Supplementary Table S1
|
|
|
- Erick Ellis
- 5 years ago
- Views:
Transcription
1 Primers used in RT-qPCR, ChIP and Bisulphite-Sequencing. Quantitative real-time RT-PCR primers Supplementary Table S1 gene Forward primer sequence Reverse primer sequence Product TRAIL CAACTCCGTCAGCTCGTTAGAAAG CGGCCCAGAGCCTTTTCATTC 200 IRF1 GCAGGCCCTGACTCCAGCAC TGCCACTCCGACTGCTCCAA 213 STAT1 CTGCTCCCTCTCTGGAATGATGGGTGCATC GAAGTCAGGTTCGCCTCCGTTCTGGGAC 180 DR5 TCCACCTGGACACCATATCTCAGAA TCCACTTCACCTGAATCACACCTG 132 DR4 CTGCTGCAGGTCGTACCTAGCTC GATCCTGGTGGACACAACTCTCC 110 APAF1 GCAGCCAGCTTCAGGATCTAC CAAAGTTCCTTGTGCATCTTGG 154 BID TGGACTGTGAGGTCAACAACG AGTCTGCAGCTCATCGTAGCC 176 CASPASE 8 TTTCTGCCTACAGGGTCATGCTC GTTCATGTCATCATCCAGTTTGC 123 XAF1 GCTCGGGAAAGGGGAAAGAATTT TCTGAGTCTGGACAACATTTACCC 120 DcR1 TCCTGCTGCCAGTCCTAGCTTAC TGAGATCCTGCTGGACACTCCTC 126 DcR2 GCCGGTCCGGGTTGACTC TGAGATCCTGCTGGACACTCCT 117 BCL2 TGGGATGCCTTTGTGGAACTGTA ATATTTGTTTGGGGCAGGCATGT 176 ciap1 TCAGCTAGTCTGGGATCCACCTC AGGGTTTGGAGAAAGGCTGGAGT 120 ciap2 TTCCGGGTACAGAAAACAGTGGA TGGTAGGAACTTCTCATCAAGGCAGA 115 XIAP GCAAGAGCTGGATTTTATGCTTTAGG CAGATATTTGCACCCTGGATACCAT 138 cmyc CCCGGAGTTGGAAAACAATGAAA GTCGTTTCCGCAACAAGTCCTCT 128 EGFR GGTGGCATTTAGGGGTGACTC CTGACCATGTTGCTTGGTCCT 187 ERBB2 GACTGCCTGTCCCTACAACTACC CCCAGACCATAGCACACTCG 150 CCND1 CCCTCGGTGTCCTACTTCAAA AGGAAGCGGTCCAGGTAGTTC 149 SHH CCAGAAACTCCGAGCGATTTA ATGGCCAAAGCGTTCAACTT 133 PAX-2 CAAATCAGAACAGGGGAACGA GCCATGTCACGACCAGTCAC 153 cmet GGTTGTGGTTTCTCGATCAGG TTCGTGATCTTCTTCCCAGTGA 144 GAPDH CATGGCACCGTCAAGGCTGA TGGTGGTGCAGGAGGCATTG 295
2 TRAIL (mouse) TCCAATCTCCAAGGATGGAAAGA GATGTAATACAGGCCCTCCTGCTC 139 DR5 (mouse) TGCTGTGCTACAGGCTGTCTTTG CTGACAGGTACTGGCCTGCTAGA B4 (mouse) AATCTCCAGAGGCACCATTG CCGATCTGCAGACACACACT 252 ChIP primers gene Position relative to the TSS Forward primer sequence Reverse primer sequence Product TRAIL > GCCGACTCTTGTAACTCCTCAAAT TATGATGTACGTGAAAGGCATGAA > GATAGAAGGCAAGGGCAGGAAGTG GTTTAGTTATTCCATTCTTGACCCACTG > -607 CATCTGATAGTGGGGAGATTTGG TGTTAGGCTGGACAGGTAGGAAG > +51 AGCTTCTTTCAGTTTCCCTCCTTT TCACTGAAGCCCTTCCTTCTCTAT > CAGGTTCTTTGGTGCCCATT TCGCAGGAATTCTCATGTCC > CTACCTTCTGCCCCCAATGC GAACTTGGCCAATGCTGGTG 203 MYO > TGGCACATAAAGTTCTTCCCAGTA TAGGAGTTTGTTCAGAGGTGATGG 170 MYOD > +654 CCGCCTGAGCAAAGTAAATGA GGCAACCGCTGGTTTGG 75 IRF > -44 CGGAGCAGCCGCCCTGTACTTCCCCT TCGCC GCGCCACCGAGCAATCCAAACACTTAGCG 165
3 Primers used for PCR amplification before bisulphite sequencing Sequence T A ( C) PCR product Region amplified TRAIL Intron1/Exon1 Region A Forward 5 -GATTATGGTTATGATGGAGGTTTAG Reverse 5 -AAAACAAAAAAATCTTCAAAAAAAA-3 TRAIL CpG island 3 Region B Forward 5 -AATATAGGGGAGAAGATTAAAGAAA Reverse 5 -CCTCCTAAAATACTAAAATTACAAA-3 TRAIL CpG island 5 Region C Forward 5 -TGTAATTTTAGTATTTTAGGAGGT Reverse 5 -TAAAACTACAAACACCCACCAC-3 Sequences of the primers used for amplification of bisulphite-modified DNA are listed with the annealing temperature (T A ) used. In addition, the length of the PCR product and the location of the region amplified relative to the transcriptional start site (TSS) (+1) are indicated.
4 Supplementary Figure Legends Lund et al. Transformation-dependent Silencing of Tumor-selective Apoptogenic TRAIL by DNA Hypermethylation is Antagonized by Decitabine Figure S1. Transcription-factor binding and histone-modifications at the TRAIL promoter of HA1E and HA1ER. A, Illustration of the TRAIL proximal promoter with the positions of transcription factor binding sites relative to the transcription start site (TSS; +1) indicating the locations of the regions analyzed by qpcr after ChIP. B, ChIP assays on the TRAIL promoter performed with antibodies against STAT1. The fold enrichment is the relative abundance of DNA fragments at the indicated regions over a transcriptional inactive control region (TSS of Myoglobin), n = 2. E-J, ChIP assays on the TRAIL promoter performed with antibodies against E, H3K27me3; F, H3; G, H3K4me1; H, H3K9me1; I, H3K9me3 and J, H3K27me1. The fold enrichment at known target sites is shown to the right hand of the graphs for B, the STAT1 binding at the IRF1 promoter, for C and D, the SP1 and SP3 binding at the DHFR promoter and for E, the H3K27me3 enrichment at the MyoD1 promoter. HA1E and HA1ER cells were either untreated (gray bars) or incubated with IFNγ (10 ng/ml; 18h; black bars). Bars indicate the fold of enrichment that is the relative abundance of DNA fragments at the indicated regions over a transcriptional inactive control region (TSS of Myoglobin), n = 2. Data are means ± SD. Figure S2. TRAIL induction through DNA demethylation of promoter regions by decitabine. A, Time-course of TRAIL mrna expression after treatment with decitabine in HA1E and HA1ER measured by RT-qPCR, n = 2. Data are means ± SEM. B, Bisulphite-Sequencing of the Exon1/Intron1 TRAIL promoter region (right) and the region covering the 5 part of the CpG-island 2kb upstream of the TSS (left). Comparison of per cent methylation for each
5 individual CpG (numbered in 5 to 3 direction) in untreated HA1ER (gray bars) versus decitabine treated HA1ER (black bars) is shown. C, Representation of the Bisulphite sequencing results displaying all single clones analysed. The individual CpG-sites (circles) are depicted as methylated (black) or unmethylated (white). Figure S3. Synergistic apoptosis induction by combination of TRAIL and decitabine. Light microscopic images of HA1ER treated with TRAIL, decitabine or a combination of both drugs demonstrates synergistic apoptosis induction by the combinatorial treatment; 20x magnification. Figure S4. TRAIL induction and antitumor effect of decitabine in syngeneic mice models. A, A, RT-qPCR analysis for TRAIL- and DR5-induction after treatment with decitabine, n = 3 and corresponding Western blot for TRAIL in C26 mouse colon tumor cells. B, Decitabine reduces tumor growth in a syngeneic BALB/c tumor model. Upper Left: Normalized tumor volume of C26-grafted mice after treatment with decitabine. Upper right: RT-qPCR for TRAIL across tumor samples, n = 3. Bottom: Normalized animal weight profile in in C26- syngeneic tumors. Administration of decitabine (1.5) (black squares) does not cause toxicity as observed by the lack of change in body weight of animals. The fold of mrna induction is normalized to the untreated cells (set to 1). Samples were taken at the day of sacrifice. Data are means ± SEM. *P <0.05, **P <0.01, ***P < Figure S5. Regulation of proto-oncogenes by decitabine. RT-qPCR analysis for regulation of proto-oncogene mrna levels after treatment with decitabine (10µM) for 96 hours, n = 3. The fold change is indicated relative to the untreated cells (>1 = Up-regulation, 1 = unchanged, < 1 = down-regulation). Data are means ± SEM.
