robert G. Roeder Input HeLa NE IP lot:
Supplementary Information Regulation of transcription by the complex and MLL complex-associated Hao Jiang, Xiangdong Lu, Miho Shimada, Yali Dou, Zhanyun Tang, and Robert G. Roeder Input HeLa NE IP lot:
Nature Immunology: doi: /ni.3015
 Supplementary Figure 1 Role of RIP1-RIP3 and PGAM5 in RNA virus induced inflammasome activation. (a) LDH release from LPS-primed BMDMs from wild-type mice (WT), Rip3 -/- or Nlrp3 -/- mice infected with
Supplementary Figure 1 Role of RIP1-RIP3 and PGAM5 in RNA virus induced inflammasome activation. (a) LDH release from LPS-primed BMDMs from wild-type mice (WT), Rip3 -/- or Nlrp3 -/- mice infected with
Supplementary data. sienigma. F-Enigma F-EnigmaSM. a-p53
 Supplementary data Supplemental Figure 1 A sienigma #2 sienigma sicontrol a-enigma - + ++ - - - - - - + ++ - - - - - - ++ B sienigma F-Enigma F-EnigmaSM a-flag HLK3 cells - - - + ++ + ++ - + - + + - -
Supplementary data Supplemental Figure 1 A sienigma #2 sienigma sicontrol a-enigma - + ++ - - - - - - + ++ - - - - - - ++ B sienigma F-Enigma F-EnigmaSM a-flag HLK3 cells - - - + ++ + ++ - + - + + - -
Supplementary Figure 1. Immunoprecipitation of synthetic SUMOm-remnant peptides using UMO monoclonal antibody. (a) LC-MS analyses of tryptic
 Supplementary Figure 1. Immunoprecipitation of synthetic SUMOm-remnant peptides using UMO 1-7-7 monoclonal antibody. (a) LC-MS analyses of tryptic digest from HEK293 cells spiked with 6 SUMOmremnant peptides
Supplementary Figure 1. Immunoprecipitation of synthetic SUMOm-remnant peptides using UMO 1-7-7 monoclonal antibody. (a) LC-MS analyses of tryptic digest from HEK293 cells spiked with 6 SUMOmremnant peptides
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/4/167/ra20/dc1 Supplementary Materials for Poly(ADP-Ribose) (PAR) Binding to Apoptosis-Inducing Factor Is Critical for PAR Polymerase-1 Dependent Cell Death (Parthanatos)
www.sciencesignaling.org/cgi/content/full/4/167/ra20/dc1 Supplementary Materials for Poly(ADP-Ribose) (PAR) Binding to Apoptosis-Inducing Factor Is Critical for PAR Polymerase-1 Dependent Cell Death (Parthanatos)
p53 increases MHC class I expression by upregulating the endoplasmic reticulum aminopeptidase ERAP1
 Supplementary Information p53 increases MHC class I expression by upregulating the endoplasmic reticulum aminopeptidase ERAP1 Bei Wang 1, Dandan Niu 1, Liyun Lai 1, & Ee Chee Ren 1,2 1 Singapore Immunology
Supplementary Information p53 increases MHC class I expression by upregulating the endoplasmic reticulum aminopeptidase ERAP1 Bei Wang 1, Dandan Niu 1, Liyun Lai 1, & Ee Chee Ren 1,2 1 Singapore Immunology
transcription and the promoter occupancy of Smad proteins. (A) HepG2 cells were co-transfected with the wwp-luc reporter, and FLAG-tagged FHL1,
 Supplementary Data Supplementary Figure Legends Supplementary Figure 1 FHL-mediated TGFβ-responsive reporter transcription and the promoter occupancy of Smad proteins. (A) HepG2 cells were co-transfected
Supplementary Data Supplementary Figure Legends Supplementary Figure 1 FHL-mediated TGFβ-responsive reporter transcription and the promoter occupancy of Smad proteins. (A) HepG2 cells were co-transfected
Supplementary information
 Supplementary information Supplementary Figure 1 (a) EM image of the pyramidal layer of wt mice. CA3 pyramidal neurons were selected according to their typical alignment, size and shape for subsequent
Supplementary information Supplementary Figure 1 (a) EM image of the pyramidal layer of wt mice. CA3 pyramidal neurons were selected according to their typical alignment, size and shape for subsequent
Figure 1: TDP-43 is subject to lysine acetylation within the RNA-binding domain a) QBI-293 cells were transfected with TDP-43 in the presence or
 Figure 1: TDP-43 is subject to lysine acetylation within the RNA-binding domain a) QBI-293 cells were transfected with TDP-43 in the presence or absence of the acetyltransferase CBP and acetylated TDP-43
Figure 1: TDP-43 is subject to lysine acetylation within the RNA-binding domain a) QBI-293 cells were transfected with TDP-43 in the presence or absence of the acetyltransferase CBP and acetylated TDP-43
Supplementary Figure 1. Co-localization of GLUT1 and DNAL4 in BeWo cells cultured
 Supplementary Figure 1. Co-localization of GLUT1 and DNAL4 in BeWo cells cultured under static conditions. Cells were seeded in the chamber area of the device and cultured overnight without medium perfusion.
