T H E J O U R N A L O F C E L L B I O L O G Y
|
|
|
- Jasmine Poole
- 5 years ago
- Views:
Transcription
1 Supplemental material Wang et al., T H E J O U R N A L O F C E L L B I O L O G Y Figure S1. Generation and characterization of unc-40 alleles. (A and B) Schematic diagram of unc-40 mutant alleles. (A) The genomic locus of unc-40 shows the exons (alternating black and dark gray boxes) and introns (lines between boxes). The locations of putative unc-40 null alleles are indicated with red arrows for nonsense mutations and a red line for the deletion. The blue arrow indicates the location of the TM domain. (B) The diagram shows the corresponding locations (red arrows) of these unc-40 alleles in the predicted transcript of the unc-40 gene, and the predicted UNC-40 proteins encoded by the wild type and each of the unc-40 mutant alleles. Netrin orients oscillatory DCC clustering Wang et al. S1
2 Figure S2. INA-1/PAT-3 (integrin) mediates unc-40 independent polarizing and F-actin promoting activity at the invasive cell membrane. Anterior is left; ventral is down. (A C) All animals were examined at the P6.p four-cell stage. (A) In wild-type animals, F-actin polarizes to the invasive cell membrane (arrowhead). Bar, 5 µm. (B) In unc-40 mutants, there was a reduction, but not loss, of F-actin at the invasive cell membrane of the AC in all cases (arrowhead). (C) RNAi targeting pat-3, the subunit of integrin heterodimer INA-1/ PAT-3, reduced F-actin volume and polarization at the invasive membrane in unc-40 mutants. (D) Quantification of F-actin polarity in the ACs of wild-type (closed diamonds), unc-40 (open circles), and unc-40; pat-3(rnai) (open squares) animals at the P6.p one-, two-, and four-cell stages (n 15 for each stage per genotype). Significant differences relative to wild-type animals are indicated. (E) Quantification of the total F-actin volume in the ACs in wild type (as a control for normalization), unc-40, and unc-40;pat-3(rnai) at the P6.p four-cell stage (n 15 per genotype). *, P < 0.05; **, P < 0.01; ***, P < N.S., no significant difference (P > 0.05, Student s t test). Error bars represent ± SEM. S2
3 Figure S3. SLT-1, MADD-4, and UNC-129 do not activate UNC-40. Anterior is left; ventral is down. (A C) Images show 3D reconstructions generated from confocal z stacks. Fluorescence (left), corresponding dense F-actin network rendered with isosurfaces (middle), and an overlay (right) are shown. Similar to unc-6 single mutants, F-actin formed and was mislocalized (arrowheads) to the apical and lateral membranes of the AC in slt-1 unc-6, madd-4; unc-6, and unc-129; unc-6 double mutants. Bar, 5 µm. (D) The percentage of the volume of F-actin that localized apicolaterally at the P6.p four-cell stage (n 12 per genotype). (E) Quantification of the total F-actin volume in the AC in wild-type animals (control for normalization) and in each of the mutant backgrounds at the P6.p four-cell stage (n 12 per genotype). Netrin orients oscillatory DCC clustering Wang et al. S3
4 Figure S4. F-actin is stably polarized in unc-40 mutants. Anterior is left; ventral is down. (A) Time-lapse series shows F-actin dynamics in an unc-40 mutant AC. Time points are indicated in minutes. The broken line denotes the position of the basement membrane in the first frame. Bar, 5 µm. (B) The volume of individual F-actin patches over time within the AC. Each colored line represents a distinct F-actin patch. The red line represents the dominant patch in the basal membrane of the AC. Similar results were observed in n = 10/10 unc-40 mutants. Table S1. Primer sequences and templates used for PCR fusions and cloning Primer sequence (5 3 ) Primer type Amplicon Template TAATGTGAGTTAGCTCACTCATTAGG Forward cdh-3 promoter ppd107.94/mk62-63 AACGATGGATACGCTAACAACTTGG Forward nested cdh-3 promoter ppd107.94/mk62-63 TTTCTGAGCTCGGTACCCTCCAAG Reverse cdh-3 promoter ppd107.94/mk62-63 ATGAGTAAAGGAGAAGAACTTTTCAC Forward GFP ppd95.81 (GFP) GGAAACAGTTATGTTTGGTATATTGGG Reverse nested GFP ppd95.81 (GFP) AAGGGCCCGTACGGCCGACTA Reverse GFP ppd95.81 (GFP) TTTGTATAGTTCATCCATGCCATGTG Reverse for GFP extension to N GFP Plasmid cdh-3 > GFP terminus of protein of interest /5Phos/TTAGCGGCTCCACCAAGTTC Forward unc-40( FN4/5) for unc-40( FN4/5)::GFP Plasmid unc-40::gfp /5Phos/AAGTGTTCGTTCTTCTTTATCTTGTCC Reverse unc-40( FN4/5) for unc-40( FN4/5)::GFP Plasmid unc-40::gfp TACGGCCGACTAGTAGGAAA Reverse for fusion unc-40( FN4/5) for unc-40( FN4/5)::GFP Plasmid unc-40::gfp AATACGTAATGATCACATCAGTATTGCG Forward unc-6::nlg-1 TM Plasmid unc-6 > unc-6:: nlg-1 TM::mCherry AAGGTACCCCGTTCTTGTGCGATGCGGATA Reverse unc-6::nlg-1 TM Plasmid unc-6 > unc-6:: nlg-1 TM::mCherry AAGCTTATGATTTTGCGACATTTCGG Forward unc-40 cdna for unc-40::mcherry N2 cdna AACCCGGGCTTATCCATACTCGTCTCAA Reverse unc-40 cdna for unc-40::mcherry N2 cdna Table S2. Extrachromosomal array and integrated strain generation Strain designation PCR fusion or plasmids Injection concentration Coinjection marker ng/µl qyex257 zmp-5 > unc-6::nlg-1 TM::GFP a 100 unc-119+, myo-2 > GFP qyis155 cdh-3 > unc-40( FN4/5)::GFP b 10 unc-119+, myo-2 > GFP qyis262 cdh-3 > unc-40::mcherry b 10 unc-119+ a Plasmid. b PCR fusion product. S4
5 Table S3. Primers used in the screen for unc-40(qy2) deletion mutant Primer sequence (5 3 ) Primer type unc-40 genomic location of the amplicon AGTGAGCAGTGCCGAAGAAG Forward (outer primer) Region I (4,129 5,345 bp) CGACAAGTGTATAGCCCAGT Reverse (outer primer) Region I (4,129 5,345 bp) CGTACACCTATACGTCCCTT Reverse (poison primer) Region I (4,129 5,345 bp) GTAGAAGGTGCTGGGAAGAG Forward (poison primer) Region I (4,129 5,345 bp) GCGCTGTACATGTGACTACA Forward (inner primer) Region I (4,129 5,345 bp) CGATAGAAGCACGTGAGACA Reverse (inner primer) Region I (4,129 5,345 bp) TGTCTCACGTGCTTCTATCG Forward (outer primer) Region II (5,301 6,694 bp) AGGTTGTCCTCCACTTGGTG Reverse (outer primer) Region II (5,301 6,694 bp) GCAGCCACTGATGATGAATC Reverse (poison primer) Region II (5,301 6,694 bp) CCTCAGGAGTTCCAATGACT Forward (poison primer) Region II (5,301 6,694 bp) ACTGGGCTATACACTTGTCG Forward (inner primer) Region II (5,301 6,694 bp) GCTCCAACGGATGTCAATCG Reverse (inner primer) Region II (5,301 6,694 bp) CGATTGACATCCGTTGGAGC Forward (outer primer) Region III (6,654 8,226 bp) GTAGTCCTTCCACAGTAGCC Reverse (outer primer) Region III (6,654 8,226 bp) GCTGGTGGAAGCCATGATAC Reverse (poison primer) Region III (6,654 8,226 bp) GGATGGGGATTGAGTGTTCC Forward (poison primer) Region III (6,654 8,226 bp) CACCAAGTGGAGGACAACCT