Supplemental Figure 1.
 Supplemental Data. Charron et al. Dynamic landscapes of four histone modifications during de-etiolation in Arabidopsis. Plant Cell (2009). 10.1105/tpc.109.066845 Supplemental Figure 1. Immunodetection
Supplemental Data. Charron et al. Dynamic landscapes of four histone modifications during de-etiolation in Arabidopsis. Plant Cell (2009). 10.1105/tpc.109.066845 Supplemental Figure 1. Immunodetection
Suppl. Table S1. Characteristics of DHS regions analyzed by bisulfite sequencing. No. CpGs analyzed in the amplicon. Genomic location specificity
 Suppl. Table S1. Characteristics of DHS regions analyzed by bisulfite sequencing. DHS/GRE Genomic location Tissue specificity DHS type CpG density (per 100 bp) No. CpGs analyzed in the amplicon CpG within
Suppl. Table S1. Characteristics of DHS regions analyzed by bisulfite sequencing. DHS/GRE Genomic location Tissue specificity DHS type CpG density (per 100 bp) No. CpGs analyzed in the amplicon CpG within
Supplementary Figure 1. Expressions of stem cell markers decreased in TRCs on 2D plastic. TRCs were cultured on plastic for 1, 3, 5, or 7 days,
 Supplementary Figure 1. Expressions of stem cell markers decreased in TRCs on 2D plastic. TRCs were cultured on plastic for 1, 3, 5, or 7 days, respectively, and their mrnas were quantified by real time
Supplementary Figure 1. Expressions of stem cell markers decreased in TRCs on 2D plastic. TRCs were cultured on plastic for 1, 3, 5, or 7 days, respectively, and their mrnas were quantified by real time
ChampionChIP Quick, High Throughput Chromatin Immunoprecipitation Assay System
 ChampionChIP Quick, High Throughput Chromatin Immunoprecipitation Assay System Liyan Pang, Ph.D. Application Scientist 1 Topics to be Covered Introduction What is ChIP-qPCR? Challenges Facing Biological
ChampionChIP Quick, High Throughput Chromatin Immunoprecipitation Assay System Liyan Pang, Ph.D. Application Scientist 1 Topics to be Covered Introduction What is ChIP-qPCR? Challenges Facing Biological
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb3240 Supplementary Figure 1 GBM cell lines display similar levels of p100 to p52 processing but respond differentially to TWEAK-induced TERT expression according to TERT promoter mutation
DOI: 10.1038/ncb3240 Supplementary Figure 1 GBM cell lines display similar levels of p100 to p52 processing but respond differentially to TWEAK-induced TERT expression according to TERT promoter mutation
CRISPR-dCas9 mediated TET1 targeting for selective DNA demethylation at BRCA1 promoter
 CRISPR-dCas9 mediated TET1 targeting for selective DNA demethylation at BRCA1 promoter SUPPLEMENTARY DATA See Supplementary Sequence File: 1 Supplementary Figure S1: The total protein was extracted from
CRISPR-dCas9 mediated TET1 targeting for selective DNA demethylation at BRCA1 promoter SUPPLEMENTARY DATA See Supplementary Sequence File: 1 Supplementary Figure S1: The total protein was extracted from
T-iPSC. Gra-iPSC. B-iPSC. TTF-iPSC. Supplementary Figure 1. Nature Biotechnology: doi: /nbt.1667
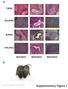 a T-iPSC Gra-iPSC B-iPSC TTF-iPSC Ectoderm Endoderm Mesoderm b Nature Biotechnology: doi:.38/nbt.667 Supplementary Figure Klf4 transgene expression Oct4 transgene expression.7 Fold GAPDH.6.5.4.3 Fold GAPDH
a T-iPSC Gra-iPSC B-iPSC TTF-iPSC Ectoderm Endoderm Mesoderm b Nature Biotechnology: doi:.38/nbt.667 Supplementary Figure Klf4 transgene expression Oct4 transgene expression.7 Fold GAPDH.6.5.4.3 Fold GAPDH
Supplementary Figure 1. jmj30-2 and jmj32-1 produce null mutants. (a) Schematic drawing of JMJ30 and JMJ32 genome structure showing regions amplified
 Supplementary Figure 1. jmj30-2 and jmj32-1 produce null mutants. (a) Schematic drawing of JMJ30 and JMJ32 genome structure showing regions amplified by primers used for mrna expression analysis. Gray
Supplementary Figure 1. jmj30-2 and jmj32-1 produce null mutants. (a) Schematic drawing of JMJ30 and JMJ32 genome structure showing regions amplified by primers used for mrna expression analysis. Gray
Figure S1. nuclear extracts. HeLa cell nuclear extract. Input IgG IP:ORC2 ORC2 ORC2. MCM4 origin. ORC2 occupancy
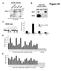 A nuclear extracts B HeLa cell nuclear extract Figure S1 ORC2 (in kda) 21 132 7 ORC2 Input IgG IP:ORC2 32 ORC C D PRKDC ORC2 occupancy Directed against ORC2 C-terminus (sc-272) MCM origin 2 2 1-1 -1kb
A nuclear extracts B HeLa cell nuclear extract Figure S1 ORC2 (in kda) 21 132 7 ORC2 Input IgG IP:ORC2 32 ORC C D PRKDC ORC2 occupancy Directed against ORC2 C-terminus (sc-272) MCM origin 2 2 1-1 -1kb
File name: Supplementary Information Description: Supplementary figures and supplementary tables. File name: Peer review file Description:
 File name: Supplementary Information Description: Supplementary figures and supplementary tables. File name: Peer review file Description: Supplementary Figure 1. dcas9-mq1 fusion protein induces de novo
File name: Supplementary Information Description: Supplementary figures and supplementary tables. File name: Peer review file Description: Supplementary Figure 1. dcas9-mq1 fusion protein induces de novo
Fig. S1. Effect of p120-catenin overexpression on the interaction of SCUBE2 with E-cadherin. The expression plasmid encoding FLAG.