Supplementary Figure 1. Co-localization of GLUT1 and DNAL4 in BeWo cells cultured under static conditions. Cells were seeded in the chamber area of the device and cultured overnight without medium perfusion.
Supplementary Fig. 1. (A) Working model. The pluripotency transcription factor OCT4
 SUPPLEMENTARY FIGURE LEGENDS Supplementary Fig. 1. (A) Working model. The pluripotency transcription factor OCT4 directly up-regulates the expression of NIPP1 and CCNF that together inhibit protein phosphatase
SUPPLEMENTARY FIGURE LEGENDS Supplementary Fig. 1. (A) Working model. The pluripotency transcription factor OCT4 directly up-regulates the expression of NIPP1 and CCNF that together inhibit protein phosphatase
SUPPLEMENTARY INFORMATION
 Supplementary Figure 1 sirna and shrna mediated depletion of ATP7A results in loss of melanosomal ATP7A staining. a-h, sirna mediated ATP7A depletion. Immunofluorescence microscopy (IFM) analysis of ATP7A
Supplementary Figure 1 sirna and shrna mediated depletion of ATP7A results in loss of melanosomal ATP7A staining. a-h, sirna mediated ATP7A depletion. Immunofluorescence microscopy (IFM) analysis of ATP7A
Supplementary Table 1. Sequences for BTG2 and BRCA1 sirnas.
 Supplementary Table 1. Sequences for BTG2 and BRCA1 sirnas. Target Gene Non-target / Control BTG2 BRCA1 NFE2L2 Target Sequence ON-TARGET plus Non-targeting sirna # 1 (Cat# D-001810-01-05) sirna1: GAACCGACAUGCUCCCGGA
Supplementary Table 1. Sequences for BTG2 and BRCA1 sirnas. Target Gene Non-target / Control BTG2 BRCA1 NFE2L2 Target Sequence ON-TARGET plus Non-targeting sirna # 1 (Cat# D-001810-01-05) sirna1: GAACCGACAUGCUCCCGGA
Supplementary Figure 1. Confirmation of sirna in PC3 and H1299 cells PC3 (a) and H1299 (b) cells were transfected with sirna oligonucleotides
 Supplementary Figure 1. Confirmation of sirna in PC3 and H1299 cells PC3 (a) and H1299 (b) cells were transfected with sirna oligonucleotides targeting RCP (SMARTPool (RCP) or two individual oligos (RCP#1
Supplementary Figure 1. Confirmation of sirna in PC3 and H1299 cells PC3 (a) and H1299 (b) cells were transfected with sirna oligonucleotides targeting RCP (SMARTPool (RCP) or two individual oligos (RCP#1
SUPPLEMENTAL EXPERIMENTAL PROCEDURES
 SUPPLEMENTAL EXPERIMENTAL PROCEDURES Luciferase Assays Cells were seeded on 24well plates and grown to 7% confluency. Cells were then transfected with ng of reporter constructs and 1 ng of the renilla
SUPPLEMENTAL EXPERIMENTAL PROCEDURES Luciferase Assays Cells were seeded on 24well plates and grown to 7% confluency. Cells were then transfected with ng of reporter constructs and 1 ng of the renilla
Nature Immunology: doi: /ni Supplementary Figure 1
 Supplementary Figure 1 EPRS expression in multiple cell lines upon viral infection. (a,b) EPRS is slightly induced upon viral induction. qpcr of Eprs mrna (a) and immunoblot analysis of corresponding endogenous
Supplementary Figure 1 EPRS expression in multiple cell lines upon viral infection. (a,b) EPRS is slightly induced upon viral induction. qpcr of Eprs mrna (a) and immunoblot analysis of corresponding endogenous
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/8/404/ra120/dc1 Supplementary Materials for The subcellular localization and activity of cortactin is regulated by acetylation and interaction with Keap1 Akihiro
www.sciencesignaling.org/cgi/content/full/8/404/ra120/dc1 Supplementary Materials for The subcellular localization and activity of cortactin is regulated by acetylation and interaction with Keap1 Akihiro