Forward (inner primer) Region III (6,654 8,226 bp) GTCTCGGTGCTATTCACGTA Reverse (inner primer) Region III (6,654 8,226 bp) TACGTGAATAGCACCGAGAC Forward (outer primer) Region IV (8,184 9,554 bp) GTGACTACAGCCTTGGCTCT Reverse (outer primer) Region IV (8,184 9,554 bp) CCTATTCCACCTCCACTTCC Reverse (poison primer) Region IV (8,184 9,554 bp) GGTCTTGGAGGTCCACCTAA Forward (poison primer) Region IV (8,184 9,554 bp) GGCTACTGTGGAAGGACTAC Forward (inner primer) Region IV (8,184 9,554 bp) ACTAATGCTTCAACCGGCTC Reverse (inner primer) Region IV (8,184 9,554 bp) Video 1. F-actin dynamics in wild-type, unc-40, and unc-6 mutant ACs. The time lapse shows F-actin (visualized with cdh-3 > mcherry::moeabd in grayscale [left] and isosurface renderings in magenta [right]) in the C. elegans AC. In wild-type and unc- 40 mutants, F-actin is dynamic, but localized tightly to the basal cell membrane. In contrast, in unc-6 mutants, F-actin localizes in dynamic clusters that form and then break down in all membrane domains of the cell. Images in movies were created from 3D reconstructions generated from confocal z stacks (CSU-10 spinning disc confocal microscope; Yokogawa Electric Corporation). Frames were acquired at 1-min intervals for a total of 60 min. The movies correspond to animals shown in Fig. S4 (A and B) and Fig. 4 D, respectively. Bar, 5 µm. Anterior is left and ventral is down. Video 2. F-actin colocalizes with UNC-40 in unc-6 mutants. The time lapse shows that UNC-40::GFP (left) and F-actin (cdh-3 > mcherry::moeabd; middle) have persistent colocalization (highlighted in magenta, right; colocalization merged with UNC-40:: GFP, far right) in the C. elegans AC s apical and lateral membranes in unc-6(ev400) mutants. Frames were acquired at 1-min intervals for a total of 75 min using a spinning disc confocal microscope (CSU-10; Yokogawa Electric Corporation). Imaris colocalization function (Coloc) was used to set a threshold for each fluorophore in reconstructed 3D images. The voxels containing both GFP and mcherry signal were considered colocalized. Bar, 5 µm. Video 3. Dorsally localized UNC-6 orients and stabilizes UNC-40 in unc-6 mutants. The time lapse shows that ectopic membrane-tethered UNC-6 (zmp-5 > unc-6::nlg-1 TM::GFP; magenta) expressed in the dorsal C. elegans uterine cells (yellow broken lines; the asterisk indicates the cell with high constant expression of UNC-6) stably orients F-actin (grayscale; white arrow) to the AC s apical cell membrane in contact with UNC-6 in unc-6(ev400). Frames were acquired at 1-min intervals for a total of 58 min using a spinning disc confocal microscope (CSU-10; Yokogawa Electric Corporation). The basement membrane is indicated with a white broken line. Bar, 5 µm. Netrin orients oscillatory DCC clustering Wang et al. S5
6 Video 4. UNC-40 reorients F-actin toward a changing source of UNC-6. The time lapse shows an AC initially directing polarity (visualized with the F-actin probe cdh-3 > mcherry::moeabd; grayscale) toward a dorsal C. elegans uterine cell expressing membrane-tethered UNC-6 (magenta; orange arrow). When the AC made contact with a new source of localized UNC-6 on a neighboring anterior dorsal uterine cell, a polarized response (yellow arrow) was directed toward this new source of UNC-6. Further, as UNC-6 protein was lost on the posterior dorsal uterine cell, polarity was lost here (orange arrowhead). The basement membrane is indicated with a white broken line. Frames were acquired at 1-min intervals for a total of 72 min using a spinning disc confocal microscope (CSU-10; Yokogawa Electric Corporation). The movie covers the time series shown in Fig. 6 (D and E). Bar, 5 µm. Video 5. MADD-2 promotes UNC-40 mediated F-actin polarization toward UNC-6. The time lapse shows F-actin (visualized with cdh-3 > mcherry::moeabd in grayscale [left] and overlaid with isosurface renderings in magenta [right]) in the C. elegans AC. In a representative wild-type animal, F-actin was dynamic but consistently polarized to the basal AC membrane. In contrast, in a madd-2 animal, transient F-actin patches were mislocalized to the apical and lateral AC membrane. Frames were acquired at 1-min intervals for a total of 60 min using a spinning disc confocal microscope (CSU-10; Yokogawa Electric Corporation). The movies cover the time series shown in Fig. 8 (B and E, respectively). Bars, 5 µm. S6
(A) Schematic depicting morphology of cholinergic DA and DB motor neurons in an L1
 SUPPLEMENTAL MATERIALS Supplemental Figure 1 Figure S1. Expression of the non-alpha nachr subunit ACR-2. (A) Schematic depicting morphology of cholinergic DA and DB motor neurons in an L1 animal. Wide-field
SUPPLEMENTAL MATERIALS Supplemental Figure 1 Figure S1. Expression of the non-alpha nachr subunit ACR-2. (A) Schematic depicting morphology of cholinergic DA and DB motor neurons in an L1 animal. Wide-field
mod-1::mcherry unc-47::gfp RME ser-4::gfp vm2 vm2 vm2 vm2 VNC
 A mod-1::mcherry unc-47::gfp RME B ser-4::gfp vm2 vm2 VNC vm2 vm2 VN Figure S1 Further details of mod- 1 and ser- 4 reporter expression patterns. (A) Adult head region showing mod- 1::mCherry and unc-
A mod-1::mcherry unc-47::gfp RME B ser-4::gfp vm2 vm2 VNC vm2 vm2 VN Figure S1 Further details of mod- 1 and ser- 4 reporter expression patterns. (A) Adult head region showing mod- 1::mCherry and unc-
Supplementary Figure 1. Espn-1 knockout characterization. (a) The predicted recombinant Espn-1 -/- allele was detected by PCR of the left (5 )
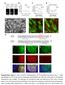 Supplementary Figure 1. Espn-1 knockout characterization. (a) The predicted recombinant Espn-1 -/- allele was detected by PCR of the left (5 ) homologous recombination arm (left) and of the right (3 )
Supplementary Figure 1. Espn-1 knockout characterization. (a) The predicted recombinant Espn-1 -/- allele was detected by PCR of the left (5 ) homologous recombination arm (left) and of the right (3 )
Mitotic cell rounding and epithelial thinning regulate lumen
 Mitotic cell rounding and epithelial thinning regulate lumen growth and shape Esteban Hoijman, Davide Rubbini, Julien Colombelli and Berta Alsina SUPPLEMENTARY FIGURES Supplementary Figure 1 (a-c) Localization
Mitotic cell rounding and epithelial thinning regulate lumen growth and shape Esteban Hoijman, Davide Rubbini, Julien Colombelli and Berta Alsina SUPPLEMENTARY FIGURES Supplementary Figure 1 (a-c) Localization
Two classes of silencing RNAs move between Caenorhabditis elegans tissues.