 Fig. S1. Effect of p120-catenin overexpression on the interaction of SCUBE2 with E-cadherin. The expression plasmid encoding FLAG.SCUBE2, E-cadherin.Myc, or HA.p120-catenin was transfected in a combination
Fig. S1. Effect of p120-catenin overexpression on the interaction of SCUBE2 with E-cadherin. The expression plasmid encoding FLAG.SCUBE2, E-cadherin.Myc, or HA.p120-catenin was transfected in a combination
Supplemental Table 1 Gene Symbol FDR corrected p-value PLOD1 CSRP2 PFKP ADFP ADM C10orf10 GPI LOX PLEKHA2 WIPF1
 Supplemental Table 1 Gene Symbol FDR corrected p-value PLOD1 4.52E-18 PDK1 6.77E-18 CSRP2 4.42E-17 PFKP 1.23E-14 MSH2 3.79E-13 NARF_A 5.56E-13 ADFP 5.56E-13 FAM13A1 1.56E-12 FAM29A_A 1.22E-11 CA9 1.54E-11
Supplemental Table 1 Gene Symbol FDR corrected p-value PLOD1 4.52E-18 PDK1 6.77E-18 CSRP2 4.42E-17 PFKP 1.23E-14 MSH2 3.79E-13 NARF_A 5.56E-13 ADFP 5.56E-13 FAM13A1 1.56E-12 FAM29A_A 1.22E-11 CA9 1.54E-11
Supplementary Tables. Primers and probes. Target Primer / probe sequences Chemistry Human HBB F: AACTGTGTTCACTAGCAACCTCAAA
 Supplementary Tables Primers and probes Target Primer / probe sequences Chemistry H F: AACTGTGTTCACTAGCAACCTCAAA promoter R: ACAGGGCAGTAACGGCAGACT H Downstream Actb alpha (162) alpha (162.) (Pro) (16)
Supplementary Tables Primers and probes Target Primer / probe sequences Chemistry H F: AACTGTGTTCACTAGCAACCTCAAA promoter R: ACAGGGCAGTAACGGCAGACT H Downstream Actb alpha (162) alpha (162.) (Pro) (16)
Apoptosis assay: Apoptotic cells were identified by Annexin V-Alexa Fluor 488 and Propidium
 Apoptosis assay: Apoptotic cells were identified by Annexin V-Alexa Fluor 488 and Propidium Iodide (Invitrogen, Carlsbad, CA) staining. Briefly, 2x10 5 cells were washed once in cold PBS and resuspended
Apoptosis assay: Apoptotic cells were identified by Annexin V-Alexa Fluor 488 and Propidium Iodide (Invitrogen, Carlsbad, CA) staining. Briefly, 2x10 5 cells were washed once in cold PBS and resuspended
Supplementary Data. Flvcr1a TCTAAGGCCCAGTAGGACCC GGCCTCAACTGCCTGGGAGC AGAGGGCAACCTCGGTGTCC
 Supplementary Data Supplementary Materials and Methods Measurement of reactive oxygen species accumulation in fresh intestinal rings Accumulation of reactive oxygen species in fresh intestinal rings was
Supplementary Data Supplementary Materials and Methods Measurement of reactive oxygen species accumulation in fresh intestinal rings Accumulation of reactive oxygen species in fresh intestinal rings was
SUPPLEMENTARY INFORMATION
 (Supplementary Methods and Materials) GST pull-down assay GST-fusion proteins Fe65 365-533, and Fe65 538-700 were expressed in BL21 bacterial cells and purified with glutathione-agarose beads (Sigma).
(Supplementary Methods and Materials) GST pull-down assay GST-fusion proteins Fe65 365-533, and Fe65 538-700 were expressed in BL21 bacterial cells and purified with glutathione-agarose beads (Sigma).
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Endogenous gene tagging to study subcellular localization and chromatin binding. a, b, Schematic of experimental set-up to endogenously tag RNAi factors using the CRISPR Cas9 technology,
Supplementary Figure 1 Endogenous gene tagging to study subcellular localization and chromatin binding. a, b, Schematic of experimental set-up to endogenously tag RNAi factors using the CRISPR Cas9 technology,
Genomic DNA fragments identified by ChIP-chip assay for MR [28] corresponding to two
![Genomic DNA fragments identified by ChIP-chip assay for MR [28] corresponding to two Genomic DNA fragments identified by ChIP-chip assay for MR [28] corresponding to two](/thumbs/89/98549522.jpg) Supplemental Materials and Methods Genomic DNA fragments identified by ChIP-chip assay for MR [28] corresponding to two upstream regions of the human Klf9 gene (-5139 to -5771 bp and -3875 to -4211 bp)
Supplemental Materials and Methods Genomic DNA fragments identified by ChIP-chip assay for MR [28] corresponding to two upstream regions of the human Klf9 gene (-5139 to -5771 bp and -3875 to -4211 bp)
Jung-Nam Cho, Jee-Youn Ryu, Young-Min Jeong, Jihye Park, Ji-Joon Song, Richard M. Amasino, Bosl Noh, and Yoo-Sun Noh
 Developmental Cell, Volume 22 Supplemental Information Control of Seed Germination by Light-Induced Histone Arginine Demethylation Activity Jung-Nam Cho, Jee-Youn Ryu, Young-Min Jeong, Jihye Park, Ji-Joon
Developmental Cell, Volume 22 Supplemental Information Control of Seed Germination by Light-Induced Histone Arginine Demethylation Activity Jung-Nam Cho, Jee-Youn Ryu, Young-Min Jeong, Jihye Park, Ji-Joon
The RT-qPCR analysis of selected type-i IFNs related genes, IRF7 and Oas3. The
 SUPPLEMENTARY MATERIALS AND METHODS Real time quantitative PCR The RT-qPCR analysis of selected type-i IFNs related genes, IRF7 and Oas3. The RT-qPCR was performed on the Applied Biosystems StepOne TM
SUPPLEMENTARY MATERIALS AND METHODS Real time quantitative PCR The RT-qPCR analysis of selected type-i IFNs related genes, IRF7 and Oas3. The RT-qPCR was performed on the Applied Biosystems StepOne TM
Nature Genetics: doi: /ng.3556 INTEGRATED SUPPLEMENTARY FIGURE TEMPLATE. Supplementary Figure 1
 INTEGRATED SUPPLEMENTARY FIGURE TEMPLATE Supplementary Figure 1 REF6 expression in transgenic lines. (a,b) Expression of REF6 in REF6-HA ref6 and REF6ΔZnF-HA ref6 plants detected by RT qpcr (a) and immunoblot
INTEGRATED SUPPLEMENTARY FIGURE TEMPLATE Supplementary Figure 1 REF6 expression in transgenic lines. (a,b) Expression of REF6 in REF6-HA ref6 and REF6ΔZnF-HA ref6 plants detected by RT qpcr (a) and immunoblot
Cell proliferation was measured with Cell Counting Kit-8 (Dojindo Laboratories, Kumamoto, Japan).
 1 2 3 4 5 6 7 8 Supplemental Materials and Methods Cell proliferation assay Cell proliferation was measured with Cell Counting Kit-8 (Dojindo Laboratories, Kumamoto, Japan). GCs were plated at 96-well
1 2 3 4 5 6 7 8 Supplemental Materials and Methods Cell proliferation assay Cell proliferation was measured with Cell Counting Kit-8 (Dojindo Laboratories, Kumamoto, Japan). GCs were plated at 96-well
Nature Biotechnology: doi: /nbt Supplementary Figure 1
 Supplementary Figure 1 Schematic and results of screening the combinatorial antibody library for Sox2 replacement activity. A single batch of MEFs were plated and transduced with doxycycline inducible
Supplementary Figure 1 Schematic and results of screening the combinatorial antibody library for Sox2 replacement activity. A single batch of MEFs were plated and transduced with doxycycline inducible
Partial list of differentially expressed genes from cdna microarray, comparing MUC18-
 Supplemental Figure legends Table-1 Partial list of differentially expressed genes from cdna microarray, comparing MUC18- silenced and NT-transduced A375SM cells. Supplemental Figure 1 Effect of MUC-18
Supplemental Figure legends Table-1 Partial list of differentially expressed genes from cdna microarray, comparing MUC18- silenced and NT-transduced A375SM cells. Supplemental Figure 1 Effect of MUC-18
Technical Review. Real time PCR
 Technical Review Real time PCR Normal PCR: Analyze with agarose gel Normal PCR vs Real time PCR Real-time PCR, also known as quantitative PCR (qpcr) or kinetic PCR Key feature: Used to amplify and simultaneously
Technical Review Real time PCR Normal PCR: Analyze with agarose gel Normal PCR vs Real time PCR Real-time PCR, also known as quantitative PCR (qpcr) or kinetic PCR Key feature: Used to amplify and simultaneously
Supplementary Fig. 1 related to Fig. 1 Clinical relevance of lncrna candidate
 Supplementary Figure Legends Supplementary Fig. 1 related to Fig. 1 Clinical relevance of lncrna candidate BC041951 in gastric cancer. (A) The flow chart for selected candidate lncrnas in 660 up-regulated
Supplementary Figure Legends Supplementary Fig. 1 related to Fig. 1 Clinical relevance of lncrna candidate BC041951 in gastric cancer. (A) The flow chart for selected candidate lncrnas in 660 up-regulated
Fig. S1. eif6 expression in HEK293 transfected with shrna against eif6 or pcmv-eif6 vector.