JCB. Supplemental material THE JOURNAL OF CELL BIOLOGY. Kimura et al.,
 Supplemental material JCB Kimura et al., http://www.jcb.org/cgi/content/full/jcb.201503023/dc1 THE JOURNAL OF CELL BIOLOGY Figure S1. TRIMs regulate IFN-γ induced autophagy. (A and B) HC image analysis
Supplemental material JCB Kimura et al., http://www.jcb.org/cgi/content/full/jcb.201503023/dc1 THE JOURNAL OF CELL BIOLOGY Figure S1. TRIMs regulate IFN-γ induced autophagy. (A and B) HC image analysis
Supplementary information. Supplementary Figures
 Supplementary information Supplementary Figures Supplementary Figure 1. A. i. HA-JMY expressing U2OS cells were treated with SAHA (6h). DAPI was used to visualise nuclei. ii. U2OS cells stably expressing
Supplementary information Supplementary Figures Supplementary Figure 1. A. i. HA-JMY expressing U2OS cells were treated with SAHA (6h). DAPI was used to visualise nuclei. ii. U2OS cells stably expressing
ENCODE RBP Antibody Characterization Guidelines
 ENCODE RBP Antibody Characterization Guidelines Approved on November 18, 2016 Background An integral part of the ENCODE Project is to characterize the antibodies used in the experiments. This document
ENCODE RBP Antibody Characterization Guidelines Approved on November 18, 2016 Background An integral part of the ENCODE Project is to characterize the antibodies used in the experiments. This document
Rer1 and calnexin regulate endoplasmic reticulum retention of a peripheral myelin protein 22 mutant that causes type 1A Charcot-Marie-Tooth disease
 Rer1 and calnexin regulate endoplasmic reticulum retention of a peripheral myelin protein mutant that causes type 1A Charcot-Marie-Tooth disease Taichi Hara, Yukiko Hashimoto, Tomoko Akuzawa, Rika Hirai,
Rer1 and calnexin regulate endoplasmic reticulum retention of a peripheral myelin protein mutant that causes type 1A Charcot-Marie-Tooth disease Taichi Hara, Yukiko Hashimoto, Tomoko Akuzawa, Rika Hirai,
Description: Nuclear morphology and dynamics in nontargeting sirna transfected cells. HeLa Kyoto
 Title of file for HTML: Supplementary Information Description: Supplementary Figures and Supplementary Tables Title of file for HTML: Supplementary Movie 1 Description: Nuclear morphology and dynamics
Title of file for HTML: Supplementary Information Description: Supplementary Figures and Supplementary Tables Title of file for HTML: Supplementary Movie 1 Description: Nuclear morphology and dynamics
Supplementary Figure Legends
 Supplementary Figure Legends Figure S1 gene targeting strategy for disruption of chicken gene, related to Figure 1 (f)-(i). (a) The locus and the targeting constructs showing HpaI restriction sites. The
Supplementary Figure Legends Figure S1 gene targeting strategy for disruption of chicken gene, related to Figure 1 (f)-(i). (a) The locus and the targeting constructs showing HpaI restriction sites. The
A RRM1 H2AX DAPI. RRM1 H2AX DAPI Merge. Cont. sirna RRM1
 A H2AX DAPI H2AX DAPI Merge Cont sirna Figure S1: Accumulation of RRM1 at DNA damage sites (A) HeLa cells were subjected to in situ detergent extraction without IR irradiation, and immunostained with the
A H2AX DAPI H2AX DAPI Merge Cont sirna Figure S1: Accumulation of RRM1 at DNA damage sites (A) HeLa cells were subjected to in situ detergent extraction without IR irradiation, and immunostained with the
Supplementary Figure 1. The Hsp70 acetylation level is related to the co-chaperone binding of Hsp70 under various stress conditions.