 Two classes of silencing RNAs move between Caenorhabditis elegans tissues. Antony M Jose, Giancarlo A Garcia, and Craig P Hunter. Supplementary Figures, Figure Legends, and Tables. Supplementary Figure
Two classes of silencing RNAs move between Caenorhabditis elegans tissues. Antony M Jose, Giancarlo A Garcia, and Craig P Hunter. Supplementary Figures, Figure Legends, and Tables. Supplementary Figure
SUPPLEMENTARY INFORMATION
 Figure S1. lev-9 mutants are resistant to levamisole. The levamisole dose-response curves indicate that lev-9 mutants are partially resistant to levamisole similar to lev-10(kr26) mutants. unc-29(x29)
Figure S1. lev-9 mutants are resistant to levamisole. The levamisole dose-response curves indicate that lev-9 mutants are partially resistant to levamisole similar to lev-10(kr26) mutants. unc-29(x29)
JCB. Supplemental material THE JOURNAL OF CELL BIOLOGY. Paul et al.,
 Supplemental material JCB Paul et al., http://www.jcb.org/cgi/content/full/jcb.201502040/dc1 THE JOURNAL OF CELL BIOLOGY Figure S1. Mutant p53-expressing cells display limited retrograde actin flow at
Supplemental material JCB Paul et al., http://www.jcb.org/cgi/content/full/jcb.201502040/dc1 THE JOURNAL OF CELL BIOLOGY Figure S1. Mutant p53-expressing cells display limited retrograde actin flow at
Nature Methods: doi: /nmeth Supplementary Figure 1. Neither driver nor effector alone displays expression of GFP
 Supplementary Figure 1 Neither driver nor effector alone displays expression of GFP Comparison of GFP fluorescence in a driver- only strain (a) or an effector-only strain (b) to their driver + effector
Supplementary Figure 1 Neither driver nor effector alone displays expression of GFP Comparison of GFP fluorescence in a driver- only strain (a) or an effector-only strain (b) to their driver + effector
Nature Biotechnology: doi: /nbt.4166
 Supplementary Figure 1 Validation of correct targeting at targeted locus. (a) by immunofluorescence staining of 2C-HR-CRISPR microinjected embryos cultured to the blastocyst stage. Embryos were stained
Supplementary Figure 1 Validation of correct targeting at targeted locus. (a) by immunofluorescence staining of 2C-HR-CRISPR microinjected embryos cultured to the blastocyst stage. Embryos were stained
unc-33/crmp and ankyrin organize microtubules and localize kinesin to polarize
 Supplementary Information unc-33/crmp and ankyrin organize microtubules and localize kinesin to polarize axon-dendrite sorting Tapan A. Maniar 1, Miriam Kaplan 1, George J. Wang 2, Kang Shen 2, Li Wei
Supplementary Information unc-33/crmp and ankyrin organize microtubules and localize kinesin to polarize axon-dendrite sorting Tapan A. Maniar 1, Miriam Kaplan 1, George J. Wang 2, Kang Shen 2, Li Wei
Supplemental Data. Farmer et al. (2010) Plant Cell /tpc
 Supplemental Figure 1. Amino acid sequence comparison of RAD23 proteins. Identical and similar residues are shown in the black and gray boxes, respectively. Dots denote gaps. The sequence of plant Ub is
Supplemental Figure 1. Amino acid sequence comparison of RAD23 proteins. Identical and similar residues are shown in the black and gray boxes, respectively. Dots denote gaps. The sequence of plant Ub is
T H E J O U R N A L O F C E L L B I O L O G Y
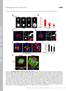 T H E J O U R N A L O F C E L L B I O L O G Y Supplemental material Rodríguez-Fraticelli et al., http://www.jcb.org/cgi/content/full/jcb.201203075/dc1 Figure S1. Cell spreading and lumen formation in confined
T H E J O U R N A L O F C E L L B I O L O G Y Supplemental material Rodríguez-Fraticelli et al., http://www.jcb.org/cgi/content/full/jcb.201203075/dc1 Figure S1. Cell spreading and lumen formation in confined
sides of the aleurone (Al) but it is excluded from the basal endosperm transfer layer
 Supplemental Data. Gómez et al. (2009). The maize transcription factor MRP-1 (Myb-Related-Protein-1) is a key regulator of the differentiation of transfer cells. Supplemental Figure 1. Expression analyses
Supplemental Data. Gómez et al. (2009). The maize transcription factor MRP-1 (Myb-Related-Protein-1) is a key regulator of the differentiation of transfer cells. Supplemental Figure 1. Expression analyses
Fig. S1. Molecular phylogenetic analysis of AtHD-ZIP IV family. A phylogenetic tree was constructed using Bayesian analysis with Markov Chain Monte
 Fig. S1. Molecular phylogenetic analysis of AtHD-ZIP IV family. A phylogenetic tree was constructed using Bayesian analysis with Markov Chain Monte Carlo algorithm for one million generations to obtain
Fig. S1. Molecular phylogenetic analysis of AtHD-ZIP IV family. A phylogenetic tree was constructed using Bayesian analysis with Markov Chain Monte Carlo algorithm for one million generations to obtain
Liu et al., http :// /cgi /content /full /jcb /DC1
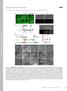 Supplemental material JCB Liu et al., http ://www.jcb.org /cgi /content /full /jcb.201506081 /DC1 THE JOU RNAL OF CELL BIO LOGY Figure S1. Identification of sorf-1 and sorf-2. (A) RNAi of ZK563.5 increases
Supplemental material JCB Liu et al., http ://www.jcb.org /cgi /content /full /jcb.201506081 /DC1 THE JOU RNAL OF CELL BIO LOGY Figure S1. Identification of sorf-1 and sorf-2. (A) RNAi of ZK563.5 increases
b number of motif occurrences
 supplementary information DOI: 1.138/ncb194 a b number of motif occurrences number of motif occurrences number of motif occurrences I II III IV V X TTGGTCAGTGCA 3 8 23 4 13 239 TTGGTCAGTGCT 7 5 8 1 3 83
supplementary information DOI: 1.138/ncb194 a b number of motif occurrences number of motif occurrences number of motif occurrences I II III IV V X TTGGTCAGTGCA 3 8 23 4 13 239 TTGGTCAGTGCT 7 5 8 1 3 83
embryos. Asterisk represents loss of or reduced expression. Brackets represent
 Supplemental Figures Supplemental Figure 1. tfec expression is highly enriched in tail endothelial cells (A- B) ISH of tfec at 15 and 16hpf in WT embryos. (C- D) ISH of tfec at 36 and 38hpf in WT embryos.
Supplemental Figures Supplemental Figure 1. tfec expression is highly enriched in tail endothelial cells (A- B) ISH of tfec at 15 and 16hpf in WT embryos. (C- D) ISH of tfec at 36 and 38hpf in WT embryos.
Figure S1. Verification of ihog Mutation by Protein Immunoblotting Figure S2. Verification of ihog and boi
 Figure S1. Verification of ihog Mutation by Protein Immunoblotting Extracts from S2R+ cells, embryos, and adults were analyzed by immunoprecipitation and immunoblotting with anti-ihog antibody. The Ihog
Figure S1. Verification of ihog Mutation by Protein Immunoblotting Extracts from S2R+ cells, embryos, and adults were analyzed by immunoprecipitation and immunoblotting with anti-ihog antibody. The Ihog
Figure S1a. The modular dcas9 fusion system works efficiently to suppress and activate endogenous gene expression in C. elegans
 Figure S1a. The modular dcas9 fusion system works efficiently to suppress and activate endogenous gene expression in C. elegans A, a normal wild-type hermaphrodite N2 is shown. B, a dpy-5-supressed F1
Figure S1a. The modular dcas9 fusion system works efficiently to suppress and activate endogenous gene expression in C. elegans A, a normal wild-type hermaphrodite N2 is shown. B, a dpy-5-supressed F1
File name: Supplementary Information Description: Supplementary figures and supplementary tables. File name: Peer review file Description:
 File name: Supplementary Information Description: Supplementary figures and supplementary tables. File name: Peer review file Description: Supplementary Figure 1. dcas9-mq1 fusion protein induces de novo
File name: Supplementary Information Description: Supplementary figures and supplementary tables. File name: Peer review file Description: Supplementary Figure 1. dcas9-mq1 fusion protein induces de novo
Supporting Information
 Supporting Information Park et al. 10.1073/pnas.1410555111 5 -TCAAGTCCATCTACATGGCC-3 5 -CAGCTGCCCGGCTACTACTA-3 5 -TGCAGCTGCCCGGCTACTAC-3 5 -AAGCTGGACATCACCTCCCA-3 5 -TGACAGGAACACCTACAAGT-3 5 -AAGGCACCTTTCTGTCTCCA-3