 Fig. S1. eif6 expression in HEK293 transfected with shrna against eif6 or pcmv-eif6 vector. (a) Western blotting analysis and (b) qpcr analysis of eif6 expression in HEK293 T cells transfected with either
Fig. S1. eif6 expression in HEK293 transfected with shrna against eif6 or pcmv-eif6 vector. (a) Western blotting analysis and (b) qpcr analysis of eif6 expression in HEK293 T cells transfected with either
supplementary information
 DOI: 1.138/ncb1839 a b Control 1 2 3 Control 1 2 3 Fbw7 Smad3 1 2 3 4 1 2 3 4 c d IGF-1 IGF-1Rβ IGF-1Rβ-P Control / 1 2 3 4 Real-time RT-PCR Relative quantity (IGF-1/ mrna) 2 1 IGF-1 1 2 3 4 Control /
DOI: 1.138/ncb1839 a b Control 1 2 3 Control 1 2 3 Fbw7 Smad3 1 2 3 4 1 2 3 4 c d IGF-1 IGF-1Rβ IGF-1Rβ-P Control / 1 2 3 4 Real-time RT-PCR Relative quantity (IGF-1/ mrna) 2 1 IGF-1 1 2 3 4 Control /
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1.
 Supplementary Figure 1. Characterization and high expression of Lnc-β-Catm in liver CSCs. (a) Heatmap of differently expressed lncrnas in Liver CSCs (CD13 + CD133 + ) and non-cscs (CD13 - CD133 - ) according
Supplementary Figure 1. Characterization and high expression of Lnc-β-Catm in liver CSCs. (a) Heatmap of differently expressed lncrnas in Liver CSCs (CD13 + CD133 + ) and non-cscs (CD13 - CD133 - ) according
Allele-specific locus binding and genome editing by CRISPR at the
 Supplementary Information Allele-specific locus binding and genome editing by CRISPR at the p6ink4a locus Toshitsugu Fujita, Miyuki Yuno, and Hodaka Fujii Supplementary Figure Legends Supplementary Figure
Supplementary Information Allele-specific locus binding and genome editing by CRISPR at the p6ink4a locus Toshitsugu Fujita, Miyuki Yuno, and Hodaka Fujii Supplementary Figure Legends Supplementary Figure
SUPPLEMENTARY INFORMATION
 Supplementary Discussion Interestingly, a recent study demonstrated that knockdown of Tet1 alone would lead to dysregulation of differentiation genes, but only minor defects in ES maintenance and no change
Supplementary Discussion Interestingly, a recent study demonstrated that knockdown of Tet1 alone would lead to dysregulation of differentiation genes, but only minor defects in ES maintenance and no change
Supplementary Information
 Supplementary Information Table of contents: Pages: Supplementary Methods 2-3 Supplementary Figure 1 4 Supplementary Figure 2 5 Supplementary Figure 3 6 Supplementary Figure 4 7 Supplementary Figure 5
Supplementary Information Table of contents: Pages: Supplementary Methods 2-3 Supplementary Figure 1 4 Supplementary Figure 2 5 Supplementary Figure 3 6 Supplementary Figure 4 7 Supplementary Figure 5
EPIGENTEK. EpiQuik Methylated DNA Immunoprecipitation Kit. Base Catalog # P-2019 PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE
 EpiQuik Methylated DNA Immunoprecipitation Kit Base Catalog # PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE The EpiQuik MeDIP Kit can be used for immunoprecipitating the methylated DNA from a broad range
EpiQuik Methylated DNA Immunoprecipitation Kit Base Catalog # PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE The EpiQuik MeDIP Kit can be used for immunoprecipitating the methylated DNA from a broad range
Nature Structural and Molecular Biology: doi: /nsmb Supplementary Figure 1. Validation of CDK9-inhibitor treatment.
 Supplementary Figure 1 Validation of CDK9-inhibitor treatment. (a) Schematic of GAPDH with the middle of the amplicons indicated in base pairs. The transcription start site (TSS) and the terminal polyadenylation
Supplementary Figure 1 Validation of CDK9-inhibitor treatment. (a) Schematic of GAPDH with the middle of the amplicons indicated in base pairs. The transcription start site (TSS) and the terminal polyadenylation
Chapter 20 Recombinant DNA Technology. Copyright 2009 Pearson Education, Inc.
 Chapter 20 Recombinant DNA Technology Copyright 2009 Pearson Education, Inc. 20.1 Recombinant DNA Technology Began with Two Key Tools: Restriction Enzymes and DNA Cloning Vectors Recombinant DNA refers
Chapter 20 Recombinant DNA Technology Copyright 2009 Pearson Education, Inc. 20.1 Recombinant DNA Technology Began with Two Key Tools: Restriction Enzymes and DNA Cloning Vectors Recombinant DNA refers
b alternative classical none
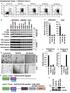 Supplementary Figure. 1: Related to Figure.1 a d e b alternative classical none NIK P-IkBa Total IkBa Tubulin P52 (Lighter) P52 (Darker) RelB (Lighter) RelB (Darker) HDAC1 Control-Sh RelB-Sh NF-kB2-Sh
Supplementary Figure. 1: Related to Figure.1 a d e b alternative classical none NIK P-IkBa Total IkBa Tubulin P52 (Lighter) P52 (Darker) RelB (Lighter) RelB (Darker) HDAC1 Control-Sh RelB-Sh NF-kB2-Sh
Mock. Control. sirna1
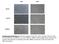 siha CaSki Mock Control sirna1 Supplementary Data Figure 1: Photomicrographs of siha and CaSki cells taken 48 hours after mock transfection, transfection of control sirna or sirna1. (100X magnification).
siha CaSki Mock Control sirna1 Supplementary Data Figure 1: Photomicrographs of siha and CaSki cells taken 48 hours after mock transfection, transfection of control sirna or sirna1. (100X magnification).
Nature Biotechnology: doi: /nbt Supplementary Figure 1. Design and sequence of system 1 for targeted demethylation.
 Supplementary Figure 1 Design and sequence of system 1 for targeted demethylation. (a) Design of system 1 for targeted demethylation. TET1CD was fused to a catalytic inactive Cas9 nuclease (dcas9) and
Supplementary Figure 1 Design and sequence of system 1 for targeted demethylation. (a) Design of system 1 for targeted demethylation. TET1CD was fused to a catalytic inactive Cas9 nuclease (dcas9) and
Supplementary Figure 2
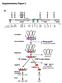 Supplementary Figure 2 a SBS-C1 SBS-C2 SBS-C3 SBS-C4 SBS-C5 SBS-C6 SBS-C7 SBS-C8 SBS-C9 LCR CNS-1 CNS-2 Il5 Rad50 Il13 Il4 Kif3a Sept8 0 50 100 150 200kb Sau3AI 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17
Supplementary Figure 2 a SBS-C1 SBS-C2 SBS-C3 SBS-C4 SBS-C5 SBS-C6 SBS-C7 SBS-C8 SBS-C9 LCR CNS-1 CNS-2 Il5 Rad50 Il13 Il4 Kif3a Sept8 0 50 100 150 200kb Sau3AI 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17
Table S1. Primers used in RT-PCR studies (all in 5 to 3 direction)
 Table S1. Primers used in RT-PCR studies (all in 5 to 3 direction) Epo Fw CTGTATCATGGACCACCTCGG Epo Rw TGAAGCACAGAAGCTCTTCGG Jak2 Fw ATCTGACCTTTCCATCTGGGG Jak2 Rw TGGTTGGGTGGATACCAGATC Stat5A Fw TTACTGAAGATCAAGCTGGGG
Table S1. Primers used in RT-PCR studies (all in 5 to 3 direction) Epo Fw CTGTATCATGGACCACCTCGG Epo Rw TGAAGCACAGAAGCTCTTCGG Jak2 Fw ATCTGACCTTTCCATCTGGGG Jak2 Rw TGGTTGGGTGGATACCAGATC Stat5A Fw TTACTGAAGATCAAGCTGGGG
AD BD TOC1. Supplementary Figure 1: Yeast two-hybrid assays showing the interaction between
 AD X BD TOC1 AD BD X PIFΔAD PIF TOC1 TOC1 PIFΔAD PIF N TOC1 TOC1 C1 PIFΔAD PIF C1 TOC1 TOC1 C PIFΔAD PIF C TOC1 Supplementary Figure 1: Yeast two-hybrid assays showing the interaction between PIF and TOC1
AD X BD TOC1 AD BD X PIFΔAD PIF TOC1 TOC1 PIFΔAD PIF N TOC1 TOC1 C1 PIFΔAD PIF C1 TOC1 TOC1 C PIFΔAD PIF C TOC1 Supplementary Figure 1: Yeast two-hybrid assays showing the interaction between PIF and TOC1
EpiQuik Methyl-CpG Binding Domain Protein 2 ChIP Kit
 EpiQuik Methyl-CpG Binding Domain Protein 2 ChIP Kit Base Catalog # PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE The EpiQuik Methyl-CpG Binding Domain Protein 2 ChIP Kit is suitable for combining the
EpiQuik Methyl-CpG Binding Domain Protein 2 ChIP Kit Base Catalog # PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE The EpiQuik Methyl-CpG Binding Domain Protein 2 ChIP Kit is suitable for combining the
Supplementary Materials and Methods:
 Supplementary Materials and Methods: Preclinical chemoprevention experimental design Mice were weighed and each mammary tumor was manually palpated and measured with digital calipers once a week from 4
Supplementary Materials and Methods: Preclinical chemoprevention experimental design Mice were weighed and each mammary tumor was manually palpated and measured with digital calipers once a week from 4
Nature Methods: doi: /nmeth Supplementary Figure 1
 Supplementary Figure 1 Workflow for multimodal analysis using sc-gem on a programmable microfluidic device (Fluidigm). 1) Cells are captured and lysed, 2) RNA from lysed single cell is reverse-transcribed
Supplementary Figure 1 Workflow for multimodal analysis using sc-gem on a programmable microfluidic device (Fluidigm). 1) Cells are captured and lysed, 2) RNA from lysed single cell is reverse-transcribed
Spermatazoa. Sertoli cells. Morc1 WT adult. Morc1 KO adult. Morc1 WT P14.5. Morc1 KO P14.5. Germ cells. Germ cells. Spermatids.