 Supplementary Figure 1. The Hsp70 acetylation level is related to the co-chaperone binding of Hsp70 under various stress conditions. 1 (a) Etoposide treatment gradually changes acetylation level and co-chaperone
Supplementary Figure 1. The Hsp70 acetylation level is related to the co-chaperone binding of Hsp70 under various stress conditions. 1 (a) Etoposide treatment gradually changes acetylation level and co-chaperone
Supplementary Information
 Supplementary Information stability is regulated by CK2-dependent interaction with R2TP complex Patrick von Morgen 1,2, Kamila Burdova 1, Thomas G. Flower 3, Nicola J. O'Reilly 4, Simon J. Boulton 5, Stephen
Supplementary Information stability is regulated by CK2-dependent interaction with R2TP complex Patrick von Morgen 1,2, Kamila Burdova 1, Thomas G. Flower 3, Nicola J. O'Reilly 4, Simon J. Boulton 5, Stephen
 DOI: 10.1038/ncb3259 A Ismail et al. Supplementary Figure 1 B 60000 45000 SSC 30000 15000 Live cells 0 0 15000 30000 45000 60000 FSC- PARR 60000 45000 PARR Width 30000 FSC- 15000 Single cells 0 0 15000
DOI: 10.1038/ncb3259 A Ismail et al. Supplementary Figure 1 B 60000 45000 SSC 30000 15000 Live cells 0 0 15000 30000 45000 60000 FSC- PARR 60000 45000 PARR Width 30000 FSC- 15000 Single cells 0 0 15000
Supplementary Fig. 1 related to Fig. 1 Clinical relevance of lncrna candidate
 Supplementary Figure Legends Supplementary Fig. 1 related to Fig. 1 Clinical relevance of lncrna candidate BC041951 in gastric cancer. (A) The flow chart for selected candidate lncrnas in 660 up-regulated
Supplementary Figure Legends Supplementary Fig. 1 related to Fig. 1 Clinical relevance of lncrna candidate BC041951 in gastric cancer. (A) The flow chart for selected candidate lncrnas in 660 up-regulated
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature09732 Supplementary Figure 1: Depletion of Fbw7 results in elevated Mcl-1 abundance. a, Total thymocytes from 8-wk-old Lck-Cre/Fbw7 +/fl (Control) or Lck-Cre/Fbw7 fl/fl (Fbw7 KO) mice
doi:10.1038/nature09732 Supplementary Figure 1: Depletion of Fbw7 results in elevated Mcl-1 abundance. a, Total thymocytes from 8-wk-old Lck-Cre/Fbw7 +/fl (Control) or Lck-Cre/Fbw7 fl/fl (Fbw7 KO) mice
Mannen et al., http :// /cgi /content /full /jcb /DC1
 Supplemental material JCB Mannen et al., http ://www.jcb.org /cgi /content /full /jcb.201601024 /DC1 THE JOURNAL OF CELL BIOLOGY Figure S1. Characterization of SNB components. (A) SNB localization of Venus-tagged
Supplemental material JCB Mannen et al., http ://www.jcb.org /cgi /content /full /jcb.201601024 /DC1 THE JOURNAL OF CELL BIOLOGY Figure S1. Characterization of SNB components. (A) SNB localization of Venus-tagged
HPV E6 oncoprotein targets histone methyltransferases for modulating specific. Chih-Hung Hsu, Kai-Lin Peng, Hua-Ci Jhang, Chia-Hui Lin, Shwu-Yuan Wu,
 1 HPV E oncoprotein targets histone methyltransferases for modulating specific gene transcription 3 5 Chih-Hung Hsu, Kai-Lin Peng, Hua-Ci Jhang, Chia-Hui Lin, Shwu-Yuan Wu, Cheng-Ming Chiang, Sheng-Chung
1 HPV E oncoprotein targets histone methyltransferases for modulating specific gene transcription 3 5 Chih-Hung Hsu, Kai-Lin Peng, Hua-Ci Jhang, Chia-Hui Lin, Shwu-Yuan Wu, Cheng-Ming Chiang, Sheng-Chung
Supplementary Figure 1. APP cleavage assay. HEK293 cells were transfected with various
 Supplementary Figure 1. APP cleavage assay. HEK293 cells were transfected with various GST-tagged N-terminal truncated APP fragments including GST-APP full-length (FL), APP (123-695), APP (189-695), or
Supplementary Figure 1. APP cleavage assay. HEK293 cells were transfected with various GST-tagged N-terminal truncated APP fragments including GST-APP full-length (FL), APP (123-695), APP (189-695), or
Primers used for PCR of conductin, SGK1 and GAPDH have been described in (Dehner et al,
 Supplementary METHODS Flow Cytometry (FACS) For FACS analysis, trypsinized cells were fixed in ethanol, rehydrated in PBS and treated with 40μg/ml propidium iodide and 10μ/ml RNase for 30 min at room temperature.