Supporting Information Park et al. 10.1073/pnas.1410555111 5 -TCAAGTCCATCTACATGGCC-3 5 -CAGCTGCCCGGCTACTACTA-3 5 -TGCAGCTGCCCGGCTACTAC-3 5 -AAGCTGGACATCACCTCCCA-3 5 -TGACAGGAACACCTACAAGT-3 5 -AAGGCACCTTTCTGTCTCCA-3
Supplemental Data. Na Xu et al. (2016). Plant Cell /tpc
 Supplemental Figure 1. The weak fluorescence phenotype is not caused by the mutation in At3g60240. (A) A mutation mapped to the gene At3g60240. Map-based cloning strategy was used to map the mutated site
Supplemental Figure 1. The weak fluorescence phenotype is not caused by the mutation in At3g60240. (A) A mutation mapped to the gene At3g60240. Map-based cloning strategy was used to map the mutated site
Supplemental Fig. 1. Mcr alleles show defects in tracheal tube size and luminal protein accumulation. (A-F) Confocal projections of living stage 15
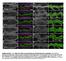 Supplemental Fig. 1. Mcr alleles show defects in tracheal tube size and luminal protein accumulation. (A-F) Confocal projections of living stage 15 embryos expressing GFP and Verm-RFP in tracheal cells
Supplemental Fig. 1. Mcr alleles show defects in tracheal tube size and luminal protein accumulation. (A-F) Confocal projections of living stage 15 embryos expressing GFP and Verm-RFP in tracheal cells
A CRISPR/Cas9 Vector System for Tissue-Specific Gene Disruption in Zebrafish
 Developmental Cell Supplemental Information A CRISPR/Cas9 Vector System for Tissue-Specific Gene Disruption in Zebrafish Julien Ablain, Ellen M. Durand, Song Yang, Yi Zhou, and Leonard I. Zon % larvae
Developmental Cell Supplemental Information A CRISPR/Cas9 Vector System for Tissue-Specific Gene Disruption in Zebrafish Julien Ablain, Ellen M. Durand, Song Yang, Yi Zhou, and Leonard I. Zon % larvae
Fig. S1. Phosphorylation of eif2 by PEK-1 is increased in ire-1 mutants. Fig. S2. ire-1
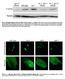 Fig. S1. Phosphorylation of eif2a by PEK-1 is increased in ire-1 mutants. Representative western blot of phosphorylated eif2a and tubulin of day-0 wild-type animals and ire-1 mutants treated with control,
Fig. S1. Phosphorylation of eif2a by PEK-1 is increased in ire-1 mutants. Representative western blot of phosphorylated eif2a and tubulin of day-0 wild-type animals and ire-1 mutants treated with control,
Fig. S1. Endocytosis of extracellular cargo does not depend on dia. (A) To visualize endocytosis, fluorescently labelled wheat germ agglutinin
 Fig. S1. Endocytosis of extracellular cargo does not depend on dia. (A) To visualize endocytosis, fluorescently labelled wheat germ agglutinin (WGA-Alexa555) was injected into the extracellular perivitteline
Fig. S1. Endocytosis of extracellular cargo does not depend on dia. (A) To visualize endocytosis, fluorescently labelled wheat germ agglutinin (WGA-Alexa555) was injected into the extracellular perivitteline
- 1 - Supplemental Data
 - 1-1 Supplemental Data 2 3 4 5 6 7 8 9 Supplemental Figure S1. Differential expression of AtPIP Genes in DC3000-inoculated plants. Gene expression in leaves was analyzed by real-time RT-PCR and expression
- 1-1 Supplemental Data 2 3 4 5 6 7 8 9 Supplemental Figure S1. Differential expression of AtPIP Genes in DC3000-inoculated plants. Gene expression in leaves was analyzed by real-time RT-PCR and expression
Supplemental Information
 Supplemental Information Itemized List Materials and Methods, Related to Supplemental Figures S5A-C and S6. Supplemental Figure S1, Related to Figures 1 and 2. Supplemental Figure S2, Related to Figure
Supplemental Information Itemized List Materials and Methods, Related to Supplemental Figures S5A-C and S6. Supplemental Figure S1, Related to Figures 1 and 2. Supplemental Figure S2, Related to Figure
Nature Methods: doi: /nmeth Supplementary Figure 1. DMS-MaPseq data are highly reproducible at elevated DMS concentrations.
 Supplementary Figure 1 DMS-MaPseq data are highly reproducible at elevated DMS concentrations. a, Correlation of Gini index for 202 yeast mrna regions with 15x coverage at 2.5% or 5% v/v DMS concentrations
Supplementary Figure 1 DMS-MaPseq data are highly reproducible at elevated DMS concentrations. a, Correlation of Gini index for 202 yeast mrna regions with 15x coverage at 2.5% or 5% v/v DMS concentrations
Mos1 insertion. MosTIC protocol-11/2006. repair template - Mos1 transposase expression - Mos1 excision - DSB formation. homolog arm.
 MosTIC (Mos1 excision induced Transgene Instructed gene Conversion) Valérie Robert (vrobert@biologie.ens.fr) and Jean-Louis Bessereau (jlbesse@biologie.ens.fr) (November 2006) Introduction: MosTIC (Robert
MosTIC (Mos1 excision induced Transgene Instructed gene Conversion) Valérie Robert (vrobert@biologie.ens.fr) and Jean-Louis Bessereau (jlbesse@biologie.ens.fr) (November 2006) Introduction: MosTIC (Robert
Supplementary Information. Isl2b regulates anterior second heart field development in zebrafish
 Supplementary Information Isl2b regulates anterior second heart field development in zebrafish Hagen R. Witzel 1, Sirisha Cheedipudi 1, Rui Gao 1, Didier Y.R. Stainier 2 and Gergana D. Dobreva 1,3* 1 Origin
Supplementary Information Isl2b regulates anterior second heart field development in zebrafish Hagen R. Witzel 1, Sirisha Cheedipudi 1, Rui Gao 1, Didier Y.R. Stainier 2 and Gergana D. Dobreva 1,3* 1 Origin
Erhard et al. (2013). Plant Cell /tpc
 Supplemental Figure 1. c1-hbr allele structure. Diagram of the c1-hbr allele found in stocks segregating 1:1 for rpd1-1 and rpd1-2 homozygous mutants showing the presence of a 363 base pair (bp) Heartbreaker
Supplemental Figure 1. c1-hbr allele structure. Diagram of the c1-hbr allele found in stocks segregating 1:1 for rpd1-1 and rpd1-2 homozygous mutants showing the presence of a 363 base pair (bp) Heartbreaker
Supplementary Figure S1. Immunodetection of full-length XA21 and the XA21 C-terminal cleavage product.
 Supplementary Information Supplementary Figure S1. Immunodetection of full-length XA21 and the XA21 C-terminal cleavage product. Total protein extracted from Kitaake wild type and rice plants carrying
Supplementary Information Supplementary Figure S1. Immunodetection of full-length XA21 and the XA21 C-terminal cleavage product. Total protein extracted from Kitaake wild type and rice plants carrying
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb3374 Supplementary Figure 1 staccato mutations affect a late step of the tube fusion process. (a) Immunostaining of stage 15 embryos labeled with anti- Dys (green in merge) and a probe
DOI: 10.1038/ncb3374 Supplementary Figure 1 staccato mutations affect a late step of the tube fusion process. (a) Immunostaining of stage 15 embryos labeled with anti- Dys (green in merge) and a probe
Neuronal Activity and CaMKII Regulate Kinesin-Mediated Transport of Synaptic AMPARs
 Neuron Supplemental Information Neuronal Activity and CaMKII Regulate Kinesin-Mediated Transport of Synaptic AMPARs Frédéric J. Hoerndli, Rui Wang, Jerry E. Mellem, Angy Kallarackal, Penelope J. Brockie,
Neuron Supplemental Information Neuronal Activity and CaMKII Regulate Kinesin-Mediated Transport of Synaptic AMPARs Frédéric J. Hoerndli, Rui Wang, Jerry E. Mellem, Angy Kallarackal, Penelope J. Brockie,
Supplementary Figure 1. Serial deletion mutants of BLITz.
 Supplementary Figure 1 Serial deletion mutants of BLITz. (a) Design of TEVseq insertion into J -helix. C-terminal end of J -helix was serially deleted and replaced by TEVseq. TEV cleavage site is labeled
Supplementary Figure 1 Serial deletion mutants of BLITz. (a) Design of TEVseq insertion into J -helix. C-terminal end of J -helix was serially deleted and replaced by TEVseq. TEV cleavage site is labeled
Supplemental Data. Liu et al. (2013). Plant Cell /tpc
 Supplemental Figure 1. The GFP Tag Does Not Disturb the Physiological Functions of WDL3. (A) RT-PCR analysis of WDL3 expression in wild-type, WDL3-GFP, and WDL3 (without the GFP tag) transgenic seedlings.
Supplemental Figure 1. The GFP Tag Does Not Disturb the Physiological Functions of WDL3. (A) RT-PCR analysis of WDL3 expression in wild-type, WDL3-GFP, and WDL3 (without the GFP tag) transgenic seedlings.