 a. Spermatazoa Sertoli cells Morc1 WT adult c. Morc1 KO adult Morc1 WT P1.5 Morc1 KO P1.5 Morc1 WT adult Morc1 KO adult Germ cells Germ cells Spermatids Spermatozoa Supplementary Figure 1. Confirmation
a. Spermatazoa Sertoli cells Morc1 WT adult c. Morc1 KO adult Morc1 WT P1.5 Morc1 KO P1.5 Morc1 WT adult Morc1 KO adult Germ cells Germ cells Spermatids Spermatozoa Supplementary Figure 1. Confirmation
Supplementary Methods Plasmid constructs
 Supplementary Methods Plasmid constructs. Mouse cdna encoding SHP-1, amplified from mrna of RAW264.7 macrophages with primer 5'cgtgcctgcccagacaaactgt3' and 5'cggaattcagacgaatgcccagatcacttcc3', was cloned
Supplementary Methods Plasmid constructs. Mouse cdna encoding SHP-1, amplified from mrna of RAW264.7 macrophages with primer 5'cgtgcctgcccagacaaactgt3' and 5'cggaattcagacgaatgcccagatcacttcc3', was cloned
Transcriptional Regulation of Pro-apoptotic Protein Kinase C-delta: Implications for Oxidative Stress-induced Neuronal Cell Death
 SUPPLEMENTAL DATA Transcriptional Regulation of Pro-apoptotic Protein Kinase C-delta: Implications for Oxidative Stress-induced Neuronal Cell Death Huajun Jin 1, Arthi Kanthasamy 1, Vellareddy Anantharam
SUPPLEMENTAL DATA Transcriptional Regulation of Pro-apoptotic Protein Kinase C-delta: Implications for Oxidative Stress-induced Neuronal Cell Death Huajun Jin 1, Arthi Kanthasamy 1, Vellareddy Anantharam
Figure S1. Replication initiation sites at efficient ORIs do not coincide with nucleosomedepleted
 Figure S1. Replication initiation sites at efficient ORIs do not coincide with nucleosomedepleted regions in the mouse genome. Typical examples of SNS profiles (Cayrou et al., 11) and nucleosome occupancies
Figure S1. Replication initiation sites at efficient ORIs do not coincide with nucleosomedepleted regions in the mouse genome. Typical examples of SNS profiles (Cayrou et al., 11) and nucleosome occupancies
Table S1. Primer sequences
 Table S1. Primer sequences Primers for quantitative PCR Tgf 1 Forward Tgf 1 Reverse Tgf 2Forward Tgf 2Reverse Tgf 3 Forward Tgf 3 Reverse Tgf r1 Forward Tgf r1 Reverse Tgf r2 Forward Tgf r2 Reverse Thbs1
Table S1. Primer sequences Primers for quantitative PCR Tgf 1 Forward Tgf 1 Reverse Tgf 2Forward Tgf 2Reverse Tgf 3 Forward Tgf 3 Reverse Tgf r1 Forward Tgf r1 Reverse Tgf r2 Forward Tgf r2 Reverse Thbs1
A Repressor Complex Governs the Integration of
 Developmental Cell 15 Supplemental Data A Repressor Complex Governs the Integration of Flowering Signals in Arabidopsis Dan Li, Chang Liu, Lisha Shen, Yang Wu, Hongyan Chen, Masumi Robertson, Chris A.
Developmental Cell 15 Supplemental Data A Repressor Complex Governs the Integration of Flowering Signals in Arabidopsis Dan Li, Chang Liu, Lisha Shen, Yang Wu, Hongyan Chen, Masumi Robertson, Chris A.