Supplementary METHODS Flow Cytometry (FACS) For FACS analysis, trypsinized cells were fixed in ethanol, rehydrated in PBS and treated with 40μg/ml propidium iodide and 10μ/ml RNase for 30 min at room temperature.
Supplementary Fig.1 Luton
 Supplementary Fig.1 Luton a 175 Brain Thymus Spleen Small Intestine Kidney Testis HeLa b 250 Lung Kidney MDCK c EFA6B si Control si Mismatch #637 #1564 #1770 83 62 47.5 175 IB: anti-efa6b #B1 130 66 Lysates
Supplementary Fig.1 Luton a 175 Brain Thymus Spleen Small Intestine Kidney Testis HeLa b 250 Lung Kidney MDCK c EFA6B si Control si Mismatch #637 #1564 #1770 83 62 47.5 175 IB: anti-efa6b #B1 130 66 Lysates
T H E J O U R N A L O F C E L L B I O L O G Y
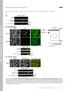 T H E J O U R N A L O F C E L L B I O L O G Y Supplemental material Nakajima and Tanoue, http://www.jcb.org/cgi/content/full/jcb.201104118/dc1 Figure S1. DLD-1 cells exhibit the characteristic morphology
T H E J O U R N A L O F C E L L B I O L O G Y Supplemental material Nakajima and Tanoue, http://www.jcb.org/cgi/content/full/jcb.201104118/dc1 Figure S1. DLD-1 cells exhibit the characteristic morphology
Fig. S1. eif6 expression in HEK293 transfected with shrna against eif6 or pcmv-eif6 vector.
 Fig. S1. eif6 expression in HEK293 transfected with shrna against eif6 or pcmv-eif6 vector. (a) Western blotting analysis and (b) qpcr analysis of eif6 expression in HEK293 T cells transfected with either
Fig. S1. eif6 expression in HEK293 transfected with shrna against eif6 or pcmv-eif6 vector. (a) Western blotting analysis and (b) qpcr analysis of eif6 expression in HEK293 T cells transfected with either
Nature Biotechnology: doi: /nbt Supplementary Figure 1
 Supplementary Figure 1 Schematic and results of screening the combinatorial antibody library for Sox2 replacement activity. A single batch of MEFs were plated and transduced with doxycycline inducible
Supplementary Figure 1 Schematic and results of screening the combinatorial antibody library for Sox2 replacement activity. A single batch of MEFs were plated and transduced with doxycycline inducible
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full//59/ra7/dc1 Supplementary Materials for Dok-7 ctivates the Muscle Receptor Kinase MuSK and Shapes Synapse Formation kane Inoue, Kiyoko Setoguchi, Yosuke Matsubara,
www.sciencesignaling.org/cgi/content/full//59/ra7/dc1 Supplementary Materials for Dok-7 ctivates the Muscle Receptor Kinase MuSK and Shapes Synapse Formation kane Inoue, Kiyoko Setoguchi, Yosuke Matsubara,