SUPPLEMENTARY INFORMATION
 AS-NMD modulates FLM-dependent thermosensory flowering response in Arabidopsis NATURE PLANTS www.nature.com/natureplants 1 Supplementary Figure 1. Genomic sequence of FLM along with the splice sites. Sequencing
AS-NMD modulates FLM-dependent thermosensory flowering response in Arabidopsis NATURE PLANTS www.nature.com/natureplants 1 Supplementary Figure 1. Genomic sequence of FLM along with the splice sites. Sequencing
i-stop codon positions in the mcherry gene
 Supplementary Figure 1 i-stop codon positions in the mcherry gene The grnas (green) that can potentially generate stop codons from Trp (63 th and 98 th aa, upper panel) and Gln (47 th and 114 th aa, bottom
Supplementary Figure 1 i-stop codon positions in the mcherry gene The grnas (green) that can potentially generate stop codons from Trp (63 th and 98 th aa, upper panel) and Gln (47 th and 114 th aa, bottom
Figure S1 is related to Figure 1B, showing more details of outer segment of
 Supplemental Information Supplementary Figure legends and Figures Figure S1. Electron microscopic images in Sema4A +/+ and Sema4A / retinas Figure S1 is related to Figure 1B, showing more details of outer
Supplemental Information Supplementary Figure legends and Figures Figure S1. Electron microscopic images in Sema4A +/+ and Sema4A / retinas Figure S1 is related to Figure 1B, showing more details of outer
Nature Genetics: doi: /ng Supplementary Figure 1
 Supplementary Figure 1 Ihh interacts preferentially with its upstream neighboring gene Nhej1. Genes are indicated by gray lines, and Ihh and Nhej1 are highlighted in blue. 4C seq performed in E14.5 limbs
Supplementary Figure 1 Ihh interacts preferentially with its upstream neighboring gene Nhej1. Genes are indicated by gray lines, and Ihh and Nhej1 are highlighted in blue. 4C seq performed in E14.5 limbs
Sperm cells are passive cargo of the pollen tube in plant fertilization
 In the format provided by the authors and unedited. SUPPLEMENTARY INFORMATION VOLUME: 3 ARTICLE NUMBER: 17079 Sperm cells are passive cargo of the pollen tube in plant fertilization Jun Zhang 1, Qingpei
In the format provided by the authors and unedited. SUPPLEMENTARY INFORMATION VOLUME: 3 ARTICLE NUMBER: 17079 Sperm cells are passive cargo of the pollen tube in plant fertilization Jun Zhang 1, Qingpei
Live-cell visualization of excitation energy dynamics in chloroplast thylakoid structures
 Supplementary Information Live-cell visualization of excitation energy dynamics in chloroplast thylakoid structures Masakazu Iwai, Makio Yokono, Kazuo Kurokawa, Akira Ichihara & Akihiko Nakano Supplementary
Supplementary Information Live-cell visualization of excitation energy dynamics in chloroplast thylakoid structures Masakazu Iwai, Makio Yokono, Kazuo Kurokawa, Akira Ichihara & Akihiko Nakano Supplementary
Grb2-Mediated Alteration in the Trafficking of AβPP: Insights from Grb2-AICD Interaction
 Journal of Alzheimer s Disease 20 (2010) 1 9 1 IOS Press Supplementary Material Grb2-Mediated Alteration in the Trafficking of AβPP: Insights from Grb2-AICD Interaction Mithu Raychaudhuri and Debashis
Journal of Alzheimer s Disease 20 (2010) 1 9 1 IOS Press Supplementary Material Grb2-Mediated Alteration in the Trafficking of AβPP: Insights from Grb2-AICD Interaction Mithu Raychaudhuri and Debashis
T H E J O U R N A L O F C E L L B I O L O G Y
 Supplemental material Prashar et al., http://www.jcb.org/cgi/content/full/jcb.201304095/dc1 T H E J O U R N A L O F C E L L B I O L O G Y Figure S1. FBT phagocytosis in cells expressing PM-GFP. (A) T-PC
Supplemental material Prashar et al., http://www.jcb.org/cgi/content/full/jcb.201304095/dc1 T H E J O U R N A L O F C E L L B I O L O G Y Figure S1. FBT phagocytosis in cells expressing PM-GFP. (A) T-PC
Supplemental Data. Wang et al. (2017). Plant Cell /tpc NIP1;2 NIP7;1
 Supplemental Data. Wang et al. (217). Plant Cell 1.115/tpc.16.825 NIP1;1 NIP1;2 NIP2;1 NIP3;1 NIP4;1 NIP5;1 NIP6;1 NIP7;1 Supplemental Figure 1. Distinct Localization of NIPs in Root Epidermal Cells. (Supports
Supplemental Data. Wang et al. (217). Plant Cell 1.115/tpc.16.825 NIP1;1 NIP1;2 NIP2;1 NIP3;1 NIP4;1 NIP5;1 NIP6;1 NIP7;1 Supplemental Figure 1. Distinct Localization of NIPs in Root Epidermal Cells. (Supports
Supplementary Methods
 Supplementary Methods Reverse transcribed Quantitative PCR. Total RNA was isolated from bone marrow derived macrophages using RNeasy Mini Kit (Qiagen), DNase-treated (Promega RQ1), and reverse transcribed
Supplementary Methods Reverse transcribed Quantitative PCR. Total RNA was isolated from bone marrow derived macrophages using RNeasy Mini Kit (Qiagen), DNase-treated (Promega RQ1), and reverse transcribed
Revised: RG-RV2 by Fukuhara et al.
 Supplemental Figure 1 The generation of Spns2 conditional knockout mice. (A) Schematic representation of the wild type Spns2 locus (Spns2 + ), the targeted allele, the floxed allele (Spns2 f ) and the
Supplemental Figure 1 The generation of Spns2 conditional knockout mice. (A) Schematic representation of the wild type Spns2 locus (Spns2 + ), the targeted allele, the floxed allele (Spns2 f ) and the
Supplemental Figure 1 Human REEP family of proteins can be divided into two distinct subfamilies. Residues (single letter amino acid code) identical
 Supplemental Figure Human REEP family of proteins can be divided into two distinct subfamilies. Residues (single letter amino acid code) identical in all six REEPs are highlighted in green. Additional
Supplemental Figure Human REEP family of proteins can be divided into two distinct subfamilies. Residues (single letter amino acid code) identical in all six REEPs are highlighted in green. Additional
SUPPLEMENTARY INFORMATION
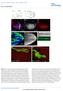 DOI: 10.1038/ncb2806 Supplemental Material: Figure S1 Wnt4 PCP phenotypes are not mediated through canonical Wnt signaling. (a-a ) Illustration of wg, dwnt4 and dwnt6 expression patterns in wing (a) and
DOI: 10.1038/ncb2806 Supplemental Material: Figure S1 Wnt4 PCP phenotypes are not mediated through canonical Wnt signaling. (a-a ) Illustration of wg, dwnt4 and dwnt6 expression patterns in wing (a) and
T H E J O U R N A L O F C E L L B I O L O G Y
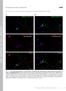 T H E J O U R N A L O F C E L L B I O L O G Y Supplemental material Kanaani et al., http://www.jcb.org/cgi/content/full/jcb.200912101/dc1 Figure S1. The K2 rabbit polyclonal antibody is specific for GAD67,
T H E J O U R N A L O F C E L L B I O L O G Y Supplemental material Kanaani et al., http://www.jcb.org/cgi/content/full/jcb.200912101/dc1 Figure S1. The K2 rabbit polyclonal antibody is specific for GAD67,
Regulation of Acetylcholine Receptor Clustering by ADF/Cofilin- Directed Vesicular Trafficking
 Regulation of Acetylcholine Receptor Clustering by ADF/Cofilin- Directed Vesicular Trafficking Chi Wai Lee, Jianzhong Han, James R. Bamburg, Liang Han, Rachel Lynn, and James Q. Zheng Supplementary Figures
Regulation of Acetylcholine Receptor Clustering by ADF/Cofilin- Directed Vesicular Trafficking Chi Wai Lee, Jianzhong Han, James R. Bamburg, Liang Han, Rachel Lynn, and James Q. Zheng Supplementary Figures
Map-Based Cloning of Qualitative Plant Genes
 Map-Based Cloning of Qualitative Plant Genes Map-based cloning using the genetic relationship between a gene and a marker as the basis for beginning a search for a gene Chromosome walking moving toward
Map-Based Cloning of Qualitative Plant Genes Map-based cloning using the genetic relationship between a gene and a marker as the basis for beginning a search for a gene Chromosome walking moving toward
Supporting information
 Supporting information Construction of strains and plasmids To create ptc67, a PCR product obtained with primers cc2570-162f (gcatgggcaagcttgaggacggcgtcatgt) and cc2570+512f (gaggccgtggtaccatagaggcgggcg),
Supporting information Construction of strains and plasmids To create ptc67, a PCR product obtained with primers cc2570-162f (gcatgggcaagcttgaggacggcgtcatgt) and cc2570+512f (gaggccgtggtaccatagaggcgggcg),
T H E J O U R N A L O F C E L L B I O L O G Y
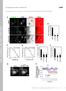 T H E J O U R N A L O F C E L L B I O L O G Y Supplemental material Yamaguchi et al., http://www.jcb.org/cgi/content/full/jcb.201009126/dc1 S1 Figure S2. The expression levels of GFP-Akt1-PH and localization
T H E J O U R N A L O F C E L L B I O L O G Y Supplemental material Yamaguchi et al., http://www.jcb.org/cgi/content/full/jcb.201009126/dc1 S1 Figure S2. The expression levels of GFP-Akt1-PH and localization
Schematic representation of the endogenous PALB2 locus and gene-disruption constructs
 Supplementary Figures Supplementary Figure 1. Generation of PALB2 -/- and BRCA2 -/- /PALB2 -/- DT40 cells. (A) Schematic representation of the endogenous PALB2 locus and gene-disruption constructs carrying
Supplementary Figures Supplementary Figure 1. Generation of PALB2 -/- and BRCA2 -/- /PALB2 -/- DT40 cells. (A) Schematic representation of the endogenous PALB2 locus and gene-disruption constructs carrying
A Repressor Complex Governs the Integration of
 Developmental Cell 15 Supplemental Data A Repressor Complex Governs the Integration of Flowering Signals in Arabidopsis Dan Li, Chang Liu, Lisha Shen, Yang Wu, Hongyan Chen, Masumi Robertson, Chris A.