SUPPLEMENTARY INFORMATION
 doi:.38/nature08267 Figure S: Karyotype analysis of ips cell line One hundred individual chromosomal spreads were counted for each of the ips samples. Shown here is an spread at passage 5, predominantly
doi:.38/nature08267 Figure S: Karyotype analysis of ips cell line One hundred individual chromosomal spreads were counted for each of the ips samples. Shown here is an spread at passage 5, predominantly
Supplementary Figure 1. Generation of B2M -/- ESCs. Nature Biotechnology: doi: /nbt.3860
 Supplementary Figure 1 Generation of B2M -/- ESCs. (a) Maps of the B2M alleles in cells with the indicated B2M genotypes. Probes and restriction enzymes used in Southern blots are indicated (H, Hind III;
Supplementary Figure 1 Generation of B2M -/- ESCs. (a) Maps of the B2M alleles in cells with the indicated B2M genotypes. Probes and restriction enzymes used in Southern blots are indicated (H, Hind III;
MeCP2. MeCP2/α-tubulin. GFP mir1-1 mir132
 Conservation Figure S1. Schematic showing 3 UTR (top; thick black line), mir132 MRE (arrow) and nucleotide sequence conservation (vertical black lines; http://genome.ucsc.edu). a GFP mir1-1 mir132 b GFP
Conservation Figure S1. Schematic showing 3 UTR (top; thick black line), mir132 MRE (arrow) and nucleotide sequence conservation (vertical black lines; http://genome.ucsc.edu). a GFP mir1-1 mir132 b GFP
Pyrosequencing. Alix Groom
 Pyrosequencing Alix Groom Pyrosequencing high-throughput CpG methylation analysis platform real-time, sequence-based detection and quantification % methylation at multiple adjacent CpG sites 80-100 bases
Pyrosequencing Alix Groom Pyrosequencing high-throughput CpG methylation analysis platform real-time, sequence-based detection and quantification % methylation at multiple adjacent CpG sites 80-100 bases
Supplementary Fig. 1 Proteomic analysis of ATR-interacting proteins. ATR, ARID1A and
 Supplementary Figure Legend: Supplementary Fig. 1 Proteomic analysis of ATR-interacting proteins. ATR, ARID1A and ATRIP protein peptides identified from our mass spectrum analysis were shown. Supplementary
Supplementary Figure Legend: Supplementary Fig. 1 Proteomic analysis of ATR-interacting proteins. ATR, ARID1A and ATRIP protein peptides identified from our mass spectrum analysis were shown. Supplementary
Supplementary Information
 Supplementary Information Supplementary Figure 1: Identification of new regulators of MuSC by a proteome-based shrna screen. (a) FACS plots of GFP + and GFP - cells from Pax7 ICN -Z/EG (upper panel) and
Supplementary Information Supplementary Figure 1: Identification of new regulators of MuSC by a proteome-based shrna screen. (a) FACS plots of GFP + and GFP - cells from Pax7 ICN -Z/EG (upper panel) and
PrimePCR Assay Validation Report
 Gene Information Gene Name Gene Symbol Organism Gene Summary Gene Aliases RefSeq Accession No. UniGene ID Ensembl Gene ID caspase 9, apoptosis-related cysteine peptidase CASP9 Human This gene encodes a
Gene Information Gene Name Gene Symbol Organism Gene Summary Gene Aliases RefSeq Accession No. UniGene ID Ensembl Gene ID caspase 9, apoptosis-related cysteine peptidase CASP9 Human This gene encodes a
(A-B) P2ry14 expression was assessed by (A) genotyping (upper arrow: WT; lower
 Supplementary Figures S1. (A-B) P2ry14 expression was assessed by (A) genotyping (upper arrow: ; lower arrow: KO) and (B) q-pcr analysis with Lin- cells, The white vertical line in panel A indicates that
Supplementary Figures S1. (A-B) P2ry14 expression was assessed by (A) genotyping (upper arrow: ; lower arrow: KO) and (B) q-pcr analysis with Lin- cells, The white vertical line in panel A indicates that
STAT3 signaling controls satellite cell expansion and skeletal muscle repair
 SUPPLEMENTARY INFORMATION STAT3 signaling controls satellite cell expansion and skeletal muscle repair Matthew Timothy Tierney 1 *, Tufan Aydogdu 2,3 *, David Sala 2, Barbora Malecova 2, Sole Gatto 2,
SUPPLEMENTARY INFORMATION STAT3 signaling controls satellite cell expansion and skeletal muscle repair Matthew Timothy Tierney 1 *, Tufan Aydogdu 2,3 *, David Sala 2, Barbora Malecova 2, Sole Gatto 2,
SUPPLEMENTAL MATERIAL. Carvajal et al. Supplemental Table 1. List of quantitative PCR primers used for cdna analyses and
 UPPLEMEAL MATERIAL Carvajal et al. upplemental Table 1. List of quantitative PCR primers used for cdna analyses and chromatin immunoprecipitation assays. Figure 1. DNA damage-induced transcriptional repression
UPPLEMEAL MATERIAL Carvajal et al. upplemental Table 1. List of quantitative PCR primers used for cdna analyses and chromatin immunoprecipitation assays. Figure 1. DNA damage-induced transcriptional repression
mir-24-mediated down-regulation of H2AX suppresses DNA repair
 Supplemental Online Material mir-24-mediated down-regulation of H2AX suppresses DNA repair in terminally differentiated blood cells Ashish Lal 1,4, Yunfeng Pan 2,4, Francisco Navarro 1,4, Derek M. Dykxhoorn
Supplemental Online Material mir-24-mediated down-regulation of H2AX suppresses DNA repair in terminally differentiated blood cells Ashish Lal 1,4, Yunfeng Pan 2,4, Francisco Navarro 1,4, Derek M. Dykxhoorn
H3K36me3 polyclonal antibody
 H3K36me3 polyclonal antibody Cat. No. C15410192 Type: Polyclonal ChIP-grade/ChIP-seq grade Source: Rabbit Lot #: A1845P Size: 50 µg/32 µl Concentration: 1.6 μg/μl Specificity: Human, mouse, Arabidopsis,
H3K36me3 polyclonal antibody Cat. No. C15410192 Type: Polyclonal ChIP-grade/ChIP-seq grade Source: Rabbit Lot #: A1845P Size: 50 µg/32 µl Concentration: 1.6 μg/μl Specificity: Human, mouse, Arabidopsis,
Nature Genetics: doi: /ng Supplementary Figure 1
 Supplementary Figure 1 Ihh interacts preferentially with its upstream neighboring gene Nhej1. Genes are indicated by gray lines, and Ihh and Nhej1 are highlighted in blue. 4C seq performed in E14.5 limbs
Supplementary Figure 1 Ihh interacts preferentially with its upstream neighboring gene Nhej1. Genes are indicated by gray lines, and Ihh and Nhej1 are highlighted in blue. 4C seq performed in E14.5 limbs
Supplementary information to accompany: A novel role for the DNA repair gene Rad51 in Netrin-1 signalling
 Supplementary information to accompany: A novel role for the DNA repair gene Rad51 in Netrin-1 signalling Glendining KA 1, Markie D 2, Gardner RJM 4, Franz EA 3, Robertson SP 4, Jasoni CL 1 Supplementary
Supplementary information to accompany: A novel role for the DNA repair gene Rad51 in Netrin-1 signalling Glendining KA 1, Markie D 2, Gardner RJM 4, Franz EA 3, Robertson SP 4, Jasoni CL 1 Supplementary
Supplementary Figure Legends. Figure-S1 Molecular basis of ASS1 deficiency in myxofibrosarcoma: (A)
 Supplementary Figure Legends Figure-S1 Molecular basis of ASS1 deficiency in myxofibrosarcoma: (A) High-resolution oligonucleotide-based array comparative genomic hybridization shows normal DNA copy number
Supplementary Figure Legends Figure-S1 Molecular basis of ASS1 deficiency in myxofibrosarcoma: (A) High-resolution oligonucleotide-based array comparative genomic hybridization shows normal DNA copy number
Electrophoretic Mobility Shift Assay (EMSA). Nuclear extracts were. oligonucleotide spanning the NF-kB site (5 -GATCC-
 SUPPLEMENTARY MATERIALS AND METHODS Electrophoretic Mobility Shift Assay (EMSA). Nuclear extracts were prepared as previously described. (1) A [ 32 P] datp-labeled doublestranded oligonucleotide spanning
SUPPLEMENTARY MATERIALS AND METHODS Electrophoretic Mobility Shift Assay (EMSA). Nuclear extracts were prepared as previously described. (1) A [ 32 P] datp-labeled doublestranded oligonucleotide spanning
Supplementary Figure 1 An overview of pirna biogenesis during fetal mouse reprogramming. (a) (b)
 Supplementary Figure 1 An overview of pirna biogenesis during fetal mouse reprogramming. (a) A schematic overview of the production and amplification of a single pirna from a transposon transcript. The
Supplementary Figure 1 An overview of pirna biogenesis during fetal mouse reprogramming. (a) A schematic overview of the production and amplification of a single pirna from a transposon transcript. The
Genome-wide CRISPR screen reveals novel host factors required for Staphylococcus aureus α-hemolysin-mediated toxicity
 Genome-wide CRISPR screen reveals novel host factors required for Staphylococcus aureus α-hemolysin-mediated toxicity Sebastian Virreira Winter, Arturo Zychlinsky and Bart W. Bardoel Department of Cellular
Genome-wide CRISPR screen reveals novel host factors required for Staphylococcus aureus α-hemolysin-mediated toxicity Sebastian Virreira Winter, Arturo Zychlinsky and Bart W. Bardoel Department of Cellular
Nature Biotechnology: doi: /nbt Supplementary Figure 1
 Supplementary Figure 1 Generation of the AARE-Gene system construct. (a) Position and sequence alignment of AAREs extracted from human Trb3, Chop or Atf3 promoters. AARE core sequences are boxed in grey.
Supplementary Figure 1 Generation of the AARE-Gene system construct. (a) Position and sequence alignment of AAREs extracted from human Trb3, Chop or Atf3 promoters. AARE core sequences are boxed in grey.