B. ADM: C. D. Apoptosis: 1.68% 2.99% 1.31% Figure.S1,Li et al. number. invaded cells. HuH7 BxPC-3 DLD-1.
 A. - Figure.S1,Li et al. B. : - + - + - + E-cadherin CK19 α-sma vimentin β -actin C. D. Apoptosis: 1.68% 2.99% 1.31% - : - + - + - + Apoptosis: 48.33% 45.32% 44.59% E. invaded cells number 400 300 200
A. - Figure.S1,Li et al. B. : - + - + - + E-cadherin CK19 α-sma vimentin β -actin C. D. Apoptosis: 1.68% 2.99% 1.31% - : - + - + - + Apoptosis: 48.33% 45.32% 44.59% E. invaded cells number 400 300 200
Supplementary Figure 1. Additional RNAi screen data
 Supplementary Figure 1. Additional RNAi screen data A. Cisplatin induced ATR autophosphorylation. Western blot illustrating ATR and phospho-atr (T1989) in cells exposed to 1 µm cisplatin for 24 hours prior
Supplementary Figure 1. Additional RNAi screen data A. Cisplatin induced ATR autophosphorylation. Western blot illustrating ATR and phospho-atr (T1989) in cells exposed to 1 µm cisplatin for 24 hours prior
SUPPLEMENTARY INFORMATION
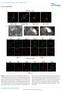 DOI: 10.1038/ncb2386 Figure 1 Src-containing puncta are not focal adhesions, podosomes or endosomes. (a) FAK-/- were stained with anti-py416 Src (green) and either (in red) the focal adhesion protein paxillin,
DOI: 10.1038/ncb2386 Figure 1 Src-containing puncta are not focal adhesions, podosomes or endosomes. (a) FAK-/- were stained with anti-py416 Src (green) and either (in red) the focal adhesion protein paxillin,
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/5/233/ra50/dc1 Supplementary Materials for Epidermal Growth Factor Receptor Is Essential for Toll-Like Receptor 3 Signaling Michifumi Yamashita, Saurabh Chattopadhyay,
www.sciencesignaling.org/cgi/content/full/5/233/ra50/dc1 Supplementary Materials for Epidermal Growth Factor Receptor Is Essential for Toll-Like Receptor 3 Signaling Michifumi Yamashita, Saurabh Chattopadhyay,
Supplemental Data Supplementary Figure Legends and Scheme Figure S1.
 Supplemental Data Supplementary Figure Legends and Scheme Figure S1. UTK1 inhibits the second EGF-induced wave of lamellipodia formation in TT cells. A and B, EGF-induced lamellipodia formation in TT cells,
Supplemental Data Supplementary Figure Legends and Scheme Figure S1. UTK1 inhibits the second EGF-induced wave of lamellipodia formation in TT cells. A and B, EGF-induced lamellipodia formation in TT cells,
Supporting Information
 Supporting Information Su et al. 10.1073/pnas.1211604110 SI Materials and Methods Cell Culture and Plasmids. Tera-1 and Tera-2 cells (ATCC: HTB- 105/106) were maintained in McCoy s 5A medium with 15% FBS
Supporting Information Su et al. 10.1073/pnas.1211604110 SI Materials and Methods Cell Culture and Plasmids. Tera-1 and Tera-2 cells (ATCC: HTB- 105/106) were maintained in McCoy s 5A medium with 15% FBS
Confocal immunofluorescence microscopy
 Confocal immunofluorescence microscopy HL-6 and cells were cultured and cytospun onto glass slides. The cells were double immunofluorescence stained for Mt NPM1 and fibrillarin (nucleolar marker). Briefly,
Confocal immunofluorescence microscopy HL-6 and cells were cultured and cytospun onto glass slides. The cells were double immunofluorescence stained for Mt NPM1 and fibrillarin (nucleolar marker). Briefly,
SUPPLEMENTARY INFORMATION
 DOI:.38/ncb327 a b Sequence coverage (%) 4 3 2 IP: -GFP isoform IP: GFP IP: -GFP IP: GFP Sequence coverage (%) 4 3 2 IP: -GFP IP: GFP 33 52 58 isoform 2 33 49 47 IP: Control IP: Peptide Sequence Start
DOI:.38/ncb327 a b Sequence coverage (%) 4 3 2 IP: -GFP isoform IP: GFP IP: -GFP IP: GFP Sequence coverage (%) 4 3 2 IP: -GFP IP: GFP 33 52 58 isoform 2 33 49 47 IP: Control IP: Peptide Sequence Start