Developmental Cell 15 Supplemental Data A Repressor Complex Governs the Integration of Flowering Signals in Arabidopsis Dan Li, Chang Liu, Lisha Shen, Yang Wu, Hongyan Chen, Masumi Robertson, Chris A.
Supplementary Figure 1. jmj30-2 and jmj32-1 produce null mutants. (a) Schematic drawing of JMJ30 and JMJ32 genome structure showing regions amplified
 Supplementary Figure 1. jmj30-2 and jmj32-1 produce null mutants. (a) Schematic drawing of JMJ30 and JMJ32 genome structure showing regions amplified by primers used for mrna expression analysis. Gray
Supplementary Figure 1. jmj30-2 and jmj32-1 produce null mutants. (a) Schematic drawing of JMJ30 and JMJ32 genome structure showing regions amplified by primers used for mrna expression analysis. Gray
Supplemental figures (1-9) Gu et al. ADF/Cofilin-Mediated Actin Dynamics Regulate AMPA Receptor Trafficking during Synaptic Plasticity
 Supplemental figures (1-9) Gu et al. ADF/Cofilin-Mediated Actin Dynamics Regulate AMPA Receptor Trafficking during Synaptic Plasticity Supplemental figure 1. The quantification method to determine the
Supplemental figures (1-9) Gu et al. ADF/Cofilin-Mediated Actin Dynamics Regulate AMPA Receptor Trafficking during Synaptic Plasticity Supplemental figure 1. The quantification method to determine the
Supplement Figure S1:
 Supplement Figure S1: A, Sequence of Xcadherin-11 Morpholino 1 (Xcad-11MO) and 2 (Xcad-11 MO2) and control morpholino in comparison to the Xcadherin-11 sequence. The Xcad-11MO binding sequence spans the
Supplement Figure S1: A, Sequence of Xcadherin-11 Morpholino 1 (Xcad-11MO) and 2 (Xcad-11 MO2) and control morpholino in comparison to the Xcadherin-11 sequence. The Xcad-11MO binding sequence spans the
Supplemental material JCB. Pitaval et al., Cell shape and ciliogenesis Pitaval et al.
 Supplemental material THE JOURNAL OF CELL BIOLOGY Pitaval et al., http://www.jcb.org/cgi/content/full/jcb.201004003/dc1 S1 Figure S1. Cell cycle exit analysis. (A) Experimental procedure. RPE1 cells were
Supplemental material THE JOURNAL OF CELL BIOLOGY Pitaval et al., http://www.jcb.org/cgi/content/full/jcb.201004003/dc1 S1 Figure S1. Cell cycle exit analysis. (A) Experimental procedure. RPE1 cells were
T H E J O U R N A L O F C E L L B I O L O G Y
 T H E J O U R N A L O F C E L L B I O L O G Y Supplemental material Monteiro et al., http://www.jcb.org/cgi/content/full/jcb.201306162/dc1 Figure S1. 3D deconvolution microscopy analysis of WASH and exocyst
T H E J O U R N A L O F C E L L B I O L O G Y Supplemental material Monteiro et al., http://www.jcb.org/cgi/content/full/jcb.201306162/dc1 Figure S1. 3D deconvolution microscopy analysis of WASH and exocyst
Supplemental Data. Borg et al. Plant Cell (2014) /tpc
 Supplementary Figure 1 - Alignment of selected angiosperm DAZ1 and DAZ2 homologs Multiple sequence alignment of selected DAZ1 and DAZ2 homologs. A consensus sequence built using default parameters is shown
Supplementary Figure 1 - Alignment of selected angiosperm DAZ1 and DAZ2 homologs Multiple sequence alignment of selected DAZ1 and DAZ2 homologs. A consensus sequence built using default parameters is shown
Supplementary Figure 1.
 Supplementary Figure 1. (A) UCSC Genome Browser view of region immediately downstream of the NEUROG3 start codon. All candidate grna target sites which meet the G(N 19 )NGG constraint are aligned to illustrate
Supplementary Figure 1. (A) UCSC Genome Browser view of region immediately downstream of the NEUROG3 start codon. All candidate grna target sites which meet the G(N 19 )NGG constraint are aligned to illustrate
Supplementary Materials for
 www.sciencemag.org/cgi/content/full/science.1245533/dc1 Supplementary Materials for A Mechanism for Reorientation of Cortical Microtubule Arrays Driven by Microtubule Severing Jelmer J. Lindeboom, Masayoshi
www.sciencemag.org/cgi/content/full/science.1245533/dc1 Supplementary Materials for A Mechanism for Reorientation of Cortical Microtubule Arrays Driven by Microtubule Severing Jelmer J. Lindeboom, Masayoshi
SUPPLEMENTARY INFORMATION
 Gene replacements and insertions in rice by intron targeting using CRISPR Cas9 Table of Contents Supplementary Figure 1. sgrna-induced targeted mutations in the OsEPSPS gene in rice protoplasts. Supplementary
Gene replacements and insertions in rice by intron targeting using CRISPR Cas9 Table of Contents Supplementary Figure 1. sgrna-induced targeted mutations in the OsEPSPS gene in rice protoplasts. Supplementary
LysoTracker Red DND-99 (Invitrogen) was used as a marker of lysosome or acidic
 information MATERIAL AND METHODS Lysosome staining LysoTracker Red DND-99 (Invitrogen) was used as a marker of lysosome or acidic compartments, according to the manufacturer s protocol. Plasmid independent
information MATERIAL AND METHODS Lysosome staining LysoTracker Red DND-99 (Invitrogen) was used as a marker of lysosome or acidic compartments, according to the manufacturer s protocol. Plasmid independent
Expanded View Figures
 Expanded View Figures Figure EV1. AM3-CLE45 control experiments and bam3 alleles. A Relative primary root length of indicated genotypes at 9 dag, in response to increasing amounts of CLE45 in the media.
Expanded View Figures Figure EV1. AM3-CLE45 control experiments and bam3 alleles. A Relative primary root length of indicated genotypes at 9 dag, in response to increasing amounts of CLE45 in the media.