EpiQuik Tissue Methyl-CpG Binding Domain Protein 2 ChIP Kit
 EpiQuik Tissue Methyl-CpG Binding Domain Protein 2 ChIP Kit Base Catalog # PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE The EpiQuik Tissue Methyl-CpG Binding Domain Protein 2 ChIP Kit is suitable for
EpiQuik Tissue Methyl-CpG Binding Domain Protein 2 ChIP Kit Base Catalog # PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE The EpiQuik Tissue Methyl-CpG Binding Domain Protein 2 ChIP Kit is suitable for
Supplementary Information
 Supplementary Information Supplementary Figures Supplementary Figure 1. MLK1-4 phosphorylate MEK in the presence of RAF inhibitors. (a) H157 cells were transiently transfected with Flag- or HA-tagged MLK1-4
Supplementary Information Supplementary Figures Supplementary Figure 1. MLK1-4 phosphorylate MEK in the presence of RAF inhibitors. (a) H157 cells were transiently transfected with Flag- or HA-tagged MLK1-4
Electromagnetic Fields Mediate Efficient Cell Reprogramming Into a Pluripotent State
 Electromagnetic Fields Mediate Efficient Cell Reprogramming Into a Pluripotent State Soonbong Baek, 1 Xiaoyuan Quan, 1 Soochan Kim, 2 Christopher Lengner, 3 Jung- Keug Park, 4 and Jongpil Kim 1* 1 Lab
Electromagnetic Fields Mediate Efficient Cell Reprogramming Into a Pluripotent State Soonbong Baek, 1 Xiaoyuan Quan, 1 Soochan Kim, 2 Christopher Lengner, 3 Jung- Keug Park, 4 and Jongpil Kim 1* 1 Lab
Supplementary information for: Ten-Eleven Translocation-2 (Tet2) Is Involved in Myogenic Differentiation of Skeletal Myoblast Cells in
 Supplementary information for: Ten-Eleven Translocation-2 (Tet2) Is Involved in Myogenic Differentiation of Skeletal Myoblast Cells in Vitro Xia Zhong*, Qian-Qian Wang*, Jian-Wei Li, Yu-Mei Zhang, Xiao-Rong
Supplementary information for: Ten-Eleven Translocation-2 (Tet2) Is Involved in Myogenic Differentiation of Skeletal Myoblast Cells in Vitro Xia Zhong*, Qian-Qian Wang*, Jian-Wei Li, Yu-Mei Zhang, Xiao-Rong
Supplementary Figure 1. NORAD expression in mouse (A) and dog (B). The black boxes indicate the position of the regions alignable to the 12 repeat
 Supplementary Figure 1. NORAD expression in mouse (A) and dog (B). The black boxes indicate the position of the regions alignable to the 12 repeat units in the human genome. Annotated transposable elements
Supplementary Figure 1. NORAD expression in mouse (A) and dog (B). The black boxes indicate the position of the regions alignable to the 12 repeat units in the human genome. Annotated transposable elements
Table S1. Oligonucleotide primer sequences
 Table S1. Oligonucleotide primer sequences Primer Name DNA Sequence (5 -> 3 ) Description CD19c CD19d Cre7 Sfpi1lox1 Sfpi1lox2 Sfpi1lox3 5 -AACCAGTCAACACCCTTCC-3 5 -CCAGACTAGATACAGACCAG-3 5 -TCAGCTACACCAGAGACGG-3
Table S1. Oligonucleotide primer sequences Primer Name DNA Sequence (5 -> 3 ) Description CD19c CD19d Cre7 Sfpi1lox1 Sfpi1lox2 Sfpi1lox3 5 -AACCAGTCAACACCCTTCC-3 5 -CCAGACTAGATACAGACCAG-3 5 -TCAGCTACACCAGAGACGG-3
Spironolactone ameliorates PIT1-dependent vascular osteoinduction in klotho-hypomorphic mice
 Spironolactone ameliorates PIT1-dependent vascular osteoinduction in klotho-hypomorphic mice Supplementary Material Supplementary Methods Materials Spironolactone, aldosterone and β-glycerophosphate were
Spironolactone ameliorates PIT1-dependent vascular osteoinduction in klotho-hypomorphic mice Supplementary Material Supplementary Methods Materials Spironolactone, aldosterone and β-glycerophosphate were
Reagents and cell culture Mcl-1 gene expression: real-time quantitative RT-PCR In vitro PP2A phosphatase assay Detection of Mcl-1 in vivo
 Reagents and cell culture Antibodies specific for caspase 3, PARP and GAPDH were purchased from Cell Signaling Technology Inc. (Beverly, MA). Caspase inhibitor z-vad-fmk and ROS scavenger N-acetyl-Lcysteine
Reagents and cell culture Antibodies specific for caspase 3, PARP and GAPDH were purchased from Cell Signaling Technology Inc. (Beverly, MA). Caspase inhibitor z-vad-fmk and ROS scavenger N-acetyl-Lcysteine
Supplementary information.
 Supplementary information. Construction of JMJD3 plasmids and other vectors plasmid, pfksd346, encoding the ORF of human JMJD3 (KI346) was obtained from the Kazusa DN Research Institute, Japan. Sequence
Supplementary information. Construction of JMJD3 plasmids and other vectors plasmid, pfksd346, encoding the ORF of human JMJD3 (KI346) was obtained from the Kazusa DN Research Institute, Japan. Sequence
EPIGENTEK. EpiQuik Tissue Chromatin Immunoprecipitation Kit. Base Catalog # P-2003 PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE
 EpiQuik Tissue Chromatin Immunoprecipitation Kit Base Catalog # P-2003 PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE The EpiQuik Tissue Chromatin Immunoprecipitation Kit is suitable for combining the specificity
EpiQuik Tissue Chromatin Immunoprecipitation Kit Base Catalog # P-2003 PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE The EpiQuik Tissue Chromatin Immunoprecipitation Kit is suitable for combining the specificity
Chemical and/or enzymatic deamination across various sequencing methods to localize modified cytosines.
 Supplementary Figure 1 Chemical and/or enzymatic deamination across various sequencing methods to localize modified cytosines. (a) Upon treatment with bisulfite under acidic conditions and at elevated
Supplementary Figure 1 Chemical and/or enzymatic deamination across various sequencing methods to localize modified cytosines. (a) Upon treatment with bisulfite under acidic conditions and at elevated
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb3562 In the format provided by the authors and unedited. Supplementary Figure 1 Glucose deficiency induced FH-ATF2 interaction. In b-m, immunoblotting or immunoprecipitation analyses were
DOI: 10.1038/ncb3562 In the format provided by the authors and unedited. Supplementary Figure 1 Glucose deficiency induced FH-ATF2 interaction. In b-m, immunoblotting or immunoprecipitation analyses were
EPIGENTEK. EpiQuik Chromatin Immunoprecipitation Kit. Base Catalog # P-2002 PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE
 EpiQuik Chromatin Immunoprecipitation Kit Base Catalog # P-2002 PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE The EpiQuik Chromatin Immunoprecipitation Kit is suitable for combining the specificity of
EpiQuik Chromatin Immunoprecipitation Kit Base Catalog # P-2002 PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE The EpiQuik Chromatin Immunoprecipitation Kit is suitable for combining the specificity of
Programmed necrosis - a new mechanism of steroidogenic luteal cell death and elimination during luteolysis in cows
 Supplementary information Programmed necrosis - a new mechanism of steroidogenic luteal cell death and elimination during luteolysis in cows Takuo Hojo, Marta J. Siemieniuch, Karolina Lukasik, Katarzyna
Supplementary information Programmed necrosis - a new mechanism of steroidogenic luteal cell death and elimination during luteolysis in cows Takuo Hojo, Marta J. Siemieniuch, Karolina Lukasik, Katarzyna
administration of tamoxifen. Bars show mean ± s.e.m (n=10-11). P-value was determined by
 Supplementary Figure 1. Chimerism of CD45.2 + GFP + cells at 1 month post transplantation No significant changes were detected in chimerism of CD45.2 + GFP + cells between recipient mice repopulated with
Supplementary Figure 1. Chimerism of CD45.2 + GFP + cells at 1 month post transplantation No significant changes were detected in chimerism of CD45.2 + GFP + cells between recipient mice repopulated with