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/5/244/ra72/dc1 Supplementary Materials for An Interaction Between BZR1 and DELLAs Mediates Direct Signaling Crosstalk Between Brassinosteroids and Gibberellins
www.sciencesignaling.org/cgi/content/full/5/244/ra72/dc1 Supplementary Materials for An Interaction Between BZR1 and DELLAs Mediates Direct Signaling Crosstalk Between Brassinosteroids and Gibberellins
Construct Design and Cloning Guide for Cas9-triggered homologous recombination
 Construct Design and Cloning Guide for Cas9-triggered homologous recombination Written by Dan Dickinson (ddickins@live.unc.edu) and last updated December 2013. Reference: Dickinson DJ, Ward JD, Reiner
Construct Design and Cloning Guide for Cas9-triggered homologous recombination Written by Dan Dickinson (ddickins@live.unc.edu) and last updated December 2013. Reference: Dickinson DJ, Ward JD, Reiner
J. Cell Sci. 128: doi: /jcs : Supplementary Material. Supplemental Figures. Journal of Cell Science Supplementary Material
 Supplemental Figures Figure S1. Trio controls endothelial barrier function. (A) TagRFP-shTrio constructs were expressed in ECs. Western blot shows efficient Trio knockdown in TagRFP-expressing ECs. (B)
Supplemental Figures Figure S1. Trio controls endothelial barrier function. (A) TagRFP-shTrio constructs were expressed in ECs. Western blot shows efficient Trio knockdown in TagRFP-expressing ECs. (B)
Supplementary Figure 1
 Supplementary Figure 1 Supplementary Fig. 1 shrna mediated knockdown of ZRSR2 in K562 and 293T cells. (a) ZRSR2 transcript levels in stably transduced K562 cells were determined using qrt-pcr. GAPDH was
Supplementary Figure 1 Supplementary Fig. 1 shrna mediated knockdown of ZRSR2 in K562 and 293T cells. (a) ZRSR2 transcript levels in stably transduced K562 cells were determined using qrt-pcr. GAPDH was
Supplemental Figure 1. Immunoblot analysis of wild-type and mutant forms of JAZ3
 Supplemental Data. Chung and Howe (2009). A Critical Role for the TIFY Motif in Repression of Jasmonate Signaling by a Stabilized Splice Variant of the JASMONATE ZIM-domain protein JAZ10 in Arabidopsis.
Supplemental Data. Chung and Howe (2009). A Critical Role for the TIFY Motif in Repression of Jasmonate Signaling by a Stabilized Splice Variant of the JASMONATE ZIM-domain protein JAZ10 in Arabidopsis.
Optimizing Cas9 and sgrna concentrations for increasing mutation efficiency. Nature Methods: doi: /nmeth.3360
 Supplementary Figure 1 Optimizing Cas9 and sgrna concentrations for increasing mutation efficiency. (a) Wild-type, 2-dpf slc24a5 b1 embryos and a series of slc24a5 CRISPR-injected embryos; anterior is
Supplementary Figure 1 Optimizing Cas9 and sgrna concentrations for increasing mutation efficiency. (a) Wild-type, 2-dpf slc24a5 b1 embryos and a series of slc24a5 CRISPR-injected embryos; anterior is
T H E J O U R N A L O F C E L L B I O L O G Y
 T H E J O U R N A L O F C E L L B I O L O G Y Supplemental material Rainero et al., http://www.jcb.org/cgi/content/full/jcb.201109112/dc1 Figure S1. The expression of DGK- is reduced upon transfection
T H E J O U R N A L O F C E L L B I O L O G Y Supplemental material Rainero et al., http://www.jcb.org/cgi/content/full/jcb.201109112/dc1 Figure S1. The expression of DGK- is reduced upon transfection
To assess the localization of Citrine fusion proteins, we performed antibody staining to
 Trinh et al 1 SUPPLEMENTAL MATERIAL FlipTraps recapitulate endogenous protein localization To assess the localization of Citrine fusion proteins, we performed antibody staining to compare the expression
Trinh et al 1 SUPPLEMENTAL MATERIAL FlipTraps recapitulate endogenous protein localization To assess the localization of Citrine fusion proteins, we performed antibody staining to compare the expression
A Nucleus-Encoded Chloroplast Protein YL1 Is Involved in Chloroplast. Development and Efficient Biogenesis of Chloroplast ATP Synthase in Rice
 A Nucleus-Encoded Chloroplast Protein YL1 Is Involved in Chloroplast Development and Efficient Biogenesis of Chloroplast ATP Synthase in Rice Fei Chen 1,*, Guojun Dong 2,*, Limin Wu 1, Fang Wang 3, Xingzheng
A Nucleus-Encoded Chloroplast Protein YL1 Is Involved in Chloroplast Development and Efficient Biogenesis of Chloroplast ATP Synthase in Rice Fei Chen 1,*, Guojun Dong 2,*, Limin Wu 1, Fang Wang 3, Xingzheng
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb3363 Supplementary Figure 1 Several WNTs bind to the extracellular domains of PKD1. (a) HEK293T cells were co-transfected with indicated plasmids. Flag-tagged proteins were immunoprecipiated
DOI: 10.1038/ncb3363 Supplementary Figure 1 Several WNTs bind to the extracellular domains of PKD1. (a) HEK293T cells were co-transfected with indicated plasmids. Flag-tagged proteins were immunoprecipiated
amplification of the 5 flanking region of CAT1 with a tail for the geneticin resistance gene cassette fusion CAT1-3F
 Supplementary materials Table S1. Primer list. Primer Sequence a (5-3 ) Description CAT1-5F CAT1-5R ATACGGATAAGGAAGCGATAGCAGCA gcacaggtacacttgtttagagagcgatccggattttaagtgaacg amplification of the 5 flanking
Supplementary materials Table S1. Primer list. Primer Sequence a (5-3 ) Description CAT1-5F CAT1-5R ATACGGATAAGGAAGCGATAGCAGCA gcacaggtacacttgtttagagagcgatccggattttaagtgaacg amplification of the 5 flanking
Supplemental Data. Sethi et al. (2014). Plant Cell /tpc
 Supplemental Data Supplemental Figure 1. MYC2 Binds to the E-box but not the E1-box of the MPK6 Promoter. (A) E1-box and E-box (wild type) containing MPK6 promoter fragment. The region shown in red denotes
Supplemental Data Supplemental Figure 1. MYC2 Binds to the E-box but not the E1-box of the MPK6 Promoter. (A) E1-box and E-box (wild type) containing MPK6 promoter fragment. The region shown in red denotes
Agouti C57BL/6N embryonic stem cells for mouse genetic resources
 nature methods Agouti N embryonic stem cells for mouse genetic resources Stephen J Pettitt, Qi Liang, Xin Y Rairdan, Jennifer L Moran, Haydn M Prosser, David R Beier, Kent Lloyd, Allan Bradley & William
nature methods Agouti N embryonic stem cells for mouse genetic resources Stephen J Pettitt, Qi Liang, Xin Y Rairdan, Jennifer L Moran, Haydn M Prosser, David R Beier, Kent Lloyd, Allan Bradley & William
Supplementary Figure 1. Nature Structural & Molecular Biology: doi: /nsmb.3494
 Supplementary Figure 1 Pol structure-function analysis (a) Inactivating polymerase and helicase mutations do not alter the stability of Pol. Flag epitopes were introduced using CRISPR/Cas9 gene targeting
Supplementary Figure 1 Pol structure-function analysis (a) Inactivating polymerase and helicase mutations do not alter the stability of Pol. Flag epitopes were introduced using CRISPR/Cas9 gene targeting
(phosphatase tensin) domain is shown in dark gray, the FH1 domain in black, and the
 Supplemental Figure 1. Predicted Domain Organization of the AFH14 Protein. (A) Schematic representation of the predicted domain organization of AFH14. The PTEN (phosphatase tensin) domain is shown in dark
Supplemental Figure 1. Predicted Domain Organization of the AFH14 Protein. (A) Schematic representation of the predicted domain organization of AFH14. The PTEN (phosphatase tensin) domain is shown in dark
Sol narae (Sona) is a Drosophila ADAMTS involved in Wg signaling
 Sol narae (Sona) is a Drosophila ADAMTS involved in Wg signaling Go-Woon Kim, Jong-Hoon Won, Ok-Kyung Lee, Sang-Soo Lee, Jeong-Hoon Han, Orkhon Tsogtbaatar, Sujin Nam, Yeon Kim, and Kyung-Ok Cho* Supplementary
Sol narae (Sona) is a Drosophila ADAMTS involved in Wg signaling Go-Woon Kim, Jong-Hoon Won, Ok-Kyung Lee, Sang-Soo Lee, Jeong-Hoon Han, Orkhon Tsogtbaatar, Sujin Nam, Yeon Kim, and Kyung-Ok Cho* Supplementary