 Figure S1. Generation of HspA4 -/- mice. (A) Structures of the wild-type (Wt) HspA4 -/- allele, targeting vector and targeted allele are shown together with the relevant restriction sites. The filled rectangles
Figure S1. Generation of HspA4 -/- mice. (A) Structures of the wild-type (Wt) HspA4 -/- allele, targeting vector and targeted allele are shown together with the relevant restriction sites. The filled rectangles
Supplementary Figure Legends
 Supplementary Figure Legends Figure S1 gene targeting strategy for disruption of chicken gene, related to Figure 1 (f)-(i). (a) The locus and the targeting constructs showing HpaI restriction sites. The
Supplementary Figure Legends Figure S1 gene targeting strategy for disruption of chicken gene, related to Figure 1 (f)-(i). (a) The locus and the targeting constructs showing HpaI restriction sites. The
Mapping long-range promoter contacts in human cells with high-resolution capture Hi-C
 CORRECTION NOTICE Nat. Genet. 47, 598 606 (2015) Mapping long-range promoter contacts in human cells with high-resolution capture Hi-C Borbala Mifsud, Filipe Tavares-Cadete, Alice N Young, Robert Sugar,
CORRECTION NOTICE Nat. Genet. 47, 598 606 (2015) Mapping long-range promoter contacts in human cells with high-resolution capture Hi-C Borbala Mifsud, Filipe Tavares-Cadete, Alice N Young, Robert Sugar,
Gene Expression Microarrays. For microarrays, purity of the RNA was further assessed by
 Supplemental Methods Gene Expression Microarrays. For microarrays, purity of the RNA was further assessed by an Agilent 2100 Bioanalyzer. 500 ng of RNA was reverse transcribed into crna and biotin-utp
Supplemental Methods Gene Expression Microarrays. For microarrays, purity of the RNA was further assessed by an Agilent 2100 Bioanalyzer. 500 ng of RNA was reverse transcribed into crna and biotin-utp
The Polymerase Chain Reaction. Chapter 6: Background
 The Polymerase Chain Reaction Chapter 6: Background PCR Amplify= Polymerase Chain Reaction (PCR) Invented in 1984 Applications Invention of PCR Kary Mullis Mile marker 46.58 in April of 1983 Pulled off
The Polymerase Chain Reaction Chapter 6: Background PCR Amplify= Polymerase Chain Reaction (PCR) Invented in 1984 Applications Invention of PCR Kary Mullis Mile marker 46.58 in April of 1983 Pulled off
Hyper sensitive protein detection by Tandem-HTRF reveals Cyclin D1
 Hyper sensitive protein detection by Tandem-HTRF reveals Cyclin D1 dynamics in adult mouse Alexandre Zampieri, Julien Champagne, Baptiste Auzemery, Ivanna Fuentes, Benjamin Maurel and Frédéric Bienvenu
Hyper sensitive protein detection by Tandem-HTRF reveals Cyclin D1 dynamics in adult mouse Alexandre Zampieri, Julien Champagne, Baptiste Auzemery, Ivanna Fuentes, Benjamin Maurel and Frédéric Bienvenu
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature09861 & &' -(' ()*+ ')(+,,(','-*+,&,,+ ',+' ' 23,45/0*6787*9:./09 ;78?4?@*+A786?B- &' )*+*(,-* -(' ()*+ ')(+,,(','-*+,&,,+ ',+'./)*+*(,-*..)*+*(,-*./)*+*(,-*.0)*+*(,-*..)*+*(,-*
doi:10.1038/nature09861 & &' -(' ()*+ ')(+,,(','-*+,&,,+ ',+' ' 23,45/0*6787*9:./09 ;78?4?@*+A786?B- &' )*+*(,-* -(' ()*+ ')(+,,(','-*+,&,,+ ',+'./)*+*(,-*..)*+*(,-*./)*+*(,-*.0)*+*(,-*..)*+*(,-*
QPCR ASSAYS FOR MIRNA EXPRESSION PROFILING
 TECH NOTE 4320 Forest Park Ave Suite 303 Saint Louis, MO 63108 +1 (314) 833-9764 mirna qpcr ASSAYS - powered by NAWGEN Our mirna qpcr Assays were developed by mirna experts at Nawgen to improve upon previously
TECH NOTE 4320 Forest Park Ave Suite 303 Saint Louis, MO 63108 +1 (314) 833-9764 mirna qpcr ASSAYS - powered by NAWGEN Our mirna qpcr Assays were developed by mirna experts at Nawgen to improve upon previously
Supplementary Information
 Supplementary Information Supplementary Figure 1. ZBTB20 expression in the developing DRG. ZBTB20 expression in the developing DRG was detected by immunohistochemistry using anti-zbtb20 antibody 9A10 on
Supplementary Information Supplementary Figure 1. ZBTB20 expression in the developing DRG. ZBTB20 expression in the developing DRG was detected by immunohistochemistry using anti-zbtb20 antibody 9A10 on
Supporting Information
 Supporting Information Lujambio et al. 10.1073/pnas.0803055105 SI Materials and Methods RNA Isolation and mirna Expression Analysis. Total RNA was isolated from SW620,, and SIHN-0 cells, before and after
Supporting Information Lujambio et al. 10.1073/pnas.0803055105 SI Materials and Methods RNA Isolation and mirna Expression Analysis. Total RNA was isolated from SW620,, and SIHN-0 cells, before and after
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/4/198/ra74/dc1 Supplementary Materials for Short RNA Duplexes Elicit RIG-I Mediated Apoptosis in a Cell Type and Length-Dependent Manner Osamu Ishibashi, Md. Moksed
www.sciencesignaling.org/cgi/content/full/4/198/ra74/dc1 Supplementary Materials for Short RNA Duplexes Elicit RIG-I Mediated Apoptosis in a Cell Type and Length-Dependent Manner Osamu Ishibashi, Md. Moksed
Flowcytometry-based purity analysis of peritoneal macrophage culture.
 Liao et al., KLF4 regulates macrophage polarization Revision of Manuscript 45444-RG- Supplementary Figure Legends Figure S Flowcytometry-based purity analysis of peritoneal macrophage culture. Thioglycollate
Liao et al., KLF4 regulates macrophage polarization Revision of Manuscript 45444-RG- Supplementary Figure Legends Figure S Flowcytometry-based purity analysis of peritoneal macrophage culture. Thioglycollate
transcription and the promoter occupancy of Smad proteins. (A) HepG2 cells were co-transfected with the wwp-luc reporter, and FLAG-tagged FHL1,
 Supplementary Data Supplementary Figure Legends Supplementary Figure 1 FHL-mediated TGFβ-responsive reporter transcription and the promoter occupancy of Smad proteins. (A) HepG2 cells were co-transfected
Supplementary Data Supplementary Figure Legends Supplementary Figure 1 FHL-mediated TGFβ-responsive reporter transcription and the promoter occupancy of Smad proteins. (A) HepG2 cells were co-transfected
Supplementary Figure 1. Characterization of hipscs derived from primary human fibroblasts. a,b. Morphology of hipscs. hipscs exhibit hesc-like
 Supplementary Figure 1. Characterization of hipscs derived from primary human fibroblasts. a,b. Morphology of hipscs. hipscs exhibit hesc-like morphology in co-culture with mouse embryonic feeder fibroblasts
Supplementary Figure 1. Characterization of hipscs derived from primary human fibroblasts. a,b. Morphology of hipscs. hipscs exhibit hesc-like morphology in co-culture with mouse embryonic feeder fibroblasts
Supplementary Figure 1. GST pull-down analysis of the interaction of GST-cIAP1 (A, B), GSTcIAP1
 Legends Supplementary Figure 1. GST pull-down analysis of the interaction of GST- (A, B), GST mutants (B) or GST- (C) with indicated proteins. A, B, Cell lysate from untransfected HeLa cells were loaded
Legends Supplementary Figure 1. GST pull-down analysis of the interaction of GST- (A, B), GST mutants (B) or GST- (C) with indicated proteins. A, B, Cell lysate from untransfected HeLa cells were loaded
Supplementary Figure 1. Phenotype and genotype of cultured and transplanted S1 KCST (A) Brightfield and mcherry fluorescence images of the spheres
 Supplementary Figure 1. Phenotype and genotype of cultured and transplanted S1 KCST (A) Brightfield and mcherry fluorescence images of the spheres generated from the CD133-positive cells infected with
Supplementary Figure 1. Phenotype and genotype of cultured and transplanted S1 KCST (A) Brightfield and mcherry fluorescence images of the spheres generated from the CD133-positive cells infected with