Supplementary Figure 1
 Supplementary Figure 1 Supplementary Figure 1. Activation capacity of tettale-ad compared to tet trans-activator (ttas) using different teto variants. (A) HeLa cells were co-transfected with activation
Supplementary Figure 1 Supplementary Figure 1. Activation capacity of tettale-ad compared to tet trans-activator (ttas) using different teto variants. (A) HeLa cells were co-transfected with activation
scgem Workflow Experimental Design Single cell DNA methylation primer design
 scgem Workflow Experimental Design Single cell DNA methylation primer design The scgem DNA methylation assay uses qpcr to measure digestion of target loci by the methylation sensitive restriction endonuclease
scgem Workflow Experimental Design Single cell DNA methylation primer design The scgem DNA methylation assay uses qpcr to measure digestion of target loci by the methylation sensitive restriction endonuclease
indicated numbers of pups at day of life (DOL) 10, or embryonic day (ED) B. Male mice of
 SUPPLEMENTRY FIGURE LEGENDS Figure S1. USP44 loss leads to chromosome missegregation.. Genotypes obtained from the indicated numbers of pups at day of life (DOL) 10, or embryonic day (ED) 13.5.. Male mice
SUPPLEMENTRY FIGURE LEGENDS Figure S1. USP44 loss leads to chromosome missegregation.. Genotypes obtained from the indicated numbers of pups at day of life (DOL) 10, or embryonic day (ED) 13.5.. Male mice
B. Transgenic plants with strong phenotype (%)
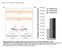 A. TCTAGTTGTTGTTGTTATGGTCTAGTTGTTGTTGTTATGGTCTAATTT AAATATGGTCTAAAGAAGAAGAATATGGTCTAAAGAAGAAGAATATGG 2XP35S STTM165 5 GGGGGATGAAGctaCCTGGTCCGA3 3 CCCCCUACUUC---GGACCAGGCU5 mir165 HindIII mir165 96 nt GTTGTTGTTGTTATGGTCTAGTTGTTGTTGTTATGGTCTAATTT
A. TCTAGTTGTTGTTGTTATGGTCTAGTTGTTGTTGTTATGGTCTAATTT AAATATGGTCTAAAGAAGAAGAATATGGTCTAAAGAAGAAGAATATGG 2XP35S STTM165 5 GGGGGATGAAGctaCCTGGTCCGA3 3 CCCCCUACUUC---GGACCAGGCU5 mir165 HindIII mir165 96 nt GTTGTTGTTGTTATGGTCTAGTTGTTGTTGTTATGGTCTAATTT
To investigate the heredity of the WFP gene, we selected plants that were homozygous
 Supplementary information Supplementary Note ST-12 WFP allele is semi-dominant To investigate the heredity of the WFP gene, we selected plants that were homozygous for chromosome 1 of Nipponbare and heterozygous
Supplementary information Supplementary Note ST-12 WFP allele is semi-dominant To investigate the heredity of the WFP gene, we selected plants that were homozygous for chromosome 1 of Nipponbare and heterozygous
Supporting Information Defined Bilayer Interactions of DNA Nanopores Revealed with a Nuclease-Based Nanoprobe Strategy
 Supporting Information Defined Bilayer Interactions of DNA Nanopores Revealed with a Nuclease-Based Nanoprobe Strategy Jonathan R. Burns* & Stefan Howorka* 1 Contents 1. Design of DNA nanopores... 3 1.1.
Supporting Information Defined Bilayer Interactions of DNA Nanopores Revealed with a Nuclease-Based Nanoprobe Strategy Jonathan R. Burns* & Stefan Howorka* 1 Contents 1. Design of DNA nanopores... 3 1.1.
Fluorescent labeling of the nuclear envelope by localizing GFP on the inner nuclear membrane
 Supporting Information Fluorescent labeling of the nuclear envelope by localizing GFP on the inner nuclear membrane Toshiyuki Taniyama 1, Natsumi Tsuda 1, and Shinji Sueda 1,2,* 1 Department of Bioscience
Supporting Information Fluorescent labeling of the nuclear envelope by localizing GFP on the inner nuclear membrane Toshiyuki Taniyama 1, Natsumi Tsuda 1, and Shinji Sueda 1,2,* 1 Department of Bioscience
Supplemental Material
 Supplemental Material 1 Figure S1. Phylogenetic analysis of Cep72 and Lrrc36, comparative localization of Cep72 and Lrrc36 and Cep72 antibody characterization (A) Phylogenetic alignment of Cep72 and Lrrc36
Supplemental Material 1 Figure S1. Phylogenetic analysis of Cep72 and Lrrc36, comparative localization of Cep72 and Lrrc36 and Cep72 antibody characterization (A) Phylogenetic alignment of Cep72 and Lrrc36
SUPPLEMENTARY INFORMATION
 DOI:.38/ncb327 a b Sequence coverage (%) 4 3 2 IP: -GFP isoform IP: GFP IP: -GFP IP: GFP Sequence coverage (%) 4 3 2 IP: -GFP IP: GFP 33 52 58 isoform 2 33 49 47 IP: Control IP: Peptide Sequence Start
DOI:.38/ncb327 a b Sequence coverage (%) 4 3 2 IP: -GFP isoform IP: GFP IP: -GFP IP: GFP Sequence coverage (%) 4 3 2 IP: -GFP IP: GFP 33 52 58 isoform 2 33 49 47 IP: Control IP: Peptide Sequence Start
Integrated Omics Study Delineates the Dynamics of Lipid Droplets in Rhodococcus Opacus PD630
 School of Natural Sciences and Mathematics 2013-10-22 Integrated Omics Study Delineates the Dynamics of Lipid Droplets in Rhodococcus Opacus PD630 UTD AUTHOR(S): Michael Qiwei Zhang 2013 The Authors This
School of Natural Sciences and Mathematics 2013-10-22 Integrated Omics Study Delineates the Dynamics of Lipid Droplets in Rhodococcus Opacus PD630 UTD AUTHOR(S): Michael Qiwei Zhang 2013 The Authors This
Supplemental Data. Hachez et al. Plant Cell (2014) /tpc Suppl. Figure 1A
 Suppl. Figure 1A Suppl. Figure 1B Supplemental Figure 1: Results of the commercial screening of interactants using split ubiquitin technique. (A) Isolated preys (192) using the bait construct pbt3-n- as
Suppl. Figure 1A Suppl. Figure 1B Supplemental Figure 1: Results of the commercial screening of interactants using split ubiquitin technique. (A) Isolated preys (192) using the bait construct pbt3-n- as
7.012 Problem Set 5. Question 1
 Name Section 7.012 Problem Set 5 Question 1 While studying the problem of infertility, you attempt to isolate a hypothetical rabbit gene that accounts for the prolific reproduction of rabbits. After much
Name Section 7.012 Problem Set 5 Question 1 While studying the problem of infertility, you attempt to isolate a hypothetical rabbit gene that accounts for the prolific reproduction of rabbits. After much
Supplementary Table 1. The Q-PCR primer sequence is summarized in the following table.
 Supplementary Table 1. The Q-PCR primer sequence is summarized in the following table. Name Sequence (5-3 ) Application Flag-u ggactacaaggacgacgatgac Shared upstream primer for all the amplifications of
Supplementary Table 1. The Q-PCR primer sequence is summarized in the following table. Name Sequence (5-3 ) Application Flag-u ggactacaaggacgacgatgac Shared upstream primer for all the amplifications of
Supplementary Figures
 Supplementary Figures Supplementary Figure S1. Inhibition kinetics of p53/hdm2 binding induced by Nutlin 3 on live cells. Reproducibility of the p53/hdm2 binding disruption kinetics induced by Nutlin 3
Supplementary Figures Supplementary Figure S1. Inhibition kinetics of p53/hdm2 binding induced by Nutlin 3 on live cells. Reproducibility of the p53/hdm2 binding disruption kinetics induced by Nutlin 3
Garrity_SupplementaryFig.1 A
 Garrity_SupplementaryFig.1 dtrp1 dtrp1 ins otential protein roducts of dtrp1ins Supplementary Figure 1 : The dtrp ins mutant. a, dtrp1 ins was created via a sitedirected insertional disruption. This loop-in
Garrity_SupplementaryFig.1 dtrp1 dtrp1 ins otential protein roducts of dtrp1ins Supplementary Figure 1 : The dtrp ins mutant. a, dtrp1 ins was created via a sitedirected insertional disruption. This loop-in
Supplemental Figure 1.
 Supplemental Data. Charron et al. Dynamic landscapes of four histone modifications during de-etiolation in Arabidopsis. Plant Cell (2009). 10.1105/tpc.109.066845 Supplemental Figure 1. Immunodetection
Supplemental Data. Charron et al. Dynamic landscapes of four histone modifications during de-etiolation in Arabidopsis. Plant Cell (2009). 10.1105/tpc.109.066845 Supplemental Figure 1. Immunodetection
